MDR Pumps as Crossroads of Resistance: Antibiotics and Bacteriophages
Abstract
:1. Time of Global Crisis: Antibiotic Resistance
2. Antibiotic Resistance: The Role of MDR Pumps
3. Alternative Resistance Mechanisms Using MDR Pumps
4. MDR Pumps in Bacteria
5. Role of MDR Pumps in Biofilm Protection and Bacterial Persistence
6. Bacteriophage as a Natural Antibacterial Weapon
7. Mechanisms of Phage Penetration into a Bacterial Cell
8. MDR Pumps as Phage Receptors
9. Conclusions
Funding
Acknowledgments
Conflicts of Interest
References
- WHO. Rolling Updates on Coronavirus Disease (COVID-19). Available online: https://www.who.int/emergencies/diseases/novel-coronavirus-2019/events-as-they-happen (accessed on 22 April 2021).
- Haque, A.; Pant, A.B. Mitigating Covid-19 in the face of emerging virus variants, breakthrough infections and vaccine hesitancy. J. Autoimmun. 2022, 127, 102792. [Google Scholar] [CrossRef] [PubMed]
- Yong, D.; Toleman, M.A.; Giske, C.G.; Cho, H.S.; Sundman, K.; Lee, K.; Walsh, T.R. Characterization of a New Metallo-β-Lactamase Gene, bla NDM-1, and a Novel Erythromycin Esterase Gene Carried on a Unique Genetic Structure in Klebsiella pneumoniae Sequence Type 14 from India. Antimicrob. Agents Chemother. 2009, 53, 5046–5054. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McCann, C.M.; Christgen, B.; Roberts, J.A.; Su, J.-Q.; Arnold, K.E.; Gray, N.; Zhu, Y.-G.; Graham, D.W. Understanding drivers of antibiotic resistance genes in High Arctic soil ecosystems. Environ. Int. 2019, 125, 497–504. [Google Scholar] [CrossRef]
- Liu, Y.-Y.; Wang, Y.; Walsh, T.R.; Yi, L.-X.; Zhang, R.; Spencer, J.; Doi, Y.; Tian, G.; Dong, B.; Huang, X.; et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: A microbiological and molecular biological study. Lancet Infect. Dis. 2016, 16, 161–168. [Google Scholar] [CrossRef]
- Wang, R.; van Dorp, L.; Shaw, L.P.; Bradley, P.; Wang, Q.; Wang, X.; Jin, L.; Zhang, Q.; Liu, Y.; Rieux, A.; et al. The global distribution and spread of the mobilized colistin resistance gene mcr-1. Nat. Commun. 2018, 9, 1179. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Plackett, B. Why big pharma has abandoned antibiotics. Nature 2020, 586, S50–S52. [Google Scholar] [CrossRef]
- Randall, J.R.; Davies, B.W. Mining for novel antibiotics. Curr. Opin. Microbiol. 2021, 63, 66–69. [Google Scholar] [CrossRef] [PubMed]
- Theuretzbacher, U.; Gottwalt, S.; Beyer, P.; Butler, M.; Czaplewski, L.; Lienhardt, C.; Moja, L.; Paul, M.; Paulin, S.; Rex, J.; et al. Analysis of the clinical antibacterial and antituberculosis pipeline. Lancet Infect. Dis. 2018, 19, e40–e50. [Google Scholar] [CrossRef]
- Nazarov, P.A. Alternatives to antibiotics: Phage lytic enzymes and phage therapy. Bull. RSMU 2018, 1, 5–15. [Google Scholar] [CrossRef]
- Baltz, R.H. Antimicrobials from Actinomycetes: Back to the Future. Microbe 2007, 2, 125–131. [Google Scholar]
- Jones, D. Tuberculosis success. Nat. Rev. Drug Discov. 2013, 12, 175–176. [Google Scholar] [CrossRef]
- Khailova, L.S.; Nazarov, P.A.; Sumbatyan, N.V.; Korshunova, G.A.; Rokitskaya, T.I.; Dedukhova, V.I.; Antonenko, Y.N.; Sku-lachev, V.P. Uncoupling and toxic action of alkyltriphenylphosphonium cations on mitochondria and the bacterium Bacillus subtilis as a function of alkyl chain length. Biochemistry 2015, 80, 1589–1597. [Google Scholar] [CrossRef]
- Nazarov, P.A.; Osterman, I.; Tokarchuk, A.; Karakozova, M.V.; Korshunova, G.A.; Lyamzaev, K.; Skulachev, M.V.; Kotova, E.A.; Skulachev, V.; Antonenko, Y.N. Mitochondria-targeted antioxidants as highly effective antibiotics. Sci. Rep. 2017, 7, 1394. [Google Scholar] [CrossRef]
- Nazarov, P.A.; Kotova, E.A.; Skulachev, V.; Antonenko, Y.N. Genetic Variability of the AcrAB-TolC Multidrug Efflux Pump Underlies SkQ1 Resistance in Gram-Negative Bacteria. Acta Nat. 2019, 11, 93–98. [Google Scholar] [CrossRef]
- Pavlova, J.; Khairullina, Z.; Tereshchenkov, A.; Nazarov, P.; Lukianov, D.; Volynkina, I.; Skvortsov, D.; Makarov, G.; Abad, E.; Murayama, S.; et al. Triphenilphosphonium Analogs of Chloramphenicol as Dual-Acting Antimicrobial and Antiproliferating Agents. Antibiotics 2021, 10, 489. [Google Scholar] [CrossRef]
- Singh, K.S.; Sharma, R.; Reddy, P.A.N.; Vonteddu, P.; Good, M.; Sundarrajan, A.; Choi, H.; Muthumani, K.; Kossenkov, A.; Goldman, A.R.; et al. RETRACTED ARTICLE: IspH inhibitors kill Gram-negative bacteria and mobilize immune clearance. Nature 2020, 589, 597–602. [Google Scholar] [CrossRef]
- Pisárčik, M.; Lukáč, M.; Jampílek, J.; Bilka, F.; Bilková, A.; Pašková, Ľ.; Devínsky, F.; Horáková, R.; Opravil, T. Silver nano-particles stabilised with cationic single-chain surfactants. Structure-physical properties-biological activity re-lationship study. J. Mol. Liq. 2018, 272, 60–72. [Google Scholar] [CrossRef]
