VaccineDesigner: A Web-Based Tool for Streamlined Multi-Epitope Vaccine Design
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Epitope Prediction and Evaluation
2.1.1. B-Cell Epitope Prediction
2.1.2. CTL Epitope Prediction
2.1.3. HTL Epitope Prediction
2.1.4. Epitope Evaluation
2.2. Multi-Epitope Vaccine Sequences
2.2.1. Vaccine Sequence Construction
2.2.2. Vaccine Candidate Sequence Evaluation and Selection
3. Results
3.1. Usage Scenario
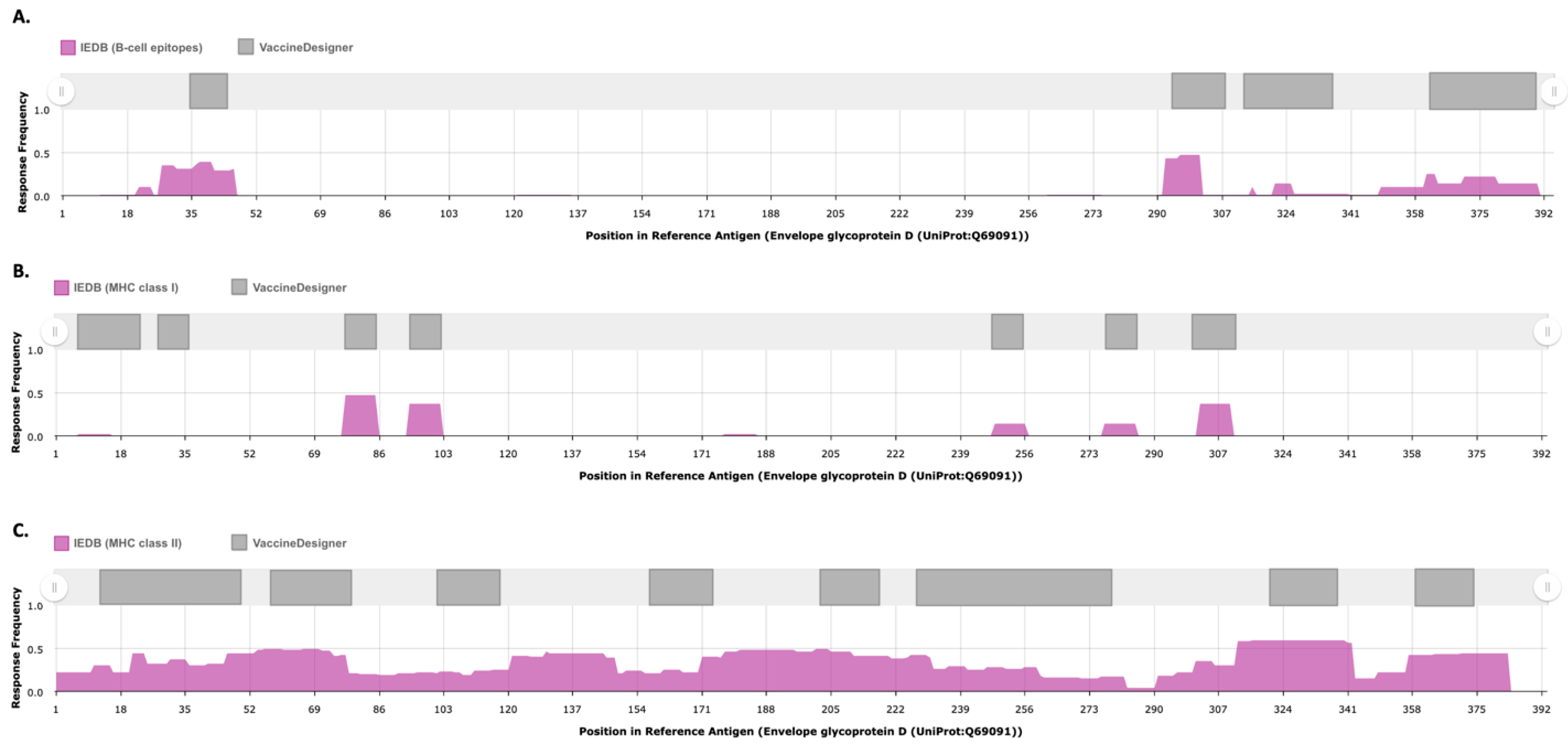
3.2. Evaluation Against Experimentally Validated Epitopes
3.3. Comparison with Other RV Pipelines
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| APCs | Antigen-presenting cells |
| CTL | Cytotoxic T cells |
| HTL | Helper T cells |
| IEDB | Immune Epitope Database |
| IEDB-AR | Immune Epitope Database and Analysis Resource |
| MHC | Major histocompatibility complex |
| RV | Reverse vaccinology |
References
- Sette, A.; Rappuoli, R. Reverse Vaccinology: Developing Vaccines in the Era of Genomics. Immunity 2010, 33, 530–541. [Google Scholar] [CrossRef]
- Rappuoli, R. Reverse vaccinology. Curr. Opin. Microbiol. 2000, 3, 445–450. [Google Scholar] [CrossRef]
- Soria-Guerra, R.E.; Nieto-Gomez, R.; Govea-Alonso, D.O.; Rosales-Mendoza, S. An overview of bioinformatics tools for epitope prediction: Implications on vaccine development. J. Biomed. Inform. 2015, 53, 405–414. [Google Scholar] [CrossRef] [PubMed]
- Suhrbier, A. Multi-epitope DNA vaccines. Immunol. Cell Biol. 1997, 75, 402–488. [Google Scholar] [CrossRef] [PubMed]
- Negahdaripour, M.; Nezafat, N.; Eslami, M.; Ghoshoon, M.B.; Shoolian, E.; Najafipour, S.; Morowvat, M.H.; Dehshahri, A.; Erfani, N.; Ghasemi, Y. Structural vaccinology considerations for in silico designing of a multi-epitope vaccine. Infect. Genet. Evol. 2018, 58, 96–109. [Google Scholar] [CrossRef]
- Chen, X.; Zaro, J.L.; Shen, W.-C. Fusion protein linkers: Property, design and functionality. Adv. Drug Deliv. Rev. 2013, 65, 1357–1369. [Google Scholar] [CrossRef] [PubMed]
- Khatoon, N.; Pandey, R.K.; Prajapati, V.K. Exploring Leishmania secretory proteins to design B and T cell multi-epitope subunit vaccine using immunoinformatics approach. Sci. Rep. 2017, 7, 8285. [Google Scholar] [CrossRef] [PubMed]
- Shey, R.A.; Ghogomu, S.M.; Esoh, K.K.; Nebangwa, N.D.; Shintouo, C.M.; Nongley, N.F.; Asa, B.F.; Ngale, F.N.; Vanhamme, L.; Souopgui, J. In-silico design of a multi-epitope vaccine candidate against onchocerciasis and related filarial diseases. Sci. Rep. 2019, 9, 4409. [Google Scholar] [CrossRef] [PubMed]
- Enayatkhani, M.; Hasaniazad, M.; Faezi, S.; Gouklani, H.; Davoodian, P.; Ahmadi, N.; Einakian, M.A.; Karmostaji, A.; Ahmadi, K. Reverse vaccinology approach to design a novel multi-epitope vaccine candidate against COVID-19: An in silico study. J. Biomol. Struct. Dyn. 2021, 39, 2857–2872. [Google Scholar] [CrossRef]
