Genetic Monitoring of Grey Wolves in Latvia Shows Adverse Reproductive and Social Consequences of Hunting
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
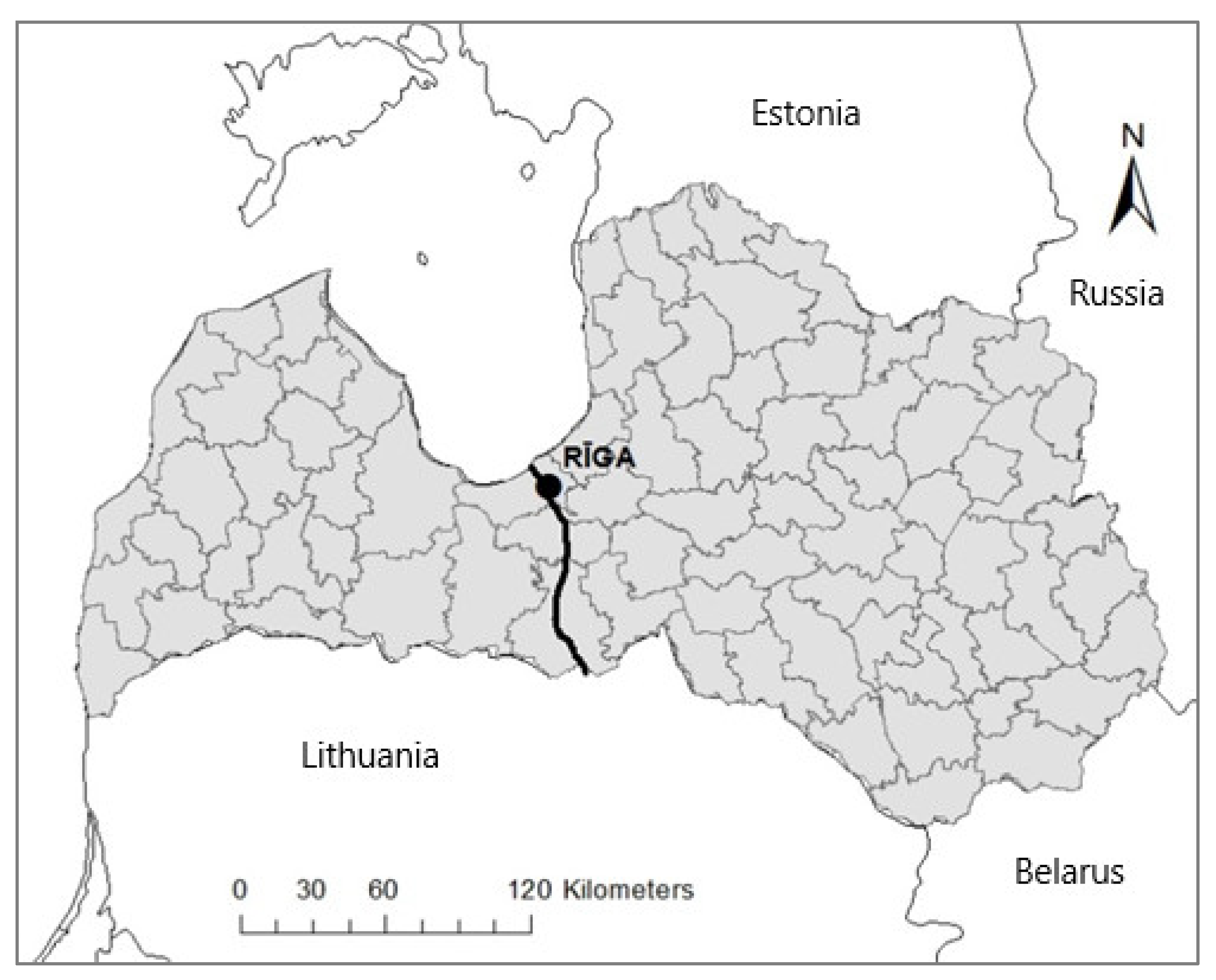
2.1. Study Area and Wolf Harvest in Latvia
2.2. Sample Collection and Analyses
3. Results
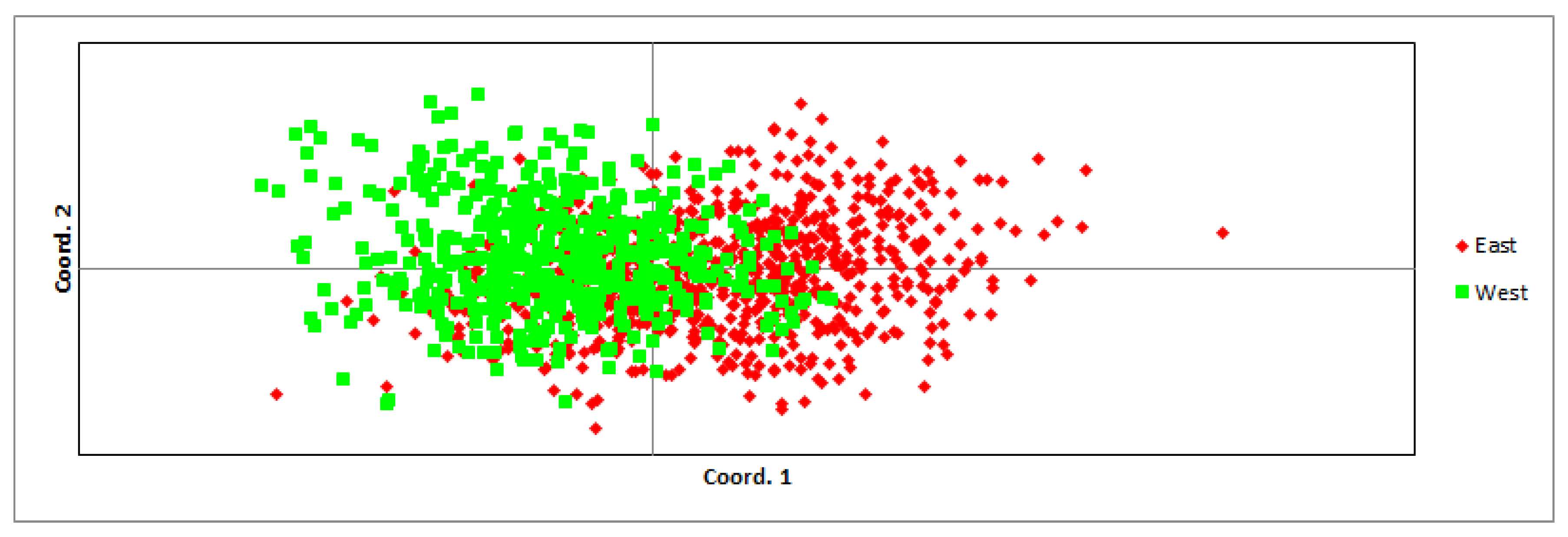
3.1. Population’s Genetic Characteristics
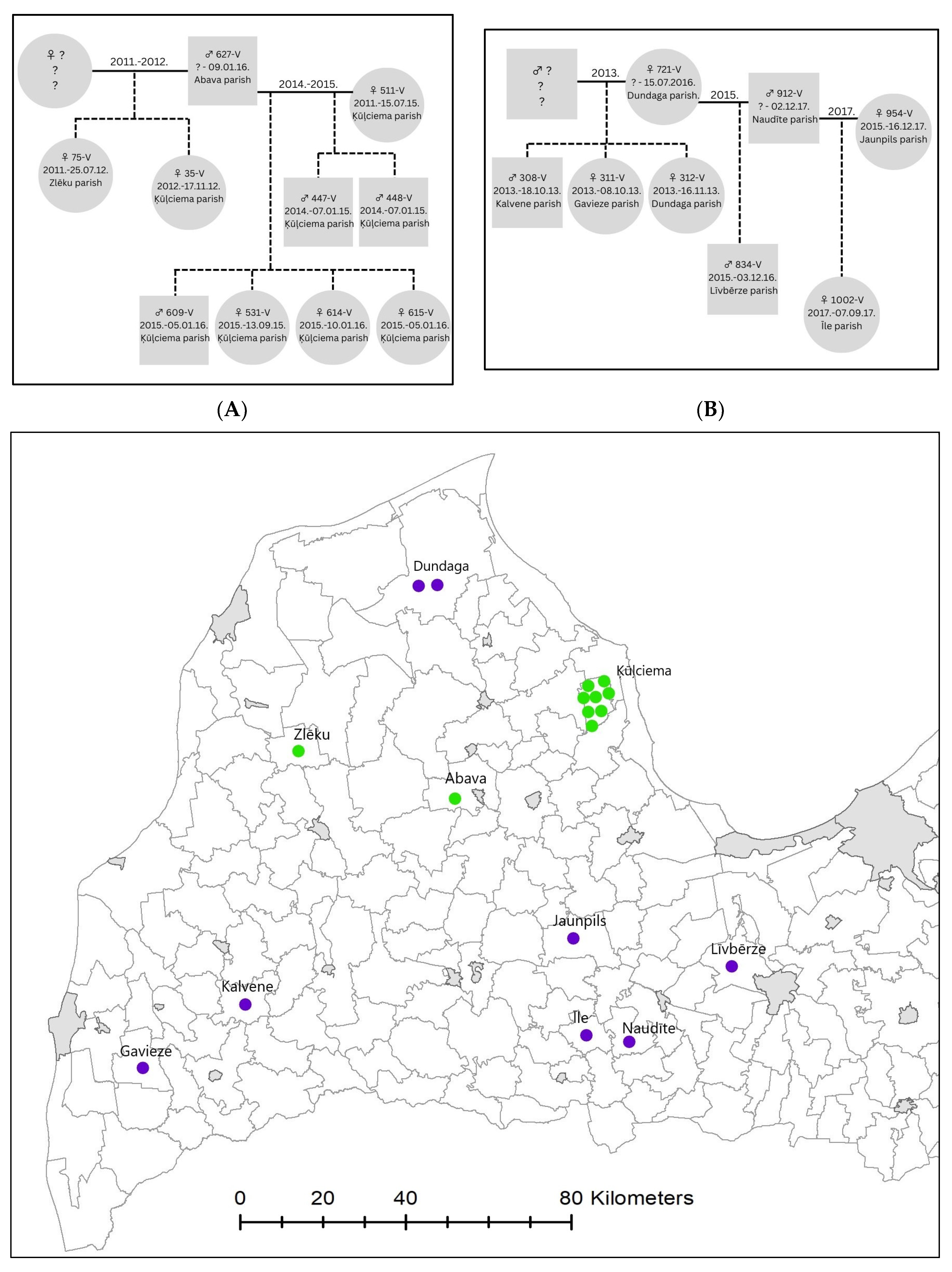
3.2. Population Kinship Structure and Pack Dynamics
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Taberlet, P.; Luikart, G.; Geffen, E. New methods for obtaining and analyzing genetic data from free-ranging carnivores. In Carnivore Conservation; Gittleman, J.L., Funk, S.M., Macdonald, D.W., Wayne, R.K., Eds.; Cambridge University Press: Cambridge, UK, 2001; pp. 313–334. [Google Scholar]
- Mills, S.L. Conservation of Wildlife Populations: Demography, Genetics, and Management; Wiley-Blackwell Publishing: Malden, MA, USA, 2007. [Google Scholar]
- Caniglia, R.; Fabbri, E.; Galaverni, M.; Milanesi, P.; Randi, E. Noninvasive sampling and genetic variability, pack structure, and dynamics in an expanding wolf population. J. Mammal. 2014, 95, 41–59. [Google Scholar] [CrossRef]
- Reinhardt, I.; Kluth, G.; Nowak, S.; Mysłajek, R.W. Standards for the Monitoring of the Central European Wolf Population in Germany and Poland; Bundesamt für Naturschutz: Bonn, Germany, 2015. [Google Scholar]
- Waser, P.M.; Strobeck, C.; Paetkau, D. Estimating interpopular dispersal rates. In Carnivore Conservation; Gittleman, J.L., Funk, S.M., Macdonald, D.W., Wayne, R.K., Eds.; Cambridge University Press: Cambridge, UK, 2001; pp. 484–497. [Google Scholar]
- Lucchini, V.; Fabbri, E.; Marucco, F.; Ricci, S.; Boitani, L.; Randi, E. Noninvasive molecular tracking of colonizing wolf (Canis lupus) packs in the western Italian Alps. Mol. Ecol. 2002, 11, 857–868. [Google Scholar] [CrossRef] [PubMed]
- Valière, N.; Fumagalli, L.; Gielly, L.; Miquel, C.; Lequette, B.; Poulle, M.L.; Weber, J.; Arlettaz, R.; Taberlet, P. Long-distance wolf recolonization of France and Switzerland inferred from non-invasive genetic sampling over a period of 10 years. Anim. Conserv. 2003, 6, 83–92. [Google Scholar] [CrossRef]
- Andersen, L.W.; Harms, V.; Caniglia, R.; Czarnomska, S.D.; Fabbri, E.; Jędrzejewska, B.; Kluth, G.; Madsen, A.B.; Nowak, C.; Pertoldi, C.; et al. Long-distance dispersal of a wolf, Canis lupus, in northwestern Europe. Mammal Res. 2015, 60, 163–168. [Google Scholar] [CrossRef]
- Liberg, O.; Aronson, A.; Sand, H.; Wabakken, P.; Maartmann, E.; Svensson, L.; Åkesson, M. Monitoring of wolves in Scandinavia. Hystrix Ital. J. Mammal. 2012, 23, 29–34. [Google Scholar] [CrossRef]
