SAM Composition and Electrode Roughness Affect Performance of a DNA Biosensor for Antibiotic Resistance
Abstract
:1. Introduction
2. Materials and Methods
3. Results and Discussion
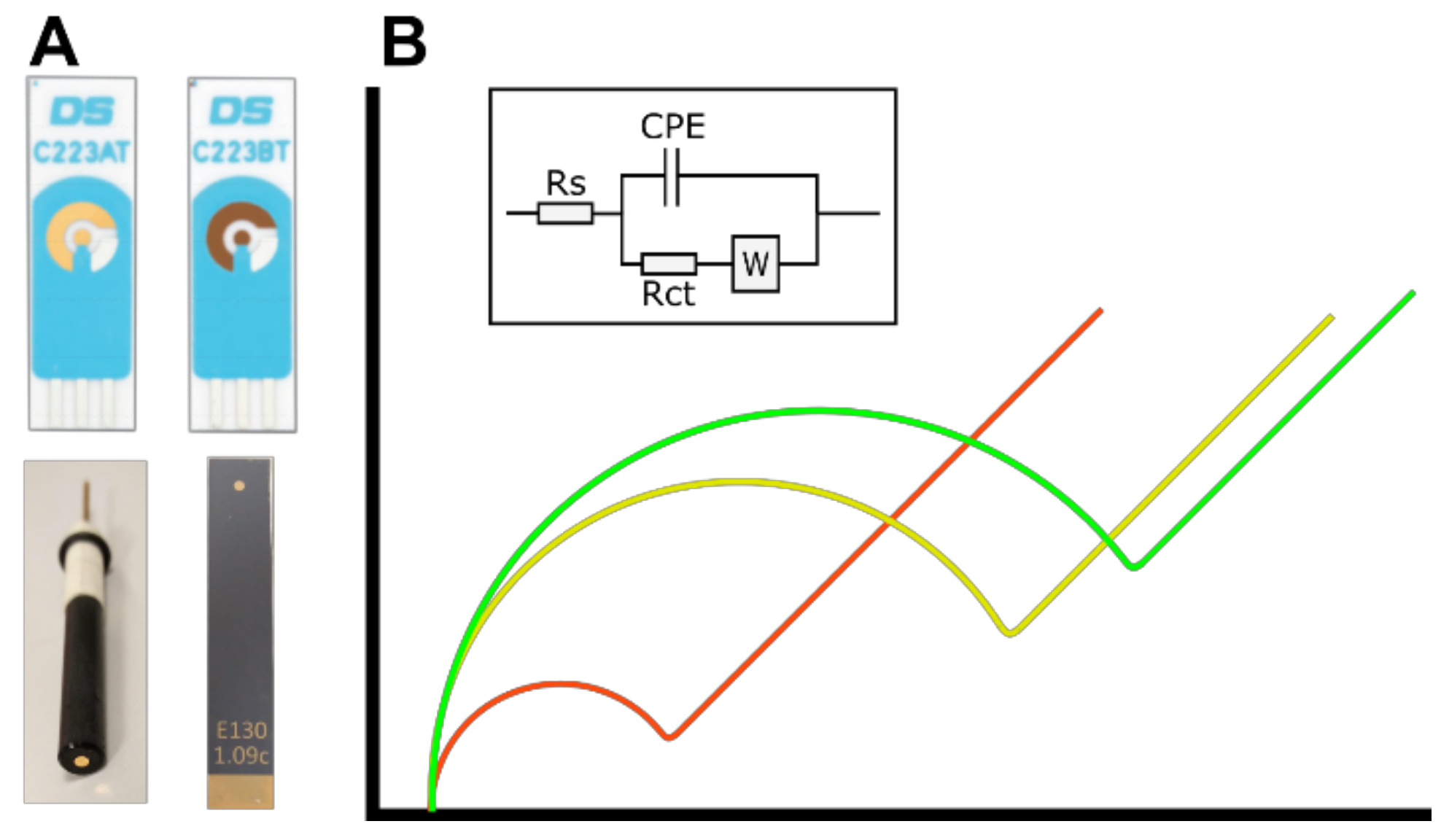
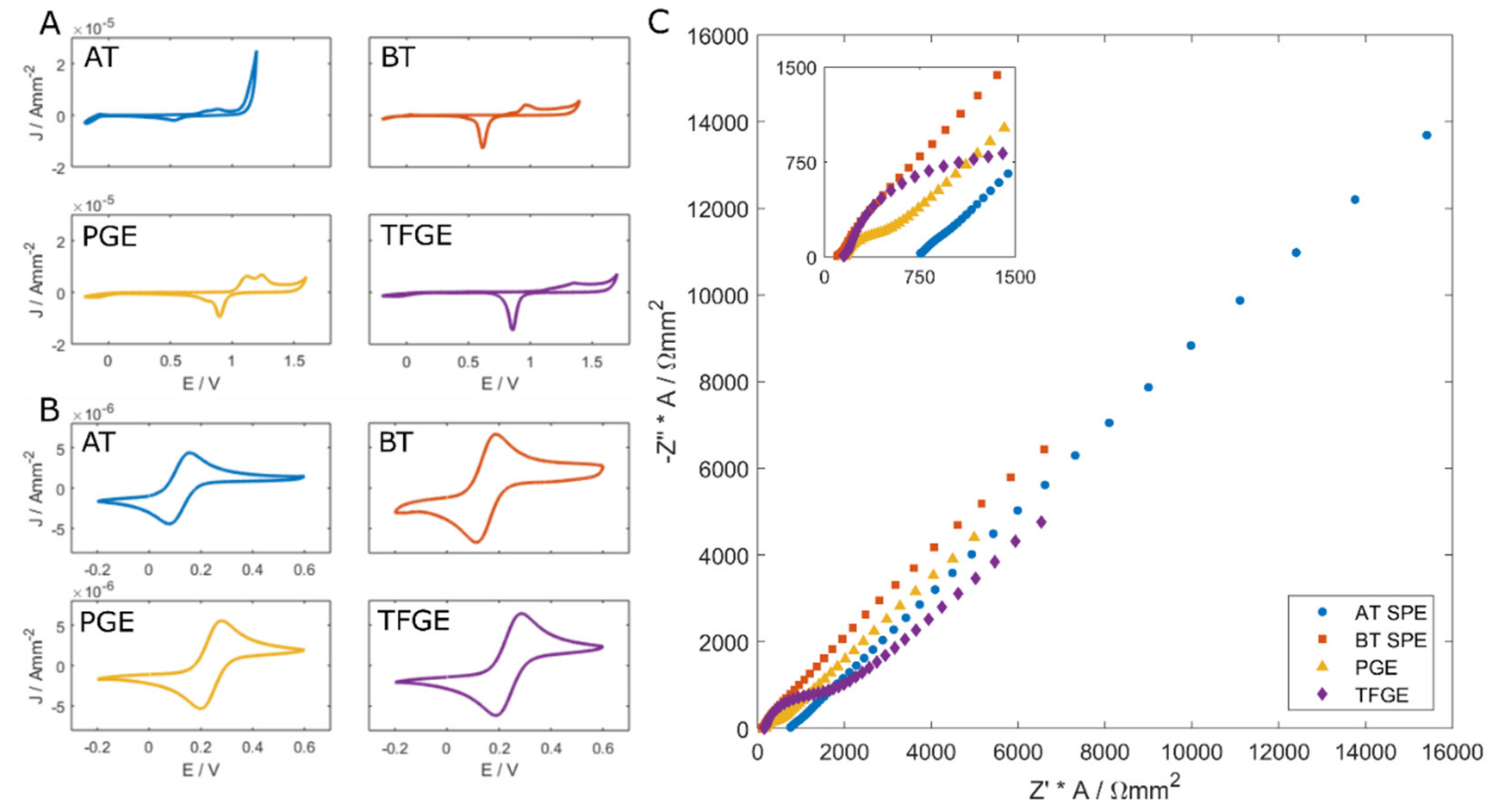
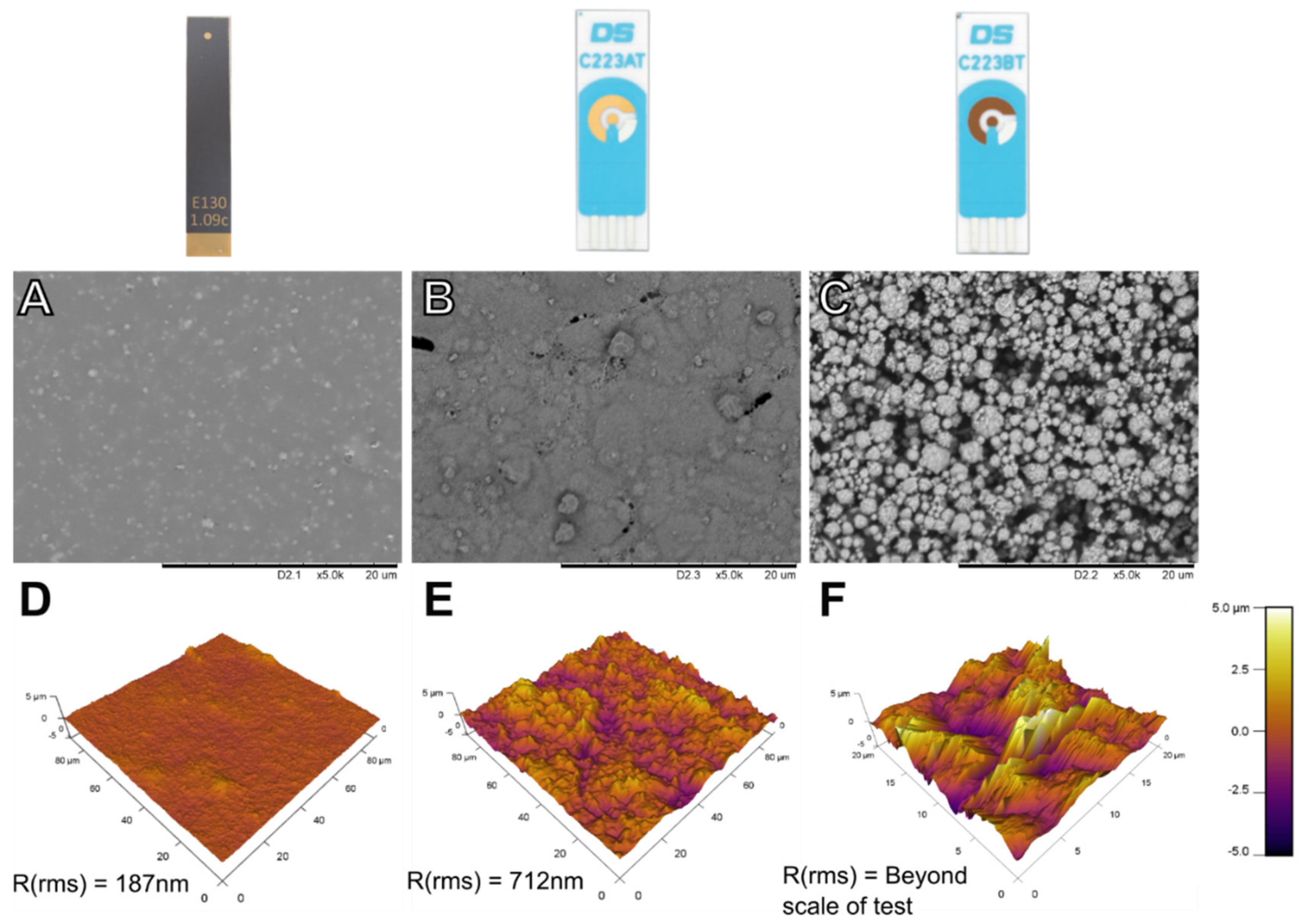
3.1. Electrode Performance—Initial Characterisation
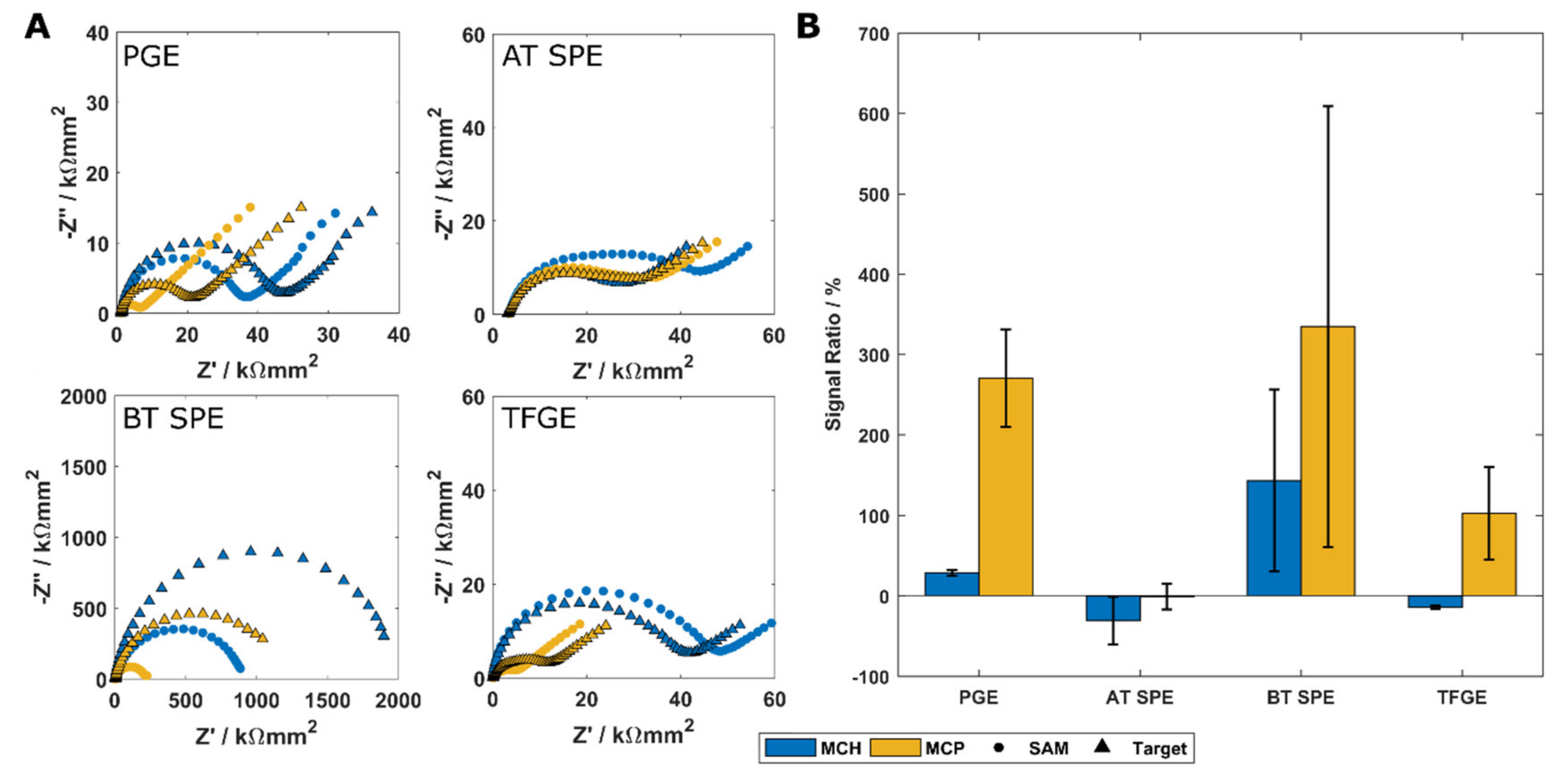
3.2. Electrode Performance Following Cleaning and Immobilisation of SAM Layers
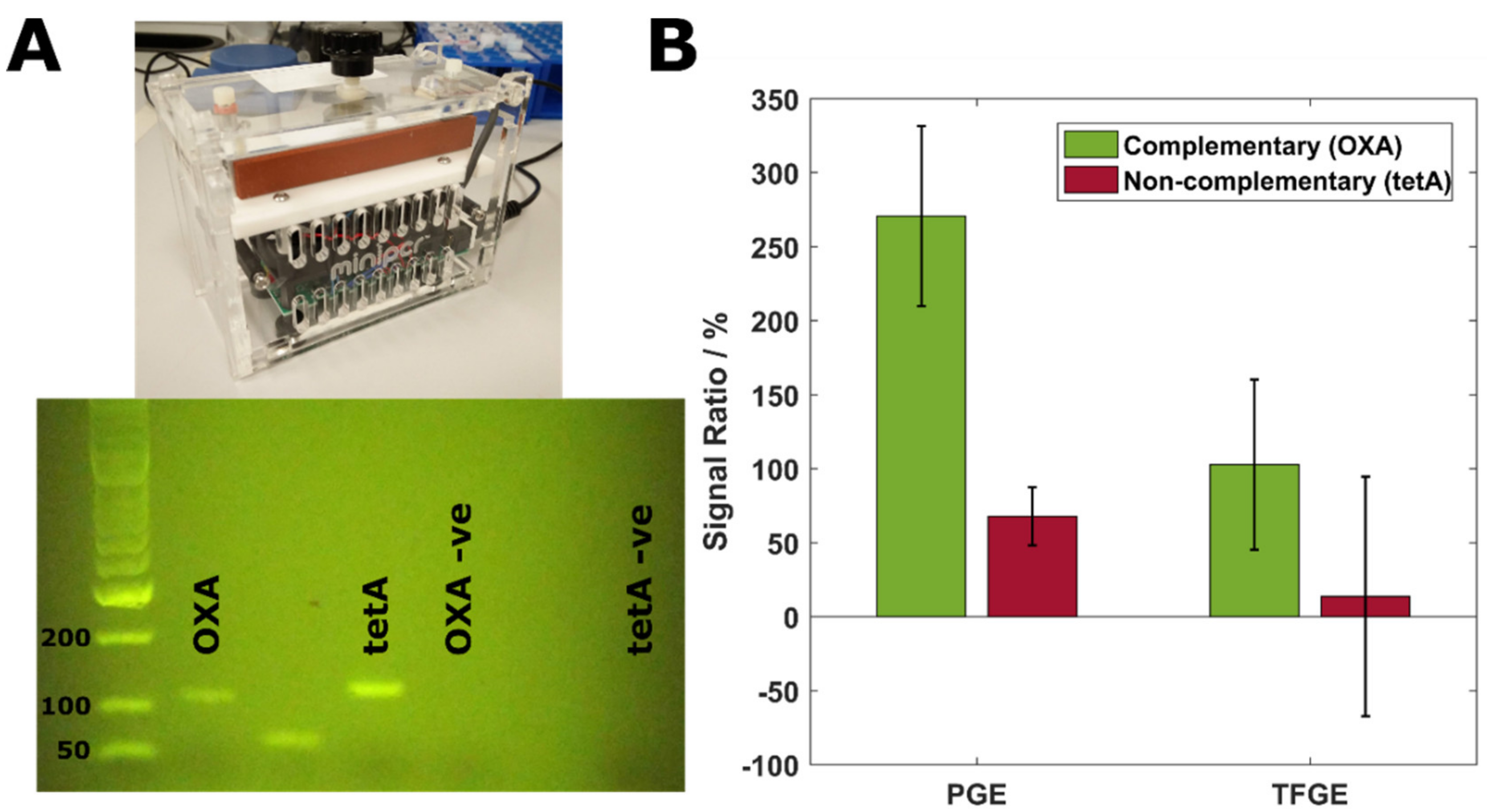
3.3. Specificity and Sensitivity of a TFGE-Based Biosensor
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Mehrotra, P. Biosensors and their applications—A review. J. Oral. Biol. Craniofac. Re.s 2016, 6, 153–159. [Google Scholar] [CrossRef] [PubMed]
