Efficacy of Ceftazidime and Cefepime in the Management of COVID-19 Patients: Single Center Report from Egypt
Abstract
:1. Introduction
2. Materials and Methods
2.1. Inclusion Criteria
- Patients within an age group of more than 18 years;
- Positive cases of COVID-19 by RT-PCR test;
- Moderate or severe cases with typical symptoms;
- Patients who were treated according to the national standard treatment protocol.
2.2. Exclusion Criteria
- Patients with severe hepatic diseases;
- Pregnant and lactating patients;
- Patients within an age group of less than 18 years.
2.3. Procedure
2.4. In Vitro MPro Inhibition Assay
2.5. In Silico Investigation
2.5.1. Ensemble Docking
2.5.2. Molecular Dynamic Simulation
3. Results
3.1. Clinical Outcomes
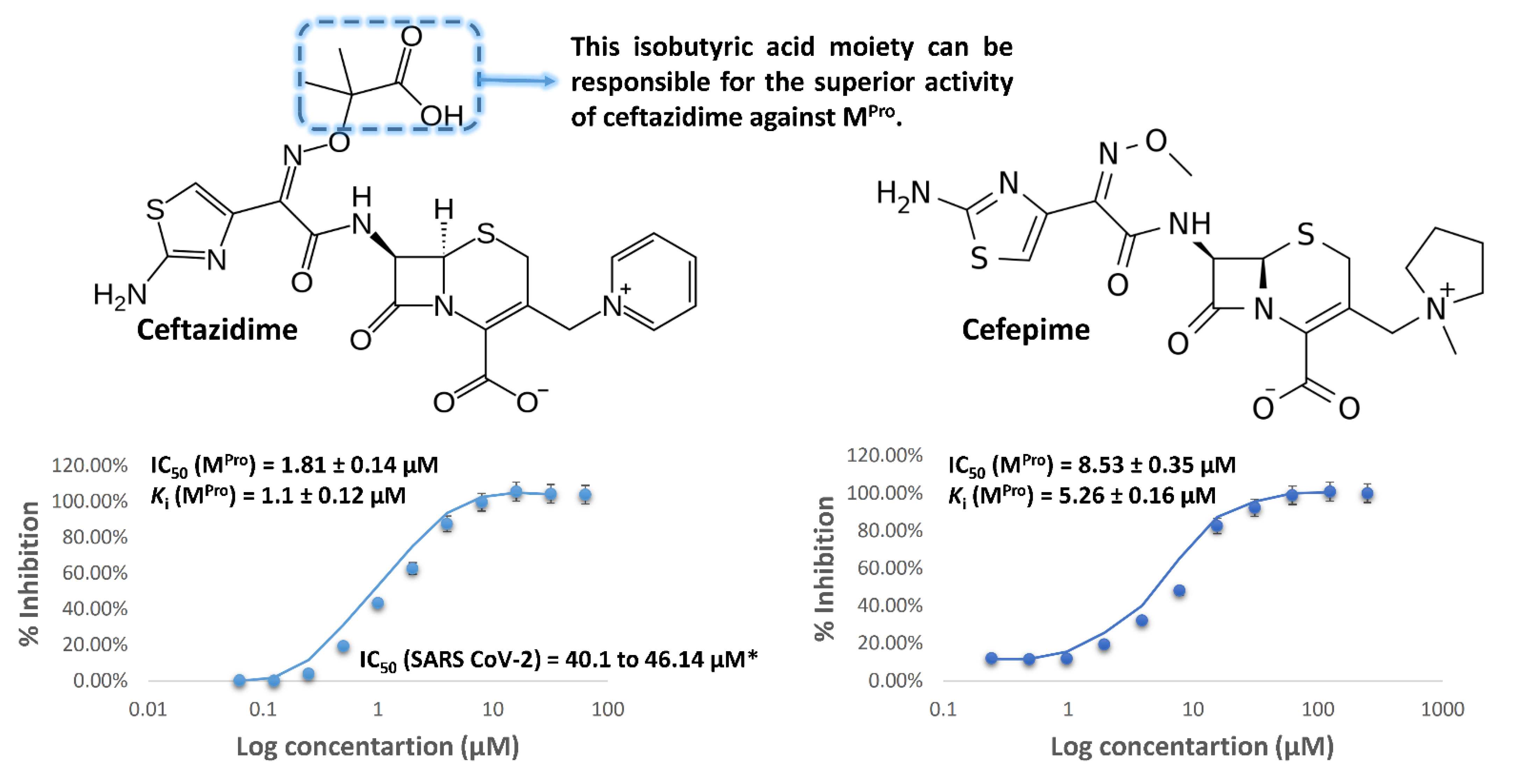
3.2. MPro Inhibitory Activity of the Used Antibiotics
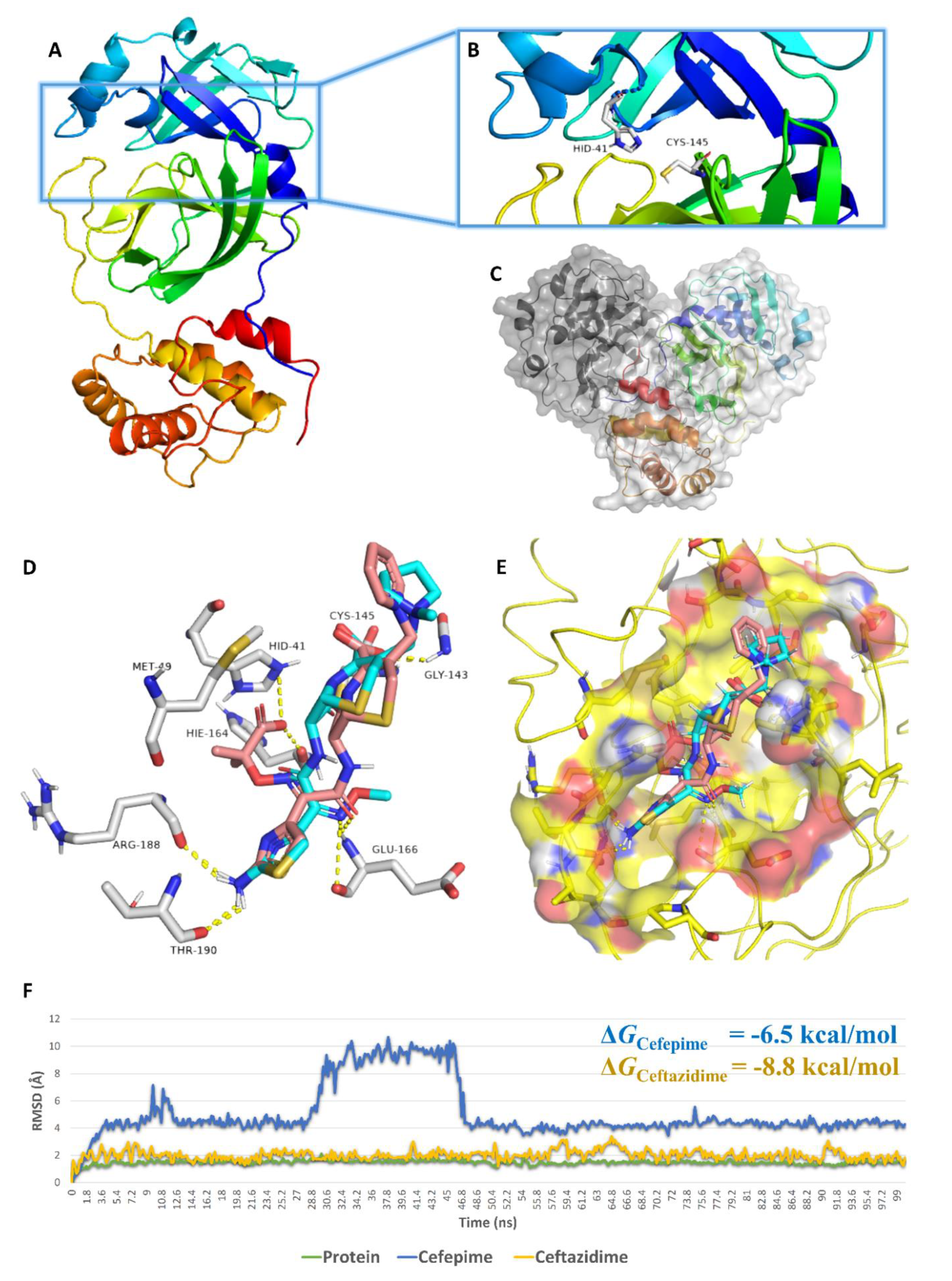
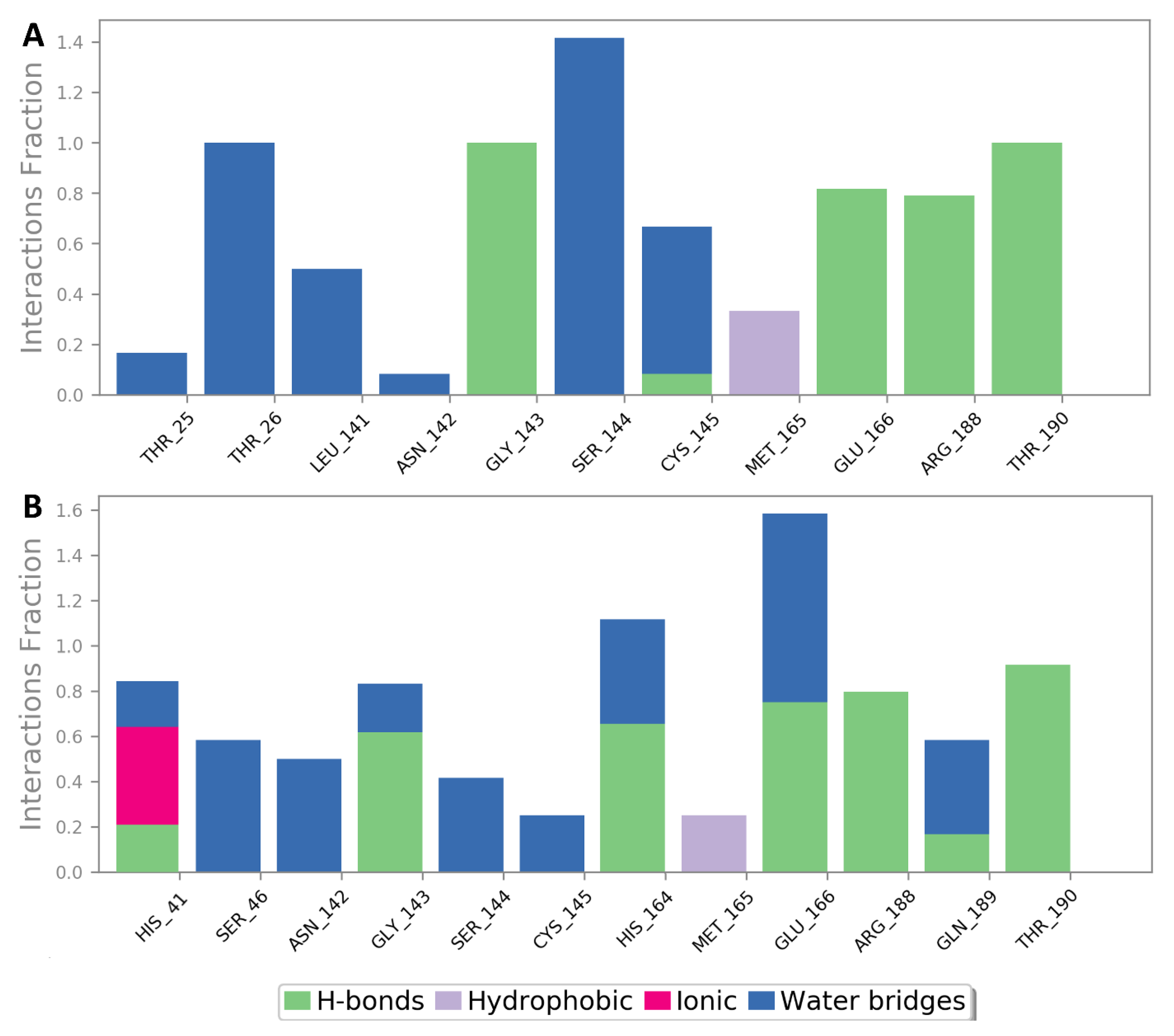
3.3. In-Silico-Based Study of the Mode of Enzyme Inhibition
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hisham Shady, N.; Youssif, K.A.; Sayed, A.M.; Belbahri, L.; Oszako, T.; Hassan, H.M.; Abdelmohsen, U.R. Sterols and triterpenes: Antiviral potential supported by in-silico analysis. Plants 2012, 10, 41. [Google Scholar] [CrossRef]
- Rajter, J.C.; Sherman, M.S.; Fatteh, N.; Vogel, F.; Sacks, J.; Rajter, J.-J. Use of ivermectin is associated with lower mortality in hospitalized patients with coronavirus disease 2019: The ivermectin in COVID nineteen study. Chest 2021, 159, 85–92. [Google Scholar] [CrossRef]
