Actinomycetes, an Inexhaustible Source of Naturally Occurring Antibiotics
Abstract
:1. Introduction
2. Historical Discovery of Novel Compounds from Actinomycetes by the Kitasato Ōmura-Drug Discovery Group
- Isolation of microorganisms, identification, and microbial cultivation.
- Discovery of substances from microbial metabolites.
- Optimization of compounds by organic synthesis.
3. Novel Compounds Discovered by Physicochemical (PC) Screening of Cultured Broths of Actinomycetes
4. Novel Compounds Discovered from the Kitasato Microbial Library (KML)
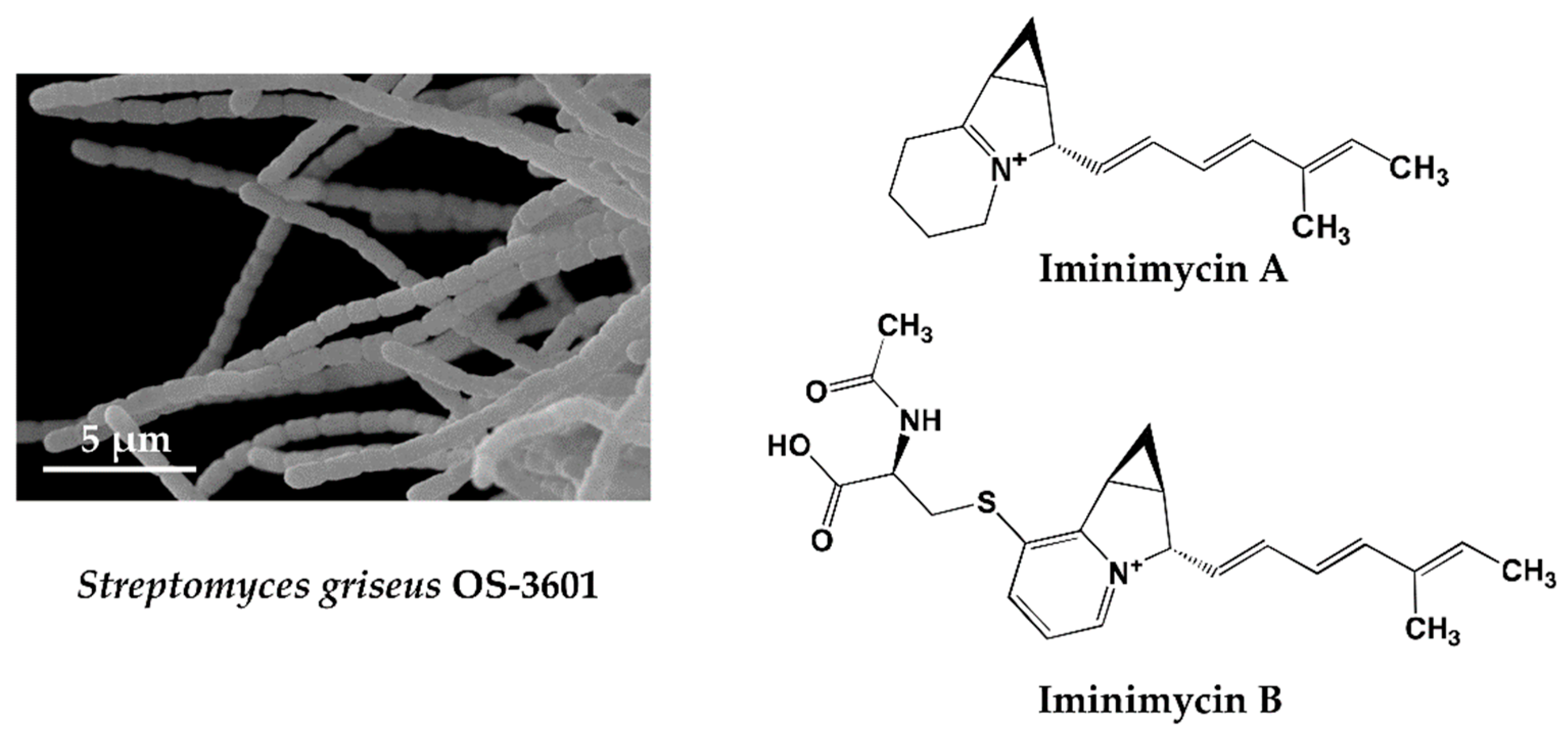
4.1. Iminimycin A by Streptomyces griseus OS-3601, a Streptomycin Producing Strain
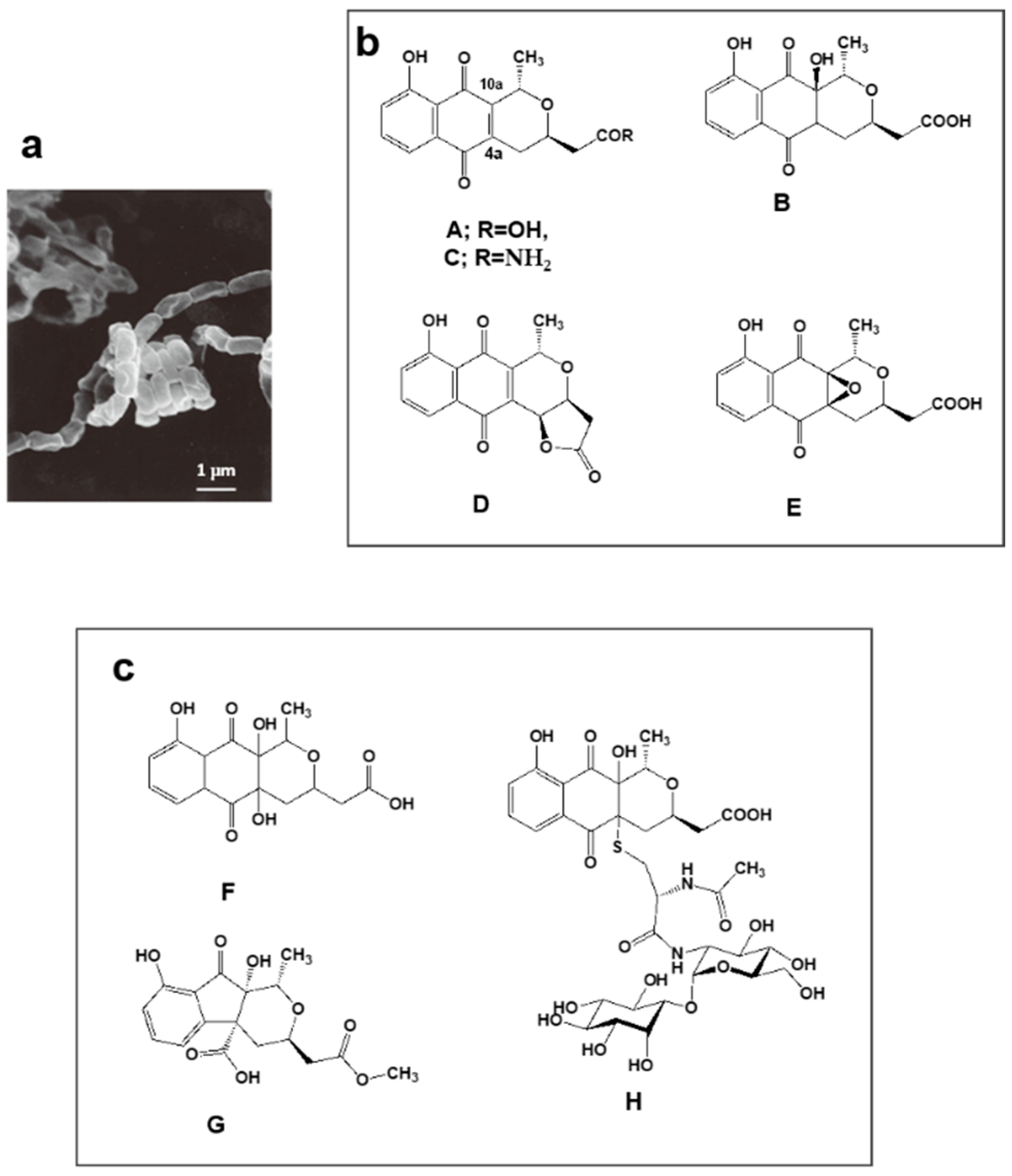
4.2. Nanaomycins F, G and H Derived from Nanaomycins A–E Produced by “Streptomyces rosa subsp. notoensis” OS-3966
5. Novel Compounds Found from Fresh Isolates
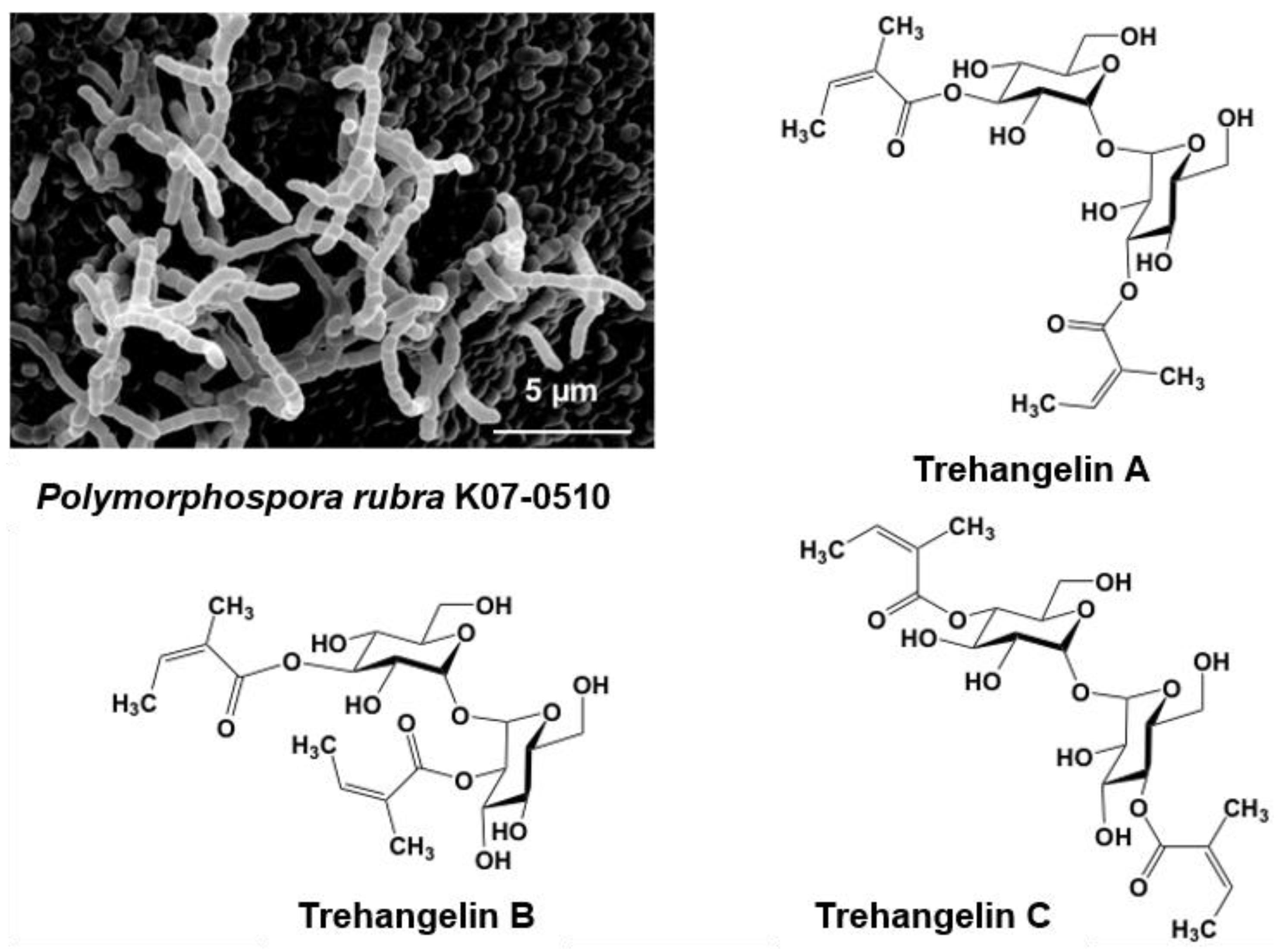
5.1. Novel Substances, Trehangelins Found from Metabolites of the Plant-Derived Rare Actinomycete Polymorphospora Rubra K07-0510
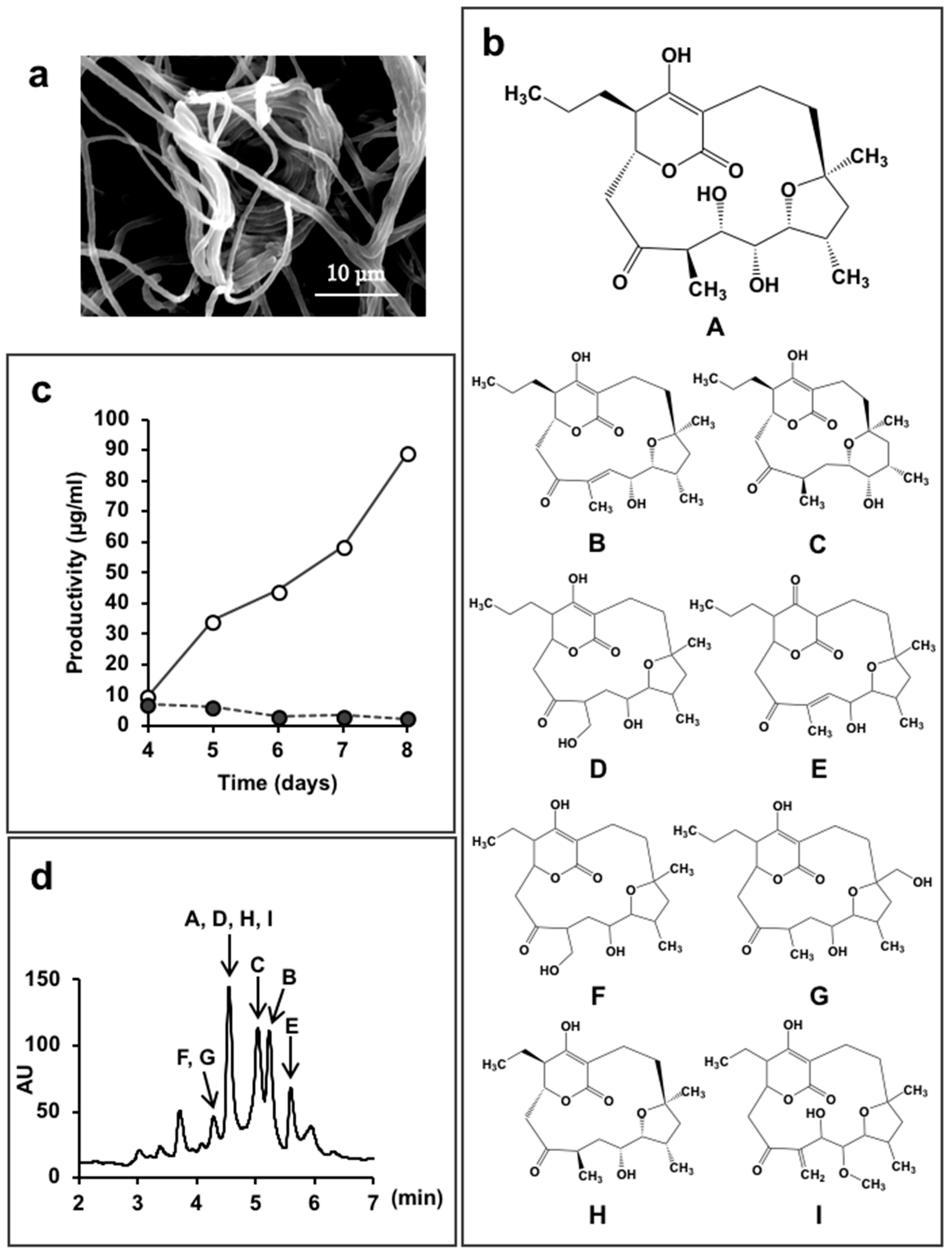
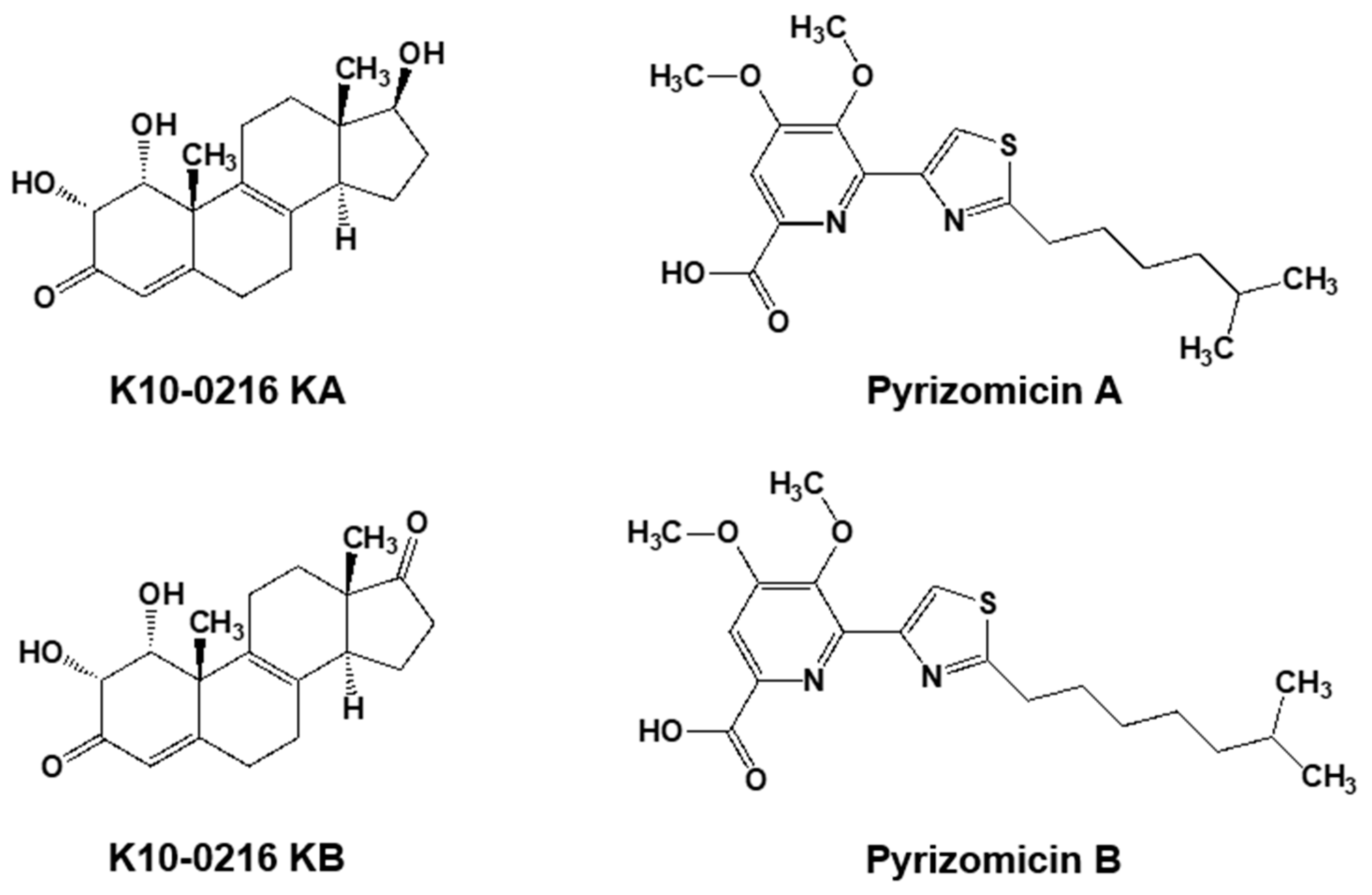
5.2. New Compounds Produced by Lechevalieria Aerocolonigenes K10-0216, Mangromycin A-I, K10-0216 KA & KB, and Pyrizomicin A & B
6. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Newman, D.J.; Cragg, G.M. Natural Products as sources of new drugs from 1981 to 2014. J. Nat. Prod. 2016, 79, 629–661. [Google Scholar] [CrossRef] [PubMed]
- Bérdy, J. Thought and facts about antibiotics: Where we are now and where we are heading. J. Antibiot. 2012, 65, 385–395. [Google Scholar] [CrossRef] [PubMed]
- Van Norman, G.A. Drugs, Devices, and the FDA: Part 1 an Overview of Approval Processes for Drugs. JACC Basic Transl. Sci. 2016, 1, 170–179. [Google Scholar] [CrossRef]
- Morgan, S.; Grootendorst, P.; Lexchin, J.; Cunningham, C.; Greyson, D. The cost of drug development: A systematic review. Health Policy 2011, 100, 4–17. [Google Scholar] [CrossRef] [PubMed]
- Dimasi, J.A.; Grabowski, G.H.; Hansen, R.W. Innovation in the pharmaceutical industry: New estimates of R&D costs. J. Health Econ. 2016, 47, 20–33. [Google Scholar] [PubMed]
- World Health Organization. Global Action Plan on Antimicrobial Resistance; World Health Organization: Geneva, Switzerland, 2015. [Google Scholar]
- World Health Organization. United Nations General Assembly High-Level Meeting on Antimicrobial Resistance; United Nations: New York, NY, USA, 2016. [Google Scholar]
