Termite Societies Promote the Taxonomic and Functional Diversity of Archaeal Communities in Mound Soils
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Site and Sample Collection
2.2. Soil Physical and Chemical Parameters
2.3. DNA Extraction
2.4. Metagenomic DNA Sequencing
2.5. Metagenomic Data Analysis
2.6. Statistical Analysis
3. Results
3.1. Physical and Chemical Characterization of Termite Mound Soils and Their Surrounding Soils
3.2. Sequencing Data Analyses of the Samples Collected from Termite Mounds and the Surrounding Soils
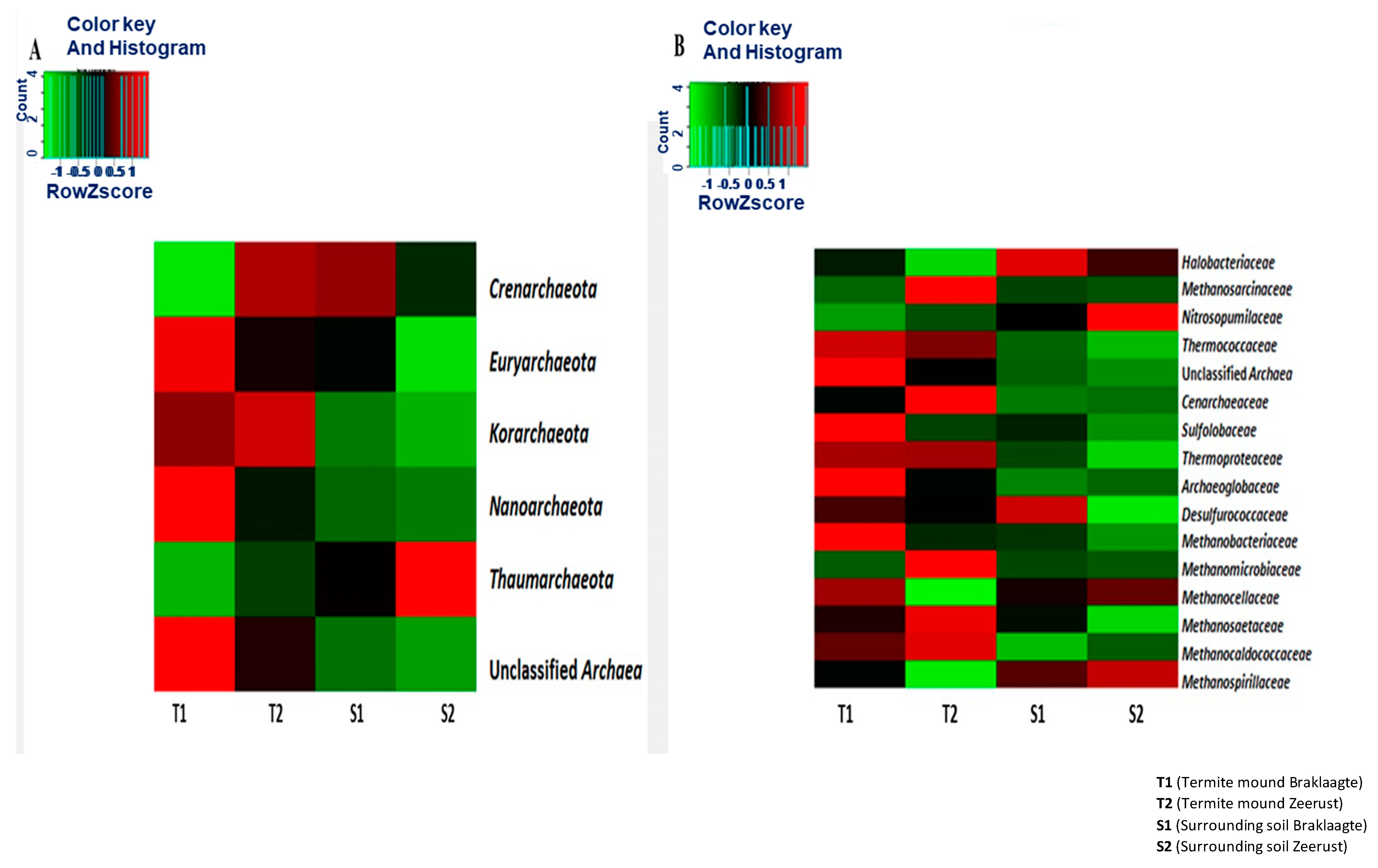
3.3. Structural Composition Analysis of the Archaeal Community
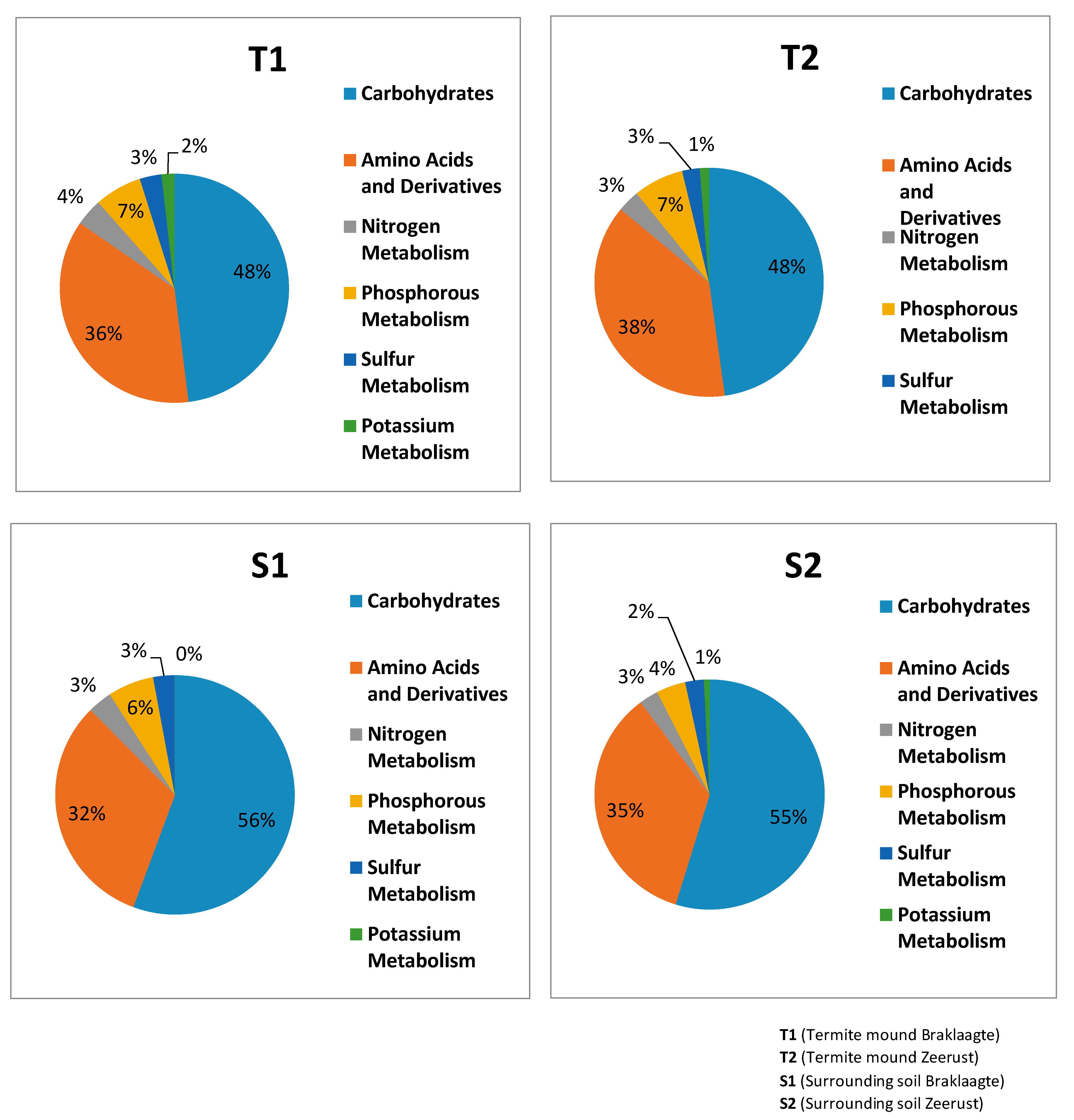
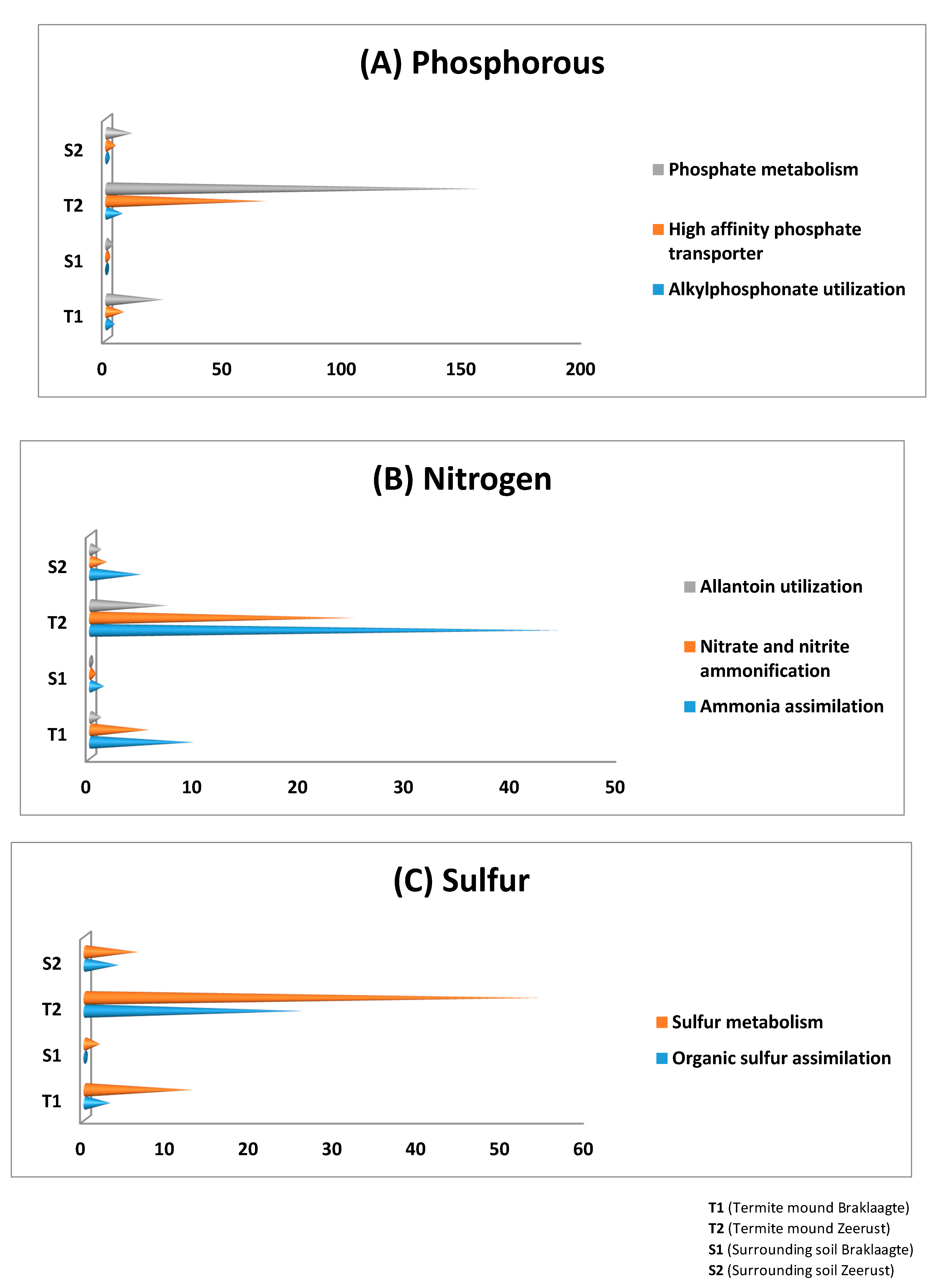
3.4. General Metabolism Categories
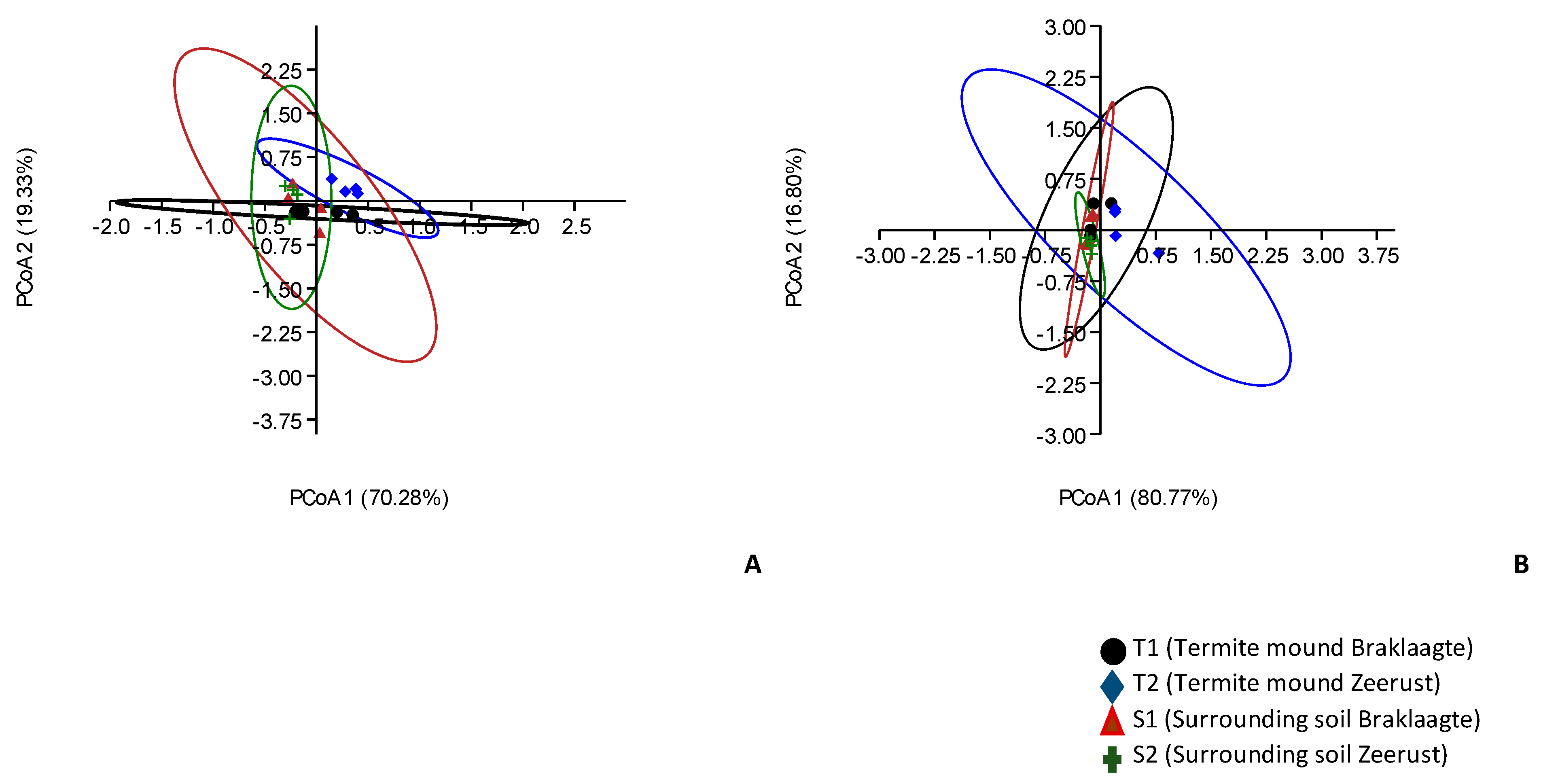
3.5. Alpha and Beta Diversity Estimations for Archaeal Communities from Termite Mounds and Surrounding Soils
3.6. Influence of Environmental Factors on Archaeal Community
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Availability of Data and Materials
References
- Bhardwaj, N.; Tomar, S. Indigenous knowledge of termite (Odontotermes obesus) control: A Review. Int. J. Adv. Sci. Res. Manag. 2018, 1, 124–127. [Google Scholar]
- Scharf, M.E. Termites as Targets and Models for Biotechnology. Annu. Rev. Entomol. 2015, 60, 77–102. [Google Scholar] [CrossRef]
- Enagbonma, B.J.; Babalola, O.O. Environmental sustainability: A review of termite mound soil material and its bacteria. Sustainability 2019, 11, 3847. [Google Scholar] [CrossRef]
