Comparative Genomics of DH5α-Inhibiting Escherichia coli Isolates from Feces of Healthy Individuals Reveals Common Co-Occurrence of Bacteriocin Genes with Virulence Factors and Antibiotic Resistance Genes
Abstract
1. Introduction
2. Results
3. Discussion
4. Materials and Methods
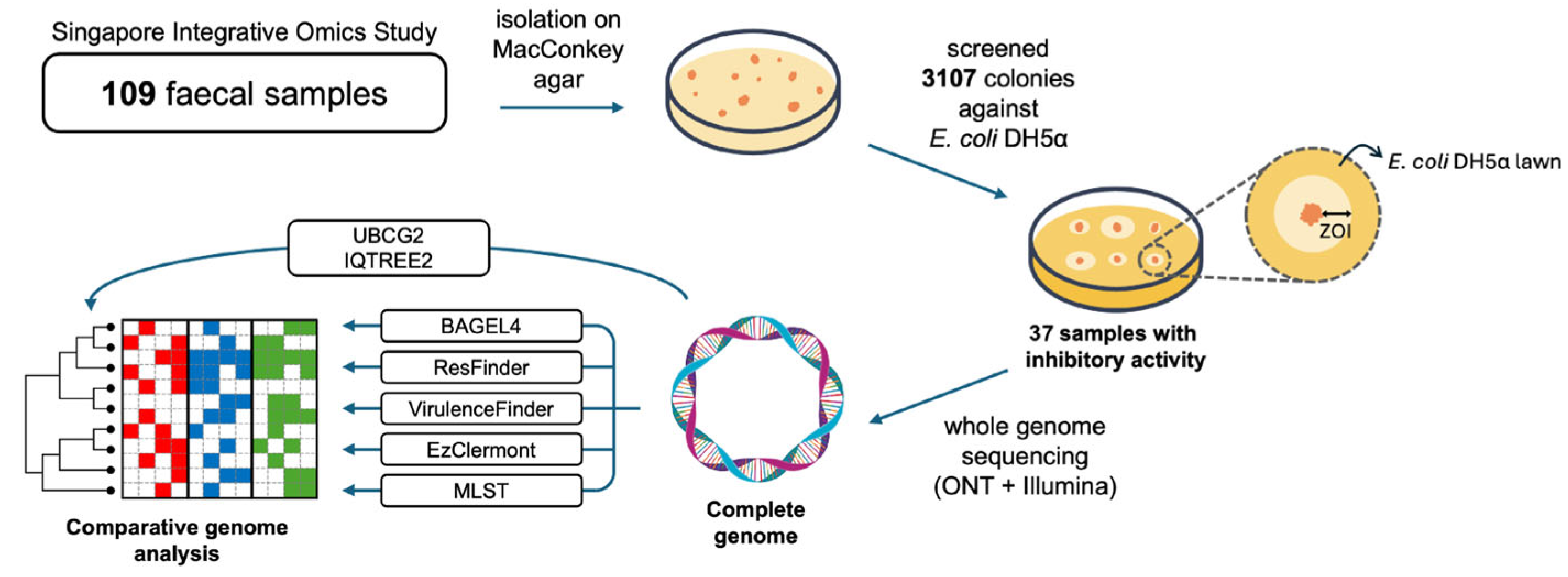
4.1. Isolation and Screening
4.2. DNA Extraction and Sequencing
4.3. Bioinformatics
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ding, Y.; Er, S.; Tan, A.; Gounot, J.-S.; Saw, W.-Y.; Tan, L.W.L.; Teo, Y.Y.; Nagarajan, N.; Seedorf, H. Comparison of tet(X4)-containing contigs assembled from metagenomic sequencing data with plasmid sequences of isolates from a cohort of healthy subjects. Microbiol. Spectr. 2024, 12, e0396923. [Google Scholar] [CrossRef]
- Ding, Y.; Saw, W.-Y.; Tan, L.W.L.; Moong, D.K.N.; Nagarajan, N.; Teo, Y.Y.; Seedorf, H. Emergence of tigecycline- and eravacycline-resistant Tet(X4)-producing Enterobacteriaceae in the gut microbiota of healthy Singaporeans. J. Antimicrob. Chemother. 2020, 75, 3480–3484. [Google Scholar] [CrossRef]
- Ding, Y.; Saw, W.-Y.; Tan, L.W.L.; Moong, D.K.N.; Nagarajan, N.; Teo, Y.Y.; Seedorf, H. Extended-Spectrum β-Lactamase-Producing and mcr-1-Positive Escherichia coli from the Gut Microbiota of Healthy Singaporeans. Appl. Environ. Microbiol. 2021, 87, e0048821. [Google Scholar] [CrossRef]
- Cotter, P.D.; Ross, R.P.; Hill, C. Bacteriocins—A viable alternative to antibiotics? Nat. Rev. Microbiol. 2013, 11, 95–105. [Google Scholar] [CrossRef]
- Heilbronner, S.; Krismer, B.; Brötz-Oesterhelt, H.; Peschel, A. The microbiome-shaping roles of bacteriocins. Nat. Rev. Microbiol. 2021, 19, 726–739. [Google Scholar] [CrossRef] [PubMed]
- Riley, M.A.; Wertz, J.E. Bacteriocins: Evolution, ecology, and application. Annu. Rev. Microbiol. 2002, 56, 117–137. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; Pan, Z.; Huang, J.; Sun, M.; Lu, C.; Yao, H. The Hcp proteins fused with diverse extended-toxin domains represent a novel pattern of antibacterial effectors in type VI secretion systems. Virulence 2017, 8, 1189–1202. [Google Scholar] [CrossRef]
- Magiorakos, A.P.; Srinivasan, A.; Carey, R.B.; Carmeli, Y.; Falagas, M.E.; Giske, C.G.; Harbarth, S.; Hindler, J.F.; Kahlmeter, G.; Olsson-Liljequist, B.; et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 2012, 18, 268–281. [Google Scholar] [CrossRef]
