Prevalence, Phylogroups and Antimicrobial Susceptibility of Escherichia coli Isolates from Food Products
Abstract
1. Introduction
2. Results
2.1. Isolation and Identification of E. coli in Food Products
2.2. Antimicrobial Susceptibility Evaluation the of E. coli Isolates
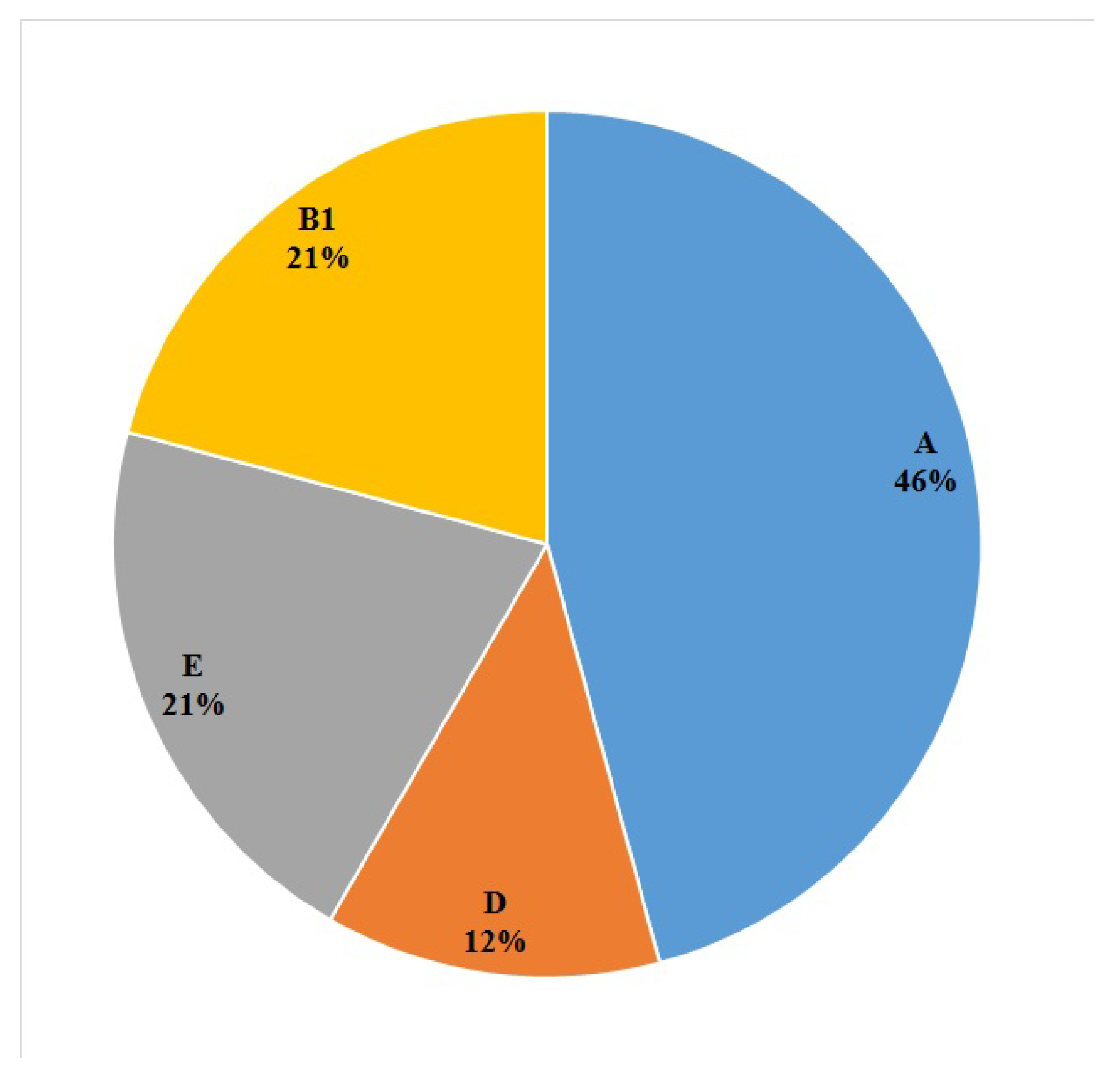
2.3. Phylogroups of the E. coli Isolates
3. Discussion
4. Materials and Methods
4.1. Collection of Food Samples
4.2. Isolation and Identification of E. coli
4.3. Antimicrobial Resistance Testing
4.4. DNA Extraction
4.5. Phylogroups Determination
4.6. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Smith, J.L.; Fratamico, P.M. Emerging and re-emerging foodborne pathogens. Foodborne Pathog. Dis. 2018, 15, 737–757. [Google Scholar] [CrossRef]
- Dewey-Mattia, D.; Manikonda, K.; Hall, A.J.; Wise, M.E.; Crowe, S.J. Surveillance for foodborne disease outbreaks—United States, 2009–2015. MMWR Surveill. Summ. 2018, 67, 1. [Google Scholar] [CrossRef]
- World Health Organization. WHO Estimates of the Global Burden of Foodborne Diseases: Foodborne Disease Burden Epidemiology Reference Group 2007–2015; World Health Organization: Geneva, Switzerland, 2015; Available online: https://apps.who.int/iris/handle/10665/199350 (accessed on 21 April 2015).
