Abstract
Alzheimer’s disease (AD) is a neurological disease that affects numerous people. The condition causes brain atrophy, which leads to memory loss, cognitive impairment, and death. In its early stages, Alzheimer’s disease is tricky to predict. Therefore, treatment provided at an early stage of AD is more effective and causes less damage than treatment at a later stage. Although AD is a common brain condition, it is difficult to recognize, and its classification requires a discriminative feature representation to separate similar brain patterns. Multimodal neuroimage information that combines multiple medical images can classify and diagnose AD more accurately and comprehensively. Magnetic resonance imaging (MRI) has been used for decades to assist physicians in diagnosing Alzheimer’s disease. Deep models have detected AD with high accuracy in computing-assisted imaging and diagnosis by minimizing the need for hand-crafted feature extraction from MRI images. This study proposes a multimodal image fusion method to fuse MRI neuroimages with a modular set of image preprocessing procedures to automatically fuse and convert Alzheimer’s disease neuroimaging initiative (ADNI) into the BIDS standard for classifying different MRI data of Alzheimer’s subjects from normal controls. Furthermore, a 3D convolutional neural network is used to learn generic features by capturing AlD biomarkers in the fused images, resulting in richer multimodal feature information. Finally, a conventional CNN with three classifiers, including Softmax, SVM, and RF, forecasts and classifies the extracted Alzheimer’s brain multimodal traits from a normal healthy brain. The findings reveal that the proposed method can efficiently predict AD progression by combining high-dimensional MRI characteristics from different public sources with an accuracy range from 88.7% to 99% and outperforming baseline models when applied to MRI-derived voxel features.
1. Introduction
The human brain is often considered one of the most crucial and intricate organs in the body, necessary for developing ideas, resolving issues, formulating judgments, exercising imagination, and storing memories [1,2]. Memory is capable of accumulating knowledge and experiences and retrieving them. Since it contains the complete record of a person’s life, physical memory is a vital component in forming personality and identity. Losing one’s memory due to dementia and the inability to identify one’s surroundings are terrible situations [1,2]. Alzheimer’s disease (AD) is the kind of dementia that affects most people. People’s concerns about AD tend to intensify as they age [3]. The gradual death of brain cells characterizes AD.
Consequently, patients eventually become disconnected from everything around them and experience a loss of loving memories, memories from childhood, the capacity to recognize family members, and even the ability to understand and carry out straightforward instructions. In the latter stages, they will also lose the power to swallow, cough, and breathe normally. There are approximately 50 million people throughout the world living with dementia, and the cost of providing health and social care for these individuals equals the size of the 18th biggest economy [3,4]. Through tracking its progression, early and precise identification of AD plays a crucial role in preventing and treating the disease and providing patient care. Imaging techniques of the brain, such as magnetic resonance imaging (MRI), are the subject of several different research initiatives. Such an approach can determine the number of cells in the brain and their size. Additionally, it can demonstrate the shrinkage of the parietal lobe associated with AD [5].
MRI may detect the brain abnormalities linked with mild cognitive impairment (MCI), and this information can predict which MCI patients will progress to AD in the future [3]. The utilization of MRI images largely depends on the availability of qualified doctors or healthcare professionals who will search for any anomalies in the MRI pictures. Researchers have investigated MRI images of patients to find that some unknown features, such as a size reduction in various brain regions (mainly affecting the temporal and parietal lobes), help detect AD [3,5]. Nevertheless, analyzing every MRI will delay detection, which, in turn, slows treatment. Furthermore, an insufficient number of radiologists in rural areas makes timely detection and intervention difficult. Hence, an automatic AD detection system, or, more accurately, a computer-aided AD detection system, is essential. It can be shown that paying closer attention to the differences between healthy, MCI, and AD people by combining their data from publicly available sources can considerably aid in the early detection and monitoring of Alzheimer’s disease [6,7]. It is well known that MRI brain images are assembled from many sources, such as hospitals, medical institutions, and others. Therefore, various imaging methods and different brain regions and textures can reveal certain principles due to their different modalities [6,7]. By learning these characteristics, patients with AD can be classified and identified with greater accuracy, allowing for faster detection and treatment of diseases.
A significant amount of information about AD can be extracted from neuroimaging data through machine learning (ML) and deep learning (DL). However, more data is produced by brain-imaging techniques [7,8,9]. This situation occurs because brain-imaging techniques generate an increasing amount of data. Traditional learning-based methods include the following three stages: stage one involves regionalized MRI of the brain based on predetermined regions of interest (ROIs), stage two selects features from the ROIs, and stage three consists of the construction and evaluation of classification models [10,11].
The process of hand-crafted ROIs identification and selection, also known as manual selection and extraction, constitutes the primary weakness of ML approaches [12,13]. This flaw significantly impacts the quality of the results produced by the model. DL has developed into a revolutionary system in recent decades compared to the classic ML methods [11]. Images can be directly analyzed using deep learning, so it no longer requires human experts for feature extraction. However, most CNN models for AD prediction are trained on individual modal image data (i.e., cohort), which cannot be applied to other cohorts.
A great deal of research has focused on individual modal image data. By integrating a mix of MRI models from multiple cohorts, this research proposes a MULTforAD, a new multimodal feature fusion model based on a 3D-CNN, to predict AD progression without using any pre-trained networks or transfer learning. The high-dimensional MRI neuroimaging fusion is achieved by combining and assembling image processing techniques. A fully connected neural network then classifies the output images. The suggested model undergoes training and testing on various public datasets with the following contributions:
- A robust three-dimensional CNN is proposed with three distinct classifiers (Softmax, SVM, and RF) for detecting multimodal-fused features for the prediction of AD;
- A new method of image fusion, MULTforAD, is presented and evaluated for brain MRI information preprocessing and fusing, which has improved network classification and performance in AD diagnosis;
- Multiple features and details for patients who are MCI or AD were collected and analyzed using the MULTforAD method, making the results easier to interpret.
2. Literature Review
We can now investigate the facets that play a significant part in our work. This is made possible by imaging methods applied to the brain. Scientists can now explore not just individual brain areas but also the dynamic pattern of connections between them due to recent developments in the field [12]. Several studies have taken place to develop deep learning (DL) models that can classify Alzheimer’s patients based on the segmentation of medical images [13,14,15,16]. An X-shaped network structure (X-Net) has been proposed by Li et al. [17]. It represents a viable alternative compared to pure convolutional networks for medical image segmentation. Additionally, local and global features can be extracted to obtain better results. Ren et al. [18] proposed a faster RCNN, which suggests the region proposal network (RPN), a separate network that predicts the network’s regions. RPN is a supervised method that evaluates bounding boxes using a loss function based on intersection over union (IoU) and requires a ground truth pixel-level label. A scheme for image fusion based on image cartoon texture decomposition and sparse representation was proposed by Zhu et al. [19].
