Simple Summary
This study represents the most comprehensive phylogenetic reconstruction of the plant subtribe Ditaxinae and related taxa within Acalyphoideae (Euphorbiaceae). The taxonomy of this group, mainly based in morphology, has long been controversial. Here, we present a new taxonomic classification at the genus and tribe ranks using a solid phylogenetic framework. We also provide key morphological synapomorphies supporting the main recovered clades.
Abstract
The subtribe Ditaxinae in the plant family Euphorbiaceae is composed of five genera (Argythamnia, Caperonia, Chiropetalum, Ditaxis and Philyra) and approximately 120 species of perennial herbs (rarely annual) to treelets. The subtribe is distributed throughout the Americas, with the exception of Caperonia, which also occurs in tropical Africa and Madagascar. Under the current classification, Ditaxinae includes genera with a questionable morphology-based taxonomy, especially Argythamnia, Chiropetalum and Ditaxis. Moreover, phylogenetic relationships among genera are largely unexplored, with previous works sampling <10% of taxa, showing Ditaxinae as paraphyletic. In this study, we inferred the phylogenetic relationships within Ditaxinae and related taxa using a dataset of nuclear (ETS, ITS) and plastid (petD, trnLF, trnTL) DNA sequences and a wide taxon sampling (60%). We confirmed the paraphyly of Ditaxinae and Ditaxis, both with high support. Following our phylogenetic results, we combined Ditaxis in Argythamnia and upgraded Ditaxinae to the tribe level (Ditaxeae). We also established and described the tribe Caperonieae based on Caperonia, and transferred Philyra to the tribe Adelieae, along with Adelia, Garciadelia, Lasiocroton and Leucocroton. Finally, we discuss the main morphological synapomorphies for the genera and tribes and provide a taxonomic treatment, including all species recognized under each genus.
Keywords:
Adelieae; Argythamnia; Caperonia; Caperonieae; Chiropetalum; Ditaxeae; Ditaxis; phylogenetics; Philyra 1. Introduction
The systematics of Euphorbiaceae Juss. have undergone substantial changes in the last two decades stemming from studies in molecular systematics. The family is currently classified into four subfamilies (Acalyphoideae Beilschmied, Cheilosoideae K.Wurdack & Petra Hoffm., Crotonoideae Beilschmied and Euphorbioideae) [1,2,3,4,5]. Phylogenetic studies have led to updates in the systematics of Euphorbiaceae, where two biovulate subfamilies were segregated and elevated to the family level, Phyllanthaceae Martinov, Picrodendraceae Small and Putranjivaceae Endl. [2,6]. Acalyphoideae was recognized as a subfamily in 1975 [7] and currently comprises 14 tribes, 23 subtribes, 99 genera and approximately 1860 species [5,8]. The subfamily is distributed worldwide, except in polar regions, with greater diversity in tropical and subtropical areas [5,9,10,11].
In the classification proposed by Webster [5], the tribe Chrozophoreae (Müll.Arg.) Pax & K.Hoffm was composed of the subtribes Ditaxinae Griseb., Speranskiinae G.L.Webster and Chrozophorinae (Müll.Arg.) Pax & K.Hoffm. Ditaxinae was proposed in 1859 [12], with the genera Argythamnia P.Browne (=Chiropetalum A.Juss.), Caperonia A.St.-Hil. and Ditaxis Vahl ex A.Juss. Later, Müller [13] presented a classification for the tribe Acalypheae that consisted of 11 subtribes, including Chrozophorinae Müll.Arg. (containing Argythamnia, Chiropetalum, Ditaxis and Philyra Klotzsch) and Caperoniinae Müll.Arg. (including only Caperonia). Müller differentiated these subtribes based on the staminate flowers having a rudimentary ovary present at the apex of the staminal column in Caperonia and absent in the other genera.
In 1912, a new classification system called “Chrozophorinarum’’ was put forward by Pax and Hoffmann, wherein Ditaxinae was treated as a synonym of Chrozophorinae-regularis, which had been circumscribed with the genera Aonikena Speg., Argythamnia, Caperonia, Chiropetalum, Chrozophora Neck. ex A.Juss., Ditaxis and Philyra [14]. Webster re-established Ditaxinae to include Argythamnia, Caperonia, Chiropetalum (= Aonikena), Ditaxis and Philyra [7].
In its current circumscription, Ditaxinae consists of five genera (Argythamnia, Caperonia, Chiropetalum, Ditaxis and Philyra) and around 120 species of herbs, subshrubs, shrubs, and small trees, widely distributed in the New World (all genera) [5,15,16,17,18,19,20,21] and continental Africa and Madagascar (only Caperonia) [5,17,21].
Caperonia has approximately 35 herbaceous and subshrub species, of which 29 are distributed in the New World and seven in Africa/Madagascar. In the New World, it occurs from Mexico to central Argentina, with one species introduced in the southern United States. Caperonia has its greatest diversity in tropical and subtropical regions, and is the only genus of the tribe occurring in the Amazonian region, exclusively in marshy environments [5,14,17,21]. Argythamnia is composed of 19 species of perennial herbs, subshrubs to shrubs distributed in Central America (Caribbean and Mexico), where it is restricted to seasonally dry tropical forests and coastal vegetation [5,22]. Chiropetalum consists of 21 species of herbs and subshrubs and is disjunct between Mexico (two species) and South America (19 species), where it occurs from Peru to Patagonia, with its highest diversity in northern Argentina and southern Brazil [15,20,23]. Chiropetalum occurs in a variety of habitats, including a range of dry and humid forests, arid environments, grasslands and coastal vegetation. Ditaxis is the most species-rich genus of the subtribe with approximately 45 species, ranging from herbs to shrubs, and is widely distributed from the southern United States to northern Patagonia, in Argentina [18,24,25,26]. Ditaxis occupies various habitats, such as deserts and grasslands, but most species occur in seasonally dry tropical forests [18,25,26]. Finally, the monotypic genus Philyra (P. brasiliensis Klotzsch) is a shrub or a small tree. The genus is restricted to central and eastern South America, growing exclusively in seasonally dry tropical forests [5,19].
Argythamnia, Chiropetalum and Ditaxis form a group of great morphological complexity that has undergone many taxonomic changes. However, few studies have approached the three genera all together to understand their phylogenetic relationships [13,14,22,23,27]. Argythamnia, Chiropetalum and Ditaxis have sometimes been treated as subgenera of Argythamnia s.l. [22,23,24,27]. Currently, these taxa are treated at the genus level, but there is still disagreement among taxonomists. Recent studies, using DNA sequence data, have attempted to resolve the relationship among Argythamnia, Chiropetalum and Ditaxis [15,28], but their phylogenetic analyses revealed topologies with low support in some clades, preventing any taxonomic changes or updates. Similarly, two other phylogenetic studies have included terminals of Ditaxinae, but these did not exceed 10% of taxon sampling and yielded low-resolution phylogenies [3,29].
Cervantes and collaborators reconstructed the biogeographic history of Acalyphoideae based on a molecular phylogenetic analysis using the petD, trnL-F and matK/trnK genetic regions [30]. Ditaxinae, even though represented by ~10% of the species, emerged as paraphyletic. Their results recovered Philyra as a sister to the tribe Adelieae G.L.Webster, and this clade was a sister to the Argythamnia + Chiropetalum + Ditaxis clade, as shown by Jestrow [29,31]. Caperonia emerged as a sister to all above taxa, albeit with low support [30].
Given the need for a solid phylogenetic and systematic framework for the subtribe Ditaxinae, in this study we established the following aims: (1) test the monophyly of Ditaxinae and its currently recognized genera, Argythamnia, Caperonia, Chiropetalum, Ditaxis and Philyra, using a comprehensive taxonomic and geographical sampling, including multiple accessions per species when possible; (2) circumscribe the recovered clades and identify potential morphological synapomorphies; (3) establish a suprageneric classification in the subfamily Acalyphoideae based on the recovered phylogenetic pattern in this study.
2. Material and Methods
2.1. Taxon Sampling and Outgroup Selection
Our sampling covered all currently recognized genera of subtribe Ditaxinae: Argythamnia (11 spp., 61% of the total), Caperonia (10 spp., 30%), Chiropetalum (18 spp., 86%), Ditaxis (35 spp., 77%) and Philyra (1 sp., 100%). Thus, our dataset included a total of 75 species, representing 60% of Ditaxinae. We also included five representatives of tribe Adelieae, the latter based on Jestrow’s circumscription [29]. We used Acalypha lanceolata Willd., Enriquebeltrania crenatifolia (Miranda) Rzed., Bernardia dichotoma (Willd.) Müll.Arg., Plukenetia penninervia Müll.Arg., P. volubilis L. and Seidelia triandra (E.Mey.) Pax as outgroups, based on previous phylogenetic analyses [15,28,30]. Overall, our study sampled 86 species of the subfamily Acalyphoideae (Supplementary File S1: Table S1). The choice of outgroups also aimed to reconstruct the clades close to Ditaxinae following the study of Cervantes and collaborators [30]. The type species of each genus in Ditaxinae was sampled in our dataset. For the taxonomic treatment, type specimens were also analyzed to infer morphological similarities, mainly of taxa not represented in the phylogenetic analyses, in order to assess the preliminary generic assignment based on morphological similarities with the taxa represented in the phylogeny, as has been done in other complex groups within Euphorbiaceae [32,33,34].