- Barka, E.A.; Vatsa, P.; Sanchez, L.; Gaveau-Vaillant, N.; Jacquard, C.; Meier-Kolthoff, J.P.; Klenk, H.P.; Clément, C.; Ouhdouch, Y.; van Wezel, G.P. Taxonomy, physiology, and natural products of Actinobacteria. Microbiol. Mol. Biol. Rev. 2015, 80, 1–43. [Google Scholar] [CrossRef] [PubMed]
- Waksman, S.A.; Schatz, A.; Reynolds, D.M. Production of antibiotic substances by actinomycetes. Ann. N. Y. Acad. Sci. 2010, 1213, 112–124. [Google Scholar] [CrossRef] [PubMed]
- Felling, R.H.; Buchanan, G.O.; Mincer, T.J.; Kauffman, C.A.; Jensen, P.R.; Fenical, P.R. Salinosporamide A: A highly cytotoxic proteasome inhibitor from a novel microbial source, a marine bacterium of the new genus Salinospora. Angew. Chem. Int. Ed. 2003, 42, 355–357. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, K.; Gupta, R.K. Rare actinomycetes: A potential storehouse for novel antibiotics. Crit. Rev. Biotechnol. 2012, 32, 108–132. [Google Scholar] [CrossRef] [PubMed]
- Burg, R.W.; Miller, B.M.; Baker, J.; Birnbaum, E.E.; Currie, S.A.; Hartman, R.R.; Kong, Y.L.; Monaghan, R.L.; Olsen, G.; Putter, I.; et al. Avermectins, new family of potent anthelmintic agents: Producing organism and fermentation. Antimicrob. Agents Chemother. 1979, 15, 361–367. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, Y.; Matsumoto, A.; Seino, A.; Ueno, J.; Iwai, Y.; Ōmura, S. Streptomyces avermectinius sp. nov., an avermectin-producing strain. Int. J. Syst. Evolut. Microbiol. 2002, 52, 2163–2168. [Google Scholar]
- Campbell, W.C.; Fisher, M.H.; Stapley, E.O.; Albers-Schönberg, G.; Jacobs, T.A. Ivermectin: A potent new antiparasitic agent. Science 1983, 221, 823–828. [Google Scholar] [CrossRef] [PubMed]
- Crump, A.; Ōmura, S. Ivermectin, ‘wonder drug’ from Japan: The human use perspective. Proc. Jpn. Acad. Ser. B Phys. Biol. Sci. 2011, 87, 13–28. [Google Scholar] [CrossRef] [PubMed]
- Crump, A. Ivermectin: Enigmatic multifaceted ‘wonder’ drug continues to surprise and exceed expectations. J. Antibiot. 2017, 70, 495–505. [Google Scholar] [CrossRef] [PubMed]
- The Nobel Assembly at Karolinska Institute, The Nobel Prize in Physiology or Medicine 2015. Available online: https://www.nobelprize.org/nobel_prizes/medicine/prize_awarder/ (accessed on 24 May 2018).
- Satoshi Ōmura-Nobel Lecture: “A Splendid Gift from the Earth: The Origins and Impact of Avermectin”. Nobelprize.org. Available online: http://www.nobelprize.org/nobel _prizes/medicine/laureates/2015/omura-lecture.html (accessed on 24 May 2018).