- Jouquet, P.; Guilleux, N.; Caner, L.; Chintakunta, S.; Ameline, M.; Shanbhag, R. Influence of soil pedological properties on termite mound stability. Geoderma 2016, 262, 45–51. [Google Scholar] [CrossRef]
- Friedrich, M.W.; Schmitt-Wagner, D.; Lueders, T.; Brune, A. Axial Differences in Community Structure ofCrenarchaeota and Euryarchaeota in the Highly Compartmentalized Gut of the Soil-Feeding TermiteCubitermes orthognathus. Appl. Environ. Microbiol. 2001, 67, 4880–4890. [Google Scholar] [CrossRef] [PubMed]
- Menichetti, L.; Landi, L.; Nannipieri, P.; Katterer, T.; Kirchmann, H.; Renella, G. Chemical properties and biochemical activity of colonized and abandoned litter-feeding termite (Macrotermes spp.) mounds in chromic Cambisol Area on the Borana Plateau, Ethiopia. Pedosphere 2014, 24, 399–407. [Google Scholar] [CrossRef]
- Jouquet, P.; Bottinelli, N.; Shanbhag, R.R.; Bourguignon, T.; Traoré, S.; Abbasi, S.A. Termites: The neglected soil engineers of tropical soils. Soil Sci. 2016, 181, 157–165. [Google Scholar] [CrossRef]
- Dhembare, A.; Pokale, A. Physico-chemical properties of termite mound soil. Arch. Appl. Sci. Res. 2013, 5, 123–126. [Google Scholar]
- Deke, A.L.; Adugna, W.T.; Fite, A.T. Soil physic-chemical properties in termite mounds and adjacent control soil in Miyo and Yabello Districts of Borana Zone, Southern Ethiopia. Am. J. Agric. For. 2016, 4, 69–74. [Google Scholar]
- Jouquet, P.; Traoré, S.; Choosai, C.; Hartmann, C.; Bignell, D. Influence of termites on ecosystem functioning. Ecosystem services provided by termites. Eur. J. Soil Biol. 2011, 47, 215–222. [Google Scholar] [CrossRef]