- Devleesschauwer, B.; Haagsma, J.A.; Mangen, M.-J.J.; Lake, R.J.; Havelaar, A.H. The global burden of foodborne disease. In Food Safety Economics; Springer: Berlin/Heidelberg, Germany, 2018; pp. 107–122. [Google Scholar] [CrossRef]
- Mainil, J. Escherichia coli virulence factors. Vet. Immunol. Immunopathol. 2013, 152, 2–12. [Google Scholar] [CrossRef] [PubMed]
- Tenaillon, O.; Skurnik, D.; Picard, B.; Denamur, E. The population genetics of commensal Escherichia coli. Nat. Rev. Microbiol. 2010, 8, 207–217. [Google Scholar] [CrossRef] [PubMed]
- Croxen, M.A.; Finlay, B.B. Molecular mechanisms of Escherichia coli pathogenicity. Nat. Rev. Microbiol. 2010, 8, 26–38. [Google Scholar] [CrossRef] [PubMed]
- Ercumen, A.; Pickering, A.J.; Kwong, L.H.; Arnold, B.F.; Parvez, S.M.; Alam, M.; Sen, D.; Islam, S.; Kullmann, C.; Chase, C. Animal feces contribute to domestic fecal contamination: Evidence from E. coli measured in water, hands, food, flies, and soil in Bangladesh. Environ. Sci. Technol. 2017, 51, 8725–8734. [Google Scholar] [CrossRef]
- Jang, J.; Hur, H.G.; Sadowsky, M.J.; Byappanahalli, M.; Yan, T.; Ishii, S. Environmental Escherichia coli: Ecology and public health implications—A review. J. Appl. Microbiol. 2017, 123, 570–581. [Google Scholar] [CrossRef]
- Clermont, O.; Bonacorsi, S.; Bingen, E. Rapid and simple determination of the Escherichia coli phylogenetic group. Appl. Environ. Microbiol. 2000, 66, 4555–4558. [Google Scholar] [CrossRef]
- Clermont, O.; Christenson, J.K.; Denamur, E.; Gordon, D.M. The C lermont Escherichia coli phylo-typing method revisited: Improvement of specificity and detection of new phylo-groups. Environ. Microbiol. Rep. 2013, 5, 58–65. [Google Scholar] [CrossRef] [PubMed]
- Beghain, J.; Bridier-Nahmias, A.; Le Nagard, H.; Denamur, E.; Clermont, O. ClermonTyping: An easy-to-use and accurate in silico method for Escherichia genus strain phylotyping. Microb. Genom. 2018, 4. [Google Scholar] [CrossRef] [PubMed]
- Stoppe, N.d.C.; Silva, J.S.; Carlos, C.; Sato, M.I.; Saraiva, A.M.; Ottoboni, L.M.; Torres, T.T. Worldwide phylogenetic group patterns of Escherichia coli from commensal human and wastewater treatment plant isolates. Front. Microbiol. 2017, 8, 2512. [Google Scholar] [CrossRef]
- Caniça, M.; Manageiro, V.; Abriouel, H.; Moran-Gilad, J.; Franz, C.M. Antibiotic resistance in foodborne bacteria. Trends Food Sci. Technol. 2019, 84, 41–44. [Google Scholar] [CrossRef]
- Walsh, C.; Fanning, S. Antimicrobial resistance in foodborne pathogens—A cause for concern? Curr. Drug Targets 2008, 9, 808–815. [Google Scholar] [CrossRef] [PubMed]
- Magiorakos, A.-P.; Srinivasan, A.; Carey, R.; Carmeli, Y.; Falagas, M.; Giske, C.; Harbarth, S.; Hindler, J.; Kahlmeter, G.; Olsson-Liljequist, B. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 2012, 18, 268–281. [Google Scholar] [CrossRef]
