Antimicrobial Resistance in Escherichia coli Strains Isolated from Humans and Pet Animals
Abstract
1. Introduction
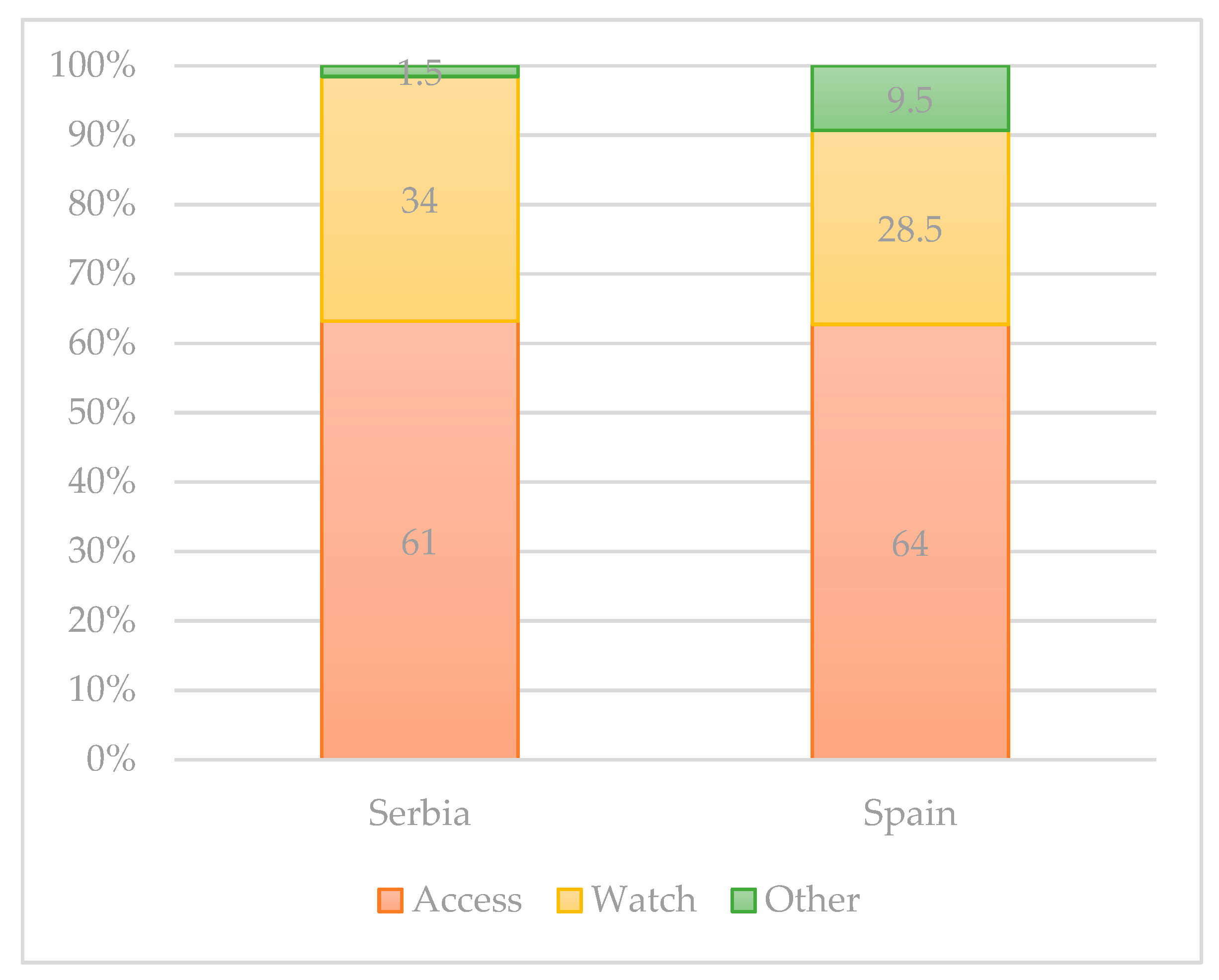
2. Usage of Antibiotics in Different Countries of EU Region and Spread of E. coli Resistance to Antibiotics
3. Inappropriate Prescribing of Antibiotics
4. Mechanisms of β-Lactams Resistance towards E. coli
- Production of β-lactamases, which render β-lactams ineffective
- Inhibited penetration of antibiotics to the intended location
- Modification of the target site PBPs
- Activation of efflux pumps
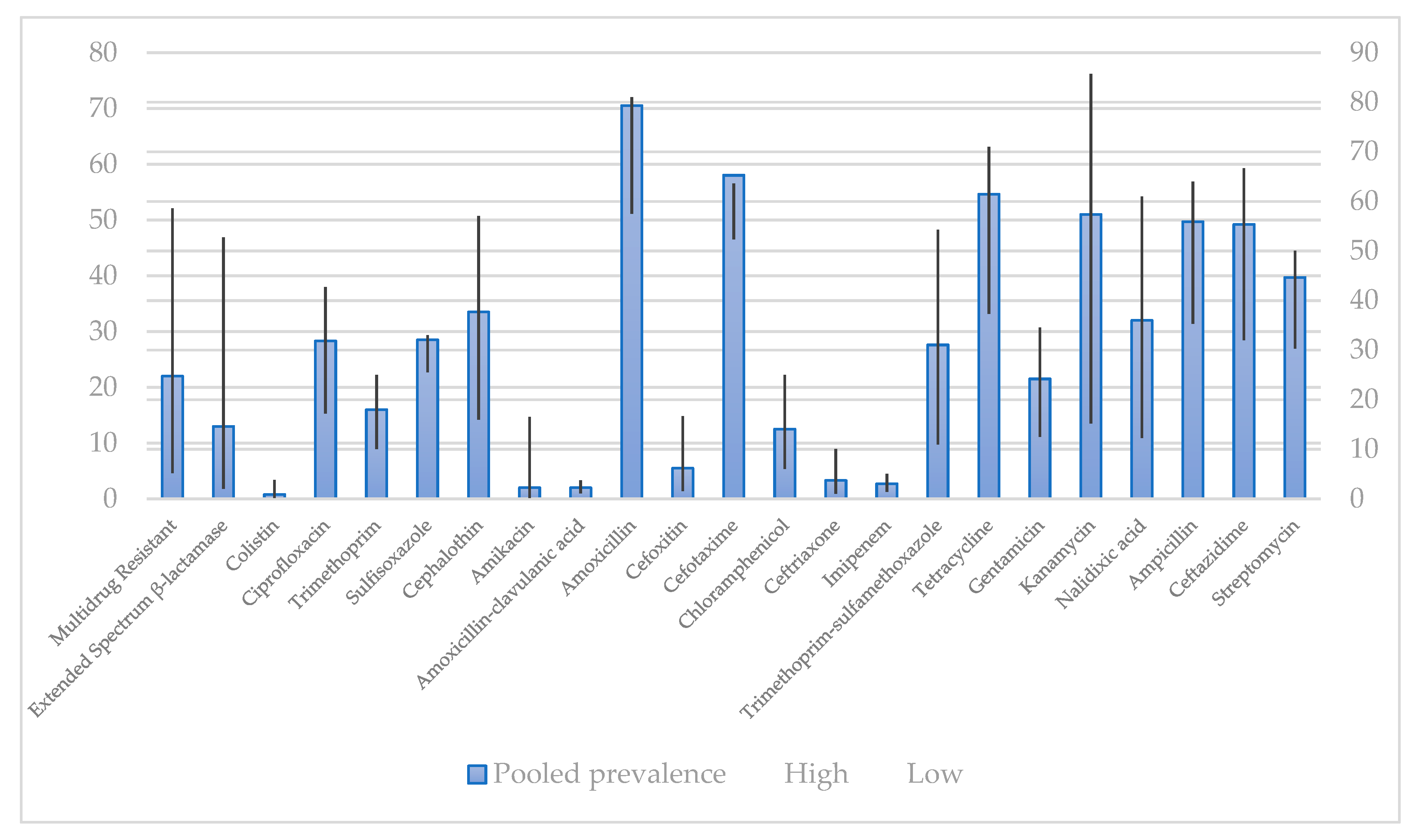
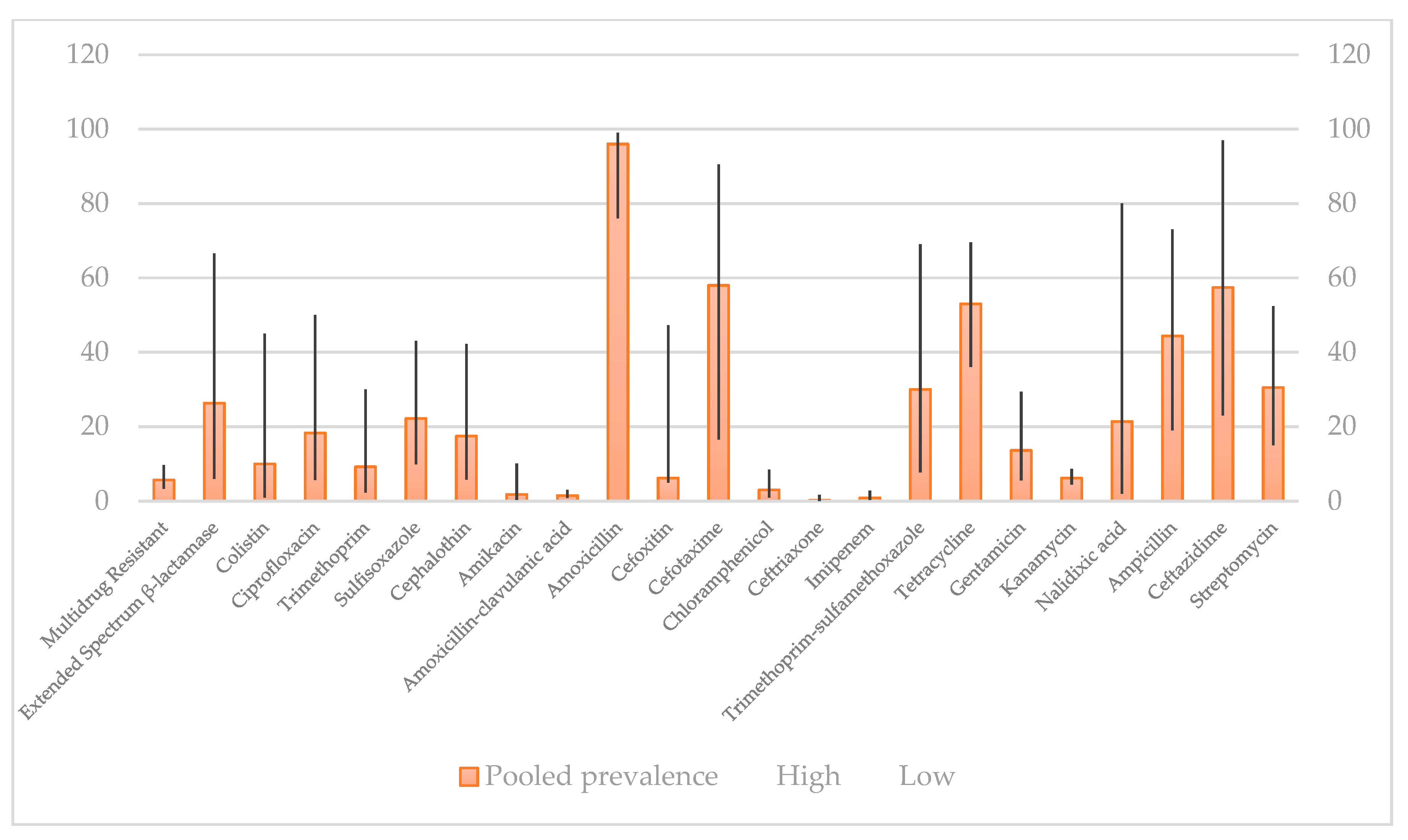
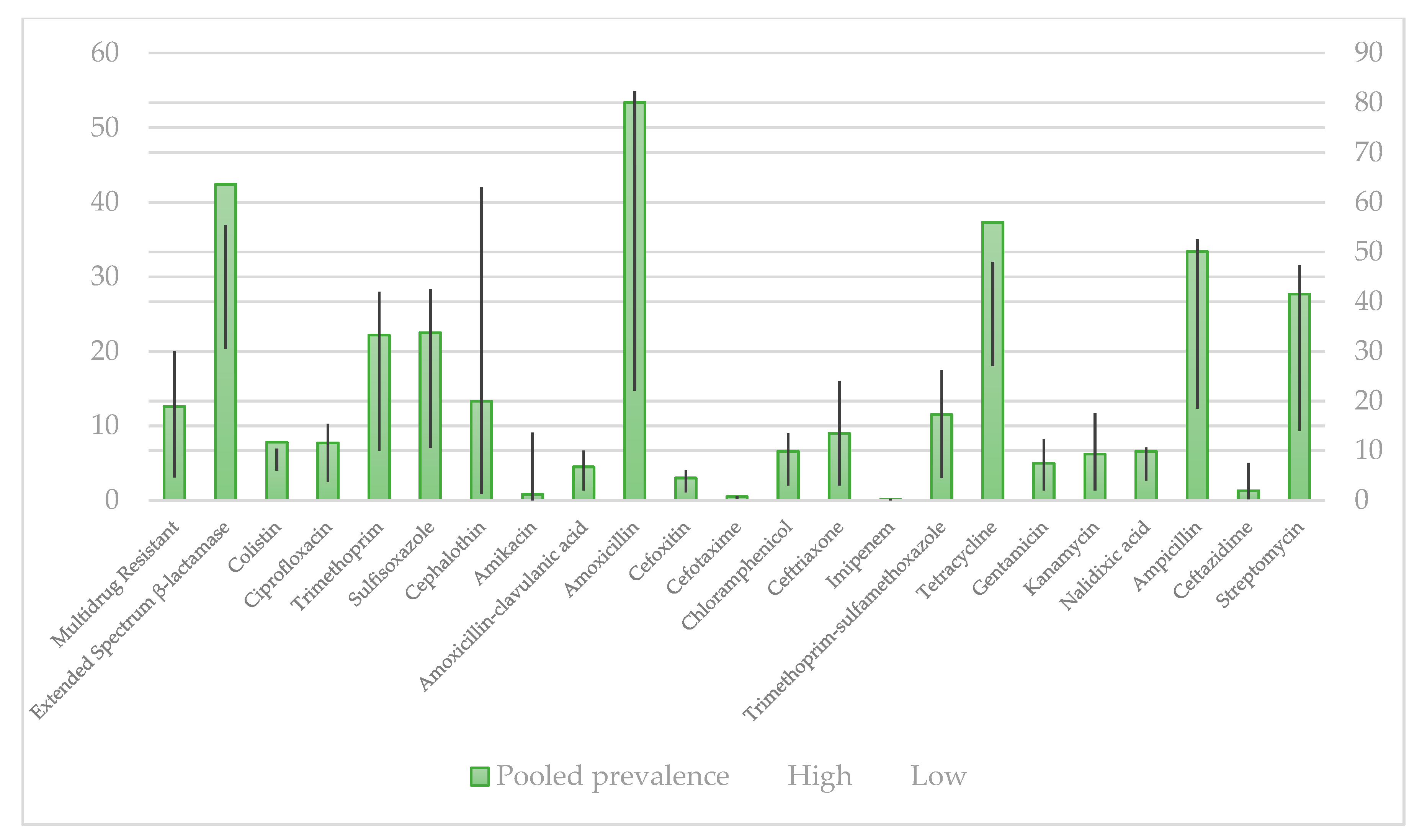
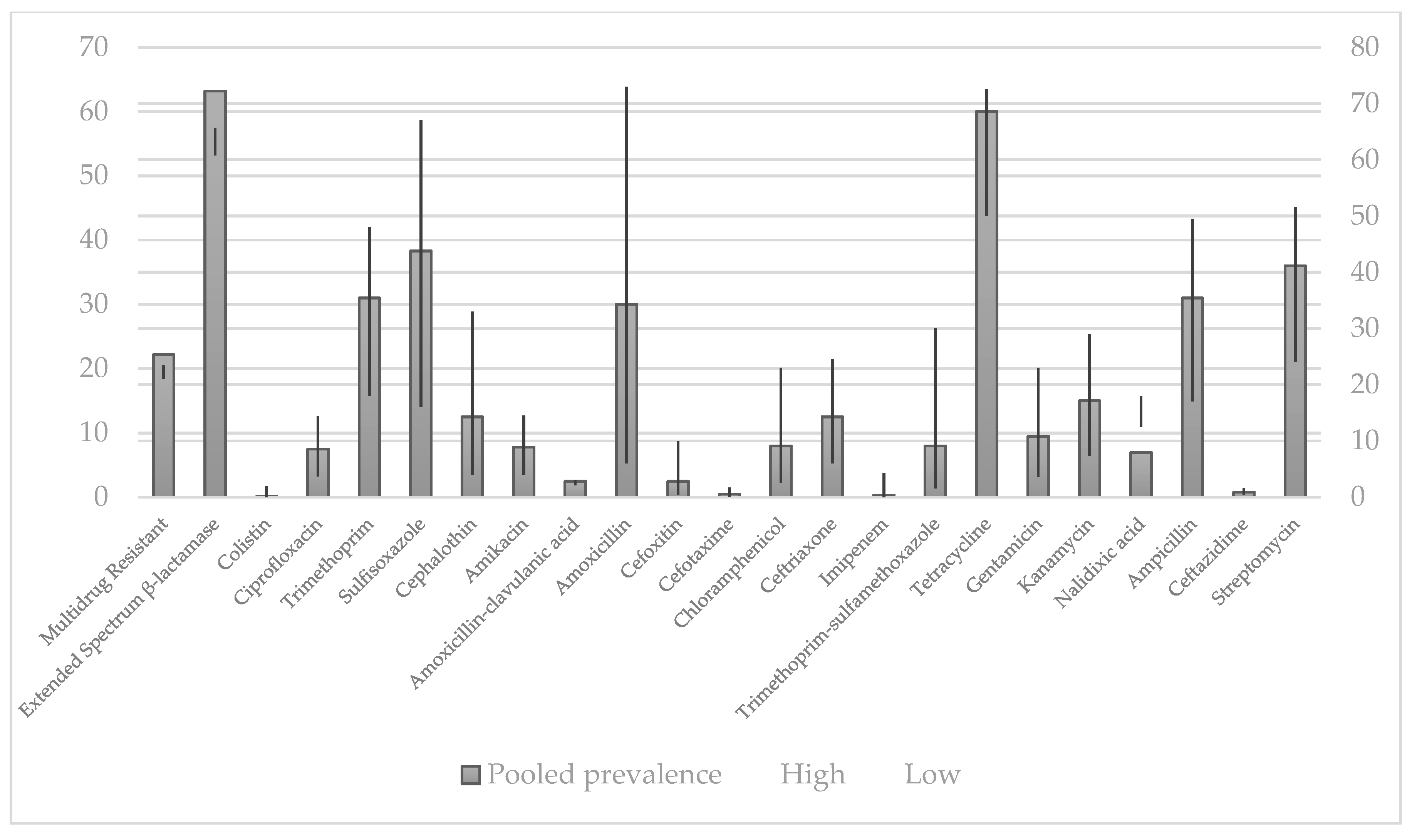
4.1. Prevalence of Antibiotic Resistance in E. coli Isolates by Disk Diffusion Method
4.2. Prevalence of Antibiotic Resistance in E. coli Isolates by Minimum Inhibitory Concentration
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hill, R.A.; Hunt, J.; Sanders, E.; Tran, M.; Burk, G.A.; Mlsna, T.E.; Fitzkee, N.C. Effect of Biochar on Microbial Growth: A Metabolomics and Bacteriological Investigation in E. Coli. Environ. Sci. Technol. 2019, 53, 2635–2646. [Google Scholar] [CrossRef] [PubMed]
- Lovley, D.R.; Holmes, D.E. Protein Nanowires: The Electrification of the Microbial World and Maybe Our Own. J. Bacteriol. 2020, 202, e00331-20. [Google Scholar] [CrossRef] [PubMed]
- Ranganathan, S.; Smith, E.M.; Abel, J.D.F.; Barry, E.M. Research in a Time of Enteroids and Organoids: How the Human Gut Model Has Transformed the Study of Enteric Bacterial Pathogens. Gut Microbes 2020, 12, 1795492. [Google Scholar] [CrossRef] [PubMed]
- Macklin, D.N.; Horst, T.A.A.; Choi, H.; Ruggero, N.A.; Carrera, J.; Mason, J.C.; Sun, G.; Agmon, E.; DeFelice, M.M.; Maayan, I.; et al. Simultaneous Cross-Evaluation of Heterogeneous E. Coli Datasets via Mechanistic Simulation. Science 2020, 369, eaav3751. [Google Scholar] [CrossRef] [PubMed]
- Micenková, L.; Bosák, J.; Smatana, S.; Novotný, A.; Budinská, E.; Šmajs, D. Administration of the Probiotic Escherichia Coli Strain A0 34/86 Resulted in a Stable Colonization of the Human Intestine During the First Year of Life. Probiot. Antimicrob. Prot. 2020, 12, 343–350. [Google Scholar] [CrossRef] [PubMed]
- Bittinger, K.; Zhao, C.; Li, Y.; Ford, E.; Friedman, E.S.; Ni, J.; Kulkarni, C.V.; Cai, J.; Tian, Y.; Liu, Q.; et al. Bacterial Colonization Reprograms the Neonatal Gut Metabolome. Nat. Microbiol. 2020, 5, 838–847. [Google Scholar] [CrossRef]
- Secher, T.; Brehin, C.; Oswald, E. Early Settlers: Which E. Coli Strains Do You Not Want at Birth? Am. J. Physiol. Gastrointest. Liver Physiol. 2016, 311, G123–G129. [Google Scholar] [CrossRef]
- Rossi, E.; Cimdins, A.; Lüthje, P.; Brauner, A.; Sjöling, Å.; Landini, P.; Römling, U. “It’s a Gut Feeling”—Escherichia Coli Biofilm Formation in the Gastrointestinal Tract Environment. Crit. Rev. Microbiol. 2018, 44, 1–30. [Google Scholar] [CrossRef]
- Zhang, S.; Abbas, M.; Rehman, M.U.; Huang, Y.; Zhou, R.; Gong, S.; Yang, H.; Chen, S.; Wang, M.; Cheng, A. Dissemination of Antibiotic Resistance Genes (ARGs) via Integrons in Escherichia Coli: A Risk to Human Health. Environ. Pollut. 2020, 266, 115260. [Google Scholar] [CrossRef]
- Abebe, E.; Gugsa, G.; Ahmed, M. Review on Major Food-Borne Zoonotic Bacterial Pathogens. J. Trop. Med. 2020, 2020, 4674235. [Google Scholar] [CrossRef]
- Isla, A.L.; Polo, J.M.; Isa, M.A.; Sala, R.B.; Sutil, R.S.; Quintas, J.J.; Moradillo, J.G.; Padilla, D.A.G.; Rojo, E.G.; Martínez, J.B.P.; et al. Urinary Infections in Patients with Catheters in the Upper Urinary Tract: Microbiological Study. Urol. Int. 2017, 98, 442–448. [Google Scholar] [CrossRef] [PubMed]
- Rodrigues, W.; Miguel, C.; Nogueira, A.; Vieira, C.U.; Paulino, T.; Soares, S.; De Resende, E.; Chica, J.L.; Araújo, M.; Oliveira, C. Antibiotic Resistance of Bacteria Involved in Urinary Infections in Brazil: A Cross-Sectional and Retrospective Study. Int. J. Environ. Res. Public Health 2016, 13, 918. [Google Scholar] [CrossRef]
- Song, D.W.; Park, B.K.; Suh, S.W.; Lee, S.E.; Kim, J.W.; Park, J.-M.; Kim, H.R.; Lee, M.-K.; Choi, Y.S.; Kim, B.G.; et al. Bacterial Culture and Antibiotic Susceptibility in Patients with Acute Appendicitis. Int. J. Colorectal Dis. 2018, 33, 441–447. [Google Scholar] [CrossRef] [PubMed]
- Park, J.; Kim, S.; Lim, H.; Liu, A.; Hu, S.; Lee, J.; Zhuo, H.; Hao, Q.; Matthay, M.A.; Lee, J.-W. Therapeutic Effects of Human Mesenchymal Stem Cell Microvesicles in an Ex Vivo Perfused Human Lung Injured with Severe E. Coli Pneumonia. Thorax 2019, 74, 43–50. [Google Scholar] [CrossRef]
- Zhao, W.-D.; Liu, D.-X.; Wei, J.-Y.; Miao, Z.-W.; Zhang, K.; Su, Z.-K.; Zhang, X.-W.; Li, Q.; Fang, W.-G.; Qin, X.-X.; et al. Caspr1 Is a Host Receptor for Meningitis-Causing Escherichia Coli. Nat. Commun. 2018, 9, 2296. [Google Scholar] [CrossRef] [PubMed]
- Akuzawa, N.; Kurabayashi, M. Native Valve Endocarditis Due to Escherichia Coli Infection: A Case Report and Review of the Literature. BMC Cardiovasc. Disord. 2018, 18, 195. [Google Scholar] [CrossRef]
- Sarowska, J.; Koloch, B.F.; Kmiecik, A.J.; Madrzak, M.F.; Ksiazczyk, M.; Ploskonska, G.B.; Krol, I.C. Virulence Factors, Prevalence and Potential Transmission of Extraintestinal Pathogenic Escherichia Coli Isolated from Different Sources: Recent Reports. Gut Pathog. 2019, 11, 10. [Google Scholar] [CrossRef]
- Poolman, J.T.; Anderson, A.S. Escherichia Coli and Staphylococcus Aureus: Leading Bacterial Pathogens of Healthcare Associated Infections and Bacteremia in Older-Age Populations. Expert Rev. Vaccines 2018, 17, 607–618. [Google Scholar] [CrossRef]
