Aptasensors Based on Whispering Gallery Mode Resonators
Abstract
:1. Introduction
2. Materials and Methods
2.1. Surface Functionalisation
2.1.1. Materials and Silica Microspherical WGMR Functionalisation
Dry Protocol
Wet Protocol
- 5′-HO-(CH2)3-S-S-(CH2)3- GGT TGG TGT GGT TGG-3′ (TBA-15)
- 5′-HO-(CH2)3-S-S-(CH2)3-AG TCC GTG GTA GGG CAG GTT GGG GTG ACT-3′ (TBA-29)
- 5′-HO-(CH2)3-S-S-(CH2)3-CCG TCG AGC AGA GTT-3′ (NS, Non Sense)
- 5′-HO-(CH2)3-S-S-(CH2)3-CC GTC TTC CAG ACA AGA GTG CAG GG-3′ (hereafter called VEGF-25)
2.1.2. Materials and SiON WGMR Functionalization
2.2. Experimental Set-up
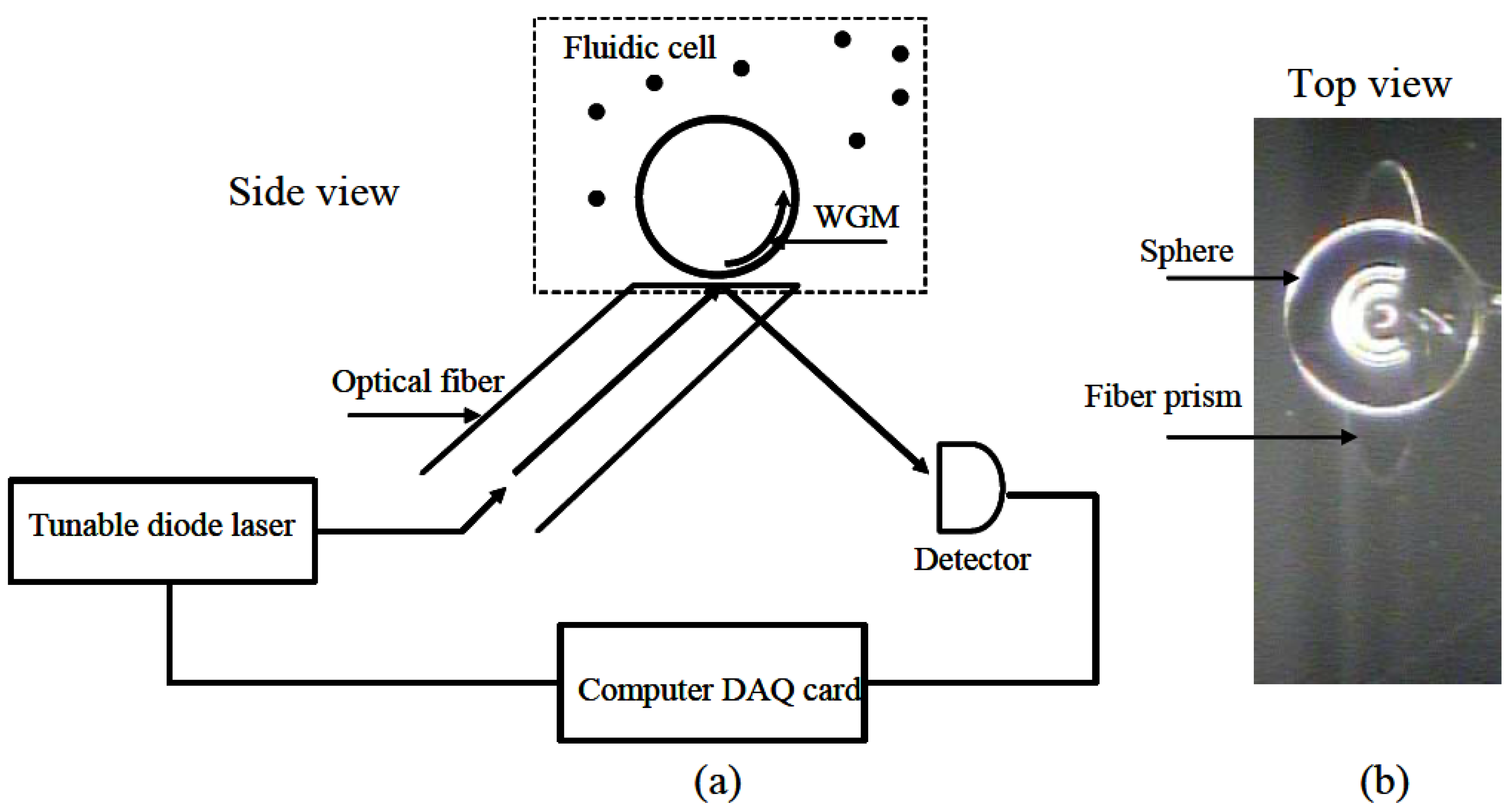
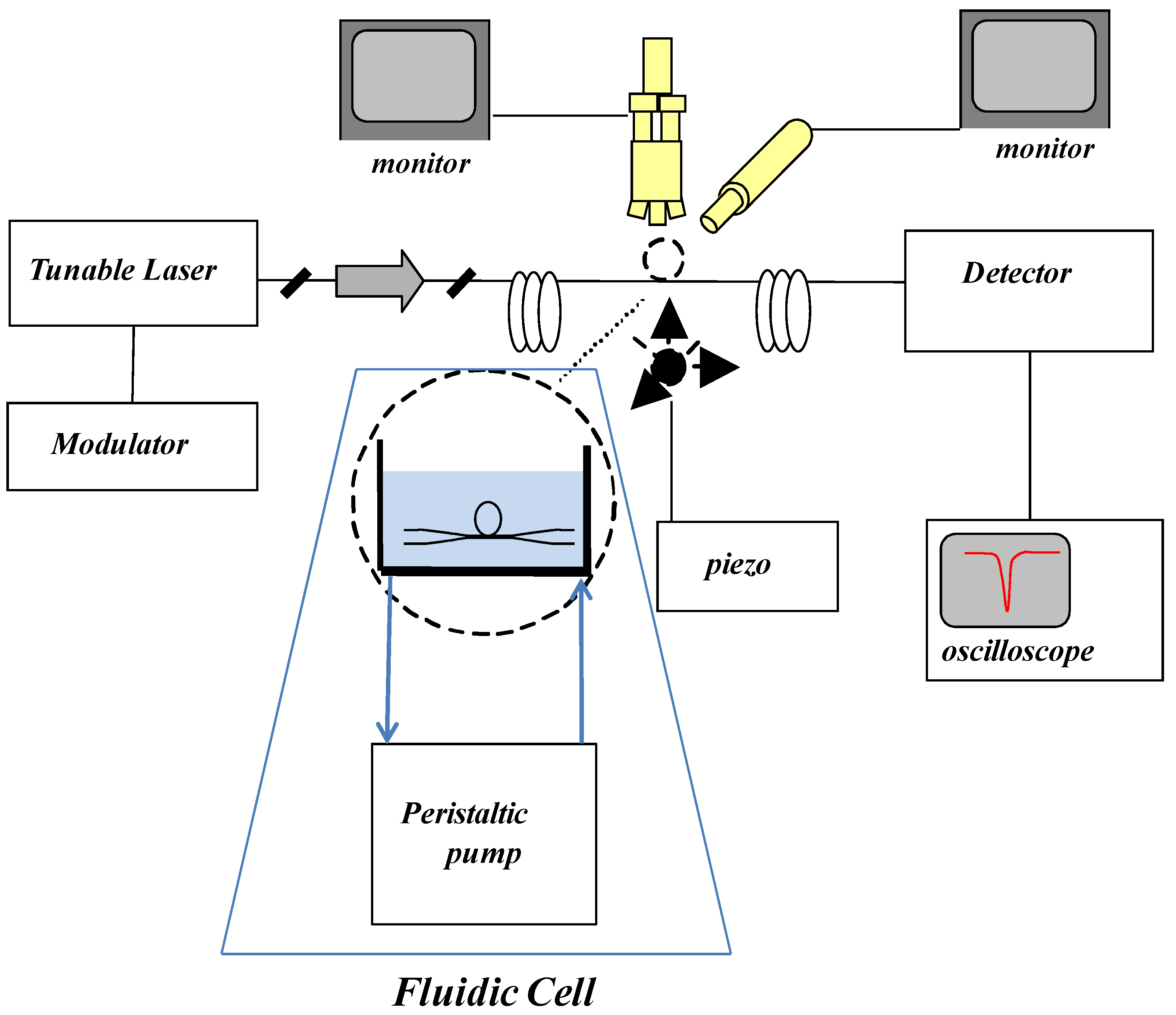
2.2.1. Experimental Set-ups for Microspheres Based Aptasensors
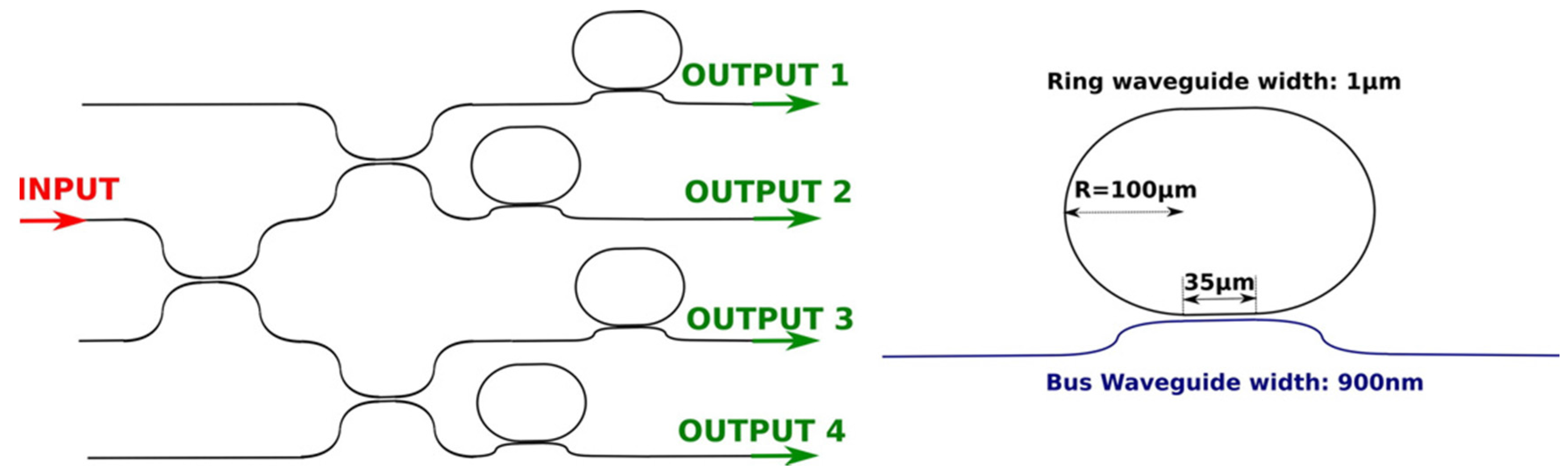
2.2.2. Experimental Set-ups for Racetrack Microring Based Aptasensors
3. Results
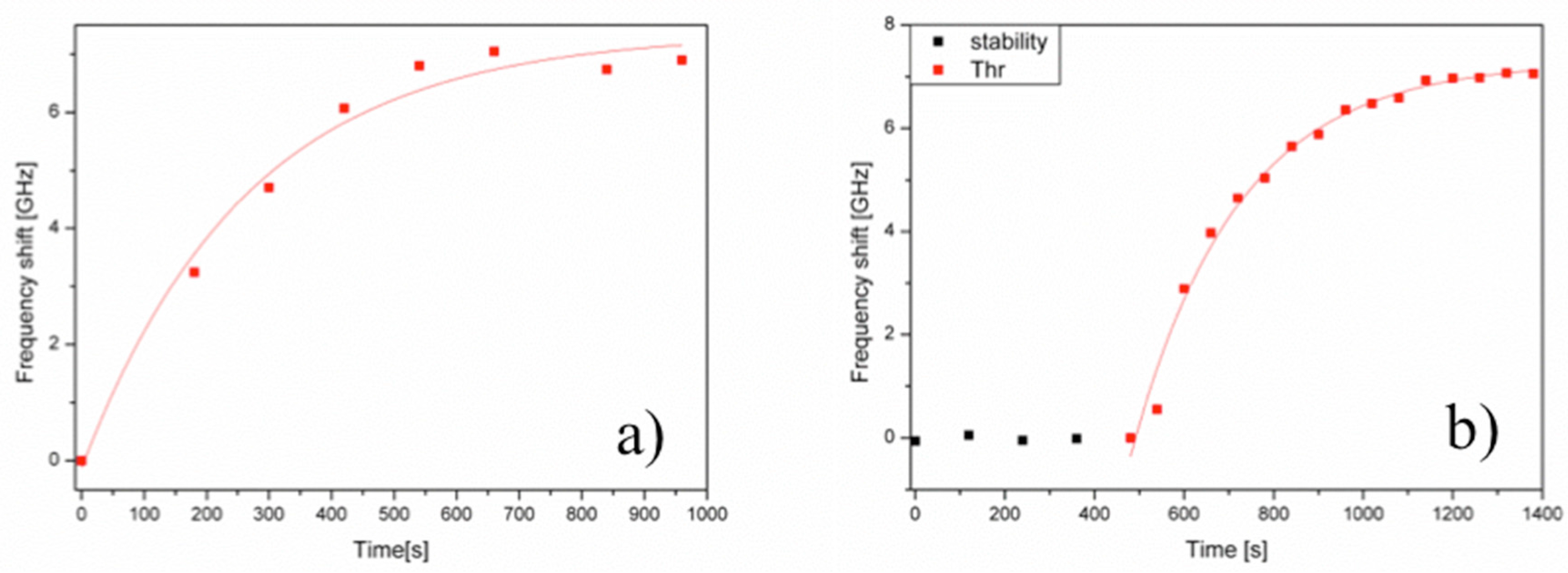
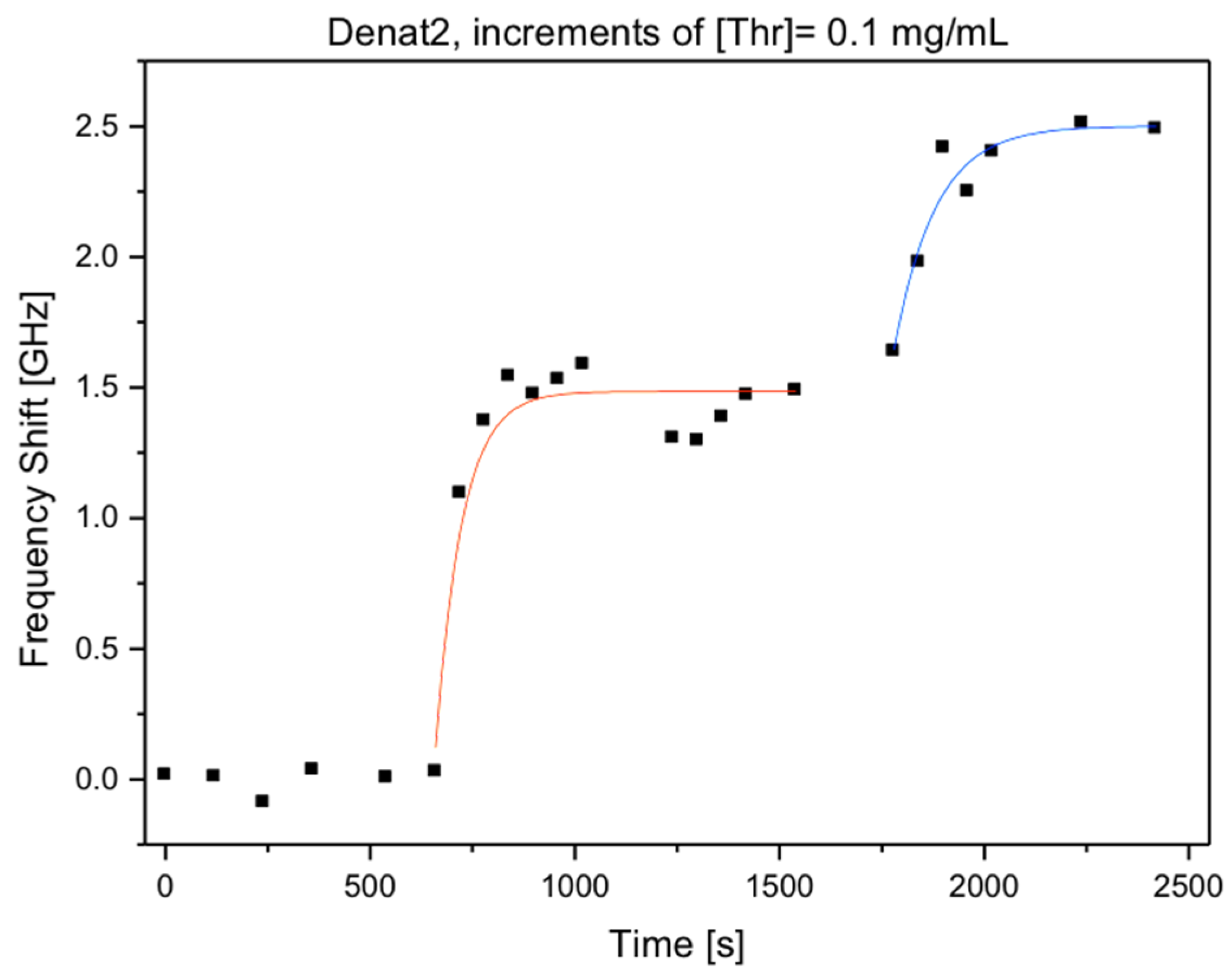
3.1. Microspherical Aptasensors for Thrombin and VEGF
3.2. Racetrack Ring Aptasensors for Aflatoxin M1
4. Discussion
δRmax = (Δνmax R)/νopt
5. Conclusions
Acknowledgments
Conflicts of Interest
Abbreviations
| WGMR | Whispering gallery mode resonators |
| SiON | silicon oxynitrid |
| SPR | surface plasmon resonators |
| VEGF | Vascular endothelial growth factor |
| AFM1 | aflatoxin M1 |
| ELISA | Enzyme-LinkedImmunoSorbent Assay |