- Wayne, R.K.; Vilà, C. Molecular genetic studies of wolves. In Wolves: Behavior, Ecology and Conservation; Mech, L.D., Boitani, L., Eds.; The University of Chicago Press: Chicago, IL, USA, 2003; pp. 218–238. [Google Scholar]
- Rutledge, L.Y.; Patterson, B.R.; Mills, K.J.; Loveless, K.M.; Murray, D.L.; White, B.N. Protection from harvesting restores the natural social structure of eastern wolf packs. Biol. Conserv. 2010, 143, 332–339. [Google Scholar] [CrossRef]
- Rick, J.A.; Moen, R.A.; Erb, J.D.; Strasburg, J.L. Population structure and gene flow in a newly harvested gray wolf (Canis lupus) population. Conserv. Genet. 2017, 18, 1091–1104. [Google Scholar] [CrossRef]
- Mysłajek, R.W.; Tracz, M.; Tracz, M.; Tomczak, P.; Szewczyk, M.; Niedźwiecka, N.; Nowak, S. Spatial organization in wolves Canis lupus recolonizing north-west Poland: Large territories at low population density. Mamm. Biol. 2018, 92, 37–44. [Google Scholar] [CrossRef]
- Pilot, M.; Greco, C.; Von Holdt, B.M.; Jedrzejewska, B.; Randi, E.; Jedrzejewski, W.; Sidorovich, V.E.; Ostrander, E.A.; Wayne, R.K. Genome-wide signatures of population bottlenecks and diversifying selection in European wolves. Heredity 2014, 112, 428–442. [Google Scholar] [CrossRef] [PubMed]
- Szewczyk, M.; Nowak, S.; Niedźwiecka, N.; Hulva, P.; Špinkytė-Bačkaitienė, R.; Demjanovičová, K.; Bolfíková, B.Č.; Antal, V.; Fenchuk, V.; Tomczak, M.P.; et al. Dynamic range expansion leads to establishment of a new, genetically distinct wolf population in Central Europe. Sci. Rep. 2019, 9, 19003. [Google Scholar] [CrossRef]
- Wayne, R.K.; Brown, D.M. Hybridization and conservation of carnivores. In Carnivore Conservation; Gittleman, J.L., Funk, S.M., Macdonald, D.W., Wayne, R.K., Eds.; Cambridge University Press: Cambridge, UK, 2001; pp. 145–162. [Google Scholar]
- Andersone, Z.; Lucchini, V.; Randi, E.; Ozolins, J. Hybridisation between wolves and dogs in Latvia as documented using mitochondrial and microsatellite DNA markers. Mammal. Biol. 2002, 67, 79–90. [Google Scholar] [CrossRef]
- Hindrikson, M.; Männil, P.; Ozolins, J.; Krzywinski, A.; Saarma, U. Bucking the Trend in Wolf-Dog Hybridization: First Evidence from Europe of Hybridization between Female Dogs and Male Wolves. PLoS ONE 2012, 7, e46465. [Google Scholar] [CrossRef] [PubMed]
- Stenglein, J.L.; Waits, L.P.; Ausband, D.E.; Zager, P.; Mack, C.M. Estimating gray wolf pack size and family relationships using noninvasive genetic sampling at rendezvous sites. J. Mammal. 2011, 92, 784–795. [Google Scholar] [CrossRef]