- Kubone, P.Z.; Mlisana, K.P.; Govinden, U.; Abia, A.L.K.; Essack, S.Y. Antibiotic Susceptibility and Molecular Characterization of Uropathogenic Escherichia Coli Associated with Community-Acquired Urinary Tract Infections in Urban and Rural Settings in South Africa. Trop. Med. Infect. Dis. 2020, 5, 176. [Google Scholar] [CrossRef]
- Djordjevic, Z.; Folic, M.; Jankovic, S. Community-Acquired Urinary Tract Infections: Causative Agents and Their Resistance to Antimicrobial Drugs. Vojnosanit. Pregl. 2016, 73, 1109–1115. [Google Scholar] [CrossRef]
- Gaynes, R. The Discovery of Penicillin—New Insights After More Than 75 Years of Clinical Use. Emerg. Infect. Dis. 2017, 23, 849–853. [Google Scholar] [CrossRef]
- Hutchings, M.I.; Truman, A.W.; Wilkinson, B. Antibiotics: Past, Present and Future. Curr. Opin. Microbiol. 2019, 51, 72–80. [Google Scholar] [CrossRef] [PubMed]
- Dodds, D.R. Antibiotic Resistance: A Current Epilogue. Biochem. Pharmacol. 2017, 134, 139–146. [Google Scholar] [CrossRef] [PubMed]
- Aminov, R. History of Antimicrobial Drug Discovery: Major Classes and Health Impact. Biochem. Pharmacol. 2017, 133, 4–19. [Google Scholar] [CrossRef] [PubMed]
- de Opitz, C.L.M.; Sass, P. Tackling Antimicrobial Resistance by Exploring New Mechanisms of Antibiotic Action. Future Microbiol. 2020, 15, 703–708. [Google Scholar] [CrossRef]
- Gajdács, M.; Albericio, F. Antibiotic Resistance: From the Bench to Patients. Antibiotics 2019, 8, 129. [Google Scholar] [CrossRef]
- Coates, A.R.M.; Hu, Y.; Holt, J.; Yeh, P. Antibiotic Combination Therapy against Resistant Bacterial Infections: Synergy, Rejuvenation and Resistance Reduction. Expert Rev. Anti Infect. Ther. 2020, 18, 5–15. [Google Scholar] [CrossRef]
- Wong, J.W.; Ip, M.; Tang, A.; Wei, V.W.; Wong, S.Y.; Riley, S.; Read, J.M.; Kwok, K.O. Prevalence and Risk Factors of Community-Associated Methicillin-Resistant Staphylococcus Aureus Carriage in Asia-Pacific Region from 2000 to 2016: A Systematic Review and Meta-Analysis. Clin. Epidemiol. 2018, 10, 1489–1501. [Google Scholar] [CrossRef]
- Adeiza, S.S.; Onaolapo, J.A.; Olayinka, B.O. Prevalence, Risk-Factors, and Antimicrobial Susceptibility Profile of Methicillin-Resistant Staphylococcus Aureus (MRSA) Obtained from Nares of Patients and Staff of Sokoto State-Owned Hospitals in Nigeria. GMS Hyg. Infect. Control 2020, 15, Doc25. [Google Scholar] [CrossRef]
- Queenan, K.; Häsler, B.; Rushton, J. A One Health Approach to Antimicrobial Resistance Surveillance: Is There a Business Case for It? Int. J. Antimicrob. Agents 2016, 48, 422–427. [Google Scholar] [CrossRef]
- Tillotson, G.S.; Zinner, S.H. Burden of Antimicrobial Resistance in an Era of Decreasing Susceptibility. Expert Rev. Anti Infect. Ther. 2017, 15, 663–676. [Google Scholar] [CrossRef] [PubMed]
- Heward, E.; Cullen, M.; Hobson, J. Microbiology and Antimicrobial Susceptibility of Otitis Externa: A Changing Pattern of Antimicrobial Resistance. J. Laryngol. Otol. 2018, 132, 314–317. [Google Scholar] [CrossRef]
- Cillóniz, C.; Ardanuy, C.; Vila, J.; Torres, A. What Is the Clinical Relevance of Drug-Resistant Pneumococcus? Curr. Opin. Pulm. Med. 2016, 22, 227–234. [Google Scholar] [CrossRef] [PubMed]
- Roman, A.C.; Roman, J.V.; Flores, M.A.V.; Villaseñor, H.F.; Vidal, J.E.; Amador, S.M.; Llanos, A.M.G.; Nuñez, E.G.; Serrano, J.M.; Pastrana, G.T.; et al. Detection of Antimicrobial-Resistance Diarrheagenic Escherichia Coli Strains in Surface Water Used to Irrigate Food Products in the Northwest of Mexico. Int. J. Food Microbiol. 2019, 304, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Relhan, N.; Pathengay, A.; Schwartz, S.G.; Flynn, H.W. Emerging Worldwide Antimicrobial Resistance, Antibiotic Stewardship and Alternative Intravitreal Agents for the Treatment of Endophthalmitis. Retina 2017, 37, 811–818. [Google Scholar] [CrossRef] [PubMed]
- Tomičić, Z.; Čabarkapa, I.; Čolović, R.; Đuragić, O.; Tomičić, R. Salmonella in the Feed Industry: Problems and Potential Solutions. J. Agron. Technol. Eng. Manag. 2019, 2, 130–137. [Google Scholar]
- Serwecińska, L.; Kiedrzyńska, E.; Kiedrzyński, M. A Catchment-Scale Assessment of the Sanitary Condition of Treated Wastewater and River Water Based on Fecal Indicators and Carbapenem-Resistant Acinetobacter Spp. Sci. Total Environ. 2021, 750, 142266. [Google Scholar] [CrossRef] [PubMed]
- Tagliabue, A.; Rappuoli, R. Changing Priorities in Vaccinology: Antibiotic Resistance Moving to the Top. Front. Immunol. 2018, 9, 1068. [Google Scholar] [CrossRef] [PubMed]
- Mutairi, R.A.; Tovmasyan, A.; Haberle, I.B.; Benov, L. Sublethal Photodynamic Treatment Does Not Lead to Development of Resistance. Front. Microbiol. 2018, 9, 1699. [Google Scholar] [CrossRef] [PubMed]
- Hong, J.; Hu, J.; Ke, F. Experimental Induction of Bacterial Resistance to the Antimicrobial Peptide Tachyplesin I and Investigation of the Resistance Mechanisms. Antimicrob. Agents Chemother. 2016, 60, 6067–6075. [Google Scholar] [CrossRef]
- van den Bergh, B.; Michiels, J.E.; Fauvart, M.; Michiels, J. Should We Develop Screens for Multi-Drug Antibiotic Tolerance? Expert Rev. Anti Infect. Ther. 2016, 14, 613–616. [Google Scholar] [CrossRef] [PubMed]
- Spagnolo, F.; Rinaldi, C.; Sajorda, D.R.; Dykhuizen, D.E. Evolution of Resistance to Continuously Increasing Streptomycin Concentrations in Populations of Escherichia Coli. Antimicrob. Agents Chemother. 2016, 60, 1336–1342. [Google Scholar] [CrossRef] [PubMed]