- Takahashi, Y. Continuing fascination of exploration in natural substances from microorganisms. Biosci. Biotechnol. Biochem. 2017, 81, 6–12. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, A.; Takahashi, Y. Endophytic actinomycetes: Promising source of novel bioactive compounds. J. Antibiot. 2017, 70, 514–519. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, A. Potential of endophytic actinomycetes for agricultural applications. Agric. Biotechnol. 2018, 2, 180–185. [Google Scholar]
- Ōmura, S. Splendid Gifts from Microorganisms—The Achievements of Satoshi Ōmura and Collaborators, 5th ed.; Kitasato Institute for Life Sciences, Kitasato University: Tokyo, Japan, 2015. [Google Scholar]
- Ōmura, S.; Iwai, Y.; Hirano, A.; Nakagawa, A.; Awaya, J.; Tsuchiya, H.; Takahashi, Y.; Masuma, R. A new alkaloid AM-2282 of Streptomyces origin. J. Antibiot. 1977, 30, 275–282. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, Y.; Shinose, M.; Seino, A.; Iwai, Y.; Ōmura, S. Transfer of staurosporine-producing strain Streptomyces staurosporeus AM-2282 to the genus Saccharothrix as, Saccharothrix aerocolonigenes (labeda 1986) subsp. staurosporeus subsp. nov. Actinomycetologica 1995, 9, 19–26. [Google Scholar] [CrossRef]
- Xie, Q.; Wang, Y.; Huang, Y.; Wu, Y.; Ba, F.; Liu, Z. Description of Lentzea flaviverrucosa sp. nov. and transfer of the type strain of Saccharothrix aerocolonigenes subsp. staurosporea to Lentzea albida. Int. J. Syst. Evolut. Microbiol. 2002, 52, 1815–1820. [Google Scholar]
- Ōmura, S.; Sasaki, Y.; Iwai, Y.; Takeshima, H. Staurosporine, a potentially important gift from a microorganism. J. Antibiot. 1995, 48, 535–548. [Google Scholar] [CrossRef] [PubMed]
- Nakano, H.; Ōmura, S. Chemical biology of natural indolocarbazole products: 30 years since the discovery of staurosporine. J. Antibiot. 2009, 62, 17–26. [Google Scholar] [CrossRef] [PubMed]
- Nakashima, T.; Takahashi, Y.; Ōmura, S. Search for new compounds from Kitasato microbial library by physicochemical screening. Biochem. Pharmacol. 2017, 134, 42–55. [Google Scholar] [CrossRef] [PubMed]
- Ōmura, S.; Iwai, Y.; Takahashi, Y.; Sadakane, N.; Nakagawa, A.; Oiwa, R.; Hasegawa, Y.; Ikai, T. Herbimycin, a new antibiotic produced by a strain of Streptomyces. J. Antibiot. 1979, 32, 255–261. [Google Scholar] [CrossRef] [PubMed]
- Ōmura, S.; Otoguro, K.; Nishikiori, T.; Oiwa, R.; Iwai, Y. Setamycin, a new antibiotic. J. Antibiot. 1981, 34, 1253–1256. [Google Scholar] [CrossRef] [PubMed]