| HPLC | High-Performance Liquid Chromatography |
| APS | Aminopropyltrimethoxysilane |
| MPTMS | mercaptopropyltrimethoxysilane |
| GPTMS | glycidoxypropyl methyldiethoxy silane |
| PDC | phenylene diisothiocyanate |
| DMF | dimethyl formamide |
| CB | Coupling Buffer |
| MB | Modification Buffer |
| BRB | BioRecognition Buffer |
| TBA | Thrombin Binding Aptamer |
| PECVD | plasma enhanced chemical vapor deposition |
| MES | 2-(Nmorpholino) ethanesulfonic acid |
| DSMO | dimethyl sulfoxide |
References
- Ellington, A.D.; Szostak, J.W. In vitro selection of RNA molecules that bind specific ligands. Nature 1990, 346, 818–822. [Google Scholar] [CrossRef] [PubMed]
- Song, S.; Wang, L.; Li, J.; Zhao, J.; Fan, C. Aptamer-based biosensors. TrAC Trend Anal. Chem. 2008, 27, 108–117. [Google Scholar] [CrossRef]
- Mascini, M.; Palchetti, I.; Tombelli, S. Nucleic Acid and Peptide Aptamers: Fundamentals and Bioanalytical Aspects. Angew. Chem. Int. Ed. 2012, 51, 1316–1332. [Google Scholar] [CrossRef] [PubMed]
- Sassolas, A.; Blum, L.J.; Leca-Bouvier, B.D. Optical detection systems using immobilized aptamers. Biosens. Bioelectron. 2011, 26, 3725–3736. [Google Scholar] [CrossRef] [PubMed]
- Soria, S.; Berneschi, S.; Brenci, M.; Cosi, F.; Nunzi Conti, G.; Pelli, S.; Righini, G.C. Optical microspherical resonators for biomedical sensing. Sensors 2011, 11, 785–805. [Google Scholar] [CrossRef] [PubMed]