We included samples collected in Africa, the Caribbean region, Central America, North America and South America. Plant tissues were preserved in silica gel, and vouchers were deposited in the herbaria BAA, FLOR, HUEFS, ICN, MA, MEXU, RB, SP, SPF and US (acronyms follow Thiers, continuously updated) [35]. Other tissue samples were obtained from herbarium specimens at BA, BAA, CA, CORD, CPAP, F, HUEFS, IEB, K, LPD, MA, MEXU, MO, MOL, RB, RSA, SI, SP, US and XAL (Supplementary File S1: Table S1). We also used 61 sequences (representing 22 species) from the US National Center for Biotechnology Information (NCBI) GenBank repository (https://www.ncbi.nlm.nih.gov/genbank). Voucher information and GenBank accession data are provided in Supplementary File S1: Table S1.
2.2. DNA Extraction, Amplification and Sequencing
DNA was extracted from silica-dried leaf tissue and herbarium material using the CTAB method [36] with some modifications [36] (see Supplementary File S2). The extracted DNA was quantified using a Qubit™ dsDNA BR Standard (Invitrogen). Samples with high concentrations (>20 ng/μL) were diluted (1:20, 1:50) depending on the concentration.
Three plastid (trnL-F, trnT-L, petD) and two nuclear (ITS, ETS) genetic regions were sequenced (see Supplementary File S2: Table S2). PCR amplifications were conducted with 25 μL reactions (for thermocycler temperature protocols, see Supplementary File S2: Table S3). Each reaction tube included MyTaq Red Mix (Bioline), H2O, primers and genomic DNA. For samples that were difficult to amplify, PuReTaq Ready-To-Go PCR Beads (GE Healthcare) were used. PCR products were purified with ExoSap PCR Purification and sent for sequencing at MACROGEN (Macrogen, Madrid, Spain), using the same amplification primers (Supplementary File S2: Table S3).
The pherograms were edited manually in UGENE [37] and automatically aligned with MUSCLE, using the default parameters. Manual adjustments were made to each alignment matrix in UGENE, employing the similarity criterion. A 120 bp region was excluded from the analysis of the trnT-L data matrix due to an uncertain homology assessment in the alignment.
2.3. Phylogenetic Analyses
Evolutionary models of nucleotide substitution were selected based on maximum likelihood (ML) using the Akaike (AIC) [38] information criterion implemented in jModelTest v.2.1.10 [39,40]. Each marker was analyzed individually, and the models were GTR + I + G for ETS and ITS, TVM + I + G for trnL-F, TPM1uf + G for trnT-L and GTR + G for petD. MrBayes does not allow implementing all of these models, and thus, we used the nearest and slightly more complex model, which was GTR + I + G for the nuclear regions and GTR + G for the plastid markers [41]. Bayesian inference (BI) appears to be more robust with respect to over-parametrization and more sensitive to infra-parametrization than the ML optimization used in jModelTest [42]. Each genetic region was analyzed individually based on BI and ML. Concatenated matrices with nuclear (ITS + ETS) and plastid markers (trnL-F + trnT-L + petD) were also analyzed separately to check for possible incongruences in the topology, and finally, a matrix with all markers was analyzed with BI and ML approaches. Topological incongruence between nuclear and plastid regions was defined as the presence of clades with a posterior probability (PP) ≥ 0.95 in IB and bootstrapping support (BS) ≥ 70% in ML [43]. In the combined analysis using only one terminal per species, we prioritized keeping the terminals with at least one nuclear and one plastid region. Bayesian analyses consisted of two independent Markov Chain Monte Carlo (MCMC) runs of 50 million generations in MrBayes v.3.1.2 [44], sampling every 1000th generation, with 20% (first 10 million trees) discarded as burn-in. Output files were summarized with TreeAnnotator v.1.6.1 [45], and the performance of each analysis (effective sample sizes, ESS > 200) was evaluated using Tracer v.1.6 [46]. Phylogenetic trees for individual and combined markers reconstructed with BI and ML are presented. Maximum-likelihood analyses were performed with RAxML [47] on the concatenated supermatrix, under a GTRGAMA model with 1000 bootstrap replicates. All analyses were hosted at CIPRES Science Gateway [48].
3. Results
The aligned DNA matrix combining the five regions (ETS, ITS, petD, trnT-L, trnL-F) was 3985 bp long and included 86 species (75 of Ditaxinae s.l.) and 223 terminals (there were species represented by more than one specimen and unidentified/unnamed specimens labeled as “sp.”). A summary of each data partition and combined matrices is provided in Table 1. The marker petD proved informative for the group. However, it was the region with the lowest taxonomic representation, as only recent tissue samples dried in silica gel could be amplified (Table 1). The analyses of the individual markers showed few cases of topological incongruences between the plastid and nuclear genome. However, in most cases, these incongruences did not have high support, and thus, the matrices (nuclear plus plastid datasets) were combined for the final analysis. Figure 1 represents the phylogenetic tree reconstructed when combining the five markers and the inclusion of one terminal per species. The phylogenetic analyses using all terminals (including multiple accessions) and individual and combined datasets are presented in Supplementary File S2: Figures S1–S9. The ML analysis did not show significant differences in tree topology when compared to the BI (Supplementary File S2: Figure S9).

Table 1.
Descriptive statistics of the separate and combined DNA datasets used in the phylogenetic analyses.
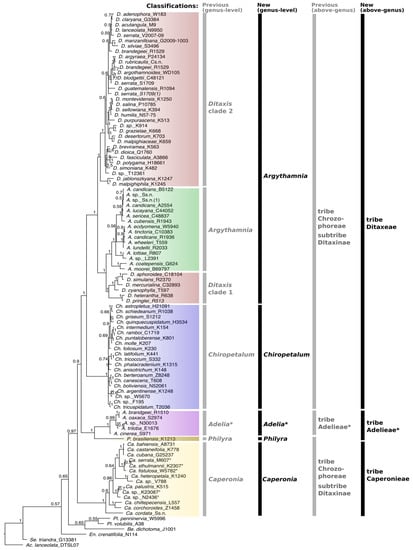
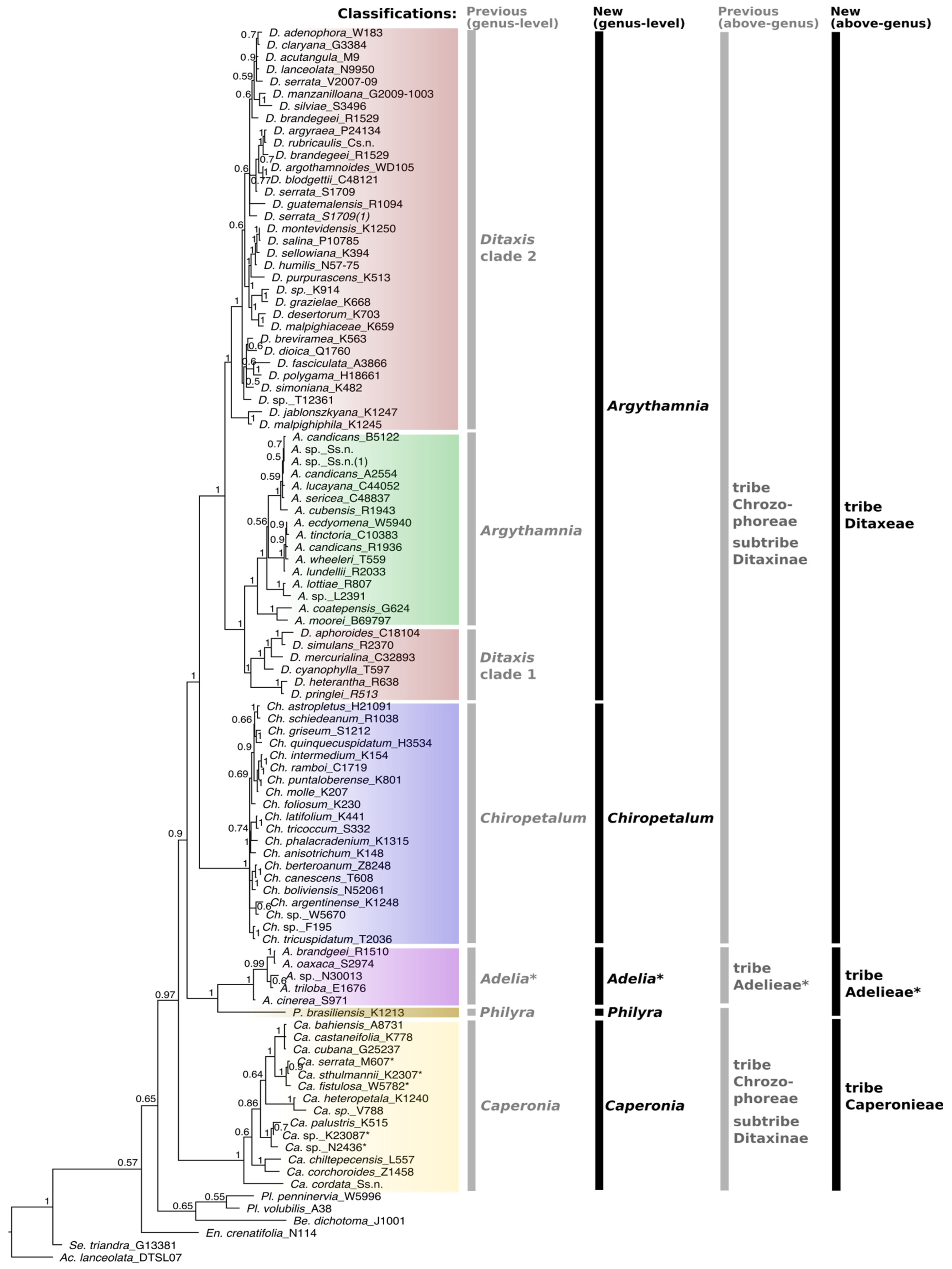
Figure 1.