- Dong, R.; Chu, Z.; Yu, F.; Zha, Y. Contriving Multi-Epitope Subunit of Vaccine for COVID-19: Immunoinformatics Approaches. Front. Immunol. 2020, 11, 1784. [Google Scholar] [CrossRef]
- Alizadeh, M.; Amini-Khoei, H.; Tahmasebian, S.; Ghatrehsamani, M.; Ghatreh Samani, K.; Edalatpanah, Y.; Rostampur, S.; Salehi, M.; Ghasemi-Dehnoo, M.; Azadegan-Dehkordi, F.; et al. Designing a novel multi-epitope vaccine against Ebola virus using reverse vaccinology approach. Sci. Rep. 2022, 12, 7757. [Google Scholar] [CrossRef]
- Khan, M.; Khan, S.; Ali, A.; Akbar, H.; Sayaf, A.M.; Khan, A.; Wei, D.Q. Immunoinformatics approaches to explore Helicobacter Pylori proteome (Virulence Factors) to design B and T cell multi-epitope subunit vaccine. Sci. Rep. 2019, 9, 13321. [Google Scholar] [CrossRef]
- Dar, H.A.; Zaheer, T.; Shehroz, M.; Ullah, N.; Naz, K.; Muhammad, S.A.; Zhang, T.; Ali, A. Immunoinformatics-aided design and evaluation of a potential multi-epitope vaccine against klebsiella pneumoniae. Vaccines 2019, 7, 88. [Google Scholar] [CrossRef]
- Ayyagari, V.S.; Venkateswarulu, T.C.; Abraham Peele, K.; Srirama, K. Design of a multi-epitope-based vaccine targeting M-protein of SARS-CoV2: An immunoinformatics approach. J. Biomol. Struct. Dyn. 2022, 40, 2963–2977. [Google Scholar] [CrossRef]
- Potocnakova, L.; Bhide, M.; Pulzova, L.B. An Introduction to B-Cell Epitope Mapping and in Silico Epitope Prediction. J. Immunol. Res. 2016, 2016, 6760830. [Google Scholar] [CrossRef]
- Shepherd, F.R.; McLaren, J.E. T cell immunity to bacterial pathogens: Mechanisms of immune control and bacterial evasion. Int. J. Mol. Sci. 2020, 21, 6144. [Google Scholar] [CrossRef]
- Zhang, L. Multi-epitope vaccines: A promising strategy against tumors and viral infections. Cell. Mol. Immunol. 2018, 15, 182–184. [Google Scholar] [CrossRef] [PubMed]
- Sette, A.; Livingston, B.; McKinney, D.; Appella, E.; Fikes, J.; Sidney, J.; Newman, M.; Chesnut, R. The Development of Multi-epitope Vaccines: Epitope Identification, Vaccine Design and Clinical Evaluation. Biologicals 2001, 29, 271–276. [Google Scholar] [CrossRef] [PubMed]
- Clifford, J.N.; Høie, M.H.; Deleuran, S.; Peters, B.; Nielsen, M.; Marcatili, P. BepiPred-3.0: Improved B-cell epitope prediction using protein language models. Protein Sci. 2022, 31, e4497. [Google Scholar] [CrossRef] [PubMed]
- Saha, S.; Raghava, G.P.S. BcePred: Prediction of continuous B-cell epitopes in antigenic sequences using physico-chemical properties. Lect. Notes Comput. Sci. 2004, 3239, 197–204. [Google Scholar] [CrossRef]