- Foreman, M.R.; Swaim, J.D.; Vollmer, F. Whispering gallery mode sensors. Adv. Opt. Photonics 2015, 7, 168–240. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Suter, J.D.; White, I.M.; Fan, X. Aptamer based microsphere biosensor for thrombin detection. Sensors 2006, 6, 785–795. [Google Scholar] [CrossRef]
- Pasquardini, L.; Berneschi, S.; Barucci, A.; Cosi, F.; Dallpiccola, R.; Lunelli, L.; Nunzi Conti, G.; Pederzolli, C.; Soria, S. Whispering Gallery Modes Aptasensors for detection of blood proteins. J. Biophotonics 2013, 6, 178–187. [Google Scholar] [CrossRef] [PubMed]
- Guider, R.; Gandolfi, D.; Chalyan, T.; Pasquardini, L.; Samusenko, A.; Pucker, G.; Pederzolli, C.; Pavesi, L. Design and Optimization of SiON Ring Resonator-Based Biosensors for Aflatoxin M1 Detection. Sensors 2015, 15, 17300–17312. [Google Scholar] [CrossRef] [PubMed]
- Pimková, K.; Bocková, M.; Hegnerová, K.; Suttnar, J.; Cermák, J.; Homola, J.; Dyr, J.E. Surface plasmon resonance biosensor for the detection of VEGFR-1--a protein marker of myelodysplastic syndromes. Anal. Bioanal. Chem. 2012, 402, 381–387. [Google Scholar] [CrossRef] [PubMed]
- Vollmer, F.; Arnold, S.; Keng, D. Single virus detection from the reactive shift of a whispering-gallery mode. Proc. Natl. Acad. Sci. USA 2008, 105, 20701–20704. [Google Scholar] [CrossRef] [PubMed]