- Iakovides, I.C.; Kordatou, I.M.; Moreira, N.F.F.; Ribeiro, A.R.; Fernandes, T.; Pereira, M.F.R.; Nunes, O.C.; Manaia, C.M.; Silva, A.M.T.; Kassinos, D.F. Continuous Ozonation of Urban Wastewater: Removal of Antibiotics, Antibiotic-Resistant Escherichia Coli and Antibiotic Resistance Genes and Phytotoxicity. Water Res. 2019, 159, 333–347. [Google Scholar] [CrossRef] [PubMed]
- Dass, S.C.; Bosilevac, J.M.; Weinroth, M.; Elowsky, C.G.; Zhou, Y.; Anandappa, A.; Wang, R. Impact of Mixed Biofilm Formation with Environmental Microorganisms on E. Coli O157:H7 Survival against Sanitization. NPJ Sci. Food 2020, 4, 16. [Google Scholar] [CrossRef]
- Khan, S.; Imran, A.; Malik, A.; Chaudhary, A.A.; Rub, A.; Jan, A.T.; Syed, J.B.; Rolfo, C. Bacterial Imbalance and Gut Pathologies: Association and Contribution of E. Coli in Inflammatory Bowel Disease. Crit. Rev. Clin. Lab. Sci. 2019, 56, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Danson, A.E.; McStea, A.; Wang, L.; Pollitt, A.Y.; Fernandez, M.L.M.; Moraes, I.; Walsh, M.A.; MacIntyre, S.; Watson, K.A. Super-Resolution Fluorescence Microscopy Reveals Clustering Behaviour of Chlamydia Pneumoniae’s Major Outer Membrane Protein. Biology 2020, 9, 344. [Google Scholar] [CrossRef]
- Pereira, R.V.; Altier, C.; Siler, J.D.; Mann, S.; Jordan, D.; Warnick, L.D. Longitudinal Effects of Enrofloxacin or Tulathromycin Use in Preweaned Calves at High Risk of Bovine Respiratory Disease on the Shedding of Antimicrobial-Resistant Fecal Escherichia Coli. J. Dairy Sci. 2020, 103, 10547–10559. [Google Scholar] [CrossRef]
- Ellis, S.J.; Crossman, L.C.; McGrath, C.J.; Chattaway, M.A.; Hölken, J.M.; Brett, B.; Bundy, L.; Kay, G.L.; Wain, J.; Schüller, S. Identification and Characterisation of Enteroaggregative Escherichia Coli Subtypes Associated with Human Disease. Sci. Rep. 2020, 10, 7475. [Google Scholar] [CrossRef]
- Kwong, L.H.; Ercumen, A.; Pickering, A.J.; Arsenault, J.E.; Islam, M.; Parvez, S.M.; Unicomb, L.; Rahman, M.; Davis, J.; Luby, S.P. Ingestion of Fecal Bacteria along Multiple Pathways by Young Children in Rural Bangladesh Participating in a Cluster-Randomized Trial of Water, Sanitation, and Hygiene Interventions (WASH Benefits). Environ. Sci. Technol. 2020, 54, 13828–13838. [Google Scholar] [CrossRef]
- Lauridsen, H.C.M.; Vallance, B.A.; Krogfelt, K.A.; Petersen, A.M. Escherichia Coli Pathobionts Associated with Inflammatory Bowel Disease. Clin. Microbiol. Rev. 2019, 32, e00060-18. [Google Scholar] [CrossRef]
- Lopes, J.G.; Sourjik, V. Chemotaxis of Escherichia Coli to Major Hormones and Polyamines Present in Human Gut. ISME J. 2018, 12, 2736–2747. [Google Scholar] [CrossRef] [PubMed]
- Poirel, L.; Madec, J.-Y.; Lupo, A.; Schink, A.-K.; Kieffer, N.; Nordmann, P.; Schwarz, S. Antimicrobial Resistance in Escherichia coli. In Antimicrobial Resistance in Bacteria from Livestock and Companion Animals; Schwarz, S., Cavaco, L.M., Shen, J., Eds.; ASM Press: Washington, DC, USA, 2018; pp. 289–316. ISBN 978-1-68367-052-0. [Google Scholar]
- Card, R.M.; Cawthraw, S.A.; Garcia, J.N.; Ellis, R.J.; Kay, G.; Pallen, M.J.; Woodward, M.J.; Anjum, M.F. An In Vitro Chicken Gut Model Demonstrates Transfer of a Multidrug Resistance Plasmid from Salmonella to Commensal Escherichia Coli. mBio 2017, 8, e00777-17. [Google Scholar] [CrossRef] [PubMed]
- Ramiro, R.S.; Durão, P.; Bank, C.; Gordo, I. Low Mutational Load and High Mutation Rate Variation in Gut Commensal Bacteria. PLoS Biol. 2020, 18, e3000617. [Google Scholar] [CrossRef] [PubMed]
- Villa, E.B.; de la Peña, C.F.M.; Zaraín, P.L.; Cevallos, M.A.; Torres, C.; Torres, A.G.; Gracia, R.d.C.R. Comparative Genomics of a Subset of Adherent/Invasive Escherichia Coli Strains Isolated from Individuals without Inflammatory Bowel Disease. Genomics 2020, 112, 1813–1820. [Google Scholar] [CrossRef] [PubMed]
- Papouskova, A.; Masarikova, M.; Valcek, A.; Senk, D.; Cejkova, D.; Jahodarova, E.; Cizek, A. Genomic Analysis of Escherichia Coli Strains Isolated from Diseased Chicken in the Czech Republic. BMC Vet. Res. 2020, 16, 189. [Google Scholar] [CrossRef]
- Aslam, B.; Wang, W.; Arshad, M.I.; Khurshid, M.; Muzammil, S.; Rasool, M.H.; Nisar, M.A.; Alvi, R.F.; Aslam, M.A.; Qamar, M.U.; et al. Antibiotic Resistance: A Rundown of a Global Crisis. Infect. Drug Resist. 2018, 11, 1645–1658. [Google Scholar] [CrossRef]
- Mihankhah, A.; Khoshbakht, R.; Raeisi, M.; Raeisi, V. Prevalence and Antibiotic Resistance Pattern of Bacteria Isolated from Urinary Tract Infections in Northern Iran. J. Res. Med. Sci. 2017, 22, 108. [Google Scholar] [CrossRef]
- Millan, A.S. Evolution of Plasmid-Mediated Antibiotic Resistance in the Clinical Context. Trends Microbiol. 2018, 26, 978–985. [Google Scholar] [CrossRef]
- Ljubojević, D.; Velhner, M.; Todorović, D.; Pajić, M.; Milanov, D. Tetracycline Resistance in Escherichia Coli Isolates from Poultry. Arch. Vet. Med. 2016, 9, 61–81. [Google Scholar] [CrossRef]
- Puvača, N.; Lika, E.; Tufarelli, V.; Bursić, V.; Ljubojević Pelić, D.; Nikolova, N.; Petrović, A.; Prodanović, R.; Vuković, G.; Lević, J.; et al. Influence of Different Tetracycline Antimicrobial Therapy of Mycoplasma (Mycoplasma Synoviae) in Laying Hens Compared to Tea Tree Essential Oil on Table Egg Quality and Antibiotics Residues. Foods 2020, 9, 612. [Google Scholar] [CrossRef]
- Abdelhalim, K.A.; Uzel, A.; Ünal, N.G. Virulence Determinants and Genetic Diversity of Adherent-Invasive Escherichia Coli (AIEC) Strains Isolated from Patients with Crohn’s Disease. Microb. Pathog. 2020, 145, 104233. [Google Scholar] [CrossRef] [PubMed]