- Saha, S.; Raghava, G.P.S. Prediction of Continuous B-Cell Epitopes in an Antigen Using Recurrent Neural Network. Proteins Struct. Funct. Bioinform. 2006, 65, 40–48. [Google Scholar] [CrossRef] [PubMed]
- Ansari, H.R.; Raghava, G.P. Identification of conformational B-cell Epitopes in an antigen from its primary sequence. Immunome Res. 2010, 6, 6. [Google Scholar] [CrossRef] [PubMed]
- Høie, M.H.; Gade, F.S.; Johansen, J.M.; Würtzen, C.; Winther, O.; Nielsen, M.; Marcatili, P. DiscoTope-3.0: Improved B-cell epitope prediction using inverse folding latent representations. Front. Immunol. 2024, 15, 1322712. [Google Scholar] [CrossRef] [PubMed]
- Ponomarenko, J.; Bui, H.H.; Li, W.; Fusseder, N.; Bourne, P.E.; Sette, A.; Peters, B. ElliPro: A new structure-based tool for the prediction of antibody epitopes. BMC Bioinform. 2008, 9, 514. [Google Scholar] [CrossRef]
- Reynisson, B.; Alvarez, B.; Paul, S.; Peters, B.; Nielsen, M. NetMHCpan-4.1 and NetMHCIIpan-4.0: Improved predictions of MHC antigen presentation by concurrent motif deconvolution and integration of MS MHC eluted ligand data. Nucleic Acids Res. 2021, 48, W449–W454. [Google Scholar] [CrossRef]
- Singh, H.; Raghava, G.P.S. ProPred: Prediction of HLA-DR binding sites. Bioinformatics 2001, 17, 1236–1237. [Google Scholar] [CrossRef]
- Wilkins, M.R.; Gasteiger, E.; Bairoch, A.; Sanchez, J.C.; Williams, K.L.; Appel, R.D.; Hochstrasser, D.F. Protein identification and analysis tools in the ExPASy server. Methods Mol. Biol. 1999, 112, 531–552. [Google Scholar] [CrossRef]
- Doytchinova, I.A.; Flower, D.R. VaxiJen: A server for prediction of protective antigens, tumour antigens and subunit vaccines. BMC Bioinform. 2007, 8, 4. [Google Scholar] [CrossRef]
- Ong, E.; Wang, H.; Wong, M.U.; Seetharaman, M.; Valdez, N.; He, Y. Vaxign-ML: Supervised machine learning reverse vaccinology model for improved prediction of bacterial protective antigens. Bioinformatics 2020, 36, 3185–3191. [Google Scholar] [CrossRef]
- Dhanda, S.K.; Vir, P.; Raghava, G.P.S. Designing of interferon-gamma inducing MHC class-II binders. Biol. Direct 2013, 8, 30. [Google Scholar] [CrossRef]
- Sharma, N.; Patiyal, S.; Dhall, A.; Pande, A.; Arora, C.; Raghava, G.P.S. AlgPred 2.0: An improved method for predicting allergenic proteins and mapping of IgE epitopes. Brief. Bioinform. 2021, 22, bbaa294. [Google Scholar] [CrossRef] [PubMed]
- Sharma, N.; Naorem, L.D.; Jain, S.; Raghava, G.P.S. ToxinPred2: An improved method for predicting toxicity of proteins. Brief. Bioinform. 2022, 23, bbac174. [Google Scholar] [CrossRef] [PubMed]