- Chaccour, C.; Casellas, A.; Blanco-Di Matteo, A.; Pineda, I.; Fernandez-Montero, A.; Ruiz-Castillo, P.; Richardson, M.-A.; Rodríguez-Mateos, M.; Jordán-Iborra, C.; Brew, J. The effect of early treatment with ivermectin on viral load, symptoms and humoral response in patients with non-severe COVID-19: A pilot, double-blind, placebo-controlled, randomized clinical trial. EClinicalMedicine 2021, 32, 100720. [Google Scholar] [CrossRef]
- Luis, A.; Wilmer, B.; Hector, C.; Haroldo, D.; Roberto, H. Ivermectin in COVID-19 Patients a Multicenter: Retrospective Study. J. Infect. Dis. Ther. 2021, 9, 1005. [Google Scholar]
- Shamim, K.; Xu, M.; Hu, X.; Lee, E.M.; Lu, X.; Huang, R.; Shah, P.; Xu, X.; Chen, C.Z.; Shen, M. Application of niclosamide and analogs as small molecule inhibitors of Zika virus and SARS-CoV-2 infection. Bioorganic Med. Chem. Lett. 2021, 40, 127906. [Google Scholar] [CrossRef] [PubMed]
- Braga, L.; Ali, H.; Secco, I.; Chiavacci, E.; Neves, G.; Goldhill, D.; Penn, R.; Jimenez-Guardeño, J.M.; Ortega-Prieto, A.M.; Bussani, R. Drugs that inhibit TMEM16 proteins block SARS-CoV-2 Spike-induced syncytia. Nature 2021, 594, 88–93. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Wang, T.; Cai, D.; Hu, Z.; Liao, H.; Zhi, L.; Wei, H.; Zhang, Z.; Qiu, Y.; Wang, J. Cytokine storm intervention in the early stages of COVID-19 pneumonia. Cytokine Growth Factor Rev. 2020, 53, 38–42. [Google Scholar] [CrossRef]