- Makonde, H.M.; Mwirichia, R.; Osiemo, Z.; Boga, H.I.; Klenk, H.P. 454 Pyrosequencing-based assessment of bacterial diversity and community structure in termite guts, mounds and surrounding soils. SpringerPlus 2015, 4, 471. [Google Scholar] [CrossRef] [PubMed]
- Makonde, H.M.; Mwirichia, R.; Muwawa, E.M.; Klenk, H.-P.; Boga, H.I. Fungal diversity and community structure in gut, mound and surrounding soil of fungus-cultivating termites. Afr. J. Microbiol. Res. 2017, 11, 504–515. [Google Scholar]
- Fall, S.; Hamelin, J.; Ndiaye, F.; Assigbetse, K.; Aragno, M.; Chotte, J.L.; Brauman, A. Differences between bacterial communities in the gut of a soil-feeding termite (Cubitermes niokoloensis) and its mounds. Appl. Environ. Microbiol. 2007, 73, 5199–5208. [Google Scholar] [CrossRef]
- Manjula, A.; Pushpanathan, M.; Sathyavathi, S.; Gunasekaran, P.; Rajendhran, J. Comparative analysis of microbial diversity in termite gut and termite nest using ion sequencing. Curr. Microbiol. 2016, 72, 267–275. [Google Scholar] [CrossRef]
- Ibarbalz, F.M.; Pérez, M.V.; Figuerola, E.L.; Erijman, L. The bias associated with amplicon sequencing does not affect the quantitative assessment of bacterial community dynamics. PLoS ONE 2014, 9, e99722. [Google Scholar] [CrossRef]
- Lan, Y.; Rosen, G.; Hershberg, R. Marker genes that are less conserved in their sequences are useful for predicting genome-wide similarity levels between closely related prokaryotic strains. Microbiome 2016, 4, 18. [Google Scholar] [CrossRef]
- Enagbonma, B.J.; Babalola, O.O. Unveiling plant-beneficial function as seen in bacteria genes from termite mound soil. J. Soil Sci. Plant Nutr. 2020, 20, 421–430. [Google Scholar] [CrossRef]