- Kilani, H.; Ferjani, S.; Mansouri, R.; Benboubaker, I.B.; Abbassi, M.S. Occurrence of Plasmid-Mediated Quinolone Resistance Determinants among Escherichia Coli Strains Isolated from Animals in Tunisia: Specific Pathovars Acquired Qnr Genes. J. Glob. Antimicrob. Resist. 2020, 20, 50–55. [Google Scholar] [CrossRef] [PubMed]
- Yu, Z.N.; Wang, J.; Ho, H.; Wang, Y.T.; Huang, S.N.; Han, R.W. Prevalence and Antimicrobial-Resistance Phenotypes and Genotypes of Escherichia Coli Isolated from Raw Milk Samples from Mastitis Cases in Four Regions of China. J. Glob. Antimicrob. Resist. 2020, 22, 94–101. [Google Scholar] [CrossRef] [PubMed]
- Farahat, E.M.; Hassuna, N.A.; Hammad, A.M.; Abdel Fattah, M.; Khairalla, A.S. Distribution of Integrons and Phylogenetic Groups among Escherichia Coli Causing Community-acquired Urinary Tract Infection in Upper Egypt. Can. J. Microbiol. 2020, cjm-2020-0292. [Google Scholar] [CrossRef] [PubMed]
- Ramay, B.M.; Caudell, M.A.; Rosales, C.C.; Archila, L.D.; Palmer, G.H.; Jarquin, C.; Moreno, P.; McCracken, J.P.; Rosenkrantz, L.; Amram, O.; et al. Antibiotic Use and Hygiene Interact to Influence the Distribution of Antimicrobial-Resistant Bacteria in Low-Income Communities in Guatemala. Sci. Rep. 2020, 10, 13767. [Google Scholar] [CrossRef]
- Farahani, O.; Ranjbar, R.; Jahromy, S.H.; Arabzadeh, B. Multilocus Variable-Number Tandem-Repeat Analysis for Genotyping of Escherichia Coli Strains Isolated from Hospital Wastewater, Tehran, Iran. Iran J. Public Health 2020, 49, 4829. [Google Scholar] [CrossRef]
- Riquelme, F.M.; Hernández, E.C.; Soto, M.G.; Ruiz, M.E.; Marí, J.M.N.; Fernández, J.G. Clinical Relevance of Antibiotic Susceptibility Profiles for Screening Gram-Negative Microorganisms Resistant to Beta-Lactam Antibiotics. Microorganisms 2020, 8, 1555. [Google Scholar] [CrossRef]
- Kapoor, G.; Saigal, S.; Elongavan, A. Action and Resistance Mechanisms of Antibiotics: A Guide for Clinicians. J. Anaesthesiol. Clin. Pharmacol. 2017, 33, 300–305. [Google Scholar] [CrossRef]
- Rahman, S.U.; Ali, T.; Ali, I.; Khan, N.A.; Han, B.; Gao, J. The Growing Genetic and Functional Diversity of Extended Spectrum Beta-Lactamases. BioMed Res. Int. 2018, 2018, 9519718. [Google Scholar] [CrossRef]
- Doma, A.O.; Popescu, R.; Mitulețu, M.; Muntean, D.; Dégi, J.; Boldea, M.V.; Radulov, I.; Dumitrescu, E.; Muselin, F.; Puvača, N.; et al. Comparative Evaluation of QnrA, QnrB, and QnrS Genes in Enterobacteriaceae Ciprofloxacin-Resistant Cases, in Swine Units and a Hospital from Western Romania. Antibiotics 2020, 9, 698. [Google Scholar] [CrossRef]
- Rilo, M.P.; Martín, C.-B.G.; Fernández, E.P.; Vilaró, A.; Fraile, L.; Martínez, S.M. Antimicrobial Resistance Genes in Porcine Pasteurella Multocida Are Not Associated with Its Antimicrobial Susceptibility Pattern. Antibiotics 2020, 9, 614. [Google Scholar] [CrossRef] [PubMed]
- Torban, A.S.; Venezia, S.N.; Kelmer, E.; Cohen, A.; Paitan, Y.; Arielly, H.; Steinman, A. Extended-Spectrum β-Lactamase-Producing Enterobacterles Shedding by Dogs and Cats Hospitalized in an Emergency and Critical Care Department of a Veterinary Teaching Hospital. Antibiotics 2020, 9, 545. [Google Scholar] [CrossRef] [PubMed]
- Falgenhauer, L.; Schwengers, O.; Schmiedel, J.; Baars, C.; Lambrecht, O.; Heß, S.; Berendonk, T.U.; Falgenhauer, J.; Chakraborty, T.; Imirzalioglu, C. Multidrug-Resistant and Clinically Relevant Gram-Negative Bacteria Are Present in German Surface Waters. Front. Microbiol. 2019, 10, 2779. [Google Scholar] [CrossRef] [PubMed]
- Santos, A.C.d.M.; Santos, F.F.; Silva, R.M.; Gomes, T.A.T. Diversity of Hybrid- and Hetero-Pathogenic Escherichia Coli and Their Potential Implication in More Severe Diseases. Front. Cell Infect. Microbiol. 2020, 10, 339. [Google Scholar] [CrossRef] [PubMed]
- Qiu, W.; Sun, J.; Fang, M.; Luo, S.; Tian, Y.; Dong, P.; Xu, B.; Zheng, C. Occurrence of Antibiotics in the Main Rivers of Shenzhen, China: Association with Antibiotic Resistance Genes and Microbial Community. Sci. Total Environ. 2019, 653, 334–341. [Google Scholar] [CrossRef]
- Sanganyado, E.; Gwenzi, W. Antibiotic Resistance in Drinking Water Systems: Occurrence, Removal, and Human Health Risks. Sci. Total Environ. 2019, 669, 785–797. [Google Scholar] [CrossRef]
- Mölstad, S.; Löfmark, S.; Carlin, K.; Erntell, M.; Aspevall, O.; Blad, L.; Hanberger, H.; Hedin, K.; Hellman, J.; Norman, C.; et al. Lessons Learnt during 20 Years of the Swedish Strategic Programme against Antibiotic Resistance. Bull. World Health Organ. 2017, 95, 764–773. [Google Scholar] [CrossRef]
- Osińska, A.; Korzeniewska, E.; Harnisz, M.; Niestępski, S. The Prevalence and Characterization of Antibiotic-Resistant and Virulent Escherichia Coli Strains in the Municipal Wastewater System and Their Environmental Fate. Sci. Total Environ. 2017, 577, 367–375. [Google Scholar] [CrossRef]
- Montealegre, M.C.; Roy, S.; Böni, F.; Hossain, M.I.; Daneshmand, T.N.; Caduff, L.; Faruque, A.S.G.; Islam, M.A.; Julian, T.R. Risk Factors for Detection, Survival, and Growth of Antibiotic-Resistant and Pathogenic Escherichia Coli in Household Soils in Rural Bangladesh. Appl. Environ. Microbiol. 2018, 84, e01978-18. [Google Scholar] [CrossRef]
- Kaesbohrer, A.; Lebl, K.B.; Irrgang, A.; Fischer, J.; Kämpf, P.; Schiffmann, A.; Werckenthin, C.; Busch, M.; Kreienbrock, L.; Hille, K. Diversity in Prevalence and Characteristics of ESBL/PAmpC Producing E. Coli in Food in Germany. Vet. Microbiol. 2019, 233, 52–60. [Google Scholar] [CrossRef]