- Fadel, R.; Morrison, A.R.; Vahia, A.; Smith, Z.R.; Chaudhry, Z.; Bhargava, P.; Miller, J.; Kenney, R.M.; Alangaden, G.; Ramesh, M.S. Early short-course corticosteroids in hospitalized patients with COVID-19. Clin. Infect. Dis. 2020, 71, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Godino, C.; Scotti, A.; Maugeri, N.; Mancini, N.; Fominskiy, E.; Margonato, A.; Landoni, G. Antithrombotic therapy in patients with COVID-19? Rationale and Evidence. Int. J. Cardiol. 2021, 324, 261–266. [Google Scholar] [CrossRef]
- Aly, R.; Gupta, S.; Singh, B.; Kaur, P.; Kim, K.; Gupta, S. The use of direct acting oral anticoagulants in patients with COVID-19 infection. J. Community Hosp. Intern. Med. Perspect. 2021, 11, 184–186. [Google Scholar] [CrossRef]
- Huttner, B.; Catho, G.; Pano-Pardo, J.; Pulcini, C.; Schouten, J. COVID-19: Don’t neglect antimicrobial stewardship principles! Clin. Microbiol. Infect. 2020, 26, 808. [Google Scholar] [CrossRef]
- Rawson, T.M.; Moore, L.S.; Zhu, N.; Ranganathan, N.; Skolimowska, K.; Gilchrist, M.; Satta, G.; Cooke, G.; Holmes, A. Bacterial and fungal coinfection in individuals with coronavirus: A rapid review to support COVID-19 antimicrobial prescribing. Clin. Infect. Dis. 2020, 71, 2459–2468. [Google Scholar] [CrossRef]
- Zhou, F.; Yu, T.; Du, R.; Fan, G.; Liu, Y.; Liu, Z.; Xiang, J.; Wang, Y.; Song, B.; Gu, X. Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: A retrospective cohort study. Lancet 2020, 395, 1054–1062. [Google Scholar] [CrossRef]
- Zhou, P.; Liu, Z.; Chen, Y.; Xiao, Y.; Huang, X.; Fan, X.-G. Bacterial and fungal infections in COVID-19 patients: A matter of concern. Infect. Control Hosp. Epidemiol. 2020, 41, 1124–1125. [Google Scholar] [CrossRef] [Green Version]
