Reactive Oxygen Species, Antioxidant Responses and Implications from a Microbial Modulation Perspective
Simple Summary
Abstract
1. Introduction
2. Enzymatic Defensive Mechanisms
2.1. Superoxide Dismutase
2.2. Catalase
2.3. Ascorbate Peroxidase
2.4. Guaiacol Peroxidase (GPX)
2.5. Glutathione Reductase (GR)
2.6. Monodehydroascorbate Reductase (MDHAR)
2.7. Dehydroascorbate Reductase (DHAR)
3. Non-Enzymatic Defensive Mechanisms
3.1. Ascorbic Acid
3.2. Glutathione
3.3. Proline
3.4. α-Tocopherols
3.5. Carotenoids
3.6. Phenolic Compounds
4. Antioxidant Machinery in Plant Systems and Microbial Mediation in Promoting Plant Tolerance
5. Conclusions and Future Prospects
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Demirel, U.; Morris, W.L.; Ducreux, L.J.M.; Yavuz, C.; Asim, A.; Tindas, I.; Campbell, R.; Morris, J.A.; Verrall, S.R.; Hedley, P.; et al. Physiological, Biochemical, and Transcriptional Responses to Single and Combined Abiotic Stress in Stress-Tolerant and Stress-Sensitive Potato Genotypes. Front. Plant Sci. 2020, 11, 169. [Google Scholar] [CrossRef] [PubMed]
- Rahman, K.; Rahman, M.; Ahmed, N.; Alam, M.; Rahman, A.; Islam, M.; Hasanuzzaman, M. Morphophysiological changes and reactive oxygen species metabolism in Corchorus olitorius L. under different abiotic stresses. Open Agric. 2021, 6, 549–562. [Google Scholar] [CrossRef]
- Ahsan, H.; Ali, A.; Ali, R. Oxygen free radicals and systemic autoimmunity. Clin. Exp. Immunol. 2003, 131, 398–404. [Google Scholar] [CrossRef] [PubMed]
- Jaleel, C.A.; Riadh, K.; Gopi, R.; Manivannan, P.; Inès, J.; Al-Juburi, H.J.; Chang-Xing, Z.; Hong-Bo, S.; Panneerselvam, R. Antioxidant defense responses: Physiological plasticity in higher plants under abiotic constraints. Acta Physiol. Plant. 2009, 31, 427–436. [Google Scholar] [CrossRef]
- Sewelam, N.; Kazan, K.; Schenk, P.M. Global Plant Stress Signaling: Reactive Oxygen Species at the Cross-Road. Front. Plant Sci. 2016, 7, 187. [Google Scholar] [CrossRef]
- Bhattacharjee, S. Reactive oxygen species and oxidative burst: Roles in stress, senescence and signal. Curr. Sci. 2005, 89, 1113–1121. Available online: https://www.jstor.org/stable/24110963 (accessed on 10 November 2021).
- Marino, D.; Andrio, E.; Danchin, E.G.J.; Oger, E.; Gucciardo, S.; Lambert, A.; Puppo, A.; Pauly, N. A Medicago truncatula NADPH oxidase is involved in symbiotic nodule functioning. New Phytol. 2010, 189, 580–592. [Google Scholar] [CrossRef]
- Kadota, Y.; Shirasu, K.; Zipfel, C. Regulation of the NADPH Oxidase RBOHD During Plant Immunity. Plant Cell Physiol. 2015, 56, 1472–1480. [Google Scholar] [CrossRef]
- Stankovic-Valentin, N.; Melchior, F. Control of SUMO and Ubiquitin by ROS: Signaling and disease implications. Mol. Asp. Med. 2018, 63, 3–17. [Google Scholar] [CrossRef]
- Mittler, R. Oxidative stress, antioxidants and stress tolerance. Trends Plant Sci. 2002, 7, 405–410. [Google Scholar] [CrossRef]
- Dvořák, P.; Krasylenko, Y.; Zeiner, A.; Šamaj, J.; Takáč, T. Signaling Toward Reactive Oxygen Species-Scavenging Enzymes in Plants. Front. Plant Sci. 2021, 11, 618835. [Google Scholar] [CrossRef] [PubMed]
- Gleason, C.; Huang, S.; Thatcher, L.; Foley, R.C.; Anderson, C.R.; Carroll, A.J.; Millar, A.H.; Singh, K.B. Mitochondrial complex II has a key role in mitochondrial-derived reactive oxygen species influence on plant stress gene regulation and defense. Proc. Natl. Acad. Sci. USA 2011, 108, 10768–10773. [Google Scholar] [CrossRef]
- Foyer, C.H. Reactive oxygen species, oxidative signaling and the regulation of photosynthesis. Environ. Exp. Bot. 2018, 154, 134–142. [Google Scholar] [CrossRef]
- Del Río, L.A.; Lopez-Huertas, E. ROS Generation in Peroxisomes and its Role in Cell Signaling. Plant Cell Physiol. 2016, 57, 1364–1376. [Google Scholar] [CrossRef]
- Bindschedler, L.V.; Dewdney, J.; Blee, K.A.; Stone, J.M.; Asai, T.; Plotnikov, J.; Denoux, C.; Hayes, T.; Gerrish, C.; Davies, D.R.; et al. Peroxidase-dependent apoplastic oxidative burst in Arabidopsis required for pathogen resistance. Plant J. 2006, 47, 851–863. [Google Scholar] [CrossRef] [PubMed]
- Roychoudhury, A.; Basu, S. Ascorbate-Glutathione and plant tolerance to various abiotic stresses. In Oxidative Stress in Plants: Causes, Consequences and Tolerance; Anjum, N.A., Umar, S., Ahmad, A., Eds.; IK International Publishers: New Delhi, India, 2012; pp. 177–258. [Google Scholar]
- Foyer, C.H.; Noctor, G. Redox homeostasis and antioxidant signaling: A metabolic interface between stress perception and physiological responses. Plant Cell 2005, 17, 1866–1875. [Google Scholar] [CrossRef]
- Mhamdi, A.; Van Breusegem, F. Reactive oxygen species in plant development. Development 2018, 145, dev164376. [Google Scholar] [CrossRef]
- Xia, X.J.; Zhou, Y.H.; Shi, K.; Zhou, J.; Foyer, C.H.; Yu, J.Q. Interplay between reactive oxygen species and hormones in the control of plant development and stress tolerance. J. Exp. Bot. 2015, 66, 2839–2856. [Google Scholar] [CrossRef] [PubMed]
- Gill, S.S.; Tuteja, N. Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol. Biochem. 2010, 48, 909–930. [Google Scholar] [CrossRef]
- Nath, M.; Bhatt, D.; Prasad, R.; Gill, S.S.; Anjum, N.A.; Tuteja, N. Reactive Oxygen Species Generation-Scavenging and Signaling during Plant-Arbuscular Mycorrhizal and Piriformospora indica Interaction under Stress Condition. Front. Plant Sci. 2016, 7, 1574. [Google Scholar] [CrossRef]
- Baxter, A.; Mittler, R.; Suzuki, N. ROS as key players in plant stress signalling. J. Exp. Bot. 2013, 65, 1229–1240. [Google Scholar] [CrossRef]
- Huang, H.; Ullah, F.; Zhou, D.X.; Yi, M.; Zhao, Y. Mechanisms of ROS regulation of plant development and stress responses. Front. Plant Sci. 2019, 10, 800. [Google Scholar] [CrossRef]
- Petrov, V.; Hille, J.; Mueller-Roeber, B.; Gechev, T.S. ROS-mediated abiotic stress-induced programmed cell death in plants. Front. Plant Sci. 2015, 6, 69. [Google Scholar] [CrossRef]
- Mittler, R.; Vanderauwera, S.; Gollery, M.; Van Breusegem, F. Reactive oxygen gene network of plants. Trends Plant Sci. 2004, 9, 490–498. [Google Scholar] [CrossRef] [PubMed]
- Das, K.; Roychoudhury, A. Reactive oxygen species (ROS) and response of antioxidants as ROS-scavengers during environmental stress in plants. Front. Environ. Sci. 2014, 2, 53. [Google Scholar] [CrossRef]
- Miller, G.; Suzuki, N.; Ciftci-Yilmaz, S.; Mittler, R. Reactive oxygen species homeostasis and signalling during drought and salinity stresses. Plant Cell Environ. 2010, 33, 453–467. [Google Scholar] [CrossRef]
- Rasool, S.; Ahmad, A.; Siddiqi, T.O.; Ahmad, P. Changes in growth, lipid peroxidation and some key antioxidant enzymes in chickpea genotypes under salt stress. Acta Physiol. Plant. 2012, 35, 1039–1050. [Google Scholar] [CrossRef]
- Talbi, S.; Romero-Puertas, M.C.; Hernández, A.; Terrón, L.; Ferchichi, A.; Sandalio, L.M. Drought tolerance in a Saharian plant Oudneya africana: Role of antioxidant defences. Environ. Exp. Bot. 2015, 111, 114–126. [Google Scholar] [CrossRef]
- Yasar, F.; Ellialtioglu, S.; Yildiz, K. Effect of salt stress on antioxidant defense systems, lipid peroxidation, and chlorophyll content in green bean. Russ. J. Plant Physiol. 2008, 55, 782–786. [Google Scholar] [CrossRef]
- Nahar, S.; Vemireddy, L.R.; Sahoo, L.; Tanti, B. Antioxidant Protection Mechanisms Reveal Significant Response in Drought-Induced Oxidative Stress in Some Traditional Rice of Assam, India. Rice Sci. 2018, 25, 185–196. [Google Scholar] [CrossRef]
- Contreras, R.A.; Pizarro, M.; Köhler, H.; Sáez, C.A.; Zúñiga, G.E. Copper stress induces antioxidant responses and accumulation of sugars and phytochelatins in Antarctic Colobanthus quitensis (Kunth) Bartl. Biol. Res. 2018, 51, 48. [Google Scholar] [CrossRef]
- Yogendra, S.G.; Singh, U.S.; Sharma, A.K. Bacterial mediated amelioration of drought stress in drought tolerant and susceptible cultivars of rice (Oryza sativa L.). Afr. J. Biotechnol. 2015, 14, 764–773. [Google Scholar] [CrossRef]
- Grover, M.; Ali, S.Z.; Sandhya, V.; Rasul, A.; Venkateswarlu, B. Role of microorganisms in adaptation of agriculture crops to abiotic stresses. World J. Microbiol. Biotechnol. 2010, 27, 1231–1240. [Google Scholar] [CrossRef]
- Rolli, E.; Marasco, R.; Vigani, G.; Ettoumi, B.; Mapelli, F.; DeAngelis, M.L.; Gandolfi, C.; Casati, E.; Previtali, F.; Gerbino, R.; et al. Improved plant resistance to drought is promoted by the root-associated microbiome as a water stress-dependent trait. Environ. Microbiol. 2014, 17, 316–331. [Google Scholar] [CrossRef]
- Berg, G.; Zachow, C.; Müller, H.; Philipps, J.; Tilcher, R. Next-Generation Bio-Products Sowing the Seeds of Success for Sustainable Agriculture. Agronomy 2013, 3, 648–656. [Google Scholar] [CrossRef]
- Sandhya, V.; Ali, S.Z.; Grover, M.; Reddy, G.; Venkateswarlu, B. Effect of plant growth promoting Pseudomonas spp. on compatible solutes, antioxidant status and plant growth of maize under drought stress. Plant Growth Regul. 2010, 62, 21–30. [Google Scholar] [CrossRef]
- Jha, Y.; Subramanian, R.B. PGPR regulate caspase-like activity, programmed cell death, and antioxidant enzyme activity in paddy under salinity. Physiol. Mol. Biol. Plants 2014, 20, 201–207. [Google Scholar] [CrossRef]
- Suárez, R.; Wong, A.; Ramírez, M.; Barraza, A.; Orozco, M.D.C.; Cevallos, M.A.; Lara, M.; Hernández, G.; Iturriaga, G. Improvement of Drought Tolerance and Grain Yield in Common Bean by Overexpressing Trehalose-6-Phosphate Synthase in Rhizobia. Mol. Plant-Microbe Interact. 2008, 21, 958–966. [Google Scholar] [CrossRef]
- Kasim, W.A.; Osman, M.E.; Omar, M.N.; El-Daim, I.A.A.; Bejai, S.; Meijer, J. Control of Drought Stress in Wheat Using Plant-Growth-Promoting Bacteria. J. Plant Growth Regul. 2012, 32, 122–130. [Google Scholar] [CrossRef]
- Pereyra, M.A.; Zalazar, C.; Barassi, C. Root phospholipids in Azospirillum-inoculated wheat seedlings exposed to water stress. Plant Physiol. Biochem. 2006, 44, 873–879. [Google Scholar] [CrossRef] [PubMed]
- Dimkpa, C.; Weinand, T.; Asch, F. Plant-rhizobacteria interactions alleviate abiotic stress conditions. Plant Cell Environ. 2009, 32, 1682–1694. [Google Scholar] [CrossRef]
- Hartmann, A.; Klink, S.; Rothballer, M. Plant Growth Promotion and Induction of Systemic Tolerance to Drought and Salt Stress of Plants by Quorum Sensing Auto-Inducers of the N-acyl-homoserine Lactone Type: Recent Developments. Front. Plant Sci. 2021, 12, 683546. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Dias, M.C.; Freitas, H. Drought and Salinity Stress Responses and Microbe-Induced Tolerance in Plants. Front. Plant Sci. 2020, 11, 591911. [Google Scholar] [CrossRef]
- Wang, C.-J.; Yang, W.; Wang, C.; Gu, C.; Niu, D.-D.; Liu, H.-X.; Wang, Y.-P.; Guo, J.-H. Induction of Drought Tolerance in Cucumber Plants by a Consortium of Three Plant Growth-Promoting Rhizobacterium Strains. PLoS ONE 2012, 7, e52565. [Google Scholar] [CrossRef]
- Dvořák, P.; Krasylenko, Y.; Ovečka, M.; Basheer, J.; Zapletalová, V.; Šamaj, J.; Takáč, T. In vivo light-sheet microscopy resolves localisation patterns of FSD1, a superoxide dismutase with function in root development and osmoprotection. Plant Cell Environ. 2020, 44, 68–87. [Google Scholar] [CrossRef] [PubMed]
- Noctor, G.; Reichheld, J.-P.; Foyer, C.H. ROS-related redox regulation and signaling in plants. Semin. Cell Dev. Biol. 2018, 80, 3–12. [Google Scholar] [CrossRef]
- Foyer, C.H.; Noctor, G. Redox Homeostasis and Signaling in a Higher-CO2World. Annu. Rev. Plant Biol. 2020, 71, 157–182. [Google Scholar] [CrossRef]
- Pilon, M.; Ravet, K.; Tapken, W. The biogenesis and physiological function of chloroplast superoxide dismutases. Biochim. Biophys. Acta Bioenerg. 2011, 1807, 989–998. [Google Scholar] [CrossRef]
- Kliebenstein, D.J.; Monde, R.-A.; Last, R.L. Superoxide Dismutase in Arabidopsis: An Eclectic Enzyme Family with Disparate Regulation and Protein Localization. Plant Physiol. 1998, 118, 637–650. [Google Scholar] [CrossRef] [PubMed]
- Alscher, R.G.; Erturk, N.; Heath, L.S. Role of superoxide dismutases (SODs) in controlling oxidative stress in plants. J. Exp. Bot. 2002, 53, 1331–1341. [Google Scholar] [CrossRef]