The fused cartoon and texture components are combined from MRI scans for medical research using texture enhancement fusion rules. For classifying and predicting whole-brain PET images, Silveira et al. [20] used the boosting approach. Despite its simplicity, this method proves 90.97% accurate on AD (Alzheimer’s disease) and NC (normal cohort).
Additionally, Liu et al. [21] used sagittal, coronal, and cross-sectional slices of the whole FDG-PET 3D image to extract features utilizing convolutional neural networks. A pre-trained 2D-CNN deep learning model, ResNet50, automatically extracted features from ADNI MRI images for AD diagnosis. The CNN then underwent evaluation using conventional Softmax, SVM, and RF metrics, such as accuracy. Pre-trained models achieve higher accuracy, ranging between 85.7% and 99%. Finally, using a recurrent neural network, the three directions of features were combined to produce the final classification prediction. These methods use only single-modality medical images to diagnose and classify AD. Additionally, a significant calculation procedure helps optimize the pre-trained network.
One of the major drawbacks of those methodologies is that they study only one (or a few) brain regions, whereas AD alterations affect multiple brain regions. However, this method may reduce the risk of model overfitting because of the smaller and fewer inputs than those methods that allow patch combinations. Multimodal images offer numerous suitable features for AD pathology, and combining images of different modalities can help diagnose AD earlier and more accurately [22,23]. By slicing the 3D brain images, the 2D convolution operation extracts features, thereby weakening the spatial correlation and potentially enhancing the loss of spatial features. An MRI 3D segmentation using a 3D fully convolutional neural network was performed by Biswas et al. [24]. According to the findings, simple 3D convolutional operations on 3D brain images (MRI images in the experiments) demonstrate great potential. The lightweight 3D convolutional model has excellent performance compared to complex models that require tedious preprocessing of data. In addition to considering ambiguity zones, the authors believe that using multiple binary classifications instead of multiple classification tasks might lead to “ambiguity zones.” Even though the above methods prove capable of extracting features from 3D images, they remain limited in specific ways. Multimodal images can demonstrate more comprehensive and significant experimental results. Different imaging principles are employed in medical brain imaging, making it possible for modal images to highlight certain pathological characteristics associated with specific brain diseases. Brain MRI images represent, for instance, receptor distribution, cortex thickness, or the functional activity of the brain [25,26]. This way, multiple modal images for the same subject can provide more comprehensive pathological information and features.
Furthermore, Liu et al. [21] developed a network model designed explicitly for AD classification based on MRI and PET images, and their results proved satisfactory. The operation of this method on 3D images requires many complex preprocessing steps, including rigid alignment, non-rigid alignment, and ROI extraction. In [21], the authors proposed a multimodal image fusion approach that combines complementary information from different PET + MRI images. Therefore, the composite modality gives more accurate information than a single input picture. The proposed image fusion approach recovers the GM area from FDG-PET based on an anatomical mask from an MRI scan. This GM is then used to combine structural MRI and functional PET data using a trained sparse autoencoder to develop 3D-CNN convolution layers. According to experimental results, the proposed model is 93.21% accurate in binary classification and 87.67% accurate in multi-class AD classifications.
Table 1 summarizes some of the recent work in the area of brain image analysis. Compared to existing feature fusion strategies, multimodal medical image fusion is more intuitive. Multiple images are fused to enhance the accuracy of diagnosis and treatment. In addition to strengthening modal features, the fusion of images also improves information representations. Inspired by this fact, the proposed research intends to implement and evaluate a multimodal MRI feature fusion method to automatically classify Alzheimer’s illnesses. This evaluation will be based on the performance of different image processing methods and a 3D convolutional neural network approach on MRI images collected from various sources. Using three distinct classifiers (Softmax, SVM, and RF), the 3D-CNN-based model detects Alzheimer’s disease on MRI images via deep learning. In this study, full-connected and convolutional layers were compared for their performance, and the result was compared with state-of-the-art benchmarked models for AD detection and classification. This paper is organized as follows: first, the details of the data, the preprocessing steps, and the tools used for the implementation are given in the “AD data description” section then the “Proposed Model” describes the proposed model architecture and testing/validation settings. After providing a quick overview of the experiment’s setup, covering the software and hardware configurations, the experimental section and the results undergo analysis in the “Implementation Results” section. Finally, the conclusion and discussion are explained in the “Conclusions” section.

Table 1.
Related work summary.
3. Proposed MULTforAD Image Fusion
Neuroimaging encounters a significant problem regarding the availability of many scans associated with AD patients since limited image samples are available. Therefore, a DL model is often more effective when trained on more data. Unfortunately, medical research is hindered by privacy concerns, making it tricky to access large datasets [12], especially when classifying cancer and Alzheimer’s disease. Furthermore, during the model’s training, the small and imbalanced dataset creates overfitting problems that affect the model’s efficiency [28,29]. To overcome this issue, we proposed a multimodal fusion MULTforAD method to fuse MRI images from three online sources. These dataset sources contain complimentary MRI information that is most prominent in Alzheimer’s disease and helps accurately forecast the development from MCI to AD.
In addition, different modalities of these images can highlight the anatomical structure and texture abnormalities more precisely in pathological regions. Figure 1 presents a multimodal coronal view from participants with varying cognitive states.
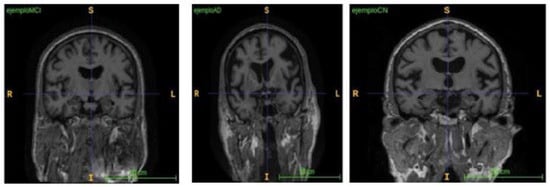
Figure 1.
Sample multimodal coronal MRI collected from ADNI dataset for the same patient.
By incorporating all such features from different sources, the composite modality more accurately reflects the information than a single input image. Utilizing the combined modality, the subject is diagnosed using a single-channel network.
The number of design variables dramatically reduces compared to multi-channel input networks with feature fusion. Figure 2 shows the suggested MULTforAD. The critical components comprise unprocessed multimodal MRI data collection, image fusion, and preprocessing, followed by a neural network-based classification. The following subsection provides the details of the proposed model to highlight each step.
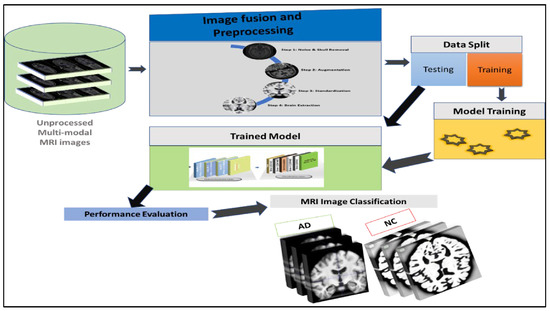
Figure 2.
Proposed MULTforAD multimodal image fusion method for AD diagnosis framework.
3.1. Multimodal MRI Data Collection
As MRI measures the energy released by protons within various tissues, such as white matter (WM), gray matter (GM), and cerebrospinal fluid (CF), it can provide detailed images of the brain, allowing significant damage and complex changes to be detected [12]. Therefore, public datasets such as ADNI, Kaggle, and OASIS provide tremendous value to the AD research community. The current study uses three datasets to consider multimodal features from MRI neuroimaging for different cohorts. Table 2 describes the collected samples as follows:

Table 2.