- Ōmura, S.; Fujimoto, K.; Otoguro, K.; Matsuzaki, K.; Moriguchi, R.; Tanaka, H.; Sasaki, Y. Lactacystin, a novel microbial metabolite, induces neuritogenesis of neuroblastoma cells. J. Antibiot. 1991, 44, 113–116. [Google Scholar] [CrossRef] [PubMed]
- Koomsiri, W.; Inahashi, Y.; Kimura, T.; Shiomi, K.; Takahashi, Y.; Ōmura, S.; Thamchaipenet, A.; Nakashima, T. Bisoxazolomycin A: A new natural product from ‘Streptomyces subflavus subsp. Irumaensis’ AM-3603. J. Antibiot. 2017, 70, 1142–1145. [Google Scholar] [PubMed]
- Nakashima, T.; Miyano, R.; Iwatsuki, M.; Shirahata, T.; Kimura, T.; Asami, Y.; Kobayashi, Y.; Shiomi, K.; Petersson, G.A.; Takahashi, Y.; Ōmura, S. Iminimycin A, the new iminium metabolite produced by Streptomyces griseus OS-3601. J. Antibiot. 2016, 69, 611–615. [Google Scholar] [CrossRef] [PubMed]
- Nakashima, T.; Miyano, R.; Matsuo, H.; Iwatsuki, M.; Shirahata, T.; Kobayashi, Y.; Shiomi, K.; Petersson, G.A.; Takahashi, Y.; Ōmura, S. Absolute configuration of iminimycin B, the new indolizidine alkaloid from Streptomyces griseus OS-3601. Tetrahedron Lett. 2016, 57, 3284–3286. [Google Scholar] [CrossRef]
- Nakashima, T.; Boonsnongcheep, P.; Kimura, T.; Iwatsuki, M.; Sato, N.; Nonaka, K.; Prathanturarug, S.; Takahashi, Y.; Ōmura, S. New compounds, nanaomycin F and G, discovered by physicochemical screening from a culture broth of Streptomyces rosa subsp. notoensis OS-3966. J. Biosci. Bioeng. 2015, 120, 596–600. [Google Scholar] [PubMed]
- Nakashima, T.; Kimura, T.; Miyano, R.; Matsuo, H.; Hirose, T.; Kimishima, A.; Nonaka, K.; Iwatsuki, M.; Nakanishi, J.; Takahashi, Y.; et al. Nanaomycin H: A new nanaomycin analog. J. Biosci. Bioeng. 2017, 123, 765–770. [Google Scholar] [CrossRef] [PubMed]
- Inahashi, Y.; Iwatsuki, M.; Ishiyama, A.; Matsumoto, A.; Hirose, T.; Oshita, J.; Sunazuka, T.; Watanalai, P.W.; Takahashi, Y.; Kaiser, M.; et al. Actinoallolides A-E, New anti-trypanosomal macrolides, produced by an endophytic actinomycete, Actinoallomurus fulvus MK10-036. Org. Lett. 2015, 17, 864–867. [Google Scholar] [CrossRef] [PubMed]
- Suga, T.; Kimura, T.; Inahashi, Y.; Iwatsuki, M.; Nonaka, K.; Také, A.; Matsumoto, A.; Takahashi, Y.; Ōmura, S.; Nakashima, T. Hamuramicins A and B, 22-membered macrolides, produced by an endophytic actinomycete Allostreptomyces sp. K12-0794. J. Antibiot. 2018. [Google Scholar] [CrossRef] [PubMed]