- Myouga, F.; Hosoda, C.; Umezawa, T.; Iizumi, H.; Kuromori, T.; Motohashi, R.; Shono, Y.; Nagata, N.; Ikeuchi, M.; Shinozaki, K. A Heterocomplex of Iron Superoxide Dismutases Defends Chloroplast Nucleoids against Oxidative Stress and Is Essential for Chloroplast Development in Arabidopsis. Plant Cell 2008, 20, 3148–3162. [Google Scholar] [CrossRef]
- Morgan, M.J.; Lehmann, M.; Schwarzländer, M.; Baxter, C.J.; Sienkiewicz-Porzucek, A.; Williams, T.C.; Schauer, N.; Fernie, A.R.; Fricker, M.D.; Ratcliffe, R.G.; et al. Decrease in Manganese Superoxide Dismutase Leads to Reduced Root Growth and Affects Tricarboxylic Acid Cycle Flux and Mitochondrial Redox Homeostasis. Plant Physiol. 2008, 147, 101–114. [Google Scholar] [CrossRef]
- Jamdhade, A.R.; Sunkar, R.; Hivrale, V.K. Zymographic Method for Distinguishing Different Classes of Superoxide Dismutases in Plants. Methods Mol. Biol. 2017, 1631, 221–227. [Google Scholar] [CrossRef] [PubMed]
- Sales, C.R.G.; Ribeiro, R.V.; Silveira, J.A.G.; Machado, E.C.; Martins, M.O.; Lagôa, A.M.M.A. Superoxide dismutase and ascorbate peroxidase improve the recovery of photosynthesis in sugarcane plants subjected to water deficit and low substrate temperature. Plant Physiol. Biochem. 2013, 73, 326–336. [Google Scholar] [CrossRef] [PubMed]
- Tewari, R.K.; Kumar, P.; Tewari, N.; Srivastava, S.; Sharma, P.N. Macronutrient deficiencies and differential antioxidant responses—Influence on the activity and expression of superoxide dismutase in maize. Plant Sci. 2004, 166, 687–694. [Google Scholar] [CrossRef]
- Yordanova, R.Y. Antioxidative enzymes in barley plants subjected to soil flooding. Environ. Exp. Bot. 2004, 51, 93–101. [Google Scholar] [CrossRef]
- Stephenie, S.; Chang, Y.P.; Gnanasekaran, A.; Esa, N.M.; Gnanaraj, C. An insight on superoxide dismutase (SOD) from plants for mammalian health enhancement. J. Funct. Foods 2020, 68, 103917. [Google Scholar] [CrossRef]
- Li, J.; Arkorful, E.; Cheng, S.; Zhou, Q.; Li, H.; Chen, X.; Sun, K.; Li, X. Alleviation of cold damage by exogenous application of melatonin in vegetatively propagated tea plant (Camellia sinensis L. O. Kuntze). Sci. Hortic. 2018, 238, 356–362. [Google Scholar] [CrossRef]
- Gupta, A.S.; Webb, R.P.; Holaday, A.S.; Allen, R.D. Overexpression of Superoxide Dismutase Protects Plants from Oxidative Stress (Induction of Ascorbate Peroxidase in Superoxide Dismutase-Overexpressing Plants). Plant Physiol. 1993, 103, 1067–1073. [Google Scholar] [CrossRef]
- Rady, M.O.A.; Semida, W.M.; El-Mageed, T.A.A.; Hemida, K.A.; Rady, M.M. Up-regulation of antioxidative defense systems by glycine betaine foliar application in onion plants confer tolerance to salinity stress. Sci. Hortic. 2018, 240, 614–622. [Google Scholar] [CrossRef]
- Yi, X.-P.; Zhang, Y.-L.; Yao, H.-S.; Luo, H.-H.; Gou, L.; Chow, W.S.; Zhang, W.-F. Rapid recovery of photosynthetic rate following soil water deficit and re-watering in cotton plants (Gossypium herbaceum L.) is related to the stability of the photosystems. J. Plant Physiol. 2016, 194, 23–34. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Lin, R.; Zhang, Z.; Zhou, M. Purification and characterisation of superoxide dismutase from tartary buckwheat leaves. Fagopyrum 1993, 13, 31–34. [Google Scholar]
- Szőllősi, R. Superoxide dismutase (SOD) and abiotic stress tolerance in plants: An overview. In Oxidative Damage to Plants; Ahmad, A., Ed.; Academic Press: New York, NY, USA, 2014; pp. 89–129. [Google Scholar] [CrossRef]
- Río, L.A.D.; Corpas, F.J.; López-Huertas, E.; Palma, J.M. Plant superoxide dismutases: Function under abiotic stress conditions. In Antioxidants and Antioxidant Enzymes in Higher Plants, 1st ed.; Gupta, D., Palma, J., Corpas, F., Eds.; Springer: Cham, Switzerland, 2018; pp. 1–26. [Google Scholar] [CrossRef]
- Bowler, C.; Van Camp, W.; Van Montagu, M.; Inzé, D.; Asada, P.K. Superoxide Dismutase in Plants. Crit. Rev. Plant Sci. 1994, 13, 199–218. [Google Scholar] [CrossRef]
- Lima, C.S.; Ferreira-Silva, S.L.; Carvalho, F.E.L.; Neto, M.C.L.; Aragão, R.M.; Silva, E.N.; Sousa, R.M.J.; Silveira, J.A.G. Antioxidant protection and PSII regulation mitigate photo-oxidative stress induced by drought followed by high light in cashew plants. Environ. Exp. Bot. 2018, 149, 59–69. [Google Scholar] [CrossRef]
- Lin, K.-H.; Kuo, W.-S.; Chiang, C.-M.; Hsiung, T.-C.; Chiang, M.-C.; Lo, H.-F. Study of sponge gourd ascorbate peroxidase and winter squash superoxide dismutase under respective flooding and chilling stresses. Sci. Hortic. 2013, 162, 333–340. [Google Scholar] [CrossRef]
- Ju, Y.-L.; Yue, X.-F.; Zhao, X.-F.; Zhao, H.; Fang, Y.-L. Physiological, micro-morphological and metabolomic analysis of grapevine (Vitis vinifera L.) leaf of plants under water stress. Plant Physiol. Biochem. 2018, 130, 501–510. [Google Scholar] [CrossRef]
- Dias, M.C.; Mariz-Ponte, N.; Santos, C. Lead induces oxidative stress in Pisum sativum plants and changes the levels of phytohormones with antioxidant role. Plant Physiol. Biochem. 2019, 137, 121–129. [Google Scholar] [CrossRef]
- Boguszewska, D.; Grudkowska, M.; Zagdanska, B. Drought-Responsive Antioxidant Enzymes in Potato (Solanum tuberosum L.). Potato Res. 2010, 53, 373–382. [Google Scholar] [CrossRef]
- Shafi, A.; Chauhan, R.; Gill, T.; Swarnkar, M.K.; Sreenivasulu, Y.; Kumar, S.; Kumar, N.; Shankar, R.; Ahuja, P.S.; Singh, A.K. Expression of SOD and APX genes positively regulates secondary cell wall biosynthesis and promotes plant growth and yield in Arabidopsis under salt stress. Plant Mol. Biol. 2015, 87, 615–631. [Google Scholar] [CrossRef]
- Xing, Y.; Cao, Q.; Zhang, Q.; Qin, L.; Jia, W.; Zhang, J. MKK5 Regulates High Light-Induced Gene Expression of Cu/Zn Superoxide Dismutase 1 and 2 in Arabidopsis. Plant Cell Physiol. 2013, 54, 1217–1227. [Google Scholar] [CrossRef] [PubMed]
- Rizhsky, L.; Liang, H.; Mittler, R. The Water-Water Cycle Is Essential for Chloroplast Protection in the Absence of Stress. J. Biol. Chem. 2003, 278, 38921–38925. [Google Scholar] [CrossRef]
- Tuzet, A.; Rahantaniaina, M.-S.; Noctor, G. Analyzing the Function of Catalase and the Ascorbate–Glutathione Pathway in H2O2 Processing: Insights from an Experimentally Constrained Kinetic Model. Antioxid. Redox Signal. 2019, 30, 1238–1268. [Google Scholar] [CrossRef]
- Frugoli, J.A.; Zhong, H.H.; Nuccio, M.L.; McCourt, P.; McPeek, M.A.; Thomas, T.L.; McClung, C.R. Catalase is encoded by a multigene family in Arabidopsis thaliana (L.) Heynh. Plant Physiol. 1996, 112, 327–336. [Google Scholar] [CrossRef] [PubMed]
- Scandalios, J.G. Oxidative stress: Molecular perception and transduction of signals triggering antioxidant gene defenses. Braz. J. Med. Biol. Res. 2005, 38, 995–1014. [Google Scholar] [CrossRef]
- Wang, W.; Cheng, Y.; Chen, D.; Liu, D.; Hu, M.; Dong, J.; Zhang, X.; Song, L.; Shen, F. The Catalase Gene Family in Cotton: Genome-Wide Characterization and Bioinformatics Analysis. Cells 2019, 8, 86. [Google Scholar] [CrossRef]
- Hu, L.; Yang, Y.; Jiang, L.; Liu, S. The catalase gene family in cucumber: Genome-wide identification and organization. Genet. Mol. Biol. 2016, 39, 408–415. [Google Scholar] [CrossRef]
- Joo, J.; Lee, Y.H.; Song, S.I. Rice CatA, CatB, and CatC are involved in environmental stress response, root growth, and photorespiration, respectively. J. Plant Biol. 2014, 57, 375–382. [Google Scholar] [CrossRef]
- Du, Y.-Y.; Wang, P.; Chen, J.; Song, C.-P. Comprehensive Functional Analysis of the Catalase Gene Family in Arabidopsis thaliana. J. Integr. Plant Biol. 2008, 50, 1318–1326. [Google Scholar] [CrossRef]
- Esaka, M.; Yamada, N.; Kitabayashi, M.; Setoguchi, Y.; Tsugeki, R.; Kondo, M.; Nishimura, M. cDNA cloning and differential gene expression of three catalases in pumpkin. Plant Mol. Biol. 1997, 33, 141–155. [Google Scholar] [CrossRef] [PubMed]
- Guan, L.; Scandalios, J.G. Developmentally related responses of maize catalase genes to salicylic acid. Proc. Natl. Acad. Sci. USA 1995, 92, 5930–5934. [Google Scholar] [CrossRef] [PubMed]
- Willekens, H.; Villarroel, R.; Van Montagu, M.; Inzé, D.; Van Camp, W. Molecular identification of catalases from Nicotiana plumbaginifolia (L.). FEBS Lett. 1994, 352, 79–83. [Google Scholar] [CrossRef]
- Skadsen, R.W.; Schulze-Lefert, P.; Herbst, J.M. Molecular cloning, characterization and expression analysis of two catalase isozyme genes in barley. Plant Mol. Biol. 1995, 29, 1005–1014. [Google Scholar] [CrossRef]
- Drory, A.; Woodson, W.R. Molecular cloning and nucleotide sequence of a cDNA encoding catalase from tomato. Plant Physiol. 1992, 100, 1605–1606. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.-J.; Wu, S.-D.; Huang, G.-J.; Shen, C.-Y.; Afiyanti, M.; Li, W.-J.; Lin, Y.-H. Expression of a cloned sweet potato catalase SPCAT1 alleviates ethephon-mediated leaf senescence and H2O2 elevation. J. Plant Physiol. 2012, 169, 86–97. [Google Scholar] [CrossRef] [PubMed]
- González, E. The C-terminal domain of plant catalases Implications for a glyoxysomal targeting sequence. JBIC J. Biol. Inorg. Chem. 1991, 199, 211–215. [Google Scholar] [CrossRef]
- Raza, A.; Su, W.; Gao, A.; Mehmood, S.; Hussain, M.; Nie, W.; Lv, Y.; Zou, X.; Zhang, X. Catalase (CAT) Gene Family in Rapeseed (Brassica napus L.): Genome-Wide Analysis, Identification, and Expression Pattern in Response to Multiple Hormones and Abiotic Stress Conditions. Int. J. Mol. Sci. 2021, 22, 4281. [Google Scholar] [CrossRef] [PubMed]
- Eising, R.; Trelease, R.N.; Ni, W. Biogenesis of catalase in glyoxysomes and leaf-type peroxisomes of sunflower cotyledons. Arch. Biochem. Biophys. 1990, 278, 258–264. [Google Scholar] [CrossRef]
- Kleff, S.; Trelease, R.N.; Eising, R. Nucleotide and deduced amino acid sequence of a putative higher molecular weight precursor for catalase in sunflower cotyledons. Biochim. Biophys. Acta-Bioenerg. 1994, 1224, 463–466. [Google Scholar] [CrossRef]
- Azpilicueta, C.E.; Pena, L.B.; Tomaro, M.L.; Gallego, S.M. Modifications in catalase activity and expression in developing sunflower seedlings under cadmium stress. Redox Rep. 2008, 13, 40–46. [Google Scholar] [CrossRef]
- Ni, W.; Trelease, R.N. Post-Transcriptional Regulation of Catalase Isozyme Expression in Cotton Seeds. Plant Cell 1991, 3, 737–744. [Google Scholar] [CrossRef]
- Isin, S.H.; Allen, R.D. Isolation and characterization of a pea catalase cDNA. Plant Mol. Biol. 1991, 17, 1263–1265. [Google Scholar] [CrossRef]
- Corpas, F.J.; Palma, J.M.; Sandalio, L.M.; Lopez-Huertas, E.; Romero-Puertas, M.C.; Barroso, J.B.; Del Río, L.A. Purification of Catalase from Pea Leaf Peroxisomes: Identification of Five Different Isoforms. Free. Radic. Res. 1999, 31, 235–241. [Google Scholar] [CrossRef]
- Morita, S.; Tasaka, M.; Fujisawa, H.; Ushimaru, T.; Tsuji, H. A cDNA Clone Encoding a Rice Catalase Isozyme. Plant Physiol. 1994, 105, 1015–1016. [Google Scholar] [CrossRef]
- Alam, N.B.; Ghosh, A. Comprehensive analysis and transcript profiling of Arabidopsis thaliana and Oryza sativa catalase gene family suggests their specific roles in development and stress responses. Plant Physiol. Biochem. 2018, 123, 54–64. [Google Scholar] [CrossRef]
- Chevalier, C.; Yamaguchi, J.; McCourt, P. Nucleotide Sequence of a cDNA for Catalase from Arabidopsis thaliana. Plant Physiol. 1992, 99, 1726–1728. [Google Scholar] [CrossRef][Green Version]
- Liu, J.; Cui, L.; Xie, Z.; Zhang, Z.; Liu, E.; Peng, X. Two NCA1 isoforms interact with catalase in a mutually exclusive manner to redundantly regulate its activity in rice. BMC Plant Biol. 2019, 19, 105. [Google Scholar] [CrossRef] [PubMed]
- Contento, A.L.; Bassham, D.C. Increase in catalase-3 activity as a response to use of alternative catabolic substrates during sucrose starvation. Plant Physiol. Biochem. 2010, 48, 232–238. [Google Scholar] [CrossRef]
- Mhamdi, A.; Queval, G.; Chaouch, S.; Vanderauwera, S.; Van Breusegem, F.; Noctor, G. Catalase function in plants: A focus on Arabidopsis mutants as stress-mimic models. J. Exp. Bot. 2010, 61, 4197–4220. [Google Scholar] [CrossRef] [PubMed]