Majority rule consensus tree of Ditaxinae (Euphorbiaceae) and related taxa based on the combined five markers (cpDNA [trnLF, trnTL, petD] and nDNA [ETS, ITS]) obtained through Bayesian inference. Bayesian posterior probabilities (PPs) are indicated on each branch. Vertical bars with labels on the right indicate the old (gray) [5] and the new (black) generic and suprageneric classifications. Asterisks in Adelia and Adelieae indicate that this clade is not fully represented here (several unsampled genera); we followed the classification proposed by Jestrow [29] (based on a complete generic sampling of Adelieae). To avoid confusion, some genera are abbreviated using two initial letters: Ac. = Acalypha, Be. = Bernardia, Ca. = Caperonia, Ch. = Chiropetalum, En. = Enriquebeltrania, Pl. = Plukeneria, Se. = Seidelia.
Despite minor incongruences between different reconstructions, the genera Argythamnia, Caperonia, Chiropetalum, Philyra and Adelia were confirmed as monophyletic, whereas Ditaxis was paraphyletic in all reconstructions (Figure 1 and Supplementary File S2: Figures S1–S8). In contrast, the phylogenetic trees obtained from the analyses of the combined and individual markers presented some incongruence regarding the positioning of Philyra, Adelia and Caperonia. In the analyses of the trnL-F and cpDNA-combined datasets, Philyra emerged as a sister (PP = 1) to the clade Caperonia + Adelia + Chiropetalum + Argythamnia + Ditaxis (Supplementary File S2: Figures S3 and S6), whereas in trnT-L, Philyra formed a polytomy with Adelia (Supplementary File S2: Figure S1). In petD and ETS, Adelia + Philyra was a sister of Caperonia + Chiropetalum + Argythamnia + Ditaxis with maximum support (Supplementary File S2: Figures S2 and S5). In the reconstructions based on ITS, ITS + ETS and the matrix with all markers combined, Philyra + Adelia emerged as a sister to Argythamnia + Chiropetalum + Ditaxis, while Caperonia emerged as a sister to the clade formed by all the five genera above (Figure 1 and Supplementary File S2: Figures S4 and S7–S9). Based on ETS only, Caperonia emerged as a sister to Argythamnia + Ditaxis, while Chiropetalum was recovered as a sister to Caperonia + Argythamnia + Ditaxis, both with low support (Supplementary File S2: Figure S5). In all other analyses, Chiropetalum emerged as a sister to Argythamnia + Ditaxis with high support (Figure 1 and Supplementary File S2: Figures S1–S4 and S6–S9). In all reconstructions, Ditaxis species were grouped into two clades (Ditaxis 1 and Ditaxis 2) separated by Argythamnia s.s. (Figure 1), leaving Ditaxis paraphyletic in its current circumscription. In ETS and ITS + ETS reconstructions (Supplementary File S2: Figures S5 and S7), the Andean species Ditaxis jablonszkyana Pax & K.Hoffm. and D. malpighipila (Hicken) L.C.Wheeler emerged as sisters to all other Ditaxis + Argythamnia species (PP = 1), whereas in all plastid reconstructions, these two species were recovered as sisters to clade Ditaxis 2 (PP = 1, Figure 1; Supplementary File S2: Figures S2–S4, S6 and S8–S9).
Phylogenetic trees generated from nuclear and plastid datasets, based on both BI and ML, supported the paraphyly of Ditaxinae as currently circumscribed (Figure 1 and Supplementary File S2: Figures S1–S8) due to the position of the representatives of the Adelieae tribe between the terminals of Ditaxinae. The results also reinforce that the Chrozophoreae tribe is polyphyletic in the current circumscription (Figure 1).
All species of Chiropetalum formed a single clade, and the geographically disjunct Mexican species emerged together with South American species. The largest clade of Ditaxinae (Argythamnia + Ditaxis) was recovered as the sister of Chiropetalum. Argythamnia species, all from the central region of the Americas (Caribbean, Central America and southern Mexico), resulted as the sister clade of Ditaxis species (Ditaxis 1 clade) with North American distribution (Figure 1). The clade Ditaxis 2, the sister of Ditaxis 1 + Argythamnia, included North American and all Central and South American species. The five African species/specimens of Caperonia sampled in the phylogeny (identified with * in Figure 1) were placed in two different clades (Figure 1). Species of the tribe Adelieae, exclusive to Central and South America, emerged as the sister clade of the monospecific genus Philyra (Figure 1).
4. Discussion
This study presents the most comprehensive taxonomic and geographical sampling of Ditaxinae (ca. 60%) to date. In an attempt to solve the generic relationships among Argythamnia, Chiropetalum and Ditaxis, Ramírez-Amezcua [28] and Külkamp [15] sampled approximately 30% and 25% of the Ditaxinae species, respectively. Furthermore, the sampling of related groups (Caperonia and Adelieae) was less than 5%, precluding any suprageneric taxonomic decisions. As a result, our research provides a solid phylogenetic framework for new taxonomic delimitations at the genus and tribe levels.
4.1. Changes in Generic Delimitation
The relationship between Argythamnia and Ditaxis could not be resolved in previous studies, probably because of the relatively low (30%) taxon sampling and lack of phylogenetic support for some clades [15,28], while the genus Chiropetalum, albeit with low support, emerged as a separated clade in both studies. A recent phylogenetic reconstruction using a large representation of subfamily Acalyphoideae [30] also recovered a monophyletic Chiropetalum, in this case with maximum support. Here, in all reconstructions, Chiropetalum emerged as monophyletic and a sister to the clade containing Ditaxis and Argythamnia, with maximum support (PP = 1) (Figure 1). The high taxon sampling of Chiropetalum (90%) in our phylogenetic analyses gives us confidence in circumscribing the genus as a distinct taxon. However, further phylogenetic studies should sample Chiropetalum patagonicum (Speg.) O’Donell & Lourteig, since the species presents a remarkable divergent morphology (prostrate habit, absence of trichomes, petals of the staminate flower slightly lobed) from that of the rest of Chiropetalum. Ingram treated this species in the genus Aonikena Apeg. [23], whereas O’Donell & Lourteig classified it in Chiropetalum sect. Aonikena (Speg.) O’Donell & Lourt [49]. Aonikena patagonica would be well placed in Chiropetalum based on comparative morphology, but nevertheless, the inclusion of this species in further phylogenetic analyses is still required to definitively clarify its taxonomic placement. Based on our phylogenetic reconstruction and morphology studies, we identified several synapomorphies of Chiropetalum, including lobed petals in the staminate flowers (Figure 2C), stamens disposed in a whorl and fused at the base forming a column (Figure 2C) and the absence of petals in the pistillate flowers (Figure 2D), except for C. tricuspidatum (Lam.) A.Juss. and C. argentinense Skottsb., which have vestigial petals [23,26,49]. A few species of Argythamnia s.s. also have pistillate flowers without petals [22]. The presence of stellate trichomes is also a unique feature of Chiropetalum, but these trichomes are present in only 10 species (50%) [23,26].
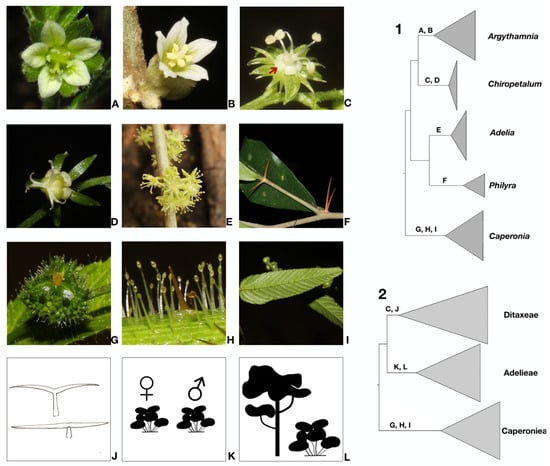
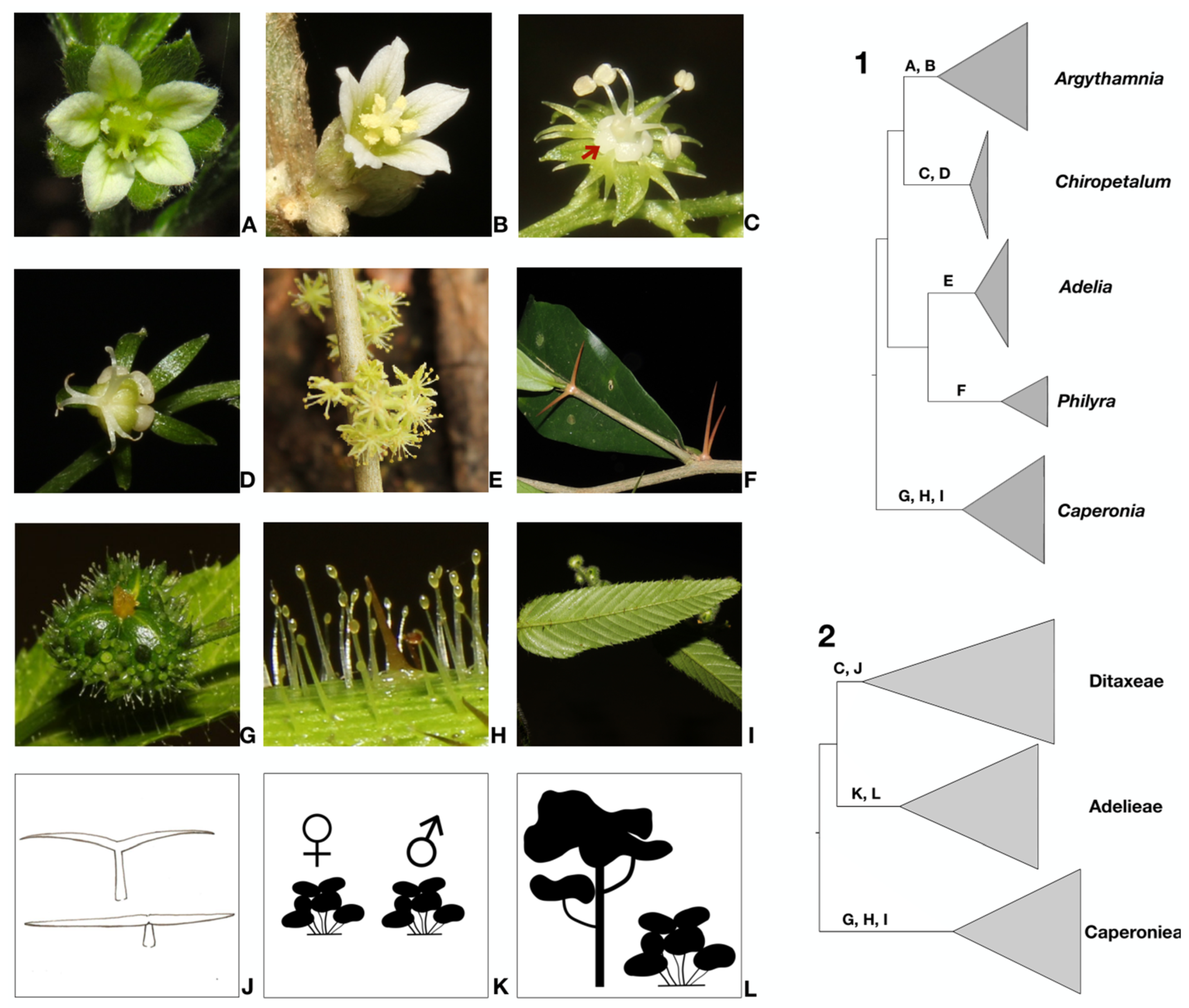
Figure 2.