- Marano, R.B.M.; Fernandes, T.; Manaia, C.M.; Nunes, O.; Morrison, D.; Berendonk, T.U.; Kreuzinger, N.; Tenson, T.; Corno, G.; Kassinos, D.F.; et al. A Global Multinational Survey of Cefotaxime-Resistant Coliforms in Urban Wastewater Treatment Plants. Environ. Int. 2020, 144, 106035. [Google Scholar] [CrossRef] [PubMed]
- Gardy, J.L.; Loman, N.J. Towards a Genomics-Informed, Real-Time, Global Pathogen Surveillance System. Nat. Rev. Genet 2018, 19, 9–20. [Google Scholar] [CrossRef] [PubMed]
- Ferran, A.A.; Lacroix, M.Z.; Mélou, A.B.; Duhil, I.; Roques, B.B. Levers to Improve Antibiotic Treatment of Lambs via Drinking Water in Sheep Fattening Houses: The Example of the Sulfadimethoxine/Trimethoprim Combination. Antibiotics 2020, 9, 561. [Google Scholar] [CrossRef] [PubMed]
- Vilaró, A.; Novell, E.; Tarancón, V.E.; Balielles, J.; Vilalta, C.; Martinez, S.; Fraile Sauce, L.J. Antimicrobial Susceptibility Pattern of Porcine Respiratory Bacteria in Spain. Antibiotics 2020, 9, 402. [Google Scholar] [CrossRef] [PubMed]
- Mileva, R.; Karadaev, M.; Fasulkov, I.; Petkova, T.; Rusenova, N.; Vasilev, N.; Milanova, A. Oxytetracycline Pharmacokinetics After Intramuscular Administration in Cows with Clinical Metritis Associated with Trueperella Pyogenes Infection. Antibiotics 2020, 9, 392. [Google Scholar] [CrossRef]
- Martens, E.; Demain, A.L. The Antibiotic Resistance Crisis, with a Focus on the United States. J. Antibiot. 2017, 70, 520–526. [Google Scholar] [CrossRef]
- Padmini, N.; Ajilda, A.A.K.; Sivakumar, N.; Selvakumar, G. Extended Spectrum β-Lactamase Producing Escherichia Coli and Klebsiella Pneumoniae: Critical Tools for Antibiotic Resistance Pattern. J. Basic Microbiol. 2017, 57, 460–470. [Google Scholar] [CrossRef]
- Sharaha, U.; Diaz, E.R.; Riesenberg, K.; Bigio, I.J.; Huleihel, M.; Salman, A. Using Infrared Spectroscopy and Multivariate Analysis to Detect Antibiotics’ Resistant Escherichia Coli Bacteria. Anal. Chem. 2017, 89, 8782–8790. [Google Scholar] [CrossRef]
- Moradigaravand, D.; Palm, M.; Farewell, A.; Mustonen, V.; Warringer, J.; Parts, L. Prediction of Antibiotic Resistance in Escherichia Coli from Large-Scale Pan-Genome Data. PLoS Comput. Biol. 2018, 14, e1006258. [Google Scholar] [CrossRef]
- Lukačišinová, M.; Fernando, B.; Bollenbach, T. Highly Parallel Lab Evolution Reveals That Epistasis Can Curb the Evolution of Antibiotic Resistance. Nat. Commun. 2020, 11, 3105. [Google Scholar] [CrossRef]
- Swain, S.S.; Paidesetty, S.K.; Padhy, R.N. Phytochemical Conjugation as a Potential Semisynthetic Approach toward Reactive and Reuse of Obsolete Sulfonamides against Pathogenic Bacteria. Drug Dev. Res. 2020, ddr.21746. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. WHO Report on Surveillance of Antibiotic Consumption: 2016–2018 Early Implementation; World Health Organization: Geneva, Switzerland, 2018; p. 128. [Google Scholar]
- Colson, A.R.; Megiddo, I.; Uria, G.A.; Gandra, S.; Bedford, T.; Morton, A.; Cooke, R.M.; Laxminarayan, R. Quantifying Uncertainty about Future Antimicrobial Resistance: Comparing Structured Expert Judgment and Statistical Forecasting Methods. PLoS ONE 2019, 14, e0219190. [Google Scholar] [CrossRef] [PubMed]
- Sartelli, M.C.; Hardcastle, T.; Catena, F.; Mefire, A.C.; Coccolini, F.; Dhingra, S.; Haque, M.; Hodonou, A.; Iskandar, K.; Labricciosa, F.M.; et al. Antibiotic Use in Low and Middle-Income Countries and the Challenges of Antimicrobial Resistance in Surgery. Antibiotics 2020, 9, 497. [Google Scholar] [CrossRef]
- Hsia, Y.; Sharland, M.; Jackson, C.; Wong, I.C.K.; Magrini, N.; Bielicki, J.A. Consumption of Oral Antibiotic Formulations for Young Children According to the WHO Access, Watch, Reserve (AWaRe) Antibiotic Groups: An Analysis of Sales Data from 70 Middle-Income and High-Income Countries. Lancet Infect. Dis. 2019, 19, 67–75. [Google Scholar] [CrossRef]
- Yan, W.; Xiao, Y.; Yan, W.; Ding, R.; Wang, S.; Zhao, F. The Effect of Bioelectrochemical Systems on Antibiotics Removal and Antibiotic Resistance Genes: A Review. Chem. Eng. J. 2019, 358, 1421–1437. [Google Scholar] [CrossRef]
- Xie, J.; Jin, L.; He, T.; Chen, B.; Luo, X.; Feng, B.; Huang, W.; Li, J.; Fu, P.; Li, X. Bacteria and Antibiotic Resistance Genes (ARGs) in PM 2.5 from China: Implications for Human Exposure. Environ. Sci. Technol. 2019, 53, 963–972. [Google Scholar] [CrossRef]
- Palme, J.B.; Kristiansson, E.; Larsson, D.G.J. Environmental Factors Influencing the Development and Spread of Antibiotic Resistance. FEMS Microbiol. Rev. 2018, 42, fux053. [Google Scholar] [CrossRef]
- Merrikh, H.; Kohli, R.M. Targeting Evolution to Inhibit Antibiotic Resistance. FEBS J. 2020, 287, 4341–4353. [Google Scholar] [CrossRef]
- Marine, J.-C.; Dawson, S.-J.; Dawson, M.A. Non-Genetic Mechanisms of Therapeutic Resistance in Cancer. Nat. Rev. Cancer 2020, 20, 743–756. [Google Scholar] [CrossRef]
- Heir, E.; Møretrø, T.; Simensen, A.; Langsrud, S. Listeria Monocytogenes Strains Show Large Variations in Competitive Growth in Mixed Culture Biofilms and Suspensions with Bacteria from Food Processing Environments. Int. J. Food Microbiol. 2018, 275, 46–55. [Google Scholar] [CrossRef]
- Stubbendieck, R.M.; May, D.S.; Chevrette, M.G.; Temkin, M.I.; Pienkowski, E.W.; Cagnazzo, J.; Carlson, C.M.; Gern, J.E.; Currie, C.R. Competition among Nasal Bacteria Suggests a Role for Siderophore-Mediated Interactions in Shaping the Human Nasal Microbiota. Appl. Environ. Microbiol. 2018, 85, e02406-18. [Google Scholar] [CrossRef] [PubMed]
- de Filippis, F.; Pasolli, E.; Ercolini, D. The Food-Gut Axis: Lactic Acid Bacteria and Their Link to Food, the Gut Microbiome and Human Health. FEMS Microbiol. Rev. 2020, 44, 454–489. [Google Scholar] [CrossRef] [PubMed]
