Mutational Analysis Reveals Functional Roles of METTL16 Domains and Residues
Simple Summary
Abstract
1. Introduction
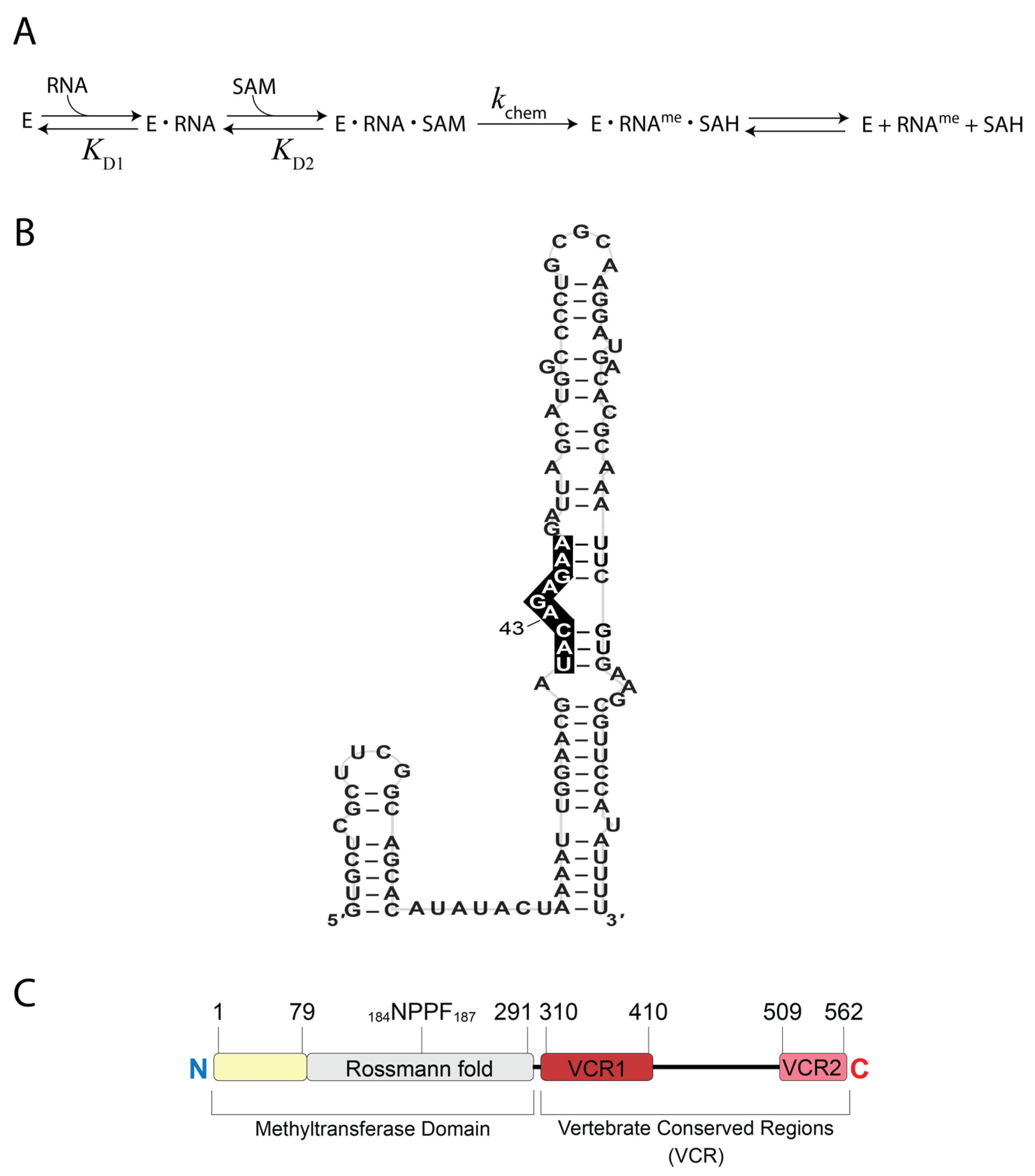
2. Materials and Methods
2.1. Expression and Purification of Wild Type and Mutant METTL16
2.2. Electrophoretic Mobility Shift Assays (EMSAs)
2.3. Circular Dichroism (CD) Spectroscopy
2.4. Methyltransferase Assays
3. Results
3.1. Arginine-Rich Region Is Critical for METTL16 to Bind to U6 snRNA
3.2. SAM Binding Improves with Small, Neutral Side Chains in K-Loop
3.3. Stabilizing Adenosyl Moiety in Binding Pocket Is Critical to SAM Binding
3.4. Mutations to Catalytic Core Greatly Reduce Activity of METTL16
3.5. Catalytic Activity of METTL16 Cancer-Associated Mutants Varies from Innocuous to Inactive
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CD | Circular dichroism |
| EMSA | Electrophoretic mobility shift assay |
| FL | Full-length |
| hp | Hairpin |
| hps | Hairpins |
| kchem | Rate constant for methylation |
| KD | Equilibrium dissociation constant |
| m6A | N6-methyladenosine |
| MAT2A | Methionine adenosyltransferase 2A |
| METTL | Methyltransferase-like protein |
| MTAP | Methylthioadenosine phosphorylase |
| PDB ID | Protein Data Bank identifier |
| RNP | Ribonucleoprotein |
| SAM | S-adenosylmethionine |
| SAH | S-adenosylhomocysteine |
| snRNA | Small nuclear RNA |
| Soga | Suppressor of glucose autophagy |
| SSB | Small RNA binding exonuclease protection factor La |
| TLC | Thin layer chromatography |
| VCR | Vertebrate conserved region |
References
- Xiao, W.; Adhikari, S.; Dahal, U.; Chen, Y.-S.; Hao, Y.-J.; Sun, B.-F.; Sun, H.-Y.; Li, A.; Ping, X.-L.; Lai, W.-Y.; et al. Nuclear m(6)A Reader YTHDC1 Regulates MRNA Splicing. Mol. Cell 2016, 61, 507–519. [Google Scholar] [CrossRef] [PubMed]
- Zhou, K.I.; Shi, H.; Lyu, R.; Wylder, A.C.; Matuszek, Ż.; Pan, J.N.; He, C.; Parisien, M.; Pan, T. Regulation of Co-Transcriptional Pre-MRNA Splicing by m6A through the Low-Complexity Protein HnRNPG. Mol. Cell 2019, 76, 70–81.e9. [Google Scholar] [CrossRef] [PubMed]
- Ma, C.; Chang, M.; Lv, H.; Zhang, Z.-W.; Zhang, W.; He, X.; Wu, G.; Zhao, S.; Zhang, Y.; Wang, D.; et al. RNA m6A Methylation Participates in Regulation of Postnatal Development of the Mouse Cerebellum. Genome Biol. 2018, 19, 68. [Google Scholar] [CrossRef] [PubMed]
- Roundtree, I.A.; Luo, G.-Z.; Zhang, Z.; Wang, X.; Zhou, T.; Cui, Y.; Sha, J.; Huang, X.; Guerrero, L.; Xie, P.; et al. YTHDC1 Mediates Nuclear Export of N6-Methyladenosine Methylated MRNAs. eLife 2017, 6, e31311. [Google Scholar] [CrossRef]
- Hsu, P.J.; Shi, H.; Zhu, A.C.; Lu, Z.; Miller, N.; Edens, B.M.; Ma, Y.C.; He, C. The RNA-Binding Protein FMRP Facilitates the Nuclear Export of N6-Methyladenosine-Containing MRNAs. J. Biol. Chem. 2019, 294, 19889–19895. [Google Scholar] [CrossRef]