Schematic phylogenies of Ditaxinae and related taxa based on Bayesian inference using the combined five markers (cpDNA [trnLF, trnTL, petD] and nDNA [ETS, ITS]). 1 & 2. Generic (1) and suprageneric (2) classifications proposed here. Letters on branches indicate the morphological synapomorphies supporting each clade corresponding to the following illustrations. (A) Dichlamydeous pistillate flower of Argythamnia desertorum. (B) Staminate flower with entire petals of Argythamnia desertorum. (C) Staminate flower with lobed petals of Chiropetalum phalacradenium for cladogram 1 and floral nectaries for cladogram 2. (D) Monochlamydeous pistillate flower of Chiropetalum phalacradenium. (E) Monochlamydeous staminate flowers of Adelia membranifolia. (F) Pair of thorns below the leaves in Philyra brasiliensis. (G) Ovary with muricate surface in Caperonia heteropetala. (H) Glandular trichomes in Caperonia heteropetala. (I) Leaves with craspedodromous secondary veins in Caperonia heteropetala. (J) Malpighiaceous trichomes. (K) Dioecious sexual system. (L) Arboreal and shrubby habit.
Our results show that Ditaxis as currently recognized is paraphyletic, because the species of Argythamnia s.s. are nested within Ditaxis (Figure 1), a topology similar to the phylogenetic reconstruction in Ramirez-Amezcua [28]. The staminate flowers in Argythamnia have four (rarely five) petals and four (rarely five) free stamens, whereas in Ditaxis the staminate flowers present with five petals and 8–10 stamens united in a column. Thus, to avoid describing a new genus lacking morphological synapomorphies or a clear set of distinguishing characteristics, we expanded the circumscription of Argythamnia s.s. with the inclusion of the two clades of Ditaxis (clade 1 & 2; Figure 1) following, in part, Ingram’s classification system [27]. Thus, Argythamnia in the circumscription proposed here is monophyletic and composed of three well-supported clades (Figure 1): (i) Argythamnia s.s., (ii) Ditaxis clade 1, exclusive to North America, and (iii) Ditaxis clade 2, the most diverse clade of Ditaxis s.s., with a distribution from North America to southern South America. In this new classification framework, Argythamnia s.l. is supported by the presence of petals in pistillate flowers (Figure 2A) (rarely absent) and entire petals (unlobed) in staminate flowers (Figure 2B). The presence of an apiculum on the seeds of Argythamnia s.l. should be studied further. Due to the lack of specimens with seeds for nine species of Argythamnia s.l., that structure was little explored in this study. The seeds of the other genera of the tribes Ditaxeae, Adelieae and Caperonieae are globose rather than apiculate.
Caperonia sensu Webster [5] is the only genus of Ditaxinae with an extra-New World distribution, with seven species occurring in tropical Africa and Madagascar. Here, we confirmed Caperonia as monophyletic, as suggested by Cervantes and collaborators [30], but with a broader taxon sampling. Pax & Hoffmann proposed two sections for Caperonia, C. sect. Eucaperonia ([nom. invalid.], autonym section = sect. Caperonia) and C. sect. Aculeolatae Pax & K.Hoffm. (taxa with prickles sampled in our phylogeny, C. corchoroides Müll.Arg., C. cordata A.St.-Hil., C. heteropetala Didr., C. linearifolia A.St.-Hil.) [14]. Based on morphology, we would have expected that these sections to be recovered in two clades in our phylogenetic analyses, but the presence of prickles appears to represent a plesiomorphic state, and some of the taxa studied have lost this state independently. However, we emphasize that Caperonia requires additional research with a larger taxonomic representation to clarify phylogenetic relationships, explore the need to establish an infrageneric classification and understand the origin and nature (multiple or single colonization events) of its amphi-Atlantic distribution pattern. When comparing Caperonia with phylogenetically closely related genera, its morphological divergence is marked by the presence of glandular trichomes (Figure 2H), a muricate ovary surface (Figure 2G) and parallel secondary veins (Figure 2I). These features are absent in all the other genera and are recognized here as synapomorphies for Caperonia. Another contrasting characteristic of Caperonia is its exclusive occurrence in marshy habitats [17,21], while all other related genera are found in desert or seasonally dry environments [15,16,19,22,23,24,25].
Philyra brasiliensis was originally the only species described in Philyra; however, the species was combined in Ditaxis by Baillon [50] and later transferred to Argythamnia by Müller [13]. Morphology does not support these classifications because Philyra lacks the synapomorphies recognized for Argythamnia + Chiropetalum (presence of floral nectaries and malpighiaceous trichomes). Moreover, Philyra is the only genus of the focal taxa having a pair of spines inserted on branches beneath the leaves (Figure 2F). Because of these unique characteristics, the species was treated again in Philyra [26]. The phylogenetic analyses of Jestrow and collaborators [31], Cervantes and collaborators [30] and our own results also support the circumscription of Philyra as a monospecific genus. The genus Adelia (sister to Philyra) includes some species with pointed branches, but it lacks the pair of spines below the leaves. Adelia is also distinguished from Philyra by its apetalous staminate flowers clustered in glomerules (Figure 2E), whereas in Philyra, the staminate flowers are dichlamydeous and grouped in racemes and the stamens (10–12) form a column with two whorls. Detailed phylogenetic information about Adelia can be found in previous studies focused on Adelieae that included a larger taxonomic representation [29,31,51,52].
4.2. Tribe Delimitation
Before our study, taxonomic affinities and phylogenetic relationships of subtribe Ditaxinae were uncertain mainly due to the poor taxon sampling in previous phylogenetic analyses [15,28,30,31,52]. Our results showed a robust topology (Figure 1), allowing us to propose a new classification. Ditaxinae has traditionally been assigned to the Chrozophoreae tribe [5,7,10]. Other phylogenetic analyses, however, revealed Chrozophoreae to be polyphyletic and Ditaxinae to be paraphyletic [2,30,31,51]. Here, we confirmed both results, with tribe Adelieae recovered as embedded among the terminals of Ditaxinae (Figure 1). Argythamnia (including Ditaxis) and Chiropetalum are part of Ditaxinae, which appear to be more closely related to each other than to Caperonia and Philyra (Figure 1 and Figure 2).
Following our phylogenetic framework, we elevated Ditaxinae to the rank of tribe (Ditaxeae), including the genera Argythamnia (including Ditaxis) and Chiropetalum (Figure 1 and Figure 2) and excluding Caperonia and Philyra (see the taxonomic treatment below). Tribe Ditaxeae is supported by two synapomorphies: the presence of floral nectaries (Figure 2C) and malpighiaceous trichomes (Figure 2J). Another important characteristic is the presence of a basal and suprabasal actinodromous venation pattern, which is very similar among taxa, but with small variations regarding the number of basal secondary veins (2–4) and the intensity of their impression on the leaf’s surface. However, this character is not exclusive to Ditaxeae; some taxa in the tribe Adelieae also present a similar venation pattern. With the exclusion of subtribe Ditaxinae, the tribe Chrozophoreae is now circumscribed to include subtribes Speranskiinae and Chrozophorinae, which are exclusively paleotropical in their distribution.
We propose to circumscribe Philyra within tribe Adelieae (Figure 1 and Figure 2), as suggested by Jestrow [31,51]. Traditionally, Philyra was circumscribed in Chrozophoreae and not in Adelieae, supported by the presence of petals in the pistillate and staminate flowers [5,10]. Now, tribe Adelieae comprises the genera Adelia, Garciadelia Jestrow & Jiménez Rodr., Lasiocroton Griseb., Leucocroton Griseb. and Philyra, which are united by two synapomorphies: the dioecious sexual system (rarely monoecious in Leucocroton) and the arborescent to shrubby habit (Figure 2K,L).
