Significance of Ubiad1 for Epidermal Keratinocytes Involves More Than CoQ10 Synthesis: Implications for Skin Aging
Abstract
:1. Introduction
2. Materials and Methods
2.1. Structure of Ubiad1 Inducer Peptide
2.2. Antibodies
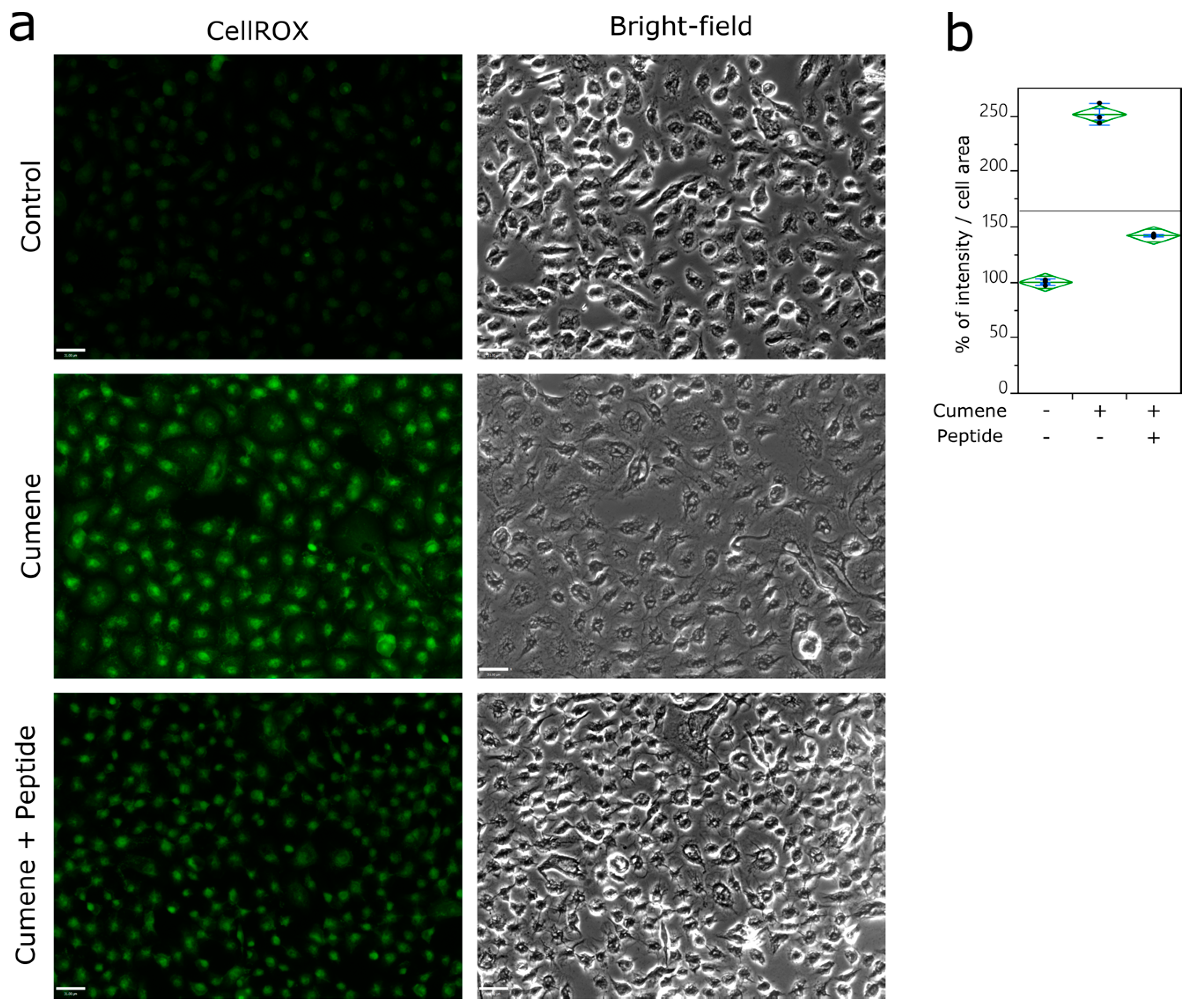
2.3. Detection of ROS
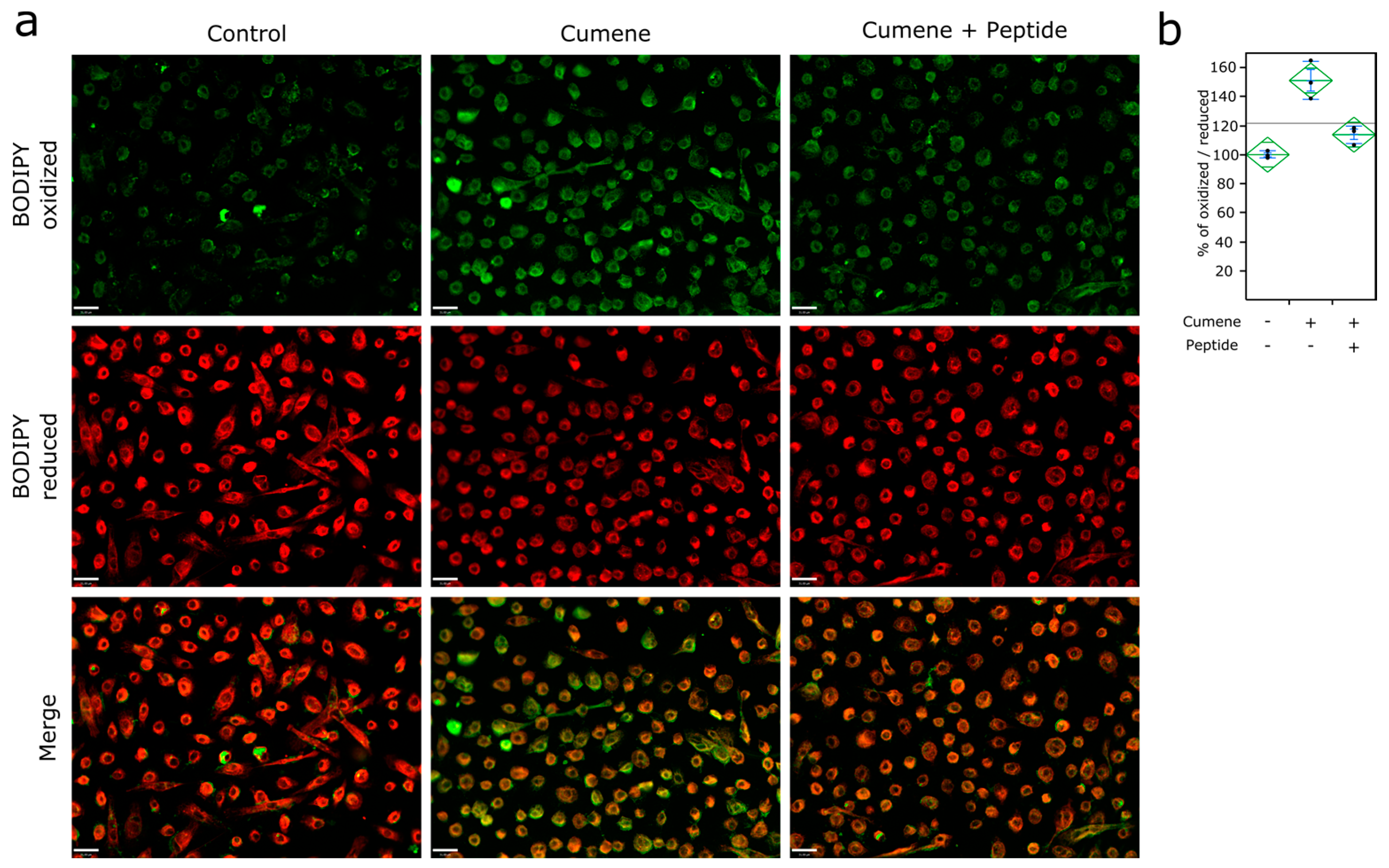
2.4. Lipid Peroxidation Monitoring
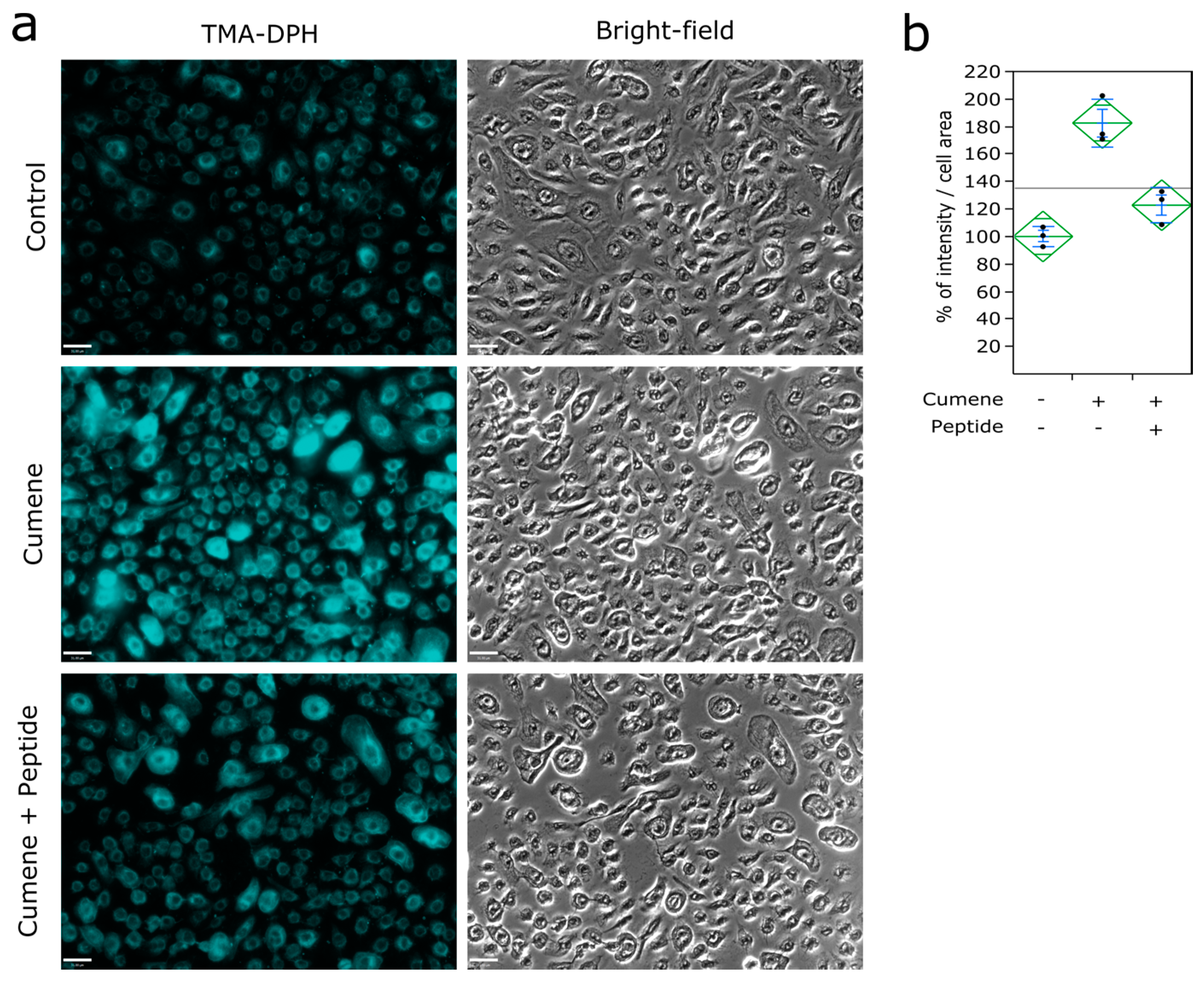
2.5. Labeling of the Plasma Membrane
2.6. HPLC Analysis
2.7. RNA Interference
2.8. Cell Culture
2.9. Reverse Transcription-PCR Assays
2.10. Western Blot Analysis
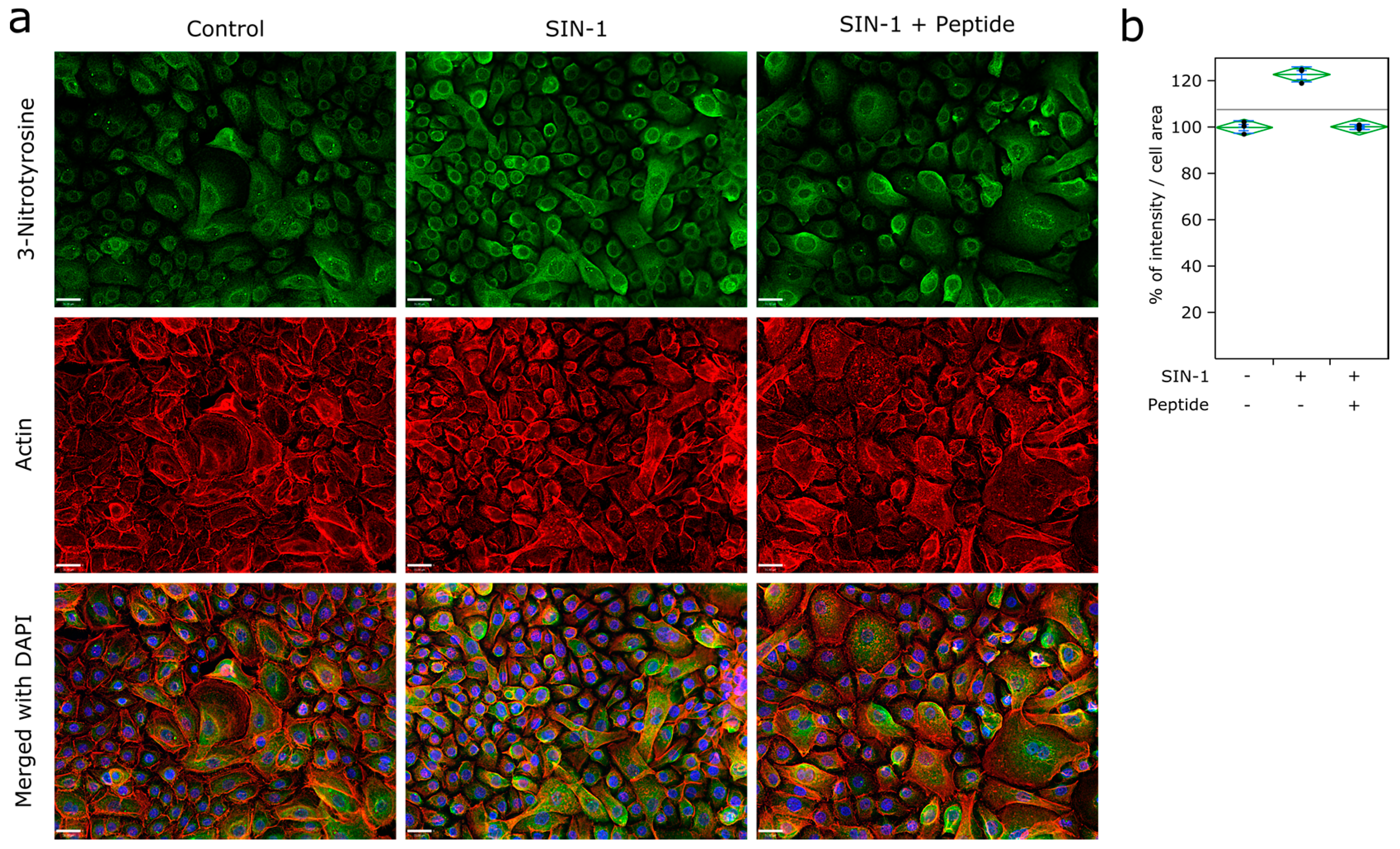
2.11. Immunofluorescence
2.12. Immunohistological Fluorescence
2.13. Electron Microscopy
2.14. Statistical Analysis
3. Results
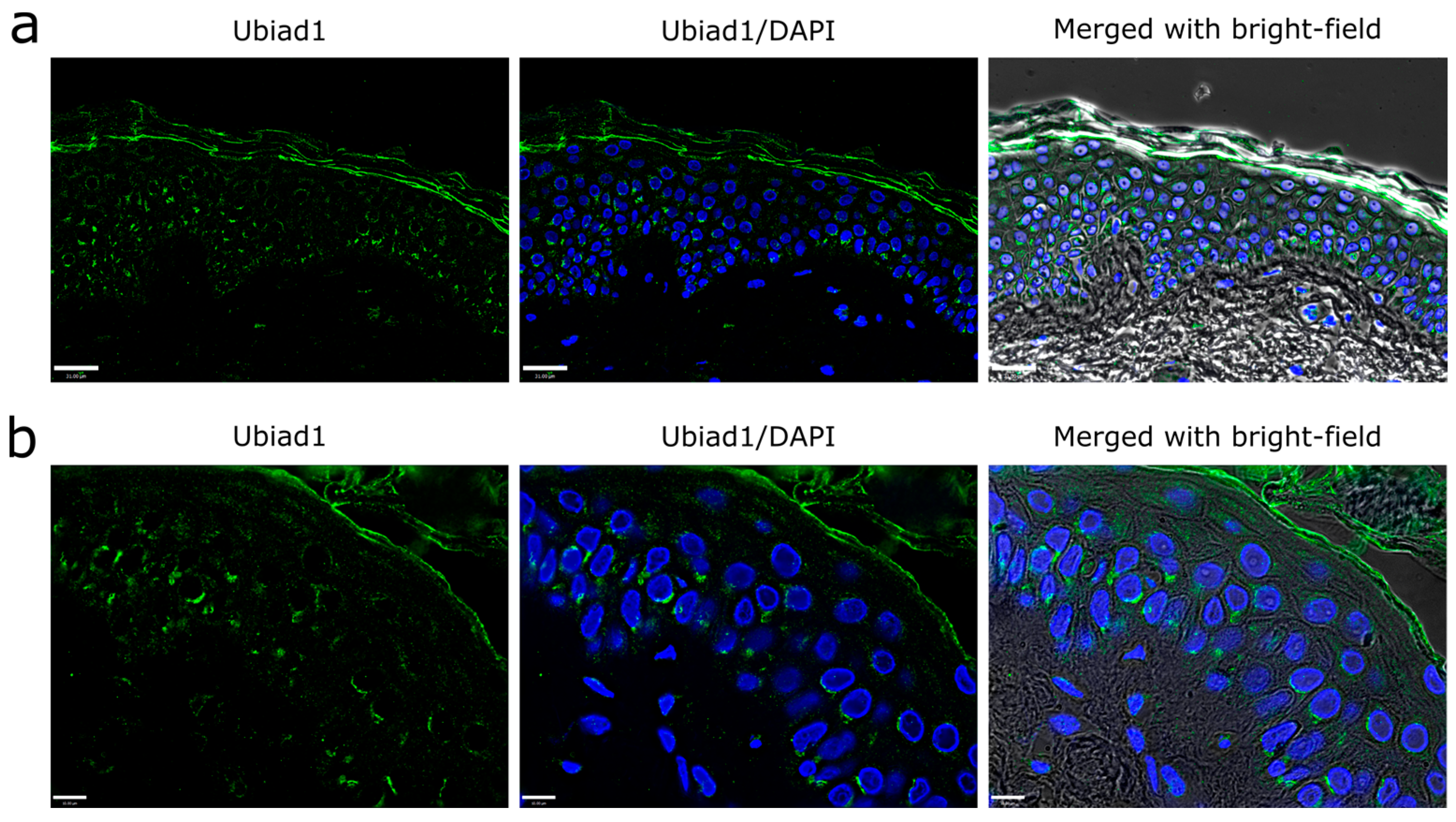
3.1. Ubiad1 Expression and Localization in Human Skin Epidermis
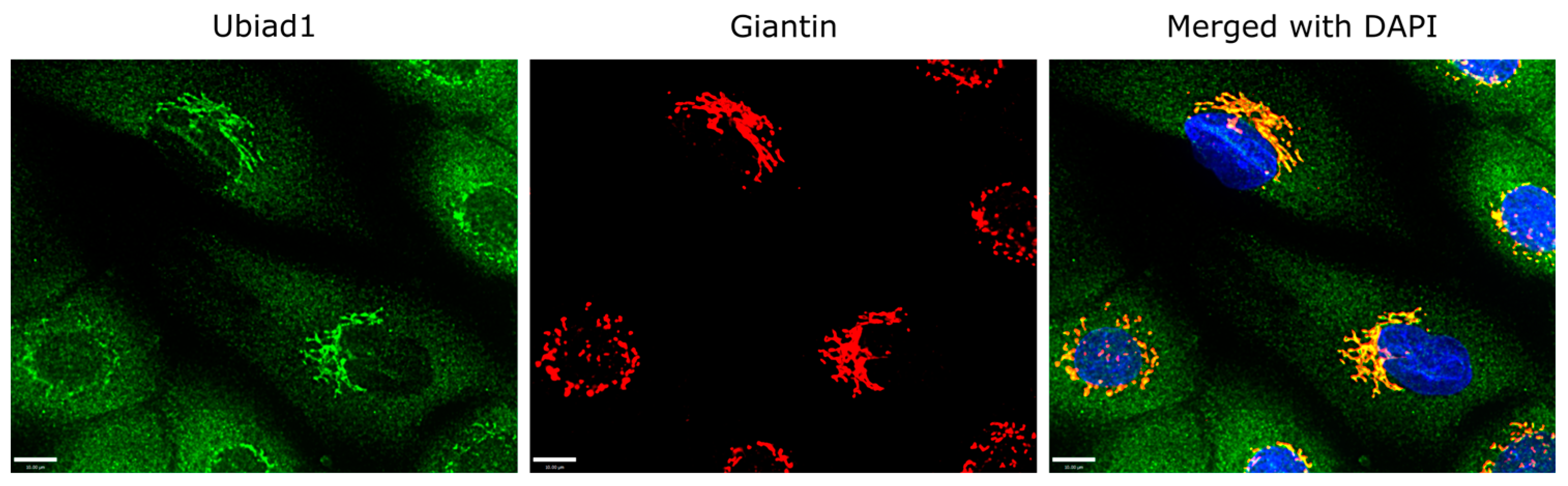
3.2. Ubiad1 Expression and Localization within Keratinocytes In Vitro
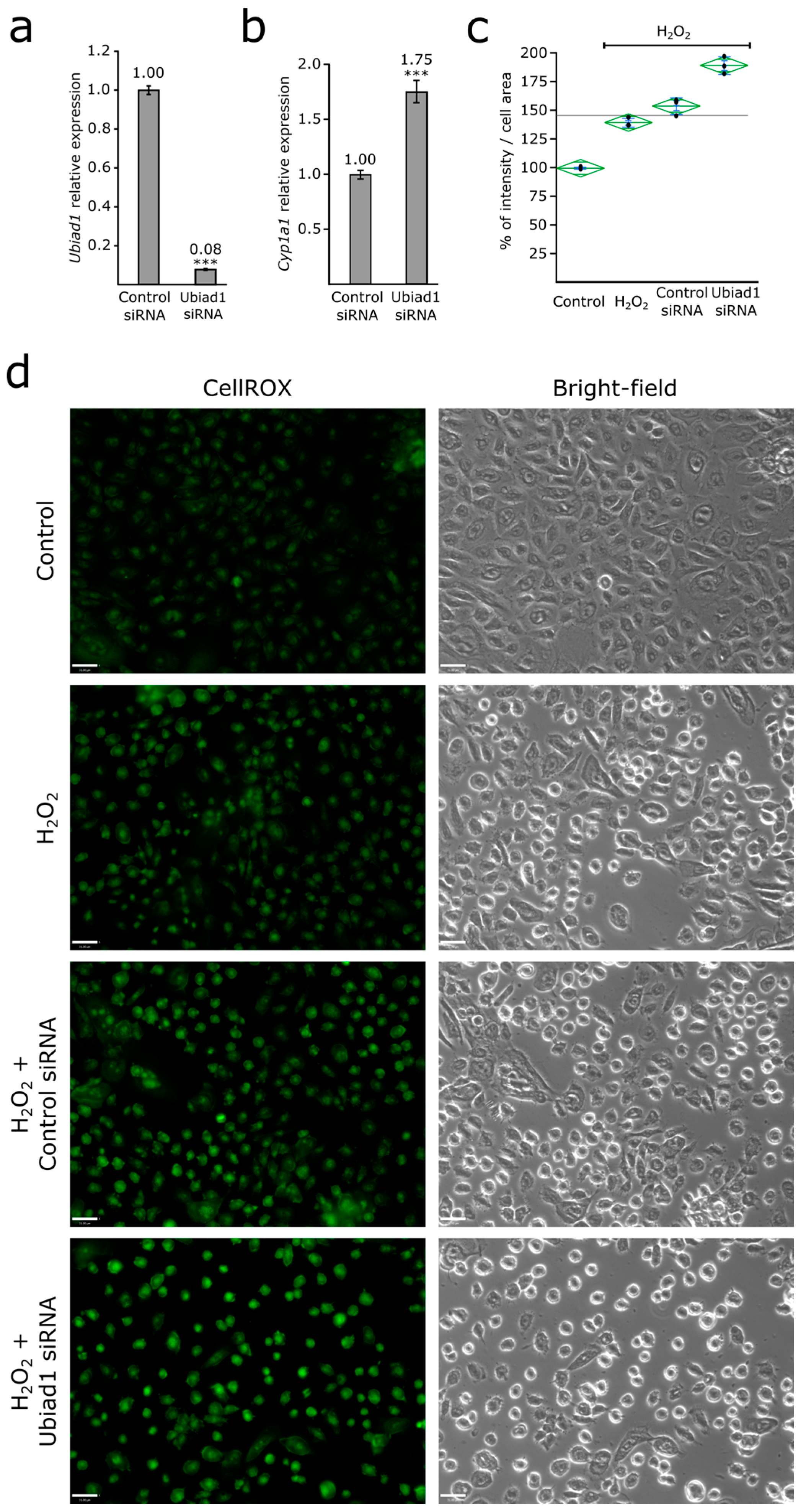
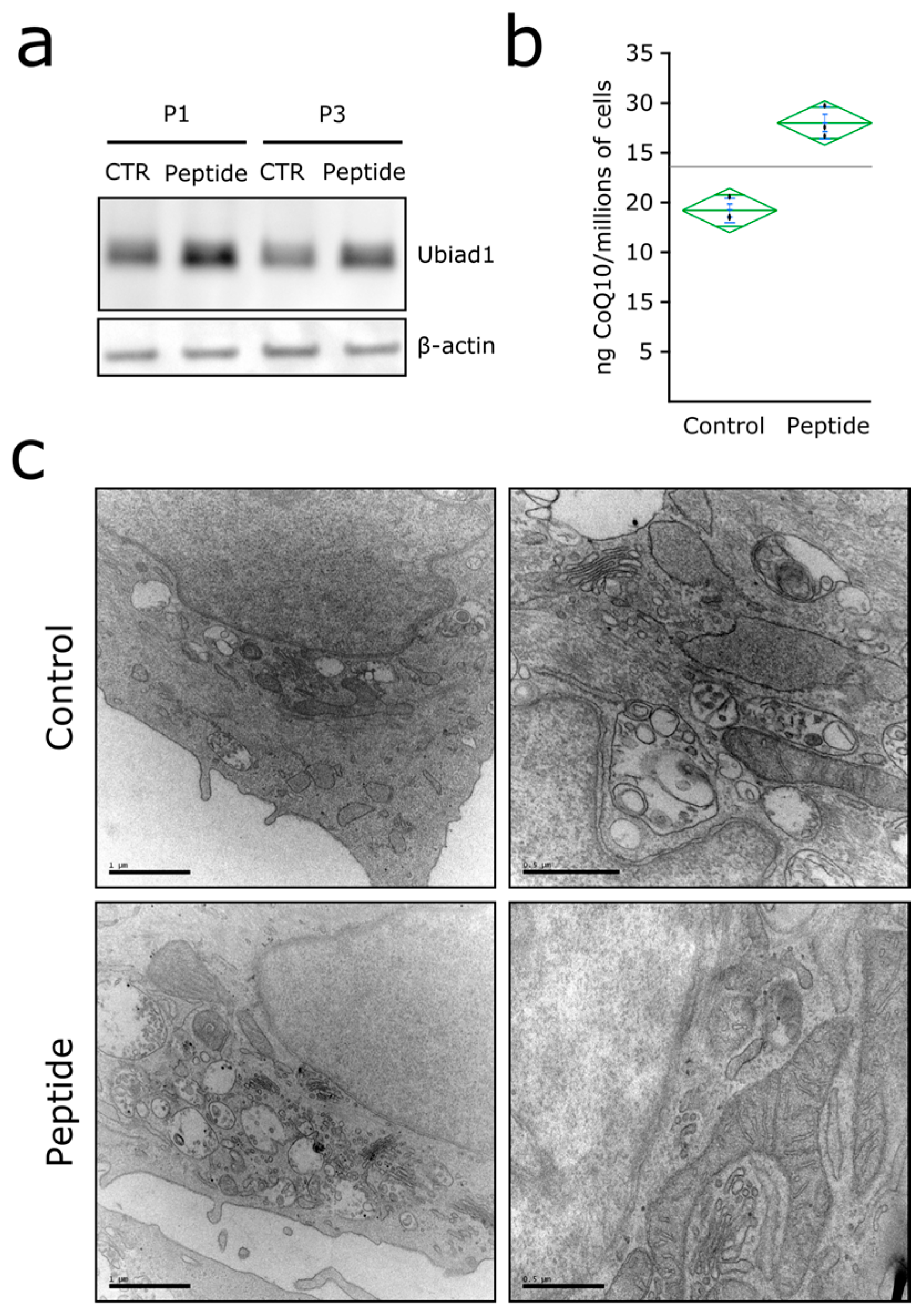
3.3. Ubiad1 Has a Key Protective Role in Keratinocyte Physiology
4. Discussion
5. Conclusions
6. Patents
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Halliwell, B.; Whiteman, M. Measuring reactive species and oxidative damage in vivo and in cell culture: How should you do it and what do the results mean? Br. J. Pharmacol. 2004, 142, 231–255. [Google Scholar] [CrossRef] [PubMed]