Fused MRI dataset description.
An authenticated user ID and password are necessary to see the MRI images on the ADNI website. An authenticated username was used for logging into the ADNI website (credentials available on direct request with the author). All images were in the nii format. The dataset comprises 5982 MRI images. For this study, only AD (Alzheimer’s Disease) (1896 images) and NC (Normal Cohort) (4086 images) class images are incorporated for the analysis of the effectiveness of the proposed model.
Additionally, the Kaggle dataset contains a mild-to-moderate dementia dataset that comprises 72 subsets of data related to three different classes, including NC, MCI (mild cognitive impairment), and AD data. These are assigned by a physician after a series of clinical tests, while there are only two diagnosis classes for OASIS (the MCI subjects are labeled as AD), which are picked up solely from the clinical data report. Therefore, there is no clear organization of the raw data downloaded, making them difficult to use. Hence, the raw data was converted to the BIDS format [30] using various preprocessing techniques. In addition, all outputs of the experiments are standardized according to the BIDS format.
3.2. Image Fusing and Preprocessing
Different sources of images show different shapes, brightness, and contrast since the images were gathered from various sources. To boost the contrast of all images, we applied different image processing techniques to resize, standardize, and augment [31] all MRI images (Figure 3). However, the nonlinear light intensity can cause the addition of undesirable information to the image during the acquisition. This may affect the accuracy of the overall image processing [32]. Therefore, deterioration of MRI images may occur during the formation process such as low variation resulting from poor brightness produced by the visual devices. So, image enhancement approaches were applied to MRI scans to upgrade the pixel distribution over a wide range of intensities.
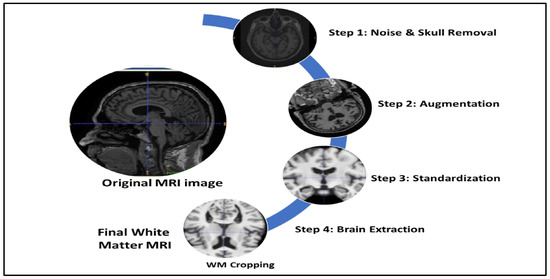
Figure 3.
The original MRI processing method. Pictures from top to bottom are MRI after removing the skull, flip, and rotation; image registration; the White Matter (WM) extraction MRI extraction.
For the set of fused MRI neuroimages, applying a noise mask distorts high- and low-frequency components and increases sample variability to overcome this limitation. Essentially, the original image is enhanced by adding a Gaussian distribution array. Figure 4 compares an MRI image with Gaussian noise to the original image. Removing unnecessary objects from the neuroimage MRI scan is crucial to improving accuracy, particularly during the classification phase. For example, a skull may negatively affect pattern recognition and significantly increase input complexity [3,7]. AD only affects brain tissue, so all objects and tissues other than the brain are undesirable. We used Pincram’s [31] skull-stripping software to process structural MRI images, as shown in Figure 5. Watershed brain mask segmentation can reduce brain size by removing bones and other non-brain material, resulting in less distortion and redundant data. This method preserves only intracranial tissue structure while removing unnecessary anatomical organs, as predicted in [32]. After skull removal, the FLIRT package accurately transforms MRI images into a global brain function map model, known as the MNI152 space. The FLIRT technique recognizes brain objects in both in-mode and inter-mode completely automated, reliably, and accurately.
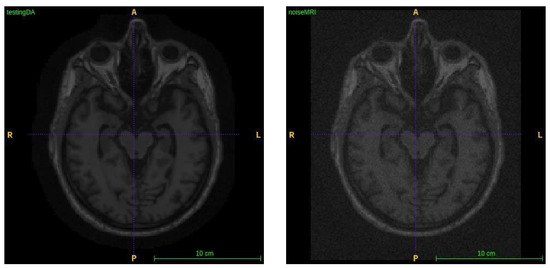
Figure 4.
Original image (left) and noise image with std = 0.2 (right).
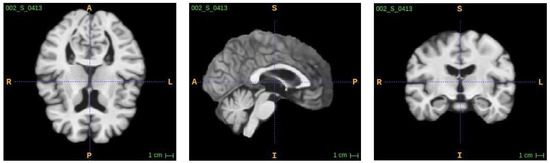
Figure 5.
Axial, sagittal, and coronal views of an MRI without a skull.
The amount of data available often improves neural network performance. Training data’s availability, quality, and labelling constitute significant constraints for training effective models. Making minor changes to our existing dataset is a popular way to acquire more data. Convolutional neural networks are not affected by translation, viewpoint, size, or lighting. As part of image data augmentation, images from the training dataset are transformed into transformed versions that belong to the same class as the original images. In neural network models, this method can prevent overfitting by explicitly adding familiar sources of variation to training samples. A model that has been overfitted performs poorly on new data as a result of memorizing its training data [33]. When selecting transformations, consideration must be given to the training dataset and the knowledge of the domain of the problem. To increase the variability of the collected datasets, all MRI scans that have been fused are flipped. It is possible to turn the images horizontally and vertically. By reversing the rows or columns of pixels in an image, a vertical flip is equivalent to rotating the image 180 degrees. Figure 6 depicts horizontal, vertical, and both flips.
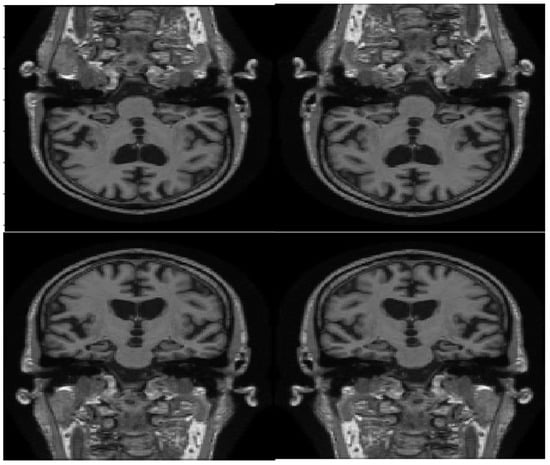
Figure 6.
Sample image flips. Above, original, and horizontal flip.
Afterwards, a cropping process is randomly applied to the original image and resized to the original size. As a result, CNNs are translation and size invariant, which can help improve the model’s robustness.
Finally, a random rotation process of the image is used as another possible transformation. The rotation of an image can introduce artifacts in areas where new information must be presented after the rotation, depending on the image. The collected data is processed using several filling techniques, including adding zeroes, reflecting, and wrapping. The background in an MRI is always black (zero), meaning this issue is avoided. Figure 7 depicts an example of rotation applied over the collected datasets.
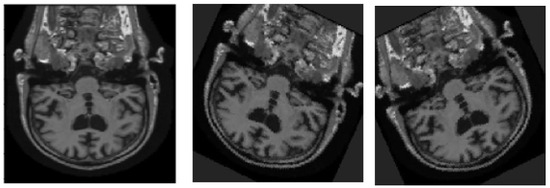
Figure 7.