- Isin, S.H. Regulation of Catalase Gene Expression in Soybean. Ph.D. Thesis, Texas Tech. University, Lubbock, TX, USA, 1992. [Google Scholar]
- Redinbaugh, M.G.; Wadsworth, G.J.; Scandalios, J.G. Characterization of catalase transcripts and their differential expression in maize. Biochim. Biophys. Acta-Gene Struct. Expr. 1988, 951, 104–116. [Google Scholar] [CrossRef]
- Wadsworth, G.J.; Scandalios, J.G. Differential expression of the maize catalase genes during kernel development: The role of steady-state mRNA levels. Dev. Genet. 1989, 10, 304–310. [Google Scholar] [CrossRef]
- Guan, L.; Scandalios, J.G. Characterization of the catalase antioxidant defense gene Cat1 of maize, and its developmentally regulated expression in transgenic tobacco. Plant J. 1993, 3, 527–536. [Google Scholar] [CrossRef] [PubMed]
- Guan, L.; Polidoros, A.N.; Scandalios, J.G. Isolation, characterization and expression of the maize Cat2 catalase gene. Plant Mol. Biol. 1996, 30, 913–924. [Google Scholar] [CrossRef]
- Abler, M.L.; Scandalios, J.G. The CAT-2 null phenotype in maize is likely due to a DNA insertion into the Cat2 gene. Theor. Appl. Genet. 1991, 81, 635–640. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.H.; An, C.S. Differential expression of three catalase genes in hot pepper (Capsicum annuum L.). Mol. Cells 2005, 20, 247–255. [Google Scholar]
- Schultes, N.P.; Zelitch, I.; Mcgonigle, B.; Nelson, T. The Primary Leaf Catalase Gene from Nicotiana tabacum and Nicotiana sylvestris. Plant Physiol. 1994, 106, 399–400. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Niewiadomska, E.; Polzien, L.; Desel, C.; Rozpądek, P.; Miszalski, Z.; Krupinska, K. Spatial patterns of senescence and development-dependent distribution of reactive oxygen species in tobacco (Nicotiana tabacum) leaves. J. Plant Physiol. 2009, 166, 1057–1068. [Google Scholar] [CrossRef]
- Suzuki, M.; Ario, T.; Hattori, T.; Nakamura, K.; Asahi, T. Isolation and characterization of two tightly linked catalase genes from castor bean that are differentially regulated. Plant Mol. Biol. 1994, 25, 507–516. [Google Scholar] [CrossRef]
- Luna, C.M.; Pastori, G.M.; Driscoll, S.; Groten, K.; Bernard, S.; Foyer, C.H. Drought controls on H2O2 accumulation, catalase (CAT) activity and CAT gene expression in wheat. J. Exp. Bot. 2005, 56, 417–423. [Google Scholar] [CrossRef]
- Wu, G.; Shah, D.M. Isolation and characterization of a potato catalase cDNA. Plant Physiol. 1995, 108, 1748–1749. [Google Scholar]
- Almeida, J.M.; Fidalgo, F.; Confraria, A.; Santos, A.; Pires, H.; Santos, I. Effect of hydrogen peroxide on catalase gene expression, isoform activities and levels in leaves of potato sprayed with homobrassinolide and ultrastructural changes in mesophyll cells. Funct. Plant Biol. 2005, 32, 707–720. [Google Scholar] [CrossRef]
- Kerdnaimongkol, K.; Woodson, W.R. Inhibition of Catalase by Antisense RNA Increases Susceptibility to Oxidative Stress and Chilling Injury in Transgenic Tomato Plants. J. Am. Soc. Hortic. Sci. 1999, 124, 330–336. [Google Scholar] [CrossRef]
- Meena, M.; Zehra, A.; Dubey, M.K.; Aamir, M.; Gupta, V.K.; Upadhyay, R.S. Comparative Evaluation of Biochemical Changes in Tomato (Lycopersicon esculentum Mill.) Infected by Alternaria alternata and Its Toxic Metabolites (TeA, AOH, and AME). Front. Plant Sci. 2016, 7, 1408. [Google Scholar] [CrossRef] [PubMed]
- Havir, E.A.; McHale, N.A. Regulation of Catalase Activity in Leaves of Nicotiana sylvestris by High CO2. Plant Physiol. 1989, 89, 952–957. [Google Scholar] [CrossRef] [PubMed]
- Mori, H.; Imaseki, H. cDNA for Catalase from Etiolated Mung Bean (Vigna radiata) Hypocotyls. Plant Physiol. 1993, 102, 691–692. [Google Scholar] [CrossRef]
- Schmidt, M.; Grief, J.; Feierabend, J. Mode of translational activation of the catalase (cat1) mRNA of rye leaves (Secale cereale L.) and its control through blue light and reactive oxygen. Planta 2006, 223, 835–846. [Google Scholar] [CrossRef]
- Sakajo, S.; Nakamura, K.; Asahi, T. Molecular cloning and nucleotide sequence of full-length cDNA for sweet potato catalase mRNA. JBIC J. Biol. Inorg. Chem. 1987, 165, 437–442. [Google Scholar] [CrossRef]
- Su, T.; Wang, P.; Li, H.; Zhao, Y.; Lu, Y.; Dai, P.; Ren, T.; Wang, X.; Li, X.; Shao, Q.; et al. The Arabidopsiscatalase triple mutant reveals important roles of catalases and peroxisome-derived signaling in plant development. J. Integr. Plant Biol. 2018, 60, 591–607. [Google Scholar] [CrossRef]
- Yang, Z.; Mhamdi, A.; Noctor, G. Analysis of catalase mutants underscores the essential role of CATALASE2 for plant growth and day length-dependent oxidative signalling. Plant Cell Environ. 2018, 42, 688–700. [Google Scholar] [CrossRef] [PubMed]
- Palma, J.M.; Mateos, R.M.; Lopez-Jaramillo, F.J.; Ruiz, M.R.; González-Gordo, S.; Lechuga-Sancho, A.; Corpas, F.J. Plant catalases as NO and H2S targets. Redox Biol. 2020, 34, 101525. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Li, C.; Ren, H.; Zhao, T.; Li, Q.; Wang, S.; Zhang, Y.; Xiao, F.; Wang, X. BAK1 Mediates Light Intensity to Phosphorylate and Activate Catalases to Regulate Plant Growth and Development. Int. J. Mol. Sci. 2020, 21, 1437. [Google Scholar] [CrossRef]
- Zhang, Y.; Ji, T.-T.; Li, T.-T.; Tian, Y.-Y.; Wang, L.-F.; Liu, W.-C. Jasmonic acid promotes leaf senescence through MYC2-mediated repression of CATALASE2 expression in Arabidopsis. Plant Sci. 2020, 299, 110604. [Google Scholar] [CrossRef]
- Wang, L.; Chen, W.J.; Wang, Q.; Eneji, A.E.; Li, Z.H.; Duan, L.S. Coronatine Enhances Chilling Tolerance in Cucumber (Cucumis sativus L.) Seedlings by Improving the Antioxidative Defence System. J. Agron. Crop. Sci. 2009, 195, 377–383. [Google Scholar] [CrossRef]
- Mylona, P.; Polidoros, A.; Scandalios, J.G. Modulation of antioxidant responses by arsenic in maize. Free Radic. Biol. Med. 1998, 25, 576–585. [Google Scholar] [CrossRef]
- Vandenabeele, S.; Vanderauwera, S.; Vuylsteke, M.; Rombauts, S.; Langebartels, C.; Seidlitz, H.K.; Zabeau, M.; Van Montagu, M.; Inzé, D.; Van Breusegem, F. Catalase deficiency drastically affects gene expression induced by high light in Arabidopsis thaliana. Plant J. 2004, 39, 45–58. [Google Scholar] [CrossRef]
- Xing, Y.; Jia, W.; Zhang, J. AtMEK1 mediates stress-induced gene expression of CAT1 catalase by triggering H2O2 production in Arabidopsis. J. Exp. Bot. 2007, 58, 2969–2981. [Google Scholar] [CrossRef]
- Corpas, F.J.; Barroso, J.B. Lead-induced stress, which triggers the production of nitric oxide (NO) and superoxide anion (O2−) in Arabidopsis peroxisomes, affects catalase activity. Nitric Oxide 2017, 68, 103–110. [Google Scholar] [CrossRef] [PubMed]
- Ono, M.; Isono, K.; Sakata, Y.; Taji, T. CATALASE2 plays a crucial role in long-term heat tolerance of Arabidopsis thaliana. Biochem. Biophys. Res. Commun. 2020, 534, 747–751. [Google Scholar] [CrossRef] [PubMed]
- Bueso, E.; Alejandro, S.; Carbonell, P.; Perez-Amador, M.A.; Fayos, J.; Bellés, J.M.; Rodriguez, P.L.; Serrano, R.; Carbonell-Bejerano, P. The lithium tolerance of the Arabidopsiscat2mutant reveals a cross-talk between oxidative stress and ethylene. Plant J. 2007, 52, 1052–1065. [Google Scholar] [CrossRef]
- Zou, J.-J.; Li, X.-D.; Ratnasekera, D.; Wang, C.; Liu, W.-X.; Song, L.-F.; Zhang, W.-Z.; Wu, W.-H. Arabidopsis calcium-dependent protein kinase8 and CATALASE3 Function in Abscisic Acid-Mediated Signaling and H2O2 Homeostasis in Stomatal Guard Cells under Drought Stress. Plant Cell 2015, 27, 1445–1460. [Google Scholar] [CrossRef] [PubMed]
- Sharma, P.; Dubey, R. Ascorbate peroxidase from rice seedlings: Properties of enzyme isoforms, effects of stresses and protective roles of osmolytes. Plant Sci. 2004, 167, 541–550. [Google Scholar] [CrossRef]
- Maruta, T.; Inoue, T.; Noshi, M.; Tamoi, M.; Yabuta, Y.; Yoshimura, K.; Ishikawa, T.; Shigeoka, S. Cytosolic ascorbate peroxidase 1 protects organelles against oxidative stress by wounding-and jasmonate-induced H2O2 in Arabidopsis plants. Biochim. Biophys. Acta-Gen. Subj. 2012, 1820, 1901–1907. [Google Scholar] [CrossRef]
- Maruta, T.; Sawa, Y.; Shigeoka, S.; Ishikawa, T. Diversity and Evolution of Ascorbate Peroxidase Functions in Chloroplasts: More Than Just a Classical Antioxidant Enzyme? Plant Cell Physiol. 2016, 57, pcv203. [Google Scholar] [CrossRef] [PubMed]
- Chew, O.; Whelan, J.; Millar, A.H. Molecular Definition of the Ascorbate-Glutathione Cycle in Arabidopsis Mitochondria Reveals Dual Targeting of Antioxidant Defenses in Plants. J. Biol. Chem. 2003, 278, 46869–46877. [Google Scholar] [CrossRef]
- Granlund, I.; Storm, P.; Schubert, M.; García-Cerdán, J.G.; Funk, C.; Schröder, W.P. The TL29 Protein is Lumen Located, Associated with PSII and Not an Ascorbate Peroxidase. Plant Cell Physiol. 2009, 50, 1898–1910. [Google Scholar] [CrossRef] [PubMed]
- Van Buer, J.; Cvetkovic, J.; Baier, M. Cold regulation of plastid ascorbate peroxidases serves as a priming hub controlling ROS signaling in Arabidopsis thaliana. BMC Plant Biol. 2016, 16, 163. [Google Scholar] [CrossRef]
- Rossel, J.B.; Walter, P.B.; Hendrickson, L.; Chow, W.S.; Poole, A.; Mullineaux, P.M.; Pogson, B.J. A mutation affecting ascorbate peroxidase 2 gene expression reveals a link between responses to high light and drought tolerance. Plant Cell Environ. 2005, 29, 269–281. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, N.; Miller, G.; Sejima, H.; Harper, J.; Mittler, R. Enhanced seed production under prolonged heat stress conditions inArabidopsis thalianaplants deficient in cytosolic ascorbate peroxidase 2. J. Exp. Bot. 2012, 64, 253–263. [Google Scholar] [CrossRef]
- Pnueli, L.; Liang, H.; Rozenberg, M.; Mittler, R. Growth suppression, altered stomatal responses, and augmented induction of heat shock proteins in cytosolic ascorbate peroxidase (Apx1)-deficient Arabidopsis plants. Plant J. 2003, 34, 187–203. [Google Scholar] [CrossRef]
- Davletova, S.; Rizhsky, L.; Liang, H.; Shengqiang, Z.; Oliver, D.J.; Coutu, J.; Shulaev, V.; Schlauch, K.; Mittler, R. Cytosolic Ascorbate Peroxidase 1 Is a Central Component of the Reactive Oxygen Gene Network of Arabidopsis. Plant Cell 2005, 17, 268–281. [Google Scholar] [CrossRef]
- Pandey, S.; Fartyal, D.; Agarwal, A.; Shukla, T.; James, D.; Kaul, T.; Negi, Y.K.; Arora, S.; Reddy, M.K. Abiotic Stress Tolerance in Plants: Myriad Roles of Ascorbate Peroxidase. Front. Plant Sci. 2017, 8, 581. [Google Scholar] [CrossRef]
- Chin, D.-C.; Kumar, R.S.; Suen, C.-S.; Chien, C.-Y.; Hwang, M.-J.; Hsu, C.-H.; Xuhan, X.; Lai, Z.X.; Yeh, K.-W. Plant Cytosolic Ascorbate Peroxidase with Dual Catalytic Activity Modulates Abiotic Stress Tolerances. iScience 2019, 16, 31–49. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Letnik, I.; Hacham, Y.; Dobrev, P.; Ben-Daniel, B.-H.; Vanková, R.; Amir, R.; Miller, G. ASCORBATE PEROXIDASE6 Protects Arabidopsis Desiccating and Germinating Seeds from Stress and Mediates Cross Talk between Reactive Oxygen Species, Abscisic Acid, and Auxin. Plant Physiol. 2014, 166, 370–383. [Google Scholar] [CrossRef]
- Parida, A.K.; Das, A.B. Salt tolerance and salinity effects on plants: A review. Ecotoxicol. Environ. Saf. 2005, 60, 324–349. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, M.; Ma, L.Q.; Singh, N.; Singh, S. Antioxidant responses of hyper-accumulator and sensitive fern species to arsenic. J. Exp. Bot. 2005, 56, 1335–1342. [Google Scholar] [CrossRef] [PubMed]
- Asada, K. Ascorbate peroxidase—A hydrogen peroxide-scavenging enzyme in plants. Physiol. Plant. 1992, 85, 235–241. [Google Scholar] [CrossRef]
- Caverzan, A.; Casassola, A.; Brammer, S.P. Reactive oxygen species and antioxidant enzymes involved in plant tolerance to stress. In Abiotic and Biotic Stress in Plants-Recent Advances and Future Perspectives; Shanker, A.K., Shanker, C., Eds.; IntechOpen: London, UK, 2016. [Google Scholar] [CrossRef]
- Asada, K. The water-water cycle in chloroplasts: Scavenging of Active Oxygens and Dissipation of Excess Photons. Annu. Rev. Plant Biol. 1999, 50, 601–639. [Google Scholar] [CrossRef]