- Ling, L.L.; Schneider, T.; Peoples, A.J.; Spoering, A.L.; Engels, I.; Conlon, B.P.; Mueller, A.; Schäberle, T.F.; Hughes, D.E.; Epstein, S.; et al. A new antibiotic kills pathogens without detectable resistance. Nature 2015, 517, 455–459. [Google Scholar] [CrossRef]
- Imai, Y.; Meyer, K.J.; Iinishi, A.; Favre-Godal, Q.; Green, R.; Manuse, S.; Caboni, M.; Mori, M.; Niles, S.; Ghiglieri, M.; et al. A new antibiotic selectively kills Gram-negative pathogens. Nature 2019, 576, 459–464. [Google Scholar] [CrossRef]
- Zipperer, A.; Konnerth, M.C.; Laux, C.; Berscheid, A.; Janek, D.; Weidenmaier, C.; Burian, M.; Schilling, N.A.; Slavetinsky, C.; Marschal, M.; et al. Human commensals producing a novel antibiotic impair pathogen colonization. Nature 2016, 535, 511–516. [Google Scholar] [CrossRef]
- Lewis, K. The Science of Antibiotic Discovery. Cell 2020, 181, 29–45. [Google Scholar] [CrossRef]
- Mendoza, N.; Ravanfar, P.; Satyaprakah, A.; Pillai, S.; Creed, R. Existing antibacterial vaccines. Dermatol. Ther. 2009, 22, 129–142. [Google Scholar] [CrossRef]
- Zurawski, D.V.; McLendon, M.K. Monoclonal Antibodies as an Antibacterial Approach against Bacterial Pathogens. Antibiotics 2020, 9, 155. [Google Scholar] [CrossRef] [Green Version]
- Silva, D.R.; Sardi, J.D.C.O.; Pitangui, N.D.S.; Roque, S.M.; da Silva, A.C.B.; Rosalen, P.L. Probiotics as an alternative antimicrobial therapy: Current reality and future directions. J. Funct. Foods 2020, 73, 104080. [Google Scholar] [CrossRef]
- Hang, B.T.B.; Phuong, N.T.; Kestemont, P. Can immunostimulants efficiently replace antibiotic in striped catfish (Pangasianodon hypophthalmus) against bacterial infection by Edwardsiella ictaluri? Fish Shellfish Immunol. 2014, 40, 556–562. [Google Scholar] [CrossRef]
- Hamblin, M.R. Antimicrobial photodynamic inactivation: A bright new technique to kill resistant microbes. Curr. Opin. Microbiol. 2016, 33, 67–73. [Google Scholar] [CrossRef] [Green Version]
- Omarova, E.O.; Nazarov, P.A.; Firsov, A.M.; Strakhovskaya, M.G.; Arkhipova, A.Y.; Moisenovich, M.M.; Agapov, I.I.; Ol’Shevskaya, V.A.; Zaitsev, A.; Kalinin, V.N.; et al. Carboranyl-Chlorin e6 as a Potent Antimicrobial Photosensitizer. PLoS ONE 2015, 10, e0141990. [Google Scholar] [CrossRef]
- Krylov, V.; Bourkaltseva, M.; Pleteneva, E. Bacteriophage’s Dualism in Therapy. Trends Microbiol. 2019, 27, 566–567. [Google Scholar] [CrossRef]
- Danis-Wlodarczyk, K.M.; Wozniak, D.J.; Abedon, S.T. Treating Bacterial Infections with Bacteriophage-Based Enzybiotics: In Vitro, In Vivo and Clinical Application. Antibiotics 2021, 10, 1497. [Google Scholar] [CrossRef]
- Miroshnikov, K.A.; Chertkov, O.V.; Nazarov, P.A.; Mesyanzhinov, V.V. Peptido-glikanliziruyushchie fermenty bakteriofagov -perspektivnye protivobakterial’nye agenty. Uspekhi Biol. Khimii 2006, 46, 65–98. (In Russian) [Google Scholar]
- Dunne, M.; Rupf, B.; Tala, M.; Qabrati, X.; Ernst, P.; Shen, Y.; Sumrall, E.; Heeb, L.; Plückthun, A.; Loessner, M.J.; et al. Reprogramming Bacteriophage Host Range through Structure-Guided Design of Chimeric Receptor Binding Proteins. Cell Rep. 2019, 29, 1336–1350.e4. [Google Scholar] [CrossRef] [Green Version]
- Huan, Y.; Kong, Q.; Mou, H.; Yi, H. Antimicrobial Peptides: Classification, Design, Application and Research Progress in Multiple Fields. Front. Microbiol. 2020, 11, 582779. [Google Scholar] [CrossRef]
- Diamond, G.; Beckloff, N.; Weinberg, A.; Kisich, K. The Roles of Antimicrobial Peptides in Innate Host Defense. Curr. Pharm. Des. 2009, 15, 2377–2392. [Google Scholar] [CrossRef] [Green Version]
- Mishra, R.; Panda, A.K.; De Mandal, S.; Shakeel, M.; Bisht, S.S.; Khan, J. Natural Anti-biofilm Agents: Strategies to Control Biofilm-Forming Pathogens. Front. Microbiol. 2020, 11, 566325. [Google Scholar] [CrossRef]
- Lamut, A.; Mašič, L.P.; Kikelj, D.; Tomašič, T. Efflux pump inhibitors of clinically relevant multidrug resistant bacteria. Med. Res. Rev. 2019, 39, 2460–2504. [Google Scholar] [CrossRef]
- Ebbensgaard, A.E.; Løbner-Olesen, A.; Frimodt-Møller, J. The Role of Efflux Pumps in the Transition from Low-Level to Clinical Antibiotic Resistance. Antibiotics 2020, 9, 855. [Google Scholar] [CrossRef]
- Berdaguer, S.; Bautista, J.; Brunet, M.; Cisneros, J.M. Antimicrobial and immunosuppressive drug interactions in solid organ transplant recipients. Enferm. Infecc. Microbiol. Clin. 2012, 30, 86–92. [Google Scholar] [CrossRef]
- Rao, L.; Tian, R.; Chen, X. Cell-Membrane-Mimicking Nanodecoys against Infectious Diseases. ACS Nano 2020, 14, 2569–2574. [Google Scholar] [CrossRef] [Green Version]
- Santos, A.L.; Sodre, C.L.; Valle, R.S.; Silva, B.A.; Abi-Chacra, E.A.; Silva, L.V.; Souza-Goncalves, A.L.; Sangenito, L.S.; Goncalves, D.S.; Souza, L.O.; et al. Antimicrobial Action of Chelating Agents: Repercussions on the Microorganism Development, Virulence and Pathogenesis. Curr. Med. Chem. 2012, 19, 2715–2737. [Google Scholar] [CrossRef]