- Ong, E.; Cooke, M.F.; Huffman, A.; Xiang, Z.; Wong, M.U.; Wang, H.; Seetharaman, M.; Valdez, N.; He, Y. Vaxign2: The second generation of the first Web-based vaccine design program using reverse vaccinology and machine learning. Nucleic Acids Res. 2021, 49, W671–W678. [Google Scholar] [CrossRef]
- Moise, L.; Gutierrez, A.; Kibria, F.; Martin, R.; Tassone, R.; Liu, R.; Terry, F.; Martin, B.; De Groot, A.S. Ivax: An integrated toolkit for the selection and optimization of antigens and the design of epitope-driven vaccines. Hum. Vaccines Immunother. 2015, 11, 2312–2321. [Google Scholar] [CrossRef]
- Goodswen, S.J.; Kennedy, P.J.; Ellis, J.T. Vacceed: A high-throughput in silico vaccine candidate discovery pipeline for eukaryotic pathogens based on reverse vaccinology. Bioinformatics 2014, 30, 2381–2383. [Google Scholar] [CrossRef] [PubMed]
- Dhanda, S.K.; Mahajan, S.; Paul, S.; Yan, Z.; Kim, H.; Jespersen, M.C.; Jurtz, V.; Andreatta, M.; Greenbaum, J.A.; Marcatili, P.; et al. IEDB-AR: Immune epitope database—Analysis resource in 2019. Nucleic Acids Res. 2019, 47, W502–W506. [Google Scholar] [CrossRef]
- Dalsass, M.; Brozzi, A.; Medini, D.; Rappuoli, R. Comparison of Open-Source Reverse Vaccinology Programs for Bacterial Vaccine Antigen Discovery. Front. Immunol. 2019, 10, 113. [Google Scholar] [CrossRef]
- Srivastava, P.; Jain, C.K. Computer Aided Reverse Vaccinology: A Game-changer Approach for Vaccine Development. Comb. Chem. High Throughput Screen. 2023, 26, 1813–1821. [Google Scholar] [CrossRef]
- Schubert, B.; de la Garza, L.; Mohr, C.; Walzer, M.; Kohlbacher, O. ImmunoNodes—Graphical development of complex immunoinformatics workflows. BMC Bioinform. 2017, 18, 242. [Google Scholar] [CrossRef]
- Raoufi, E.; Hemmati, M.; Eftekhari, S.; Khaksaran, K.; Mahmodi, Z.; Farajollahi, M.M.; Mohsenzadegan, M. Epitope Prediction by Novel Immunoinformatics Approach: A State-of-the-art Review. Int. J. Pept. Res. Ther. 2020, 26, 1155–1163. [Google Scholar] [CrossRef]
- Paul, S.; Sidney, J.; Sette, A.; Peters, B. TepiTool: A Pipeline for Computational Prediction of T Cell Epitope Candidates. Curr. Protoc. Immunol. 2016, 114, 18.19.1–18.19.24. [Google Scholar] [CrossRef]
- Yang, Y.; Sun, W.; Guo, J.; Zhao, G.; Sun, S.; Yu, H.; Guo, Y.; Li, J.; Jin, X.; Du, L.; et al. In silico design of a DNA-based HIV-1 multi-epitope vaccine for Chinese populations. Hum. Vaccines Immunother. 2015, 11, 795–805. [Google Scholar] [CrossRef]
- Livingston, B.; Crimi, C.; Newman, M.; Higashimoto, Y.; Appella, E.; Sidney, J.; Sette, A. A rational strategy to design multiepitope immunogens based on multiple Th lymphocyte epitopes. J. Immunol. 2002, 168, 5499–5506. [Google Scholar] [CrossRef]
- Li, X.; Guo, L.; Kong, M.; Su, X.; Yang, D.; Zou, M.; Liu, Y.; Lu, L. Design and Evaluation of a Multi-Epitope Peptide of Human Metapneumovirus. Intervirology 2016, 58, 403–412. [Google Scholar] [CrossRef]