- Rashid, M.; Kotwal, S.K.; Malik, M.; Singh, M. Prevalence, genetic profile of virulence determinants and multidrug resistance of Escherichia coli isolates from foods of animal origin. Vet. World 2013, 6, 139–142. [Google Scholar] [CrossRef]
- Oh, S.-S.; Song, J.; Kim, J.; Shin, J. Increasing prevalence of multidrug-resistant mcr-1-positive Escherichia coli isolates from fresh vegetables and healthy food animals in South Korea. Int. J. Infect. Dis. 2020, 92, 53–55. [Google Scholar] [CrossRef] [PubMed]
- Gomes, T.A.; Elias, W.P.; Scaletsky, I.C.; Guth, B.E.; Rodrigues, J.F.; Piazza, R.M.; Ferreira, L.C.; Martinez, M.B. Diarrheagenic escherichia coli. Braz. J. Microbiol. 2016, 47, 3–30. [Google Scholar] [CrossRef]
- Beauchamp, C.S.; Sofos, J.N. Diarrheagenic Escherichia coli. In Pathogens and Toxins in Foods: Challenges and Interventions 2009; ASM Press: Washington, DC, USA, 2009; pp. 71–94. [Google Scholar]
- Kim, H.J.; Oh, T.; Baek, S.-Y. Multidrug resistance, biofilm formation, and virulence of Escherichia coli isolates from commercial meat and vegetable products. Foodborne Pathog. Dis. 2018, 15, 782–789. [Google Scholar] [CrossRef]
- Pormohammad, A.; Nasiri, M.J.; Azimi, T. Prevalence of antibiotic resistance in Escherichia coli strains simultaneously isolated from humans, animals, food, and the environment: A systematic review and meta-analysis. Infect. Drug Resist. 2019, 12, 1181. [Google Scholar] [CrossRef]
- Canizalez-Roman, A.; Gonzalez-Nuñez, E.; Vidal, J.E.; Flores-Villaseñor, H.; León-Sicairos, N. Prevalence and antibiotic resistance profiles of diarrheagenic Escherichia coli strains isolated from food items in northwestern Mexico. Int. J. Food Microbiol. 2013, 164, 36–45. [Google Scholar] [CrossRef]
- Rúgeles, L.C.; Bai, J.; Martínez, A.J.; Vanegas, M.C.; Gómez-Duarte, O.G. Molecular characterization of diarrheagenic Escherichia coli strains from stools samples and food products in Colombia. Int. J. Food Microbiol. 2010, 138, 282–286. [Google Scholar] [CrossRef]
- Lee, G.Y.; Jang, H.I.; Hwang, I.G.; Rhee, M.S. Prevalence and classification of pathogenic Escherichia coli isolated from fresh beef, poultry, and pork in Korea. Int. J. Food Microbiol. 2009, 134, 196–200. [Google Scholar] [CrossRef] [PubMed]
- Mazaheri, S.; Ahrabi, S.S.; Aslani, M.M. Shiga toxin-producing Escherichia coli isolated from lettuce samples in Tehran, Iran. Jundishapur J. Microbiol. 2014, 7. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Nakamura, H.; Kage-Nakadai, E.; Hara-Kudo, Y.; Nishikawa, Y. Prevalence, antimicrobial resistance and multiple-locus variable-number tandem-repeat analysis profiles of diarrheagenic Escherichia coli isolated from different retail foods. Int. J. Food Microbiol. 2017, 249, 44–52. [Google Scholar] [CrossRef]
