1. Introduction
Magnetic Resonance Imaging (MRI) is preferred for the structural and functional analysis of several organs in the clinical setting thanks to its non-ionizing nature and ability to highlight structures with high contrast. In particular, MR neuroimaging is widely employed in the screening and diagnosis of brain cancers and neurodegenerative dysfunctions such as Alzheimer’s disease and multiple sclerosis [
1]. MRI can highlight tissue with various contrasts using different sequences of Radio-Frequency (RF) pulses. Specific pathologies are accurately analyzed and interpreted when captured using a particular RF pulse sequence. For instance, ‘substantia nigra’, a brain area affected due to Parkinson’s disease can be visualized clearly on T2-w images compared to T1-w [
2], whereas, T1-w images are preferred in the quantification of atrophy, an irreversible loss of neurons associated with multiple sclerosis [
3]. However, certain pathologies possess uncertain features and assorted topography, whose existence needs to be validated by multiple modalities especially if their surgical resection is essential. A cohort study comprising 200 surgically treated Craniopharyngiomas (CPs), an infiltrative brain tumor concluded that several key radiological variables recognized on both T1-w and T2-w MR images correctly predicted the CP topography in 86% of cases [
4].
During MRI acquisition, noise is mainly introduced due to motion of charged particles in the radio frequency coils. This noise affects the reliability of diagnosis and image analysis tasks including feature extraction and segmentation [
5,
6]. Denoising of the images then becomes indispensable to make them suitable for further analysis.
and
Y denote the ideal and observed MR images, respectively, and
N is the noise contained in the MRI signal. The noisy observation
Y of
X can be expressed in the case of an additive model as:
The objective of denoising algorithms is to reduce the noise content
N in
Y to obtain an estimate of the original image
X. The noise in MR images follows Rician distribution whose probability density function is expressed as:
In the above equation, represents the 0th order Bessel function, N is a Rician distributed random variable. is the unit step Heaviside step function indicating that the pdf expression is valid for non-negative values of N. X is a non-noisy signal as stated above and is the noise variance. The Rician noise is a signal dependant noise and demonstrates gaussian distribution when Signal-to-Noise Ratio (SNR) is high and rayleigh distribution when SNR is low.
Despite the considerable amount of work devoted to image denoising during the two last decades, it is still a challenging problem particularly in the case of signal-dependant and correlated noise [
7,
8]. This is the case in medical imaging. Most often simplifying assumptions are made to make the denoising problem more or less tractable. This has led to a variety of denoising methods applied to various imaging modalities. Several denoising approaches have been proposed in the past that can be broadly grouped into two types: conventional methods and deep learning-based approaches. The conventional denoising methods include spatial domain methods such as bilateral filter [
9], Non-Local Means filter (NLM) [
10] and anisotropic filter [
11] to name a few. Among these filters, the NLM filter specifically demonstrates superior performance when the image contains regions of various types of textures. Wavelet domain approaches were also widely researched for image quality enhancement [
12,
13,
14]; one such approach applies thresholding on the detail coefficients. The wavelet-based denoising methods well preserve sharp edges in the images compared to spatial domain methods. Optimization-based denoising techniques including total-variation denoising [
15] provide more control over preserving details in the image and the extent of noise reduction. Recently, data driven machine learning approaches, particularly deep learning methods are gaining incredible attention due to their promising performance in various areas such as biomedicine [
16,
17,
18,
19], video processing. These methods are able to mimic human cognition [
20,
21]. Similarly, these approaches clearly outperform the conventional approaches in the area of denoising [
22,
23,
24].
Indeed, acquisition of multi-modal medical imaging data during therapeutics is becoming increasingly common [
25,
26]. Since these diagnostic imaging techniques are one of the largest sources of big data [
27,
28,
29,
30,
31], their automated analysis is highly desirable to facilitate the computer aided diagnosis of several diseases [
32,
33,
34,
35]. For instance, Computed Tomography (CT) and positron emission tomography (PET) are concurrently acquired as a standard treatment protocol in oncology. Similarly, T1 and T2-w MRI provides anatomical and pathological information, respectively. The combination of this complementary information plays a significant role in therapy and surgical planning. The concept of ‘weak learnability’ in ensemble learning further motivates to exploit the strength of this complimentarity. According to this concept, the learner (imaging modality here) can be incorporated into the learning system to elevate its performance, provided it can perform slightly better than random guessing [
36].