- Yano, A.; Onozuka, A.; Asahi-Ozaki, Y.; Imai, S.; Hanada, N.; Miwa, Y.; Nisizawa, T. An ingenious design for peptide vaccines. Vaccine 2005, 23, 2322–2326. [Google Scholar] [CrossRef]
- Willimsky, G.; Beier, C.; Immisch, L.; Papafotiou, G.; Scheuplein, V.; Goede, A.; Holzhütter, H.-G.; Blankenstein, T.; Kloetzel, P.M. In vitro proteasome processing of neo-splicetopes does not predict their presentation in vivo. Biochem. Chem. Biol. Immunol. Inflamm. 2021, 10, e62019. [Google Scholar] [CrossRef] [PubMed]
- Díaz-Dinamarca, D.A.; Salazar, M.L.; Castillo, B.N.; Manubens, A.; Vasquez, A.E.; Salazar, F.; Becker, M.I. Protein-Based Adjuvants for Vaccines as Immunomodulators of the Innate and Adaptive Immune Response: Current Knowledge, Challenges, and Future Opportunities. Pharmaceutics 2022, 14, 1671. [Google Scholar] [CrossRef] [PubMed]
- Toussi, D.N.; Carraway, M.; Wetzler, L.M.; Lewis, L.A.; Liu, X.; Massari, P. The Amino Acid Sequence of Neisseria lactamica PorB Surface-Exposed Loops Influences Toll-Like Receptor 2-Dependent Cell Activation. Infect. Immun. 2012, 80, 3417–3428. [Google Scholar] [CrossRef]
- Ghaffari-Nazari, H.; Tavakkol-Afshari, J.; Jaafari, M.R.; Tahaghoghi-Hajghorbani, S.; Masoumi, E.; Jalali, S.A. Improving Multi-Epitope Long Peptide Vaccine Potency by Using a Strategy that Enhances CD4+ T Help in BALB/c Mice. PLoS ONE 2015, 10, e0142563. [Google Scholar] [CrossRef] [PubMed]
- Giudice, G. Del Hsp70: A carrier molecule with built-in adjuvanticity. Experientia 1994, 50, 1061–1066. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Bogdan, P.; Nazarian, S. An in silico deep learning approach to multi-epitope vaccine design: A SARS-CoV-2 case study. Sci. Rep. 2021, 11, 3238. [Google Scholar] [CrossRef] [PubMed]
- Bairoch, A.; Apweiler, R. The SWISS-PROT protein sequence database and its supplement TrEMBL in 2000. Nucleic Acids Res. 2000, 28, 45–48. [Google Scholar] [CrossRef]
- McGinnis, S.; Madden, T.L. BLAST: At the core of a powerful and diverse set of sequence analysis tools. Nucleic Acids Res. 2004, 1, W20–W25. [Google Scholar] [CrossRef]
- Saxová, P.; Buus, S.; Brunak, S.; Keşmir, C. Predicting proteasomal cleavage sites: A comparison of available methods. Int. Immunol. 2003, 15, 781–787. [Google Scholar] [CrossRef] [PubMed]
- Stranger-Jones, Y.K.; Bae, T.; Schneewind, O. Vaccine assembly from surface proteins of Staphylococcus aureus. Proc. Natl. Acad. Sci. USA 2006, 103, 16942–16947. [Google Scholar] [CrossRef]
- Dhanda, S.K.; Vita, R.; Ha, B.; Grifoni, A.; Peters, B.; Sette, A. ImmunomeBrowser: A tool to aggregate and visualize complex and heterogeneous epitopes in reference proteins. Bioinformatics 2018, 34, 3931–3933. [Google Scholar] [CrossRef]