- Shindo, Y.; Witt, E.; Han, D.; Epstein, W.; Packer, L. Enzymic and non-enzymic antioxidants in epidermis and dermis of human skin. J. Investig. Dermatol. 1994, 102, 122–124. [Google Scholar] [CrossRef] [PubMed]
- Rinnerthaler, M.; Bischof, J.; Streubel, M.K.; Trost, A.; Richter, K. Oxidative stress in aging human skin. Biomolecules 2015, 5, 545–589. [Google Scholar] [CrossRef] [PubMed]
- Crane, F.L.; Hatefi, Y.; Lester, R.L.; Widmer, C. Isolation of a quinone from beef heart mitochondria. Biochim. Biophys. Acta 1957, 25, 220–221. [Google Scholar] [CrossRef]
- Ernster, L.; Dallner, G. Biochemical, physiological and medical aspects of ubiquinone function. Biochim. Biophys. Acta 1995, 1271, 195–204. [Google Scholar] [CrossRef]
- Hoppe, U.; Bergemann, J.; Diembeck, W.; Ennen, J.; Gohla, S.; Harris, I.; Jacob, J.; Kielholz, J.; Mei, W.; Pollet, D.; et al. Coenzyme Q10, a cutaneous antioxidant and energizer. Biofactors 1999, 9, 371–378. [Google Scholar] [CrossRef] [PubMed]
- Inui, M.; Ooe, M.; Fujii, K.; Matsunaka, H.; Yoshida, M.; Ichihashi, M. Mechanisms of inhibitory effects of CoQ10 on UVB-induced wrinkle formation in vitro and in vivo. Biofactors 2008, 32, 237–243. [Google Scholar] [CrossRef] [PubMed]
- Crane, F.L. Biochemical Functions of Coenzyme Q10. J. Am. Coll. Nutr. 2001, 20, 591–598. [Google Scholar] [CrossRef] [PubMed]