- Wang, X.; Zhao, B.S.; Roundtree, I.A.; Lu, Z.; Han, D.; Ma, H.; Weng, X.; Chen, K.; Shi, H.; He, C. N6-Methyladenosine Modulates Messenger RNA Translation Efficiency. Cell 2015, 161, 1388–1399. [Google Scholar] [CrossRef]
- Zhou, K.I.; Parisien, M.; Dai, Q.; Liu, N.; Diatchenko, L.; Sachleben, J.R.; Pan, T. N6-Methyladenosine Modification in a Long Noncoding RNA Hairpin Predisposes Its Conformation to Protein Binding. J. Mol. Biol. 2016, 428, 822–833. [Google Scholar] [CrossRef]
- Meyer, K.D.; Patil, D.P.; Zhou, J.; Zinoviev, A.; Skabkin, M.A.; Elemento, O.; Pestova, T.V.; Qian, S.-B.; Jaffrey, S.R. 5′ UTR m6A Promotes Cap-Independent Translation. Cell 2015, 163, 999–1010. [Google Scholar] [CrossRef]
- Poh, H.X.; Mirza, A.H.; Pickering, B.F.; Jaffrey, S.R. Alternative Splicing of METTL3 Explains Apparently METTL3-Independent m6A Modifications in MRNA. PLoS Biol. 2022, 20, e3001683. [Google Scholar] [CrossRef]
- Flaherty, J.N.; Sivasudhan, E.; Tegowski, M.; Xing, Z.; McGinnis, M.M.; Hunter, O.V.; Featherston, K.M.; Sethia, K.; Tu, B.P.; Meyer, K.D.; et al. The Catalytic Efficiency of METTL16 Affects Cellular Processes by Governing the Intracellular S-Adenosylmethionine Setpoint. Cell Rep. 2025, 44, 115966. [Google Scholar] [CrossRef]
- Pendleton, K.E.; Chen, B.; Liu, K.; Hunter, O.V.; Xie, Y.; Tu, B.P.; Conrad, N.K. The U6 SnRNA m6A Methyltransferase METTL16 Regulates SAM Synthetase Intron Retention. Cell 2017, 169, 824–835.e14. [Google Scholar] [CrossRef]
- Mendel, M.; Chen, K.-M.; Homolka, D.; Gos, P.; Pandey, R.R.; McCarthy, A.A.; Pillai, R.S. Methylation of Structured RNA by the m6A Writer METTL16 Is Essential for Mouse Embryonic Development. Mol. Cell 2018, 71, 986–1000.e11. [Google Scholar] [CrossRef]
- Yoshinaga, M.; Han, K.; Morgens, D.W.; Horii, T.; Kobayashi, R.; Tsuruyama, T.; Hia, F.; Yasukura, S.; Kajiya, A.; Cai, T.; et al. The N6-Methyladenosine Methyltransferase METTL16 Enables Erythropoiesis through Safeguarding Genome Integrity. Nat. Commun. 2022, 13, 6435. [Google Scholar] [CrossRef]
- Breger, K.; Brown, J.A. Elucidating the Kinetic Mechanism of Human METTL16. Biochemistry 2023, 62, 494–506. [Google Scholar] [CrossRef] [PubMed]
- Aoyama, T.; Yamashita, S.; Tomita, K. Mechanistic Insights into m6A Modification of U6 SnRNA by Human METTL16. Nucleic Acids Res. 2020, 48, 5157–5168. [Google Scholar] [CrossRef] [PubMed]
- Yu, D.; Kaur, G.; Blumenthal, R.M.; Zhang, X.; Cheng, X. Enzymatic Characterization of Three Human RNA Adenosine Methyltransferases Reveals Diverse Substrate Affinities and Reaction Optima. J. Biol. Chem. 2021, 296, 100270. [Google Scholar] [CrossRef]
- Doxtader, K.A.; Wang, P.; Scarborough, A.M.; Seo, D.; Conrad, N.K.; Nam, Y. Structural Basis for Regulation of METTL16, an S-Adenosylmethionine Homeostasis Factor. Mol. Cell 2018, 71, 1001–1011.e4. [Google Scholar] [CrossRef]