- Lima, S.S.d.; Pereira, M.G.; Santos, G.L.d.; Pontes, R.d.M.; Diniz, A.R. Soil Physical and Chemical Properties in Epigeal Termite Mounds in Pastures. Floresta E Ambiente 2018, 25. [Google Scholar] [CrossRef]
- Enagbonma, B.J.; Aremu, B.R.; Babalola, O.O. Profiling the functional diversity of termite mound soil bacteria as revealed by shotgun sequencing. Genes 2019, 10, 637. [Google Scholar] [CrossRef]
- Shi, J.-Y.; Yuan, X.-F.; Lin, H.-R.; Yang, Y.-Q.; Li, Z.-Y. Differences in soil properties and bacterial communities between the rhizosphere and bulk soil and among different production areas of the medicinal plant Fritillaria thunbergii. Int. J. Mol. Sci. 2011, 12, 3770–3785. [Google Scholar] [CrossRef]
- Meyer, F.; Paarmann, D.; D’Souza, M.; Olson, R.; Glass, E.M.; Kubal, M.; Paczian, T.; Rodriguez, A.; Stevens, R.; Wilke, A. The metagenomics RAST server—A public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinform. 2008, 9, 386. [Google Scholar] [CrossRef] [PubMed]
- Kent, W.J. BLAT—The BLAST-like alignment tool. Genome Res. 2002, 12, 656–664. [Google Scholar] [CrossRef] [PubMed]
- Wilke, A.; Harrison, T.; Wilkening, J.; Field, D.; Glass, E.M.; Kyrpides, N.; Mavrommatis, K.; Meyer, F. The M5nr: A novel non-redundant database containing protein sequences and annotations from multiple sources and associated tools. BMC Bioinform. 2012, 13, 141. [Google Scholar] [CrossRef]
- Hammer, Ř.; Harper, D.; Ryan, P. PAST: Paleontological statistics software package for education and data analysis–palaeontol. Electron 2001, 4, 9–15. [Google Scholar]
- Clarke, K.; Green, R. Statistical design and analysis for a “biological effects” study. Mar. Ecol. Prog. Ser. 1988, 46, 213–226. [Google Scholar] [CrossRef]