- Sharma, P.; Jha, A.B.; Dubey, R.S.; Pessarakli, M. Reactive Oxygen Species, Oxidative Damage, and Antioxidative Defense Mechanism in Plants under Stressful Conditions. J. Bot. 2012, 2012, 217037. [Google Scholar] [CrossRef]
- Passardi, F.; Longet, D.; Penel, C.; Dunand, C. The class III peroxidase multigenic family in rice and its evolution in land plants. Phytochemistry 2004, 65, 1879–1893. [Google Scholar] [CrossRef]
- Ivanov, S.; Shopova, E.; Kerchev, P.; Sergiev, I.; Miteva, L.; Polizoev, D.; Alexieva, V. Long-term impact of sublethal atrazine perturbs the redox homeostasis in pea (Pisum sativum L.) plants. Protoplasma 2012, 250, 95–102. [Google Scholar] [CrossRef]
- Mishra, S.; Jha, A.B.; Dubey, R.S. Arsenite treatment induces oxidative stress, upregulates antioxidant system, and causes phytochelatin synthesis in rice seedlings. Protoplasma 2010, 248, 565–577. [Google Scholar] [CrossRef]
- Srivastava, S.; Dubey, R.S. Manganese-excess induces oxidative stress, lowers the pool of antioxidants and elevates activities of key antioxidative enzymes in rice seedlings. Plant Growth Regul. 2010, 64, 1–16. [Google Scholar] [CrossRef]
- Song, H.; Wang, Y.-S.; Sun, C.-C.; Wang, Y.-T.; Peng, Y.-L.; Cheng, H. Effects of pyrene on antioxidant systems and lipid peroxidation level in mangrove plants, Bruguiera gymnorrhiza. Ecotoxicology 2012, 21, 1625–1632. [Google Scholar] [CrossRef]
- Li, C.; Wang, T.; Li, Y.; Zheng, Y.-H.; Jiang, G.-M. Flixweed Is More Competitive than Winter Wheat under Ozone Pollution: Evidences from Membrane Lipid Peroxidation, Antioxidant Enzymes and Biomass. PLoS ONE 2013, 8, e60109. [Google Scholar] [CrossRef] [PubMed]
- Kataya, A.R.A.; Reumann, S. Arabidopsis glutathione reductase 1 is dually targeted to peroxisomes and the cytosol. Plant Signal. Behav. 2010, 5, 171–175. [Google Scholar] [CrossRef]
- Berkholz, D.S.; Faber, H.R.; Savvides, S.N.; Karplus, P.A. Catalytic Cycle of Human Glutathione Reductase Near 1 Å Resolution. J. Mol. Biol. 2008, 382, 371–384. [Google Scholar] [CrossRef]
- Karpinski, S.; Escobar, C.; Karpinska, B.; Creissen, G.; Mullineaux, P.M. Photosynthetic electron transport regulates the expression of cytosolic ascorbate peroxidase genes in Arabidopsis during excess light stress. Plant Cell 1997, 9, 627–640. [Google Scholar] [CrossRef]
- Müller-Schüssele, S.J.; Wang, R.; Gütle, D.D.; Romer, J.; Rodriguez-Franco, M.; Scholz, M.; Buchert, F.; Lüth, V.M.; Kopriva, S.; Dörmann, P.; et al. Chloroplasts require glutathione reductase to balance reactive oxygen species and maintain efficient photosynthesis. Plant J. 2020, 103, 1140–1154. [Google Scholar] [CrossRef]
- Csiszár, J.; Brunner, S.; Horváth, E.; Bela, K.; Ködmön, P.; Riyazuddin, R.; Gallé, Á.; Hurton, Á.; Papdi, C.; Szabados, L.; et al. Exogenously applied salicylic acid maintains redox homeostasis in salt-stressed Arabidopsis gr1 mutants expressing cytosolic roGFP1. Plant Growth Regul. 2018, 86, 181–194. [Google Scholar] [CrossRef]
- Wang, Q.; Pu, Y.; Yang, D.; Yin, X.; He, Z.; Yang, Y.; Yang, Y. Molecular cloning and characterization of the glutathione reductase gene from Stipa purpurea. Biochem. Biophys. Res. Commun. 2018, 495, 1851–1857. [Google Scholar] [CrossRef]
- Kornyeyev, D.; Logan, B.A.; Payton, P.R.; Allen, R.D.; Holaday, A.S. Elevated chloroplastic glutathione reductase activities decrease chilling-induced photoinhibition by increasing rates of photochemistry, but not thermal energy dissipation, in transgenic cotton. Funct. Plant Biol. 2003, 30, 101–110. [Google Scholar] [CrossRef] [PubMed]
- Logan, B.A.; Monteiro, G.; Kornyeyev, D.; Payton, P.; Allen, R.D.; Holaday, A.S. Transgenic overproduction of glutathione reductase does not protect cotton, Gossypium hirsutum (Malvaceae), from photoinhibition during growth under chilling conditions. Am. J. Bot. 2003, 90, 1400–1403. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Ding, H.; Chen, Q.; Ouyang, L.; Li, S.; Zhang, J. Enhanced Tolerance to Methyl Viologen-Mediated Oxidative Stress via AtGR2 Expression From Chloroplast Genome. Front. Plant Sci. 2019, 10, 1178. [Google Scholar] [CrossRef]
- Guo, B.; Liu, C.; Li, H.; Yi, K.; Ding, N.; Li, N.; Lin, Y.; Fu, Q. Endogenous salicylic acid is required for promoting cadmium tolerance of Arabidopsis by modulating glutathione metabolisms. J. Hazard. Mater. 2016, 316, 77–86. [Google Scholar] [CrossRef] [PubMed]
- Yin, L.; Mano, J.; Tanaka, K.; Wang, S.; Zhang, M.; Deng, X.; Zhang, S. High level of reduced glutathione contributes to detoxification of lipid peroxide-derived reactive carbonyl species in transgenic Arabidopsis overexpressing glutathione reductase under aluminum stress. Physiol. Plant. 2017, 161, 211–223. [Google Scholar] [CrossRef] [PubMed]
- Marty, L.; Bausewein, D.; Müller, C.; Bangash, S.A.K.; Moseler, A.; Schwarzländer, M.; Müller-Schüssele, S.J.; Zechmann, B.; Riondet, C.; Balk, J.; et al. Arabidopsis glutathione reductase 2 is indispensable in plastids, while mitochondrial glutathione is safeguarded by additional reduction and transport systems. New Phytol. 2019, 224, 1569–1584. [Google Scholar] [CrossRef] [PubMed]
- Sumugat, M.R.; Donahue, J.L.; Cortes, D.F.; Stromberg, V.K.; Grene, R.; Shulaev, V.; Welbaum, G.E. Seed Development and Germination in anArabidopsis thalianaLine Antisense to Glutathione Reductase 2. J. New Seeds 2010, 11, 104–126. [Google Scholar] [CrossRef]
- Yu, X.; Pasternak, T.; Eiblmeier, M.; Ditengou, F.; Kochersperger, P.; Sun, J.; Wang, H.; Rennenberg, H.; Teale, W.; Paponov, I.; et al. Plastid-Localized Glutathione Reductase2-Regulated Glutathione Redox Status Is Essential for Arabidopsis Root Apical Meristem Maintenance. Plant Cell 2013, 25, 4451–4468. [Google Scholar] [CrossRef] [PubMed]
- Jiang, M. Water stress-induced abscisic acid accumulation triggers the increased generation of reactive oxygen species and up-regulates the activities of antioxidant enzymes in maize leaves. J. Exp. Bot. 2002, 53, 2401–2410. [Google Scholar] [CrossRef]
- Romero-Puertas, M.C.; Corpas, F.J.; Sandalio, L.M.; Leterrier, M.; Rodriguez-Serrano, M.; Rio, L.A.; Palma, J.M. Glutathione reductase from pea leaves: Response to abiotic stress and characterization of the peroxisomal isozyme. New Phytol. 2006, 170, 43–52. [Google Scholar] [CrossRef]
- Kuźniak, E.; Skłodowska, M. Ascorbate, glutathione and related enzymes in chloroplasts of tomato leaves infected by Botrytis cinerea. Plant Sci. 2001, 160, 723–731. [Google Scholar] [CrossRef]
- Gill, S.S.; Anjum, N.A.; Hasanuzzaman, M.; Gill, R.; Trivedi, D.K.; Ahmad, I.; Pereira, E.; Tuteja, N. Glutathione and glutathione reductase: A boon in disguise for plant abiotic stress defense operations. Plant Physiol. Biochem. 2013, 70, 204–212. [Google Scholar] [CrossRef]
- Sudhakar, C.; Lakshmi, A.; Giridarakumar, S. Changes in the antioxidant enzyme efficacy in two high yielding genotypes of mulberry (Morus alba L.) under NaCl salinity. Plant Sci. 2001, 161, 613–619. [Google Scholar] [CrossRef]
- Bandeoğlu, E.; Eyidoğan, F.; Yücel, M.; Öktem, H.A. Antioxidant responses of shoots and roots of lentil to NaCl-salinity stress. Plant Growth Regul. 2004, 42, 69–77. [Google Scholar] [CrossRef]
- Wang, L.-J.; Li, S.-H. Salicylic acid-induced heat or cold tolerance in relation to Ca2+ homeostasis and antioxidant systems in young grape plants. Plant Sci. 2006, 170, 685–694. [Google Scholar] [CrossRef]
- Hossain, M.A.; Asada, K. Monodehydroascorbate reductase from cucumber is a flavin adenine dinucleotide enzyme. J. Biol. Chem. 1985, 260, 12920–12926. [Google Scholar] [CrossRef]
- Suman, S.; Bagal, D.; Jain, D.; Singh, R.; Singh, I.K.; Singh, A. Biotic stresses on plants: Reactive oxygen species generation and antioxidant mechanism, Chapter 14. In Frontiers in Plant-Soil Interactions; Aftab, T., Hakeem, K.R., Eds.; Academic Press: New York, NY, USA, 2021; pp. 381–411. [Google Scholar] [CrossRef]
- Hasanuzzaman, M.; Bhuyan, M.H.M.B.; Anee, T.I.; Parvin, K.; Nahar, K.; Mahmud, J.A.; Fujita, M. Regulation of Ascorbate-Glutathione Pathway in Mitigating Oxidative Damage in Plants under Abiotic Stress. Antioxidants 2019, 8, 384. [Google Scholar] [CrossRef]
- Obara, K.; Sumi, K.; Fukuda, H. The use of multiple transcription starts causes the dual targeting of Arabidopsis putative monodehydroascorbate reductase to both mitochondria and chloroplasts. Plant Cell Physiol. 2002, 43, 697–705. [Google Scholar] [CrossRef] [PubMed]
- Kaur, N.; Hu, J. Defining the Plant Peroxisomal Proteome: From Arabidopsis to Rice. Front. Plant Sci. 2011, 2, 103. [Google Scholar] [CrossRef]
- Eastmond, P.J. Monodehyroascorbate reductase4 is Required for Seed Storage Oil Hydrolysis and Postgerminative Growth in Arabidopsis. Plant Cell 2007, 19, 1376–1387. [Google Scholar] [CrossRef] [PubMed]
- Eltayeb, A.E.; Kawano, N.; Badawi, G.H.; Kaminaka, H.; Sanekata, T.; Shibahara, T.; Inanaga, S.; Tanaka, K. Overexpression of monodehydroascorbate reductase in transgenic tobacco confers enhanced tolerance to ozone, salt and polyethylene glycol stresses. Planta 2007, 225, 1255–1264. [Google Scholar] [CrossRef]
- Sultana, S.; Khew, C.Y.; Morshed, M.; Namasivayam, P.; Napis, S.; Ho, C.-L. Overexpression of monodehydroascorbate reductase from a mangrove plant (AeMDHAR) confers salt tolerance on rice. J. Plant Physiol. 2011, 169, 311–318. [Google Scholar] [CrossRef]
- Rahantaniaina, M.-S.; Li, S.; Chatel-Innocenti, G.; Tuzet, A.; Issakidis-Bourguet, E.; Mhamdi, A.; Noctor, G. Cytosolic and Chloroplastic DHARs Cooperate in Oxidative Stress-Driven Activation of the Salicylic Acid Pathway. Plant Physiol. 2017, 174, 956–971. [Google Scholar] [CrossRef]
- Chen, Z.; Gallie, D.R. Dehydroascorbate Reductase Affects Leaf Growth, Development, and Function. Plant Physiol. 2006, 142, 775–787. [Google Scholar] [CrossRef]
- Ding, H.; Wang, B.; Han, Y.; Li, S. The pivotal function of dehydroascorbate reductase in glutathione homeostasis in plants. J. Exp. Bot. 2020, 71, 3405–3416. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Xiao, Y.; Chen, W.; Tang, K.; Zhang, L. Increased Vitamin C Content Accompanied by an Enhanced Recycling Pathway Confers Oxidative Stress Tolerance in Arabidopsis. J. Integr. Plant Biol. 2010, 52, 400–409. [Google Scholar] [CrossRef]
- Noshi, M.; Yamada, H.; Hatanaka, R.; Tanabe, N.; Tamoi, M.; Shigeoka, S. Arabidopsis dehydroascorbate reductase 1 and 2 modulate redox states of ascorbate-glutathione cycle in the cytosol in response to photooxidative stress. Biosci. Biotechnol. Biochem. 2017, 81, 523–533. [Google Scholar] [CrossRef]
- Yoshida, S.; Tamaoki, M.; Shikano, T.; Nakajima, N.; Ogawa, D.; Ioki, M.; Aono, M.; Kubo, A.; Kamada, H.; Inoue, Y.; et al. Cytosolic Dehydroascorbate Reductase is Important for Ozone Tolerance in Arabidopsisthaliana. Plant Cell Physiol. 2006, 47, 304–308. [Google Scholar] [CrossRef]
- Noshi, M.; Hatanaka, R.; Tanabe, N.; Terai, Y.; Maruta, T.; Shigeoka, S. Redox regulation of ascorbate and glutathione by a chloroplastic dehydroascorbate reductase is required for high-light stress tolerance in Arabidopsis. Biosci. Biotechnol. Biochem. 2016, 80, 870–877. [Google Scholar] [CrossRef]
- Sánchez-Viveros, G.; Ferrera-Cerrato, R.; Alarcón, A. Short-Term Effects of Arsenate-Induced Toxicity on Growth, Chlorophyll and Carotenoid Contents, and Total Content of Phenolic Compounds of Azolla filiculoides. Water Air Soil Pollut. 2010, 217, 455–462. [Google Scholar] [CrossRef]
- Do, H.; Kim, I.-S.; Jeon, B.W.; Lee, C.W.; Park, A.K.; Wi, A.R.; Shin, S.C.; Park, H.; Kim, Y.-S.; Yoon, H.-S.; et al. Structural understanding of the recycling of oxidized ascorbate by dehydroascorbate reductase (OsDHAR) from Oryza sativa L. japonica. Sci. Rep. 2016, 6, 19498. [Google Scholar] [CrossRef] [PubMed]
- Dell’Aglio, E.; Mahmdi, A. What are the Roles for Dehydroascorbate Reductases and Glutathione in Sustaining Ascorbate Accumulation? Available online: https://plantae.org/what-are-the-roles-for-dehydroascorbate-reductases-and-glutathione-in-sustaining-ascorbate-accumulation (accessed on 10 October 2021).