- Galaverni, M.; Palumbo, D.; Fabbri, E.; Caniglia, R.; Greco, C.; Randi, E. Monitoring wolves (Canis lupus) by non-invasive genetics and camera trapping: A small-scale pilot study. Eur. J. Wildl. Res. 2012, 58, 47–58. [Google Scholar] [CrossRef]
- Ellegren, H. Inbreeding and relatedness in Scandinavian grey wolves Canis lupus. Hereditas 1999, 130, 239–244. [Google Scholar] [CrossRef]
- Gomerčić, T.; Sindičić, M.; Galov, A.; Arbanasić, H.; Kusak, J.; Kocijan, I.; Gomerčić, M.; Huber, Ð. High genetic variability of the grey wolf (Canis lupus L.) population from Croatia as revealed by mitochondrial DNA control region sequences. Zool. Stud. 2010, 49, 816–823. [Google Scholar]
- Shakarashvili, M.; Kopaliani, N.; Gurielidze, Z.; Dekanoidze, D.; Ninua, L.; Tarkhnishvili, D. Population genetic structure and dispersal patterns of grey wolf (Canis lupus) and golden jackal (Canis aureus) in Georgia, the Caucasus. J. Zool. 2020, 312, 227–238. [Google Scholar] [CrossRef]
- VonHoldt, B.M.; Stahler, D.R.; Smith, D.W.; Earl, D.A.; Pollinger, J.P.; Wayne, R.K. The genealogy and genetic viability of reintroduced Yellowstone grey wolves. Mol. Ecol. 2008, 17, 252–274. [Google Scholar] [CrossRef] [PubMed]
- Pilot, M.; Jedrzejewski, W.; Branicki, W.; Sidorovich, V.E.; Jedrzejewska, B.; Stachura, K.; Funk, S. Ecological factors influence population genetic structure of European grey wolves. Mol. Ecol. 2006, 15, 4533–4553. [Google Scholar] [CrossRef]
- Hindrikson, M.; Remm, J.; Männil, P.; Ozolins, J.; Tammeleht, E.; Saarma, U. Spatial Genetic Analyses Reveal Cryptic Population Structure and Migration Patterns in a Continuously Harvested Grey Wolf (Canis lupus) Population in North-Eastern Europe. PLoS ONE 2013, 8, e75765. [Google Scholar] [CrossRef]
- Stronen, A.V.; Jędrzejewska, B.; Pertoldi, C.; Demontis, D.; Randi, E.; Niedziałkowska, M.; Pilot, M.; Sidorovich, V.E.; Dykyy, I.; Kusak, J.; et al. North-South Differentiation and a Region of High Diversity in European Wolves (Canis lupus). PLoS ONE 2013, 8, e76454. [Google Scholar] [CrossRef] [PubMed]
- Ozoliņš, J.; Andersone, Ž. Management Plan for Wolf (Canis lupus) in Latvia; LSFRI Silava: Salaspils, Latvia, 2002. [Google Scholar]
- Ozoliņš, J.; Žunna, A.; Ornicāns, A.; Done, G.; Stepanova, A.; Pilāte, D.; Šuba, J.; Lūkins, M.; Howlett, S.J.; Bagrade, G. Action Plan for Grey Wolf Canis lupus Conservation and Management; LSFRI Silava: Salaspils, Latvia, 2017. [Google Scholar]
- Šuba, J.; Žunna, A.; Bagrade, G.; Done, G.; Lūkins, M.; Ornicāns, A.; Pilāte, D.; Stepanova, A.; Ozoliņš, J. Closer to Carrying Capacity: Analysis of the Internal Demographic Structure Associated with the Management and Density Dependence of a Controlled Wolf Population in Latvia. Sustainability 2021, 13, 9783. [Google Scholar] [CrossRef]