- Khomtchouk, B.B.; Hennessy, J.R.; Wahlestedt, C. Shinyheatmap: Ultra fast low memory heatmap web interface for big data genomics. PLoS ONE 2017, 12, e0176334. [Google Scholar] [CrossRef]
- Castañeda, L.E.; Barbosa, O. Metagenomic analysis exploring taxonomic and functional diversity of soil microbial communities in Chilean vineyards and surrounding native forests. PeerJ 2017, 5, e3098. [Google Scholar] [CrossRef]
- Ranjan, R.; Rani, A.; Metwally, A.; McGee, H.S.; Perkins, D.L. Analysis of the microbiome: Advantages of whole genome shotgun versus 16S amplicon sequencing. Biochem. Biophys. Res. Commun. 2016, 469, 967–977. [Google Scholar] [CrossRef]
- Enagbonma, B.J.; Amoo, A.E.; Babalola, O.O. Deciphering the microbiota data from termite mound soil in South Africa using shotgun metagenomics. Data Brief 2020, 28, 104802. [Google Scholar] [CrossRef]
- Dinsdale, E.A.; Edwards, R.A.; Hall, D.; Angly, F.; Breitbart, M.; Brulc, J.M.; Furlan, M.; Desnues, C.; Haynes, M.; Li, L. Functional metagenomic profiling of nine biomes. Nature 2008, 452, 629–632. [Google Scholar] [CrossRef]
- Li, Y.; Zhu, X.; Zhang, W.; Zhu, D.; Zhou, X.; Zhang, L. Archaeal communities in the deep-sea sediments of the South China Sea revealed by Illumina high-throughput sequencing. Ann. Microbiol. 2019, 69, 839–848. [Google Scholar] [CrossRef]
- Tripathi, B.M.; Kim, M.; Lai-Hoe, A.; Shukor, N.A.; Rahim, R.A.; Go, R.; Adams, J.M. pH dominates variation in tropical soil archaeal diversity and community structure. FEMS Microbiol. Ecol. 2013, 86, 303–311. [Google Scholar] [CrossRef] [PubMed]