Systematists have always had difficulty placing Caperonia. Klotzsch [53] classified Caperonia in tribe Crotoneae Dumort., whereas Müller [13] placed it within tribe Acalypheae Dumort., subtribe Caperoniinae Müll.Arg. Pax & Hoffmann [14], including the genus in subtribe Chrozophorinae, and Webster [7] classified Caperonia as part of tribe Chrozophoreae, subtribe Ditaxinae, where it remained until now. Here, we circumscribe Caperonia as a monogeneric tribe based on strong phylogenetic and morphological evidence. In the most recent phylogenetic reconstruction, based on plastid data only, Caperonia emerged as a sister to Argythamnia + Chiropetalum + Ditaxis + Adelieae [30]. Although we found that the position of Caperonia was incongruent (but with low support) among phylogenetic reconstructions based on individual plastid and nuclear markers (Supplementary File S2: Figures S1–S9), our combined analysis provides strong support for its position as a sister to Adelieae + Ditaxineae (as circumscribed here), justifying its treatment as a monogeneric tribe, Caperonieae.
The new tribe Caperonieae (see taxonomic treatment below) is supported by the presence of glandular trichomes (Figure 2H) and a muricate ovary surface (Figure 2G). We also highlight the presence of leaves with craspedodromous secondary veins (Figure 2I), heteromorphic petals in staminate flowers in most species and a thickened structure at the apex of the staminal column, identified by some authors as a rudimentary ovary (pistillode) [5]. However, ontogenetic studies are needed to understand the origin of this floral structure.
4.3. Taxonomic Treatment
The molecular phylogenetic results presented here support the establishment of a new classification for Ditaxinae, raising it from the subtribe to the tribe level (Ditaxeae), and including two well-supported clades composed of genera Chiropetalum and Argythamnia. We maintain tribe Adelieae, extending its circumscription to include the genus Philyra. We also elevate subtribe Caperoniinae to the tribe level, adding two new tribes to the subfamily Acalyphoideae. Furthermore, we expanded the circumscription of Argythamnia to include the two well-supported clades of Ditaxis, representatives that emerged as paraphyletic in our analyses. Future studies will be directed at refining this delimitation and possibly proposing infrageneric classification systems for Argythamnia, Caperonia and Chiropetalum. Here, we present the names and diagnosis of the tribes and genera recognized, as well as a list of all species recognized under each genus. The necessary infrageneric nomenclature combinations will be presented in future taxonomic studies. Species with phylogenetic data used in this study are marked with an asterisk (*) in the “species recognized” section of each genus below. In Supplementary File S3: Table S1, we present a summary of the new and previous classification of all taxa treated here.
1. CAPERONIEAE Külkamp & Riina, stat. nov.
Basionym: Caperoniinae Müll.Arg. (as ‘Caperonieae’), Linnaea 34: 152. 1865.
Type, designated here: Caperonia A.St.-Hil.
Caperonia A.St.-Hil. Histoire des plantes les plus remarquables du Bresil et fu Paraguay 3/4: 244–247. 1825. Ditaxis sect. Caperonia (A.St.-Hil.) Baill. Adansonia, 4: 272. 1865.
Description: Monoecious, rarely dioecious; herbs, rarely subshrubs, annual or perennial; stems hollow; trichomes simple and glandular, sometimes prickly; stipules present; leaves alternate, petiolate or subsessile, penninerved, rarely palmatinerved, with craspedodromous secondary veins, margins serrate; inflorescences racemiform, bisexual or unisexual, bracteoles uniflorous, flowers dichlamydeous; staminate flowers with articulated pedicels; sepals 5, lanceolate, margin entire, pubescent or glabrous; petals 5, often unequal, glabrous, rarely pubescent, basally adnate to the staminal column; stamens 8–10 in two whorls, and pistillode on the column apex; floral nectaries absent; pistillate flowers proximal, dichlamydeous, sepals 5–6, equal or unequal, lanceolate to ovate, margin entire, pubescent, persistent in fruit; petals 5, usually equal, unequal or reduced; floral nectaries absent; ovary 3–locular, surface muricate, covered by glandular trichomes; style multifid; capsule verrucose, columella persistent; seeds one per locule, orbicular, foveolate, gray to black.
Distribution: Caperonia is distributed in the New World and Africa (continental Africa and Madagascar). The greatest diversity of Caperonia occurs in South America, mainly Brazil, with approximately 40% of the taxa (14). All Caperonia species occur in marshy environments [5,17,21].
Species recognized (35). Africa/Madagascar (7): Caperonia fistulosa Beille*, C. latifolia Pax, C. palustris (L.) A.St.-Hil.*, C. rutenbergii Müll.Arg., C. serrata (Turcz.) C.Presl.*, C. stuhlmannii Pax*, C. subrotunda Chiov. America (29): Caperonia aculeolata Müll.Arg., C. altissima Eskuche, C. amarumayu Külkamp & Cordeiro, C. angustissima Klotzsch, C. bahiensis Müll.Arg.*, C. buettneriacea Müll.Arg., C. capiibariensis Eskuche, C. castaneifolia (L.) A.St.-Hil.*, C. castrobarrosiana Paula & Hamburgo, C. chiltepecensis Croizat*, C. corchoroides Müll.Arg.*, C. cordata A.St.-Hil.*, C. cubana Pax & K.Hoffm.*, C. gardneri Müll.Arg., C. glabrata Pax & K.Hoffm., C. heteropetala Didr.*, C. hystrix Pax & K.Hoffm., C. langsdorffii Müll.Arg., C. linearifolia A.St.-Hil.*, C. lutea Pax & K.Hoffm., C. maracaibensis Külkamp & Cordeiro, C. multicostata Müll.Arg., C. neglecta G.L.Webster, C. palustris (L.) A.St.-Hil.*, C. paraguayensis Pax & K.Hoffm., C. regnellii Müll.Arg., C. similis Pax & K.Hoffm., C. stenophylla Müll.Arg. and C. zaponzeta Mansf.
2. DITAXEAE Külkamp & Riina, stat. nov.
Basionym: Ditaxinae Griseb., Fl. Brit. W. I., 43. 1859.
Type, designated here: Ditaxis Vahl ex A.Juss.
Description: Monoecious rarely dioecious; herbs, annual or perennial, and shrubs; branches erect, decumbent or prostrate; stipules present; leaves simple, alternate; venation actinodromous basal and suprabasal; margins serrate to entire; trichomes malpighiaceous, simple or stellate in both surfaces; racemes axillary, bisexual, rarely unisexual; pistillate flowers proximal and staminate distal; bracteoles uniflorous, lanceolate to ovate, pubescent, rarely glabrous; staminate flowers dichlamydeous; sepals (4–)5, linear to lanceolate, margin entire or serrated, pubescent or glabrous; petals (4–)5, entire, erose, laciniate or lobed, glabrous or pubescent, adnate to the staminal column; stamens 4–10, distinct or connate forming a column, stamens in one or two whorls; staminodes 0–5 at the top of the staminal column, pubescent or glabrous; floral nectaries 4–5, pubescent or glabrous; pistillate flowers dichlamydeous or monochlamydeous; sepals (4–)5(–6), linear, lanceolate, ovate or elliptic, pubescent rarely glabrous; petals 0–5, linear, lanceolate, oval, elliptic or rhomboid, pubescent or glabrous, margins entire, erose or laciniate; floral nectaries 5, adnate to the receptacle at the base of the ovary, glabrous, ciliate or pubescent; ovary pubescent, rarely glabrous; styles bifid or trifid, pubescent or glabrous; capsule 3–locular, smooth, pubescent rarely glabrous; seeds one per locule, orbicular to ovoid, apiculate or not, surface foveolate, smooth, undulate or reticulate, gray to black.
Distribution: Species of Ditaxeae are distributed throughout the New World, from the southern United States to Patagonia in the south of Argentina. There are two main centers of diversity for Ditaxeae: the first comprising southern North America, the Caribbean Islands and northern and central South America, and the second in northeastern Brazil [15,16,18,20,22,23,24,25,28].
Argythamnia P.Browne, Civ. Nat. Hist. Jamaica: 338. 1756.—Type: Argythamnia candicans Sw. = Ditaxis Vahl ex A.Juss., Euphorb. Gen. 27. 1824.—Type: Ditaxis fasciculata Vahl ex A.Juss.
Description: Monoecious, rarely dioecious herbs to shrubs, annual or perennial; trichomes malpighiaceous and simple; racemes bisexual, rarely unisexual; staminate flowers 2–15, dichlamydeous, sepals (4)5; petals (4)5, glabrous or pubescent, entire, erose or laciniate; stamens 4–10, distinct or connate, when connate arranged in two whorls; staminodes 0–5 at the top of the staminal column, pubescent or glabrous; floral nectaries 4 or 5, glabrous; pistillate flowers 1–4; dichlamydeous or monochlamydeous, sepals (4–)5(–6); petals 5, rarely 0, entire, erose or laciniate; floral nectaries 5, glabrous or ciliate; styles bifid or trifid; seeds orbicular to ovoid, apiculate, surface smooth, undulate or reticulate.
Distribution: Argythamnia is distributed throughout the New World, from southern United States to Patagonia. Greater diversity is found in southern North America, the Caribbean Islands, northern and central South America and northeastern Brazil [16,18,22,24,25,26,28].
New combination
Argythamnia grazielae (Külkamp) Külkamp & Riina comb. nov.
