The Cellular Mechanisms that Ensure an Efficient Secretion in Streptomyces
Abstract
1. Introduction
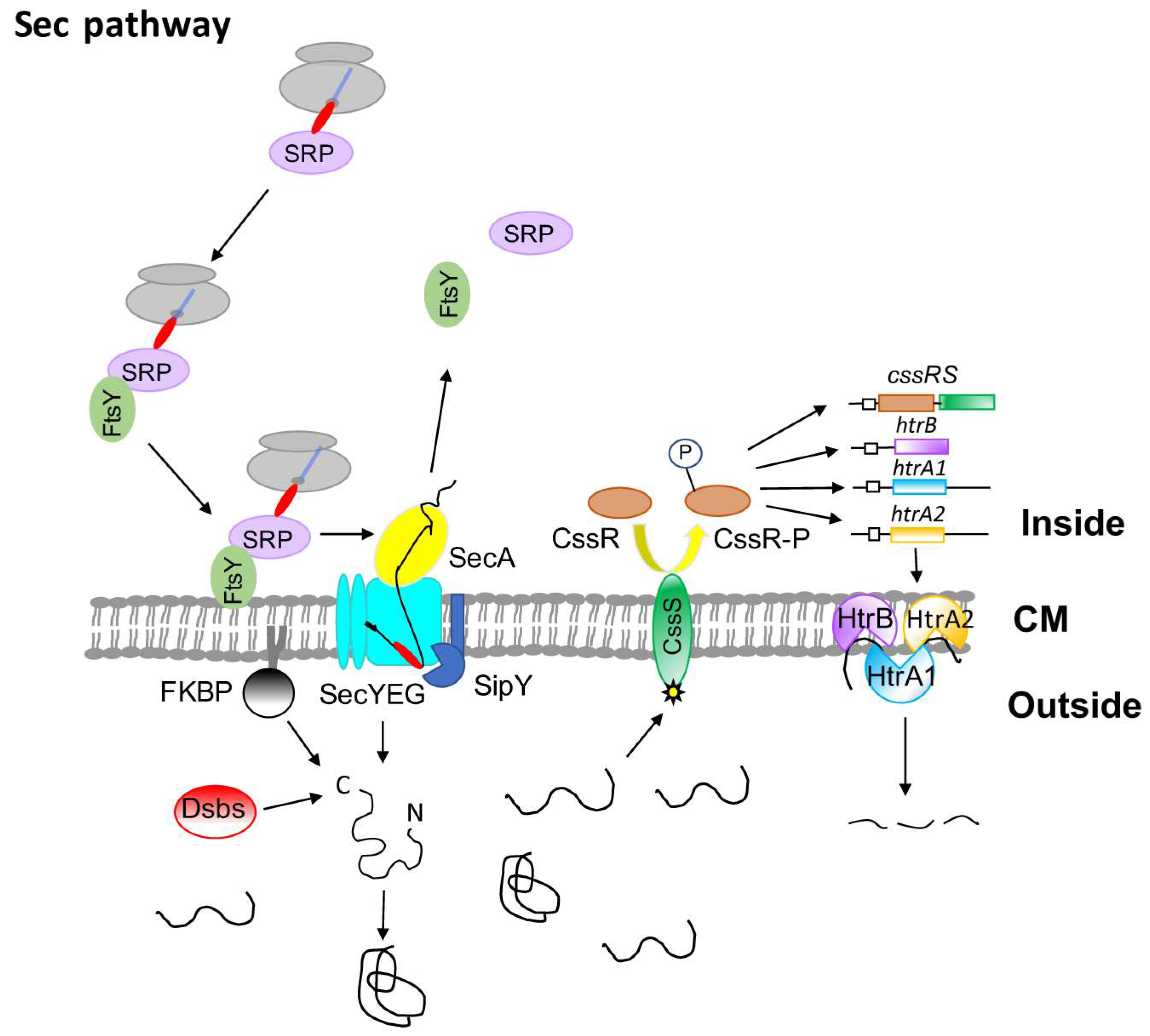
2. The Sec Pathway
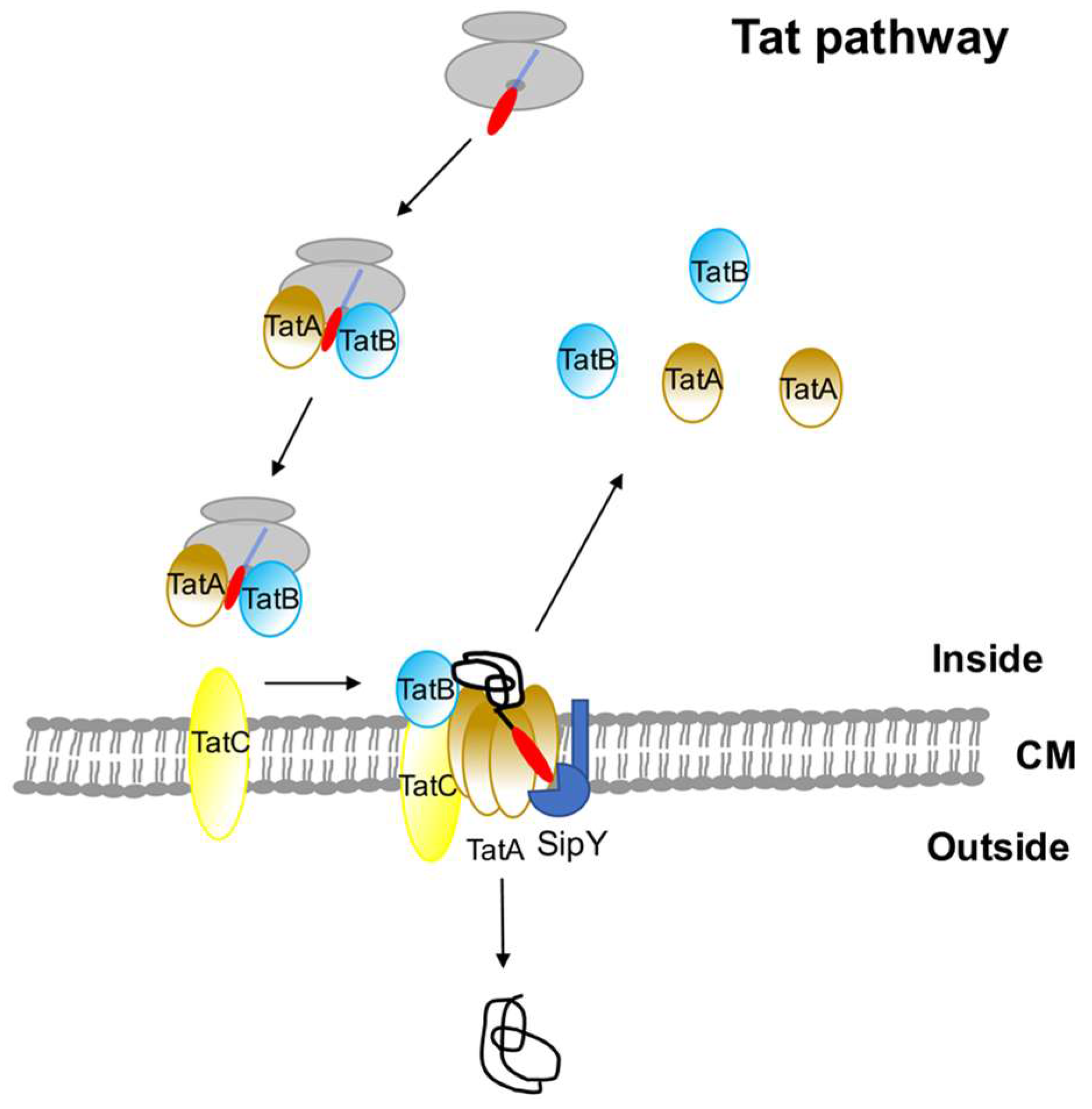
3. The Twin-Arginine (Tat) Pathway
4. Stresses Caused by Overproduction of Secretory Proteins in Streptomycetes
4.1. Temporal Translocase Blockage
4.2. Induction of the Stringent Response
5. Overproduction of Model Secretory Proteins in S. lividans
6. Extracellular Stability of Secretory Proteins
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Gilbert, M.; Morosoli, R.; Shareck, F.; Kluepfel, D. Production and secretion of proteins by streptomycetes. Crit. Rev. Biotechnol. 1995, 15, 13–19. [Google Scholar] [CrossRef] [PubMed]
- Chater, K.F. Taking a genetic scalpel to the Streptomyces colony. Microbiology 1998, 144, 1465–1478. [Google Scholar] [CrossRef]
- Cruz-Morales, P.; Vijgenboom, E.; Iruegas-Bocargo, F.; Girard, G.; Yáñez-Guerra, L.A.; Ramos-Aboites, H.E.; Pernodet, J.L.; Anné, J.; van Wezel, G.P.; Barona-Gómez, F. The genome sequence of Streptomyces lividans 66 reveals a novel tRNA-dependent peptide biosynthetic system within a metal-related genomic island. Genome Biol. Evol. 2013, 5, 1165–1175. [Google Scholar] [CrossRef] [PubMed]
- Rückert, C.; Albersmeier, A.; Busche, T.; Jaenicke, S.; Winkler, A.; Friðjónsson, Ó.H.; Hreggviðsson, G.Ó.; Lambert, C.; Badcock, D.; Bernaerts, K.; et al. Complete genome sequence of Streptomyces lividans TK24. J. Biotechnol. 2015, 199, 21–22. [Google Scholar] [CrossRef] [PubMed]
- Parro, V.; Mellado, R.P. Effect of glucose on agarase overproduction by Streptomyces. Gene 1994, 145, 49–55. [Google Scholar] [CrossRef]
- Isiegas, C.; Parro, V.; Mellado, R.P. Streptomyces lividans as a host to produce and secrete Escherichia coli TEM β-lactamase. Lett. Appl. Microbiol. 1999, 28, 321–326. [Google Scholar] [CrossRef] [PubMed]
- Anné, J.; Maldonado, B.; Van Impe, J.; Van Mellaert, L.; Bernaerts, K. Recombinant protein production and streptomycetes. J. Biotechnol. 2012, 158, 159–167. [Google Scholar] [CrossRef] [PubMed]
- Driessen, A.J.; Nouwen, N. Protein translocation across the bacterial cytoplasmic membrane. Annu. Rev. Biochem. 2008, 77, 643–667. [Google Scholar] [CrossRef] [PubMed]
- Berks, B.C.; Sargent, F.; Palmer, T. The Tat protein export pathway. Mol. Microbiol. 2000, 35, 260–274. [Google Scholar] [CrossRef] [PubMed]
