About the Analysis of 18S rDNA Sequence Data from Trypanosomes in Barcoding and Phylogenetics: Tracing a Continuation Error Occurring in the Literature
Abstract
Simple Summary
Abstract
1. Introduction
2. Results and Discussion

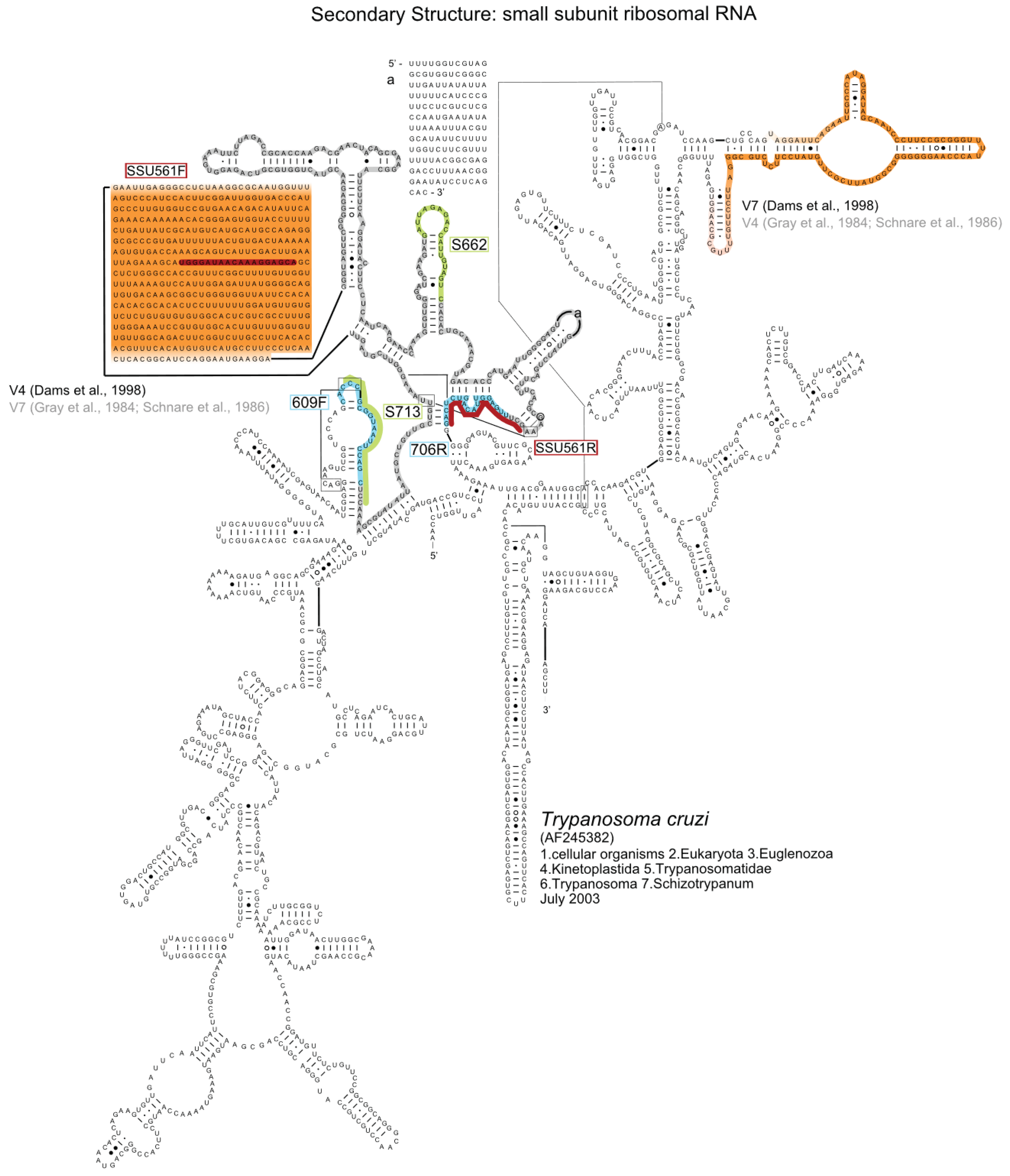
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Rothschild, L.J.; Ragan, M.A.; Coleman, A.W.; Heywood, P.; Gerbi, S.A. Are rRNA sequence comparisons the Rosetta stone of phylogenetics? Cell 1986, 47, 640. [Google Scholar] [CrossRef]
- Olsen, G.J.; Woese, C.R. Ribosomal RNA: A key to phylogeny. FASEB J. 1993, 7, 113–123. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.; Park, J.S. Comparative analyses of the V4 and V9 regions of 18S rDNA for the extant eukaryotic community using the Illumina platform. Sci. Rep. 2020, 10, 6519. [Google Scholar] [CrossRef] [PubMed]
- Stiegler, P.; Carbon, P.; Ebel, J.-P.; Ehresmann, C. A General Secondary-Structure Model for Procaryotic and Eucaryotic RNAs of the Small Ribosomal Subunits. JBIC J. Biol. Inorg. Chem. 1981, 120, 487–495. [Google Scholar] [CrossRef]
- Spencer, D.F.; Schnare, M.N.; Gray, M.W. Pronounced structural similarities between the small subunit ribosomal RNA genes of wheat mitochondria and Escherichia coli. Proc. Natl. Acad. Sci. USA 1984, 81, 493–497. [Google Scholar] [CrossRef]
- Gray, M.; Sankoff, D.; Cedergren, R.J. On the evolutionary descent of organisms and organelles: A global phylogeny based on a highly conserved structural core in small subunit ribosomal RNA. Nucleic Acids Res. 1984, 12, 5837–5852. [Google Scholar] [CrossRef]
- Schnare, M.N.; Collings, J.C.; Gray, M.W. Structure and evolution of the small subunit ribosomal RNA gene of Crithidia fasciculata. Curr. Genet. 1986, 10, 405–410. [Google Scholar] [CrossRef]
- Hernández, R.; Rios, P.; Valdes, A.; Piñero, D. Primary structure of Trypanosoma cruzi small-subunit ribosomal RNA coding region: Comparison with other trypanosomatids. Mol. Biochem. Parasitol. 1990, 41, 207–212. [Google Scholar] [CrossRef]
- Van de Peer, Y.; De Rijk, P.; Wuyts, J.; Winkelmans, T.; De Wachter, R. The European Small Subunit Ribosomal RNA database. Nucleic Acids Res. 2002, 28, 175–176. [Google Scholar] [CrossRef]
- Huysmans, E.; De Wachter, R. Compilation of small ribosomal sobunit RNA sequences. Nucleic Acids Res. 1986, 14, r73–r118. [Google Scholar] [CrossRef][Green Version]
- Dams, E.; Hendriks, L.; Van De Peer, Y.; Neefs, J.-M.; Smits, G.; Vandenbempt, I.; De Wachter, R. Compilation of small ribosomal subunit RNA sequences. Nucleic Acids Res. 1988, 16, r87–r173. [Google Scholar] [CrossRef] [PubMed]