- Iwu, C.D.; Korsten, L.; Okoh, A.I. The Incidence of Antibiotic Resistance within and beyond the Agricultural Ecosystem: A Concern for Public Health. Microbiol. Open 2020, 9, e1035. [Google Scholar] [CrossRef] [PubMed]
- Calap, P.D.; Martínez, J.D. Bacteriophages: Protagonists of a Post-Antibiotic Era. Antibiotics 2018, 7, 66. [Google Scholar] [CrossRef]
- Reygaert, W.C. An Overview of the Antimicrobial Resistance Mechanisms of Bacteria. AIMS Microbiol. 2018, 4, 482–501. [Google Scholar] [CrossRef]
- Wang, Y.; Alenazy, R.; Gu, X.; Polyak, S.W.; Zhang, P.; Sykes, M.J.; Zhang, N.; Venter, H.; Ma, S. Design and Structural Optimization of Novel 2H-Benzo[h]Chromene Derivatives That Target AcrB and Reverse Bacterial Multidrug Resistance. Eur. J. Med. Chem. 2020, 113049. [Google Scholar] [CrossRef]
- Impey, R.E.; Hawkins, D.A.; Sutton, J.M.; da Costa, T.P.S. Overcoming Intrinsic and Acquired Resistance Mechanisms Associated with the Cell Wall of Gram-Negative Bacteria. Antibiotics 2020, 9, 623. [Google Scholar] [CrossRef]
- Iskandar, K.; Molinier, L.; Hallit, S.; Sartelli, M.; Catena, F.; Coccolini, F.; Craig Hardcastle, T.; Roques, C.; Salameh, P. Drivers of Antibiotic Resistance Transmission in Low- and Middle-Income Countries from a “One Health” Perspective—A Review. Antibiotics 2020, 9, 372. [Google Scholar] [CrossRef]
- Dsani, E.; Afari, E.A.; Appiah, A.D.; Kenu, E.; Kaburi, B.B.; Egyir, B. Antimicrobial Resistance and Molecular Detection of Extended Spectrum β-Lactamase Producing Escherichia Coli Isolates from Raw Meat in Greater Accra Region, Ghana. BMC Microbiol. 2020, 20, 253. [Google Scholar] [CrossRef]
- Hassan, J.; Eddine, R.Z.; Mann, D.; Li, S.; Deng, X.; Saoud, I.P.; Kassem, I.I. The Mobile Colistin Resistance Gene, Mcr-1.1, Is Carried on IncX4 Plasmids in Multidrug Resistant E. Coli Isolated from Rainbow Trout Aquaculture. Microorganisms 2020, 8, 1636. [Google Scholar] [CrossRef]
- Montealegre, M.C.; Rodríguez, A.T.; Roy, S.; Hossain, M.I.; Islam, M.A.; Lanza, V.F.; Julian, T.R. High Genomic Diversity and Heterogenous Origins of Pathogenic and Antibiotic-Resistant Escherichia Coli in Household Settings Represent a Challenge to Reducing Transmission in Low-Income Settings. mSphere 2020, 5, e00704-19. [Google Scholar] [CrossRef] [PubMed]
- Alexander, J.; Hembach, N.; Schwartz, T. Evaluation of Antibiotic Resistance Dissemination by Wastewater Treatment Plant Effluents with Different Catchment Areas in Germany. Sci. Rep. 2020, 10, 8952. [Google Scholar] [CrossRef] [PubMed]
- Pang, X.; Li, D.; Zhu, J.; Cheng, J.; Liu, G. Beyond Antibiotics: Photo/Sonodynamic Approaches for Bacterial Theranostics. Nano Micro Lett. 2020, 12, 144. [Google Scholar] [CrossRef]
- Durão, P.; Balbontín, R.; Gordo, I. Evolutionary Mechanisms Shaping the Maintenance of Antibiotic Resistance. Trends Microbiol. 2018, 26, 677–691. [Google Scholar] [CrossRef]
- Ivanov, I.I.; Frutos, R.d.L.; Manel, N.; Yoshinaga, K.; Rifkin, D.B.; Sartor, R.B.; Finlay, B.B.; Littman, D.R. Specific Microbiota Direct the Differentiation of IL-17-Producing T-Helper Cells in the Mucosa of the Small Intestine. Cell Host Microbe. 2008, 4, 337–349. [Google Scholar] [CrossRef]
- Lobanovska, M.; Pilla, G. Penicillin’s Discovery and Antibiotic Resistance: Lessons for the Future? Yale J. Biol. Med. 2017, 90, 135–145. [Google Scholar]
- Lartey, S.F.; Yee, M.; Gaarslev, C.; Khan, R. Why Do General Practitioners Prescribe Antibiotics for Upper Respiratory Tract Infections to Meet Patient Expectations: A Mixed Methods Study. BMJ Open 2016, 6, e012244. [Google Scholar] [CrossRef]
- Chui, C.S.L.; Cowling, B.J.; Lim, W.W.; Hui, C.K.M.; Chan, E.W.; Wong, I.C.K.; Wu, P. Patterns of Inpatient Antibiotic Use Among Public Hospitals in Hong Kong from 2000 to 2015. Drug Saf. 2020, 43, 595–606. [Google Scholar] [CrossRef]
- Osoro, A.A.; Atitwa, E.B.; Moturi, J.K. Universal Health Coverage. WJSSR 2020, 7, p14. [Google Scholar] [CrossRef]
- Tsao, L.-H.; Hsin, C.-Y.; Liu, H.-Y.; Chuang, H.-C.; Chen, L.-Y.; Lee, Y.-J. Risk Factors for Healthcare-Associated Infection Caused by Carbapenem-Resistant Pseudomonas Aeruginosa. J. Microbiol. Immunol. Infect. 2018, 51, 359–366. [Google Scholar] [CrossRef]
- Dhesi, Z.; Enne, V.I.; O’Grady, J.; Gant, V.; Livermore, D.M. Rapid and Point-of-Care Testing in Respiratory Tract Infections: An Antibiotic Guardian? ACS Pharmacol. Transl. Sci. 2020, 3, 401–417. [Google Scholar] [CrossRef] [PubMed]
- David, S.; Reuter, S.; Harris, S.R.; Glasner, C.; Feltwell, T.; Argimon, S.; Abudahab, K.; Goater, R.; Giani, T.; Errico, G.; et al. Epidemic of Carbapenem-Resistant Klebsiella Pneumoniae in Europe Is Driven by Nosocomial Spread. Nat. Microbiol. 2019, 4, 1919–1929. [Google Scholar] [CrossRef] [PubMed]
- Codjoe, F.; Donkor, E. Carbapenem Resistance: A Review. Med. Sci. 2017, 6, 1. [Google Scholar] [CrossRef]
- Pitout, J.D.D.; Peirano, G.; Kock, M.M.; Strydom, K.-A.; Matsumura, Y. The Global Ascendency of OXA-48-Type Carbapenemases. Clin. Microbiol. Rev. 2019, 33, e00102-19. [Google Scholar] [CrossRef] [PubMed]
- Paterson, D.L.; Bonomo, R.A. Multidrug-Resistant Gram-Negative Pathogens: The Urgent Need for ‘Old’ Polymyxins. In Polymyxin Antibiotics: From Laboratory Bench to Bedside; Li, J., Nation, R.L., Kaye, K.S., Eds.; Advances in Experimental Medicine and Biology; Springer International Publishing: Cham, The Netherlands, 2019; Volume 1145, pp. 9–13. ISBN 978-3-030-16371-6. [Google Scholar]