- Schrempf, H.; Merling, P. Extracellular Streptomyces lividans vesicles: Composition, biogenesis and antimicrobial activity. 2015. Microb. Biotechnol. 2015, 8, 644–658. [Google Scholar] [CrossRef] [PubMed]
- Pallen, M.J. The ESAT-6/WXG100 superfamily- and a new Gram-positive secretion system? Trends Microbiol. 2002, 10, 209–212. [Google Scholar] [CrossRef]
- Chater, K.F.; Biró, S.; Lee, K.; Palmer, T.; Schrempf, H. The complex extracelular biology of Streptomyces. FEMS Microbiol. Rev. 2010, 34, 171–198. [Google Scholar] [CrossRef] [PubMed]
- Sala, A.; Bordes, P.; Genevaux, P. Multitasking SecB chaperones in bacteria. Front. Microbiol. 2014, 5, 666. [Google Scholar] [CrossRef] [PubMed]
- Kunst, F.; Ogasawara, N.; Moszer, I.; Albertini, A.M.; Alloni, G.; Azevedo, V.; Bertero, M.G.; Bessières, P.; Bolotin, A.; Borchert, S.; et al. The complete genome sequence of the Gram-positive bacterium Bacillus subtilis. Nature 1997, 390, 249–256. [Google Scholar] [CrossRef] [PubMed]
- Bentley, S.D.; Chater, K.F.; Cerdeño-Tárraga, A.M.; Challis, G.L.; Thomson, N.R.; James, K.D.; Harris, D.E.; Quail, M.A.; Kieser, H.; Harper, D.; et al. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3. Nature 2002, 417, 141–147. [Google Scholar] [CrossRef] [PubMed]
- Keenan, R.J.; Freymann, D.M.; Stroud, R.M.; Walter, P. The signal recognition particle. Annu. Rev. Biochem. 2001, 70, 755–775. [Google Scholar] [CrossRef] [PubMed]
- Cao, T.; Saier, M. The general protein secretory pathway: Phylogenetic analyses leading to evolutionary conclusions. Biochim. Biophys. Acta 2003, 1609, 115–125. [Google Scholar] [CrossRef]
- Pohlschöder, M.; Dilks, K.; Hand, N.; Wesley Rose, R. Translocation of proteins across archaeal cytoplasmic membranes. FEMS Microbiol. Rev. 2004, 28, 3–24. [Google Scholar] [CrossRef] [PubMed]
- Walter, P.; Johnson, A.E. Signal recognition and protein targeting to the endoplasmic reticulum membrane. Annu. Rev. Cell Biol. 1994, 10, 87–119. [Google Scholar] [CrossRef] [PubMed]
- Wollin, S.L. From the elephant to E. coli: SRP-dependent protein targeting. Cell 1994, 77, 787–790. [Google Scholar] [CrossRef]
- Miller, J.; Bernstein, H.D.; Walter, P. Interaction of E. coli Ffh/4.5 S ribonucleoprotein and FtsY mimics that of mammalian signal recognition particle and its receptor. Nature 1994, 366, 351–354. [Google Scholar] [CrossRef] [PubMed]
- Powers, T.; Walter, P. Co-translational protein targeting catalised by the Escherichia coli signal recognition particle and its receptor. EMBO J. 1997, 16, 4880–4886. [Google Scholar] [CrossRef] [PubMed]