- Chen, Z.; Young, T.E.; Ling, J.; Chang, S.-C.; Gallie, D.R. Increasing vitamin C content of plants through enhanced ascorbate recycling. Proc. Natl. Acad. Sci. USA 2003, 100, 3525–3530. [Google Scholar] [CrossRef]
- Smirnoff, N. Ascorbic acid metabolism and functions: A comparison of plants and mammals. Free Radic. Biol. Med. 2018, 122, 116–129. [Google Scholar] [CrossRef]
- Potters, G.; De Gara, L.; Asard, H.; Horemans, N. Ascorbate and glutathione: Guardians of the cell cycle, partners in crime? Plant Physiol. Biochem. 2002, 40, 537–548. [Google Scholar] [CrossRef]
- Ushimaru, T.; Nakagawa, T.; Fujioka, Y.; Daicho, K.; Naito, M.; Yamauchi, Y.; Nonaka, H.; Amako, K.; Yamawaki, K.; Murata, N. Transgenic Arabidopsis plants expressing the rice dehydroascorbate reductase gene are resistant to salt stress. J. Plant Physiol. 2006, 163, 1179–1184. [Google Scholar] [CrossRef]
- Yin, L.; Wang, S.; Eltayeb, A.E.; Uddin, I.; Yamamoto, Y.; Tsuji, W.; Takeuchi, Y.; Tanaka, K. Overexpression of dehydroascorbate reductase, but not monodehydroascorbate reductase, confers tolerance to aluminum stress in transgenic tobacco. Planta 2009, 231, 609–621. [Google Scholar] [CrossRef] [PubMed]
- Eltelib, H.A.; Badejo, A.A.; Fujikawa, Y.; Esaka, M. Gene expression of monodehydroascorbate reductase and dehydroascorbate reductase during fruit ripening and in response to environmental stresses in acerola (Malpighia glabra). J. Plant Physiol. 2011, 168, 619–627. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.-S.; Kim, I.-S.; Bae, M.-J.; Choe, Y.-H.; Kim, Y.-H.; Park, H.-M.; Kang, H.-G.; Yoon, H.-S. Homologous expression of cytosolic dehydroascorbate reductase increases grain yield and biomass under paddy field conditions in transgenic rice (Oryza sativa L. japonica). Planta 2013, 237, 1613–1625. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Gallie, D.R. Dehydroascorbate Reductase Affects Non-photochemical Quenching and Photosynthetic Performance. J. Biol. Chem. 2008, 283, 21347–21361. [Google Scholar] [CrossRef]
- Dowdle, J.; Ishikawa, T.; Gatzek, S.; Rolinski, S.; Smirnoff, N. Two genes in Arabidopsis thaliana encoding GDP-l-galactose phosphorylase are required for ascorbate biosynthesis and seedling viability. Plant J. 2007, 52, 673–689. [Google Scholar] [CrossRef]
- Terai, Y.; Ueno, H.; Ogawa, T.; Sawa, Y.; Miyagi, A.; Kawai-Yamada, M.; Ishikawa, T.; Maruta, T. Dehydroascorbate Reductases and Glutathione Set a Threshold for High-Light–Induced Ascorbate Accumulation. Plant Physiol. 2020, 183, 112–122. [Google Scholar] [CrossRef]
- Han, Y.; Chaouch, S.; Mhamdi, A.; Queval, G.; Zechmann, B.; Noctor, G. Functional Analysis of Arabidopsis Mutants Points to Novel Roles for Glutathione in Coupling H2O2 to Activation of Salicylic Acid Accumulation and Signaling. Antioxid. Redox Signal. 2013, 18, 2106–2121. [Google Scholar] [CrossRef]
- Niak, E.B.K. Ascorbate and ascorbate-dependent enzymes in detached tomato leaves under conditions modulating the ascorbate pool. Acta Physiol. Plant. 2004, 26, 1–6. [Google Scholar] [CrossRef]
- Liebler, D.C.; Kling, D.S.; Reed, D.J. Antioxidant protection of phospholipid bilayers by alpha-tocopherol. Control of α-tocopherol status and lipid peroxidation by ascorbic acid and glutathione. J. Biol. Chem. 1986, 261, 12114–12119. [Google Scholar] [CrossRef]
- Szarka, A.; Tomasskovics, B.; Bánhegyi, G. The ascorbate-glutathione-α-tocopherol triad in abiotic stress response. Int. J. Mol. Sci. 2012, 13, 4458–4483. [Google Scholar] [CrossRef] [PubMed]
- Mullineaux, P.M.; Rausch, T. Glutathione, photosynthesis and the redox regulation of stress-responsive gene expression. Photosynth. Res. 2005, 86, 459–474. [Google Scholar] [CrossRef] [PubMed]
- Roychoudhury, A.; Pradhan, S.; Chaudhuri, B.; Das, K. Phytoremediation of toxic metals and the involvement of Brassica species. In Phytotechnologies: Remediation of Environmental Contaminants; Anjum, N.A., Pereira, M.E., Ahmad, I., Duarte, A.C., Umar, S., Khan, N.A., Eds.; CRC Press: Boca Raton, FL, USA, 2012; pp. 219–251. [Google Scholar]
- RoyChoudhury, A.; Das, K.; Ghosh, S.; Mukherjee, R.N.; Banerjee, R. Transgenic plants: Benefits and controversies. J. Bot. Soc. Bengal 2012, 66, 29–35. [Google Scholar]
- Liang, X.; Zhang, L.; Natarajan, S.K.; Becker, D.F. Proline Mechanisms of Stress Survival. Antioxid. Redox Signal. 2013, 19, 998–1011. [Google Scholar] [CrossRef]
- Szabados, L.; Savouré, A. Proline: A multifunctional amino acid. Trends Plant Sci. 2010, 15, 89–97. [Google Scholar] [CrossRef] [PubMed]
- Trelstad, R.L.; Lawley, K.R.; Holmes, L.B. Nonenzymatic hydroxylations of proline and lysine by reduced oxygen derivatives. Nature 1981, 289, 310–312. [Google Scholar] [CrossRef]
- Kaul, S.; Sharma, S.S.; Mehta, I.K. Free radical scavenging potential of L-proline: Evidence from in vitro assays. Amino Acids 2008, 34, 315–320. [Google Scholar] [CrossRef]
- Sharma, S.S.; Dietz, K.-J. The relationship between metal toxicity and cellular redox imbalance. Trends Plant Sci. 2009, 14, 43–50. [Google Scholar] [CrossRef] [PubMed]
- Floyd, R.A.; Nagy, I.Z. Formation of long-lived hydroxyl free radical adducts of proline and hydroxyproline in a fenton reaction. Biochim. Biophys. Acta-Protein Struct. Mol. Enzym. 1984, 790, 94–97. [Google Scholar] [CrossRef]
- Saradhi, P.P.; Mohanty, P. Involvement of proline in protecting thylakoid membranes against free radical-induced photodamage. J. Photochem. Photobiol. B Biol. 1997, 38, 253–257. [Google Scholar] [CrossRef]
- Matysik, J.; Alia Bhalu, B.; Mohanty, P. Molecular mechanisms of quenching of reactive oxygen species by proline under stress in plants. Curr. Sci. 2002, 82, 525–532. [Google Scholar]
- Sharma, S.S.; Schat, H.; Vooijs, R. In vitro alleviation of heavy metal-induced enzyme inhibition by proline. Phytochemistry 1998, 49, 1531–1535. [Google Scholar] [CrossRef]
- Farago, M.; Mullen, W. Plants which accumulate metals. Part IV. A possible copper-proline complex from the roots of armeria maritima. Inorg. Chim. Acta 1979, 32, L93–L94. [Google Scholar] [CrossRef]
- Chen, C.; Dickman, M.B. From The Cover: Proline suppresses apoptosis in the fungal pathogen Colletotrichum trifolii. Proc. Natl. Acad. Sci. USA 2005, 102, 3459–3464. [Google Scholar] [CrossRef] [PubMed]
- Hoque, A.; Okuma, E.; Banu, M.N.A.; Nakamura, Y.; Shimoishi, Y.; Murata, Y. Exogenous proline mitigates the detrimental effects of salt stress more than exogenous betaine by increasing antioxidant enzyme activities. J. Plant Physiol. 2007, 164, 553–561. [Google Scholar] [CrossRef]
- Xu, J.; Yin, H.; Li, X. Protective effects of proline against cadmium toxicity in micropropagated hyperaccumulator, Solanum nigrum L. Plant Cell Rep. 2008, 28, 325–333. [Google Scholar] [CrossRef] [PubMed]
- Hoque, A.; Banu, M.N.A.; Nakamura, Y.; Shimoishi, Y.; Murata, Y. Proline and glycinebetaine enhance antioxidant defense and methylglyoxal detoxification systems and reduce NaCl-induced damage in cultured tobacco cells. J. Plant Physiol. 2008, 165, 813–824. [Google Scholar] [CrossRef]
- Islam, M.M.; Hoque, A.; Okuma, E.; Jannat, R.; Banu, M.N.A.; Jahan, S.; Nakamura, Y.; Murata, Y. Proline and Glycinebetaine Confer Cadmium Tolerance on Tobacco Bright Yellow-2 Cells by Increasing Ascorbate-Glutathione Cycle Enzyme Activities. Biosci. Biotechnol. Biochem. 2009, 73, 2320–2323. [Google Scholar] [CrossRef] [PubMed]
- Diplock, T.; Machlin, L.J.; Packer, L.; Pryor, W.A. Vitamin E: Biochemistry and Health Implications; Annals of the New York Academy of Sciences: New York, NY, USA, 1989; pp. 372–378. [Google Scholar]
- Kamal-Eldin, A.; Appelqvist, L.-Å. The chemistry and antioxidant properties of tocopherols and tocotrienols. Lipids 1996, 31, 671–701. [Google Scholar] [CrossRef]
- Kiffin, R.; Bandyopadhyay, U.; Cuervo, A.M. Oxidative Stress and Autophagy. Antioxid. Redox Signal. 2006, 8, 152–162. [Google Scholar] [CrossRef] [PubMed]
- Lichtenthaler, H.K.; Prenzel, U.; Douce, R.; Joyard, J. Localization of prenylquinones in the envelope of spinach chloroplasts. Biochim. Biophys. Acta-Biomembr. 1981, 641, 99–105. [Google Scholar] [CrossRef]
- Soll, J.; Schultz, G.; Joyard, J.; Douce, R.; Block, M.A. Localization and synthesis of prenylquinones in isolated outer and inner envelope membranes from spinach chloroplasts. Arch. Biochem. Biophys. 1985, 238, 290–299. [Google Scholar] [CrossRef]
- Kanwischer, M.; Porfirova, S.; Bergmuüller, E.; Doörmann, P. Alterations in Tocopherol Cyclase Activity in Transgenic and Mutant Plants of Arabidopsis Affect Tocopherol Content, Tocopherol Composition, and Oxidative Stress. Plant Physiol. 2005, 137, 713–723. [Google Scholar] [CrossRef]
- Shinozaki, K. A novel cis-acting element in an Arabidopsis gene is involved in responsiveness to drought, low-temperature, or high-salt stress. Plant Cell 1994, 6, 251–264. [Google Scholar] [CrossRef]
- Munné-Bosch, S.; Schwarz, K.; Alegre, L. Enhanced Formation of α-Tocopherol and Highly Oxidized Abietane Diterpenes in Water-Stressed Rosemary Plants. Plant Physiol. 1999, 121, 1047–1052. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Liu, X.; Li, X.; Chen, S.; Jin, Z.; Liu, G. Overexpression of VTE1 from Arabidopsis resulting in high vitamin E accumu-lation and salt stress tolerance increase in tobacco plant. Chin. J. Appl. Environ. Biol. 2006, 12, 468–471. (In Chinese) [Google Scholar]
- Bafeel, S.O.; Ibrahim, M.M. Antioxidants and accumulation of α-tocopherol induce chilling tolerance in Medicago sativa. Int. J. Agric. Biol. 2008, 10, 593–598. [Google Scholar]
- Li, Y.; Zhou, Y.; Wang, Z.; Sun, X.; Tang, K. Engineering tocopherol biosynthetic pathway in Arabidopsis leaves and its effect on antioxidant metabolism. Plant Sci. 2010, 178, 312–320. [Google Scholar] [CrossRef]
- Fritsche, S.; Wang, X.; Jung, C. Recent Advances in our Understanding of Tocopherol Biosynthesis in Plants: An Overview of Key Genes, Functions, and Breeding of Vitamin E Improved Crops. Antioxidants 2017, 6, 99. [Google Scholar] [CrossRef]
- DellaPenna, D.; Last, R. Progress in the dissection and manipulation of plant vitamin E biosynthesis. Physiol. Plant. 2006, 126, 356–368. [Google Scholar] [CrossRef]
- Ivanov, B.N.; Khorobrykh, S. Participation of Photosynthetic Electron Transport in Production and Scavenging of Reactive Oxygen Species. Antioxid. Redox Signal. 2003, 5, 43–53. [Google Scholar] [CrossRef]
- Matringe, M.; Ksas, B.; Rey, P.; Havaux, M. Tocotrienols, the Unsaturated Forms of Vitamin E, Can Function as Antioxidants and Lipid Protectors in Tobacco Leaves. Plant Physiol. 2008, 147, 764–778. [Google Scholar] [CrossRef]
- Mène-Saffrané, L.; DellaPenna, D. Biosynthesis, regulation and functions of tocochromanols in plants. Plant Physiol. Biochem. 2009, 48, 301–309. [Google Scholar] [CrossRef] [PubMed]
- Kagan, V.E.; Fabisiak, J.P.; Quinn, P.J. Coenzyme Q and vitamin E need each other as antioxidants. Protoplasma 2000, 214, 11–18. [Google Scholar] [CrossRef]