- Niu, X.; Zheng, W.; Yin, C.; Weng, W.; Li, G.; Sun, W.; Men, Y. Electrochemical DNA biosensor based on gold nanoparticles and partially reduced graphene oxide modified electrode for the detection of Listeria monocytogenes hly gene sequence. J. Electroanal. Chem. 2017, 806, 116–122. [Google Scholar] [CrossRef]
- Hashimoto, K.; Inada, M.; Ito, K. A novel voltammetric approach for real-time electrochemical detection of targeted nucleic acid sequences using LAMP. Anal. Biochem. 2017, 539, 113–117. [Google Scholar] [CrossRef] [PubMed]
- Cheng, W.; Zhang, W.; Yan, Y.; Shen, B.; Zhu, D.; Lei, P.; Ding, S. A novel electrochemical biosensor for ultrasensitive and specific detection of DNA based on molecular beacon mediated circular strand displacement and rolling circle amplification. Biosens. Bioelectron. 2014, 62, 274–279. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Liang, W.; Wen, Y.; Wang, L.; Yang, X.; Ren, S.; Jia, N.; Zuo, X.; Liu, G. An ultrasensitive electrochemical biosensor for the detection of mecA gene in methicillin-resistant Staphylococcus aureus. Biosens. Bioelectron. 2018, 99, 424–430. [Google Scholar] [CrossRef] [PubMed]
- Matsishin, M.; Rachkov, A.; Errachid, A.; Dzyadevych, S.; Soldatkin, A. Development of impedimetric DNA biosensor for selective detection and discrimination of oligonucleotide sequences of the rpoB gene of Mycobacterium tuberculosis. Sens. Actuators B Chem. 2016, 222, 1152–1158. [Google Scholar] [CrossRef]