With technical advancement and the availability of medical imaging techniques, using multi-modal data for the underlying computer-aided tasks is attracting several researchers. It has been exploited in segmentation, classification, super-resolution, and denoising [
36,
37,
38]. For instance, in the context of lung nodule detection, CT and PET images were combined in a CNN-based approach [
37]. Similarly, CT, PET, and MRI were also combined for tumor segmentation [
36]. It is worth mentioning here that multi-modal information-based methods showed superior performance compared to those relying on a single modality (either CT or PET) [
39].
The use of multi-modal medical imaging methods in improving segmentation and object detection motivates the researchers to employ the dual imaging in denoising as well. Few research works presented for medical image denoising [
23,
40] show improved performance over their single image denoising counterparts. Single image denoising approaches have an intrinsic limitation where the corrupted information in the original image is only hallucinated during the reconstruction process [
41]. Consequently, these approaches over smooth certain critical structures in the image at the expense of removing noise [
42]. It often leads to compromised performance of segmentation and object detection algorithms [
43]. In this context, techniques that rely on cross-modal guidance offer the potential to overcome this limitation. Conventionally, cross-modal denoising methods use an image of better perceptual quality to facilitate the restoration process. Cross-modality guided medical image denoising is a relatively under-explored area; however, there exist a few approaches for natural images. One of the traditional denoising methods attempted to denoise depth maps using corresponding RGB images [
44,
45]. Deep learning-based cross-modal denoising approaches include [
46,
47]. One of these methods uses RGB-depth data pair to denoise depth images. Their proposed method consists of two CNNs; first to extract features individually from the RGB (guidance) and depth (target) images; the features are later concatenated to be fed to the third CNN, which selectively transfers the common structures in both images to generate the denoised image [
46]. This work was further extended by adding a skip connection between the input image and the network prediction to enforce residual learning [
47]. This modification brought significant improvement in the results by leveraging accurate details from the guidance to the target image.
A few cross-modal medical image denoising methods including [
23,
48,
49] are found in the literature. One such work consolidated information from PET and MRI (T1 and T2 FLAIR) to denoise very low-dose PET images of the human brain [
23]. The proposed method ResUNet was a residual encoder-decoder network, where residual learning was combined with U-Net. The PET, T1 and T2 FLAIR slices were stacked together and fed to the network. Using 2.5D information offers a way to discriminate structural information from noise. Compared to the ResUNet (PET without MRI), the combination of both modalities not only resulted in improved denoising performance but also improved lesion segmentation. In another similar CNN-based approach, amyloid PET images were concatenated with corresponding T1, T2, and T2 FLAIR images to learn denoising ultra-low-dose PET images using standard dose PET as ground truth. U-Net with residual learning was used in their approach. A similar idea was applied to T1 and T2 brain images [
48]. The traditional guided filter was integrated with the deep learning framework, where guidance map generator takes guidance and cross-modal noisy images as input (T1 and T2 MR images). The guidance map generation component was realized using a modification of popular architecture, U-Net; where the encoding path was extended to dual branches for each modality followed by feature concatenation at the last encoding layer. The guidance filter was then incorporated as a differential layer and implemented as a linear combination of the guidance map and input image to yield the restored image. The method claimed to outperform approaches that do not include the guidance information from input image directly in the restoration process and rather only rely on the network prediction as final output.