- McGarvey, T.W.; Nguyen, T.; Tomaszewski, J.E.; Monson, F.C.; Malkowicz, S.B. Isolation and characterization of the TERE1 gene, a gene down-regulated in transitional cell carcinoma of the bladder. Oncogene 2001, 20, 1042–1051. [Google Scholar] [CrossRef] [PubMed]
- Heide, L. Prenyl transfer to aromatic substrates: Genetics and enzymology. Curr. Opin. Chem. Biol. 2009, 13, 171–179. [Google Scholar] [CrossRef] [PubMed]
- Nakagawa, K.; Hirota, Y.; Sawada, N.; Yuge, N.; Watanabe, M.; Uchino, Y.; Okuda, N.; Shimomura, Y.; Suhara, Y.; Okano, T. Identification of UBIAD1 as a novel human menaquinone-4 biosynthetic enzyme. Nature 2010, 468, 117–121. [Google Scholar] [CrossRef] [PubMed]
- Bonitz, T.; Alva, V.; Saleh, O.; Lupas, A.N.; Heide, L. Evolutionary relationships of microbial aromatic prenyltransferases. PLoS ONE 2011, 6, e27336. [Google Scholar] [CrossRef] [PubMed]
- Hirota, Y.; Nakagawa, K.; Sawada, N.; Okuda, N.; Suhara, Y.; Uchino, Y.; Kimoto, T.; Funahashi, N.; Kamao, M.; Tsugawa, N.; et al. Functional characterization of the vitamin K biosynthetic enzyme UBIAD1. PLoS ONE 2015, 10, e0125737. [Google Scholar] [CrossRef] [PubMed]
- Mugoni, V.; Medana, C.; Santoro, M.M. 13C-isotope-based protocol for prenyl lipid metabolic analysis in zebrafish embryos. Nat. Protoc. 2013, 8, 2337–2347. [Google Scholar] [CrossRef] [PubMed]