- Manzano, M.; Viezzi, S.; Mazerat, S.; Marks, R.S.; Vidic, J. Rapid and label-free electrochemical DNA biosensor for detecting hepatitis A virus. Biosens. Bioelectron. 2018, 100, 89–95. [Google Scholar] [CrossRef]
- Ben-Yoav, H.; Dykstra, P.H.; Bentley, W.E.; Ghodssi, R. A controlled microfluidic electrochemical lab-on-a-chip for label-free diffusion-restricted DNA hybridization analysis. Biosens. Bioelectron. 2015, 64, 579–585. [Google Scholar] [CrossRef]
- Bhattacharya, S.; Salamat, S.; Morisette, D.; Banada, P.; Akin, D.; Liu, Y.-S.; Bhunia, A.K.; Ladisch, M.; Bashir, R. PCR-based detection in a micro-fabricated platform. Lab Chip 2008, 8, 1130–1136. [Google Scholar] [CrossRef]
- Fang, X.; Jin, Q.; Jing, F.; Zhang, H.; Zhang, F.; Mao, H.; Xu, B.; Zhao, J. Integrated biochip for label-free and real-time detection of DNA amplification by contactless impedance measurements based on interdigitated electrodes. Biosens. Bioelectron. 2013, 44, 241–247. [Google Scholar] [CrossRef]
- Uludag, Y.; Narter, F.; Sağlam, E.; Köktürk, G.; Gök, M.Y.; Akgün, M.; Barut, S.; Budak, S. An integrated lab-on-a-chip-based electrochemical biosensor for rapid and sensitive detection of cancer biomarkers. Anal. Bioanal. Chem. 2016, 408, 7775–7783. [Google Scholar] [CrossRef] [PubMed]
- Norian, H.M.; Field, R.; Kymissis, I.L.; Shepard, K. An integrated CMOS quantitative-polymerase-chain-reaction lab-on-chip for point-of-care diagnostics. Lab Chip 2014, 14, 4076–4084. [Google Scholar] [CrossRef] [PubMed]
- Lee, T.M.-H.; Carles, M.C.; Hsing, I.-M. Microfabricated PCR-electrochemical device for simultaneous DNA amplification and detection. Lab Chip 2003, 3, 100–105. [Google Scholar] [CrossRef] [PubMed]
- Davies, J.; Davies, D. Origins and Evolution of Antibiotic Resistance. Microbiol. Mol. Biol. Rev. 2010, 74, 417–433. [Google Scholar] [CrossRef] [PubMed]
- European Centre for Disease Prevention and Control Surveillance of Antimicrobial Resistance in Europe: Annual report of the European Antimicrobial Resistance Surveillance Network (EARS-Net) 2017. Available online: https://ecdc.europa.eu/en/publications-data/surveillance-antimicrobial-resistance-europe-2017 (accessed on 3 December 2018).