- Biancardi, A. Determinazione di aflatossina M1 nel latte. Industrie Alimentari 1997, XXXVI, 870–876. (In Italian) [Google Scholar]
- Markaki, P.; Melissari, E. Occurrence of aflatoxin M1 in commercial pasteurised milk determined with ELISA and HPLC. Food Addit. Contam. 1997, 14, 451–456. [Google Scholar] [CrossRef] [PubMed]
- Ren, H.-C.; Vollmer, F.; Arnold, S.; Libchaber, A. High-Q microsphere biosensor—Analysis for adsorption of rodlike bacteria. Opt. Express 2007, 15, 17410–17423. [Google Scholar] [CrossRef] [PubMed]
- Charles, P.T.; Vora, G.J.; Andreadis, J.D.; Fortney, A.J.; Meador, C.E.; Dulcey, C.S.; Stenger, D.A. Fabrication and Surface Characterization of DNA Microarrays Using Amine- and Thiol-Terminated Oligonucleotide Probes. Langmuir 2003, 19, 1586–1591. [Google Scholar] [CrossRef]
- Hilliard, L.; Zhao, X.; Tan, W. Immobilization of oligonucleotides onto silica nanoparticles for DNA hybridization studies. Anal. Chim. Acta 2002, 470, 51–56. [Google Scholar] [CrossRef]
- Ultra-Fast mycotoxin clean-up columns. Available online: http://neoventures.ca/products/mycotoxin-testing/Ultra-Fast%20ad.pdf (accessed on 22 April 2016).
- Vollmer, F.; Braun, D.; Libchaber, A.; Khoshsima, M.; Teraoka, I.; Arnold, S. Protein detection by optical shift of a resonant microcavity. Appl. Phys. Lett. 2002, 80, 4057–4059. [Google Scholar] [CrossRef]