- Nakagawa, K.; Sawada, N.; Hirota, Y.; Uchino, Y.; Suhara, Y.; Hasegawa, T.; Amizuka, N.; Okamoto, T.; Tsugawa, N.; Kamao, M.; et al. Vitamin K2 biosynthetic enzyme, UBIAD1 is essential for embryonic development of mice. PLoS ONE 2014, 9, e104078. [Google Scholar] [CrossRef] [PubMed]
- Mugoni, V.; Postel, R.; Catanzaro, V.; De Luca, E.; Turco, E.; Digilio, G.; Silengo, L.; Murphy, M.P.; Medana, C.; Stainier, D.Y.; et al. Ubiad1 is an antioxidant enzyme that regulates eNOS activity by CoQ10 synthesis. Cell 2013, 152, 504–518. [Google Scholar] [CrossRef] [PubMed]
- Fredericks, W.J.; McGarvey, T.; Wang, H.; Lal, P.; Puthiyaveettil, R.; Tomaszewski, J.; Sepulveda, J.; Labelle, E.; Weiss, J.S.; Nickerson, M.L.; et al. The bladder tumor suppressor protein TERE1 (UBIAD1) modulates cell cholesterol: Implications for tumor progression. DNA Cell. Biol. 2011, 30, 851–864. [Google Scholar] [CrossRef] [PubMed]
- Nickerson, M.L.; Bosley, A.D.; Weiss, J.S.; Kostiha, B.N.; Hirota, Y.; Brandt, W.; Esposito, D.; Kinoshita, S.; Wessjohann, L.; Morham, S.G.; et al. The UBIAD1 prenyltransferase links menaquinone-4 synthesis to cholesterol metabolic enzymes. Hum. Mutat. 2013, 34, 317–329. [Google Scholar] [CrossRef] [PubMed]