- Neefs, J.-M.; De Wachter, R. A proposal for the secondary structure of a variable area of eukaryotic small ribosomal subunit RNA involving the existence of a pseudoknot. Nucleic Acids Res. 1990, 18, 5695–5704. [Google Scholar] [CrossRef] [PubMed]
- Neefs, J.-M.; Van de Peer, Y.; Hendriks, L.; De Wachter, R. Compilation of small ribosomal subunit RNA sequences. Nucleic Acids Res. 1990, 18, 2237–2317. [Google Scholar] [CrossRef] [PubMed]
- Neefs, J.-M.; Van de Peer, Y.; De Rijk, P.; Chapelle, S.; De Wachter, R. Compilation of small ribosomal subunit RNA structures. Nucleic Acids Res. 1993, 21, 3025–3049. [Google Scholar] [CrossRef] [PubMed]
- Neefs, J.-M.; Van de Peer, Y.; De Rijk, P.; Goris, A.; De Wachter, R. Compilation of small ribosomal subunit RNA sequences. Nucleic Acids Res. 1991, 19, 1987–2015. [Google Scholar] [CrossRef]
- De Rijk, P.; Neefs, J.-M.; Van De Peer, Y.; De Wachter, R. Compilation of small ribosomal subunit RNA sequences. Nucleic Acids Res. 1992, 20, 2075–2089. [Google Scholar] [CrossRef]
- Xie, Q.; Lin, J.; Qin, Y.; Zhou, J.; Bu, W. Structural diversity of eukaryotic 18S rRNA and its impact on alignment and phylogenetic reconstruction. Protein Cell 2011, 2, 161–170. [Google Scholar] [CrossRef]
- Ki, J.-S. Hypervariable regions (V1–V9) of the dinoflagellate 18S rRNA using a large dataset for marker considerations. J. Appl. Phycol. 2011, 24, 1035–1043. [Google Scholar] [CrossRef]
- Bininda-Emonds, O.R.P. 18S rRNA variability maps reveal three highly divergent, conserved motifs within Rotifera. BMC Ecol. Evol. 2021, 21, 118. [Google Scholar] [CrossRef]
- Bradley, I.M.; Pinto, A.J.; Guest, J.S. Design and Evaluation of Illumina MiSeq-Compatible, 18S rRNA Gene-Specific Primers for Improved Characterization of Mixed Phototrophic Communities. Appl. Environ. Microbiol. 2016, 82, 5878–5891. [Google Scholar] [CrossRef]
- Stoeck, T.; Bass, D.; Nebel, M.; Christen, R.; Jones, M.D.M.; Breiner, H.-W.; Richards, T.A. Multiple marker parallel tag environmental DNA sequencing reveals a highly complex eukaryotic community in marine anoxic water. Mol. Ecol. 2010, 19, 21–31. [Google Scholar] [CrossRef] [PubMed]
- Harder, C.B.; Rønn, R.; Brejnrod, A.; Bass, D.; Abu Al-Soud, W.; Ekelund, F. Local diversity of heathland Cercozoa explored by in-depth sequencing. ISME J. 2016, 10, 2488–2497. [Google Scholar] [CrossRef] [PubMed]
- Taib, N.; Mangot, J.-F.; Domaizon, I.; Bronner, G.; Debroas, D. Phylogenetic Affiliation of SSU rRNA Genes Generated by Massively Parallel Sequencing: New Insights into the Freshwater Protist Diversity. PLoS ONE 2013, 8, e58950. [Google Scholar] [CrossRef]