The above-mentioned approaches are not very effective since simply concatenating the images as network input as in [
23] or combining features from all encoding layers [
48] does not fully exploit the potential of cross-modal complementarity. It leaves a huge space to further explore the improvement of cross-modal denoising methods and advance in this direction. Therefore, there is a need for a more efficient way of manipulating and combining features. To address the denoising problem in MR images, we present a cross-modality-guided denoising approach
in this paper. The proposed model is inspired by the work of Fu et al. [
50], where a similar model was used to detect salient objects in RGB-Depth images. Cross-modal image denoising for brain MR images was earlier explored by Stimpel et al. [
48]; however, simple feature concatenation at the last encoding layer of their proposed method does not effectively exploit the information in the non-noisy guidance image. Unlike the previous denoising approaches,
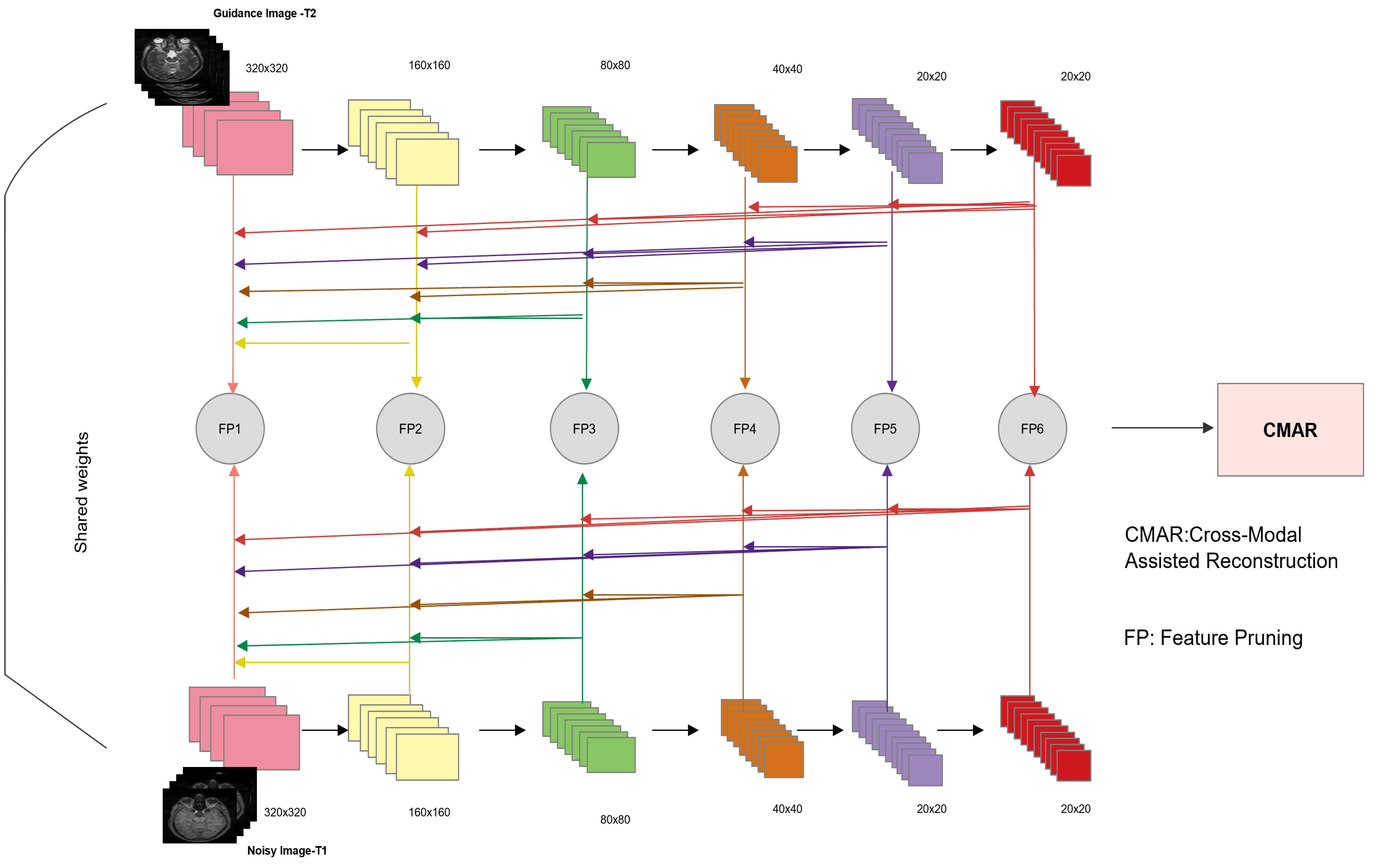
extracts hierarchical features from the input and guidance image using a siamese network (mirror backbones) that are later combined in the complimentarity-aware mechanism. Although T1 and T2 images belong to different modalities; nonetheless, they capture similar structures and analogous object contours. The guidance image (T2 in our case) has better perceptual quality (noise-free), while T1 is of lower quality due to its sensitivity to acquisition noise. This scenario renders cross-modal feature learning viable in the presence of a guidance image. Our contributions in this work are listed as follows:
A novel framework based on cross-modal guidance information is designed to denoise T1-w brain MR images. In particular, a siamese network is specifically modified to train the denoising network using both T1 and T2 MR images. By exploiting the diversity of information contained in the two modalities and in particular better perceptual quality of T2 images and the structural information contained in T1 images, the proposed approach seeks additional guidance from these images in the reconstruction process.