- Millman, J.S.; Andrews, D.W. A site-specific, membrane-dependent cleavage event defines the membrane binding domain of FtsY. J. Biol. Chem. 1999, 274, 33227–33234. [Google Scholar] [CrossRef] [PubMed]
- Herscovits, A.A.; Bochkareva, E.; Bibi, E. New prospects in studying the bacterial signal recoginiton particle pathway. Mol. Microbiol. 2000, 38, 927–939. [Google Scholar] [CrossRef]
- De Gier, J.W.; Mansournia, P.; Valent, Q.A.; Philips, G.J.; Luirink, J.; von Heijne, G. Assembly of a cytoplasmic membrane protein in Escherichia coli is dependent on the signal recognition particle. FEBS Lett. 1996, 399, 307–309. [Google Scholar] [CrossRef]
- Ulbrandt, N.D.; Newitt, J.A.; Bernstein, H.D. The E. coli signal recognition particle is required for the insertion of a subset of inner membrane proteins. Cell 1997, 88, 187–196. [Google Scholar] [CrossRef]
- Beha, D.; Deitermann, S.; Müller, M.; Koch, H.G. Export of beta-lactamase is independent of the signal recognition particle. J. Biol. Chem. 2003, 278, 22161–22167. [Google Scholar] [CrossRef] [PubMed]
- Beck, K.; Wu, L.F.; Brunner, J.; Muller, M. Discrimination between SRP-and SecA/SecB-dependent substrates involves selective recognition of nascent chains by SRP and trigger factor. EMBO J. 2000, 19, 134–143. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, D.; Yahagi, S.; Yamazaki, T.; Yamane, K. Bacillus subtilis histone-like protein HBsu, is an integral component of a SRP-like particle that can bind the Alu domain of small cytoplasmic RNA. J. Biol. Chem. 1999, 274, 13569–13576. [Google Scholar] [CrossRef] [PubMed]
- Zanen, G.; Antelmann, H.; Meima, R.; Jongbloed, J.D.; Kolkman, M.; Hecker, M.; van Dijl, J.M.; Quax, W.J. Proteomic dissection of potential signal recognition particle dependence in protein secretion by Bacillus subtilis. Proteomics 2006, 6, 3636–3648. [Google Scholar] [CrossRef] [PubMed]
- Zanen, G.; Houben, E.N.; Meima, R.; Tjalsma, J.; Jongbloed, J.D.; Westers, H.; Oudega, B.; Luirink, J.; van Dijl, J.M.; Quax, W.J. Signal peptide hydrophobibity is a critical for early stages in protein export by Bacillus subtilis. FEBS J. 2005, 272, 4617–4630. [Google Scholar] [CrossRef] [PubMed]
- Palacín, A.; de la Fuente, R.; Valle, I.; Rivas, L.A.; Mellado, R.P. Streptomyces lividans contains a minimal functional signal recognition particle that is involved in protein secretion. Microbiology 2003, 149, 2435–2442. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Palomino, C. Role of the Signal Recoginition Particle (SRP) in the Extracelular Protein Secretion in Streptomyces lividans. Ph.D. Thesis, Universidad Autónoma de Madrid, Madrid, Spain, 2006. [Google Scholar]