- Baron, S.; Fons, M.; Albrecht, T. Viral Pathogenesis. In Medical Microbiology, 4th ed.; Baron, S., Ed.; University of Texas Medical Branch at Galveston: Galveston, TX, USA, 1996. [Google Scholar]
- Rist, M.J.; Theodossis, A.; Croft, N.P.; Neller, M.A.; Welland, A.D.; Chen, Z.; Sullivan, L.C.; Burrows, J.M.; Miles, J.J.; Brennan, R.M.; et al. HLA peptide length preferences control CD8+ T cell responses. J. Immunol. 2013, 191, 561–571. [Google Scholar] [CrossRef]
- Hemmer, B.; Kondo, T.; Gran, B.; Pinilla, C.; Cortese, I.; Pascal, J.; Tzou, A.; McFarland, H.F.; Houghten, R.; Martin, R. Minimal peptide length requirements for CD4+ T cell clones—Implications for molecular mimicry and T cell survival. Int. Immunol. 2000, 12, 375–383. [Google Scholar] [CrossRef]
- Liu, G.; Carter, B.; Bricken, T.; Jain, S.; Viard, M.; Carrington, M.; Gifford, D.K. Computationally Optimized SARS-CoV-2 MHC Class I and II Vaccine Formulations Predicted to Target Human Haplotype Distributions. Cell Syst. 2020, 11, 131–144.e6. [Google Scholar] [CrossRef]
- D’Mello, A.; Ahearn, C.P.; Murphy, T.F.; Tettelin, H. ReVac: A reverse vaccinology computational pipeline for prioritization of prokaryotic protein vaccine candidates. BMC Genom. 2019, 20, 981. [Google Scholar] [CrossRef] [PubMed]
- Rizwan, M.; Naz, A.; Ahmad, J.; Naz, K.; Obaid, A.; Parveen, T.; Ahsan, M.; Ali, A. VacSol: A high throughput in silico pipeline to predict potential therapeutic targets in prokaryotic pathogens using subtractive reverse vaccinology. BMC Bioinform. 2017, 18, 106. [Google Scholar] [CrossRef]
- Vivona, S.; Bernante, F.; Filippini, F. NERVE: New Enhanced Reverse Vaccinology Environment. BMC Biotechnol. 2006, 6, 35. [Google Scholar] [CrossRef]
- Goodswen, S.J.; Kennedy, P.J.; Ellis, J.T. A guide to current methodology and usage of reverse vaccinology towards in silico vaccine discovery. FEMS Microbiol. Rev. 2023, 47, fuad004. [Google Scholar] [CrossRef]
- Hebditch, M.; Carballo-Amador, M.A.; Charonis, S.; Curtis, R.; Warwicker, J. Protein–Sol: A web tool for predicting protein solubility from sequence. Bioinformatics 2017, 33, 3098–3100. [Google Scholar] [CrossRef] [PubMed]
- Puigbo, P.; Guzman, E.; Romeu, A.; Garcia-Vallve, S. OPTIMIZER: A web server for optimizing the codon usage of DNA sequences. Nucleic Acids Res. 2007, 35, W126–W131. [Google Scholar] [CrossRef] [PubMed]
- Gruber, A.R.; Lorenz, R.; Bernhart, S.H.; Neubock, R.; Hofacker, I.L. The Vienna RNA Websuite. Nucleic Acids Res. 2008, 36, W70–W74. [Google Scholar] [CrossRef] [PubMed]
- Mirdita, M.; Schutze, K.; Moriwaki, Y.; Heo, L.; Ovchinnikov, S.; Steinegger, M. ColabFold: Making protein folding accessible to all. Nat. Methods 2022, 19, 679–682. [Google Scholar] [CrossRef] [PubMed]