Literature dictates that complementarity-aware cross-modal feature fusion is not well explored in the context of denoising, hence in this work, an effective cross-modal information fusion strategy is incorporated. The experimental results show that this fusion mechanism works well in comparison to single image denoising approaches.
Comprehensive experiments have been conducted to analyze the performance of the proposed method on different noise levels both on registered as well as unregistered data. Moreover, the role of different loss functions is inspected to analyze their impact on denoising performance.
In this work, two public datasets are customized keeping in view the requirement of denoising in medical image analysis. The dataset consists of both T1 and T2 MR images, meeting the requirement of learning models based on cross-modal guidance.
This paper consists of five sections.
Section 1 gives an introduction to and motivations for the work followed by background and related work. The dataset and proposed methodology are elaborated on in
Section 2. Experiments, comparisons with different techniques, and the results are discussed in
Section 3.
Section 4 summarizes the discussion of results. The conclusion and suggested future work are presented in
Section 5.
3. Experiments and Results
The performance of the proposed method was validated by comparing it with five state-of-the-art methods including Non-local means filter (NLM) [
10], Stein’s unbiased risk estimate (SURE) [
58], Block-matching and 3D filtering (BM3D) [
59], Multi-channel Denoising convolutional neural network (MCDnCNN), referred as MCDN in the paper [
24] and FFD-Net [
53]. Among the methods chosen, NLM [
10] is a popular denoising method that computes the weighted average of not only the local neighborhood but all pixels in the image. Wavelet-based denoising approach SURE does not rely on prior statistical modeling of wavelet coefficients [
58]. Instead, it parametrizes denoising by computing parameters that minimize this MSE estimate. BM3D is a popular approach based on stacking similar 2D image patches followed by hard thresholding and Wiener filtering to denoise 3D stacks [
59]. Although BM3D was originally developed for removing Gaussian noise in images; however, it has been applied to Rician noise removal as well [
60]. MCDN is a 10 convolution layer network embedded with residual learning taking multi-channel input; however, we modified it to take identical slices. FFD-Net [
53] is another CNN architecture that is capable of handling a variable range of noise levels in a single model.
Moreover, the denoising performance was also evaluated by using different combinations of loss functions on registered as well as unregistered data. Three metrics were used to quantitatively evaluate the performance. The first metric peak signal-to-noise ratio (PSNR) compares the root mean square error (RMSE) between the ground truth and denoised images. Another metric Structural Similarity Index (SSIM) was also included in the assessment that measures the structural affinity between denoised images and the ground truth. Feature Similarity Index (FSIM) [
61] is a full reference image quality assessment (IQA) metric that is often used to evaluate the performance of denoising methods [
62]. It computes feature similarity between the two images based on the low-level features including phase congruency and gradient magnitude.
In the following subsections, we describe in detail the experiments conducted on the brain MR images using the proposed method and state-of-the-art methods.
3.1. Configurations
The performance of the proposed method
was evaluated on unregistered and registered data with different combinations of loss functions. Different configurations of data and loss functions tested in the proposed method are mentioned in
Table 1 and briefly explained below:
3.1.1.
Under this configuration, registration was not performed between T1-w and T2-w volumes. Using MSE as loss function, the model was trained and then tested on both datasets. The results of this configuration are referred as ’’.
3.1.2.
The role of using an additional SSIM-based loss function was analyzed in case of unregistered data under this configuration. Therefore, both SSIM and MSE were combined here.
3.1.3.
In this case, registration was performed between T1 and T2 volumes. Registration was done using 3D Slicer. Rigid Registration with 12 degrees of freedom was applied in all cases where T1 volume was fixed, while T2 was moved with reference to T1 in the registration process. The effect of registration can be better comprehended by visually inspecting the registered and unregistered T2 images with reference to T1 in
Figure 4. It can be noticed that T1 and unregistered T2 slices are structurally similar; however, careful insight points to structural mismatches at various regions in the image. After applying registration, the structural similarity in the registered T2 image can be seen in the highlighted areas. In the following experimental sections, we further analyze the impact of registration on denoising and structural preservation in the presence of cross-modal image T2. The loss function used in this case is MSE.
3.1.4.
SSIM was combined with MSE for analyzing the performance of the proposed method on the registered data in this configuration.