- Boitani, L.; Kaczensky, P.; Alvares, F.; Andrén, H.; Balys, V.; Blanco, J.C.; Chapron, G.; Chiriac, S.; Cirovic, D.; Drouet-Houguet, N.; et al. Assessment of the Conservation Status of the Wolf (Canis lupus) in Europe; Council of Europe Publishing: Strasbourg, France, 2022. [Google Scholar]
- Anonymous. Wolf (Canis lupus) Protection Plan. Environment Ministry of the Republic of Lithuania, Vilnius, 2014. Available online: http://www.vilkai.lt/wp-content/uploads/LTU_Wolf_Protection_Plan_2014_en.pdf (accessed on 12 July 2023).
- Remm, J.; Hindrikson, M. Estonian Conservation and Management Plan of Large Carnivores 2022–2031. Environmental Board, Pärnu, Estonia, 2022. Available online: https://keskkonnaamet.ee/en/news/environmental-board-approved-new-action-plan-protection-and-management-large-carnivores-ten (accessed on 20 June 2023).
- Jędrzejewski, W.; Jędrzejewska, B.; Andersone-Lilley, Ž.; Balčiauskas, L.; Männil, P.; Ozoliņš, J.; Sidorovich, V.E.; Bagrade, G.; Kübarsepp, M.; Ornicāns, A.; et al. Synthesizing wolf ecology and management in Eastern Europe: Similarities and contrasts with North America. In The World of Wolves: New Perspectives on Ecology, Behaviour and Management; Musiani, M., Boitani, L., Eds.; University of Calgary Press: Calgary, AB, Canada, 2010; pp. 207–233. [Google Scholar]
- Ballard, W.B.; Whitman, J.S.; Gardner, C.L. Ecology of an Exploited Wolf Population in South-Central Alaska. Wildl. Monogr. 1987, 98, 3–54. [Google Scholar]
- Fuller, T.K.; Mech, L.D.; Cohraine, J.F. Wolf population dynamics. In Wolves: Behavior, Ecology and Conservation; Mech, L.D., Boitani, L., Eds.; The University of Chicago Press: Chicago, IL, USA, 2003; pp. 161–191. [Google Scholar]
- Adams, L.G.; Stephenson, R.O.; Dale, B.W.; Ahgook, R.T.; Demma, D.J. Population dynamics and harvest characteristics of wolves in the Central Brooks Range, Alaska. Wildl. Monogr. 2008, 170, 1–25. [Google Scholar] [CrossRef]
- Creel, S.; Rotella, J.J. Meta-Analysis of Relationships between Human Offtake, Total Mortality and Population Dynamics of Gray Wolves (Canis lupus). PLoS ONE 2010, 5, e12918. [Google Scholar] [CrossRef]
- Haber, G.C. Biological, Conservation, and Ethical Implications of Exploiting and Controlling Wolves. Conserv. Biol. 1996, 10, 1068–1081. [Google Scholar] [CrossRef]
- Johnson, W.E.; Eizirik, E.; Lento, G.M. The control, exploitation, and conservation of carnivores. In Carnivore Conservation; Gittleman, J.L., Funk, S.M., Macdonald, D.W., Wayne, R.K., Eds.; Cambridge University Press: Cambridge, UK, 2001; pp. 196–219. [Google Scholar]