- Inahashi, Y.; Matsumoto, A.; Ōmura, S.; Takahashi, Y. Streptosporangium oxazolinicum sp. nov., a novel endophytic actinomycete producing new antitrypanosomal antibiotics, spoxazomicins. J. Antibiot. 2011, 64, 297–302. [Google Scholar] [CrossRef] [PubMed]
- Inahashi, Y.; Iwatsuki, M.; Ishiyama, A.; Namatame, M.; Nishihara-Tsukashima, A.; Matsumoto, A.; Hirose, T.; Sunazuka, T.; Yamada, H.; Otoguro, K.; et al. Spoxazomicins A-C, novel antitrypanosomal alkaloids produced by an endophytic actinomycete, Streptosporangium oxazolinicum K07-0460T. J. Antibiot. 2011, 64, 303–307. [Google Scholar] [CrossRef] [PubMed]
- Nakashima, T.; Okuyama, R.; Kamiya, Y.; Matsumoto, A.; Iwatsuki, M.; Inahashi, Y.; Yamaji, K.; Takahashi, Y.; Ōmura, S. Trehangelins A, B and C, novel photo-oxidative hemolysis inhibitors produced by an endophytic actinomycete, Polymorphospora rubra K07-0510. J. Antibiot. 2013, 66, 311–317. [Google Scholar] [CrossRef] [PubMed]
- Inahashi, Y.; Shiraishi, T.; Palm, K.; Takahashi, Y.; Ōmura, S.; Kuzuyama, T.; Nakashima, T. Biosynthesis of Trehangelin in Polymorphospora rubra K07-0510: Identification of methabolic pathway to angelyl-CoA. ChemBioChem 2016, 17, 1442–1447. [Google Scholar] [CrossRef] [PubMed]
- Nakashima, T.; Iwatsuki, M.; Ochiai, J.; Kamiya, Y.; Nagai, K.; Matsumoto, A.; Ishiyama, A.; Otoguro, K.; Shiomi, K.; Takahashi, Y.; et al. Mangromicins A and B: Structure and antitrypanosomal activity of two new cyclopentadecane compounds from Lechevalieria aerocolonigenes K10-0216. J. Antibiot. 2014, 67, 253–260. [Google Scholar] [CrossRef] [PubMed]
- Nakashima, T.; Kamiya, Y.; Iwatsuki, M.; Takahashi, Y.; Ōmura, S. Mangromicins, six new anti-oxidative agents isolated from a culture broth of the actinomycete, Lechevalieria aerocolonigenes K10-0216. J. Antibiot. 2014, 67, 533–539. [Google Scholar] [CrossRef] [PubMed]
- Nakashima, T.; Kamiya, Y.; Iwatsuki, M.; Sato, N.; Takahashi, Y.; Ōmura, S. Mangromicin C, a new analog of mangromicin. J. Antibiot. 2015, 68, 220–222. [Google Scholar] [CrossRef] [PubMed]
- Nakashima, T.; Kamiya, Y.; Yamaji, K.; Iwatsuki, M.; Sato, N.; Takahashi, Y.; Ōmura, S. New steroidal compounds from an actinomycete strain, Lechevalieria aerocolonigenes K10-0216. J. Antibiot. 2015, 68, 348–350. [Google Scholar] [CrossRef] [PubMed]
- Kimura, T.; Inahashi, Y.; Matsuo, H.; Suga, T.; Iwatsuki, M.; Shiomi, K.; Takahashi, Y.; Ōmura, S.; Nakashima, T. Pyrizomicin A and B: Structure and bioactivity of new thiazolyl pyridines from Lechevalieria aerocolonigenes K10-0216. J. Antibiot. 2018, 71, 606–608. [Google Scholar] [CrossRef] [PubMed]