- Shima, H.; Matsumoto, M.; Ishigami, Y.; Ebina, M.; Muto, A.; Sato, Y.; Kumagai, S.; Ochiai, K.; Suzuki, T.; Igarashi, K. S-Adenosylmethionine Synthesis Is Regulated by Selective N6-Adenosine Methylation and MRNA Degradation Involving METTL16 and YTHDC1. Cell Rep. 2017, 21, 3354–3363. [Google Scholar] [CrossRef]
- Hunter, O.V.; Ruiz, J.C.; Flaherty, J.N.; Conrad, N.K. Functional Analysis of 3′-UTR Hairpins Supports a Two-Tiered Model for Posttranscriptional Regulation of MAT2A by METTL16. RNA 2023, 29, 1725–1737. [Google Scholar] [CrossRef]
- Shen, A.; Hencel, K.; Parker, M.T.; Scott, R.; Skukan, R.; Adesina, A.S.; Metheringham, C.L.; Miska, E.A.; Nam, Y.; Haerty, W.; et al. U6 SnRNA m6A Modification Is Required for Accurate and Efficient Splicing of C. Elegans and Human Pre-MRNAs. Nucleic Acids Res. 2024, 52, 9139–9160. [Google Scholar] [CrossRef]
- Ruszkowska, A.; Ruszkowski, M.; Dauter, Z.; Brown, J.A. Structural Insights into the RNA Methyltransferase Domain of METTL16. Sci. Rep. 2018, 8, 5311. [Google Scholar] [CrossRef]
- Necci, M.; Piovesan, D.; Dosztányi, Z.; Tosatto, S.C.E. MobiDB-Lite: Fast and Highly Specific Consensus Prediction of Intrinsic Disorder in Proteins. Bioinformatics 2017, 33, 1402–1404. [Google Scholar] [CrossRef]
- Kim, Y.; Babnigg, G.; Jedrzejczak, R.; Eschenfeldt, W.H.; Li, H.; Maltseva, N.; Hatzos-Skintges, C.; Gu, M.; Makowska-Grzyska, M.; Wu, R.; et al. High-Throughput Protein Purification and Quality Assessment for Crystallization. Methods 2011, 55, 12–28. [Google Scholar] [CrossRef]
- García-Nafría, J.; Watson, J.F.; Greger, I.H. IVA Cloning: A Single-Tube Universal Cloning System Exploiting Bacterial In Vivo Assembly. Sci. Rep. 2016, 6, 27459. [Google Scholar] [CrossRef] [PubMed]
- Hou, Y.-M. Single-Turnover Kinetics of Methyl Transfer to TRNA by Methyltransferases. Methods Mol. Biol. 2016, 1421, 79–96. [Google Scholar] [CrossRef] [PubMed]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera--A Visualization System for Exploratory Research and Analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef] [PubMed]
- Dolinsky, T.J.; Nielsen, J.E.; McCammon, J.A.; Baker, N.A. PDB2PQR: An Automated Pipeline for the Setup of Poisson-Boltzmann Electrostatics Calculations. Nucleic Acids Res. 2004, 32, W665–W667. [Google Scholar] [CrossRef]
- Ruszkowska, A. METTL16, Methyltransferase-Like Protein 16: Current Insights into Structure and Function. Int. J. Mol. Sci. 2021, 22, 2176. [Google Scholar] [CrossRef]
- Malone, T.; Blumenthal, R.M.; Cheng, X. Structure-Guided Analysis Reveals Nine Sequence Motifs Conserved among DNA Amino-Methyltransferases, and Suggests a Catalytic Mechanism for These Enzymes. J. Mol. Biol. 1995, 253, 618–632. [Google Scholar] [CrossRef]
- Martin, J.L.; McMillan, F.M. SAM (Dependent) I AM: The S-Adenosylmethionine-Dependent Methyltransferase Fold. Curr. Opin. Struct. Biol. 2002, 12, 783–793. [Google Scholar] [CrossRef]