≡ Ditaxis grazielae Külkamp, Phytotaxa 455(1): 154. 2020.
Type: BRAZIL. Bahia: Wanderley, 25 January 1996 (fl. fr.), B.R. Chagas s.n. (holotype: RB [RB00084882]!; isotypes: CEPEC [CEPEC131190]!, K [K001206888]!, MG!, NY [NY01183998]!, SPF [SPF196837]!).
Species recognized (68): Argythamnia acaulis (Herter ex Arechav.) J.W.Ingram, A. acutangula Croizat*, A. adenophora A.Gray*, A. aphoroides Müll.Arg.*, A. argentea Millsp., A. argothamnoides (Bertero ex Spreng.) J.W.Ingram*, A. argyraea Cory*, A. arlynniana J.W.Ingram, A. brandegeei Millsp.*, A. breviramea Müll.Arg.*, A. calycina Müll.Arg., A. candicans Swartz*, A. claryana Jeps.*, A. coatepensis (T.S.Brandegee) Croizat*, A. cubensis Britton & Wilson*, A. cuneifolia (Pax & K.Hoffm.) J.W.Ingram, A. cyanophylla (Wooton & Standl.) J.W.Ingram*, A. depressa (Greenm.) J.W.Ingram, A. desertorum Müll.Arg.*, A. dioica (Bonpland, Humboldt & Kunth) Müll.Arg.*, A. ecdyomena J.Ingram*, A. erubescens J.R.Johnst., A. fasciculata (Vahl ex A.Juss.) Müll.Arg.*, A. fendleri Müll.Arg., A. grazielae (Külkamp) Külkamp & Riina*, A. guatemalensis Müll.Arg.*, A. haitiensis (Urb.) J.W.Ingram, A. haplostigma Pax & K.Hoffm., A. heterantha (Zucc.) Müll.Arg.*, A. heteropilosa J.W.Ingram, A. humilis (Engelm. & A.Gray) Müll.Arg.*, A. illimaniensis (Baill.) Müll.Arg., A. ingramii Y.Ramírez-Amezcua & V.W.Steinm., A. jablonszkyana (Pax & K.Hoffm.) J.W.Ingram*, A. katharinae (Pax) Croizat, A. lanceolata (Benth.) Müll.Arg.*, A. lottiae J.W.Ingram*, A. lucayana Millsp.*, A. lundellii J.W.Ingram*, A. macrantha (Pax & K.Hoffm.) Croizat, A. macrobotrys (Pax & K.Hoffm.) J.W.Ingram, A. malpighiacea Ule*, A. malpighiphila (Hicken) J.W.Ingram*, A. manzanilloana Rose*, A. mercurialina (Nutt.) Müll.Arg.*, A. microphylla Pax, A. montevidensis (Didr.) Müll.Arg.*, A. moorei J.W.Ingram*, A. oblongifolia Urb., A. pilosissima (Benth.) Müll.Arg., A. polygama (Jacq.) Kuntze*, A. pringlei Greenm.*, A. proctorii J.W.Ingram, A. purpurascens S.Moore*, A. rubricaulis (Pax & K.Hoffm.) Croizat*, A. salina (Pax & K.Hoffm.) J.W.Ingram*, A. sellowiana (Pax & K.Hoffm.) J.W.Ingram*, A. sericea Griseb.*, A. serrata (Torr.) Müll.Arg.*, A. silviae Y.Ramírez-Amezcua & V.W.Steinm.*, A. simoniana (Casar.) Müll.Arg.*, A. simulans J.W.Ingram*, A. sitiens (T.S.Brandegee) J.W.Ingram, A. stahlii Urb., A. tinctoria Mill.* and A. wheeleri J.W.Ingram*.
Chiropetalum A.Juss., Ann. Sci. Nat. (Paris) 25: 21. 1832. —Type: Chiropetalum lanceolatum (Cav.) A.Juss.
Description: Monoecious herbs or subshrubs, perennial rarely annual; trichomes malpighiaceous, simple, stellate or rarely absent (C. patagonicum); racemes bisexual, rarely unisexual; staminate flowers 3–35, dichlamydeous, sepals 5; petals 5, glabrous, 3–7-lobed; floral nectaries 5, glabrous or pubescent; stamens 3–6, partially connate forming a column, anthers arranged in one whorl, staminodes absent; pistillate flowers 1–5, monochlamydeous, rarely dichlamydeous, sepals 5; petals usually absent, rarely 5; floral nectaries 5, glabrous or pubescent; styles bifid; capsule covered by simple, stellate and/or malpighiaceous trichomes; seeds orbicular, surface foveolate or rough.
Distribution: Chiropetalum is distributed in South America (19 species) and Mexico (2 species). Species richness is concentrated in the central region of South America, and the species presenting with the southernmost distribution is C. patagonicum, occurring in the Patagonia region of Argentina. Morphological and geographic details of each species can be found in studies of Ingram and Külkamp [15,20,23,26].
Species recognized (21): Chiropetalum anisotrichum (Müll.Arg.) Pax & K.Hoffm.*, C. argentinense Skottsb.*, C. astroplethos (J.W.Ingram) Radcl.-Sm. & Govaerts*, C. berteroanum Schltdl.*, C. boliviense (Müll.Arg.) Pax & K.Hoffm.*, C. canescens Phil.*, C. cremnophilum I.M.Johnst., C. foliosum Pax & K.Hoffm.*, C. griseum Griseb.*, C. intermedium Pax & K.Hoffm.*, C. molle (Klotzsch ex. Baill.) Pax & K.Hoffm.*, C. patagonicum (Speg.) O’Donell & Lourteig, C. pavonianum (Müll.Arg.) Pax, C. phalacradenium (J.W.Ingram) L.B.Sm. & Downs*, C. puntaloberense Alonso Paz & Bassagoda*, C. quinquecuspidatum (A.Juss.) Pax & K.Hoffm.*, C. ramboi (Allem & Irgang) Radcl.-Sm. & Govaerts*, C. ruizianum (Müll.Arg.) Pax & K.Hoffm., C. schiedeanum (Müll.Arg.) Pax*, C. tricoccum (Vell.) Chodat & Hassl.* and C. tricuspidatum (Lam.) A.Juss*.
3. ADELIEAE G.L.Webster Taxon 24: 597. 1975
Type: Adelia L.
Description: Dioecious, rarely monoecious; trees to shrubs; pair of stipular spines absent, rarely present (Philyra); leaves alternate, simple; penninerved or actinodromous, basal and suprabasal, margins dentate to entire; trichomes simple or stellate; inflorescences axillary in racemes, glomerules or subpanicles, unisexual, rarely bisexual; staminate flowers monochlamydeous or dichlamydeous; sepals 4–5; petals absent, rarely 5; entire, pubescent; stamens 8–18(–30), filaments distinct or connate at base; staminode present or absent, floral nectaries absent; pistillate flowers monochlamydeous or dichlamydeous, sepals 5–6 lanceolate, ovate or elliptic, pubescent, rarely glabrous, persistent in fruit; petals 0–5, lanceolate, oval, elliptic or rhomboid, pubescent or glabrous; floral nectaries absent; ovary 3-locular, pubescent; ovules with inner integuments thick, outer thin to thick; styles bifid to multifid, pubescent or glabrous; capsule 3–locular; columella persistent; seeds one per locule, orbicular, surface foveolate, rough or smooth.
Distribution: Adelieae taxa are found from Mexico to Argentina, with three of the five genera endemic to the West Indies (Garciadelia, Lasiocroton and Leucocroton) [19,26,29,31,51,52,54,55].
Philyra Klotzsch, Archiv für Naturgeschichte 7(1): 199. 1841.—Type: Philyra brasiliensis Klotzsch
Description: Dioecious shrubs or treelets; paired stipular spines; leaves glabrous, venation pinnate, margin entire; bracts paleaceous, pubescent to glabrescent; staminate flowers dichlamydeous; sepals 5, petals 5; stamens 10–12, connate in a column with 2 whorls; staminodes 2 at the top of column, pubescent; pistillate flowers with pedicels larger than 12 mm long, dichlamydeous, petals 5, larger than sepals, styles multifid; capsule glabrous, columella persistent; seeds orbicular, surface smooth, gray to blackish.
Distribution: Philyra is distributed in northern Argentina, central and southern Paraguay and Brazil. In Brazil, this species occurs in the central–western region and the Atlantic coast, in the southeast and northeast of the country. For additional information, see Külkamp’s studies [19,26].
Species recognized (1): Philyra brasiliensis Klotzsch*.
Adelia L., Syst. Nat. ed. 10, 2: 1298. 1759 (nom. cons.).—Type: Adelia ricinella L. (typ. cons.)
Distribution: Adelia has a continuous distribution from the southern United States to central South America. The greatest diversity of species is found in Mexico and Central America. Although a few species are widespread (e.g., Adelia membranifolia (Müll.Arg.) Chodat & Hassl.), most of them have a narrow distribution, and some are only known to be from a limited number of localities (e.g., Adelia cinerea (Wiggins & Rollins) A.Cerv., V.W.Steinm. & Flores-Olivera).
For a diagnosis, see De-Nova & Sosa and Jestrow’s studies [31,51,55].
Species recognized (9): Adelia barbinervis Cham. & Schltdl., A. brandegeei V.W. Steinm.*, A. cinerea (Wiggins & Rollins) A.Cerv., V.W.Steinm. & Flores-Olvera*, A. membranifolia (Müll.Arg.) Chodat & Hassl., A. oaxacana (Müll.Arg.) Hemsl.*, A. obovata Wiggins & Rollins, A. ricinella L., A. triloba (Müll.Arg.) Hemsl.* and A. vaseyi (J.M.Coult.) Pax & K.Hoffm.