- Yu, Z.; Wang, J.; Ho, H.; Wang, Y.; Huang, S.; Han, R. Prevalence and antimicrobial-resistance phenotypes and genotypes of Escherichia coli isolated from raw milk samples from mastitis cases in four regions of China. J. Glob. Antimicrob. Resist. 2020, 22, 94–101. [Google Scholar] [CrossRef]
- Kagambega, A.; Martikainen, O.; Siitonen, A.; Traore, A.S.; Barro, N.; Haukka, K. Prevalence of diarrheagenic E scherichia coli virulence genes in the feces of slaughtered cattle, chickens, and pigs in Burkina Faso. MicrobiologyOpen 2012, 1, 276–284. [Google Scholar] [CrossRef] [PubMed]
- Ombarak, R.A.; Hinenoya, A.; Awasthi, S.P.; Iguchi, A.; Shima, A.; Elbagory, A.-R.M.; Yamasaki, S. Prevalence and pathogenic potential of Escherichia coli isolates from raw milk and raw milk cheese in Egypt. Int. J. Food Microbiol. 2016, 221, 69–76. [Google Scholar] [CrossRef] [PubMed]
- Berge, A.; Baars, T. Raw milk producers with high levels of hygiene and safety. Epidemiol. Infect. 2020, 148. [Google Scholar] [CrossRef]
- Boor, K.J.; Wiedmann, M.; Murphy, S.; Alcaine, S. A 100-year review: Microbiology and safety of milk handling. J. Dairy Sci. 2017, 100, 9933–9951. [Google Scholar] [CrossRef]
- Oniciuc, E.-A.; Likotrafiti, E.; Alvarez-Molina, A.; Prieto, M.; López, M.; Alvarez-Ordóñez, A. Food processing as a risk factor for antimicrobial resistance spread along the food chain. Curr. Opin. Food Sci. 2019, 30, 21–26. [Google Scholar] [CrossRef]
- Hudson, J.A.; Frewer, L.J.; Jones, G.; Brereton, P.A.; Whittingham, M.J.; Stewart, G. The agri-food chain and antimicrobial resistance: A review. Trends Food Sci. Technol. 2017, 69, 131–147. [Google Scholar] [CrossRef]
- Elmonir, W.; Shalaan, S.; Tahoun, A.; Mahmoud, S.F.; Remela, E.M.A.; Eissa, R.; El-Sharkawy, H.; Shukry, M.; Zahran, R.N. Prevalence, antimicrobial resistance, and genotyping of Shiga toxin-producing Escherichia coli in foods of cattle origin, diarrheic cattle, and diarrheic humans in Egypt. Gut Pathog. 2021, 13, 1–11. [Google Scholar] [CrossRef]
- Onmaz, N.E.; Yildirim, Y.; Karadal, F.; Hizlisoy, H.; Al, S.; Gungor, C.; Disli, H.B.; Barel, M.; Dishan, A.; Tegin, R.A.A. Escherichia coli O157 in fish: Prevalence, antimicrobial resistance, biofilm formation capacity, and molecular characterization. LWT 2020, 133, 109940. [Google Scholar] [CrossRef]
- Bailey, J.K.; Pinyon, J.L.; Anantham, S.; Hall, R.M. Distribution of human commensal Escherichia coli phylogenetic groups. J. Clin. Microbiol. 2010, 48, 3455–3456. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Higgins, J.; Hohn, C.; Hornor, S.; Frana, M.; Denver, M.; Joerger, R. Genotyping of Escherichia coli from environmental and animal samples. J. Microbiol. Methods 2007, 70, 227–235. [Google Scholar] [CrossRef] [PubMed]
- Minogue, T.; Daligault, H.; Davenport, K.; Bishop-Lilly, K.; Broomall, S.; Bruce, D.; Chain, P.; Chertkov, O.; Coyne, S.; Freitas, T. Complete genome assembly of Escherichia coli ATCC 25922, a serotype O6 reference strain. Genome Announc. 2014, 2. [Google Scholar] [CrossRef] [PubMed]