- Palacín, A.; Parro, V.; Geukens, N.; Anné, J.; Mellado, R.P. SipY is the Streptomyces lividans type I signal peptidase exerting a major effect on protein secretion. J. Bacteriol. 2002, 184, 4875–4880. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Bibi, E.; Herskovits, A.A.; Bochkareva, E.; Zelazny, A. Putative integral membrane SRP receptors. Trends Biochem. Sci. 2001, 26, 15–16. [Google Scholar] [CrossRef]
- Palomino, C.; Mellado, R.P. The Streptomyces lividans cytoplasmic signal recognition particle receptor FtsY is involved in protein secretion. J. Mol. Microbiol. Biotechnol. 2005, 9, 57–62. [Google Scholar] [CrossRef] [PubMed]
- Blanco, J.; Driessen, A.J.; Coque, J.J.; Martín, J.F. Biochemical characterization of the SecA protein of Streptomyces lividans interaction with nucleotides, binding to membrane vesicles and in vitro translocation of proAmy protein. Eur. J. Biochem. 1998, 257, 472–478. [Google Scholar] [CrossRef] [PubMed]
- Palomino, C.; Mellado, R.P. Influence of a Streptomyces lividans SecG functional analogue on protein secretion. Int. Microbiol. 2008, 11, 25–31. [Google Scholar] [PubMed]
- Van Welly, K.H.M.; Swaving, J.; Broekhuizen, C.P.; Rose, M.; Quax, W.J.; Driessen, A.J.M. Functional identification of the product of Bacillus subtilis yvaL gene as a SecG homologue. J. Bacteriol. 1999, 181, 1786–1792. [Google Scholar]
- Nishiyama, K.; Hanada, M.; Tokuda, H. Disruption of the gene encoding p12 (SecG) reveals the direct involvement and important function of SecG in protein translocation of Escherichia coli at low temperature. EMBO J. 1994, 13, 3272–3277. [Google Scholar] [PubMed]
- Gullón, S.; Vicente, R.L.; Valverde, J.R.; Marín, S.; Mellado, R.P. Exploring the feasibility of the Sec route to secrete proteins using the Tat route in Streptomyces lividans. Mol. Biotechnol. 2015, 57, 931–938. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Li, Y.; Sun, N.; Sun, Z.; Lv, L.; Wang, Y.; Shen, L.; Li, Y.Q. Function and evolution of two forms of SecDF homologs in Streptomyces coelicolor. PLoS ONE 2014, 9, e105237. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Sun, N.; Wu, S.; Li, Y.Q.; Wang, Y. Genomic data mining reveals a rich repertoire of transport proteins in Streptomyces. BMC Genom. 2016, 17 (Suppl. 7), 510. [Google Scholar] [CrossRef] [PubMed]