- Kimura, T.; Tajima, A.; Inahashi, Y.; Iwatsuki, M.; Kasai, H.; Mokudai, T.; Niwano, Y.; Shiomi, K.; Takahashi, Y.; Ōmura, S.; et al. Mumiamicin: Structure and bioactivity of a new furan fatty acid from Mumia sp. YSP-2-79. J. Gen. Appl. Microbiol. 2018, 64, 62–67. [Google Scholar] [CrossRef] [PubMed]
- Kimura, T.; Iwatsuki, M.; Asami, Y.; Ishiyama, A.; Hokari, R.; Otoguro, K.; Matsumoto, A.; Sato, N.; Shiomi, K.; Takahashi, Y.; et al. Anti-trypanosoma compound, sagamilactam, a new polyene macrocyclic lactam from Actinomadura sp. K13-0306. J. Antibiot. 2016, 69, 818–824. [Google Scholar] [CrossRef] [PubMed]
- Izuta, S.; Kosaka, S.; Kawai, M.; Miyano, R.; Matsuo, H.; Matsumoto, A.; Nonaka, K.; Takahashi, Y.; Ōmura, S.; Natashima, T. Dipyrimicins A and B, microbial compounds interacted with ergosterolresin, from Amycolatopsis sp. K16-0194. J. Antibiot. 2018, 71, 535–537. [Google Scholar] [CrossRef] [PubMed]
- Bringmann, G.; Kajahn, I.; Reichert, M.; Pedersen, S.E.; Faber, J.H.; Gulder, T.; Brun, R.; Christensen, S.B.; Ponte-Sucre, A.; Moll, H.; et al. Ancistrocladinium A and B, the first N, C-coupled naphthyldihydroisoquinoline alkaloids, from a Congolese Ancistrocladus species. J. Org. Chem. 2006, 71, 9348–9356. [Google Scholar] [CrossRef] [PubMed]
- Ohnishi, Y.; Ishikawa, J.; Hara, H.; Suzuki, H.; Ikenoya, M.; Ikeda, H.; Yamashita, A.; Hattori, M.; Horinouchi, S. Genome sequence of the streptomycin-producing microorganism Streptomyces griseus IFO 13350. J. Bacteriol. 2008, 190, 4050–4060. [Google Scholar] [CrossRef] [PubMed]
- Ōmura, S.; Tanaka, H.; Koyama, Y.; Oiwa, R.; Katagiri, M. Nanaomycins A and B, new antibiotics produced by a strain of Streptomyces. J. Antibiot. 1974, 27, 363–365. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, H.; Marumo, H.; Nagai, T.; Okada, M.; Taniguchi, K. Nanaomycins, new antibiotics produced by a strain of Streptomyces. III. A new component, nanaomycin C, and biological activities of nanaomycin derivatives. J. Antibiot. 1975, 28, 925–930. [Google Scholar] [CrossRef] [PubMed]
- Kasai, M.; Shirahata, K.; Ishii, S.; Mineura, K.; Marumo, H.; Tanaka, H.; Ōmura, S. Structure of nanaomycin E, a new nanaomycin. J. Antibiot. 1979, 32, 442–445. [Google Scholar] [CrossRef] [PubMed]