- Hanumegowda, N.M.; Stica, C.J.; Patel, B.C.; White, I.M.; Fan, X. Refractometric sensors based on microsphere resonators. Appl. Phys. Lett. 2005, 87, 201107. [Google Scholar] [CrossRef]
- Brenci, M.; Calzolai, R.; Cosi, F.; Nunzi Conti, G.; Pelli, S.; Righini, G.C. Microspherical resonators for biophotonic sensors. Proc. SPIE 2006, 6158, 61580S. [Google Scholar] [CrossRef]
- Ristic, D.; Berneschi, S.; Camerini, M.; Farnesi, D.; Pelli, S.; Trono, C.; Chiappini, A.; Chiasera, A.; Ferrari, M.; Lukowiak, A.; et al. Photoluminescence and lasing in whispering gallery mode glass microspherical resonators. J. Lumin. 2016, 170, 755–760. [Google Scholar] [CrossRef]
- Arnold, S.; Ramjit, R.; Keng, D.; Kolchenko, V.; Teraoka, I. MicroParticle photophysics illuminates viral bio-sensing. Faraday Discuss. 2008, 137, 65–83; discussion 99–113. [Google Scholar] [CrossRef] [PubMed]
- Soria, S.; Baldini, F.; Berneschi, S.; Cosi, F.; Giannetti, A.; Nunzi Conti, G.; Pelli, S.; Righini, G.C.; Tiribilli, B. High-Q polymer-coated microspheres for immunosensing applications. Opt. Express 2009, 17, 14694–14699. [Google Scholar] [CrossRef] [PubMed]
- Giannetti, A.; Berneschi, S.; Baldini, F.; Cosi, F.; Nunzi Conti, G.; Soria, S. Performance of Eudragit Coated Whispering Gallery Mode Resonator-Based Immunosensors. Sensors 2012, 12, 14604–14611. [Google Scholar] [CrossRef] [PubMed]