The image rotated 25 and −25 degrees.
High-fidelity 3D information, as input to the deep learning network, requires more computational resources during the training phase. Moreover, standardizing inputs will speed up training when modeling a neural network.
To address this limitation, all the fused images are manipulated using clipping and sampling techniques to reduce the time required to calculate individual information. Figure 8 shows that each modality image contains many background areas with zero-pixel values. Properly minimizing these useless background spots reduces the amount of input information without damaging brain tissue areas. MRI is trimmed from 182 × 218 × 182 to 145 × 172 × 145, and a Z-score normalization for image dimension scaling is applied. The process of rescaling features to have a zero mean and unit variance is known as standardization (or Z-score normalization). The standardization formula is shown in Equation (1).
where µ is the mean, and σ is the standard deviation.
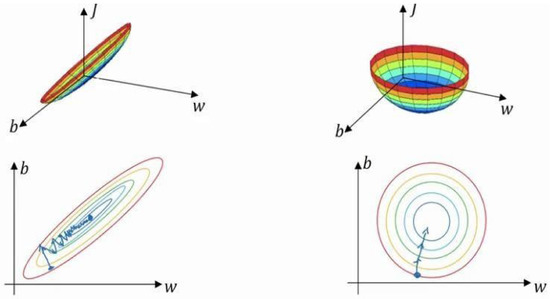
Figure 8.
The cost functions for non-optimized (left) and optimized features (right).
The standardization of the entire dataset and applying the same transformation to the training and test sets is, therefore, essential. Figure 8 illustrates the significance of data standardization in training algorithms. Using the collected datasets, the gradient-based algorithm is used to find the minimum value, identify the correct learning rate, and avoid time complexity. The final step is to apply a segmentation method for creating a brain tissue mask from the multimodality collected input. The extraction of the brain mask is essential for the subsequent phases of analysis and classification.
Several methods proposed to deal with the brain segmentation problem reflect the importance of reliable and accurate brain extraction [32,33]. The brain extraction (registration procedure) aims to exclude the individual’s spatial variation tissues, including the skull, skin, eyes, and fat, and not remove any normal orientation part of the brain. Most brain extraction methods use T1-weighted MR images since they provide excellent contrast among the different brain tissues. In MULTforAD, the white matter tissue (WM) undergoes segmentation from the input MRI image using the FSL (FMRIB Software Library V6.0) software FAST module. FAST separates 3D brain images into different tissue types, taking into account changes in geographical brightness [31] for the AD classification task. Multiplying the mask by the brain MR image removes the skull from the image. Figure 9 illustrates the brain extraction process. Figure 10 shows a sample of the brain extraction process conducted over the fused images. Having collected the data and processed the images, the dataset, which contains 5982 instances, is divided into train, validation, and test sets in a ratio of 75:15:10, respectively.
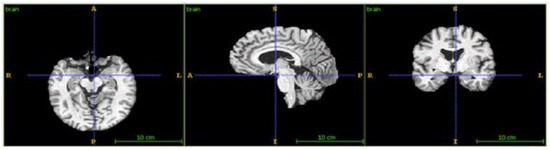
Figure 9.
The brain extraction process.
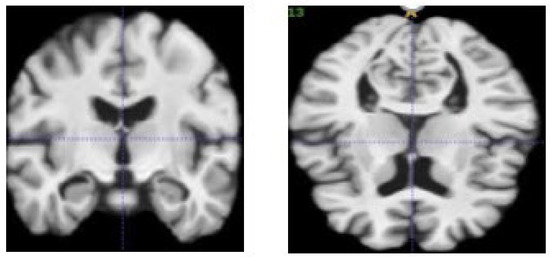
Figure 10.
MRI after the brain extraction process.
3.3. 3D-Convolutional Neural Network
Convolutional neural networks may be trained for various tasks, including image segmentation, classification, and reconstruction, using an image as an input. CNN architecture is based on what can be called the human brain and visual cortex neuronal connection pattern. CNN can do this using multiple convolutional kernels to detect spatial correlations in a picture. The architecture of the suggested 3D-CNN (shown in Figure 11) consists of the input, fully connected (FC) layer, pooling layers, and classification layer. Initially, the experiment used an overfitting model that was heavy due to its high number of FC layers. Next, the following operations were repeated iteratively. Firstly, the number of FC layers decreased until the accuracy of the validation set declined significantly. Secondly, one more convolutional block was added. For example, start with a heavy model of four convolutional blocks + five FC layers, then four convolutional blocks + two FC layers, and repeat with one FC layer. Afterwards, change the convolution blocks from four to seven and notice the accuracy.
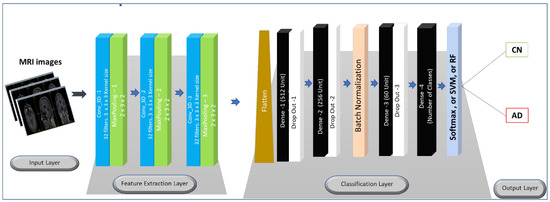
Figure 11.
The proposed 3D-CNN architecture.
Four convolutional blocks and three FC layers were chosen as the optimal architecture for multimodal MRI classification tasks. The training hyperparameters (learning rate, decay of weight) are adjusted based on the evolving training accuracy. The convolutional layer extracts relevant features from the MRI neuroimages to obtain relevant knowledge about AD and NC patients. Training images preprocessed to 145 × 172 × 145 size are provided as inputs to the CNN, and the last feature extraction layer generates feature maps. Combining all the extracted feature maps with the ReLU activation function in the convolution layer, a CNNForAD with 32 neurons is proposed, further connected to a classifier layer. A 3D convolutional layer applies to slide cuboidal convolution filters to transform 3D inputs into convolutional filters.
Convoluting the input involves moving filters vertically and horizontally and, along with the depth of the input, calculating the dot product of weights and input and then adding a bias term. In the case of a 3D MRI input, the layer’s dimensions are determined by the input layer, which contains data with five dimensions corresponding to pixels in three spatial dimensions, channels, and observations. During the feature extraction, the 3D-CNN model will create MRI feature vectors from the FC layer after being trained with the training set. The feature vectors are then fed into final classifiers. While fine-tuning the model, the validation set offers an unbiased assessment of the model’s capacity to match the training dataset. The feature extraction process in CNN employs local connections to identify local features and pooling to combine characteristics in comparable localities into a single feature. In the meantime, the FC layer is utilized in computing the result for each MRI picture input.