- Costes, S.V.; Daelemans, D.; Cho, E.H.; Dobbin, Z.; Pavlakis, G.; Lockett, S. Automatic and quantitative measurement of protein-protein colocalization in live cells. Biophys. J. 2004, 86, 3993–4003. [Google Scholar] [CrossRef] [PubMed]
- Linstedt, A.D.; Hauri, H.P. Giantin, a novel conserved Golgi membrane protein containing a cytoplasmic domain of at least 350 kDa. Mol. Biol. Cell 1993, 4, 679–693. [Google Scholar] [CrossRef] [PubMed]
- Ito, S.; Chen, C.; Satoh, J.; Yim, S.; Gonzalez, F.J. Dietary phytochemicals regulate whole-body CYP1A1 expression through an arylhydrocarbon receptor nuclear translocator-dependent system in gut. J. Clin. Investig. 2007, 117, 1940–1950. [Google Scholar] [CrossRef] [PubMed]
- Schumacher, M.M.; Elsabrouty, R.; Seemann, J.; Jo, Y.; DeBose-Boyd, R.A. The prenyltransferase UBIAD1 is the target of geranylgeraniol in degradation of HMG CoA reductase. eLIFE 2015, 4. [Google Scholar] [CrossRef] [PubMed]
- Glass, D.; Viñuela, A.; Davies, M.N.; Ramasamy, A.; Parts, L.; Knowles, D.; Brown, A.A.; Hedman, A.K.; Small, K.S.; Buil, A.; et al. Gene expression changes with age in skin, adipose tissue, blood and brain. Genome Biol. 2013, 14, R75. [Google Scholar] [CrossRef] [PubMed]
- Menon, G.K.; DalFarra, C.; Botto, J.M.; Domloge, N. Mitochondria: A new focus as an anti-aging target in skin care. J. Cosmet. Dermatol. 2010, 9, 122–131. [Google Scholar] [CrossRef] [PubMed]