- Hadziavdic, K.; Lekang, K.; Lanzén, A.; Jonassen, I.; Thompson, E.M.; Troedsson, C. Characterization of the 18S rRNA Gene for Designing Universal Eukaryote Specific Primers. PLoS ONE 2014, 9, e87624. [Google Scholar] [CrossRef]
- Hu, S.K.; Liu, Z.; Lie, A.A.Y.; Countway, P.D.; Kim, D.Y.; Jones, A.C.; Gast, R.J.; Cary, S.C.; Sherr, E.B.; Sherr, B.F.; et al. Estimating Protistan Diversity Using High-Throughput Sequencing. J. Eukaryot. Microbiol. 2015, 62, 688–693. [Google Scholar] [CrossRef]
- Latz, M.A.C.; Grujcic, V.; Brugel, S.; Lycken, J.; John, U.; Karlson, B.; Andersson, A.; Andersson, A.F. Short- and long-read metabarcoding of the eukaryotic rRNA operon: Evaluation of primers and comparison to shotgun metagenomics sequencing. Mol. Ecol. Resour. 2022, 22, 2304–2318. [Google Scholar] [CrossRef] [PubMed]
- DA Silva, F.M.; Noyes, H.; Campaner, M.; Junqueira, A.C.V.; Coura, J.R.; Añez, N.; Shaw, J.J.; Stevens, J.R.; Teixeira, M.M.G. Phylogeny, taxonomy and grouping of Trypanosoma rangeli isolates from man, triatomines and sylvatic mammals from widespread geographical origin based on SSU and ITS ribosomal sequences. Parasitology 2004, 129, 549–561. [Google Scholar] [CrossRef] [PubMed]
- Cannone, J.J.; Subramanian, S.; Schnare, M.N.; Collett, J.R.; D’Souza, L.M.; Du, Y.; Feng, B.; Lin, N.; Madabusi, L.V.; Müller, K.M.; et al. The comparative RNA web (CRW) site: An online database of comparative sequence and structure information for ribosomal, intron, and other RNAs. BMC Bioinform. 2002, 3, 2. [Google Scholar] [CrossRef]
- Votýpka, J.; Svobodová, M. Trypanosoma avium: Experimental transmission from black flies to canaries. Parasitol Res. 2004, 92, 147–151. [Google Scholar] [CrossRef]
- Votýpka, J.; Oborník, M.; Volf, P.; Svobodová, M.; Lukeš, J. Trypanosoma avium of raptors (Falconiformes): Phylogeny and identification of vectors. Parasitology 2002, 125, 253–263. [Google Scholar] [CrossRef]
- Nickrent, D.L.; Sargent, M.L. An overview of the secondary structure of the V4 region of eukaryotic small-subunit ribosomal RNA. Nucleic Acids Res. 1991, 19, 227–235. [Google Scholar] [CrossRef] [PubMed]
- Hutchinson, R.; Stevens, J. Barcoding in trypanosomes. Parasitology 2017, 145, 563–573. [Google Scholar] [CrossRef] [PubMed]
- Noyes, H.A.; Camps, A.P.; Chance, M.L. Leishmania herreri (Kinetoplastida; Trypanosomatidae) is more closely related to Endotrypanum (Kinetoplastida; Trypanosomatidae) than to Leishmania. Mol. Biochem. Parasitol. 1996, 80, 119–123. [Google Scholar] [CrossRef]
| Authors | Year of Publication | Digital Object Identifier |
|---|---|---|
| Maia da Silva et al. | 2004 | 10.1017/S0031182004005931 |
| Rodrigues et al. | 2006 | 10.1017/S0031182005008929 |