- Mech, L.D. The Wolf: The Ecology and Behaviour of an Endangered Species; University of Minnesota Press: Minneapolis, MN, USA; London, UK, 1970. [Google Scholar]
- Mech, L.D.; Boitani, L. Wolf Social Ecology. In Wolves: Behavior, Ecology and Conservation; Mech, L.D., Boitani, L., Eds.; The University of Chicago Press: Chicago, IL, USA, 2003; pp. 1–34. [Google Scholar]
- Silk, J.B. The adaptive value of sociality in mammalian groups. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2007, 362, 539–559. [Google Scholar] [CrossRef]
- Cassidy, K.A.; MacNulty, D.R.; Stahler, D.R.; Smith, D.W.; Mech, L.D. Group composition effects on aggressive inter-pack interactions of gray wolves in Yellowstone National Park. Behav. Ecol. 2015, 26, 1352–1360. [Google Scholar] [CrossRef]
- Cassidy, K.A.; McIntyre, R.T. Do gray wolves (Canis lupus) support pack mates during aggressive inter-pack interactions? Anim. Cognit. 2016, 19, 939–947. [Google Scholar] [CrossRef]
- Jędrzejewska, B.; Jędrzejewski, W.; Bunevich, A.N.; Miłkowski, L.; Okarma, H. Population Dynamics of Wolves Canis lupus in Białowieża Primeval Forest (Poland and Belarus) in relation to hunting by humans, 1847–1993. Mammal Rev. 1996, 26, 103–126. [Google Scholar] [CrossRef]
- Brainerd, S.M.; Andrén, H.; Bangs, E.E.; Bradley, E.H.; Fontaine, J.A.; Hall, W.; Iliopoulos, Y.; Jimenez, M.D.; Jozwiak, E.A.; Liberg, O. The effects of breeder loss on wolves. J. Wildl. Manag. 2008, 72, 89–98. [Google Scholar] [CrossRef]
- Ausband, D.E.; Mitchell, M.S.; Waits, L.P. Effects of breeder turnover and harvest on group composition and recruitment in a social carnivore. J. Anim. Ecol. 2017, 86, 1094–1101. [Google Scholar] [CrossRef] [PubMed]
- Wallach, A.D.; Ritchie, E.G.; Read, J.; O’Neill, A.J. More than Mere Numbers: The Impact of Lethal Control on the Social Stability of a Top-Order Predator. PLoS ONE 2009, 4, e6861. [Google Scholar] [CrossRef] [PubMed]
- Jędrzejewski, W.; Branicki, W.; Veit, C.; MeĐugorac, I.; Pilot, M.; Bunevich, A.N.; Jędrzejewska, B.; Schmidt, K.; Theuerkauf, J.; Okarma, H.; et al. Genetic diversity and relatedness within packs in an intensely hunted population of wolves Canis lupus. Acta Theriol. 2005, 50, 3–22. [Google Scholar] [CrossRef]
- Frank, L.G.; Woodroffe, R. Behaviour of carnivores in exploited and controlled populations. In Carnivore Conservation; Gittleman, J.L., Funk, S.M., Macdonald, D.W., Wayne, R.K., Eds.; Cambridge University Press: Cambridge, UK, 2001; pp. 419–442. [Google Scholar]
- Allendorf, F.W.; England, P.R.; Luikart, G.; Ritchie, P.A.; Ryman, N. Genetic effects of harvest on wild animal populations. Trends Ecol. Evol. 2008, 23, 327–337. [Google Scholar] [CrossRef]