- Breger, K.; Kunkler, C.N.; O’Leary, N.J.; Hulewicz, J.P.; Brown, J.A. Ghost Authors Revealed: The Structure and Function of Human N6-Methyladenosine RNA Methyltransferases. Wiley Interdiscip. Rev. RNA 2023, 15, e1810. [Google Scholar] [CrossRef]
- Tate, J.G.; Bamford, S.; Jubb, H.C.; Sondka, Z.; Beare, D.M.; Bindal, N.; Boutselakis, H.; Cole, C.G.; Creatore, C.; Dawson, E.; et al. COSMIC: The Catalogue of Somatic Mutations Cancer. Nucleic Acids Res. 2019, 47, D941–D947. [Google Scholar] [CrossRef]
- Grossman, R.L.; Heath, A.P.; Ferretti, V.; Varmus, H.E.; Lowy, D.R.; Kibbe, W.A.; Staudt, L.M. Toward a Shared Vision for Cancer Genomic Data. N. Engl. J. Med. 2016, 375, 1109–1112. [Google Scholar] [CrossRef] [PubMed]
- Yeon, S.Y.; Jo, Y.S.; Choi, E.J.; Kim, M.S.; Yoo, N.J.; Lee, S.H. Frameshift Mutations in Repeat Sequences of ANK3, HACD4, TCP10L, TP53BP1, MFN1, LCMT2, RNMT, TRMT6, METTL8 and METTL16 Genes in Colon Cancers. Pathol. Oncol. Res. 2018, 24, 617–622. [Google Scholar] [CrossRef]
- Wang, T.; Birsoy, K.; Hughes, N.W.; Krupczak, K.M.; Post, Y.; Wei, J.J.; Lander, E.S.; Sabatini, D.M. Identification and Characterization of Essential Genes in the Human Genome. Science 2015, 350, 1096–1101. [Google Scholar] [CrossRef] [PubMed]
- Wu, T.; Qian, T.-Y.; Lin, R.-J.; Jin, D.-D.; Xu, X.-B.; Huang, M.-X.; Ji, J.; Jiang, F.; Pan, L.-L.; Luo, L.; et al. Construction and Validation of a m6A RNA Methylation and Ferroptosis-Related Prognostic Model for Pancreatic Cancer by Integrated Bioinformatics Analysis. J. Gastrointest. Oncol. 2022, 13, 2553–2564. [Google Scholar] [CrossRef] [PubMed]
- Ye, F.; Wu, J.; Zhang, F. METTL16 Epigenetically Enhances GPX4 Expression via m6A Modification to Promote Breast Cancer Progression by Inhibiting Ferroptosis. Biochem. Biophys. Res. Commun. 2023, 638, 1–6. [Google Scholar] [CrossRef]
- Wang, J.; Xiu, M.; Wang, J.; Gao, Y.; Li, Y. METTL16-SENP3-LTF Axis Confers Ferroptosis Resistance and Facilitates Tumorigenesis in Hepatocellular Carcinoma. J. Hematol. Oncol. 2024, 17, 78. [Google Scholar] [CrossRef]
- Zhao, S.; Cao, J.; Liang, R.; Peng, T.; Wu, S.; Liu, Z.; Wu, Y.; Song, L.; Sun, C.; Liu, Y.; et al. METTL16 Suppresses Ferroptosis in Cholangiocarcinoma by Promoting ATF4 via m6A Modification. Int. J. Biol. Sci. 2025, 21, 189–203. [Google Scholar] [CrossRef]
- Su, R.; Dong, L.; Li, Y.; Gao, M.; He, P.C.; Liu, W.; Wei, J.; Zhao, Z.; Gao, L.; Han, L.; et al. METTL16 Exerts an m6A-Independent Function to Facilitate Translation and Tumorigenesis. Nat. Cell Biol. 2022, 24, 205–216. [Google Scholar] [CrossRef]
- Wang, F.; Zhang, J.; Lin, X.; Yang, L.; Zhou, Q.; Mi, X.; Li, Q.; Wang, S.; Li, D.; Liu, X.-M.; et al. METTL16 Promotes Translation and Lung Tumorigenesis by Sequestering Cytoplasmic EIF4E2. Cell Rep. 2023, 42, 112150. [Google Scholar] [CrossRef]
- Mahendran, G.; Breger, K.; McCown, P.J.; Hulewicz, J.P.; Bhandari, T.; Addepalli, B.; Brown, J.A. Multi-Omics Approach Reveals Genes and Pathways Affected in Miller-Dieker Syndrome. Mol. Neurobiol. 2025, 62, 5073–5094. [Google Scholar] [CrossRef]