- Musuuza, J.S.; Watson, L.; Parmasad, V.; Putman-Buehler, N.; Christensen, L.; Safdar, N. Prevalence and outcomes of co-infection and superinfection with SARS-CoV-2 and other pathogens: A systematic review and meta-analysis. PLoS ONE 2021, 16, e0251170. [Google Scholar] [CrossRef]
- Tobaiqy, M.; Qashqary, M.; Al-Dahery, S.; Mujallad, A.; Hershan, A.A.; Kamal, M.A.; Helmi, N. Therapeutic management of COVID-19 patients: A systematic review. Infect. Prev. Pract. 2020, 2, 100061. [Google Scholar] [CrossRef]
- Mahoney, A.R.; Safaee, M.M.; Wuest, W.M.; Furst, A.L. The silent pandemic: Emergent antibiotic resistances following the global response to SARS-CoV-2. Iscience 2021, 24, 102304. [Google Scholar] [CrossRef] [PubMed]
- Miranda, C.; Silva, V.; Capita, R.; Alonso-Calleja, C.; Igrejas, G.; Poeta, P. Implications of antibiotics use during the COVID-19 pandemic: Present and future. J. Antimicrob. Chemother. 2020, 75, 3413–3416. [Google Scholar] [CrossRef]
- Mostafa, A.; Kandeil, A.; AMM Elshaier, Y.; Kutkat, O.; Moatasim, Y.; Rashad, A.A.; Shehata, M.; Gomaa, M.R.; Mahrous, N.; Mahmoud, S.H. FDA-Approved Drugs with Potent In Vitro Antiviral Activity against Severe Acute Respiratory Syndrome Coronavirus 2. Pharmaceuticals 2020, 13, 443. [Google Scholar] [CrossRef] [PubMed]
- Malla, T.R.; Tumber, A.; John, T.; Brewitz, L.; Strain-Damerell, C.; Owen, C.D.; Lukacik, P.; Chan, H.H.; Maheswaran, P.; Salah, E. Mass spectrometry reveals potential of β-lactams as SARS-CoV-2 M pro inhibitors. Chem. Commun. 2021, 57, 1430–1433. [Google Scholar] [CrossRef] [PubMed]