- Coraça-Huber, D.C.; Dichtl, S.; Steixner, S.; Nogler, M.; Weiss, G. Iron chelation destabilizes bacterial biofilms and potentiates the antimicrobial activity of antibiotics against coagulase-negative Staphylococci. Pathog. Dis. 2018, 76, fty052. [Google Scholar] [CrossRef]
- Qiu, D.-H.; Huang, Z.-L.; Zhou, T.; Shen, C.; Hider, R. In vitro inhibition of bacterial growth by iron chelators. FEMS Microbiol. Lett. 2010, 314, 107–111. [Google Scholar] [CrossRef]
- Afrasiabi, S.; Pourhajibagher, M.; Raoofian, R.; Tabarzad, M.; Bahador, A. Therapeutic applications of nucleic acid aptamers in microbial infections. J. Biomed. Sci. 2020, 27, 6–13. [Google Scholar] [CrossRef]
- Wojciechowska, M.; Równicki, M.; Mieczkowski, A.; Miszkiewicz, J.; Trylska, J. Antibacterial Peptide Nucleic Acids—Facts and Perspectives. Molecules 2020, 25, 559. [Google Scholar] [CrossRef] [Green Version]
- Lee, H.T.; Kim, S.K.; Yoon, J.W. Antisense peptide nucleic acids as a potential anti-infective agent. J. Microbiol. 2019, 57, 423–430. [Google Scholar] [CrossRef]
- Basit, A.; Qadir, S.; Qureshi, S.; Rehman, S.U. Cloning and expression analysis of fused holin-endolysin from RL bacteriophage; Exhibits broad activity against multi drug resistant pathogens. Enzym. Microb. Technol. 2021, 149, 109846. [Google Scholar] [CrossRef]
- Aslam, B.; Arshad, M.I.; Aslam, M.A.; Muzammil, S.; Siddique, A.B.; Yasmeen, N.; Khurshid, M.; Rasool, M.; Ahmad, M.; Rasool, M.H.; et al. Bacteriophage Proteome: Insights and Potentials of an Alternate to Antibiotics. Infect. Dis. Ther. 2021, 10, 1171–1193. [Google Scholar] [CrossRef]
- Grabowski, Ł.; Łepek, K.; Stasiłojć, M.; Kosznik-Kwaśnicka, K.; Zdrojewska, K.; Maciąg-Dorszyńska, M.; Węgrzyn, G.; Węgrzyn, A. Bacteriophage-encoded enzymes destroying bacterial cell membranes and walls, and their potential use as antimicrobial agents. Microbiol. Res. 2021, 248, 126746. [Google Scholar] [CrossRef]
- Sugden, R.; Kelly, R.; Davies, S. Combatting antimicrobial resistance globally. Nat. Microbiol. 2016, 1, 16187. [Google Scholar] [CrossRef]
- Reygaert, W.C. An overview of the antimicrobial resistance mechanisms of bacteria. AIMS Microbiol. 2018, 4, 482–501. [Google Scholar] [CrossRef]
- Anisimov, V.N.; Egorov, M.V.; Krasilshchikova, M.S.; Lyamzaev, K.G.; Manskikh, V.N.; Moshkin, M.P.; Novikov, E.A.; Popovich, I.G.; Rogovin, K.A.; Shabalina, I.G.; et al. Effects of the mitochondria-targeted antioxidant SkQ1 on lifespan of rodents. Aging 2011, 3, 1110–1119. [Google Scholar] [CrossRef] [Green Version]
- Nazarov, P.A.; Sorochkina, A.I.; Karakozova, M.V. New Functional Criterion for Evaluation of Homologous MDR Pumps. Front. Microbiol. 2020, 11, 592283. [Google Scholar] [CrossRef]
- Sulavik, M.C.; Houseweart, C.; Cramer, C.; Jiwani, N.; Murgolo, N.; Greene, J.; DiDomenico, B.; Shaw, K.J.; Miller, G.H.; Hare, R.; et al. Antibiotic Susceptibility Profiles of Escherichia coli Strains Lacking Multidrug Efflux Pump Genes. Antimicrob. Agents Chemother. 2001, 45, 1126–1136. [Google Scholar] [CrossRef] [Green Version]
- Jo, J.T.H.; Brinkman, F.S.L.; Hancock, R.E.W. Aminoglycoside Efflux in Pseudomonas aeruginosa: Involvement of Novel Outer Membrane Proteins. Antimicrob. Agents Chemother. 2003, 47, 1101–1111. [Google Scholar] [CrossRef] [Green Version]
- Rosenberg, E.Y.; Ma, D.; Nikaido, H. AcrD of Escherichia coli Is an Aminoglycoside Efflux Pump. J. Bacteriol. 2000, 182, 1754–1756. [Google Scholar] [CrossRef] [Green Version]
- Biot, F.V.; Lopez, M.M.; Poyot, T.; Neulat-Ripoll, F.; Lignon, S.; Caclard, A.; Thibault, F.M.; Peinnequin, A.; Pagès, J.-M.; Valade, E. Interplay between Three RND Efflux Pumps in Doxycycline-Selected Strains of Burkholderia thailandensis. PLoS ONE 2013, 8, e84068. [Google Scholar] [CrossRef]
- Jung, Y.-H.; Shin, E.S.; Kim, O.; Yoo, J.S.; Lee, K.M.; Yoo, J.I.; Chung, G.T.; Lee, Y.S. Characterization of Two Newly Identified Genes, vgaD and vatG, Conferring Resistance to Streptogramin A in Enterococcus faecium. Antimicrob. Agents Chemother. 2010, 54, 4744–4749. [Google Scholar] [CrossRef] [Green Version]
- Hirata, K.; Suzuki, H.; Nishizawa, T.; Tsugawa, H.; Muraoka, H.; Saito, Y.; Matsuzaki, J.; Hibi, T. Contribution of efflux pumps to clarithromycin resistance in Helicobacter pylori. J. Gastroenterol. Hepatol. 2010, 25, S75–S79. [Google Scholar] [CrossRef]
- Bosnar, M.; Kelnerić, Z.; Munić, V.; Eraković, V.; Parnham, M.J. Cellular Uptake and Efflux of Azithromycin, Erythromycin, Clarithromycin, Telithromycin, and Cethromycin. Antimicrob. Agents Chemother. 2005, 49, 2372–2377. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.; Zhang, Y.; Liu, S.; Huang, N.; Zeng, W.; Xu, W.; Zhou, T.; Shen, M. Comparison of Anti-Microbic and Anti-Biofilm Activity among Tedizolid and Radezolid against Linezolid-Resistant Enterococcus faecalis Isolates. Infect. Drug Resist. 2021, 14, 4619–4627. [Google Scholar] [CrossRef]