- Xue, M.; Dong, L.; Zhang, H.; Li, Y.; Qiu, K.; Zhao, Z.; Gao, M.; Han, L.; Chan, A.K.N.; Li, W.; et al. METTL16 Promotes Liver Cancer Stem Cell Self-Renewal via Controlling Ribosome Biogenesis and MRNA Translation. J. Hematol. Oncol. 2024, 17, 7. [Google Scholar] [CrossRef] [PubMed]
- Zeng, X.; Zhao, F.; Cui, G.; Zhang, Y.; Deshpande, R.A.; Chen, Y.; Deng, M.; Kloeber, J.A.; Shi, Y.; Zhou, Q.; et al. METTL16 Antagonizes MRE11-Mediated DNA End Resection and Confers Synthetic Lethality to PARP Inhibition in Pancreatic Ductal Adenocarcinoma. Nat. Cancer 2022, 3, 1088–1104. [Google Scholar] [CrossRef] [PubMed]
- Warda, A.S.; Kretschmer, J.; Hackert, P.; Lenz, C.; Urlaub, H.; Höbartner, C.; Sloan, K.E.; Bohnsack, M.T. Human METTL16 Is a N6-Methyladenosine m6A Methyltransferase That Targets Pre-MRNAs and Various Non-Coding RNAs. EMBO Rep. 2017, 18, 2004–2014. [Google Scholar] [CrossRef] [PubMed]
- Basu, S.; Bahadur, R.P. A Structural Perspective of RNA Recognition by Intrinsically Disordered Proteins. Cell Mol. Life Sci. 2016, 73, 4075–4084. [Google Scholar] [CrossRef]
- Hennig, J.; Sattler, M. Deciphering the Protein-RNA Recognition Code: Combining Large-Scale Quantitative Methods with Structural Biology. Bioessays 2015, 37, 899–908. [Google Scholar] [CrossRef]
- Didychuk, A.L.; Butcher, S.E.; Brow, D.A. The Life of U6 Small Nuclear RNA, from Cradle to Grave. RNA 2018, 24, 437–460. [Google Scholar] [CrossRef]
- Shimba, S.; Bokar, J.A.; Rottman, F.; Reddy, R. Accurate and Efficient N-6-Adenosine Methylation in Spliceosomal U6 Small Nuclear RNA by HeLa Cell Extract in Vitro. Nucleic Acids Res. 1995, 23, 2421–2426. [Google Scholar] [CrossRef]
- Sun, L.; Zhang, Y.; Yang, B.; Sun, S.; Zhang, P.; Luo, Z.; Feng, T.; Cui, Z.; Zhu, T.; Li, Y.; et al. Lactylation of METTL16 Promotes Cuproptosis via m6A-Modification on FDX1 MRNA in Gastric Cancer. Nat. Commun. 2023, 14, 6523. [Google Scholar] [CrossRef]
- Li, H.; Wilkinson, E.; Cui, Y.-H.; Sun, M.; Lu, K.; Yang, S.; Bissonnette, M.; He, Y.-Y. SSB Cooperates with METTL16-Mediated m6A RNA Methylation to Promote Chemoresistance in Colorectal Cancer Cells. Cell Rep. 2025, 44, 115926. [Google Scholar] [CrossRef]
- Ren, W.; Lu, J.; Huang, M.; Gao, L.; Li, D.; Wang, G.G.; Song, J. Structure and Regulation of ZCCHC4 in m6A-Methylation of 28S RRNA. Nat. Commun. 2019, 10, 5042. [Google Scholar] [CrossRef]
- Wang, X.; Feng, J.; Xue, Y.; Guan, Z.; Zhang, D.; Liu, Z.; Gong, Z.; Wang, Q.; Huang, J.; Tang, C.; et al. Structural Basis of N6-Adenosine Methylation by the METTL3-METTL14 Complex. Nature 2016, 534, 575–578. [Google Scholar] [CrossRef]
- Ju, J.; Aoyama, T.; Yashiro, Y.; Yamashita, S.; Kuroyanagi, H.; Tomita, K. Structure of the Caenorhabditis Elegans m6A Methyltransferase METT10 That Regulates SAM Homeostasis. Nucleic Acids Res. 2023, 51, 2434–2446. [Google Scholar] [CrossRef] [PubMed]