- Fryer, M.J. The antioxidant effects of thylakoid Vitamin E (alpha-tocopherol). Plant Cell Environ. 1992, 15, 381–392. [Google Scholar] [CrossRef]
- Igamberdiev, A.U.; Seregelyes, C.; Manac, N.; Hill, R.D. NADH-dependent metabolism of nitric oxide in alfalfa root cultures expressing barley hemoglobin. Planta 2004, 219, 95–102. [Google Scholar] [CrossRef]
- Fukuzawa, K.; Tokumura, A.; Ouchi, S.; Tsukatani, H. Antioxidant activities of tocopherols on Fe2+-ascorbate-induced lipid peroxidation in lecithin liposomes. Lipids 1982, 17, 511–513. [Google Scholar] [CrossRef] [PubMed]
- Halliwell, B. Reactive Species and Antioxidants. Redox Biology Is a Fundamental Theme of Aerobic Life. Plant Physiol. 2006, 141, 312–322. [Google Scholar] [CrossRef]
- Halliwell, B.; Gutteridge, J.M.C. Free Radicals in Biology and Medicine, 5th ed.; Oxford University Press: Oxford, UK, 2015; pp. 381–382. [Google Scholar]
- Maeda, H.; Song, W.; Sage, T.L.; DellaPenna, D. Tocopherols Play a Crucial Role in Low-Temperature Adaptation and Phloem Loading in Arabidopsis. Plant Cell 2006, 18, 2710–2732. [Google Scholar] [CrossRef]
- Azzi, A. Molecular mechanism of α-tocopherol action. Free Radic. Biol. Med. 2007, 43, 16–21. [Google Scholar] [CrossRef]
- Hyun, T.K.; Kumar, K.; Rao, K.P.; Sinha, A.K.; Roitsch, T. Role of α-tocopherol in cellular signaling: α-tocopherol inhibits stress-induced mitogen-activated protein kinase activation. Plant Biotechnol. Rep. 2010, 5, 19–25. [Google Scholar] [CrossRef]
- Young, A.J. The photoprotective role of carotenoids in higher plants. Physiol. Plant. 1991, 83, 702–708. [Google Scholar] [CrossRef]
- Havaux, M. Carotenoid oxidation products as stress signals in plants. Plant J. 2013, 79, 597–606. [Google Scholar] [CrossRef]
- Lohr, M. Carotenoids. In The Chlamydomonas Sourcebook, 2nd ed.; Harris, E.H., Stern, D.B., Witman, G.B., Eds.; Academic Press: Cambridge, MA, USA, 2009; pp. 799–817. [Google Scholar] [CrossRef]
- Siefermann-Harms, D. The light-harvesting and protective functions of carotenoids in photosynthetic membranes. Physiol. Plant. 1987, 69, 561–568. [Google Scholar] [CrossRef]
- Cogdell, R.J.; Gardiner, A.T. Functions of carotenoids in photosynthesis. Meth. Enzymol. 1993, 214, 185–193. [Google Scholar] [CrossRef]
- Mallik, N.; Rai, L.C. Physiological responses of non-vascular plants to heavy metals, Chapter 5. In Metals Physiology and Bio-chemistry of Metal Toxicity and Tolerance in Plants; Prasad, M.N., Strzatka, K., Eds.; Springer: Dordrecht, The Netherlands, 2002; pp. 111–147. [Google Scholar]
- Johnson, M.; Havaux, M.; Triantaphylides, C.; Ksas, B.; Pascal, A.; Robert, B.; Davison, P.A.; Ruban, A.V.; Horton, P. Elevated Zeaxanthin Bound to Oligomeric LHCII Enhances the Resistance of Arabidopsis to Photooxidative Stress by a Lipid-protective, Antioxidant Mechanism. J. Biol. Chem. 2007, 282, 22605–22618. [Google Scholar] [CrossRef]
- Triantaphylidès, C.; Havaux, M. Singlet oxygen in plants: Production, detoxification and signaling. Trends Plant Sci. 2009, 14, 219–228. [Google Scholar] [CrossRef] [PubMed]
- Jahns, P.; Holzwarth, A.R. The role of the xanthophyll cycle and of lutein in photoprotection of photosystem II. Biochim. Biophys. Acta 2012, 1817, 182–193. [Google Scholar] [CrossRef]
- DellaPenna, D.; Pogson, B.J. Vitamin synthesis in plants: Tocopherols and Carotenoids. Annu. Rev. Plant Biol. 2006, 57, 711–738. [Google Scholar] [CrossRef]
- Li, F.; Vallabhaneni, R.; Yu, J.; Rocheford, T.; Wurtzel, E.T. The Maize Phytoene Synthase Gene Family: Overlapping Roles for Carotenogenesis in Endosperm, Photomorphogenesis, and Thermal Stress Tolerance. Plant Physiol. 2008, 147, 1334–1346. [Google Scholar] [CrossRef]
- Gomathi, R.; Rakkiyapan, P. Comparative lipid peroxidation, leaf membrane thermostability, and antioxidant system in four sugarcane genotypes differing in salt tolerance. Int. J. Plant Physiol. Biochem. 2011, 3, 67–74. [Google Scholar] [CrossRef]
- Grace, S.C.; Logan, B. Energy dissipation and radical scavenging by the plant phenylpropanoid pathway. Philos. Trans. R. Soc. B Biol. Sci. 2000, 355, 1499–1510. [Google Scholar] [CrossRef]
- Michalak, A. Phenolic compounds and their antioxidant activity in plants growing under heavy metal stress. Pol. J. Environ. Stud. 2006, 15, 523–530. [Google Scholar]
- Fini, A.; Brunetti, C.; Di Ferdinando, M.; Ferrini, F.; Tattini, M. Stress-induced flavonoid biosynthesis and the antioxidant machinery of plants. Plant Signal. Behav. 2011, 6, 709–711. [Google Scholar] [CrossRef]
- Agati, G.; Azzarello, E.; Pollastri, S.; Tattini, M. Flavonoids as antioxidants in plants: Location and functional significance. Plant Sci. 2012, 196, 67–76. [Google Scholar] [CrossRef]
- Sakihama, Y.; Cohen, M.F.; Grace, S.C.; Yamasaki, H. Plant phenolic antioxidant and prooxidant activities: Phenolics-induced oxidative damage mediated by metals in plants. Toxicology 2002, 177, 67–80. [Google Scholar] [CrossRef]
- Rice-Evans, C.; Miller, N.; Paganga, G. Antioxidant properties of phenolic compounds. Trends Plant Sci. 1997, 2, 152–159. [Google Scholar] [CrossRef]
- Finnegan, P.M.; Chen, W. Arsenic Toxicity: The Effects on Plant Metabolism. Front. Physiol. 2012, 3, 182. [Google Scholar] [CrossRef] [PubMed]
- Waszczak, C.; Akter, S.; Jacques, S.; Huang, J.; Messens, J.; Van Breusegem, F. Oxidative post-translational modifications of cysteine residues in plant signal transduction. J. Exp. Bot. 2015, 66, 2923–2934. [Google Scholar] [CrossRef]
- Waszczak, C.; Carmody, M.; Kangasjärvi, J. Reactive Oxygen Species in Plant Signaling. Annu. Rev. Plant Biol. 2018, 69, 209–236. [Google Scholar] [CrossRef]
- Wu, F.; Chi, Y.; Jiang, Z.; Xu, Y.; Xie, L.; Huang, F.; Wan, D.; Ni, J.; Yuan, F.; Wu, X.; et al. Hydrogen peroxide sensor HPCA1 is an LRR receptor kinase in Arabidopsis. Nature 2020, 578, 577–581. [Google Scholar] [CrossRef] [PubMed]
- Yalcinkaya, T.; Uzilday, B.; Ozgur, R.; Turkan, I.; Mano, J. Lipid peroxidation-derived reactive carbonyl species (RCS): Their interaction with ROS and cellular redox during environmental stresses. Environ. Exp. Bot. 2019, 165, 139–149. [Google Scholar] [CrossRef]
- Suzuki, Y.J.; Carini, M.; Butterfield, D.A. Protein carbonylation. Antioxid. Redox Signal. 2010, 12, 323–325. [Google Scholar] [CrossRef] [PubMed]
- Ciacka, K.; Tymiński, M.; Gniazdowska, A.; Krasuska, U. Carbonylation of proteins—An element of plant ageing. Planta 2020, 252, 12. [Google Scholar] [CrossRef]
- Cui, F.; Brosche, M.; Shapiguzov, A.; He, X.-Q.; Vainonen, J.P.; Leppälä, J.; Trotta, A.; Kangasjärvi, S.; Salojärvi, J.; Kangasjärvi, J.; et al. Interaction of methyl viologen-induced chloroplast and mitochondrial signalling in Arabidopsis. Free. Radic. Biol. Med. 2019, 134, 555–566. [Google Scholar] [CrossRef]
- Exposito-Rodriguez, M.; Laissue, P.P.; Yvon-Durocher, G.; Smirnoff, N.; Mullineaux, P.M. Photosynthesis-dependent H2O2 transfer from chloroplasts to nuclei provides a high-light signalling mechanism. Nat. Commun. 2017, 8, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Trebak, M.; Ginnan, R.; Singer, H.A.; Jourd’Heuil, D. Interplay Between Calcium and Reactive Oxygen/Nitrogen Species: An Essential Paradigm for Vascular Smooth Muscle Signaling. Antioxid. Redox Signal. 2010, 12, 657–674. [Google Scholar] [CrossRef] [PubMed]
- Hempel, N.; Trebak, M. Crosstalk between calcium and reactive oxygen species signaling in cancer. Cell Calcium 2017, 63, 70–96. [Google Scholar] [CrossRef] [PubMed]
- Niu, L.; Liao, W. Hydrogen Peroxide Signaling in Plant Development and Abiotic Responses: Crosstalk with Nitric Oxide and Calcium. Front. Plant Sci. 2016, 7, 230. [Google Scholar] [CrossRef] [PubMed]
- Farnese, F.D.S.; Menezes-Silva, P.E.; Gusman, G.S.; Oliveira, J. When Bad Guys Become Good Ones: The Key Role of Reactive Oxygen Species and Nitric Oxide in the Plant Responses to Abiotic Stress. Front. Plant Sci. 2016, 7, 471. [Google Scholar] [CrossRef]
- Lindermayr, C.; Durner, J. Interplay of Reactive Oxygen Species and Nitric Oxide: Nitric Oxide Coordinates Reactive Oxygen Species Homeostasis. Plant Physiol. 2015, 167, 1209–1210. [Google Scholar] [CrossRef]
- Del Río, L.A. ROS and RNS in plant physiology: An overview. J. Exp. Bot. 2015, 66, 2827–2837. [Google Scholar] [CrossRef]
- Piterková, J.; Luhová, L.; Navrátilová, B.; Sedlářová, M.; Petřivalský, M. Early and long-term responses of cucumber cells to high cadmium concentration are modulated by nitric oxide and reactive oxygen species. Acta Physiol. Plant. 2015, 37, 19. [Google Scholar] [CrossRef]
- Delledonne, M.; Zeier, J.; Marocco, A.; Lamb, C. Signal interactions between nitric oxide and reactive oxygen intermediates in the plant hypersensitive disease resistance response. Proc. Natl. Acad. Sci. USA 2001, 98, 13454–13459. [Google Scholar] [CrossRef]
- Wang, Y.; Loake, G.J.; Chu, C. Cross-talk of nitric oxide and reactive oxygen species in plant programed cell death. Front. Plant Sci. 2013, 4, 314. [Google Scholar] [CrossRef]
- Kohli, S.K.; Khanna, K.; Bhardwaj, R.; Allah, E.F.A.; Ahmad, P.; Corpas, F.J. Assessment of Subcellular ROS and NO Metabolism in Higher Plants: Multifunctional Signaling Molecules. Antioxidants 2019, 8, 641. [Google Scholar] [CrossRef]
- Kankofer, M. Antioxidative Defence Mechanisms Against Reactive Oxygen Species in Bovine Retained and Not-Retained Placenta: Activity of Glutathione Peroxidase, Glutathione Transferase, Catalase and Superoxide Dismutase. Placenta 2001, 22, 466–472. [Google Scholar] [CrossRef] [PubMed]
- Vardharajula, S.; Ali, S.Z.; Grover, M.; Reddy, G.; Bandi, V. Drought-tolerant plant growth promotingBacillusspp.: Effect on growth, osmolytes, and antioxidant status of maize under drought stress. J. Plant Interact. 2011, 6, 1–14. [Google Scholar] [CrossRef]
- Batool, T.; Ali, S.; Seleiman, M.F.; Naveed, N.H.; Ali, A.; Ahmed, K.; Abid, M.; Rizwan, M.; Shahid, M.R.; Alotaibi, M.; et al. Plant growth promoting rhizobacteria alleviates drought stress in potato in response to suppressive oxidative stress and antioxidant enzymes activities. Sci. Rep. 2020, 10, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Gan, Y.; Xu, B. Application of Plant-Growth-Promoting Fungi Trichoderma longibrachiatum T6 Enhances Tolerance of Wheat to Salt Stress through Improvement of Antioxidative Defense System and Gene Expression. Front. Plant Sci. 2016, 7, 1405. [Google Scholar] [CrossRef]
- Borriss, R.; Wu, H.; Gao, X. Secondary metabolites of the plant growth promoting model rhizobacterium Bacillus velezensis FZB42 are involved in direct suppression of plant pathogens and in stimulation of plant-induced systemic resistance. In Metabolites of Plant Growth Promoting Rhizomicroorganisms: Discovery and Applications, 1st ed.; Singh, H.B., Keswani, C., Reddy, M.S., Sansinenea, E., García-Estrada, C., Eds.; Springer: Singapore, 2019; pp. 147–168. [Google Scholar]
- Shrivastava, P.; Kumar, R. Soil salinity: A serious environmental issue and plant growth promoting bacteria as one of the tools for its alleviation. Saudi J. Biol. Sci. 2014, 22, 123–131. [Google Scholar] [CrossRef] [PubMed]