- Kim, H.J.; Kim, Y.; Lee, M.S.; Lee, H.S. Gene lmrB of Corynebacterium glutamicum confers efflux-mediated resistance to linco-mycin. Mol. Cells 2001, 12, 112–116. [Google Scholar]
- Lüthje, P.; Schwarz, S. Molecular analysis of constitutively expressed erm(C) genes selected in vitro in the presence of the non-inducers pirlimycin, spiramycin and tylosin. J. Antimicrob. Chemother. 2006, 59, 97–101. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Steward, C.D.; Raney, P.M.; Morrell, A.K.; Williams, P.P.; McDougal, L.K.; Jevitt, L.; McGowan, J.E.; Tenover, F.C. Testing for Induction of Clindamycin Resistance in Erythromycin-Resistant Isolates of Staphylococcus aureus. J. Clin. Microbiol. 2005, 43, 1716–1721. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sánchez, M.B.; Martínez, J.L. The Efflux Pump SmeDEF Contributes to Trimethoprim-Sulfamethoxazole Resistance in Stenotrophomonas maltophilia. Antimicrob. Agents Chemother. 2015, 59, 4347–4348. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Popella, P.; Krauss, S.; Ebner, P.; Nega, M.; Deibert, J.; Götz, F. VraH Is the Third Component of the Staphylococcus aureus VraDEH System Involved in Gallidermin and Daptomycin Resistance and Pathogenicity. Antimicrob. Agents Chemother. 2016, 60, 2391–2401. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Amiri, G.; Abbasi Shaye, M.; Bahreini, M.; Mafinezhad, A.; Ghazvini, K.; Sharifmoghadam, M.R. Determination of imipenem efflux-mediated resistance in Acinetobacter spp., using an efflux pump inhibitor. Iran. J. Microbiol. 2019, 11, 368–372. [Google Scholar] [CrossRef] [PubMed]
- Mateus, C.; Nunes, A.; Oleastro, M.; Domingues, F.; Ferreira, S. RND Efflux Systems Contribute to Resistance and Virulence of Aliarcobacter butzleri. Antibiotics 2021, 10, 823. [Google Scholar] [CrossRef] [PubMed]
- Jorth, P.; McLean, K.; Ratjen, A.; Secor, P.R.; Bautista, G.E.; Ravishankar, S.; Rezayat, A.; Garudathri, J.; Harrison, J.J.; Harwood, R.A.; et al. Evolved Aztreonam Resistance Is Multifactorial and Can Produce Hypervirulence in Pseudomonas aeruginosa. mBio 2017, 8, e00517-17. [Google Scholar] [CrossRef] [Green Version]
- Dinesh, N.; Sharma, S.; Balganesh, M. Involvement of Efflux Pumps in the Resistance to Peptidoglycan Synthesis Inhibitors in Mycobacterium tuberculosis. Antimicrob. Agents Chemother. 2013, 57, 1941–1943. [Google Scholar] [CrossRef] [Green Version]
- Elkins, C.A.; Nikaido, H. 3D structure of AcrB: The archetypal multidrug efflux transporter of Escherichia coli likely captures substrates from periplasm. Drug Resist. Updates 2003, 6, 9–13. [Google Scholar] [CrossRef]
- Nichols, R.J.; Sen, S.; Choo, Y.J.; Beltrao, P.; Zietek, M.; Chaba, R.; Lee, S.; Kazmierczak, K.M.; Lee, K.J.; Wong, A.; et al. Phenotypic Landscape of a Bacterial Cell. Cell 2011, 144, 143–156. [Google Scholar] [CrossRef] [Green Version]
- Galkina, K.V.; Besedina, E.; Zinovkin, R.; Severin, F.F.; Knorre, D.A. Penetrating cations induce pleiotropic drug resistance in yeast. Sci. Rep. 2018, 8, 8131. [Google Scholar] [CrossRef]
- Lee, A.; Mao, W.; Warren, M.S.; Mistry, A.; Hoshino, K.; Okumura, R.; Ishida, H.; Lomovskaya, O. Interplay between Efflux Pumps May Provide Either Additive or Multiplicative Effects on Drug Resistance. J. Bacteriol. 2000, 182, 3142–3150. [Google Scholar] [CrossRef] [Green Version]
- Tal, N.; Schuldiner, S. A coordinated network of transporters with overlapping specificities provides a robust survival strategy. Proc. Natl. Acad. Sci. USA 2009, 106, 9051–9056. [Google Scholar] [CrossRef] [Green Version]
- Somprasong, N.; Hall, C.M.; Webb, J.R.; Sahl, J.W.; Wagner, D.M.; Keim, P.; Currie, B.J.; Schweizer, H.P. Burkholderia ubonensis High-Level Tetracycline Resistance Is Due to Efflux Pump Synergy Involving a Novel TetA(64) Resistance Determinant. Antimicrob. Agents Chemother. 2021, 65, e01767-20. [Google Scholar] [CrossRef]
- Bergmiller, T.; Andersson, A.M.C.; Tomasek, K.; Balleza, E.; Kiviet, D.J.; Hauschild, R.; Tkačik, G.; Guet, C.C. Biased partitioning of the multidrug efflux pump AcrAB-TolC underlies long-lived phenotypic heterogeneity. Science 2017, 356, 311–315. [Google Scholar] [CrossRef]
- Snoussi, M.; Talledo, J.P.; Del Rosario, N.-A.; Mohammadi, S.; Ha, B.-Y.; Košmrlj, A.; Taheri-Araghi, S. Heterogeneous absorption of antimicrobial peptide LL37 in Escherichia coli cells enhances population survivability. eLife 2018, 7, 311–315. [Google Scholar] [CrossRef]
- Wu, F.; Tan, C. Dead bacterial absorption of antimicrobial peptides underlies collective tolerance. J. R. Soc. Interface 2019, 16, 20180701. [Google Scholar] [CrossRef] [Green Version]