- Martínez, J.L.; Baquero, F.; Andersson, D.I. Predicting antibiotic resistance. Nat. Rev. Microbiol. 2007, 5, 958–965. [Google Scholar] [CrossRef] [PubMed]
- Johnson, J.R.; Murray, A.C.; Gajewski, A.; Sullivan, M.; Snippes, P.; Kuskowski, M.A.; Smith, K.E. Isolation and molecular characterization of nalidixic acid-resistant extraintestinal pathogenic Escherichia coli from retail chicken products. Antimicrob. Agents Chemother. 2003, 47, 2161–2168. [Google Scholar] [CrossRef]
- Chakraborty, A.; Saralaya, V.; Adhikari, P.; Shenoy, S.; Baliga, S.; Hegde, A. Characterization of Escherichia coli Phylogenetic Groups Associated with Extraintestinal Infections in South Indian Population. Ann. Med. Health Sci. Res. 2015, 5, 241–246. [Google Scholar] [CrossRef]
- Wick, R.R.; Judd, L.M.; Gorrie, C.L.; Holt, K.E. Unicycler: Resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput. Biol. 2017, 13, e1005595. [Google Scholar] [CrossRef]
- Carattoli, A.; Zankari, E.; García-Fernández, A.; Voldby Larsen, M.; Lund, O.; Villa, L.; Møller Aarestrup, F.; Hasman, H. In silico detection and typing of plasmids using PlasmidFinder and plasmid multilocus sequence typing. Antimicrob. Agents Chemother. 2014, 58, 3895–3903. [Google Scholar] [CrossRef]
- van Heel, A.J.; de Jong, A.; Song, C.; Viel, J.H.; Kok, J.; Kuipers, O.P. BAGEL4: A user-friendly web server to thoroughly mine RiPPs and bacteriocins. Nucleic Acids Res. 2018, 46, W278–W281. [Google Scholar] [CrossRef]
- Bortolaia, V.; Kaas, R.S.; Ruppe, E.; Roberts, M.C.; Schwarz, S.; Cattoir, V.; Philippon, A.; Allesoe, R.L.; Rebelo, A.R.; Florensa, A.F.; et al. ResFinder 4.0 for predictions of phenotypes from genotypes. J. Antimicrob. Chemother. 2020, 75, 3491–3500. [Google Scholar] [CrossRef]
- Joensen, K.G.; Scheutz, F.; Lund, O.; Hasman, H.; Kaas, R.S.; Nielsen, E.M.; Aarestrup, F.M. Real-Time Whole-Genome Sequencing for Routine Typing, Surveillance, and Outbreak Detection of Verotoxigenic Escherichia coli. J. Clin. Microbiol. 2014, 52, 1501–1510. [Google Scholar] [CrossRef] [PubMed]
- Tetzschner, A.M.M.; Johnson, J.R.; Johnston, B.D.; Lund, O.; Scheutz, F. In Silico Genotyping of Escherichia coli Isolates for Extraintestinal Virulence Genes by Use of Whole-Genome Sequencing Data. J. Clin. Microbiol. 2020, 58, e01269-20. [Google Scholar] [CrossRef]
- Larsen, M.V.; Cosentino, S.; Rasmussen, S.; Friis, C.; Hasman, H.; Marvig, R.L.; Jelsbak, L.; Sicheritz-Pontén, T.; Ussery, D.W.; Aarestrup, F.M.; et al. Multilocus Sequence Typing of Total-Genome-Sequenced Bacteria. J. Clin. Microbiol. 2012, 50, 1355–1361. [Google Scholar] [CrossRef]