| Cortez et al. | 2006 | 10.1017/S0031182006000254 |
| Ferreira et al. | 2007 | 10.1017/S0031182007003058 |
| Maia da Silva et al. | 2007 | 10.1111/j.1365-294X.2007.03371.x |
| Martins et al. | 2008 | 10.4269/ajtmh.2008.79.427 |
| Viola et al. | 2008 | 10.1017/S0031182008004253 |
| Rodrigues et al. | 2008 | 10.1017/S0031182008004848 |
| Marcili et al. | 2009 | 10.1017/S0031182009005861 |
| Viola et al. | 2009 | 10.1017/S003118200800512X |
| Marcili et al. | 2009 | 10.1016/j.meegid.2009.07.003 |
| Marcili et al. | 2009 | 10.1016/j.ijpara.2008.09.015 |
| Averis et al. | 2009 | 10.1017/S0031182009990801 |
| Maia da Silva et al. | 2009 | 10.1016/j.actatropica.2008.11.005 |
| Maia da Silva et al. | 2010 | 10.1016/j.meegid.2010.02.005 |
| Cavazzana et al. | 2010 | 10.1016/j.ijpara.2009.08.015 |
| Teixeira et al. | 2011 | 10.1016/j.protis.2011.01.001 |
| Garcia et al. | 2011 | 10.1016/j.ijpara.2011.09.001 |
| Lima et al. | 2012 | 10.1016/j.protis.2011.12.003 |
| Martinković et al. | 2012 | 10.1111/j.1550-7408.2011.00599.x |
| Hamilton et al. | 2012 | 10.1016/j.ympev.2012.01.007 |
| Ramirez et al. | 2012 | 10.1016/j.exppara.2012.09.017 |
| Borghesan et al. | 2013 | 10.1016/j.protis.2012.06.001 |
| Marcili et al. | 2013 | 10.5402/2013/328794 |
| Lima et al. | 2013 | 10.1186/1756-3305-6-221 |
| Fermino et al. | 2013 | 10.1186/1756-3305-6-313 |
| Silva-Iturriza et al. | 2013 | 10.1016/j.parint.2012.10.003 |
| Marcili et al. | 2013 | 10.1645/12-156.1 |
| Guhl et al. | 2013 | 10.1016/j.meegid.2013.08.028 |
| Acosta et al. | 2014 | 10.1603/ME13177 |
| Marcili et al. | 2014 | 10.1016/j.meegid.2014.04.001 |
| Da Costa et al. | 2014 | 10.4172/ijbbd.1000120 |
| Lemos et al. | 2015 | 10.1186/s13071-015-1193-7 |
| Fermino et al. | 2015 | 10.1016/j.ijppaw.2015.10.005 |
| Juliana et al. | 2015 | 10.1007/s11230-015-9558-z |
| Lima et al. | 2015 | 10.1186/s13071-015-1255-x |
| Da Costa et al. | 2015 | 10.1089/vbz.2015.1771 |
| Da Costa et al. | 2015 | 10.1089/vbz.2015.1866 |
| Lima et al. | 2015 | 10.1016/j.actatropica.2015.07.015 |
| Martins et al. | 2015 | 10.1515/ap-2015-0009 |
| Dario et al. | 2016 | 10.1186/s13071-016-1754-4 |
| Attias et al. | 2016 | 10.1111/jeu.12310 |
| Zanetti et al. | 2016 | 10.1016/j.ejop.2016.09.004 |
| Szpeiter et al. | 2017 | 10.1590/s1984-29612017022 |
| Galvis-Ovallos | 2017 | 10.1186/s13071-017-2211-8 |
| Da Costa et al. | 2018 | 10.1590/0037-8682-0098-2018 |
| Ribeiro et al. | 2018 | 10.4269/ajtmh.16-0200 |
| Pacheco et al. | 2018 | 10.1590/s1984-296120180049 |
| Dos Santos et al. | 2018 | 10.1017/S0031182017001834 |
| Espinosa et al. | 2018 | 10.1017/S0031182016002092 |