A binary-class classification problem is utilized as the neuroimages undergo classification in the following two categories: AD and CN. The feature extractor convolutional layer contains 3 × 3 filters (shown in Table 3). A max-pooling layer follows the convolution layer to minimize the feature maps of the collected images by partitioning the images into sets of 2 × 2 zones that do not overlap. Subsequently, a batch normalization layer resists any incorrect weight initialization of the proposed model to accelerate the training process. Finally, a dropout layer prevents model overfitting. The dropout rate of 0.5 determines the probability of neuron loss by controlling the number of neurons eliminated from the network. The neurons are only eliminated while the training procedure is carried out. The outcome of the classification layer (the fully connected layer), also known as dense layers, is added to the model with 32 neurons, followed by the final output layer. The FC layer had several adjustments, so it could be calibrated to link the several layers in the network while also providing the Alzheimer’s disease categorization task by utilizing a normalized exponential function, Softmax, SVM, or RF.

Table 3.
Specifications of the designed model’s parameters.
3.3.1. Softmax
The Softmax function is utilized in the final layer of CNN architecture to categorize the labelled data and then transform the output values into perceptible ones, using ground-truth labels between 0 and 1.
3.3.2. SVM
The last FC layers will be replaced with an SVM classifier with several splits (the number of the folds will be set to 10, and the seed will be set to 7), demonstrating notable success in solving real-world issues. Additionally, RBF kernels are used by SVM classifiers to develop nonlinear classifiers by mapping the original dataset to a higher-dimensional space through linear regression [34].
3.3.3. Random Forest
RF can reduce the amount of variation in an estimated prediction function [34]. It is used in both classification and regression analysis. When applied to the classification task, each tree in the forest places a vote as a class. Then the input is classified based on the vote that receives the majority of the total.
When utilized for regression, the trees’ predictions are averaged when positioned at the target point x. In the course of our research, we used RF for categorization. The number estimator figure is set to 20, even though the default value is 100. This value may range from 1 to 100, and we found that 20 produced the most accurate results.
4. Experimental Result
The experiment and its setup are discussed in this section, and then the outcomes are presented. After providing an overview of the experiment’s setup, we will discuss the results of the model training and validation processes. Next, the findings achieved while using the CNN model for feature extraction with the three different classifiers undergo discussion in the third subsection (Softmax, SVM, and RF).
Finally, we will examine how the results were achieved using the suggested strategy against state-of-the-art approaches.
4.1. Experimental Setup
The experiments were conducted using the MATLAB 2021b environment with a ‘MiniBatch’ size of 12 and several ‘MaxEpoches.’ This study employed MRI neuroimages and visualization of the brain’s anatomy from a coronal plane. Test datasets should only evaluate the performance of fully specified and trained classifiers [32,33]. In light of this, the training data with 5982 instances were split into training/validation/test files. Training/validation sets were used for selected models in a CV classification task. It was decided to leave test sets untouched until the peer review was completed. A random sample of 100 subjects for each diagnosis group was selected from the ADNI test dataset, matched by age and sex (e.g., 100 NC patients and 100 AD patients). We used the remaining ADNI data as a training and validation set to determine whether the model has overfitted the training/validation set. A separate test set was created for all OASIS and Kaggle classes to ensure age and gender distribution consistency. Stochastic gradient descent with momentum (SGDM) optimization was used during the training process with an initial learning rate of 0.001.
The model’s generalization ability was tested using the Kaggle test set, and the OASIS test determined whether a dataset with various inclusion criteria and different imaging conditions could be generalized. It is crucial to note that the neuroimage labels in OASIS and ADNI/Kaggle are not based on the same criteria. Consequently, the ADNI-trained models cannot be generalized sufficiently well to OASIS. The CNNForAD model was selected based on the training/validation dataset, which included the selection of the model architecture and the fine-tuning of its training hyperparameters. Cross-validation experiments were conducted with a 10-fold increase in the learning rate, with 40% of the data used for validation and the rest for training. This data split occurred only once for the experiments with a fixed seed number (random state = 2), ensuring that all experiments used the same subjects during the classification process. Additionally, there will be no overlap.
4.2. Performance Evaluation Metrics
The accuracy (ACC) performance metric is considered essential for model assessment. Additionally, sensitivity (SPE) and specificity (SEN) are performance metrics. The positive tuples assigned to the appropriate labels by the classifier are referred to as true positives (TP). Let us call this number TP, which stands for “true positives”. False positives, abbreviated as FP, refer to the negative tuples mistakenly categorized as positive. Let’s call this figure FP for the number of false positives. The actual negatives, also known as TNs, are the negative tuples correctly categorized by the classifier. Let us call this number TN, which stands for “true negatives”. The positive tuples mistakenly classified as negatives are false negatives (FN). Let us call this occurrence the number of false negatives or FN.
Accuracy (ACC): The percentage of the number of records classified correctly versus the total records shown in the equation below:
ACC = (TP + TN)/(TP + TN + FP + FN)
Sensitivity (SEN)/recall shows the percentage of the number of records identified correctly over the total number of AD subjects, as shown in the equation below:
SEN = TP/(TP + FN)
Specificity (SPE): The percentage of the number of records. Normal control is divided by the total number of normal nodes, as shown in the equation below:
SPE = TP/(TP + FP)
F1: a measure of a test’s accuracy:
4.3. Experiments and Results
4.3.1. Image Fusion Performance
In Table 4, unimodal (a single cohort dataset with the same setting and preprocessing steps) and MultforAD multimodal imaging are presented with various network layers in categorizing AD: NC. The MultForAD technique performs better since the MRI data can be fused effectively.

Table 4.
Results of the proposed image fusion method against a single modality based on 3D-CNN in three different epoch sizes.
The MultforAD fusion approach outperforms the unimodal method in terms of overall indicators. Using 3D CNN, the accuracy of the classification amounted to (93.21 ± 5.0)%, specificity amounted to (91.43 ± 4.9)%, and the sensitivity amounted to (95.42 ± 2.5)%. There was a high sensitivity (94.44 ± 7.9)% for feature fusion but a low level of accuracy and specificity. Our image fusion approach provided the best results using the softmax activation function for the AD–NC classification test.
4.3.2. 3D-CNN Network Performance
In this section, the proposed 3D-CNN for multimodal MRI image classification undergoes evaluation using a ten-fold cross-validation matrix, as presented in Table 5, Table 6 and Table 7. Fold 1, Fold 6, Fold 7, Fold 9, and Fold 10 achieved 100% performance metrics. Using a ten-fold cross-validation strategy, it was found that the average accuracy, sensitivity, specificity, and F1 score were 99.0%, 99.6%, 98.4%, and 99.01%, respectively, using a ten-fold cross-validation strategy.

Table 5.
The suggested method’s performance metrics after being subjected to ten-fold cross-validation with the softmax classifier.

Table 6.
The suggested method’s performance metrics after being subjected to ten-fold cross-validation with the RF classifier.

Table 7.
The suggested method’s performance metrics after being subjected to ten-fold cross-validation with the SVM classifier.
Since Softmax produces the best classification results over the RF and SVM classifiers, we will examine its performance.