- Funk, S.M.; Fiorello, C.V.; Cleaveland, S.; Gompper, M.E. The role of disease in carnivore ecology and conservation. In Carnivore Conservation; Gittleman, J.L., Funk, S.M., Macdonald, D.W., Wayne, R.K., Eds.; Cambridge University Press: Cambridge, UK, 2001; pp. 443–466. [Google Scholar]
- Ausband, D.E.; Waits, L. Does harvest affect genetic diversity in grey wolves? Mol. Ecol. 2020, 29, 3187–3195. [Google Scholar] [CrossRef]
- Linnell, J.; Salvatori, V.; Boitani, L. Guidelines for Population Level Management Plans for Large Carnivores; LCIE Report Prepared for the European Commission (Contract nr. 070501/2005/424162/MAR/B2); Large Carnivore Initiative for Europe; IUCN/SSC Working Group: Rome, Italy, 2008. [Google Scholar]
- Bassing, S.B.; Ausband, D.E.; Mitchell, M.S.; Lukacs, P.; Keever, A.; Hale, G.; Waits, L. Stable pack abundance and distribution in a harvested wolf population. J. Wildl. Manag. 2019, 83, 577–590. [Google Scholar] [CrossRef]
- Haig, S.M.; Ballou, J.D. Pedigree Analyses in Wild Populations. In Population Viability Analysis; Beissinger, S.R., McCullough, D.R., Eds.; The University of Chicago Press: Chicago, IL, USA; London, UK, 2002; pp. 388–405. [Google Scholar]
- Klevezal, G.A. Age-Related Structures in Zoological Studies of Mammals; Nauka: Moscow, Russia, 1988. (In Russian) [Google Scholar]
- Francisco, L.V.; Langsten, A.A.; Mellersh, C.S.; Neal, C.L.; Ostrander, E.A. A class of highly polymorphic tetranucleotide repeats for canine genetic mapping. Mamm. Genome 1996, 7, 359–362. [Google Scholar] [CrossRef]
- Shibuya, H.; Collins, B.K.; Huang, T.M.; Johnson, G.S. A polymorphic (AGGAAT), tandem repeat in an intron of the canine von Willebrand factor gene. Anim. Genet. 1994, 25, 122. [Google Scholar] [CrossRef]
- Holmes, N.G.; Dickens, H.F.; Parker, H.L.; Binns, M.M.; Mellersh, C.S.; Sampson, J. Eighteen canine microsatellites. Anim. Genet. 1995, 26, 132–133. [Google Scholar] [CrossRef]
- Fredholm, M.; Winterø, A.K. Variation of short tandem repeats within and between species belonging to the Canidae family. Mamm. Genome 1995, 6, 11–18. [Google Scholar] [CrossRef] [PubMed]
- Ostrander, E.A.; Mapa, F.A.; Yee, M.; Rine, J. One hundred and one new simple sequence repeat-based markers for the canine genome. Mamm. Genome 1995, 6, 192–195. [Google Scholar] [CrossRef]
- Peakall, R.; Smouse, P.E. GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 2012, 28, 2537–2539. [Google Scholar] [CrossRef] [PubMed]
- Queller, D.C.; Goodnight, K.F. Estimating relatedness using genetic markers. Evolution 1989, 43, 258–275. [Google Scholar] [CrossRef] [PubMed]