- Fair, R.J.; Tor, Y. Antibiotics and Bacterial Resistance in the 21st Century. Perspect. Medicin. Chem. 2014, 6, 25–64. [Google Scholar] [CrossRef]
- Denkinger, C.M.; Grant, A.D.; Denkinger, M.; Gautam, S.; D’Agata, E.M.C. Increased multi-drug resistance among the elderly on admission to the hospital—A 12-year surveillance study. Arch. Gerontol. Geriatr. 2013, 56, 227–230. [Google Scholar] [CrossRef]
- Cardoso, A.R.; Moreira, F.T.C.; Fernandes, R.; Sales, M.G.F. Novel and simple electrochemical biosensor monitoring attomolar levels of miRNA-155 in breast cancer. Biosens. Bioelectron. 2016, 80, 621–630. [Google Scholar] [CrossRef]
- Ribeiro, J.A.; Pereira, C.M.; Silva, A.F.; Sales, M.G.F. Disposable electrochemical detection of breast cancer tumour marker CA 15-3 using poly(Toluidine Blue) as imprinted polymer receptor. Biosens. Bioelectron. 2018, 109, 246–254. [Google Scholar] [CrossRef]
- Li, Y.; Deng, L.; Deng, C.; Nie, Z.; Yang, M.; Si, S. Simple and sensitive aptasensor based on quantum dot-coated silica nanospheres and the gold screen-printed electrode. Talanta 2012, 99, 637–642. [Google Scholar] [CrossRef]
- Mousavisani, S.Z.; Raoof, J.B.; Ojani, R.; Bagheryan, Z. An impedimetric biosensor for DNA damage detection and study of the protective effect of deferoxamine against DNA damage. Bioelectrochemistry 2018, 122, 142–148. [Google Scholar] [CrossRef]
- Henihan, G.; Schulze, H.; Corrigan, D.K.; Giraud, G.; Terry, J.G.; Hardie, A.; Campbell, C.J.; Walton, A.J.; Crain, J.; Pethig, R.; et al. Label- and amplification-free electrochemical detection of bacterial ribosomal RNA. Biosens. Bioelectron. 2016, 81, 487–494. [Google Scholar] [CrossRef] [PubMed]
- Kumar Mistry, K.; Layek, K.; Mahapatra, A.; RoyChaudhuri, C.; Saha, H. A review on amperometric-type immunosensors based on screen-printed electrodes. Analyst 2014, 139, 2289–2311. [Google Scholar] [CrossRef] [PubMed]
- Elgrishi, N.; Rountree, K.J.; McCarthy, B.D.; Rountree, E.S.; Eisenhart, T.T.; Dempsey, J.L. A Practical Beginner’s Guide to Cyclic Voltammetry. J. Chem. Educ. 2018, 95, 197–206. [Google Scholar] [CrossRef]
- Bard, A.; Faulkner, L. Electrochemical Methods: Fundamentals and Applications, 2nd ed.; Wiley: Hoboken, NJ, USA, 2001; ISBN 0-471-04372-9. [Google Scholar]
- Keighley, S.D.; Li, P.; Estrela, P.; Migliorato, P. Optimization of DNA immobilization on gold electrodes for label-free detection by electrochemical impedance spectroscopy. Biosens. Bioelectron. 2008, 23, 1291–1297. [Google Scholar] [CrossRef] [PubMed]
- Štrbac, S.; Adžć, R.R.; Hamelin, A. Oxide formation on gold single crystal stepped surfaces. J. Electroanal. Chem. Interfacial. Electrochem. 1988, 249, 291–310. [Google Scholar] [CrossRef]
- Bernalte, E.; Marín-Sánchez, C.; Pinilla-Gil, E.; Brett, C.M.A. Characterisation of screen-printed gold and gold nanoparticle-modified carbon sensors by electrochemical impedance spectroscopy. J. Electroanal. Chem. 2013, 709, 70–76. [Google Scholar] [CrossRef]
- Corrigan, D.K.; Schulze, H.; Henihan, G.; Hardie, A.; Ciani, I.; Giraud, G.G.; Terry, J.J.; Walton, A.; Pethig, R.; Ghazal, P.; et al. Development of a PCR-free electrochemical point of care test for clinical detection of methicillin resistant Staphylococcus aureus (MRSA). Analyst 2013, 138, 6997–7005. [Google Scholar] [CrossRef] [PubMed]
- Corrigan, D.K.; Schulze, H.; Ciani, I.; Henihan, G.; Mount, A.R.; Bachmann, T.T. Improving performance of a rapid electrochemical MRSA assay: Optimisation of assay conditions to achieve enhanced discrimination of clinically important DNA sequences under ambient conditions. J. Electroanal. Chem. 2017, 786, 58–62. [Google Scholar] [CrossRef]