The confusion matrix for the Softmax classifier is given in Figure 12. Two AD and four NC samples were misidentified. Figure 13 and Figure 14 represent the progress of training the developed 3D-CNN network using Softmax. At the beginning of the training, the training and test accuracy scores amounted to approximately 60%. At the end of the tenth iteration, both the training and test datasets achieved 100% accuracy scores. After the tenth iteration, the loss value for the training dataset decreased from over seven to approximately one. For the test dataset, the loss value dropped to around zero after the tenth iteration and remained during the training.
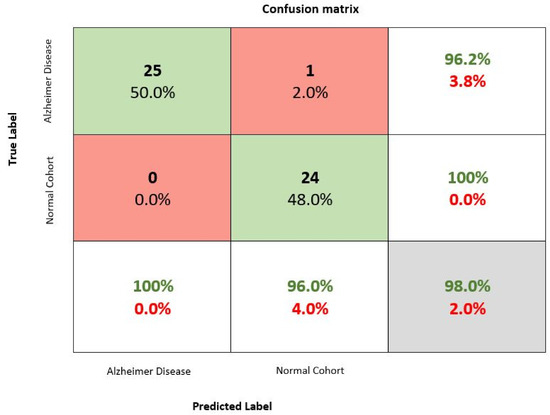
Figure 12.
Confusion matrix.
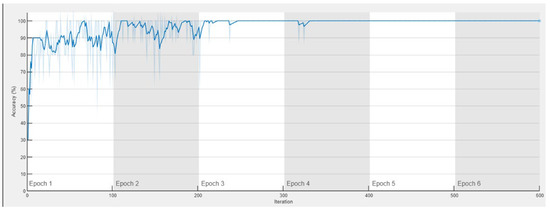
Figure 13.
Training accuracy.
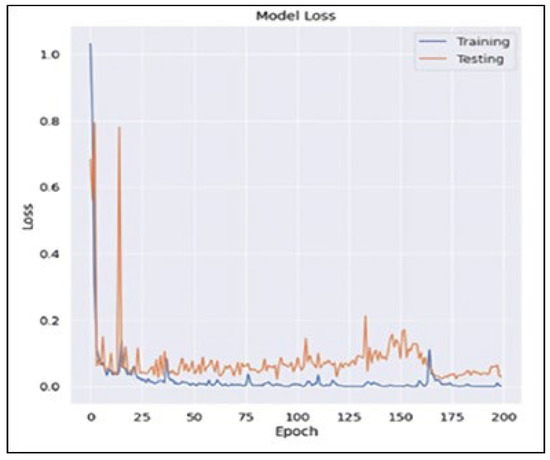
Figure 14.
Model loss.
4.3.3. Comparison with the State-of-the-Art Models
This section contrasts and evaluates the proposed MULTforAD image fusion based on the 3D-CNN efficiency with the state-of-the-art multimodal algorithms. The proposed AD diagnosis model has proven effective and outperformed the available MRI neuroimaging algorithms, with a classification rate (98.875%) and a low false alarm of 1.125%, as shown in Table 8. Moreover, the performance levels of the network with three classifiers show consistency with one another, and their hyperparameter optimization reduces significantly due to the use of single composite vector neuroimaging input rather than a set of different images. Therefore, the findings clarify that the suggested multimodal image fusion strategy provided a remarkably consistent high level of accuracy, and the computational complexity and storage cost will remain the same with the existing strategy.

Table 8.
The performance of MULTforAD in comparison with competitors.
4.4. Discussions
Here, we propose an image fusion approach based on a 3D-CNN model for diagnosing Alzheimer’s disease. Multimodal data can provide complete pathological information. This makes it possible to combine heterogeneous image information from MRI images successfully.
Moreover, while 2D MRI could have been used in the proposed model, we chose to convert the MRI image for analysis before inputting it into the model. Therefore, the model can distinguish the discriminative value of individual MRI neuroimages in diagnosing Alzheimer’s disease, although relationships among the neuroimages, which may aid expert interpretation, have not been preserved. Feature extraction (concatenation and selection operations) and classification must be fully incorporated within the model to provide an end-to-end DL solution. Due to the proposed MULTforAD data fusion model, the classification system may become more complex. For this, 3D image segments were built and placed into the data’s channels, and feature extraction and classification occurred using the 3D-CNN architecture.
Figure 2 shows that fusion images store the patient’s brain structure data from MRI and preserve the details of the patient’s metabolic data through rotation and resizing. The proposed image fusion method is also superior to techniques based on multimodal feature learning in its ability to handle the problem of cropping and skull removal of heterogeneous features between multimodal images. MULTforAD represents a more intuitive approach to combining neuroimaging features than existing methods. A fusion image combines relevant and supplemental information from multiple input images. In addition to having more powerful information representation features, the merged images also have more powerful modal features.
The suggested 3D CNN architecture would necessitate high-performance computers for high-dimensional input volumes. However, the proposed MULTforAD methodology uses a single network instead of the multi-input network used in the feature-matching process by merging multimodal image scans into a single aggregate image. Consequently, the number of CNN parameters has been significantly reduced. Moreover, the training and test times for one-fold running the algorithms on a single core-I12 CPU 8 GB RAM computer with NVIDIA GEFORCE GTX 1050 GPU amounted to 11 min and 3 s.
Using ten-fold cross-validation, the performance of the proposed image fusion approach is evaluated according to the classification results based on three different classifiers. Table 5, Table 6 and Table 7 and Figure 13 show the average accuracy rates obtained using the proposed 3D-CNN. According to the classification results in Table 4, Table 5, Table 6, Table 7 and Table 8, the MULTforAD method outperformed the unimodal methods because the multimodal approach contained more supplementary information. Additionally, 3D-CNN using the proposed multimodal neuroimaging has produced the highest performance. However, recent studies considering the unimodal pre-trained model, such as Resnet, show the highest overall accuracy (100%). However, such models do not work if the features learned from the classification layers cannot distinguish the AD classes. Thus, if the datasets are multimodal, the pre-trained characteristics will be inadequate, leading to overfitting problems. In addition, the proposed image fusion method outperforms methods based on multimodal learning with networks’ building blocks that extract features much smaller than a complex 3D convolution model.
However, the sensitivity and specificity of the model were not always optimal. To address this issue, we plan to focus on mask and ROI indicators such as WM and CSF tissues and combine them with existing preprocessing steps.
The following are the most crucial characteristics of the proposed MULTforAD model:
- (1)
- The method fused 5982 MRI neuroimages, allowing the model to learn all the features needed to distinguish AD from CN samples accurately and quickly accurately;
- (2)
- The suggested method provides anatomical and metabolic information without pre-trained models or transfers learning. In addition, it reduces noise with the scanned brain patterns based on the multimodal image fusion method and 3D-CNN;
- (3)
- Using lite models that comprise a smaller number of convolutional blocks and training parameters, the suggested model achieved the highest classification accuracy among the recent multimodal-based AD classification methods with 98.8593% accuracy.
The following are the limitations of our work:
- (1)
- Increasing the size of the input neuroimaging raises the proposed model’s computational complexity, and storage costs increase as well;
- (2)
- Designing an unbiased neuroimaging dataset is tricky and generates sensitivity and specificity artifacts. Therefore, incorporating additional tissue filters for neuroimaging could help overcome this limitation.