- Goudet, J.F. FSTAT (version 1.2): A computer program to calculate F-statistics. J. Hered. 1995, 86, 485–486. [Google Scholar] [CrossRef]
- Jones, O.R.; Wang, J. COLONY: A program for parentage and sibship inference from multilocus genotype data. Mol. Ecol. Resour. 2010, 10, 551–555. [Google Scholar] [CrossRef]
- Kalinowski, S.T.; Taper, M.L.; Marshall, T.C. Revising how the computer program Cervus accommodates genotyping error increases success in paternity assignment. Mol. Ecol. 2007, 16, 1099–1106. [Google Scholar] [CrossRef]
- Ozoliņš, J.; Stepanova, A.; Žunna, A.; Bagrade, G.; Ornicāns, A. Wolf hunting in Latvia in the light of population continuity in the Baltics. In Beiträge zur Jagd- und Wildforschung; Stubbe, M., Ed.; Band 36; Gesellschaft für Wildtier- und Jagdforschung e.V.: Halle/Saale, Germany, 2011; pp. 93–104. [Google Scholar]
- Baltrūnaité, L.; Balčiauskas, L.; Åkesson, M. The genetic structure of the Lithuanian wolf population. Centr. Eur. J. Biol. 2013, 8, 440–447. [Google Scholar] [CrossRef]
- Hindrikson, M.; Remm, J.; Pilot, M.; Godinho, R.; Stronen, A.V.; Baltrūnaité, L.; Czarnomska, S.D.; Leonard, J.A.; Randi, E.; Nowak, C.; et al. Wolf population genetics in Europe: A systematic review, meta-analysis and suggestions for conservation and management. Biol. Rev. 2017, 92, 1601–1629. [Google Scholar] [CrossRef]
- Palsbøll, P.J.; Peery, M.Z.; Bérubé, M. Detecting populations in the ‘ambiguous’ zone: Kinship-based estimation of population structure at low genetic divergence. Mol. Ecol. Resour. 2010, 10, 797–805. [Google Scholar] [CrossRef]
- Lehman, N.; Clarkson, P.; Mech, L.D.; Meier, T.J.; Wayne, R.K. A study of the genetic relationships within and among wolf packs using DNA fingerprinting and mitochondrial DNA. Behav. Ecol. Sociobiol. 1992, 30, 83–94. [Google Scholar] [CrossRef]
- Packard, J.M. Wolf behavior: Reproductive, social, and intelligent. In Wolves: Behavior, Ecology and Conservation; Mech, L.D., Boitani, L., Eds.; The University of Chicago Press: Chicago, IL, USA, 2003; pp. 35–65. [Google Scholar]
- Eklund, A.; López-Bao, J.V.; Tourani, M.; Chapron, G.; Frank, J. Limited evidence on the effectiveness of interventions to reduce livestock predation by large carnivores. Sci. Rep. 2017, 7, 2097. [Google Scholar] [CrossRef]
- Balčiauskas, L.; Balčiauskienė, L.; Litvaitis, J.A.; Tijušas, E. Adaptive monitoring: Using citizen scientists to track wolf populations when winter-track counts become unreliable. Wildl. Res. 2020, 48, 76–85. [Google Scholar] [CrossRef]
- Hayes, R.D.; Harestad, A.S. Demography of a recovering wolf population in the Yukon. Can. J. Zool. 2000, 78, 36–48. [Google Scholar] [CrossRef]