- Godman, B. Ongoing Initiatives to Improve the Prescribing of Medicines across Sectors and the Implications. Adv. Hum. Biol. 2020, 10, 85. [Google Scholar] [CrossRef]
- Mcoyi, S.; Amoako, D.G.; Somboro, A.M.; Khumalo, H.M.; Khan, R.B. The Molecular Effect of 1,4,7-triazacyclononane on Oxidative Stress Parameters in Human Hepatocellular Carcinoma (HepG2) Cells. J. Biochem. Mol. Toxicol. 2020, 34, e22607. [Google Scholar] [CrossRef] [PubMed]
- Singleton, D.A.; Pinchbeck, G.L.; Radford, A.D.; Arsevska, E.; Dawson, S.; Jones, P.H.; Noble, P.-J.M.; Williams, N.J.; Sánchez-Vizcaíno, F. Factors Associated with Prescription of Antimicrobial Drugs for Dogs and Cats, United Kingdom, 2014–2016. Emerg. Infect. Dis. 2020, 26, 1778–1791. [Google Scholar] [CrossRef]
- Robbins, S.N.; Goggs, R.; Lhermie, G.; Paul, D.F.L.; Menard, J. Antimicrobial Prescribing Practices in Small Animal Emergency and Critical Care. Front. Vet. Sci. 2020, 7, 110. [Google Scholar] [CrossRef]
- Lehner, C.; Hubbuch, A.; Schmitt, K.; Regula, G.S.; Willi, B.; Mevissen, M.; Peter, R.; Muentener, C.R.; Naegeli, H.; Schuller, S. Effect of Antimicrobial Stewardship on Antimicrobial Prescriptions for Selected Diseases of Dogs in Switzerland. J. Vet. Intern. Med. 2020, 34, 2418–2431. [Google Scholar] [CrossRef]
- Teoh, L.; Stewart, K.; Marino, R.J.; McCullough, M.J. Improvement of Dental Prescribing Practices Using Education and a Prescribing Tool: A Pilot Intervention Study. Br. J. Clin. Pharmacol. 2020, 4373. [Google Scholar] [CrossRef]
- Bansal, R.; Jain, A.; Goyal, M.; Singh, T.; Sood, H.; Malviya, H.S. Antibiotic Abuse during Endodontic Treatment: A Contributing Factor to Antibiotic Resistance. J. Fam. Med. Prim Care 2019, 8, 3518–3524. [Google Scholar] [CrossRef]
- Nadeem, S.F.; Gohar, U.F.; Tahir, S.F.; Mukhtar, H.; Pornpukdeewattana, S.; Nukthamna, P.; Moula Ali, A.M.; Bavisetty, S.C.B.; Massa, S. Antimicrobial Resistance: More than 70 Years of War between Humans and Bacteria. Crit. Rev. Microbiol. 2020, 46, 578–599. [Google Scholar] [CrossRef] [PubMed]
- Podolsky, S.H. The Evolving Response to Antibiotic Resistance (1945–2018). Palgrave Commun. 2018, 4, 124. [Google Scholar] [CrossRef]
- Choez, X.S.; Acurio, M.L.A.; Sotomayor, R.E.J. Appropriateness and Adequacy of Antibiotic Prescription for Upper Respiratory Tract Infections in Ambulatory Health Care Centers in Ecuador. BMC Pharmacol. Toxicol. 2018, 19, 46. [Google Scholar] [CrossRef]
- Brink, A.J.; van Wyk, J.; Moodley, V.M.; Corcoran, C.; Ekermans, P.; Nutt, L.; Boyles, T.; Perovic, O.; Feldman, C.; Richards, G.; et al. The Role of Appropriate Diagnostic Testing in Acute Respiratory Tract Infections: An Antibiotic Stewardship Strategy to Minimise Diagnostic Uncertainty in Primary Care. S. Afr. Med. J. 2016, 106, 554. [Google Scholar] [CrossRef] [PubMed]
- Teratani, Y.; Hagiya, H.; Koyama, T.; Adachi, M.; Ohshima, A.; Zamami, Y.; Tanaka, H.Y.; Tatebe, Y.; Tasaka, K.; Mikami, N.; et al. Pattern of Antibiotic Prescriptions for Outpatients with Acute Respiratory Tract Infections in Japan, 2013–15: A Retrospective Observational Study. Fam. Pract. 2019, 36, 402–409. [Google Scholar] [CrossRef]
- Stuart, B.; Brotherwood, H.; van’t Hoff, C.; Brown, A.; van den Bruel, A.; Hay, A.D.; Moore, M.; Little, P. Exploring the Appropriateness of Antibiotic Prescribing for Common Respiratory Tract Infections in UK Primary Care. J. Antimicrob. Chemother. 2019, 75, dkz410. [Google Scholar] [CrossRef]
- Pouwels, K.B.; Pebody, B.M.; Smieszek, T.; Hopkins, S.; Robotham, J.V. Selection and Co-Selection of Antibiotic Resistances among Escherichia Coli by Antibiotic Use in Primary Care: An Ecological Analysis. PLoS ONE 2019, 14, e0218134. [Google Scholar] [CrossRef]
- Shively, N.R.; Buehrle, D.J.; Clancy, C.J.; Decker, B.K. Prevalence of Inappropriate Antibiotic Prescribing in Primary Care Clinics within a Veterans Affairs Health Care System. Antimicrob. Agents Chemother. 2018, 62, e00337-18. [Google Scholar] [CrossRef]
- Roess, A.; Leibler, J.H.; Graham, J.P.; Lowenstein, C.; Waters, W.F. Animal Husbandry Practices and Perceptions of Zoonotic Infectious Disease Risks Among Livestock Keepers in a Rural Parish of Quito, Ecuador. Am. J. Trop. Med. Hyg. 2016, 95, 1450–1458. [Google Scholar] [CrossRef]
- Riddle, M.S.; Connor, B.A.; Beeching, N.J.; DuPont, H.L.; Hamer, D.H.; Kozarsky, P.; Libman, M.; Steffen, R.; Taylor, D.; Tribble, D.R.; et al. Guidelines for the Prevention and Treatment of Travelers’ Diarrhea: A Graded Expert Panel Report. J. Travel Med. 2017, 24, S63–S80. [Google Scholar] [CrossRef] [PubMed]
- Giallourou, N.; Medlock, G.L.; Bolick, D.T.; Medeiros, P.H.; Ledwaba, S.E.; Kolling, G.L.; Tung, K.; Guerry, P.; Swann, J.R.; Guerrant, R.L. A Novel Mouse Model of Campylobacter Jejuni Enteropathy and Diarrhea. PLoS Pathog. 2018, 14, e1007083. [Google Scholar] [CrossRef] [PubMed]
- Vergalli, J.; Bodrenko, I.V.; Masi, M.; Moynié, L.; Gutiérrez, S.A.; Naismith, J.H.; Regli, A.D.; Ceccarelli, M.; van den Berg, B.; Winterhalter, M.; et al. Porins and Small-Molecule Translocation across the Outer Membrane of Gram-Negative Bacteria. Nat. Rev. Microbiol. 2020, 18, 164–176. [Google Scholar] [CrossRef] [PubMed]
- Arzanlou, M.; Chai, W.C.; Venter, H. Intrinsic, Adaptive and Acquired Antimicrobial Resistance in Gram-Negative Bacteria. Essays Biochem. 2017, 61, 49–59. [Google Scholar] [CrossRef] [PubMed]
- Dunn, S.J.; Connor, C.; McNally, A. The Evolution and Transmission of Multi-Drug Resistant Escherichia Coli and Klebsiella Pneumoniae: The Complexity of Clones and Plasmids. Curr. Opin. Microbiol. 2019, 51, 51–56. [Google Scholar] [CrossRef]