- Corrigan, D.K.; Schulze, H.; McDermott, R.A.; Schmüser, I.; Henihan, G.; Henry, J.B.; Bachmann, T.T.; Mount, A.R. Improving electrochemical biosensor performance by understanding the influence of target DNA length on assay sensitivity. J. Electroanal. Chem. 2014, 732, 25–29. [Google Scholar] [CrossRef]
- Liu, B.; Bard, A.J.; Mirkin, M.V.; Creager, S.E. Electron Transfer at Self-Assembled Monolayers Measured by Scanning Electrochemical Microscopy. J. Am. Chem. Soc. 2004, 126, 1485–1492. [Google Scholar] [CrossRef]
- Gębala, M.; Schuhmann, W. Controlled Orientation of DNA in a Binary SAM as a Key for the Successful Determination of DNA Hybridization by Means of Electrochemical Impedance Spectroscopy. ChemPhysChem 2010, 11, 2887–2895. [Google Scholar] [CrossRef] [PubMed]
- Keighley, S.D.; Estrela, P.; Li, P.; Migliorato, P. Optimization of label-free DNA detection with electrochemical impedance spectroscopy using PNA probes. Biosens. Bioelectron. 2008, 24, 906–911. [Google Scholar] [CrossRef] [PubMed]
- Love, J.C.; Estroff, L.A.; Kriebel, J.K.; Nuzzo, R.G.; Whitesides, G.M. Self-Assembled Monolayers of Thiolates on Metals as a Form of Nanotechnology. Chem. Rev. 2005, 105, 1103–1170. [Google Scholar] [CrossRef] [PubMed]
- Cecchet, F.; Marcaccio, M.; Margotti, M.; Paolucci, F.; Rapino, S.; Rudolf, P. Redox Mediation at 11-Mercaptoundecanoic Acid Self-Assembled Monolayers on Gold. J. Phys. Chem. B 2006, 110, 2241–2248. [Google Scholar] [CrossRef] [PubMed]
- Riedel, M.; Kartchemnik, J.; Schöning, M.J.; Lisdat, F. Impedimetric DNA Detection—Steps Forward to Sensorial Application. Anal. Chem. 2014, 86, 7867–7874. [Google Scholar] [CrossRef] [PubMed]
- Kafka, J.; Pänke, O.; Abendroth, B.; Lisdat, F. A label-free DNA sensor based on impedance spectroscopy. Electrochim. Acta 2008, 53, 7467–7474. [Google Scholar] [CrossRef]
- Liu, G.; Sun, C.; Li, D.; Song, S.; Mao, B.; Fan, C.; Tian, Z. Gating of Redox Currents at Gold Nanoelectrodes via DNA Hybridization. Adv. Mater. 2010, 22, 2148–2150. [Google Scholar] [CrossRef] [PubMed]
- Malecka, K.; Stachyra, A.; Góra-Sochacka, A.; Sirko, A.; Zagórski-Ostoja, W.; Radecka, H.; Radecki, J. Electrochemical genosensor based on disc and screen printed gold electrodes for detection of specific DNA and RNA sequences derived from Avian Influenza Virus H5N1. Sens. Actuators B Chem. 2016, 224, 290–297. [Google Scholar] [CrossRef]
| Sequence Name | Forward Sequence | Reverse Sequence | Amplicon Length (bp) |
|---|---|---|---|
| OXA Primers | AACAGAAGCATGGCTCGAAA | TGGTGTTTTCTATGGCTGAGTT | 116 |
| tetA Primers | GCATGATGAAGAAGACCGCCA | GAGTCGCACAAAGGCGAAC | 121 |
| DNA Probe | [ThiC6]AACAGAAGCATGGCTCGAAA | ||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Butterworth, A.; Blues, E.; Williamson, P.; Cardona, M.; Gray, L.; Corrigan, D.K. SAM Composition and Electrode Roughness Affect Performance of a DNA Biosensor for Antibiotic Resistance. Biosensors 2019, 9, 22. https://doi.org/10.3390/bios9010022
Butterworth A, Blues E, Williamson P, Cardona M, Gray L, Corrigan DK. SAM Composition and Electrode Roughness Affect Performance of a DNA Biosensor for Antibiotic Resistance. Biosensors. 2019; 9(1):22. https://doi.org/10.3390/bios9010022
Chicago/Turabian StyleButterworth, Adrian, Elizabeth Blues, Paul Williamson, Milovan Cardona, Louise Gray, and Damion K Corrigan. 2019. "SAM Composition and Electrode Roughness Affect Performance of a DNA Biosensor for Antibiotic Resistance" Biosensors 9, no. 1: 22. https://doi.org/10.3390/bios9010022
APA StyleButterworth, A., Blues, E., Williamson, P., Cardona, M., Gray, L., & Corrigan, D. K. (2019). SAM Composition and Electrode Roughness Affect Performance of a DNA Biosensor for Antibiotic Resistance. Biosensors, 9(1), 22. https://doi.org/10.3390/bios9010022