- van Tran, N.; Ernst, F.G.M.; Hawley, B.R.; Zorbas, C.; Ulryck, N.; Hackert, P.; Bohnsack, K.E.; Bohnsack, M.T.; Jaffrey, S.R.; Graille, M.; et al. The Human 18S RRNA m6A Methyltransferase METTL5 Is Stabilized by TRMT112. Nucleic Acids Res. 2019, 47, 7719–7733. [Google Scholar] [CrossRef] [PubMed]
- Oerum, S.; Catala, M.; Atdjian, C.; Brachet, F.; Ponchon, L.; Barraud, P.; Iannazzo, L.; Droogmans, L.; Braud, E.; Ethève-Quelquejeu, M.; et al. Bisubstrate Analogues as Structural Tools to Investigate m6A Methyltransferase Active Sites. RNA Biol. 2019, 16, 798–808. [Google Scholar] [CrossRef]
- Hartstock, K.; Kueck, N.A.; Spacek, P.; Ovcharenko, A.; Hüwel, S.; Cornelissen, N.V.; Bollu, A.; Dieterich, C.; Rentmeister, A. MePMe-Seq: Antibody-Free Simultaneous m6A and m5C Mapping in MRNA by Metabolic Propargyl Labeling and Sequencing. Nat. Commun. 2023, 14, 7154. [Google Scholar] [CrossRef]
- Du, J.; Mahcene, B.; Martynov, V.; Frezza, E.; Vasnier, C.; Ponchon, L.; Coelho, D.; Bonhomme, F.; Braud, E.; Etheve-Quelquejeu, M.; et al. Investigation of a Squaramide Motif as a Bioisostere of the Amino-Acid Group of S-Adenosyl-L-Methionine and Its Functional Impact on RNA Methylation. Commun. Chem. 2025, 8, 244. [Google Scholar] [CrossRef]
- Mikutis, S.; Gu, M.; Sendinc, E.; Hazemi, M.E.; Kiely-Collins, H.; Aspris, D.; Vassiliou, G.S.; Shi, Y.; Tzelepis, K.; Bernardes, G.J.L. MeCLICK-Seq, a Substrate-Hijacking and RNA Degradation Strategy for the Study of RNA Methylation. ACS Cent. Sci. 2020, 6, 2196–2208. [Google Scholar] [CrossRef]
- Ovcharenko, A.; Weissenboeck, F.P.; Rentmeister, A. Tag-Free Internal RNA Labeling and Photocaging Based on MRNA Methyltransferases. Angew. Chem. Int. Ed. Engl. 2021, 60, 4098–4103. [Google Scholar] [CrossRef]
- Wang, P.; Doxtader, K.A.; Nam, Y. Structural Basis for Cooperative Function of Mettl3 and Mettl14 Methyltransferases. Mol. Cell 2016, 63, 306–317. [Google Scholar] [CrossRef] [PubMed]
- Baker, C.M.; Grant, G.H. Role of Aromatic Amino Acids in Protein-Nucleic Acid Recognition. Biopolymers 2007, 85, 456–470. [Google Scholar] [CrossRef] [PubMed]
- Shi, Z.; Cao, X.; Ma, Y.; Li, K.; Wang, X.; Lin, J.; Tang, H.; Zhu, X. RNA Methyltransferase METTL16: From Molecular Mechanisms to Therapeutic Prospects in Cancers. Cancer Lett. 2025, 624, 217698. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Wang, S.; Chen, X.; Luo, Q.; Wang, Z.; Li, J. Pan-Cancer Analysis of Methyltransferase-like 16 (METTL16) and Validated in Colorectal Cancer. Aging 2025, 17, 588–606. [Google Scholar] [CrossRef] [PubMed]
- Michalak, E.M.; Burr, M.L.; Bannister, A.J.; Dawson, M.A. The Roles of DNA, RNA and Histone Methylation in Ageing and Cancer. Nat. Rev. Mol. Cell Biol. 2019, 20, 573–589. [Google Scholar] [CrossRef]