- Kayastha, K.; Dhungel, B.; Karki, S.; Adhikari, B.; Banjara, M.R.; Rijal, K.R.; Ghimire, P. Extended-Spectrum β-Lactamase-Producing Escherichia Coli and Klebsiella Species in Pediatric Patients Visiting International Friendship Children’s Hospital, Kathmandu, Nepal. Infect. Dis. (Auckl.) 2020, 13, 117863372090979. [Google Scholar] [CrossRef]
- Feria, C. Patterns and Mechanisms of Resistance to Beta-Lactams and Beta-Lactamase Inhibitors in Uropathogenic Escherichia Coli Isolated from Dogs in Portugal. J. Antimicrob. Chemother. 2002, 49, 77–85. [Google Scholar] [CrossRef]
- Liu, J.; Huang, Z.; Ruan, B.; Wang, H.; Chen, M.; Rehman, S.; Wu, P. Quantitative Proteomic Analysis Reveals the Mechanisms of Polymyxin B Toxicity to Escherichia Coli. Chemosphere 2020, 259, 127449. [Google Scholar] [CrossRef]
- Sharifzadeh, S.; Dempwolff, F.; Kearns, D.B.; Carlson, E.E. Harnessing β-Lactam Antibiotics for Illumination of the Activity of Penicillin-Binding Proteins in Bacillus Subtilis. ACS Chem. Biol. 2020, 15, 1242–1251. [Google Scholar] [CrossRef]
- Decuyper, L.; Jukič, M.; Sosič, I.; Žula, A.; D’hooghe, M.; Gobec, S. Antibacterial and β-Lactamase Inhibitory Activity of Monocyclic β-Lactams. Med. Res. Rev. 2018, 38, 426–503. [Google Scholar] [CrossRef]
- Hameed, A.S.H.; Louis, G.; Karthikeyan, C.; Thajuddin, N.; Ravi, G. Impact of L-Arginine and l-Histidine on the Structural, Optical and Antibacterial Properties of Mg Doped ZnO Nanoparticles Tested against Extended-Spectrum Beta-Lactamases (ESBLs) Producing Escherichia Coli. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2019, 211, 373–382. [Google Scholar] [CrossRef] [PubMed]
- Andersson, D.I.; Balaban, N.Q.; Baquero, F.; Courvalin, P.; Glaser, P.; Gophna, U.; Kishony, R.; Molin, S.; Tønjum, T. Antibiotic Resistance: Turning Evolutionary Principles into Clinical Reality. FEMS Microbiol. Rev. 2020, 44, 171–188. [Google Scholar] [CrossRef] [PubMed]
- Bush, K. Past and Present Perspectives on β-Lactamases. Antimicrob. Agents Chemother. 2018, 62, e01076-18. [Google Scholar] [CrossRef] [PubMed]
- Silveira, M.C.; da Silva, R.A.; da Mota, F.F.; Catanho, M.; Jardim, R.; Guimarães, A.C.R.; de Miranda, A.B. Systematic Identification and Classification of β-Lactamases Based on Sequence Similarity Criteria: β-Lactamase Annotation. Evol. Bioinform. Online 2018, 14, 117693431879735. [Google Scholar] [CrossRef] [PubMed]
- Both, A.; Huang, J.; Kaase, M.; Hezel, J.; Wertheimer, D.; Fenner, I.; Günther, T.; Grundhoff, A.; Büttner, H.; Aepfelbacher, M.; et al. First Report of Escherichia Coli Co-Producing NDM-1 and OXA-232. Diagn. Microbiol. Infect. Dis. 2016, 86, 437–438. [Google Scholar] [CrossRef]
- Montso, K.P.; Dlamini, S.B.; Kumar, A.; Ateba, C.N. Antimicrobial Resistance Factors of Extended-Spectrum Beta-Lactamases Producing Escherichia Coli and Klebsiella Pneumoniae Isolated from Cattle Farms and Raw Beef in North-West Province, South Africa. BioMed Res. Int. 2019, 2019, 4318306. [Google Scholar] [CrossRef]
- Aguirre, L.; Vidal, A.; Seminati, C.; Tello, M.; Redondo, N.; Darwich, L.; Martín, M. Antimicrobial Resistance Profile and Prevalence of Extended-Spectrum Beta-Lactamases (ESBL), AmpC Beta-Lactamases and Colistin Resistance (Mcr) Genes in Escherichia Coli from Swine between 1999 and 2018. Porc. Health Manag. 2020, 6, 8. [Google Scholar] [CrossRef]
- Pormohammad, A.; Nasiri, M.J.; Azimi, T. Prevalence of Antibiotic Resistance in Escherichia Coli Strains Simultaneously Isolated from Humans, Animals, Food, and the Environment: A Systematic Review and Meta-Analysis. Infect. Drug Resist. 2019, 12, 1181–1197. [Google Scholar] [CrossRef]
| Country | DDD/1000 Inhabitants Per Day | Country | DDD/1000 Inhabitants Per Day |
|---|---|---|---|
| Albania | 16.41 | Kosovo | 20.18 |
| Armenia | 10.31 | Kyrgyzstan | 17.94 |
| Austria | 12.17 | Latvia | 13.30 |
| Azerbaijan | 7.66 | Lithuania | 15.83 |
| Belarus | 17.48 | Luxemburg | 22.31 |
| Belgium | 25.57 | Malta | 21.88 |
| Bosnia and Herzegovina | 17.85 | Montenegro | 29.33 |
| Bulgaria | 20.25 | Netherlands | 9.78 |
| Croatia | 20.28 | Norway | 16.97 |
| Cyprus | 27.14 | Poland | 24.30 |
| Czech Republic | 17.18 | Portugal | 17.72 |
| Denmark | 17.84 | North Macedonia | 13.42 |
| Estonia | 12.13 | Romania | 28.50 |
| Finland | 18.52 | Russia | 14.82 |
| France | 25.92 | Serbia | 31.57 |
| Georgia | 24.44 | Slovakia | 24.34 |
| Germany | 11.49 | Slovenia | 13.48 |
| Greece | 33.85 | Spain | 17.96 |
| Hungary | 16.31 | Sweden | 13.73 |
| Iceland | 17.87 | Tajikistan | 21.95 |
| Ireland | 23.27 | Turkey | 38.18 |
| Italy | 26.62 | United Kingdom | 20.47 |
| Kazakhstan | 17.89 | Uzbekistan | 8.56 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Puvača, N.; de Llanos Frutos, R. Antimicrobial Resistance in Escherichia coli Strains Isolated from Humans and Pet Animals. Antibiotics 2021, 10, 69. https://doi.org/10.3390/antibiotics10010069
Puvača N, de Llanos Frutos R. Antimicrobial Resistance in Escherichia coli Strains Isolated from Humans and Pet Animals. Antibiotics. 2021; 10(1):69. https://doi.org/10.3390/antibiotics10010069
Chicago/Turabian StylePuvača, Nikola, and Rosa de Llanos Frutos. 2021. "Antimicrobial Resistance in Escherichia coli Strains Isolated from Humans and Pet Animals" Antibiotics 10, no. 1: 69. https://doi.org/10.3390/antibiotics10010069
APA StylePuvača, N., & de Llanos Frutos, R. (2021). Antimicrobial Resistance in Escherichia coli Strains Isolated from Humans and Pet Animals. Antibiotics, 10(1), 69. https://doi.org/10.3390/antibiotics10010069