5. Conclusions
This study proposed a new approach for AD identification using multimodal MRI fusion based on DL. The proposed model was trained and optimized to provide more accurate and comprehensive classifications with AD and CN. Moreover, the proposed method optimizes and improves data fusion, enhancement, and oversampling tasks, considering three publicly available MRI neuroimaging datasets. Therefore, it can effectively monitor and track older people daily. A 3D-CNN network for learning the fused 3D characteristics that best represent AD biomarkers is proposed, and a series of experiments prove its effectiveness. The average achieved accuracy rate is 98.21, 90.77, and 86.01 when considering Softmax, SVM, and RF, respectively, with the proposed model for AD classifications. In the future, we plan to combine different brain scans, such as PET images, to create a composite fusion modality that may enrich the classification accuracy and extend the model to multi-class classification problems.
Author Contributions
Conceptualization, W.N.I., F.R.P. and M.A.S.A.; data curation, W.N.I. and F.R.P.; formal analysis, W.N.I., F.R.P. and M.A.S.A.; funding acquisition, W.N.I., F.R.P. and M.A.S.A.; investigation, W.N.I., F.R.P. and M.A.S.A.; methodology, W.N.I., F.R.P. and M.A.S.A.; project administration, W.N.I., F.R.P. and M.A.S.A.; resources, W.N.I.; software, W.N.I.; supervision, W.N.I., F.R.P. and M.A.S.A.; validation, W.N.I., F.R.P. and M.A.S.A.; visualization, W.N.I. and M.A.S.A.; writing—original draft, W.N.I., F.R.P. and M.A.S.A.; writing—review and editing, W.N.I., F.R.P. and M.A.S.A. All authors have read and agreed to the published version of the manuscript.
Funding
This research has been funded by The Deanship of Scientific Research, King Faisal University, Saudi Arabia, with grant Number (GRANT2094).
Data Availability Statement
The study did not report any data.
Acknowledgments
We deeply acknowledge The Deanship of Scientific Research, King Faisal University, Saudi Arabia, as they funded our project with grant Number (GRANT2094).
Conflicts of Interest
The authors declare no conflict of interest.
References
- Cunnane, S.; Nugent, S.; Roy, M.; Courchesne-Loyer, A.; Croteau, E.; Tremblay, S.; Castellano, A.; Pifferi, F.; Bocti, C.; Paquet, N.; et al. Brain fuel metabolism, aging, and Alzheimer’s disease. Nutrition 2011, 27, 3–20. [Google Scholar] [CrossRef] [PubMed]
- Schäfer, A.; zu Schwabedissen, H.M.; Grube, M. Expression and function of organic anion transporting polypeptides in the human brain: Physiological and pharmacological implications. Pharmaceutics 2021, 13, 834. [Google Scholar] [CrossRef] [PubMed]
- Scuderi, C.; Valenza, M. How useful are biomarkers for the diagnosis of Alzheimer’s disease and especially for its therapy? Neural Regen. Res. 2022, 17, 2205. [Google Scholar] [CrossRef] [PubMed]
- Mansour, H.M.; Fawzy, H.M.; El-Khatib, A.S.; Khattab, M.M. Potential Repositioning of Anti-cancer EGFR Inhibitors in Alzheimer’s Disease: Current Perspectives and Challenging Prospects. Neuroscience 2021, 469, 191–196. [Google Scholar] [CrossRef]
- Quintas-Neves, M.; Teylan, M.A.; Morais-Ribeiro, R.; Almeida, F.; Mock, C.N.; Kukull, W.A.; Crary, J.F.; Oliveira, T.G. Divergent magnetic resonance imaging atrophy patterns in Alzheimer’s disease and primary age-related tauopathy. Neurobiol. Aging 2022, 117, 1. [Google Scholar] [CrossRef]
- Zhou, Z.; Yu, L.; Tian, S.; Xiao, G. Diagnosis of Alzheimer’s disease using 2D dynamic magnetic resonance imaging. J. Ambient Intell. Humaniz. Comput. 2022, 1–11. [Google Scholar] [CrossRef]
- Odusami, M.; Maskeliūnas, R.; Damaševičius, R.; Krilavičius, T. Analysis of features of alzheimer’s disease: Detection of early stage from functional brain changes in magnetic resonance images using a fine-tuned ResNet18 network. Diagnostics 2021, 11, 1071. [Google Scholar] [CrossRef]
- Lodha, P.; Talele, A.; Degaonkar, K. Diagnosis of alzheimer’s disease using machine learning. In Proceedings of the 2018 Fourth International Conference on Computing Communication Control and Automation (ICCUBEA), Pune, India, 6–18 August 2018; IEEE: Piscataway, NJ, USA, 2019; pp. 1–4. [Google Scholar]
- Ismail, W.N.; Hassan, M.M.; Alsalamah, H.A.; Fortino, G. CNN-based health model for regular health factors analysis in internet-of-medical things environment. IEEE Access 2020, 8, 52541–52549. [Google Scholar] [CrossRef]
- Kumar, Y.; Koul, A.; Singla, R.; Ijaz, M.F. Artificial intelligence in disease diagnosis: A systematic literature review, synthesizing framework and future research agenda. J. Ambient Intell. Humaniz. Comput. 2022, 1–28. [Google Scholar] [CrossRef]
- Yuvalı, M.; Yaman, B.; Tosun, Ö. Classification Comparison of Machine Learning Algorithms Using Two Independent CAD Datasets. Mathematics 2022, 10, 311. [Google Scholar] [CrossRef]
- Alzheimer’s Association. 2019 Alzheimer’s disease facts and figures. Alzheimer’s Dement. 2019, 15, 321–387. [Google Scholar] [CrossRef]
- Ullah, Z.; Farooq, M.U.; Lee, S.-H.; An, D. A hybrid image enhancement based brain MRI images classification technique. Med. Hypotheses 2020, 143, 109922. [Google Scholar] [CrossRef]
- Amini, M.; Pedram, M.M.; Moradi, A.; Jamshidi, M.; Ouchani, M. GC-CNNnet: Diagnosis of Alzheimer’s Disease with PET Images Using Genetic and Convolutional Neural Network. Comput. Intell. Neurosci. 2022, 2022, 7413081. [Google Scholar] [CrossRef]
- Venugopalan, J.; Tong, L.; Hassanzadeh, H.R.; Wang, M.D. Multimodal deep learning models for early detection of Alzheimer’s disease stage. Sci. Rep. 2021, 11, 3254. [Google Scholar] [CrossRef]
- Hazarika, R.A.; Kandar, D.; Maji, A.K. An experimental analysis of different deep learning based models for Alzheimer’s disease classification using brain magnetic resonance images. J. King Saud Univ.-Comput. Inf. Sci. 2021, 34, 5876–8598. [Google Scholar] [CrossRef]