- Donovan, S.E.; Purdy, K.J.; Kane, M.D.; Eggleton, P. Comparison of Euryarchaea Strains in the Guts and Food-Soil of the Soil-Feeding Termite Cubitermes fungifaber across Different Soil Types. Appl. Environ. Microbiol. 2004, 70, 3884–3892. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Q.; Hu, Y.; Zhang, S.; Noll, L.; Böckle, T.; Dietrich, M.; Herbold, C.W.; Eichorst, S.A.; Woebken, D.; Richter, A.; et al. Soil multifunctionality is affected by the soil environment and by microbial community composition and diversity. Soil Biol. Biochem. 2019, 136, 107521. [Google Scholar] [CrossRef] [PubMed]
- Faoro, H.; Alves, A.; Souza, E.; Rigo, L.; Cruz, L.; Al-Janabi, S.; Monteiro, R.; Baura, V.; Pedrosa, F. Influence of soil characteristics on the diversity of bacteria in the Southern Brazilian Atlantic Forest. Appl. Environ. Microbiol. 2010, 76, 4744–4749. [Google Scholar] [CrossRef] [PubMed]

| Soil Property | T1 | T2 | S1 | S2 |
|---|---|---|---|---|
| pH | 5.10 ± 0.33 a | 4.48 ± 0.46 a | 5.80 ± 0.32 b | 5.38 ± 0.39 c |
| N (%) | 0.09 ± 0.03 a | 0.10 ± 0.03 b | 0.59 ± 0.47 c | 0.25 ± 0.04 d |
| P (mg/L) | 0.25 ± 0.50 a | 0.75 ± 0.50 a | 0.00 ± 0.00 a | 0.00 ± 0.00 a |
| K (mg/L) | 393.50 ± 120.33 a | 427.50 ± 57.93 a | 216.75 ± 48.40 b | 184.50 ± 27.72 c |
| Ca (mg/L) | 1879.50 ± 587.38 a | 2237.75 ± 318.91 a | 1493.50 ± 456.59 a | 1108.50 ± 160.48 b |
| Mg (mg/L) | 575.00 ± 262.32 a | 622.25 ± 60.84 a | 349.75 ± 159.70 a | 330.25 ± 138.75 a |
| OC (%) | 0.31 ± 0.42 a | 0.10 ± 0.00 a | 0.11 ± 0.0 a | 0.11 ± 0.01 a |
| Sand (%) | 65.00 ± 8.29 a | 47.75 ± 23.60 b | 72.00 ± 17.66 c | 76.50 ± 3.00 d |