- Bhattacharyya, S.; Walker, D.M.; Harshey, R.M. Dead cells release a ‘necrosignal’ that activates antibiotic survival pathways in bacterial swarms. Nat. Commun. 2020, 11, 4157. [Google Scholar] [CrossRef]
- Björkholm, B.; Sjölund, M.; Falk, P.G.; Berg, O.G.; Engstrand, L.; Andersson, D.I. Mutation frequency and biological cost of antibiotic resistance in Helicobacter pylori. Proc. Natl. Acad. Sci. USA 2001, 98, 14607–14612. [Google Scholar] [CrossRef] [Green Version]
- Long, H.; Miller, S.F.; Strauss, C.; Zhao, C.; Cheng, L.; Ye, Z.; Griffin, K.; Te, R.; Lee, H.; Chen, C.-C.; et al. Antibiotic treatment enhances the genome-wide mutation rate of target cells. Proc. Natl. Acad. Sci. USA 2016, 113, E2498–E2505. [Google Scholar] [CrossRef] [Green Version]
- El Meouche, I.; Dunlop, M.J. Heterogeneity in efflux pump expression predisposes antibiotic-resistant cells to mutation. Science 2018, 362, 686–690. [Google Scholar] [CrossRef] [Green Version]
- Reams, D.; Roth, J.R. Mechanisms of Gene Duplication and Amplification. Cold Spring Harb. Perspect. Biol. 2015, 7, a016592. [Google Scholar] [CrossRef] [Green Version]
- Du, D.; Wang-Kan, X.; Neuberger, A.; Van Veen, H.W.; Pos, K.M.; Piddock, L.J.V.; Luisi, B.F. Multidrug efflux pumps: Structure, function and regulation. Nat. Rev. Microbiol. 2018, 16, 523–539. [Google Scholar] [CrossRef]
- Nikaido, H. RND transporters in the living world. Res. Microbiol. 2018, 169, 363–371. [Google Scholar] [CrossRef]
- Pasqua, M.; Grossi, M.; Zennaro, A.; Fanelli, G.; Micheli, G.; Barras, F.; Colonna, B.; Prosseda, G. The Varied Role of Efflux Pumps of the MFS Family in the Interplay of Bacteria with Animal and Plant Cells. Microorganisms 2019, 7, 285. [Google Scholar] [CrossRef] [Green Version]
- Hassan, K.A.; Liu, Q.; Elbourne, L.; Ahmad, I.; Sharples, D.; Naidu, V.; Chan, C.L.; Li, L.; Harborne, S.; Pokhrel, A.; et al. Pacing across the membrane: The novel PACE family of efflux pumps is widespread in Gram-negative pathogens. Res. Microbiol. 2018, 169, 450–454. [Google Scholar] [CrossRef]
- Bot, C.T.; Prodan, C. Quantifying the membrane potential during E. coli growth stages. Biophys. Chem. 2010, 146, 133–137. [Google Scholar] [CrossRef]
- Zorova, L.D.; Popkov, V.A.; Plotnikov, E.Y.; Silachev, D.N.; Pevzner, I.B.; Jankauskas, S.S.; Babenko, V.A.; Zorov, S.D.; Balakireva, A.V.; Juhaszova, M.; et al. Mitochondrial membrane potential. Anal. Biochem. 2018, 552, 50–59. [Google Scholar] [CrossRef]
- Silhavy, T.J.; Kahne, D.; Walker, S. The Bacterial Cell Envelope. Cold Spring Harb. Perspect. Biol. 2010, 2, a000414. [Google Scholar] [CrossRef]
- Rybenkov, V.V.; Zgurskaya, H.I.; Ganguly, C.; Leus, I.V.; Zhang, Z.; Moniruzzaman, M. The Whole Is Bigger than the Sum of Its Parts: Drug Transport in the Context of Two Membranes with Active Efflux. Chem. Rev. 2021, 121, 5597–5631. [Google Scholar] [CrossRef]
- Verma, P.; Tiwari, M.; Tiwari, V. Efflux pumps in multidrug-resistant Acinetobacter baumannii: Current status and challenges in the discovery of efflux pumps inhibitors. Microb. Pathog. 2021, 152, 104766. [Google Scholar] [CrossRef] [PubMed]
- Stavri, M.; Piddock, L.J.V.; Gibbons, S. Bacterial efflux pump inhibitors from natural sources. J. Antimicrob. Chemother. 2006, 59, 1247–1260. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lomovskaya, O.; Warren, M.S.; Lee, A.; Galazzo, J.; Fronko, R.; Lee, M.; Blais, J.; Cho, D.; Chamberland, S.; Renau, T.; et al. Identification and Characterization of Inhibitors of Multidrug Resistance Efflux Pumps in Pseudomonas aeruginosa: Novel Agents for Combination Therapy. Antimicrob. Agents Chemother. 2001, 45, 105–116. [Google Scholar] [CrossRef] [Green Version]
- Lamers, R.P.; Cavallari, J.F.; Burrows, L.L. The Efflux Inhibitor Phenylalanine-Arginine Beta-Naphthylamide (PAβN) Permeabilizes the Outer Membrane of Gram-Negative Bacteria. PLoS ONE 2013, 8, e60666. [Google Scholar] [CrossRef] [Green Version]
- Radchenko, M.; Symerský, J.; Nie, R.; Lu, M. Structural basis for the blockade of MATE multidrug efflux pumps. Nat. Commun. 2015, 6, 7995. [Google Scholar] [CrossRef] [Green Version]
- Mukherjee, S.; Bassler, B.L. Bacterial quorum sensing in complex and dynamically changing environments. Nat. Rev. Microbiol. 2019, 17, 371–382. [Google Scholar] [CrossRef]
- Humphries, J.; Xiong, L.; Liu, J.; Prindle, A.; Yuan, F.; Arjes, H.A.; Tsimring, L.; Süel, G.M. Species-Independent Attraction to Biofilms through Electrical Signaling. Cell 2017, 168, 200–209.e12. [Google Scholar] [CrossRef] [Green Version]
- Wu, H.; Moser, C.; Wang, H.-Z.; Høiby, N.; Song, Z.-J. Strategies for combating bacterial biofilm infections. Int. J. Oral Sci. 2014, 7, 1–7. [Google Scholar] [CrossRef] [Green Version]