- Humphries, R.M.; Ambler, J.; Mitchell, S.L.; Castanheira, M.; Dingle, T.; Hindler, J.A.; Koeth, L.; Sei, K. CLSI methods development and standardization working group best practices for evaluation of antimicrobial susceptibility tests. J. Clin. Microbiol. 2018, 56. [Google Scholar] [CrossRef]
- Pakbin, B.; Mahmoudi, R.; Mousavi, S.; Allahyari, S.; Amani, Z.; Peymani, A.; Qajarbeygi, P.; Hoseinabadi, Z. Genotypic and antimicrobial resistance characterizations of Cronobacter sakazakii isolated from powdered milk infant formula: A comparison between domestic and imported products. Food Sci. Nutr. 2020, 8, 6708–6717. [Google Scholar] [CrossRef]

| Antibiotic class | Antibiotic Agent | n (%) a | |||
|---|---|---|---|---|---|
| Raw Milk (n = 15) | Ground Meat (n = 5) | Vegetable Salad (n = 4) | Total (n = 24) | ||
| β-Lactams | Cefoxitin | 3 (20.0) | 0 (0) | 1 (25.0) | 4 (16.6) |
| Imipenem | 1 (6.6) | 0 (0) | 0 (0) | 1 (4.1) | |
| Amoxicillin | 12 (80.0) | 5 (100) | 2 (50.0) | 19 (79.1) | |
| Ampicillin | 8 (53.3) | 2 (40.0) | 1 (25.0) | 11 (45.8) | |
| Cefepime | 2 (13.3) | 0 (0) | 1 (25.0) | 3 (12.5) | |
| Amoxicillin-clavulanic acid | 11 (73.3) | 3 (60.0) | 1 (25.0) | 15 (62.5) | |
| Aminoglycosides | Streptomycin | 7 (46.6) | 4 (80.0) | 0 (0) | 11 (45.8) |
| Kanamycin | 8 (53.3) | 0 (0) | 0 (0) | 8 (33.3) | |
| Amikacin | 1 (6.6) | 0 (0) | 0 (0) | 1 (4.1) | |
| Gentamicin | 1 (6.6) | 0 (0) | 0 (0) | 1 (4.1) | |
| Quinolones and fluoroquinolones | Nalidixic acid | 0 (0) | 1 (20.0) | 0 (0) | 1 (4.1) |
| Norfloxacin | 0 (0) | 0 (0) | 0 (0) | 0 (0) | |
| Levofloxacin | 1 (6.6) | 0 (0) | 0 (0) | 1 (4.1) | |
| Macrolides | Azithromycin | 0 (0) | 0 (0) | 0 (0) | 0 (0) |
| Tetracyclines | Tetracycline | 10 (66.6) | 3 (60.0) | 0 (0) | 13 (54.1) |
| Lipopeptides | Colistin | 4 (26.6) | 0 (0) | 0 (0) | 4 (16.6) |
| Phenicols | Chloramphenicol | 9 (60.0) | 3 (60.0) | 1 (25.0) | 13 (54.1) |
| Nitroheterocyclics | Nitrofurantoin | 10 (66.6) | 0 (0) | 3 (75.0) | 13 (54.1) |
| Folate pathway antagonists | Trimethoprim-sulfamethoxazole | 12 (80.0) | 4 (80.0) | 1 (25.0) | 17 (70.8) |
| No. Classes of Antibiotics | Multidrug Resistance Patterns a (No. Isolates in Each Pattern) | No. Total Isolates (%) b in Each Class of Antibiotic |
|---|---|---|
| One | βLs (n = 3) | 3 (12.5) |
| Two | βLs-NHCs (n = 4) | 4 (16.6) |
| Three | βLs-AGs-FPAs (n = 1) | 1 (4.1) |
| Four | βLs-TCs-PNs-FPAs (n = 1) | 4 (16.6) |
| βLs-LPs-NHCs-FPAs (n = 1) | ||
| βLs-AGs-NHCs-FPAs (n = 1) | ||
| βLs-PNs-NHCs-FPAs (n = 1) | ||
| Five | βLs-AGs-TCs-PNs-FPAs (n = 3) | 5 (20.8) |
| AGs-TCs-LPs-PNs-FPAs (n = 1) | ||
| βLs-AGs-TCs-NHCs-FPAs (n = 1) | ||
| Six | βLs-AGs-TCs-NHCs-PNs-FPAs (n = 3) | 5 (20.8) |
| βLs-AGs-TCs-QNs-PNs-FPAs (n = 2) | ||
| Seven | βLs-AGs-TCs-LPs-NHCs-PNs-FPAs (n = 2) | 2 (8.3) |
| Food Product | n (%) of the E. coli Isolates (n = 24) | |||