- Fuller, T.K.; Sievert, P.R. Carnivore demography and the consequences of changes in prey availability. In Carnivore Conservation; Gittleman, J.L., Funk, S.M., Macdonald, D.W., Wayne, R.K., Eds.; Cambridge University Press: Cambridge, UK, 2001; pp. 163–178. [Google Scholar]
- Kojola, I. Status and development of the wolf population in Finland. In Management Plan for the Wolf Population in Finland; Ministry of Agriculture and Forestry: Helsinki, Finland, 2005; pp. 8–14. [Google Scholar]
- Mech, L.D.; Barber-Meyer, S.M.; Erb, J. Wolf (Canis lupus) Generation Time and Proportion of Current Breeding Females by Age. PLoS ONE 2016, 11, e0156682. [Google Scholar] [CrossRef] [PubMed]
- Borg, B.L.; Brainerd, S.M.; Meier, T.J.; Prugh, L.R. Impacts of breeder loss on social structure, reproduction and population growth in a social canid. J. Anim. Ecol. 2015, 84, 177–187. [Google Scholar] [CrossRef] [PubMed]
- Žunna, A.; Bagrade, G.; Ozoliņš, J. Attitudes of the General Public and Hunters Towards Wolves in Latvia; Its Predictors and Changes Over Time. Proc. Latv. Acad. Sci. Sect. B 2020, 74, 280–286. [Google Scholar] [CrossRef]
- Šuba, J.; Žunna, A.; Bagrade, G.; Done, G.; Ornicāns, A.; Pilāte, D.; Stepanova, A.; Ozoliņš, J. Does Wolf Management in Latvia Decrease Livestock Depredation? An Analysis of Available Data. Sustainability 2023, 15, 8509. [Google Scholar] [CrossRef]
| Group Duration (Years) | 11 | 10 | 9 | 8 | 7 | 6 | 5 | 4 | 3 | 2 | 1 |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Proportion in the sample of kinship analyses (%) | 0.6 | 1.7 | 1.7 | 4.4 | 2.2 | 3.9 | 5.0 | 10.0 | 8.9 | 15.6 | 46.1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Žunna, A.; Ruņģis, D.E.; Ozoliņš, J.; Stepanova, A.; Done, G. Genetic Monitoring of Grey Wolves in Latvia Shows Adverse Reproductive and Social Consequences of Hunting. Biology 2023, 12, 1255. https://doi.org/10.3390/biology12091255
Žunna A, Ruņģis DE, Ozoliņš J, Stepanova A, Done G. Genetic Monitoring of Grey Wolves in Latvia Shows Adverse Reproductive and Social Consequences of Hunting. Biology. 2023; 12(9):1255. https://doi.org/10.3390/biology12091255
Chicago/Turabian StyleŽunna, Agrita, Dainis Edgars Ruņģis, Jānis Ozoliņš, Alda Stepanova, and Gundega Done. 2023. "Genetic Monitoring of Grey Wolves in Latvia Shows Adverse Reproductive and Social Consequences of Hunting" Biology 12, no. 9: 1255. https://doi.org/10.3390/biology12091255
APA StyleŽunna, A., Ruņģis, D. E., Ozoliņš, J., Stepanova, A., & Done, G. (2023). Genetic Monitoring of Grey Wolves in Latvia Shows Adverse Reproductive and Social Consequences of Hunting. Biology, 12(9), 1255. https://doi.org/10.3390/biology12091255