- Guider, R.; Gandolfi, D.; Chalyan, T.; Pasquardini, L.; Samusenko, A.; Pederzolli, C.; Pucker, G.; Pavesi, L. Sensitivity and Limit of Detection of biosensors based on ring resonators. Sens. Bio-Sens. Res. 2015, 6, 99–102. [Google Scholar] [CrossRef]
- Gandolfi, D.; Guider, R.; Chalyan, T.; Pavesi, L.; Pasquardini, L.; Pederzolli, C.; Samusenko, A.; Pucker, G. Sensitivity and Limit of detection of biosensors based on ring resonators. In Proceedings of the 2015 XVIII IEEE AISEM Annual Conference, Trento, Italy, 3–5 February 2015. [CrossRef]
- Iqbal, M.; Gleeson, M.A.; Spaugh, B.; Tybor, F.; Gunn, W.G.; Hochberg, M.; Baehr-Jones, T.; Bailey, R.C.; Gunn, L.C. Label-free biosensor arrays based on silicon ring resonators and high-speed optical scanning instrumentation. IEEE J. Sel. Top. Quantum Electron. 2010, 16, 654–661. [Google Scholar] [CrossRef]
- Freitas, R.A., Jr. Nanomedicine, Volume IIA: Biocompability, Landes Bioscience: Georgetown, TX, 2003. Available online: http://www.nanomedicine.com/NMIIA/15.2.5.htm (accessed on 14 April 2016).
- Torquato, S. Mean nearest neighbor distance in random packaging in hard D-dimensional spheres. Phys. Rev. Lett. 1995, 74, 2156–2159. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.; Ilchenko, V.S.; Nadeau, J.; Maleki, L. Biochemical Detection With Optical Whispering-Gallery Resonators. Proc. SPIE 2007, 6452. [Google Scholar] [CrossRef]