- Li, J.; Yang, F.; Wang, Z.; Zheng, S.; Zhang, S.; Wang, C.; He, B.; Wang, J.-B.; Wang, H. METTL16-Mediated N6-Methyladenosine Modification of Soga1 Enables Proper Chromosome Segregation and Chromosomal Stability in Colorectal Cancer. Cell Prolif. 2024, 57, e13590. [Google Scholar] [CrossRef]
- Bradley, R.K.; Anczuków, O. RNA Splicing Dysregulation and the Hallmarks of Cancer. Nat. Rev. Cancer 2023, 23, 135–155. [Google Scholar] [CrossRef]
- Mitra, A.; Manna, S.; Kundu, R.; Hazra, D.; Roychowdhury, A. Brute Force Virtual Drug Screening with Molecular Dynamics Simulation and MM/PBSA to Find Potent Inhibitors of METTL16. IEEE/ACM Trans. Comput. Biol. Bioinform. 2023, 20, 2356–2361. [Google Scholar] [CrossRef]
- Liu, Y.; Goebel, G.L.; Kanis, L.; Hastürk, O.; Kemker, C.; Wu, P. Aminothiazolone Inhibitors Disrupt the Protein-RNA Interaction of METTL16 and Modulate the m6A RNA Modification. JACS Au 2024, 4, 1436–1449. [Google Scholar] [CrossRef]
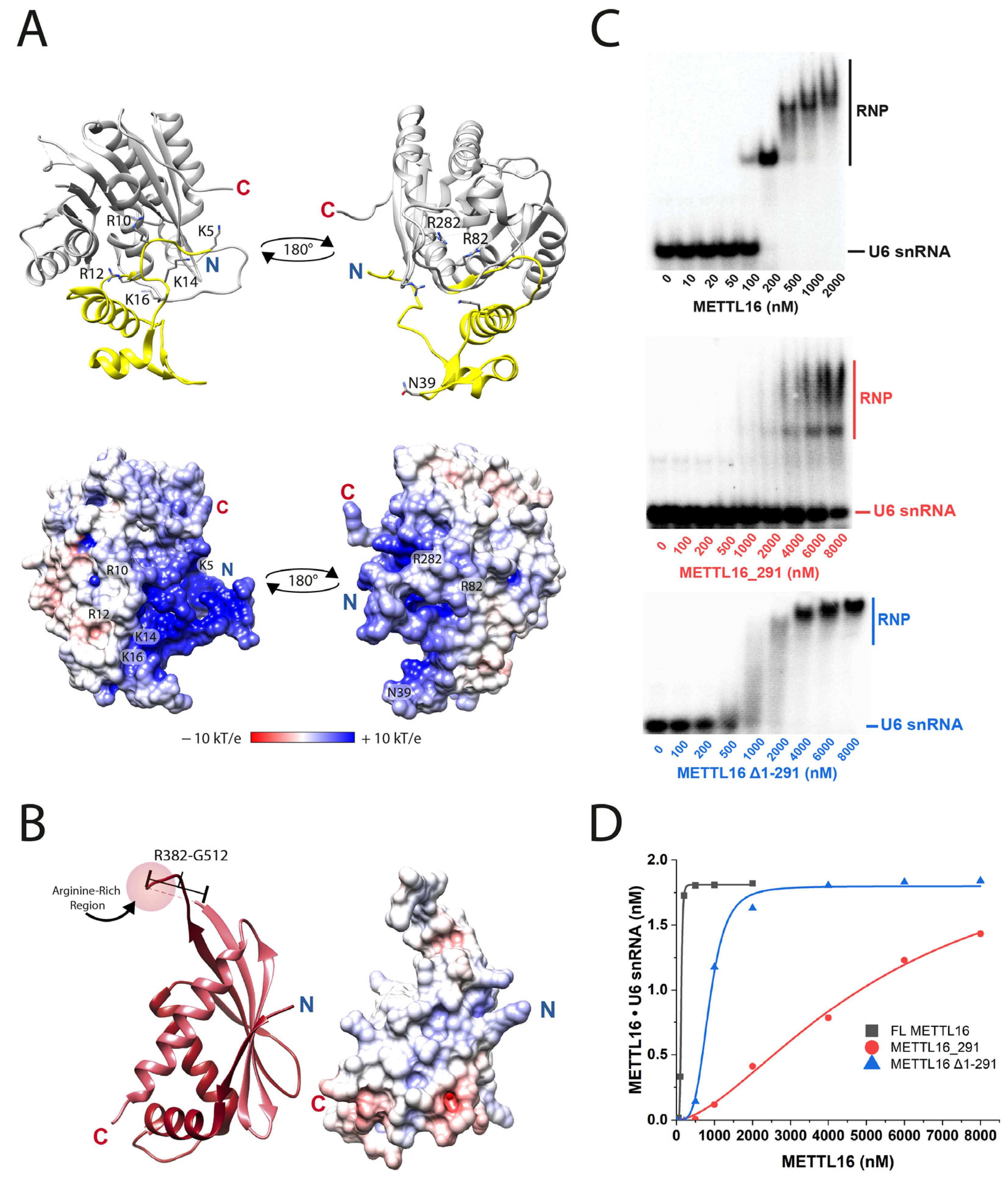
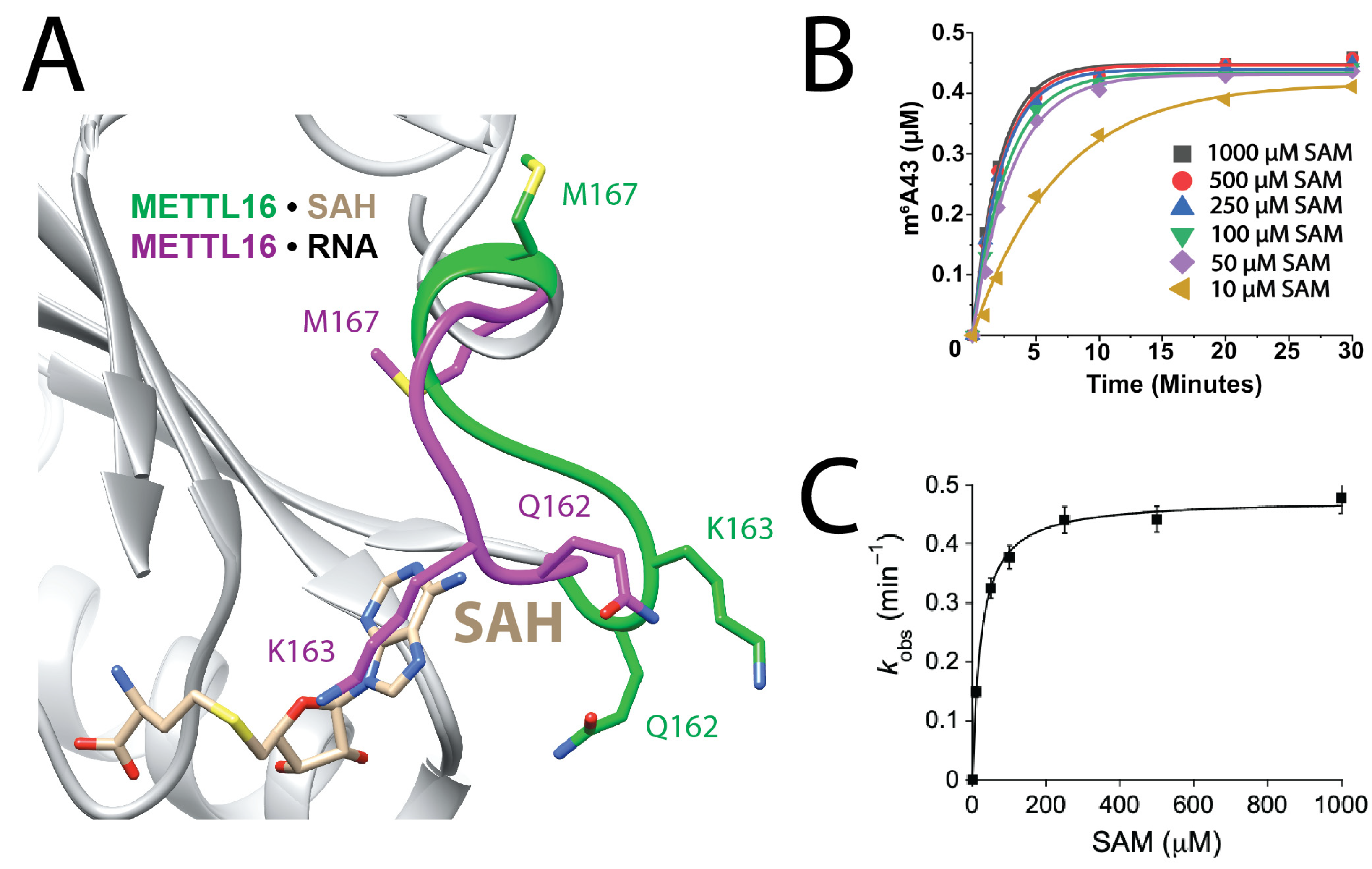
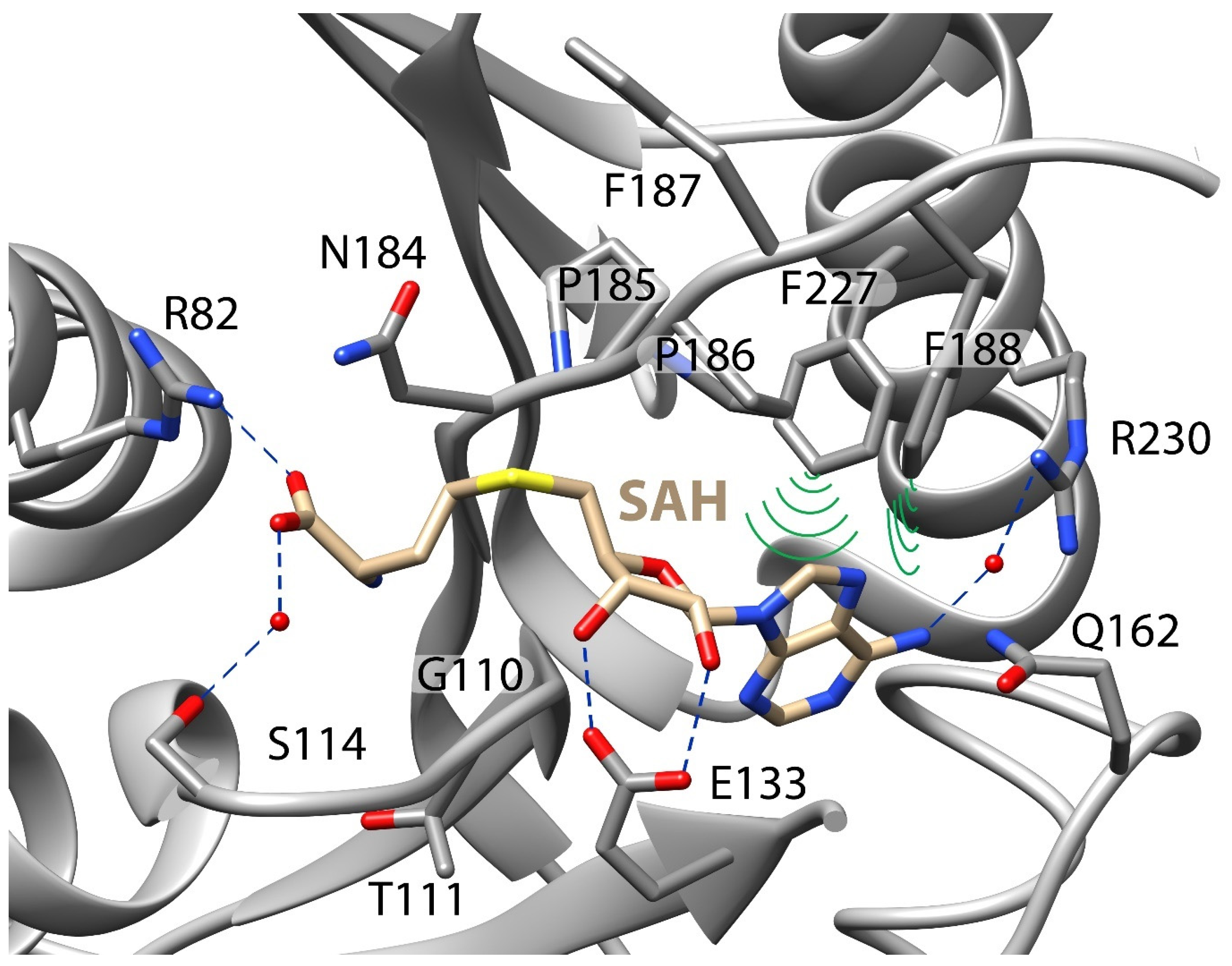
| METTL16 | KD1 (nM) | Degree of Cooperativity | Fold Weaker RNA Binding a |
|---|---|---|---|
| Full-length (FL) and truncated METTL16 | |||
| FL (1–562) | 132 ± 15 | 5 ± 1 | - |
| METTL16_291 (1–291) | 5200 ± 1300 | 1.6 ± 0.1 | 39 |
| METTL16Δ1-291 (291–562) | 814 ± 90 | 3.3 ± 0.5 | 6.2 |
| RNA-binding region (1–79) | |||
| K5A | 126 ± 10 | 7 ± 1 | 0.95 |
| K5A/R10A | 175 ± 27 | 6.8 ± 0.6 | 1.3 |
| K5A/R10A/R12A | 231 ± 58 | 4 ± 2 | 1.8 |
| K5A/R10A/R12A/K14A | 353 ± 25 | 6 ± 1 | 2.7 |
| K5A/R10A/R12A/K14A/K16A | 293 ± 35 | 4.3 ± 0.2 | 2.2 |
| N39A | 187 ± 0.5 | 7 ± 1 | 1.4 |
| Rossmann fold (80–291) | |||
| R82A | 306 ± 9 | 6.8 ± 0.6 | 2.3 |
| F187G | 330 ± 12 | 6.8 ± 0.4 | 2.5 |
| F187W | 276 ± 12 | 6.0 ± 0.8 | 2.1 |
| R282A | 307 ± 28 | 6.0 ± 0.3 | 2.3 |
| Arginine-rich region (382–388) in VCR1 | |||
| R382A | 294 ± 13 | 7 ± 1 | 2.2 |
| R382A/R383A/R386A/R388A | 649 ± 33 | 2.9 ± 0.1 | 4.9 |
| ΔR382-R388 | 1339 ± 122 | 2.7 ± 0.8 | 10.1 |
| METTL16 | KD2 (µM) | kchem (min−1) | kchem/KD2 (µM−1min−1) | Fold Tighter SAM Binding a |
|---|---|---|---|---|
| FL METTL16 b | 126 ± 6 | 0.56 ± 0.01 | 0.0044 | - |
| Q162A | 77 ± 6 | 0.49 ± 0.01 | 0.0064 | 1.6 |
| K163A | 23 ± 2 | 0.48 ± 0.01 | 0.021 | 5.5 |
| M167A | 43 ± 10 | 0.52 ± 0.03 | 0.012 | 2.9 |
| K163A/M167A | 14 ± 5 | 1.30 ± 0.07 | 0.093 | 9 |
| Q162A/K163A/M167A | 13 ± 2 | 1.98 ± 0.05 | 0.15 | 9.7 |
| METTL16 | KD2 (µM) | kchem (min−1) | kchem/KD2 (µM−1min−1) | Fold Weaker SAM Binding a |
|---|---|---|---|---|
| FL METTL16 b | 126 ± 6 | 0.56 ± 0.01 | 0.0044 | - |
| METTL16_291 b | 736 ± 94 | 0.42 ± 0.02 | 5.7 × 10−4 | 5.8 |
| R82A | 280 ± 38 | 0.012 ± 0.001 | 4.3 × 10−5 | 2.2 |
| T111A | 86 ± 9 | 0.45 ± 0.01 | 0.0052 | 0.7 |
| S114A | 286 ± 19 | 0.37 ± 0.01 | 0.0013 | 2.3 |
| E133A | >1000 | 0.04 ± 0.01 c | <4 × 10−5 | >7.9 |
| F188A | 383 ± 97 | 0.018 ± 0.002 | 4.7 × 10−5 | 3.0 |