- Alav, I.; Sutton, J.M.; Rahman, K.M. Role of bacterial efflux pumps in biofilm formation. J. Antimicrob. Chemother. 2018, 73, 2003–2020. [Google Scholar] [CrossRef] [Green Version]
- Flemming, H.-C.; Wingender, J.; Szewzyk, U.; Steinberg, P.; Rice, S.A.; Kjelleberg, S. Biofilms: An emergent form of bacterial life. Nat. Rev. Microbiol. 2016, 14, 563–575. [Google Scholar] [CrossRef]
- Arroyo-Olarte, R.; Rodríguez, R.B.; Morales-Ríos, E. Genome Editing in Bacteria: CRISPR-Cas and Beyond. Microorganisms 2021, 9, 844. [Google Scholar] [CrossRef]
- Loureiro, A.; da Silva, G.J. CRISPR-Cas: Converting A Bacterial Defence Mechanism into A State-of-the-Art Genetic Manipulation Tool. Antibiotics 2019, 8, 18. [Google Scholar] [CrossRef] [Green Version]
- Millman, A.; Bernheim, A.; Stokar-Avihail, A.; Fedorenko, T.; Voichek, M.; Leavitt, A.; Oppenheimer-Shaanan, Y.; Sorek, R. Bacterial Retrons Function in Anti-Phage Defense. Cell 2020, 183, 1551–1561.e12. [Google Scholar] [CrossRef]
- Wang, Z.; Gong, X.; Xie, J.; Xu, Z.; Liu, G.; Zhang, G. Investigation of Formation of Bacterial Biofilm upon Dead Siblings. Langmuir 2018, 35, 7405–7413. [Google Scholar] [CrossRef]
- Kumar, A.; Ting, Y.P. Streptomycin favors biofilm formation by altering cell surface properties. Appl. Microbiol. Biotechnol. 2016, 100, 8843. [Google Scholar] [CrossRef]
- Kaplan, J.B. Antibiotic-Induced Biofilm Formation. Int. J. Artif. Organs 2011, 34, 737–751. [Google Scholar] [CrossRef]
- Wen, X.; Langevin, A.M.; Dunlop, M.J. Antibiotic export by efflux pumps affects growth of neighboring bacteria. Sci. Rep. 2018, 8, 15120. [Google Scholar] [CrossRef]
- Balaban, N.Q.; Merrin, J.; Chait, R.; Kowalik, L.; Leibler, S. Bacterial Persistence as a Phenotypic Switch. Science 2004, 305, 1622–1625. [Google Scholar] [CrossRef] [Green Version]
- Lewis, K. Persister cells, dormancy and infectious disease. Nat. Rev. Genet. 2006, 5, 48–56. [Google Scholar] [CrossRef]
- Lewis, K. Persister Cells. Annu. Rev. Microbiol. 2010, 64, 357–372. [Google Scholar] [CrossRef]
- Harms, A.; Maisonneuve, E.; Gerdes, K. Mechanisms of bacterial persistence during stress and antibiotic exposure. Science 2016, 354, aaf4268. [Google Scholar] [CrossRef] [PubMed]
- Pu, Y.; Zhao, Z.; Li, Y.; Zou, J.; Ma, Q.; Zhao, Y.; Ke, Y.; Zhu, Y.; Chen, H.; Baker, M.; et al. Enhanced Efflux Activity Facilitates Drug Tolerance in Dormant Bacterial Cells. Mol. Cell 2016, 62, 284–294. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dörr, T.; Vulić, M.; Lewis, K. Ciprofloxacin Causes Persister Formation by Inducing the TisB toxin in Escherichia coli. PLoS Biol. 2010, 8, e1000317. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Berghoff, B.A.; Hoekzema, M.; Aulbach, L.; Wagner, E.G.H. Two regulatory RNA elements affect TisB-dependent depolarization and persister formation. Mol. Microbiol. 2017, 103, 1020–1033. [Google Scholar] [CrossRef] [Green Version]
- Edelmann, D.; Leinberger, F.; Schmid, N.; Oberpaul, M.; Schäberle, T.; Berghoff, B. Elevated Expression of Toxin TisB Protects Persister Cells against Ciprofloxacin but Enhances Susceptibility to Mitomycin C. Microorganisms 2021, 9, 943. [Google Scholar] [CrossRef]
- Suttle, C.A. Marine viruses—Major players in the global ecosystem. Nat. Rev. Microbiol. 2007, 5, 801–812. [Google Scholar] [CrossRef]
- Wittebole, X.; De Roock, S.; Opal, S.M. A historical overview of bacteriophage therapy as an alternative to antibiotics for the treatment of bacterial pathogens. Virulence 2013, 5, 226–235. [Google Scholar] [CrossRef]
- Mai, V.; Ukhanova, M.; Reinhard, M.K.; Li, M.; Sulakvelidze, A. Bacteriophage administration significantly reduces Shigella colonization and shedding by Shigella-challenged mice without deleterious side effects and distortions in the gut microbiota. Bacteriophage 2015, 5, e1088124. [Google Scholar] [CrossRef] [Green Version]
- Galtier, M.; De Sordi, L.; Maura, D.; Arachchi, H.; Volant, S.; Dillies, M.; Debarbieux, L. Bacteriophages to reduce gut carriage of antibiotic resistant uropathogens with low impact on microbiota composition. Environ. Microbiol. 2016, 18, 2237–2245. [Google Scholar] [CrossRef] [Green Version]
- Servick, K. Beleaguered phage therapy trial presses on. Science 2016, 352, 1506. [Google Scholar] [CrossRef]
- Flores, C.O.; Valverde, S.; Weitz, J.S. Multi-scale structure and geographic drivers of cross-infection within marine bacteria and phages. ISME J. 2012, 7, 520–532. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wolf, A.; Wiese, J.; Jost, G.; Witzel, K.-P. Wide Geographic Distribution of Bacteriophages That Lyse the Same Indigenous Freshwater Isolate (Sphingomonas sp. Strain B18). Appl. Environ. Microbiol. 2003, 69, 2395–2398. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alam Sarker, S.; Sultana, S.; Reuteler, G.; Moine, D.; Descombes, P.; Charton, F.; Bourdin, G.; McCallin, S.; Ngom-Bru, C.; Neville, T.; et al. Oral Phage Therapy of Acute Bacterial Diarrhea with Two Coliphage Preparations: A Randomized Trial in Children from Bangladesh. EBioMedicine 2016, 4, 124–137. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bourdin, G.; Navarro, A.; Sarker, S.A.; Pittet, A.-C.; Qadri, F.; Sultana, S.; Cravioto, A.; Talukder, K.A.; Reuteler, G.; Brüssow, H. Coverage of diarrhoea-associated Escherichia coli isolates from different origins with two types of phage cocktails. Microb. Biotechnol. 2014, 7, 165–176. [Google Scholar] [CrossRef]