| Borghesan et al. | 2018 | 10.3389/fmicb.2018.00131 |
| Espinosa-Álvarez et al. | 2018 | 10.1016/j.ijpara.2017.12.008 |
| Suganuma et al. | 2019 | 10.1007/s00436-019-06313-x |
| Borges et al. | 2019 | 10.1111/jeu.12678 |
| Barros et al. | 2019 | 10.1016/j.ijppaw.2018.12.009 |
| Fermino et al. | 2019 | 10.1186/s13071-019-3463-2 |
| Pérez et al. | 2019 | 10.1186/s13071-019-3726-y |
| Garcia et al. | 2019 | 10.1007/s10393-019-01440-4 |
| Latif et al. | 2019 | 10.4102/ojvr.v86i1.1634 |
| Kuhls et al. | 2019 | 10.1007/978-1-4939-9210-2_2 |
| Barros et al. | 2020 | 10.3390/pathogens9090736 |
| Garcia et al. | 2020 | 10.1186/s13071-020-04169-0 |
| Rodrigues et al. | 2020 | 10.1016/j.meegid.2019.104143 |
| Boucinha et al. | 2020 | 10.1590/0074-02760200504 |
| e Azevedo et al. | 2020 | 10.1590/0103-8478cr20200262 |
| Marcili et al. | 2020 | 10.1089/vbz.2020.2638 |
| Jaimes-Dueñez et al. | 2020 | 10.1016/j.prevetmed.2020.105159 |
| Dario et al. | 2021 | 10.3390/pathogens10060736 |
| Rosyadi et al. | 2021 | 10.1017/S0031182021001360 |
| Mule et al. | 2021 | 10.1038/s42003-021-01762-6 |
| Dario et al. | 2021 | 10.1016/j.ijppaw.2021.04.003 |
| Ardila et al. | 2022 | 10.1007/s12639-021-01459-x |
| Yasein et al. | 2022 | 10.29261/pakvetj/2022.034 |
| Chiariello et al. | 2022 | 10.1016/j.ijppaw.2021.11.006 |
| Kostygov et al. | 2022 | 10.1186/s13071-022-05212-y |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rackevei, A.S.; Borges, A.; Engstler, M.; Dandekar, T.; Wolf, M. About the Analysis of 18S rDNA Sequence Data from Trypanosomes in Barcoding and Phylogenetics: Tracing a Continuation Error Occurring in the Literature. Biology 2022, 11, 1612. https://doi.org/10.3390/biology11111612
Rackevei AS, Borges A, Engstler M, Dandekar T, Wolf M. About the Analysis of 18S rDNA Sequence Data from Trypanosomes in Barcoding and Phylogenetics: Tracing a Continuation Error Occurring in the Literature. Biology. 2022; 11(11):1612. https://doi.org/10.3390/biology11111612
Chicago/Turabian StyleRackevei, Antonia S., Alyssa Borges, Markus Engstler, Thomas Dandekar, and Matthias Wolf. 2022. "About the Analysis of 18S rDNA Sequence Data from Trypanosomes in Barcoding and Phylogenetics: Tracing a Continuation Error Occurring in the Literature" Biology 11, no. 11: 1612. https://doi.org/10.3390/biology11111612
APA StyleRackevei, A. S., Borges, A., Engstler, M., Dandekar, T., & Wolf, M. (2022). About the Analysis of 18S rDNA Sequence Data from Trypanosomes in Barcoding and Phylogenetics: Tracing a Continuation Error Occurring in the Literature. Biology, 11(11), 1612. https://doi.org/10.3390/biology11111612