- Scotti, P.A.; Urbanus, M.L.; Brunner, J.; de Gier, J.W.; von Heijne, G.; van der Does, C.; Driessen, A.J.; Oudega, B.; Luirink, J.; Yid, C. The Escherichia coli homologue of mitochondrial Oxa1p, is a component of the Sec translocase. EMBO J. 2000, 19, 542–549. [Google Scholar] [CrossRef] [PubMed]
- Van der Laan, M.; Bechtluft, P.; Kol, S.; Nouwen, N.; Driessen, A.J. F1F0 ATP synthase subunit c is a substrate of the novel YidC pathway for membrane protein biogenesis. J. Cell Biol. 2004, 165, 213–222. [Google Scholar] [CrossRef] [PubMed]
- Serek, J.; Bauer-Manz, G.; Struhalla, G.; van der Berg, L.; Kiefer, D.; Dalbey, R.; Kuhn, A. Escherichia coli YidC is a membrane insertase for Sec-independent proteins. EMBO J. 2004, 23, 294–301. [Google Scholar] [CrossRef] [PubMed]
- Keller, R.; de Keyzer, J.; Driessen, A.J.; Palmer, T. Co-operation between different targeting pathways during integration of a membrane protein. J. Cell Biol. 2012, 199, 303–315. [Google Scholar] [CrossRef] [PubMed]
- Berks, B.C.; Palmer, T.; Sargent, F. The Tat protein translocation pathway and its role in microbial physiology. Adv. Microb. Physiol. 2003, 47, 187–254. [Google Scholar] [PubMed]
- Eimer, E.; Fröbel, J.; Blümmel, A.S.; Müller, M. TatE as a regular constituent of bacterial twin-arginine protein translocases. J. Biol. Chem. 2015, 290, 29281–29289. [Google Scholar] [CrossRef] [PubMed]
- Dilks, K.; Giménez, M.I.; Pohlschröder, M. Genetic and biochemical analysis of the twin-arginine translocation pathway in halophilic archarea. J. Bacteriol. 2005, 187, 8013–8104. [Google Scholar] [CrossRef] [PubMed]
- Jongbloed, J.D.; Grieger, U.; Antelmann, H.; Hecker, M.; Nijland, R.; Bron, S.; van Dijl, J.M. Two minimal Tat translocases in Bacillus. Mol. Microbiol. 2004, 545, 1319–1325. [Google Scholar] [CrossRef] [PubMed]
- Pop, O.; Martín, U.; Abel, C.; Muller, J.P. The twin-arginine signal peptide of PhoD and the TatAd/Cd proteins of Bacillus subtilis form an autonomous Tat translocation system. J. Biol. Chem. 2002, 277, 3268–3273. [Google Scholar] [CrossRef] [PubMed]
- Goosens, V.; Monteferrante, C.G.; van Dijl, J.M. The tat system of Gram positive bacteria. Biochim. Biophys. Acta 2014, 1843, 1698–1706. [Google Scholar] [CrossRef] [PubMed]
- Tsolis, K.C.; Tsare, E.P.; Orfanoudaki, G.; Busche, T.; Kanaki, K.; Ramakrishnan, R.; Rousseau, F.; Schymkowitz, J.; Rückert, C.; Kalinowski, J.; et al. Comprehensive subcellular topologies of polypeptides in Streptomyces. Microb. Cell Fact. 2018, 17, 43. [Google Scholar] [CrossRef] [PubMed]