- Naz, K.; Naz, A.; Ashraf, S.T.; Rizwan, M.; Ahmad, J.; Baumbach, J.; Ali, A. PanRV: Pangenome-reverse vaccinology approach for identifications of potential vaccine candidates in microbial pangenome. BMC Bioinform. 2019, 20, 123. [Google Scholar] [CrossRef]
- Abramson, J.; Adler, J.; Dunger, J.; Evans, R.; Green, T.; Pritzel, A.; Ronneberger, O.; Willmore, L.; Ballard, A.J.; Bambrick, J.; et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 2024, 630, 493–500. [Google Scholar] [CrossRef]



| Name/Tool | Web Interface | Streamlined Process (Yes/No) | Epitope Types (BP/TP/BP + TP) | Non-Human Species (Yes/No) | Multi-Epitope Construction (Yes/No) | Pathogen Protein Selection (Yes/No) | Vaccine Candidate Evaluation (*) | Source Code Availability (Yes/No) |
|---|---|---|---|---|---|---|---|---|
| NERVE [63] | No | Yes | No | No | No | Yes | SL, AP, CR | Yes |
| iVAX [34] ** | Yes | Yes | TP | Yes | No | Yes | I, C, CR, A | No |
| OptiVax [60] | No | Yes | TP | Yes | Yes | Yes | I, PC, SP | Yes |
| ReVac [61] | No | Yes | BP + TP | Yes | No | Yes | A, C, SL, CR | Yes |
| IEDB-AR [36] | Yes | No | BP + TP | Yes | No | No | No | Yes |
| Vacceed [35] | No | Yes | TP | Yes | No | Yes | SL, SP, TH, T | Yes |
| VacSOL [62] | No | Yes | BP + TP | No | No | Yes | SL, V, TH, A, B + T | Yes |
| Vaxign2 [33] | Yes | Yes | TP | No | No | Yes | SL, TH, AP, B + T | Yes |
| VaccineDesigner | Yes | Yes | BP + TP | Yes | Yes | No | B + T, A, S, CR, PC | Yes |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Trygoniaris, D.; Korda, A.; Paraskeva, A.; Dushku, E.; Tzimagiorgis, G.; Yiangou, M.; Kotzamanidis, C.; Malousi, A. VaccineDesigner: A Web-Based Tool for Streamlined Multi-Epitope Vaccine Design. Biology 2025, 14, 1019. https://doi.org/10.3390/biology14081019
Trygoniaris D, Korda A, Paraskeva A, Dushku E, Tzimagiorgis G, Yiangou M, Kotzamanidis C, Malousi A. VaccineDesigner: A Web-Based Tool for Streamlined Multi-Epitope Vaccine Design. Biology. 2025; 14(8):1019. https://doi.org/10.3390/biology14081019
Chicago/Turabian StyleTrygoniaris, Dimitrios, Anna Korda, Anastasia Paraskeva, Esmeralda Dushku, Georgios Tzimagiorgis, Minas Yiangou, Charalampos Kotzamanidis, and Andigoni Malousi. 2025. "VaccineDesigner: A Web-Based Tool for Streamlined Multi-Epitope Vaccine Design" Biology 14, no. 8: 1019. https://doi.org/10.3390/biology14081019
APA StyleTrygoniaris, D., Korda, A., Paraskeva, A., Dushku, E., Tzimagiorgis, G., Yiangou, M., Kotzamanidis, C., & Malousi, A. (2025). VaccineDesigner: A Web-Based Tool for Streamlined Multi-Epitope Vaccine Design. Biology, 14(8), 1019. https://doi.org/10.3390/biology14081019