Garciadelia Jestrow & Jiménez Rodr., Taxon 59(6): 1809–1810. 2010.—Type: Croton leprosus Willd.
Distribution: Garciadelia has four species endemic to Hispaniola.
For a diagnosis, see Jestrow’s studies [29,31].
Species included (4): Garciadelia abbottii Jestrow & Jiménez Rodr., G. castilloae Jestrow & Jiménez Rodr., G. leprosa (Willd.) Jestrow & Jiménez Rodr. and G. mejiae Jestrow & Jiménez Rodr.
Lasiocroton Griseb., Fl. Brit. W. I. 46. 1864.—Type: Croton macrophyllus Sw.
Distribution: Lasiocroton occurs in Cuba, Hispaniola, Jamaica and the Bahamas.
For a diagnosis, see Jestrow’s studies [29,31].
Species recognized (7): Lasiocroton bahamensis Pax & K.Hoffm., L. fawcettii Urb., L. gracilis Britton & P.Wilson, L. gutierrezii Jestrow, L. harrisii Britton, L. macrophyllus (Sw.) Griseb. and L. microphyllus (A.Rich.) Jestrow.
Leucocroton Griseb. Abh. Königl. Ges. Wiss. Göttingen, 9: 20. 1861.—Type: Leucocroton wrightii Griseb.
Distribution: Leucocroton is restricted to serpentine soils of Cuba.
For a diagnosis, see studies by Borhidi and Jestrow [29,31,54].
Species recognized (26): Leucocroton acunae Borhidi, L. anomalus Borhidi, L. bracteosus Urb., L. brittonii Alain, L. comosus Urb., L. cordifolius (Britton & P.Wilson) Alain, L. discolor Urb., L. ekmanii Urb., L. flavicans Müll.Arg., L. havanensis Borhidi, L. incrustatus Borhidi, L. linearifolius Britton, L. longibracteatus Borhidi, L. moaensis Borhidi & O.Muñiz, L. moncadae Borhidi, L. obovatus Urb., L. pachyphylloides Borhidi, L. pachyphyllus Urb., L. pallidus Britton & P.Wilson, L. revolutus C.Wright, L. sameki Borhidi, L. saxicola Britton, L. stenophyllus Urb., L. subpeltatus (Urb.) Alain, L. virens Griseb. and L. wrightii Griseb.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/biology12020173/s1. Supplementary File S1: Supplementary table with taxa, voucher information and accessions used in this study. Supplementary File S2: Supplementary tables and figures including sequences of primers, protocols for PCR amplification and DNA extraction. Figures S1–S9 containing phylogenetic reconstructions of individual markers using Maximum Likelihood and Bayesian methods. Supplementary File S3: Supplementary table with summary of the new classification of Ditaxinae proposed here and the previous classification by Webster [5].
Author Contributions
J.K.: Conceptualization, formal analysis, investigation, discussion, writing (original draft). R.R.: Data resources, formal analysis, investigation, discussion, writing (original draft). Y.R.-A.: Data resources, investigation, discussion, writing (review and editing). J.R.V.I.: Investigation, discussion, writing (review and editing). I.C.: Investigation, discussion, writing (review and editing). R.G.-P.: Data resources, formal analysis. S.I.L.-C.: Data resources, writing (review and editing). J.F.A.B.: Investigation, discussion, writing (review and editing). All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded in part by: Coordenação de Aperfeiçoamento de Pessoal de Nível Superior—Brasil (CAPES) Finance Code 001; Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) for financial support to I.C. (process 309917/2015-8), J.K. (process 141707/2020-8; 402157/2022-2), J.I. (process 311847/2021-8; 402157/2022-2); J.K. received support from SYNTHESYS+ (Synthesis of Systematic resources-process ES-TAF-8188); R.R. was supported by project grant PID2019-108109GB-I00 from MCIN/AEI/10.13039/501100011033/ and FEDER “A way to make Europe”.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The DNA sequence datasets generated for this study can be found in the NCBI GenBank website. Data on vouchers used in the phylogenetic analyses are included as Supplementary Material.
Acknowledgments
We thank the European SYNTHESYS+ for supporting this study and Real Jardín Botánico, CSIC (Madrid, Spain) for providing access to the herbarium (MA) and the laboratory of molecular systematics. R.G.P. thanks RJB-CSIC for allowing her to conduct an undergraduate student internship under the supervision of R.R. in 2021. We are thankful to all people who contributed with field sample collection, curators and technicians of the physical and virtual herbaria consulted (A, ACOR, B, BA, BAA, BC, BM, BKL, BR, C, CAS, CEPEQ, CGMS, CM, COL, COR, CORD, CPAP, CTES, E, ECT, F, FLOR, G, GB, GH, GOET, HAL, HAS, HBG, HBR, HUEFS, ICN, IEB, K, LIL, LP, M, MA, MBM, MEXU, MO, MOL, MPU, MVFA, MVJB, MVM, NY, P, PACA, PEL, R, RB, RSA, S, SGO, SI, SP, SPF, TUB, UC, UPCB, US and XAL). The authors also thank Victor Steinmann for his valuable contributions to earlier version of this manuscript.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Chase, M.W.; Zmarzty, S.; Lledo, M.D.; Wurdack, K.J.; Swensen, S.M.; Fay, M.F. When in doubt, put it in Flacourtiaceae: A molecular phylogenetic analysis based on plastid rbcL DNA sequences. Kew Bull. 2002, 57, 141–181. [Google Scholar] [CrossRef]
- Wurdack, K.J.; Hoffman, P.; Chase, M.W. Molecular phylogenetic analysis of uniovulate Euphorbiaceae (Euphorbiaceae sensu stricto) using plastid rbcl and trnL-F DNA sequences. Am. J. Bot. 2005, 92, 1397–1420. [Google Scholar] [CrossRef]
- Tokuoka, T. Molecular phylogenetic analysis of Euphorbiaceae sensu stricto based on plastid and nuclear DNA sequences and ovule and seed character evolution. J. Plant Res. 2007, 120, 511–522. [Google Scholar] [CrossRef]
- Wurdack, K.J.; Davis, C.C. Malpighiales phylogenetics: Gaining ground on one of the most recalcitrant clades in the angiosperm tree of life. Am. J. Bot. 2009, 96, 1551–1570. [Google Scholar] [CrossRef]
- Webster, G.L. Euphorbiaceae. In The Families and Genera of Vascular Plants. Flowering Plants. Eudicots. Malpighiales; Kubitzki, K., Ed.; Springer: Berlin/Heidelberg, Germany, 2014; Volume 11, pp. 51–216. ISBN 978-3-642-39417-1. [Google Scholar]
- Wurdack, K.J.; Hoffmann, P.; Samuel, R.; Bruijn, A.; Bank, M.; Chase, M.W. Molecular phylogenetic analysis of Phyllanthaceae (Phyllanthoideae pro parte, Euphorbiaceae sensu lato) using plastid rbcL DNA sequences. Am. J. Bot. 2004, 91, 1882–1900. [Google Scholar] [CrossRef]
- Webster, G.L. Conspectus of a new classification of the Euphorbiaceae. Taxon 1975, 24, 593–601. [Google Scholar] [CrossRef]
- The Angiosperm Phylogeny Group; Chase, M.W.; Christenhusz, M.J.M.; Fay, M.F.; Byng, J.W.; Judd, W.S.; Soltis, D.E.; Mabberley, D.J.; Sennikov, A.N.; Soltis, P.S.; et al. An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG IV. Bot. J. Linn. Soc. 2016, 181, 12385. [Google Scholar] [CrossRef]
- Govaerts, R.; Frodin, D.G.; Radcliffe-Smith, A. World Checklist and Bibliography of Euphorbiaceae (and Pandaceae); Royal Botanical Gardens, Kew: London, UK, 2000; Volume 2, p. 1661, ISBN-10: 1900347849. [Google Scholar]
- Radcliffe-Smith, A. Genera Euphorbiacearum; Royal Botanic Garden, Kew: London, UK, 2001; p. 464, ISBN-10: 1842460226. [Google Scholar]
- Pscheidt, A.C.; Cordeiro, I. Synopsis of the tribe Hippomaneae (Euphorbiaceae) in São Paulo State. Hoehnea 2012, 39, 347–368. [Google Scholar] [CrossRef]
- Grisebach, A.H.R. Flora of the Britrish West Indian Island; Rovell Reeve Co.: London, UK, 1859; Volume 2, pp. 193–315. [Google Scholar]
- Müller, A. Euphorbiaceae: Vorläufige Mitteilungen aus dem für DeCandolle Prodromus bestimmten Manuscript über diese Familie. Linnaea 1863, 34, 1–225. [Google Scholar]
- Pax, F.; Hoffmann, K. Euphorbiaceae—Acalypheae—Chrozophorinae. In Das Pflanzenreich, 4th ed.; Engler, A., Ed.; Engelmann: Leipzig, Germany, 1912; Volume 147, pp. 1–142. [Google Scholar]
- Külkamp, J. Chiropetalum A. Juss. (Euphorbiaceae Juss.) no Brasil. Master’s Thesis, Universidade Federal do Rio Grande do Sul, Porto Alegre, Brazil, 2018; p. 87. [Google Scholar]
- Külkamp, J.; Iganci, J.R.V.; Cordeiro, I.; Baumgratz, J.F.A. Ditaxis (Euphorbiaceae) from the Brazilian Caatinga, including a new species. Phytotaxa 2020, 455, 152–160. [Google Scholar] [CrossRef]
- Külkamp, J.; Caperonia in Flora e Funga do Brasil. Jardim Botânico do Rio de Janeiro. 2021. Available online: http://floradobrasil.jbrj.gov.br/reflora/floradobrasil/FB17471 (accessed on 5 August 2021).