- Li, Y.; Wang, Z.; Yin, L.; Zhu, Z.; Qi, G.; Liu, Y. X-Net: A dual encoding–decoding method in medical image segmentation. Vis. Comput. 2021, 1–11. [Google Scholar] [CrossRef]
- Ren, S.; He, K.; Girshick, R.; Sun, J. Faster r-cnn: Towards real-time object detection with region proposal networks. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 39, 1137–1149. [Google Scholar] [CrossRef]
- Zhu, Z.; Yin, H.; Chai, Y.; Li, Y.; Qi, G. A novel multimodality image fusion method based on image decomposition and sparse representation. Inf. Sci. 2018, 432, 516–529. [Google Scholar] [CrossRef]
- Silveira, M.; Marques, J. Boosting Alzheimer disease diagnosis using PET images. In Proceedings of the 2010 20th International Conference on Pattern Recognition, Istanbul, Turkey, 23–26 August 2010; IEEE: Piscataway, NJ, USA, 2010; pp. 2556–2559. [Google Scholar]
- Liu, M.; Cheng, D.; Yan, W.; Alzheimer’s Disease Neuroimaging Initiative. Classification of Alzheimer’s disease by combination of convolutional and recurrent neural networks using FDG-PET images. Front. Neuroin. Form. 2018, 12, 35. [Google Scholar] [CrossRef]
- Chételat, G. Multimodal neuroimaging in Alzheimer’s disease: Early diagnosis, physiopathological mechanisms, and impact of lifestyle. J. Alzheimer’s Dis. 2018, 64, S199–S211. [Google Scholar] [CrossRef]
- Lin, A.-L.; Laird, A.R.; Fox, P.T.; Gao, J.-H. Multimodal MRI neuroimaging biomarkers for cognitive normal adults, amnestic mild cognitive impairment, and Alzheimer’s disease. Neurol. Res. Int. 2012, 2012, 907409. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Biswas, M.; Mahbub, M.; Miah, M.; Mozid, A. An Enhanced Deep Convolution Neural Network Model to Diagnose Alzheimer’s Disease Using Brain Magnetic Resonance Imaging. In International Conference on Recent Trends in Image Processing and Pattern Recognition; Springer: Cham, Switzerland, 2022; pp. 42–52. [Google Scholar]
- Kang, J.; Ullah, Z.; Gwak, J. Mri-based brain tumor classification using ensemble of deep features and machine learning classifiers. Sensors 2021, 21, 2222. [Google Scholar] [CrossRef] [PubMed]
- Ullah, Z.; Usman, M.; Jeon, M.; Gwak, J. Cascade multiscale residual attention cnns with adaptive roi for automatic brain tumor segmentation. Inf. Sci. 2022, 608, 1541–1556. [Google Scholar] [CrossRef]
- Baghdadi, N.A.; Malki, A.; Balaha, H.M.; Badawy, M.; Elhosseini, M. A3C-TL-GTO: Alzheimer Automatic Accurate Classification Using Transfer Learning and Artificial Gorilla Troops Optimizer. Sensors 2022, 22, 4250. [Google Scholar] [CrossRef]
- Kong, Z.; Zhang, M.; Zhu, W.; Yi, Y.; Wang, T.; Zhang, B. Multimodal data Alzheimer’s disease detection based on 3D convolution. Biomed. Signal Process. Control 2022, 75, 103565. [Google Scholar] [CrossRef]
- Kaur, S.; Gupta, S.; Singh, S.; Gupta, I. Detection of Alzheimer’s disease using deep convolutional neural network. Int. J. Image Graph. 2022, 22, 2140012. [Google Scholar] [CrossRef]
- Gorgolewski, K.J.; Auer, T.; Calhoun, V.D.; Craddock, R.C.; Das, S.; Duff, E.P.; Flandin, G.; Ghosh, S.S.; Glatard, T.; Halchenko, Y.O.; et al. The brain imaging data structure, a format for organizing and describing outputs of neuroimaging experiments. Sci. Data 2016, 3, 160044. [Google Scholar] [CrossRef]
- Plataniotis, K.; Venetsanopoulos, A.N. Color Image Processing and Applications; Springer Science & Business Media: Berlin/Heidelberg, Germany, 2000. [Google Scholar] [CrossRef]
- Fatima, A.; Shahid, A.R.; Raza, B.; Madni, T.M.; Janjua, U.I. State-of-the-art traditional to the machine-and deep-learning-based skull stripping techniques, models, and algorithms. J. Digit. Imaging 2020, 33, 1443–1464. [Google Scholar] [CrossRef]
- Piotrowski, A.P.; Napiorkowski, J.J. A comparison of methods to avoid overfitting in neural networks training in the case of catchment runoff modelling. J. Hydrol. 2013, 476, 97–111. [Google Scholar] [CrossRef]
- Miah, Y.; Prima, C.N.E.; Seema, S.J.; Mahmud, M.; Kaiser, M.S. Performance comparison of machine learning techniques in identifying dementia from open access clinical datasets. In Advances on Smart and Soft Computing; Springer: Singapore, 2021; pp. 79–89. [Google Scholar]
- Soliman, S.A.; El-Dahshan, E.-S.A.; Salem, A.-B.M. Deep Learning 3D Convolutional Neural Networks for Predicting Alzheimer’s Disease (ALD). In New Approaches for Multidimensional Signal Processing; Springer: Singapore, 2022; pp. 151–162. [Google Scholar]
- Liu, S.; Liu, S.; Cai, W.; Che, H.; Pujol, S.; Kikinis, R.; Feng, D.; Fulham, M.J. Multimodal neuroimaging feature learning for multi-class diagnosis of Alzheimer’s disease. IEEE Trans. Biomed. Eng. 2014, 62, 1132–1140. [Google Scholar] [CrossRef]
- Tong, T.; Gray, K.; Gao, Q.; Chen, L.; Rueckert, D.; Alzheimer’s Disease Neuroimaging Initiative. Multimodal classification of Alzheimer’s disease using nonlinear graph fusion. Pattern Recognit. 2017, 63, 171–181. [Google Scholar] [CrossRef]
- Shao, W.; Peng, Y.; Zu, C.; Wang, M.; Zhang, D.; Alzheimer’s Disease Neuroimaging Initiative. Hypergraph based multi-task feature selection for multimodal classification of Alzheimer’s disease. Comput. Med. Imaging Graph. 2020, 80, 101663. [Google Scholar] [CrossRef]
- Raees, P.C.M.; Thomas, V. Automated detection of Alzheimer’s Disease using Deep Learning in MRI. J. Phys. Conf. Ser. 2021, 1921, 012024. [Google Scholar] [CrossRef]
- Ji, H.; Liu, Z.; Yan, W.Q.; Klette, R. Early diagnosis of Alzheimer’s disease using deep learning. In Proceedings of the 2nd International Conference on Control and Computer Vision, Jeju Island, Republic of Korea, 15–18 June 2019; Association for Computing Machinery: New York, NY, USA, 2019; pp. 87–91. [Google Scholar]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).