- Moreno-Galván, A.; Romero-Perdomo, F.A.; Estrada-Bonilla, G.; Meneses, C.H.S.G.; Bonilla, R.R. Dry-Caribbean Bacillus spp. Strains Ameliorate Drought Stress in Maize by a Strain-Specific Antioxidant Response Modulation. Microorganisms 2020, 8, 826. [Google Scholar] [CrossRef] [PubMed]
- Carlson, R.; Tugizimana, F.; Steenkamp, P.A.; Dubery, I.A.; Hassen, A.I.; Labuschagne, N. Rhizobacteria-induced systemic tolerance against drought stress in Sorghum bicolor (L.) Moench. Microbiol. Res. 2019, 232, 126388. [Google Scholar] [CrossRef] [PubMed]
- Lucas, J.A.; García-Cristobal, J.; Bonilla, A.; Ramos, B.; Gutierrez-Mañero, J. Beneficial rhizobacteria from rice rhizosphere confers high protection against biotic and abiotic stress inducing systemic resistance in rice seedlings. Plant Physiol. Biochem. 2014, 82, 44–53. [Google Scholar] [CrossRef] [PubMed]
- Naseem, H.; Bano, A. Role of plant growth-promoting rhizobacteria and their exopolysaccharide in drought tolerance of maize. J. Plant Interact. 2014, 9, 689–701. [Google Scholar] [CrossRef]
- Ngumbi, E.; Kloepper, J. Bacterial-mediated drought tolerance: Current and future prospects. Appl. Soil Ecol. 2016, 105, 109–125. [Google Scholar] [CrossRef]
- Yang, J.; Kloepper, J.W.; Ryu, C.-M. Rhizosphere bacteria help plants tolerate abiotic stress. Trends Plant Sci. 2009, 14, 1–4. [Google Scholar] [CrossRef]
- Harish, S.; Kavino, M.; Kumar, N.; Saravanakumar, D.; Soorianathasundaram, K.; Samiyappan, R. Biohardening with Plant Growth Promoting Rhizosphere and Endophytic bacteria induces systemic resistance against Banana bunchy top virus. Appl. Soil Ecol. 2008, 39, 187–200. [Google Scholar] [CrossRef]
- Walters, D.R.; Fountaine, J.M. Practical application of induced resistance to plant diseases: An appraisal of effectiveness under field conditions. J. Agric. Sci. 2009, 147, 523–535. [Google Scholar] [CrossRef]
- Wang, C.; Guo, Y.; Wang, C.; Liu, H.; Niu, D.; Wang, Y.; Guo, J. Enhancement of tomato (Lycopersicon esculentum) tolerance to drought stress by plant-growth-promoting rhizobacterium (PGPR) Bacillus cereus AR156. J. Agric. Biotechnol. 2012, 20, 1097–1105. (In Chinese) [Google Scholar]
- Gallé, A.; Haldimann, P.; Feller, U. Photosynthetic performance and water relations in young pubescent oak (Quercus pubescens) trees during drought stress and recovery. New Phytol. 2007, 174, 799–810. [Google Scholar] [CrossRef]
- Banik, P.; Zeng, W.; Tai, H.; Bizimungu, B.; Tanino, K. Effects of drought acclimation on drought stress resistance in potato (Solanum tuberosum L.) genotypes. Environ. Exp. Bot. 2016, 126, 76–89. [Google Scholar] [CrossRef]
- Vurukonda, S.S.K.P.; Vardharajula, S.; Shrivastava, M.; SkZ, A. Enhancement of drought stress tolerance in crops by plant growth promoting rhizobacteria. Microbiol. Res. 2016, 184, 13–24. [Google Scholar] [CrossRef]
- Ansary, M.H.; Rahmani, H.A.; Ardakani, M.R.; Paknejad, F.; Habibi, D.; Mafakheri, S. Effect of Pseudomonas fluorescens on proline and phytohormonal status of maize (Zea mays L.) under water deficit stress. Ann. Biol. Res. 2012, 3, 1054–1062. [Google Scholar]
- Armada, E.; Roldan, A.; Azcon, R. Differential Activity of Autochthonous Bacteria in Controlling Drought Stress in Native Lavandula and Salvia Plants Species Under Drought Conditions in Natural Arid Soil. Microb. Ecol. 2013, 67, 410–420. [Google Scholar] [CrossRef]
- Shintu, P.V.; Jayaram, K.M. Phosphate solubilising bacteria (Bacillus polymyxa)—An effective approach to mitigate drought in tomato (Lycopersicon esculentum Mill). Trop. Plant Res. 2015, 2, 17–22. [Google Scholar]
- Paul, M.J.; Primavesi, L.F.; Jhurreea, D.; Zhang, Y. Trehalose Metabolism and Signaling. Annu. Rev. Plant Biol. 2008, 59, 417–441. [Google Scholar] [CrossRef]
- Mastouri, F.; Björkman, T.; Harman, G.E. Seed Treatment with Trichoderma harzianum Alleviates Biotic, Abiotic, and Physiological Stresses in Germinating Seeds and Seedlings. Phytopathology 2010, 100, 1213–1221. [Google Scholar] [CrossRef]
- Yogendra, S.G.; Singh, U.S.; Sharma, A.K. Enhance activity of stress related enzymes in rice (Oryza sativa L.) induced by plant growth promoting fungi under drought stress. Afr. J. Agric. Res. 2014, 9, 1430–1434. [Google Scholar] [CrossRef]
- Zlatev, Z.S.; Lidon, F.C.; Ramalho, J.; Yordanov, I.T. Comparison of resistance to drought of three bean cultivars. Biol. Plant. 2006, 50, 389–394. [Google Scholar] [CrossRef]
- Chang-Quan, W.; Rui-Chang, L. Enhancement of superoxide dismutase activity in the leaves of white clover (Trifolium repens L.) in response to polyethylene glycol-induced water stress. Acta Physiol. Plant. 2008, 30, 841–847. [Google Scholar] [CrossRef]
- Mafakheri, A.; Siosemardeh, A.; Bahramnejad, B.; Struik, P.C.; Sohrabi, Y. Effect of drought stress on yield, proline and chlo-rophyll contents in three chickpea cultivars. Aust. J. Crop Sci. 2010, 4, 580–585. [Google Scholar]
- Kukreja, S.; Nandwal, A.S.; Kumar, N.; Sharma, S.K.; Sharma, S.K.; Unvi, V.; Sharma, P.K. Plant water status, H2O2 scavenging enzymes, ethylene evolution and membrane integrity of Cicer arietinum roots as affected by salinity. Biol. Plant. 2005, 49, 305–308. [Google Scholar] [CrossRef]
- Eyidogan, F.; Oz, M.T. Effect of salinity on antioxidant responses of chickpea seedlings. Acta Physiol. Plant. 2007, 29, 485–493. [Google Scholar] [CrossRef]
- Gapińska, M.; Skłodowska, M.; Gabara, B. Effect of short- and long-term salinity on the activities of antioxidative enzymes and lipid peroxidation in tomato roots. Acta Physiol. Plant. 2007, 30, 11–18. [Google Scholar] [CrossRef]
- Takshak, S.; Agrawal, S.B. Effect of ultraviolet-B radiation on biomass production, lipid peroxidation, reactive oxygen species, and antioxidants in withania somnifera. Biol. Plant. 2014, 58, 328–334. [Google Scholar] [CrossRef]
- Zhang, F.; Zhang, H.; Xia, Y.; Wang, G.; Xu, L.; Shen, Z. Exogenous application of salicylic acid alleviates cadmium toxicity and reduces hydrogen peroxide accumulation in root apoplasts of Phaseolus aureus and Vicia sativa. Plant Cell Rep. 2011, 30, 1475–1483. [Google Scholar] [CrossRef] [PubMed]
- Sandalio, L.; Dalurzo, H.; Gómez, M.; Romero-Puertas, M.; del Río, L. Cadmium-induced changes in the growth and oxidative metabolism of pea plants. J. Exp. Bot. 2001, 52, 2115–2126. [Google Scholar] [CrossRef]
- Li, F.-T.; Qi, J.-M.; Zhang, G.-Y.; Lin, L.-H.; Fang, P.-P.; Tao, A.-F.; Xu, J.-T. Effect of Cadmium Stress on the Growth, Antioxidative Enzymes and Lipid Peroxidation in Two Kenaf (Hibiscus cannabinus L.) Plant Seedlings. J. Integr. Agric. 2013, 12, 610–620. [Google Scholar] [CrossRef]
- Gao, S.; Ouyang, C.; Wang, S.; Xu, Y.; Tang, L.; Chen, F. Effects of salt stress on growth, antioxidant enzyme and phenylalanine ammonia-lyase activities in Jatropha curcas L. seedlings. Plant Soil Environ. 2008, 54, 374–381. [Google Scholar] [CrossRef]
- Badawi, G.H.; Kawano, N.; Yamauchi, Y.; Shimada, E.; Sasaki, R.; Kubo, A.; Tanaka, K. Over-expression of ascorbate peroxidase in tobacco chloroplasts enhances the tolerance to salt stress and water deficit. Physiol. Plant. 2004, 121, 231–238. [Google Scholar] [CrossRef]
- Kim, S.Y.; Lim, J.-H.; Park, M.R.; Kim, Y.J.; Park, T.I.; Seo, Y.W.; Choi, K.G.; Yun, S.J. Enhanced antioxidant enzymes are associated with reduced hydrogen peroxide in barley roots under saline stress. J. Biochem. Mol. Biol. 2005, 38, 218–224. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Ying, Y.; Chen, J.; Wang, X. Transgenic Arabidopsis overexpressing Mn-SOD enhanced salt-tolerance. Plant Sci. 2004, 167, 671–677. [Google Scholar] [CrossRef]
- Rao, M.V.; Paliyath, G.; Ormrod, D.P. Ultraviolet-B- and Ozone-Induced Biochemical Changes in Antioxidant Enzymes of Arabidopsis thaliana. Plant Physiol. 1996, 110, 125–136. [Google Scholar] [CrossRef] [PubMed]
- Malar, S.; Shivendra Vikram, S.; Favas, P.J.; Perumal, V. Lead heavy metal toxicity induced changes on growth and antioxidative enzymes level in water hyacinths [Eichhornia crassipes (Mart.)]. Bot. Stud. 2016, 55, 54. [Google Scholar] [CrossRef] [PubMed]
- Roychoudhury, A.; Ghosh, S. Physiological and biochemical responses of mungbean (Vigna radiata L. Wilczek) to varying concentrations of cadmium chloride or sodium chloride. Unique J. Pharm. Biol. Sci. 2013, 1, 11–21. [Google Scholar]
- Kandziora-Ciupa, M.; Ciepał, R.; Nadgórska-Socha, A.; Barczyk, G. A comparative study of heavy metal accumulation and antioxidant responses in Vaccinium myrtillus L. leaves in polluted and non-polluted areas. Environ. Sci. Pollut. Res. 2013, 20, 4920–4932. [Google Scholar] [CrossRef]
- Aghaei, K.; Ehsanpour, A.A.; Komatsu, S. Potato Responds to Salt Stress by Increased Activity of Antioxidant Enzymes. J. Integr. Plant Biol. 2009, 51, 1095–1103. [Google Scholar] [CrossRef]
- Sharma, P.; Dubey, R.S. Drought Induces Oxidative Stress and Enhances the Activities of Antioxidant Enzymes in Growing Rice Seedlings. Plant Growth Regul. 2005, 46, 209–221. [Google Scholar] [CrossRef]
- Basu, S.; Roychoudhury, A.; Saha, P.P.; Sengupta, D.N. Differential antioxidative responses of indica rice cultivars to drought stress. Plant Growth Regul. 2009, 60, 51–59. [Google Scholar] [CrossRef]
- Roychoudhury, A.; Basu, S.; Sengupta, D.N. Antioxidants and stress-related metabolites in the seedlings of two indica rice varieties exposed to cadmium chloride toxicity. Acta Physiol. Plant. 2011, 34, 835–847. [Google Scholar] [CrossRef]
- Roychoudhury, A.; Basu, S.; Sarkar, S.N.; Sengupta, D.N. Comparative physiological and molecular responses of a common aromatic indica rice cultivar to high salinity with non-aromatic indica rice cultivars. Plant Cell Rep. 2008, 27, 1395–1410. [Google Scholar] [CrossRef]
- Vaidyanathan, H.; Sivakumar, P.; Chakrabarty, R.; Thomas, G. Scavenging of reactive oxygen species in NaCl-stressed rice (Oryza sativa L.)—differential response in salt-tolerant and sensitive varieties. Plant Sci. 2003, 165, 1411–1418. [Google Scholar] [CrossRef]
- Mishra, P.; Bhoomika, K.; Dubey, R.S. Differential responses of antioxidative defense system to prolonged salinity stress in salt-tolerant and salt-sensitive Indica rice (Oryza sativa L.) seedlings. Protoplasma 2011, 250, 3–19. [Google Scholar] [CrossRef]
- Çelik, Ö.; Çakır, B.C.; Atak, Ç. Identification of the antioxidant defense genes which may provide enhanced salt tolerance in Oryza sativa L. Physiol. Mol. Biol. Plants 2018, 25, 85–99. [Google Scholar] [CrossRef]
- Chutipaijit, S.; Cha-Um, S.; Sompornpailin, K. Differential accumulations of proline and flavonoids in indica rice varieties against salinity. Pak. J. Bot. 2009, 41, 2497–2506. [Google Scholar]
- Xu, S.; Li, J.; Zhang, X.; Wei, H.; Cui, L. Effects of heat acclimation pretreatment on changes of membrane lipid peroxidation, antioxidant metabolites, and ultrastructure of chloroplasts in two cool-season turfgrass species under heat stress. Environ. Exp. Bot. 2006, 56, 274–285. [Google Scholar] [CrossRef]