Next, we explain the experiments conducted to compare the performance of with other denoising methods.
3.2. Experiment I
The first set of experiments was conducted by comparing the proposed method (‘’ configuration was used in this set of experiments) with state-of-the art denoising methods. The experiments were conducted on the T1 images taken from two datasets, HH and Guy’s, corrupted by Rician noise in the range 5% to 13%.
3.3. Experiment II
The second experiment was conducted to investigate the impact of registration on the denoising performance; besides, the role of using different loss functions was also evaluated. Therefore, the experiments were conducted using the four configurations
,
,
, and
, in
Table 1.
3.4. Experiment III
Another experiment was conducted to investigate the impact of integrating corresponding cross-modal images in the proposed framework and analyze its impact in denoising and preserving the structural information in the image. In order to do this, a noisy input image (T1) was fed to both the branches of the PHL module instead of the combination of noisy input and cross-modal (guidance) image. The model was then trained using MSE and SSIM losses on Guy’s hospital dataset (contaminated with 13% noise).
4. Discussion
In this section, we summarize the discussion of our results. The results of Experiment I are shown in
Figure 5,
Figure 6 and
Figure 7, where the denoising performance of the proposed method is shown in comparison with state-of-the-art denoising methods. In
Figure 5, the input images were contaminated using 13% noise. All the images denoised using different approaches suppress noise to some extent; however, NLM [
10] removes important structural details in the image and oversmoothes the contents of the denoised image during the restoration. Wavelet-based technique SURE [
58] and BM3D [
59] preserve the structural details; however, they do not eradicate noise to a reasonable extent. The deep learning methods clearly show better performance compared to the traditional methods, both in removing noise and maintaining the morphology of the image. Both MCDN [
24] and FFD-Net [
53] effectively remove the noise. Similarly,