- Masoud, H.; Elassal, G.; Zaky, S.; Baki, A.; Ibrahim, H.; Amin, W. Management Protocol for COVID-19 Patients Version 1.4/30th May 2020; Ministry of Health and Population (MOHP): Cairo, Egypt, 2020. [Google Scholar]
- Seeliger, D.; de Groot, B.L. Ligand docking and binding site analysis with PyMOL and Autodock/Vina. J. Comput. Aided Mol. Des. 2010, 24, 417–422. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jin, Z.; Du, X.; Xu, Y.; Deng, Y.; Liu, M.; Zhao, Y.; Zhang, B.; Li, X.; Zhang, L.; Peng, C.; et al. Structure of M pro from SARS-CoV-2 and discovery of its inhibitors. Nature 2020, 582, 289–293. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Amaro, R.E.; Baudry, J.; Chodera, J.; Demir, Ö.; McCammon, J.A.; Miao, Y.; Smith, J.C. Ensemble docking in drug discovery. Biophys. J. 2018, 114, 2271–2278. [Google Scholar] [CrossRef] [Green Version]
- Bowers, K.J.; Chow, D.E.; Xu, H.; Dror, R.O.; Eastwood, M.P.; Gregersen, B.A.; Klepeis, J.L.; Kolossvary, I.; Moraes, M.A.; Sacerdoti, F.D.; et al. Scalable algorithms for molecular dynamics simulations on commodity clusters. In Proceedings of the SC’06: 2006 ACM/IEEE Conference on Supercomputing, Tampa, FL, USA, 11–17 November 2006; IEEE: Piscataway, NJ, USA, 2016. [Google Scholar]
- Schrödinger Release 2017-3: Desmond Molecular Dynamics System; DE Shaw Research: New York, NY, USA; Maestro-Desmond Interoperability Tools, Schrödinger: New York, NY, USA, 2017.