- Kitaura, K.; Araki, Y.; Marumo, H. The therapeutic effect of nanaomycin A against experimental Trichophyton mentagrophytes infection in guinea pigs (author’s transl). Jpn. J. Antibiot. 1980, 33, 728–732. [Google Scholar] [PubMed]
- Ōmura, S.; Takahashi, Y.; Nakashima, T.; Matsumoto, A.; Nakanishi, J.; Matsuo, H. Epithelial-Mesenchymal Transition Induced Cell Proliferation Inhibitor. Patent No. PCT/JP2017/034819, 26 September 2016. [Google Scholar]
- Bode, H.B.; Bethe, B.; Höfs, R.; Zeeck, A. Big Effects from Small Changes: Possible Ways to Explore Nature’s Chemical Diversity. ChemBioChem 2002, 3, 619–627. [Google Scholar] [CrossRef]
- Yoon, S.H.; Ha, S.M.; Kwon, S.; Lim, J.; Kim, Y.; Seo, H.; Chun, J. Introducing EzBioCloud: A taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int. J. Syst. Evol. Microbiol. 2017, 67, 1613–1617. [Google Scholar] [PubMed]
- Kamiya, Y.; Enomoto, Y.; Matsukuma, S.; Nakashima, T.; Takahashi, Y.; Omura, S. Study on anti-aging effects of trehangelin, a new trehalose compound. In Proceedings of the 136th Annual Meeting of the Pharmaceutical Society of Japan, Yokohama, Japan, 26–29 March 2016; p. 137. [Google Scholar]
- Azman, A.S.; Othman, I.; Velu, S.S.; Chan, K.G.; Lee, L.H. Mangrove rare actinobacteria: Taxonomy, natural compound, and discovery of bioactivity. Front. Microbiol. 2015, 20, 856. [Google Scholar] [CrossRef] [PubMed]
- Pozzi, R.; Simone, M.; Mazzetti, C.; Maffioli, S.; Monciardini, P.; Cavaletti, L.; Bamonte, R.; Sosio, M.; Donadio, S. The genus Actinoallomurus and some of its metabolites. J. Antibiot. 2011, 64, 133–139. [Google Scholar] [CrossRef] [PubMed]
- Takada, H.; Yamada, T.; Hirose, T.; Ishihara, T.; Nakashima, T.; Takahashi, Y.; Ōmura, S.; Sunazuka, T. Total synthesis and determination of the absolute configuration of naturally occurring mangromicin A, with potent antitrypanosomal activity. Org. Lett. 2017, 19, 230–233. [Google Scholar] [CrossRef] [PubMed]
- Amann, R.I.; Ludwig, W.; Schleifer, K.H. Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol. Rev. 1995, 59, 143–169. [Google Scholar] [PubMed]
| No. | Compound | Producing Microorganism | Source | Biological Activity | References |
|---|---|---|---|---|---|
| 1 | Bisoxazolomycin | Streptomyces subflavus subsp. irumaensis AM-3603 | * KML Irumamycin producing strain, ** 36 years | Antibacterial | [32] |
| 2 | Iminimycin A & B | Streptomyces griseus OS-3601 | * KML, Streptomycin producing strain, ** 43 years | Antibacterial | [33,34] |