- Balique, F.; Lecoq, H.; Raoult, D.; Colson, P. Can Plant Viruses cross the Kingdom Border and Be Pathogenic to Humans? Viruses 2015, 7, 2074–2098. [Google Scholar] [CrossRef] [Green Version]
- Bellas, C.M.; Schroeder, D.C.; Edwards, A.; Barker, G.; Anesio, A.M. Flexible genes establish widespread bacteriophage pan-genomes in cryoconite hole ecosystems. Nat. Commun. 2020, 11, 4403. [Google Scholar] [CrossRef]
- Bertozzi Silva, J.; Storms, Z.; Sauvageau, D. Host receptors for bacteriophage adsorption. FEMS Microbiol. Lett. 2016, 363, fnw002. [Google Scholar] [CrossRef] [Green Version]
- Harada, L.K.; Silva, E.C.; Campos, W.F.; Fiol, F.D.S.D.; Vila, M.; Dąbrowska, K.; Krylov, V.N.; Balcão, V.M. Biotechnological applications of bacteriophages: State of the art. Microbiol. Res. 2018, 212-213, 38–58. [Google Scholar] [CrossRef]
- Ackermann, H.-W. 5500 Phages examined in the electron microscope. Arch. Virol. 2007, 152, 227–243. [Google Scholar] [CrossRef]
- Hanlon, G.W. Bacteriophages: An appraisal of their role in the treatment of bacterial infections. Int. J. Antimicrob. Agents 2007, 30, 118–128. [Google Scholar] [CrossRef]
- Rakhuba, D.V.; Kolomiets, E.I.; Dey, E.S.; Novik, G.I. Bacteriophage receptors, mechanisms of phage adsorption and penetration into host cell. Pol. J. Microbiol. 2010, 59, 145–155. [Google Scholar] [CrossRef] [PubMed]
- Delbrück, M. The Growth of Bacteriophage and Lysis of the Host. J. Gen. Physiol. 1940, 23, 643–660. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Labedan, B.; Goldberg, E.B. Requirement for membrane potential in injection of phage T4 DNA. Proc. Natl. Acad. Sci. USA 1979, 76, 4669–4673. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Labedan, B.; Letellier, L. Membrane potential changes during the first steps of coliphage infection. Proc. Natl. Acad. Sci. USA 1981, 78, 215–219. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yamamoto, M.; Kanegasaki, S.; Yoshikawa, M. Role of Membrane Potential and ATP in Complex Formation between Escherichia coli Male Cells and Filamentous Phage fd. Microbiology 1981, 123, 343–349. [Google Scholar] [CrossRef] [Green Version]
- Molineux, I.J.; Panja, D. Popping the cork: Mechanisms of phage genome ejection. Nat. Rev. Microbiol. 2013, 11, 194–204. [Google Scholar] [CrossRef]
- Purohit, P.K.; Kondev, J.; Phillips, R. Mechanics of DNA packaging in viruses. Proc. Natl. Acad. Sci. USA 2003, 100, 3173–3178. [Google Scholar] [CrossRef] [Green Version]
- Molineux, I.J. Fifty-three years since Hershey and Chase; much ado about pressure but which pressure is it? Virology 2006, 344, 221–229. [Google Scholar] [CrossRef] [Green Version]
- Mindich, L. Phages with Segmented Double-Stranded RNA Genomes. In The Bacteriophages, 2nd ed.; Calendar, R., Ed.; Oxford University Press: Oxford, UK, 2006. [Google Scholar]
- Maghsoodi, A.; Chatterjee, A.; Andricioaei, I.; Perkins, N.C. How the phage T4 injection machinery works including energetics, forces, and dynamic pathway. Proc. Natl. Acad. Sci. USA 2019, 116, 25097–25105. [Google Scholar] [CrossRef]
- Abdelrahman, F.; Easwaran, M.; Daramola, O.I.; Ragab, S.; Lynch, S.; Oduselu, T.J.; Khan, F.M.; Ayobami, A.; Adnan, F.; Torrents, E.; et al. Phage-Encoded Endolysins. Antibiotics 2021, 10, 124. [Google Scholar] [CrossRef]
- Leiman, P.G.; Arisaka, F.; van Raaij, M.J.; Kostyuchenko, V.A.; Aksyuk, A.A.; Kanamaru, S.; Rossmann, M.G. Morphogenesis of the T4 tail and tail fibers. Virol. J. 2010, 7, 355. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Davison, S.; Couture-Tosi, E.; Candela, T.; Mock, M.; Fouet, A. Identification of the Bacillus anthracis γ Phage Receptor. J. Bacteriol. 2005, 187, 6742–6749. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Morona, R.; Henning, U. New locus (ttr) in Escherichia coli K-12 affecting sensitivity to bacteriophage T2 and growth on oleate as the sole carbon source. J. Bacteriol. 1986, 168, 534–540. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kortright, K.E.; Chan, B.K.; Turner, P.E. High-throughput discovery of phage receptors using transposon insertion sequencing of bacteria. Proc. Natl. Acad. Sci. USA 2020, 117, 18670–18679. [Google Scholar] [CrossRef]