|---|---|---|---|---|
| A | B1 | D | E | |
| Raw milk | 7 (29.1) | 2 (8.3) | 2 (8.3) | 4 (16.6) |
| Vegetable salad | 2 (8.3) | 2 (8.3) | 0 (0) | 0 (0) |
| Ground meat | 2 (8.3) | 1 (4.1) | 1 (4.1) | 1 (4.1) |
| No. | Isolate | Food Sample | Resistance Phenotype a | Phylogroup |
|---|---|---|---|---|
| 1 | ECS1 | Vegetable salad | FOX, NIT | B1 |
| 2 | ECS2 | Raw milk | AMC, IPM, AMX, TET, CHL, SXT | B1 |
| 3 | ECS3 | Raw milk | AMC, KAN, AMX, TET, LVX, FEP, CHL, SXT | E |
| 4 | ECS4 | Raw milk | SPT, AMC, AMX, AMP, TET, FEP, CHL, NIT, SXT | E |
| 5 | ECS5 | Raw milk | FOX, AMC, KAN, AMX, TET, CHL, NIT, SXT | E |
| 6 | ECS6 | Raw milk | AMC, CST, NIT, SXT | A |
| 7 | ECS7 | Raw milk | KAN, TET, CST, CHL, SXT | D |
| 8 | ECS8 | Vegetable salad | AMC, AMX, AMP, NIT | A |
| 9 | ECS9 | Raw milk | SPT, AMC, KAN, AMX, AMP, TET, CHL, SXT | A |
| 10 | ECS10 | Raw milk | FOX, SPT, AMX, AMP, GEN, NIT, SXT | A |
| 11 | ECS11 | Raw milk | SPT, AMC, KAN, AMX, AMP, TET, CST, CHL, NIT SXT | A |
| 12 | ECS12 | Raw milk | SPT, AMC, KAN, AMX, AMP, TET, CHL, NIT SXT | D |
| 13 | ECS13 | Raw milk | FOX, AMX, NIT | A |
| 14 | ECS14 | Ground meat | SPT, AMC, AMX, SXT | A |
| 15 | ECS15 | Ground meat | SPT, AMC, AMX, CHL, SXT | E |
| 16 | ECS16 | Ground meat | NAL, SPT, AMC, AMX, AMP, TET, CHL, SXT | D |
| 17 | ECS17 | Vegetable salad | AMX, CHL, NIT, SXT | B1 |
| 18 | ECS18 | Raw milk | AMX | A |
| 19 | ECS19 | Vegetable salad | FEP | A |
| 20 | ECS20 | Ground meat | AMX | B1 |
| 21 | ECS21 | Raw milk | SPT, AMC, KAN, AMX, AMK, AMP, TET, CST, CHL, NIT, SXT | B1 |
| 22 | ECS22 | Raw milk | SPT, AMC, KAN, AMX, AMP, TET, NIT, SXT | E |
| 23 | ECS23 | Ground meat | SPT, AMX, TET, CHL, SXT | A |
| 24 | ECS24 | Raw milk | AMC, AMX, AMP, NIT | A |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pakbin, B.; Allahyari, S.; Amani, Z.; Brück, W.M.; Mahmoudi, R.; Peymani, A. Prevalence, Phylogroups and Antimicrobial Susceptibility of Escherichia coli Isolates from Food Products. Antibiotics 2021, 10, 1291. https://doi.org/10.3390/antibiotics10111291
Pakbin B, Allahyari S, Amani Z, Brück WM, Mahmoudi R, Peymani A. Prevalence, Phylogroups and Antimicrobial Susceptibility of Escherichia coli Isolates from Food Products. Antibiotics. 2021; 10(11):1291. https://doi.org/10.3390/antibiotics10111291
Chicago/Turabian StylePakbin, Babak, Samaneh Allahyari, Zahra Amani, Wolfram Manuel Brück, Razzagh Mahmoudi, and Amir Peymani. 2021. "Prevalence, Phylogroups and Antimicrobial Susceptibility of Escherichia coli Isolates from Food Products" Antibiotics 10, no. 11: 1291. https://doi.org/10.3390/antibiotics10111291
APA StylePakbin, B., Allahyari, S., Amani, Z., Brück, W. M., Mahmoudi, R., & Peymani, A. (2021). Prevalence, Phylogroups and Antimicrobial Susceptibility of Escherichia coli Isolates from Food Products. Antibiotics, 10(11), 1291. https://doi.org/10.3390/antibiotics10111291