- Figueiredo, P.A.; Powers, S.K.; Ferreira, R.M.; Appell, H.J.; Duarte, J.A. Aging impairs skeletal muscle mitochondrial bioenergetic function. J. Gerontol. A Biol. Sci. Med. Sci. 2009, 64, 21–33. [Google Scholar] [CrossRef] [PubMed]
- Elias, P.M.; Feingold, K.R.; Fartasch, M. The Epidermal lamelar body as a multifunctional secretory organelle. In Skin Barrier; Elias, P.M., Feingold, K.R., Eds.; Taylor & Francis Group: New York, NY, USA, 2006; pp. 261–272. ISBN 978-0-8493-6129-6. [Google Scholar]
- Ghadially, R.; Brown, B.E.; Hanley, K.; Reed, J.T.; Feingold, K.R.; Elias, P.M. Decreased epidermal lipid synthesis accounts for altered barrier function in aged mice. J. Investig. Dermatol. 1996, 106, 1064–1069. [Google Scholar] [CrossRef] [PubMed]
- Bentinger, M.; Tekle, M.; Dallner, G. Coenzyme Q-biosynthesis and functions. Biochem. Biophys. Res. Commun. 2010, 396, 74–79. [Google Scholar] [CrossRef] [PubMed]
- Schniertshauer, D.; Müller, S.; Mayr, T.; Sonntag, T.; Gebhard, D.; Bergemann, J. Accelerated Regeneration of ATP Level after Irradiation in Human Skin Fibroblasts by Coenzyme Q10. Photochem. Photobiol. 2016, 92, 488–494. [Google Scholar] [CrossRef] [PubMed]
- Kammeyer, A.; Luiten, R.M. Oxidation events and skin aging. Ageing Res. Rev. 2015, 21, 16–29. [Google Scholar] [CrossRef] [PubMed]