- De Keersmaeker, S.; Vrancken, K.; Van Mellaert, L.; Anné, J.; Geukens, N. The Tat pathway in Streptomyces lividans: Interaction of Tat subunits and their role in translocation. Microbiology 2007, 153, 1087–1094. [Google Scholar] [CrossRef] [PubMed]
- Berks, B.C. A common export pathway for proteins binding complex redox cofactors? Mol. Microbiol. 1996, 22, 393–404. [Google Scholar] [CrossRef] [PubMed]
- Stanley, N.T.; Palmer, T.; Berks, B.C. The twin-arginine consensus motif of Tat signal peptides is involved in Sec-independent protein targeting in Escherichia coli. J. Biol. Chem. 2000, 275, 11591–11596. [Google Scholar] [CrossRef] [PubMed]
- Widdick, D.A.; Dilks, K.; Chandra, G.; Bottrill, A.; Naldrett, M.; Pohlschroder, M.; Palmer, T. The twin-arginine translocation pathway is a major route of protein export in Streptomyces coelicolor. Proc. Natl. Acad. Sci. USA 2006, 103, 17927–17932. [Google Scholar] [CrossRef] [PubMed]
- Palmer, T.; Berks, B.C. The twin-arginine translocation (Tat) protein export pathway. Nat. Rev. Microbiol. 2012, 10, 483–496. [Google Scholar] [CrossRef] [PubMed]
- Bendtsen, J.D.; Nielsen, H.; Widdick, D.; Palmer, T.; Brunak, S. Prediction of twin-arginine signal peptides. BMC Bioinf. 2005, 6, 167–175. [Google Scholar] [CrossRef] [PubMed]
- Dilks, K.; Rose, R.W.; Hartmann, E.; Pohschröder, M. Prokaryotic utilization of the twin-arginine translocation pathway: A genomic survey. J. Bacteriol. 2003, 185, 1478–1483. [Google Scholar] [CrossRef] [PubMed]
- Schaerlaekens, K.; van Mellaert, L.; Lammertyn, E.; Geukens, N.; Anné, J. The importance of the Tat-dependent protein secretion pathway in Streptomyces as revealed by phenotypic changes in tat deletion mutants and genome analysis. Microbiology 2004, 150, 21–31. [Google Scholar] [CrossRef] [PubMed]
- Petrus, M.L.C.; Vijgenboom, E.; Chaplin, A.K.; Worrall, J.A.R.; van Wezel, G.P.; Claessen, D. The Dyp-type peroxidase DtpA is a Tat-substrate required for GlxA maturation and morphogenesis in Streptomyces. Open Biol. 2015, 6, 150149. [Google Scholar] [CrossRef] [PubMed]
- Palmer, T.; Sargent, F.; Berks, B.C. Export of complex cofactor-containing proteins y the bacterial Tat pathway. Trends Microbiol. 2005, 13, 175–180. [Google Scholar] [CrossRef] [PubMed]