- Chakraborty, U.; Pradhan, D. High temperature-induced oxidative stress in Lens culinaris, role of antioxidants and amelioration of stress by chemical pre-treatments. J. Plant Interact. 2011, 6, 43–52. [Google Scholar] [CrossRef]
- Sairam, R.; Deshmukh, P.; Saxena, D. Role of Antioxidant Systems in Wheat Genotypes Tolerance to Water Stress. Biol. Plant. 1998, 41, 387–394. [Google Scholar] [CrossRef]
- Qayyum, A.; Al Ayoubi, S.; Sher, A.; Bibi, Y.; Ahmad, S.; Shen, Z.; Jenks, M.A. Improvement in drought tolerance in bread wheat is related to an improvement in osmolyte production, antioxidant enzyme activities, and gaseous exchange. Saudi J. Biol. Sci. 2021, 28, 5238–5249. [Google Scholar] [CrossRef]
- Khanna-Chopra, R.; Chauhan, S. Wheat cultivars differing in heat tolerance show a differential response to oxidative stress during monocarpic senescence under high temperature stress. Protoplasma 2015, 252, 1241–1251. [Google Scholar] [CrossRef]
- Almeselmani, M.; Deshmukh, P.S.; Sairam, R.K. High temperature stress tolerance in wheat genotypes: Role of antioxidant defence enzymes. Acta Agron. Hung. 2009, 57, 1–14. [Google Scholar] [CrossRef]
- Kaur, H.; Sirhindi, G.; Bhardwaj, R.; Alyemeni, M.N.; Siddique, K.; Ahmad, P. 28-homobrassinolide regulates antioxidant enzyme activities and gene expression in response to salt- and temperature-induced oxidative stress in Brassica juncea. Sci. Rep. 2018, 8, 8735. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Gupta, D.; Nayyar, H. Comparative response of maize and rice genotypes to heat stress: Status of oxidative stress and antioxidants. Acta Physiol. Plant. 2011, 34, 75–86. [Google Scholar] [CrossRef]
- Shabankareh, H.G.; Khorasaninejad, S.; Soltanloo, H.; Shariati, V. Physiological response and secondary metabolites of three lavender genotypes under water deficit. Sci. Rep. 2021, 11, 1–22. [Google Scholar] [CrossRef]
- Munné-Bosch, S.; Alegre, L. Changes in carotenoids, tocopherols and diterpenes during drought and recovery, and the biological significance of chlorophyll loss in Rosmarinus officinalis plants. Planta 2000, 210, 925–931. [Google Scholar] [CrossRef]
- Kim, T.Y.; Ku, H.; Lee, S.-Y. Crop Enhancement of Cucumber Plants under Heat Stress by Shungite Carbon. Int. J. Mol. Sci. 2020, 21, 4858. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.; Sullivan, J.; Garrett, W.M.; Caperna, T.J.; Natarajan, S. Impact of solar Ultraviolet-B on the proteome in soybean lines differing in flavonoid contents. Phytochemistry 2008, 69, 38–48. [Google Scholar] [CrossRef] [PubMed][Green Version]
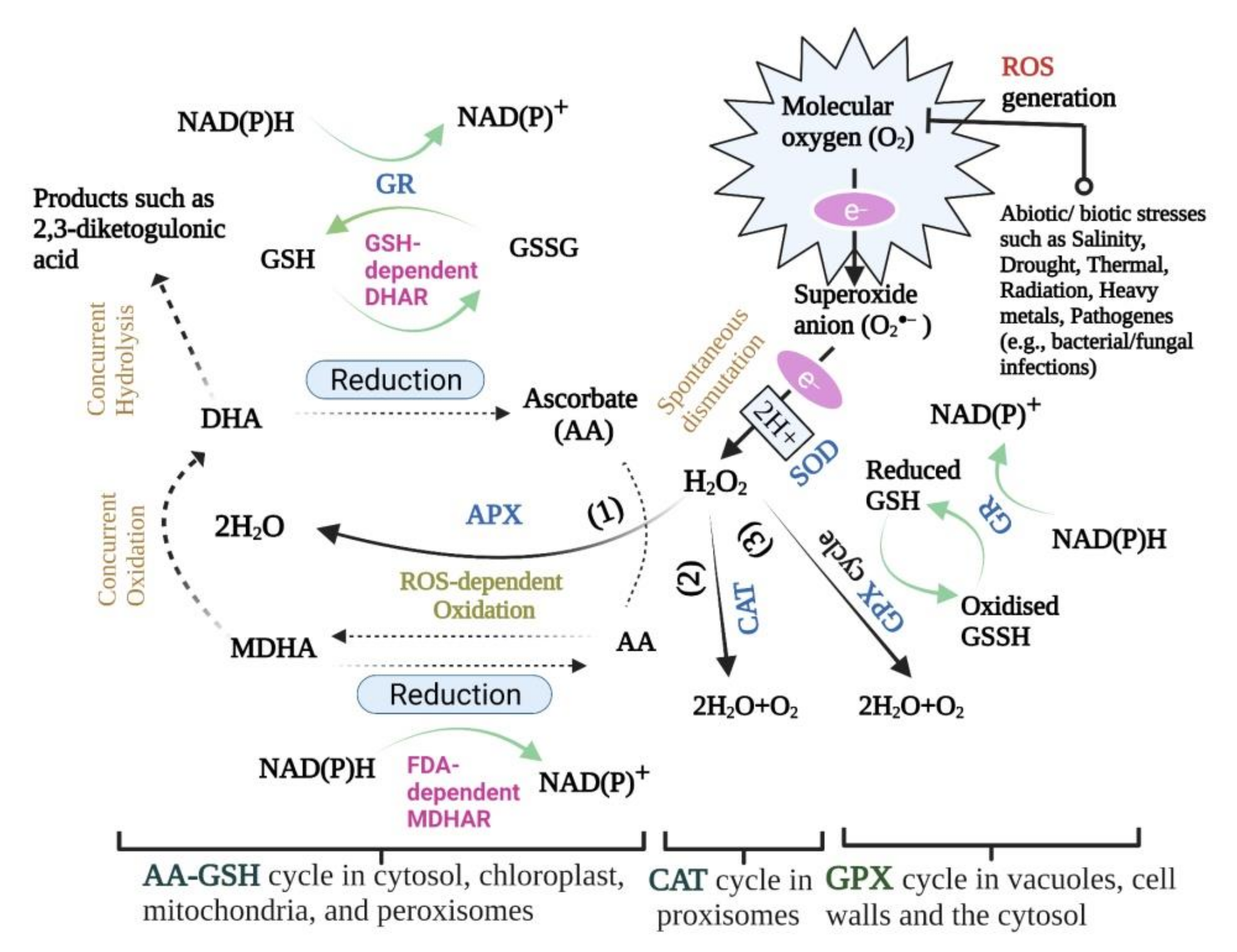
| Plant Species | Common Name | Genomic DNA or Protein | Subcellular Localisation and Site of Detection in Plant | References |
|---|---|---|---|---|
| Brassica napus | Rapeseed | CAT1–CAT14 | Peroxisomes (CAT2–CAT3, CAT6–CAT8, CAT11–CAT14), cytoskeleton (CAT1), cytoplasm (CAT4), mitochondrion (CAT9), and chloroplast (CAT5, CAT10) in root, leaf, stem, and silique samples | [89] |
| Helianthus annus | Sunflower | CAT1–CAT8 | Peroxisomes in cotyledons and roots | [90,91,92] |
| Gossypium hirsutum | Cotton | CAT1–CAT7 | Peroxisomes in leaves | [78,93] |
| Pisum sativum | Pea | CAT1–CAT5 | Peroxisomes in leaves and the whole fruit | [94,95] |
| Cucumis sativus | Cucumber | CAT1–CAT4 | Peroxisomes in roots, stem, leaves, flowers, and fruits | [79] |
| Oryza sativa L. | Rice | OsCATA–OsCATD | Cytosol (OsCATA–OsCATC), peroxisomes (OsCATB, OsCATC), and plasma membrane (OsCATD) in immature seeds and rice seedlings | [80,96,97] |
| Nicotiana plumbaginofolia | Tex-Mex tobacco | CAT1–CAT3 | Peroxisomes in leaves | [84] |
| Arabidopsis thaliana | Arabidopsis | CAT1–CAT3 | Peroxisomes in bolts, leaves, siliques (CAT1, CAT2), in pollen and seeds (CAT1), stems and roots (CAT3) | [81,98,99,100,101] |
| Cucurpita pepo | Pumpkin | CAT1–CAT3 | Glyoxysomes in seeds and early seedlings (CAT1), mature leaves, stem (CAT2), green cotyledons and green hypocotyls (CAT2, CAT3), and roots (CAT3) | [82] |
| Glycine max | Soybean | CAT1–CAT3 | Glyoxysomes in developing kernels (CAT1), green leaves (CAT2, CAT3), roots (CAT3), and epicotyl of the developing seedlings (CAT3) | [102] |
| Zea mays | Maize | CAT1–CAT3 | In scutella, milky endosperm of immature kernels, leaves and epicotyls (CAT1); Post-germinative scutella, with lower levels in leaves, epicotyls and growing kernels (CAT2); Epicotyls and, to a lesser extent, in leaves and scutella (CAT3) | [83,103,104,105,106,107] |
| Capsicum annuum | Hot pepper | CAT1–CAT3 | Peroxisomes in leaf and stem | [108] |
| Nicotiana tabacum | Tobacco | CAT1–CAT3 | Peroxisomes in leaves | [109,110] |
| Ricinus cammunis | Castor bean | CAT1, CAT2 | Glyoxysomes, peroxisomes in hypocotyls and roots (CAT2), in endosperms and cotyledons (CAT1) | [88,111] |
| Hurdeom vulgare | Barley | CAT1, CAT2 | Peroxisomes in whole endosperms, in isolated aleurones and in developing seeds (CAT1), in etiolated seedling shoots and leaf blades (CAT2) | [85] |
| Triticum aestivum | Wheat | CAT1, CAT2 | Peroxisomes in leaves | [112] |
| Solanum tuberosum | Potato | CAT1, CAT2 | Peroxisomes in leaves | [113,114] |
| Lycopersicum sculentum | Tomato | CAT1, CAT2 | Peroxisomes in leaves | [86,115,116] |
| Nicotiana sylvestris | Woodland tobacco | CAT1, CAT4 | Peroxisomes in leaves | [117] |
| Vigna radiata | Mung bean | CAT1 | Glyoxysomes, leaf peroxisomes, and nonspecialised microbodies (etiolated or nongreen tissues) | [118] |
| Secale cereale | Rye | CAT1 | Peroxisomes in leaves | [119] |
| Ipomoea batatas | Sweet potato | CAT1 | Peroxisomes in leaves | [87,120] |
| Plant Species | Stress Type | Antioxidant Reported + (Increase)/−(Decrease) | Reference |
|---|---|---|---|
| Phaseolus vulgaris L. | Drought | +SOD, +APX in leaves | [316] |
| Trifolium repens L. | Drought | +SOD in leaves | [317] |
| Cicer arietinum L. cv. ILC482 | Drought | +Proline in leaves | [318] |
| Cicer arietinum L. cv. CSG-8962 | Salinity | +SOD, +CAT, + Peroxidase (POX), +APX, +GR, and –AA in roots | [319] |
| Cicer arietinum L. cv. Gökçe | Salinity | +SOD in roots and leaves, +APX and +GR in leaves, and +CAT in roots, –CAT in leaves | [320] |
| Lycopersicon esculentum Mill. cv. Perkoz | Salinity | +SOD, +Glutathione S-transferase (GST), and +Glutathione peroxidase (GSH-PX) in roots | [321] |
| Withania somnifera L. | UV-B radiation | +CAT, +GR, +POX, +Polyphenol oxidase (PPO), and +SOD in roots > leaves | [322] |
| Vicia sativa L. | Cd stress | +SOD, +APX, and +CAT in roots | [323] |
| Pisum sativum L. | Cd stress | –SOD, –CAT, –GPX, and +Lipid peroxidation (LPO) in leaves | [324] |
| Hibiscus cannabinus L. cvs. Fuhong 991 and ZM 412 | Cd stress | +GR in leaves (Fuhong 991), +SOD, +CAT, and +POX in roots (Fuhong 991 and ZM 412), +Lipid peroxidation (LPO) in roots (ZM 412) | [325] |
| Jatropha curcas L. | Salinity | +SOD, +CAT, +POX in the cotyledons, hypocotyls and radicles | [326] |
| Nicotiana tabacum L. line Chl-APX5 | Salinity and drought | +APX in chloroplast | [327] |
| Nicotiana tabacum L. | Salinity | +SOD, +CAT, +APX, +POX, and +GR in roots | [328] |
| Nicotiana tabacum L. | Salt, O3 and polyethylene glycol (PEG) stresses | +MDHAR in leaves | [186] |
| Arabidopsis thaliana L. | Salinity | +DHAR in leaves | [201] |
| Arabidopsis thaliana L. | Salinity | +SOD, +CAT, and +POX in leaves | [329] |
| Arabidopsis thaliana L. | UV-B/Ozone (O3) radiation | +SOD, +APX, +GPX, +POX (UV-B and O3), and +GR (O3) in leaves | [330] |
| Eichhornia crassipes (water hyacinth) | Pb stress | +APX, +POX, +SOD, +CAT in leaf and root tissues | [331] |
| Vigna radiata L. Wilczek | CdCl2 stress | +MDHA +POX, +CAT, and +Proline in roots and leaves | [332] |
| Vaccinium myrtillus L. | Heavy metal stress (Cd, Pb, and Zn) | +GSH, +Non-protein thiols, +Proline, and +GPX in leaves | [333] |
| Solanum tuberosum L. | Salt stress | +APX, +CAT and +GR in roots and shoots | [334] |
| Oryza sativa L. | Drought | +SOD, +APX, +MDHAR, +DHAR, +GR, and –CAT in roots and leaves | [335] |
| Oryza sativa L. cvs. IR-29 and Pusa Basmati | Drought | +Flavonoids, +Phenolics, –CAT, –SOD, and +GPX in leaves | [336] |
| Oryza sativa L. vars. IR-29 and Nonabokra | CdCl2/NaCl stress | + Anthocyanin, + Carotenoids, +GPX, +APX, +Proline, +Polyamines (spermidine and spermine) (IR-29/Nonabokra), –Cysteine and –CAT (IR-29), +Cysteine and +CAT (Nonabokra) in leaves | [337,338] |
| Oryza sativa L. cv. Pokkali | Salt stress | +CAT, +AA, and +GSH in leaves | [339] |
| Oryza sativa L. cvs. CSR-27 and Osmancık-97 | Salt stress | +SOD, +APX, +GPX, +CAT, +MDHAR, +DHAR, +GR, and +Proline in roots and leaves | [340,341] |
| Oryza sativa L. vars. Jasmine (KDML105) and Sangyod (SY) | Salt stress | +Flavonoids, and +Proline in shoots | [342] |
| Turfgrass species | Heat stress | +GSH, and +AA in leaves | [343] |
| Lens culinaris Medik. | Heat stress | +CAT, +APX, +SOD, and +LPO in leaves | [344] |
| Triticum aestivum L. cv. C306 | Drought | +SOD, +APX, +CAT, and +AA in leaves | [345] |
| Triticum aestivum L. cv. Chakwal-50 | Drought | +Proline,+CAT, +SOD, +POX, and +MDHAR in leaves | [346] |
| Triticum aestivum L. cv. Hindi62 | Heat stress | +GSH, +SOD, +CAT, +APX, +GR, and +MDHAR in leaves | [347] |
| Triticum aestivum L.cv. C 306. | Heat stress | +SOD, +APX, +CAT, +GR and +POX in leaves | [348] |
| Brassica juncea L. | High temperature and salt stress | +SOD, +CAT, +APX, +GR, +DHAR, and +MDHAR in seedlings | [349] |
| Zea mays L. | Heat stress | +CAT, +APX, +GR, +AA, +GSH, and +Proline in leaves | [350] |
| Lavandula angustifolia cv. Hidcote, Lavandula angustifolia cv. Munstead, and Lavandula stricta | Drought | +Proline, +CAT, +APX, +POX, +SOD, +MDHA, +Flavonoids, and +Phenols in leaves | [351] |
| Rosmarinus officinalis L. | Drought | +Carotenoids, and +α-tocopherol in leaves | [352] |
| Cucumis sativus L. | Heat stress | +SOD, +CAT, and +POX in leaves | [353] |
| Glycine max L. cv. Clark | UV-B radiation | –APX, –CAT, +SOD, +DHA, and +Flavonoid in leaves | [354] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zandi, P.; Schnug, E. Reactive Oxygen Species, Antioxidant Responses and Implications from a Microbial Modulation Perspective. Biology 2022, 11, 155. https://doi.org/10.3390/biology11020155
Zandi P, Schnug E. Reactive Oxygen Species, Antioxidant Responses and Implications from a Microbial Modulation Perspective. Biology. 2022; 11(2):155. https://doi.org/10.3390/biology11020155
Chicago/Turabian StyleZandi, Peiman, and Ewald Schnug. 2022. "Reactive Oxygen Species, Antioxidant Responses and Implications from a Microbial Modulation Perspective" Biology 11, no. 2: 155. https://doi.org/10.3390/biology11020155
APA StyleZandi, P., & Schnug, E. (2022). Reactive Oxygen Species, Antioxidant Responses and Implications from a Microbial Modulation Perspective. Biology, 11(2), 155. https://doi.org/10.3390/biology11020155