| T216A | 42 ± 11 | 0.29 ± 0.01 | 0.0069 | 0.3 |
| F227A | >1000 | 0.36 ± 0.13 | <3.6 × 10−5 | >7.9 |
| R230A | 70 ± 5 | 0.54 ± 0.01 | 0.0077 | 0.6 |
| METTL16 | KD2 (µM) | kchem (min−1) | kchem/KD2 (µM−1min−1) | Relative Catalytic Efficiency a |
|---|---|---|---|---|
| FL METTL16 b | 126 ± 6 | 0.56 ± 0.01 | 0.0044 | -- |
| N184A | -- | No measurable activity | -- | -- |
| N184D | -- | No measurable activity | -- | -- |
| N184D/F187W | -- | No measurable activity | -- | -- |
| P185A/P186A | -- | No measurable activity | -- | -- |
| F187G | 227 ± 36 | 0.0034 ± 0.0002 | 1.5 × 10−5 | ↓ 293 |
| F187W | 38 ± 10 | 0.44 ± 0.02 | 0.012 | ↑ 2.7 |
| METTL16 Cancer-Associated Mutation (Cancer Type) | KD1 (nM) | KD2 (µM) | kchem (min−1) | kchem/KD2 (µM−1min−1) | Relative Catalytic Efficiency a |
|---|---|---|---|---|---|
| Degree of Cooperativity | |||||
| FL METTL16 b | 132 ± 15 | 126 ± 6 | 0.56 ± 0.01 | 0.0044 | - |
| 5 ± 1 | |||||
| METTL16_291 (1–291) | 5200 ± 1300 | 736 ± 94 | 0.42 ± 0.02 | 5.7 × 10−4 | ↓ 7.7 |
| 1.6 ± 0.1 | |||||
| Rossmann fold | |||||
| G110C (Intestinal) | 179 ± 19 | >1000 | 0.005 ± 0.003 c | <5 × 10−6 | ↓ >880 |
| 14 ± 10 | |||||
| R200Q (Intestinal) | 160 ± 6 | 66 ± 9 | 0.41 ± 0.01 | 0.0062 | ↑ 1.4 |
| 9.8 ± 0.3 | |||||
| R241Dfs*2 (Colorectal) | 5960 ± 420 | -- | No measurable activity | -- | -- |
| 4.9 ± 0.2 | |||||
| VCR1 | |||||
| E408K (Esophageal) | 181 ± 2 | 134 ± 21 | 0.48 ± 0.02 | 0.0036 | ↓ 1.2 |
| 5.6 ± 0.7 | |||||
| Disordered region | |||||
| P460L (Liver) | 131 ± 4 | 149 ± 21 | 0.53 ± 0.02 | 0.0036 | ↓ 1.2 |
| 6.4 ± 0.8 | |||||
| VCR2 | |||||
| T549A (Central Nervous System) | 148 ± 2 | 120 ± 4 | 0.44 ± 0.01 | 0.0037 | ↓ 1.2 |
| 9 ± 2 | |||||
| R552H (Stomach) | 137 ± 9 | 125 ± 10 | 0.50 ± 0.01 | 0.0040 | ↓ 1.1 |
| 7.1 ± 0.4 | |||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Breger, K.; Schowe, I.P.; Springer, N.A.; O’Leary, N.J.; Ruszkowska, A.; Resende, C.; Brown, J.A. Mutational Analysis Reveals Functional Roles of METTL16 Domains and Residues. Biology 2025, 14, 1145. https://doi.org/10.3390/biology14091145
Breger K, Schowe IP, Springer NA, O’Leary NJ, Ruszkowska A, Resende C, Brown JA. Mutational Analysis Reveals Functional Roles of METTL16 Domains and Residues. Biology. 2025; 14(9):1145. https://doi.org/10.3390/biology14091145
Chicago/Turabian StyleBreger, Kurtis, Ian P. Schowe, Noah A. Springer, Nathan J. O’Leary, Agnieszka Ruszkowska, Carlos Resende, and Jessica A. Brown. 2025. "Mutational Analysis Reveals Functional Roles of METTL16 Domains and Residues" Biology 14, no. 9: 1145. https://doi.org/10.3390/biology14091145
APA StyleBreger, K., Schowe, I. P., Springer, N. A., O’Leary, N. J., Ruszkowska, A., Resende, C., & Brown, J. A. (2025). Mutational Analysis Reveals Functional Roles of METTL16 Domains and Residues. Biology, 14(9), 1145. https://doi.org/10.3390/biology14091145