also eradicates noise with reasonable preservation of the structural information. The enlarged ROIs are also shown in the figure for careful insight into the denoising performance of all the methods.
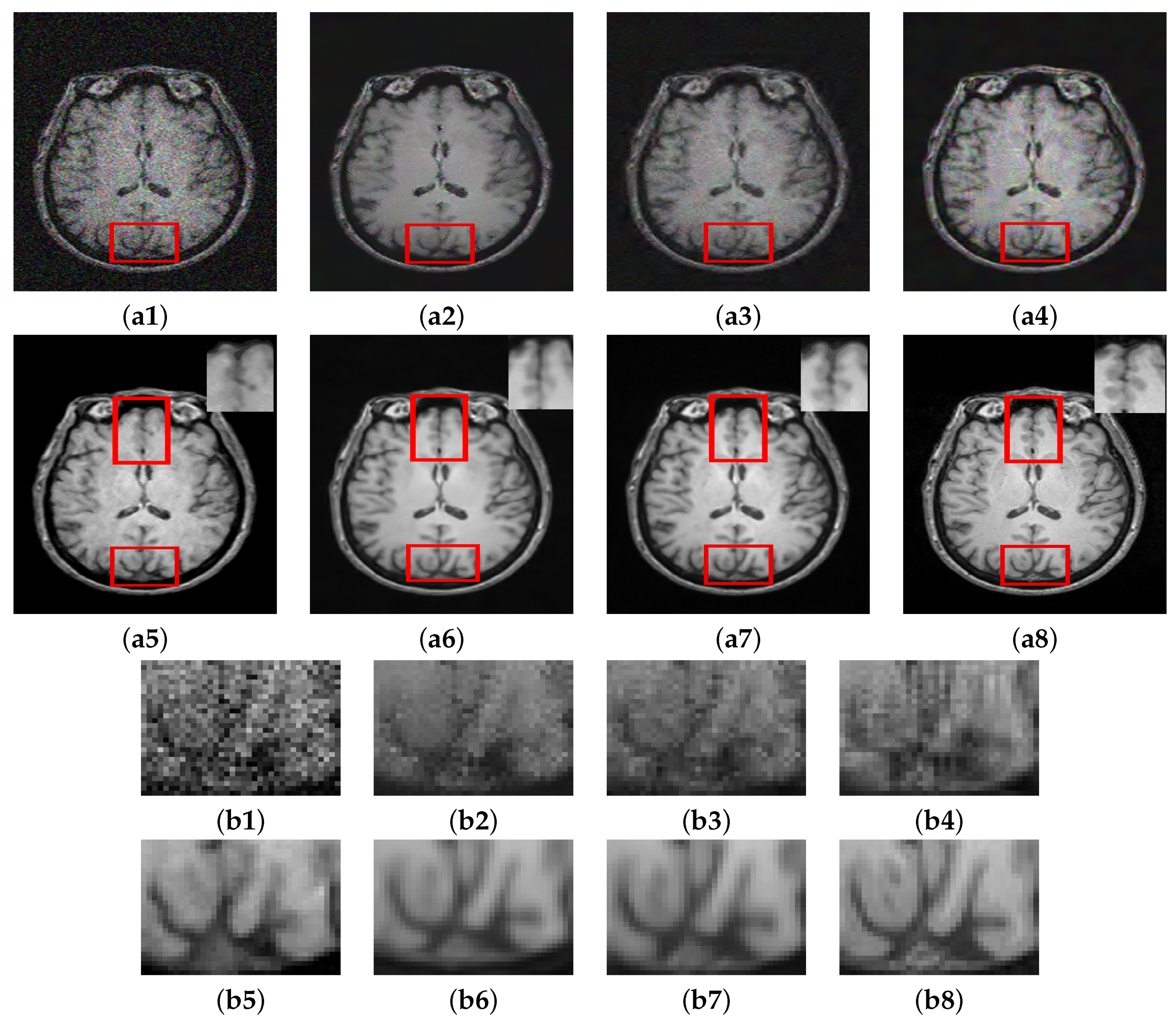
Figure 6 shows the results of denoising applied on images contaminated with 8% noise. A similar trend can be observed in this case as well where the methods MCDN [
24], FFD-Net [
53], and
preserve important structures in the denoised images. However, NLM [
10] produces over-smoothing effects. The performance was quantitatively evaluated using PSNR, SSIM, and FSIM. BM3D [
59] works better compared to NLM and SURE [
58]; this claim is also supported by the higher PSNR value in
Table 2. The performance of FFD-Net [
53] and MCDN is very similar when quantitatively evaluated. However,
performs best among all the techniques evaluated.
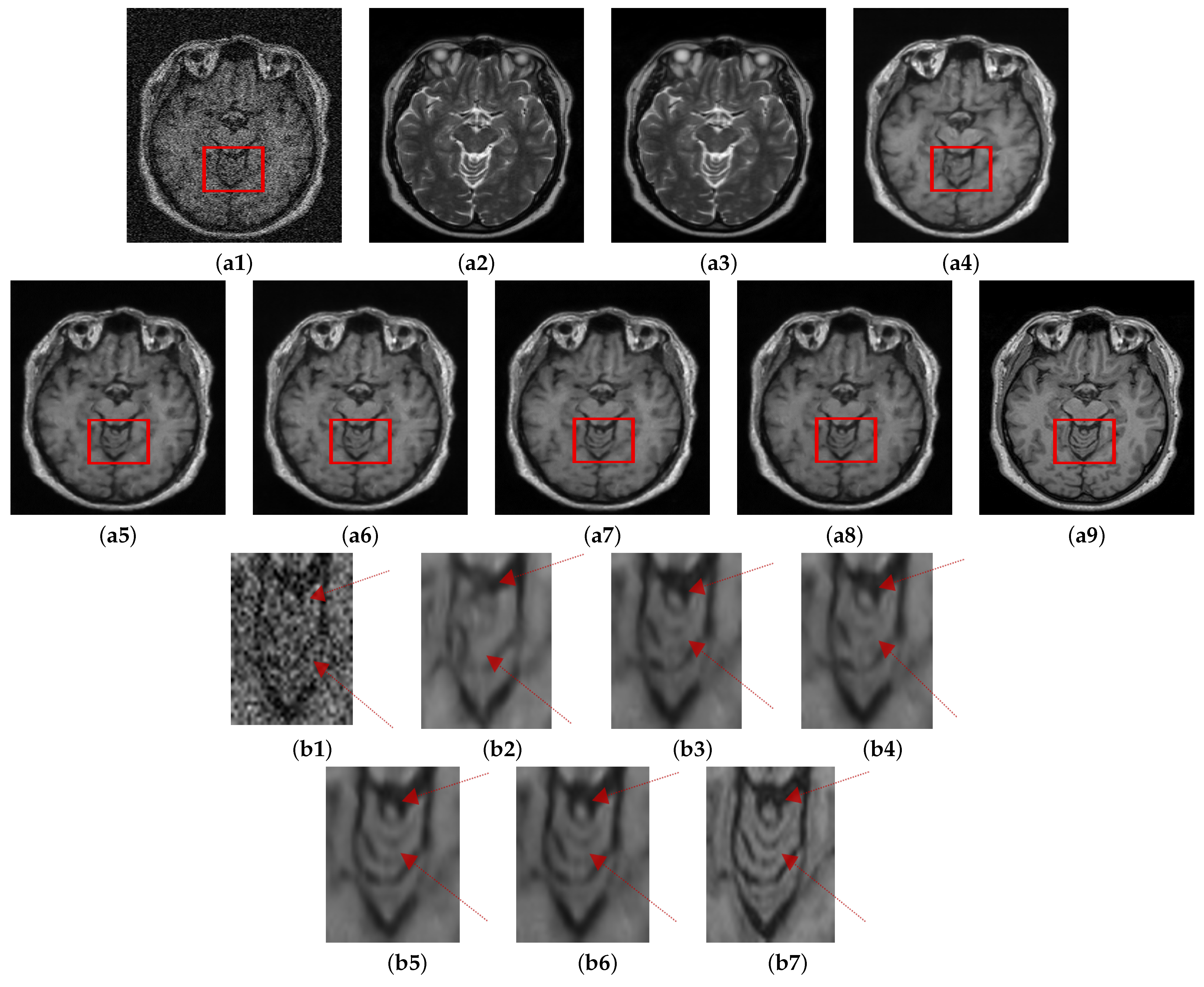
The visual comparison of the performance of the proposed method with other denoising methods conducted on the Guys dataset (13% noise) is shown in
Figure 7. NLM and SURE exhibit worse performance among all the methods tested. NLM eradicates significant details from the image while SURE removes minimal noise. BM3D performs slightly better than the two approaches. MCDN preserves structural information of the image; however, it leaves some noticeable noise in the image. The performance of FFD-Net visually in this case is comparable with
. The quantitative assessment also validates the visual observations, which are shown in
Table 3. For instance, NLM and SURE are ranked low at all the noise levels by PSNR and SSIM. BM3D performs better than both NLM and SURE. It is pertinent to mention that even the more robust conventional denoising methods such as BM3D leveraging the benefits of spatial and transform domains rely on pre-defined assumptions that do not work well under several types and levels of noise. On the other hand, deep learning approaches allow the underlying model to learn various levels of feature representations from raw to the higher level. In the context of denoising, the model thus learns the uncertain noise distributions from the input data. Consequently, these techniques can adapt to several types of noise efficiently. The deep