| Silt (%) | 9.00 ± 2.94 a | 19.75 ± 11.38 a | 11.75 ± 12.87 a | 10.25 ± 0.96 a |
| Clay (%) | 26.00 ± 6.27 a | 33.25 ± 13.52 a | 16.25 ± 4.50 b | 13.25 ± 3.20 c |
| T1 | T2 | S1 | S2 | p-Value | ||
|---|---|---|---|---|---|---|
| Taxonomic | Shannon_H | 2.69 ± 0.17 | 2.67 ± 0.18 | 2.68 ± 0.16 | 2.66 ± 0.18 | 0.99 |
| composition | Evenness_e^H/S | 0.87 ± 0.07 | 0.85 ± 0.07 | 0.86 ± 0.06 | 0.84 ± 0.07 | 0.99 |
| Functional | Shannon_H | 2.88 ± 0.13 | 2.87 ± 0.01 | 2.79 ± 1.57 | 2.39 ± 1.18 | 0.20 |
| category | Evenness_e^H/S | 0.64 ± 0.05 | 0.61 ± 0.02 | 0.63 ± 0.37 | 0.40 ± 0.28 | 0.20 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wakung’oli, M.; Amoo, A.E.; Enagbonma, B.J.; Babalola, O.O. Termite Societies Promote the Taxonomic and Functional Diversity of Archaeal Communities in Mound Soils. Biology 2020, 9, 136. https://doi.org/10.3390/biology9060136
Wakung’oli M, Amoo AE, Enagbonma BJ, Babalola OO. Termite Societies Promote the Taxonomic and Functional Diversity of Archaeal Communities in Mound Soils. Biology. 2020; 9(6):136. https://doi.org/10.3390/biology9060136
Chicago/Turabian StyleWakung’oli, Monde, Adenike Eunice Amoo, Ben Jesuorsemwen Enagbonma, and Olubukola Oluranti Babalola. 2020. "Termite Societies Promote the Taxonomic and Functional Diversity of Archaeal Communities in Mound Soils" Biology 9, no. 6: 136. https://doi.org/10.3390/biology9060136
APA StyleWakung’oli, M., Amoo, A. E., Enagbonma, B. J., & Babalola, O. O. (2020). Termite Societies Promote the Taxonomic and Functional Diversity of Archaeal Communities in Mound Soils. Biology, 9(6), 136. https://doi.org/10.3390/biology9060136