- Külkamp, J.; Ditaxis in Flora e Funga do Brasil. Jardim Botânico do Rio de Janeiro. 2021. Available online: http://floradobrasil.jbrj.gov.br/reflora/floradobrasil/FB25551 (accessed on 5 August 2021).
- Külkamp, J.; Philyra in Flora e Funga do Brasil. Jardim Botânico do Rio de Janeiro. 2021. Available online: http://floradobrasil.jbrj.gov.br/reflora/floradobrasil/FB25551 (accessed on 5 August 2021).
- Külkamp, J.; Iganci, J.R.V.; Cordeiro, I.; Chiropetalum in Flora e Funga do Brasil. Jardim Botânico do Rio de Janeiro 2021. Available online: http://floradobrasil.jbrj.gov.br/reflora/floradobrasil/FB29160 (accessed on 4 August 2021).
- Külkamp, J.; Iganci, J.R.V.; Cordeiro, I.; Baumgratz, J.F. New species and occurrences of Caperonia (Euphorbiaceae) for South America. Phytotaxa 2021, 529, 86–92. [Google Scholar] [CrossRef]
- Ingram, J.W. A revisional study of Argythamnia subgenus Argythamnia (Euphorbiaceae). Gentes Herbarum 1967, 10, 1–38. [Google Scholar]
- Ingram, J.W. A revision of Argythamnia subgenus Chiropetalum (Euphorbiaceae). Gentes Herbarum 1980, 11, 437–468. [Google Scholar]
- Ingram, J.W. A sectional revision of Argythamnia subgenus Ditaxis (Euphorbiaceae). Ph.D. Thesis, University of California, Berkeley, CA, USA, 1956; p. 115. [Google Scholar]
- Ramírez-Amezcua, Y.; Steinmann, V.W. Revisión taxonómica de Argythamnia subgénero Ditaxis (Euphorbiaceae) en México. Bot. Sci. 2013, 91, 427–459. [Google Scholar] [CrossRef]
- Külkamp, J. Sistemática da Subtribo Ditaxinae (Euphorbiaceae). Ph.D. Thesis, Escola Nacional de Botânica Tropical, Jardim Botânico do Rio de Janeiro, Rio de Janeiro, Brazil, 2022; p. 245. [Google Scholar]
- Ingram, J.W. The generic limits of Argythamnia (Euphorbiaceae) defined. Gentes Herbarum 1980, 11, 427–436. [Google Scholar]
- Ramírez-Amezcua, Y. Relaciones Filogenéticas en Argythamnia (Euphorbiaceae) Sensu Lato. Master’s Thesis, Universidad Michoacana de San Nicolás de Hidalgo, Michoacán, Mexico, 2011; p. 64. [Google Scholar]
- Jestrow, B.; Gutièrez, J.A.; Francisco-Ortega, J. Islands within islands: A molecular phylogenetic study of the Leucocroton alliance (Euphorbiaceae) across the Carribean Islands and within the serpentinite archipelago of Cuba. J. Biogeogr. 2012, 39, 452–464. [Google Scholar] [CrossRef]
- Cervantes, A.; Fuentes, S.; Gutiérrez, J.; Magallón, S.; Borsch, T. Successive arrivals since the Miocene shaped the diversity of the Caribbean Acalyphoideae (Euphorbiaceae). J. Biogeogr. 2016, 43, 1773–1785. [Google Scholar] [CrossRef]
- Jestrow, B.; Rodríguez, F.J.; Francisco-Ortega, J. Generic delimitation in the Antillean Adelieae (Euphorbiaceae) with description of the Hispaniolan endemic genus Garciadelia. Taxon 2010, 59, 1801–1814. [Google Scholar] [CrossRef]
- Yang. Y.; Riina. R.; Morawetz, J.J.; Haevermans, T.; Aubriot, X.; Berry, P.E. Molecular phylogenetics and classification of Euphorbia subgenus Chamaesyce (Euphorbiaceae). Taxon 2012, 61, 764–789. [Google Scholar] [CrossRef]
- Riina, R.; Peirson, J.A.; Geltman, D.V.; Molero, J.; Frajman, B.; Pahlevani, A.; Barres, L.; Morawetz, J.J.; Salmaki, Y.; Zarre, S.; et al. A worldwide molecular phylogeny and classification of the leafy spurges, Euphorbia subgenus Esula (Euphorbiaceae). Taxon 2013, 62, 316–342. [Google Scholar] [CrossRef]
- Van Ee, B.W.; Riina, R.; Berry, P.E. A revised infrageneric classification and molecular phylogeny of New World Croton (Euphorbiaceae). Taxon 2011, 60, 791–823. [Google Scholar] [CrossRef]
- Thiers, B. Index Herbariorum: A Global Directory of Public Herbaria and Associated Staff; New York Botanical Garden’s Virtual Herbarium: New York, NY, USA, 2022; Available online: http://sweetgum.nybg.org/science/ih/ (accessed on 20 February 2022).
- Doyle, J. DNA Protocols for Plants. In Molecular Techniques in Taxonomy; Hewitt, G.M., Johnston, A.W.B., Young, J.P.W., Eds.; Springer: Berlin/Heidelberg, Germany, 1991; pp. 283–293. ISBN 978-3-642-83962-7. [Google Scholar]
- Okonechnikov, K.; Golosova, O.; Fursov, M. Unipro UGENE: A unified bioinformatics toolkit. Bioinformatics 2012, 28, 1166–1167. [Google Scholar] [CrossRef] [PubMed]
- Akaike, H. A new look at the statistical model identification. IEEE Trans. Autom. Contr. 1974, 19, 716–723. [Google Scholar] [CrossRef]
- Guindon, S.; Gascuel, O. A simple, fast and accurate method to estimate large phylogenies by maximum- likelihood. Syst. Biol. 2003, 52, 696–704. [Google Scholar] [CrossRef]
- Darriba, D.; Taboada, G.L.; Doallo, R.; Posada, D. jModelTest 2: More models, new heuristics and parallel computing. Nat. Methods 2012, 9, 772. [Google Scholar] [CrossRef]
- Masa-Iranzo, I.; Sanmartín, I.; Caruzo, M.B.R.; Riina, R. Skipping the Dry Diagonal: Spatio-temporal evolution of Croton section Cleodora (Euphorbiaceae) in the Neotropics. Bot. J. Linn. Soc. 2021, 197, 61–84. [Google Scholar] [CrossRef]
- Ronquist, F.; Deans, A.R. Bayesian phylogenetics and its influence on insect systematics. Annu. Rev. Entomol. 2010, 55, 189–206. [Google Scholar] [CrossRef]
- Alfaro, M.E.; Zoller, S.; Lutzoni, F. Bayes or bootstrap? A simulation study comparing the performance of Bayesian Markov chain Monte Carlo sampling and bootstrapping in assessing phylogenetic confidence. Mol. Biol. Evol. 2003, 20, 255–266. [Google Scholar] [CrossRef]
- Ronquist, F.; Teslenko, M.; Van der Mark, P.; Ayres, D.; Darling, A.; Höhna, S.; Larget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef]
- Drummond, A.J.; Rambaut, A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol. Biol. 2007, 7, 214. [Google Scholar] [CrossRef]
- Rambaut, A.; Drummond, A.J.; Xie, D.; Baele, G. Posterior Summarization in Bayesian Phylogenetics Using Tracer 1.7. Syst. Biol. 2018, 67, 901–904. [Google Scholar] [CrossRef] [PubMed]
- Stamatakis, A.; Hoover, P.; Rougemont, J. A rapid bootstrap algorithm for the RAxML web servers. Syst. Biol. 2008, 57, 758–771. [Google Scholar] [CrossRef] [PubMed]
- Miller, M.A.; Pfeiffer, W.; Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In Proceedings of the Gateway Computing Environments Workshop (GCE), New Orleans, LA, USA, 14 November 2010; pp. 1–8. [Google Scholar] [CrossRef]
- O’Donell, C.A.; Lourteig, A. Chrozophorae Argentinae. Lilloa 1942, 8, 37–81. [Google Scholar]
- Baillon, H.E. D’Observations Botaniques. Adansonia 1864, 4, 289. [Google Scholar]
- Jestrow, B.; Proctor, G.; Francisco-Ortega, J. Lasiocroton trelawniensis (Euphorbiaceae), a critically endangered species from the Cockpit Country of Jamaica, belongs to Bernardia (Euphorbiaceae). Bot. Rev. 2008, 74, 166–177. [Google Scholar] [CrossRef]
- De-Nova, J.A.; Sosa, V. Phylogeny and generic delimitation of Adelia (Euphorbiaceae) inferred from molecular and morphological data. Taxon 2007, 56, 1027–1036. [Google Scholar] [CrossRef]
- Bentham, G. Contribution towards a Flora of South America—Enumeration of Plants Collected by Mr. Schomburgk in British Guiana. Lond. J. Bot. 1843, 2, 42–52. [Google Scholar]
- Borhidi, A. Taxonomic revision of genus Leucocroton (Euphorbiaceae). Acta Bot. Hung. 1991, 36, 13–40. [Google Scholar]
- De-Nova, J.A.; Sosa, V.; Wurdack, K.J. Phylogenetic relationships and the description of a new species of Enriquebeltrania (Euphorbiaceae s.s.): An enigmatic genus endemic to Mexico. Syst. Bot. 2006, 31, 533–546. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).