- Gullón, S.; Palomino, C.; Navajas, R.; Paradela, A.; Mellado, R.P. Translocase and major signal peptidase malfunctions affect aerial mycelium formation in Streptomyces lividans. J. Biotecnol. 2012, 160, 112–122. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.W.; Hesketh, A.; Kim, E.S.; Song, J.Y.; Lee, D.H.; Kin, I.S.; Chater, K.F.; Lee, K.J. Complex extracellular interactions of proteases and a protease inhibitor influence multicellular development of Streptomyces coelicolor. Mol. Microbiol. 2008, 70, 1180–1193. [Google Scholar] [CrossRef] [PubMed]
- Rozas, D.; Gullón, S.; Mellado, R.P. A novel two-component system involved in the transition to secondary metabolism in Streptomyces coelicolor. PLoS ONE 2012, 7, e31760. [Google Scholar] [CrossRef] [PubMed]
- Ogura, M.; Yamaguchi, H.; Yoshida, K.; Fujita, Y.; Tanaka, T. DNA microarray analysis of Bacillus subtilis DegU, ComA and PhoP regulons: An approach to comprehensive analysis of B. subtilis two-component regulatory systems. Nucl. Acids Res. 2001, 29, 3804–3813. [Google Scholar] [CrossRef] [PubMed]
- Mäder, U.; Antelmann, H.; Buder, T.; Dahl, M.K.; Hecker, M.; Homuth, G. Bacillus subtilis functional genomics: Genome-widw analysis of the DegS-DegU regulon by transcriptomics and proteomics. Mol. Genet. Genom. 2002, 268, 455–467. [Google Scholar] [CrossRef] [PubMed]
- Hutchings, M.I.; Palmer, T.; Harrington, D.J.; Sutcliffe, I.C. Lipoprotein biogenesis in Gram-positive bacteria: Knowing when to hold 'em, knowing when to fold 'em. Trends Microbiol. 2009, 17, 13–21. [Google Scholar] [CrossRef] [PubMed]
- Thompson, B.J.; Widdick, D.A.; Hicks, M.G.; Chandra, G.; Sutcliffe, I.C.; Palmer, T.; Hutchings, M.I. Investigating lipoprotein biogenesis and function in the model Gram-positive bacterium Streptomyces coelicolor. Mol. Microbiol. 2010, 77, 943–957. [Google Scholar] [CrossRef] [PubMed]
- Widdick, D.A.; Hicks, M.G.; Thompson, B.J.; Tschumi, A.; Chandra, G.; Sutcliffe, I.C.; Brülle, J.K.; Sander, P.; Palmer, T.; Hutchings, M.I. Dissecting the complete lipoprotein biogenesis pathway in Streptomyces scabies. Mol. Microbiol. 2011, 80, 1395–1412. [Google Scholar] [CrossRef] [PubMed]
- Gullón, S.; Arranz, E.I.G.; Mellado, R.P. Transcriptional characterisation of the negative effect exerted by a deficiency in type II signal peptidase on extracelular protein secretion in Streptomyces lividans. Appl. Microbiol. Biotechnol. 2013, 97, 10069–10080. [Google Scholar] [CrossRef] [PubMed]
- Oliveros, J.C.; VENNY. An Interactive Tool for Comparing Lists with Venn’s Diagrams. 2007. Available online: http://bioinfogp.cnb.csic.es/tools/venny/index.html (accessed on 27 February 2018).