- Waters, N.R.; Abram, F.; Brennan, F.; Holmes, A.; Pritchard, L. Easy phylotyping of Escherichia coli via the EzClermont web app and command-line tool. Access Microbiol. 2020, 2, e000143. [Google Scholar] [CrossRef] [PubMed]
- Hoang, D.T.; Chernomor, O.; von Haeseler, A.; Minh, B.Q.; Vinh, L.S. UFBoot2: Improving the Ultrafast Bootstrap Approximation. Mol. Biol. Evol. 2018, 35, 518–522. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Na, S.-I.; Kim, D.; Chun, J. UBCG2: Up-to-date bacterial core genes and pipeline for phylogenomic analysis. J. Microbiol. 2021, 59, 609–615. [Google Scholar] [CrossRef] [PubMed]
- Minh, B.Q.; Schmidt, H.A.; Chernomor, O.; Schrempf, D.; Woodhams, M.D.; von Haeseler, A.; Lanfear, R. IQ-TREE 2: New Models and Efficient Methods for Phylogenetic Inference in the Genomic Era. Mol. Biol. Evol. 2020, 37, 1530–1534. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Er, S.; Ding, Y.; Tan, L.W.L.; Teo, Y.Y.; Nagarajan, N.; Seedorf, H. Comparative Genomics of DH5α-Inhibiting Escherichia coli Isolates from Feces of Healthy Individuals Reveals Common Co-Occurrence of Bacteriocin Genes with Virulence Factors and Antibiotic Resistance Genes. Antibiotics 2025, 14, 860. https://doi.org/10.3390/antibiotics14090860
Er S, Ding Y, Tan LWL, Teo YY, Nagarajan N, Seedorf H. Comparative Genomics of DH5α-Inhibiting Escherichia coli Isolates from Feces of Healthy Individuals Reveals Common Co-Occurrence of Bacteriocin Genes with Virulence Factors and Antibiotic Resistance Genes. Antibiotics. 2025; 14(9):860. https://doi.org/10.3390/antibiotics14090860
Chicago/Turabian StyleEr, Shuan, Yichen Ding, Linda Wei Lin Tan, Yik Ying Teo, Niranjan Nagarajan, and Henning Seedorf. 2025. "Comparative Genomics of DH5α-Inhibiting Escherichia coli Isolates from Feces of Healthy Individuals Reveals Common Co-Occurrence of Bacteriocin Genes with Virulence Factors and Antibiotic Resistance Genes" Antibiotics 14, no. 9: 860. https://doi.org/10.3390/antibiotics14090860
APA StyleEr, S., Ding, Y., Tan, L. W. L., Teo, Y. Y., Nagarajan, N., & Seedorf, H. (2025). Comparative Genomics of DH5α-Inhibiting Escherichia coli Isolates from Feces of Healthy Individuals Reveals Common Co-Occurrence of Bacteriocin Genes with Virulence Factors and Antibiotic Resistance Genes. Antibiotics, 14(9), 860. https://doi.org/10.3390/antibiotics14090860