| 3 | Nanaomycin F–H | “Streptomyces rosa subsp. Notoensis” OS-3966 | * KML Nanaomycin producing strain, ** 36 years | Inhibitor of Epithelial-Mesenchymal Transition induced cells | [35,36] |
| 4 | Actinoallolide A–E | Actinoallomurus fulvus MK10-036 | Roots of Capsicum frutescents in Thailand | Antitrypanosomal | [37] |
| A. fulvus K09-0307 | Roots of mondo grass in Saitama Pref., Japan | ||||
| 5 | Hamuramicin A & B | Allostreptomyces sp. K12-0794 | Roots of fern in Hamura city, Tokyo, Japan | Antibacterial | [38] |
| 6 | Spoxazomicin A–C | Streptosporangium oxazolinicum K07-0460T | Roots of orchid in Iriomote Island, Japan | Antitrypanosomal | [39,40] |
| 7 | Trehangelin A–C | Polymorphospora rubra K07-0510 | Roots of orchid in Iriomote Island, Japan | Anti-Lipid peroxidation | [41,42] |
| Enhanced production of collagen | |||||
| 8 | Mangromicin A–I | Lechevalieria aerocolonigenes K10-0216 | Sediment from mangrove forest in Iriomote Island, Japan | Antitrypanosomal | [43,44,45] |
| Antioxidative | |||||
| 9 | K10-0216 KA & KB | Lechevalieria aerocolonigenes K10-0216 | Sediment from mangrove forest in Iriomote Island, Japan | Inhibitory effect on the lipid accumulation | [46] |
| 10 | Pyrizomicin A & B | Lechevalieria aerocolonigenes K10-0216 | Sediment from mangrove forest in Iriomote Island, Japan | Antimicrobial | [47] |
| 11 | Mumiamicin | Mumia sp. YSP-2-79 | Sea sediment, Namako Pond in Kagoshima Pref., Japan | Antibacterial | [48] |
| Antioxidative | |||||
| 12 | Sagamilactam | Actinomadura sp. K13-0306 | Soil, Kanagawa Pref., Japan | Cytotoxicity | [49] |
| Antitrypanosomal | |||||
| 13 | Dipyrimicin A & B | Amycolatopsis sp. K16-0194 | Soil, Okinawa Pref., Japan | Antibacterial | [50] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Takahashi, Y.; Nakashima, T. Actinomycetes, an Inexhaustible Source of Naturally Occurring Antibiotics. Antibiotics 2018, 7, 45. https://doi.org/10.3390/antibiotics7020045
Takahashi Y, Nakashima T. Actinomycetes, an Inexhaustible Source of Naturally Occurring Antibiotics. Antibiotics. 2018; 7(2):45. https://doi.org/10.3390/antibiotics7020045
Chicago/Turabian StyleTakahashi, Yōko, and Takuji Nakashima. 2018. "Actinomycetes, an Inexhaustible Source of Naturally Occurring Antibiotics" Antibiotics 7, no. 2: 45. https://doi.org/10.3390/antibiotics7020045
APA StyleTakahashi, Y., & Nakashima, T. (2018). Actinomycetes, an Inexhaustible Source of Naturally Occurring Antibiotics. Antibiotics, 7(2), 45. https://doi.org/10.3390/antibiotics7020045