- Gullón, S.; Marín, S.; Mellado, R.P. Overproduction of a model Sec-and Tat-Dependent secretory protein elicits different cellular responses in Streptomyces lividans. PLoS ONE 2015, 10, e0133645. [Google Scholar] [CrossRef] [PubMed]
- Tullman-Ercek, D.; DeLisa, M.P.; Kawarasaki, Y.; Iranpour, P.; Ribnicky, B.; Palmer, T.; Georgiou, G. Export pathway selectivity of Escherichia coli twin arginine translocation signal peptides. J. Biol. Chem. 2007, 282, 8309–8319. [Google Scholar] [CrossRef] [PubMed]
- Tooke, F.J.; Babot, M.; Chandra, G.; Buchanan, G.; Palmer, T. A unifying mechanism for biogenesis of membrane proteins co-operatively integrated by the Sec and Tat pathways. eLife 2017, 6, e26577. [Google Scholar] [CrossRef] [PubMed]
- Cristóbal, S.; de Gier, J.W.; Nielsen, H.; von Heijne, G. Competition between Sec-and TAT-dependent protein translocation in Escherichia coli. EMBO J. 1999, 18, 2982–2990. [Google Scholar] [CrossRef] [PubMed]
- Hyyriläinen, H.L.; Bolhuis, A.; Darmon, E.; Muukkonen, L.; Koski, P.; Vitikainen, M.; Sarvas, M.; Prágai, Z.; Bron, S.; van Dijl, M.; et al. A novel two-component regulatory system in Bacillus subtilis for the survival of severe secretion stress. Mol. Microbiol. 2001, 41, 1159–1172. [Google Scholar] [CrossRef]
- Gullón, S.; Vicente, R.L.; Mellado, R.-P. A novel two-component system involved in secretion stress response in Streptomyces lividans. PLoS ONE 2012, 7, e48987. [Google Scholar] [CrossRef] [PubMed]
- Darmon, E.; Noone, D.; Masson, A.; Bron, S.; Kuipers, O.P.; Devine, K.M.; van Dijl, J.M. A novel class of heat and secretion stress-responsive genes is controlled by the autoregulated CssRS two-component system of Bacillus subtilis. J. Bacteriol. 2002, 184, 5661–5671. [Google Scholar] [CrossRef] [PubMed]
- Lulko, A.T.; Veening, J.W.; Buist, G.; Smits, W.K.; Blom, E.J.; Beekman, A.C.; Bron, S.; Kuipers, O.P. Production and secretion stress caused by overexpression of heterologous α-amylase leads to inhibition of sporulation and a prolonged motile phase in Bacillus subtilis. Appl. Environ. Microbiol. 2007, 73, 5354–5362. [Google Scholar] [CrossRef] [PubMed]
- Vicente, R.L.; Gullón, S.; Marín, S.; Mellado, R.P. The three Streptomyces lividans HtrA-like proteases involved in the secretion stress response act in a cooperative manner. PLoS ONE 2016, 11, e0168112. [Google Scholar] [CrossRef] [PubMed]
- Westers, H.; Westers, L.; Darmon, E.; van Dijl, J.M.; Quax, W.J.; Zanen, G. The CssRS two-component regulatory system controls a general secretion stress response in B. subtilis. FEBS J. 2006, 273, 3816–3827. [Google Scholar] [CrossRef] [PubMed]
- Noone, D.; Howell, A.; Collery, R.; Devine, K.M. YkdA and YvtA, HtrA-Like serine proteases in Bacillus subtilis, engage in negative autoregulation and reciprocal cross-regulation of ykdA and yvtA gene expression. J. Bacteriol. 2001, 183, 654–663. [Google Scholar] [CrossRef] [PubMed]
- Clausen, T.; Kaiser, M.; Huber, R.; Ehrmann, M. HTRA proteases: Regulated proteolysis in protein quality control. Nat. Rev. Mol. Cell Biol. 2011, 12, 152–162. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gullón, S.; Mellado, R.P. The Cellular Mechanisms that Ensure an Efficient Secretion in Streptomyces. Antibiotics 2018, 7, 33. https://doi.org/10.3390/antibiotics7020033
Gullón S, Mellado RP. The Cellular Mechanisms that Ensure an Efficient Secretion in Streptomyces. Antibiotics. 2018; 7(2):33. https://doi.org/10.3390/antibiotics7020033
Chicago/Turabian StyleGullón, Sonia, and Rafael P. Mellado. 2018. "The Cellular Mechanisms that Ensure an Efficient Secretion in Streptomyces" Antibiotics 7, no. 2: 33. https://doi.org/10.3390/antibiotics7020033
APA StyleGullón, S., & Mellado, R. P. (2018). The Cellular Mechanisms that Ensure an Efficient Secretion in Streptomyces. Antibiotics, 7(2), 33. https://doi.org/10.3390/antibiotics7020033