- Soteropulos, C.E.; Hunt, H.K.; Armani, A.M. Determination of binding kinetics using whispering gallery mode microcavities. Appl. Phys. Lett. 2011, 99. [Google Scholar] [CrossRef] [PubMed]
- Goji, S.; Matsui, J. Direct detection of thrombin binding to 8-bromodeoxyguanosine-modified aptamer: Effects of modification on affinity and kinetics. J. Nucleic Acids 2011, 2011, 316079. [Google Scholar] [CrossRef]
- Li, X.X.; Shen, L.H.; Zhang, D.D.; Qi, H.L.; Gao, Q.; Ma, F.; Zhang, C.X. Electrochemical impedance spectroscopy for study of aptamer-thrombin interfacial interactions. Biosens. Bioelectron. 2008, 23, 1624–1630. [Google Scholar] [CrossRef] [PubMed]


© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nunzi Conti, G.; Berneschi, S.; Soria, S. Aptasensors Based on Whispering Gallery Mode Resonators. Biosensors 2016, 6, 28. https://doi.org/10.3390/bios6030028
Nunzi Conti G, Berneschi S, Soria S. Aptasensors Based on Whispering Gallery Mode Resonators. Biosensors. 2016; 6(3):28. https://doi.org/10.3390/bios6030028
Chicago/Turabian StyleNunzi Conti, Gualtiero, Simome Berneschi, and Silvia Soria. 2016. "Aptasensors Based on Whispering Gallery Mode Resonators" Biosensors 6, no. 3: 28. https://doi.org/10.3390/bios6030028
APA StyleNunzi Conti, G., Berneschi, S., & Soria, S. (2016). Aptasensors Based on Whispering Gallery Mode Resonators. Biosensors, 6(3), 28. https://doi.org/10.3390/bios6030028