- Duckworth, D.H. History and Basic Properties of Bacterial Viruses. In Phage Ecology; Goyal, S.M., Gerba, C.P., Bitton, G., Eds.; John Wiley & Sons: New York, NY, USA, 1987; Volume 1, p. 43. [Google Scholar]
- Esteves, N.C.; Porwollik, S.; McClelland, M.; Scharf, B.E. The Multidrug Efflux System AcrABZ-TolC Is Essential for Infection of Salmonella Typhimurium by the Flagellum-Dependent Bacteriophage Chi. J. Virol. 2021, 95, e00394-21. [Google Scholar] [CrossRef]
- Gurnev, P.A.; Oppenheim, A.B.; Winterhalter, M.; Bezrukov, S.M. Docking of a Single Phage Lambda to its Membrane Receptor Maltoporin as a Time-resolved Event. J. Mol. Biol. 2006, 359, 1447–1455. [Google Scholar] [CrossRef]
- Comeau, A.M.; Tétart, F.; Trojet, S.N.; Prere, M.-F.; Krisch, H.M. La “synergie phages-antibiotiques“:Un enjeu pour la phagothérapie: The discovery of a natural phenomenon, “Phage-Antibiotic Synergy“. Implic. Phage Ther. 2008, 24, 449–451. [Google Scholar] [CrossRef]
- Li, X.; He, Y.; Wang, Z.; Wei, J.; Hu, T.; Si, J.; Tao, G.; Zhang, L.; Xie, L.; Abdalla, A.E.; et al. A combination therapy of Phages and Antibiotics: Two is better than one. Int. J. Biol. Sci. 2021, 17, 3573–3582. [Google Scholar] [CrossRef]
- Burmeister, A.R.; Turner, P.E. Trading-off and trading-up in the world of bacteria–phage evolution. Curr. Biol. 2020, 30, R1120–R1124. [Google Scholar] [CrossRef]
- Valappil, S.K.; Shetty, P.; Deim, Z.; Terhes, G.; Urbán, E.; Váczi, S.; Patai, R.; Polgár, T.; Pertics, B.Z.; Schneider, G.; et al. Survival Comes at a Cost: A Coevolution of Phage and Its Host Leads to Phage Resistance and Antibiotic Sensitivity of Pseudomonas aeruginosa Multidrug Resistant Strains. Front. Microbiol. 2021, 12, 783722. [Google Scholar] [CrossRef]
- Burmeister, A.R.; Fortier, A.; Roush, C.; Lessing, A.J.; Bender, R.G.; Barahman, R.; Grant, R.; Chan, B.K.; Turner, P.E. Pleiotropy complicates a trade-off between phage resistance and antibiotic resistance. Proc. Natl. Acad. Sci. USA 2020, 117, 11207–11216. [Google Scholar] [CrossRef] [PubMed]
- Chan, B.K.; Sistrom, M.; Wertz, J.E.; Kortright, K.E.; Narayan, D.; Turner, P.E. Phage selection restores antibiotic sensitivity in MDR Pseudomonas aeruginosa. Sci. Rep. 2016, 6, 26717. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Knezevic, P.; Curcin, S.; Aleksic, V.; Petrusic, M.; Vlaški, L. Phage-antibiotic synergism: A possible approach to combatting Pseudomonas aeruginosa. Res. Microbiol. 2013, 164, 55–60. [Google Scholar] [CrossRef] [PubMed]
- Karakozova, M.; Nazarov, P. Conserved sequences of genes coding for the multidrug resistance pump AcrAB-TolC of Escherichia coli suggest their involvement into permanent cell “cleaning”. Bull. Russ. State Med. Univ. 2018, 2, 23–36. [Google Scholar] [CrossRef]
- Tamer, Y.T.; Gaszek, I.; Rodrigues, M.; Coskun, F.S.; Farid, M.; Koh, A.Y.; Russ, W.; Toprak, E. The Antibiotic Efflux Protein TolC Is a Highly Evolvable Target under Colicin E1 or TLS Phage Selection. Mol. Biol. Evol. 2021, 38, 4493–4504. [Google Scholar] [CrossRef]
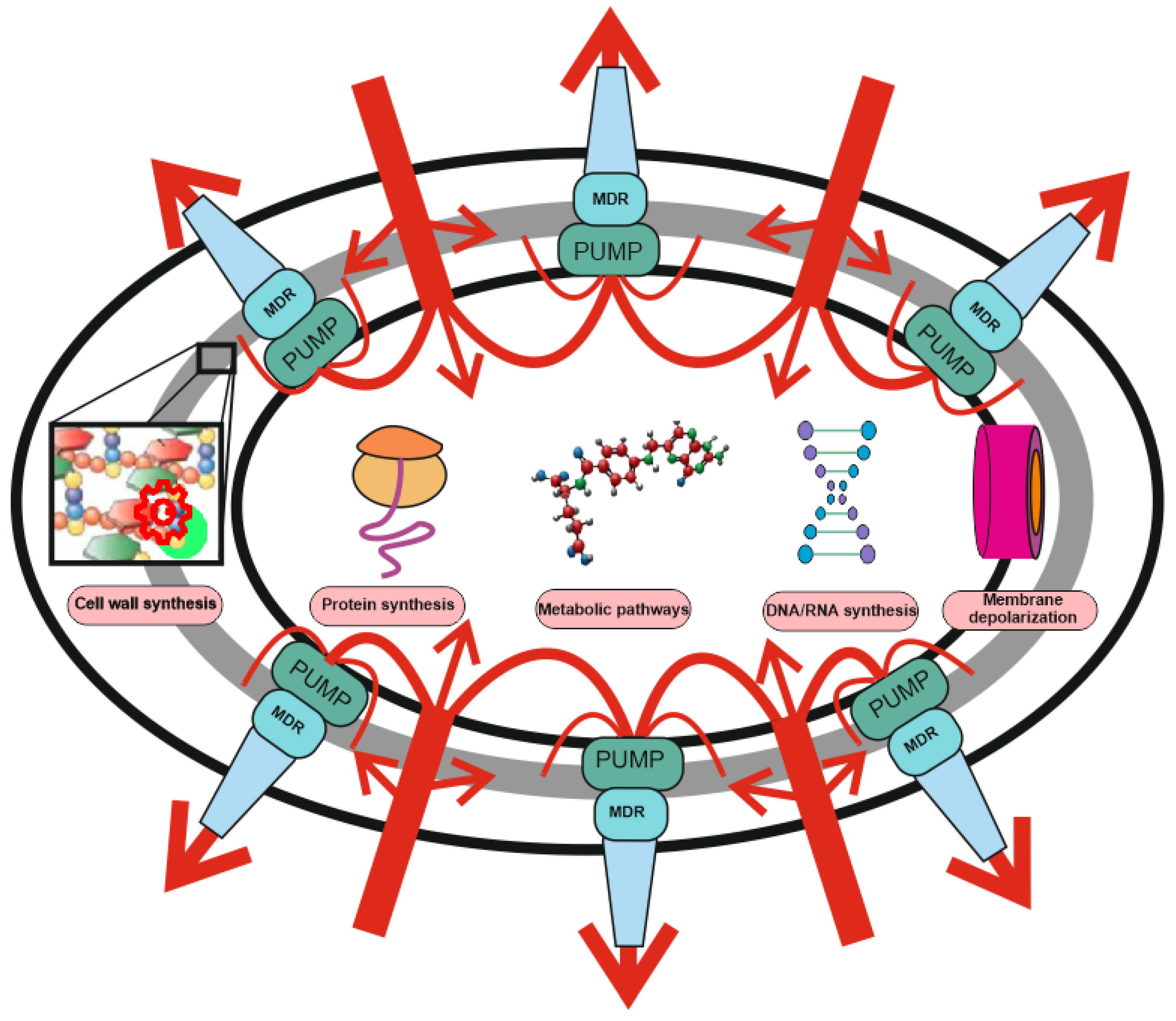
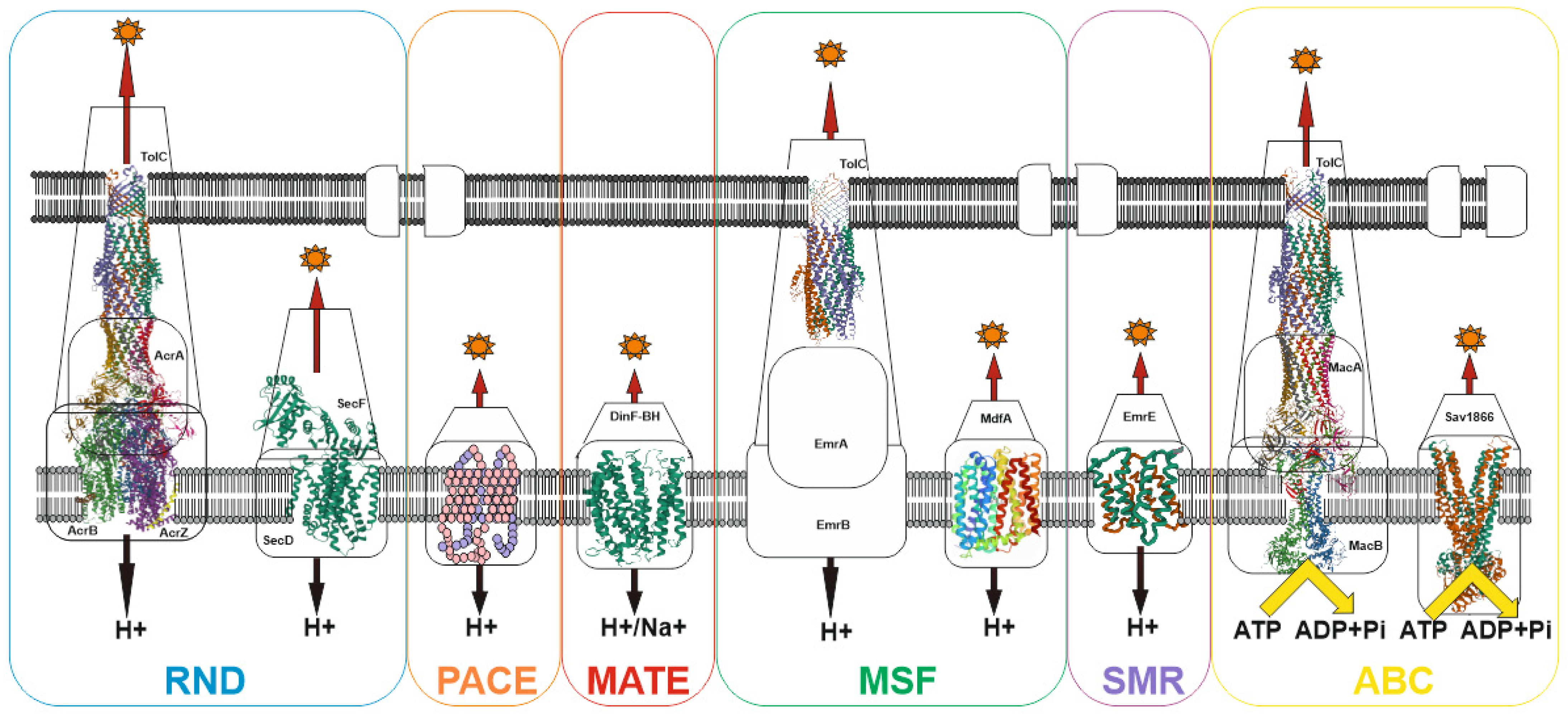
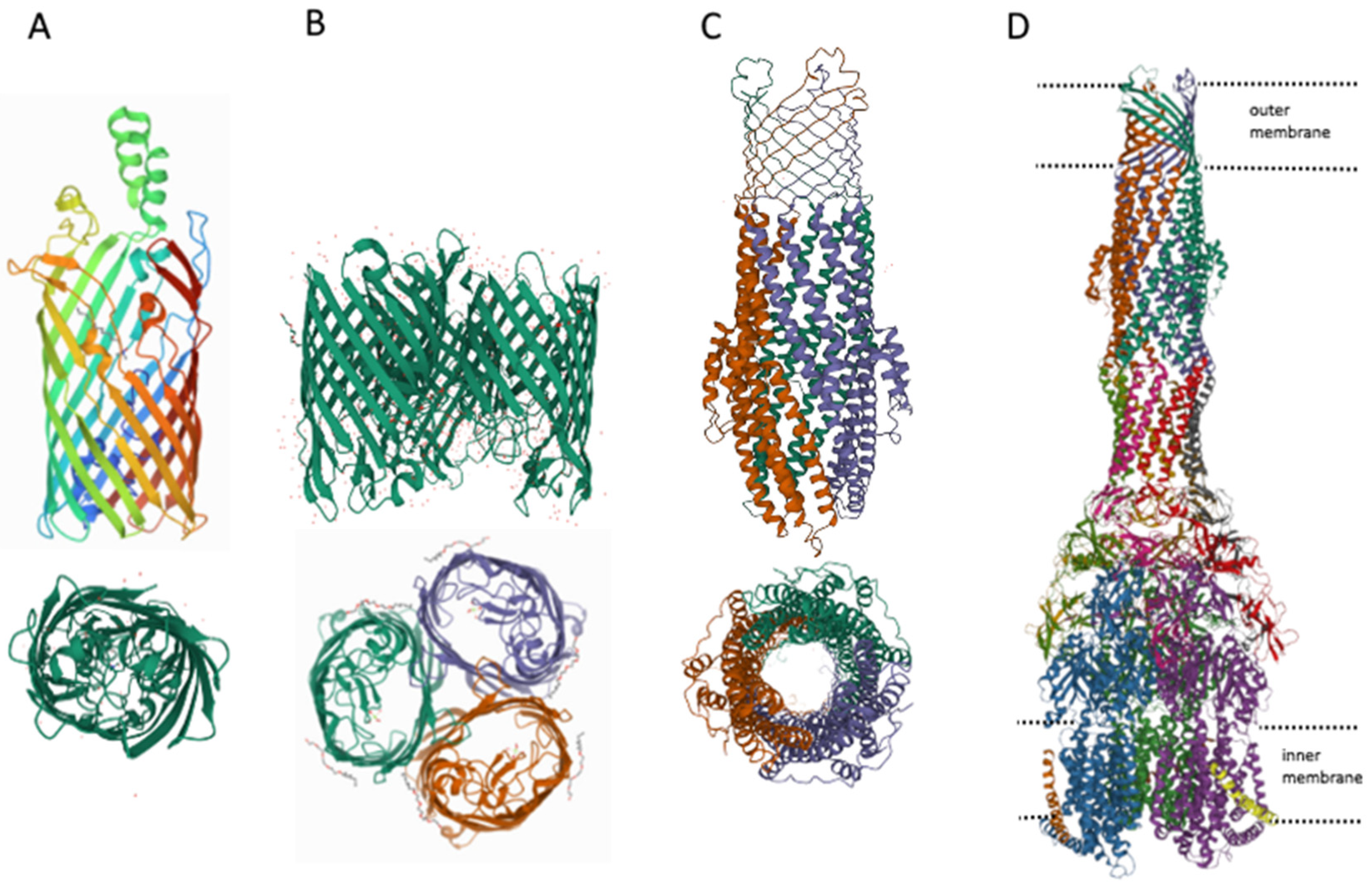
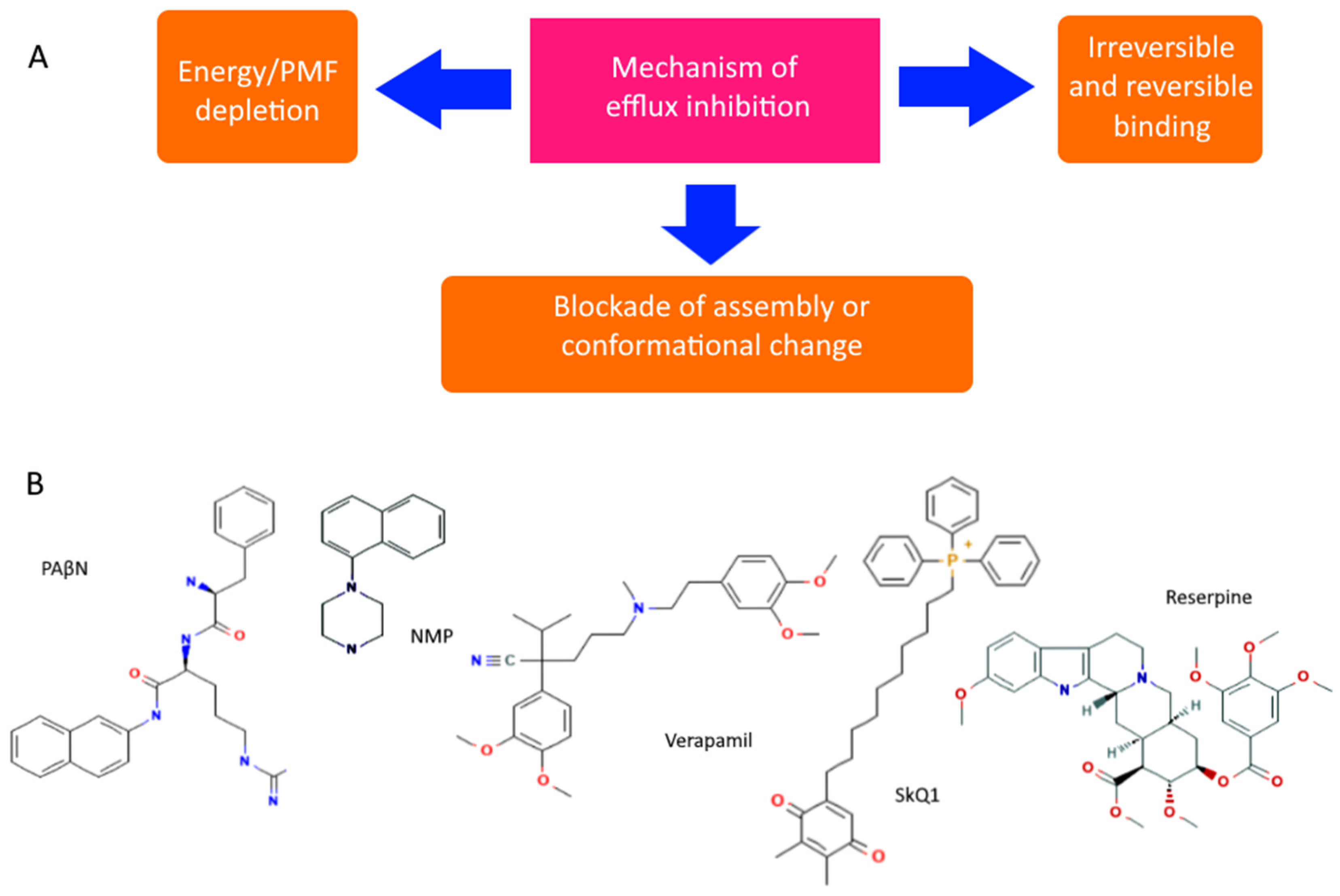
| Mechanism of Action | Antibiotic Group | Antibiotics 1 | References 2 |
|---|---|---|---|
| Protein synthesis: 30S ribosomal subunit binding | Aminoglycosides | Kan, Sm, Gm | [54,55] |
| Tetracyclines | Tet, Dox, Min | [53,56] | |
| Protein synthesis: 50S ribosomal subunit binding | Streptogramins | Q/D | [57] |
| Macrolides | Erm, Clr, Azm | [53,58,59] | |
| Oxazolidinones | Tzd, Rzd | [60] | |
| Lincosamides | Lnm, Cdm, Prm | [61,62,63] | |
| Chloramphenicol | [53] | ||
| Nucleic Acid Synthesis | Quinolones and Fluoroquinolones | Cip, Lvx, Nal | [56] |
| Metabolic Pathways | Sulfonamides | Sul, Smx | [53,64] |
| Trimethoprim | [64] | ||
| Depolarize Cell Membrane | Lipopeptides | Dap | [65] |
| Lantibiotics | Gdm | [65] | |
| Cell Wall Synthesis | β-Lactams | Amp | [53] |
| Carbapenems | Imp | [66] | |
| Cephalosporins | Cpx | [67] | |
| Monobactams | Azn | [68] | |
| Glycopeptides | Van | [69] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nazarov, P.A. MDR Pumps as Crossroads of Resistance: Antibiotics and Bacteriophages. Antibiotics 2022, 11, 734. https://doi.org/10.3390/antibiotics11060734
Nazarov PA. MDR Pumps as Crossroads of Resistance: Antibiotics and Bacteriophages. Antibiotics. 2022; 11(6):734. https://doi.org/10.3390/antibiotics11060734
Chicago/Turabian StyleNazarov, Pavel A. 2022. "MDR Pumps as Crossroads of Resistance: Antibiotics and Bacteriophages" Antibiotics 11, no. 6: 734. https://doi.org/10.3390/antibiotics11060734
APA StyleNazarov, P. A. (2022). MDR Pumps as Crossroads of Resistance: Antibiotics and Bacteriophages. Antibiotics, 11(6), 734. https://doi.org/10.3390/antibiotics11060734