- Littarru, G.P.; Langsjoen, P. Coenzyme Q10 and statins: Biochemical and clinical implications. Mitochondrion 2007, 7, S168–S174. [Google Scholar] [CrossRef] [PubMed]
- Wong-Ekkabut, J.; Xu, Z.; Triampo, W.; Tang, I.M.; Tieleman, D.P.; Monticelli, L. Effect of lipid peroxidation on the properties of lipid bilayers: A molecular dynamics study. Biophys. J. 2007, 93, 4225–4236. [Google Scholar] [CrossRef] [PubMed]
- Jovanović, Z. Effects of Oxidative Stress on the Electrophysiological Function of Neuronal Membranes. In Oxidative Stress and Diseases; Lushchak, V.I., Gospodaryo, D.V., Eds.; InTech Europe: Rijeka, Croatia, 2012; pp. 337–358. ISBN 978-953-51-0552-7. [Google Scholar] [CrossRef]
- Lingappan, K.; Jiang, W.; Wang, L.; Wang, G.; Couroucli, X.I.; Shivanna, B.; Welty, S.E.; Barrios, R.; Khan, M.F.; Nebert, D.W.; et al. Mice deficient in the gene for cytochrome P450 (CYP)1A1 are more susceptible than wild-type to hyperoxic lung injury: Evidence for protective role of CYP1A1 against oxidative stress. Toxicol. Sci. 2014, 141, 68–77. [Google Scholar] [CrossRef] [PubMed]
- Schöpfer, F.; Riobó, N.; Carreras, M.C.; Alvarez, B.; Radi, R.; Boveris, A.; Cadenas, E.; Poderoso, J.J. Oxidation of ubiquinol by peroxynitrite: Implications for protection of mitochondria against nitrosative damage. Biochem. J. 2000, 349, 35–42. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Wang, L.; Jacoby, A.M.; Jasinski, K.; Kubant, R.; Malinski, T. Ultraviolet B light-induced nitric oxide/peroxynitrite imbalance in keratinocytes—Implications for apoptosis and necrosis. Photochem. Photobiol. 2010, 86, 389–396. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Labarrade, F.; Menon, G.; Labourasse, L.; Gondran, C.; Cucumel, K.; Domloge, N. Significance of Ubiad1 for Epidermal Keratinocytes Involves More Than CoQ10 Synthesis: Implications for Skin Aging. Cosmetics 2018, 5, 9. https://doi.org/10.3390/cosmetics5010009
Labarrade F, Menon G, Labourasse L, Gondran C, Cucumel K, Domloge N. Significance of Ubiad1 for Epidermal Keratinocytes Involves More Than CoQ10 Synthesis: Implications for Skin Aging. Cosmetics. 2018; 5(1):9. https://doi.org/10.3390/cosmetics5010009
Chicago/Turabian StyleLabarrade, Florian, Gopinathan Menon, Laura Labourasse, Catherine Gondran, Karine Cucumel, and Nouha Domloge. 2018. "Significance of Ubiad1 for Epidermal Keratinocytes Involves More Than CoQ10 Synthesis: Implications for Skin Aging" Cosmetics 5, no. 1: 9. https://doi.org/10.3390/cosmetics5010009
APA StyleLabarrade, F., Menon, G., Labourasse, L., Gondran, C., Cucumel, K., & Domloge, N. (2018). Significance of Ubiad1 for Epidermal Keratinocytes Involves More Than CoQ10 Synthesis: Implications for Skin Aging. Cosmetics, 5(1), 9. https://doi.org/10.3390/cosmetics5010009