learning methods in the proposed study perform better than the conventional methods on all the metrics. However, the cross-modal image information further enhances the network learning capability. Overall, the images denoised using all the methods still look blurry compared to the ground truth. It is because it is not possible to recover the image contents completely that have been corrupted by noise without any loss of information. However, it can be sensed that the denoising at level 8% introduces less blur compared to the denoising applied to images containing 13% noise. Overall, the proposed method achieves the best performance among all the methods both in PSNR and SSIM.
exhibits an average gain of 4.7% in SSIM value compared to the second-best MCDN (0.89 against 0.85).
All the denoising methods included in this study bring improvement in preserving low-level features in the restored images when compared to the input noisy image as can be seen in terms of FSIM values (
Table 4 and
Table 5). It is worth mentioning here that the FSIM scores for all the methods are very close particularly at low noise levels (5%). However, this difference is more pronounced at the higher noise levels (13%). For instance, at 13% noise, the proposed
method shows the best performance on both datasets. The average gain in FSIM values in the case of
(FSIM value 0.903) compared to the second-best performing method FFD-Net [
53] (FSIM value 0.883) was 2.3%.
Another experiment (Experiment II) was conducted on the HH dataset using 13% noise. The denoising results are shown in
Figure 8 along with enlarged regions for careful inspection.
Table 6 shows the quantitative assessment results on different variants of data (i.e., registered and unregistered) using two different loss functions. Among the variants of the proposed method, it is observed that registration between the corresponding T1 and T2 images together with employing SSIM as loss function with MSE facilitates in improving the structural similarity between denoised image and ground truth as implied by the higher SSIM values in the case of
compared to its corresponding variants
and
; however, noticeable improvement in PSNR values was not observed under this configuration.