- Alam, M.A.; Naik, P.K. Applying linear interaction energy method for binding affinity calculations of podophyllotoxin analogues with tubulin using continuum solvent model and prediction of cytotoxic activity. J. Mol. Graph. Model. 2009, 27, 930–943. [Google Scholar] [CrossRef]
- Phillips, J.C.; Braun, R.; Wang, W.; Gumbart, J.; Tajkhorshid, E.; Villa, E.; Chipot, C.; Skeel, R.D.; Kale, L.; Schulten, K. Scalable molecular dynamics with NAMD. J. Comput. Chem. 2005, 26, 1781–1802. [Google Scholar] [CrossRef] [Green Version]
- Jo, S.; Kim, T.; Iyer, V.G.; Im, W. Charmm-Gui: A web-based graphical user interface for Charmm. J. Comput. Chem. 2008, 29, 1859–1865. [Google Scholar] [CrossRef]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: Visual molecular dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef]
- Mahmoud, D.B.; Shitu, Z.; Mostafa, A. Drug repurposing of nitazoxanide: Can it be an effective therapy for COVID-19? J. Genet. Eng. Biotechnol. 2020, 18, 35. [Google Scholar] [CrossRef]
- Lin, C.; Li, Y.; Zhang, Y.; Liu, Z.; Mu, X.; Gu, C.; Liu, J.; Li, Y.; Li, G.; Chen, J. Ceftazidime is a potential drug to inhibit SARS-CoV-2 infection in vitro by blocking spike protein–ACE2 interaction. Signal Transduct. Target. Ther. 2021, 6, 198. [Google Scholar] [CrossRef] [PubMed]
- Mishra, G.P.; Mulani, J. Corticosteroids for COVID-19: The search for an optimum duration of therapy. Lancet Respir. Med. 2021, 9, e8. [Google Scholar] [CrossRef]
- Sayed, A.M.; Khalaf, A.M.; Abdelrahim, M.E.; Elgendy, M.O. Repurposing of some anti-infective drugs for COVID-19 treatment: A surveillance study supported by an in silico investigation. Int. J. Clin. Pract. 2020, 75, e13877. [Google Scholar] [PubMed]
- Lechien, J.R.; Chiesa-Estomba, C.M.; Place, S.; Van Laethem, Y.; Cabaraux, P.; Mat, Q.; Huet, K.; Plzak, J.; Horoi, M.; Hans, S.; et al. Clinical and epidemiological characteristics of 1420 European patients with mild-to-moderate coronavirus disease 2019. J. Intern. Med. 2020, 288, 335–344. [Google Scholar] [CrossRef] [PubMed]
- Barman, M.P.; Rahman, T.; Bora, K.; Borgohain, C. COVID-19 pandemic and its recovery time of patients in India: A pilot study. Diabetes Metab. Syndr. Clin. Res. Rev. 2020, 14, 1205–1211. [Google Scholar] [CrossRef] [PubMed]
- Voinsky, I.; Baristaite, G.; Gurwitz, D. Effects of age and sex on recovery from COVID-19: Analysis of 5769 Israeli patients. J. Infect. 2020, 81, e102–e103. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Eid, R.A.; Elgendy, M.O.; El-Gendy, A.O.; Elgendy, S.O.; Belbahri, L.; Sayed, A.M.; Rateb, M.E. Efficacy of Ceftazidime and Cefepime in the Management of COVID-19 Patients: Single Center Report from Egypt. Antibiotics 2021, 10, 1278. https://doi.org/10.3390/antibiotics10111278
Eid RA, Elgendy MO, El-Gendy AO, Elgendy SO, Belbahri L, Sayed AM, Rateb ME. Efficacy of Ceftazidime and Cefepime in the Management of COVID-19 Patients: Single Center Report from Egypt. Antibiotics. 2021; 10(11):1278. https://doi.org/10.3390/antibiotics10111278
Chicago/Turabian StyleEid, Ragaey A., Marwa O. Elgendy, Ahmed O. El-Gendy, Sara O. Elgendy, Lassaad Belbahri, Ahmed M. Sayed, and Mostafa E. Rateb. 2021. "Efficacy of Ceftazidime and Cefepime in the Management of COVID-19 Patients: Single Center Report from Egypt" Antibiotics 10, no. 11: 1278. https://doi.org/10.3390/antibiotics10111278
APA StyleEid, R. A., Elgendy, M. O., El-Gendy, A. O., Elgendy, S. O., Belbahri, L., Sayed, A. M., & Rateb, M. E. (2021). Efficacy of Ceftazidime and Cefepime in the Management of COVID-19 Patients: Single Center Report from Egypt. Antibiotics, 10(11), 1278. https://doi.org/10.3390/antibiotics10111278









