Kinetic Analyses of Data from a Human Serum Albumin Assay Using the liSPR System
Abstract
:1. Introduction
2. Experimental Section
2.1. Materials
2.2. SPR Measurements
2.2.1. Preparation of the Gold Surface and SAM
2.2.2. Immobilization of HSA
2.2.3. Kinetic Measurements
3. Results and Discussion
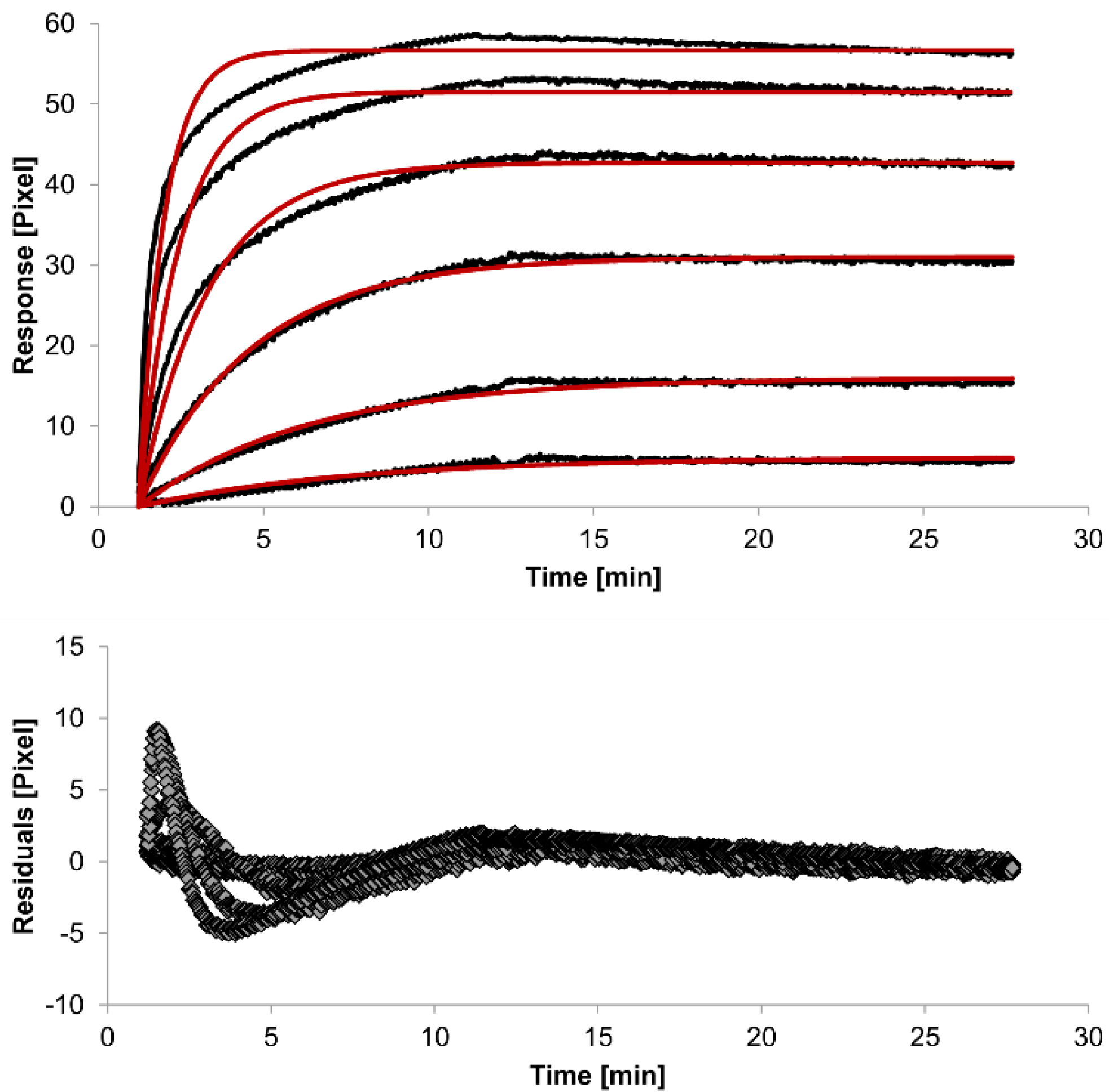
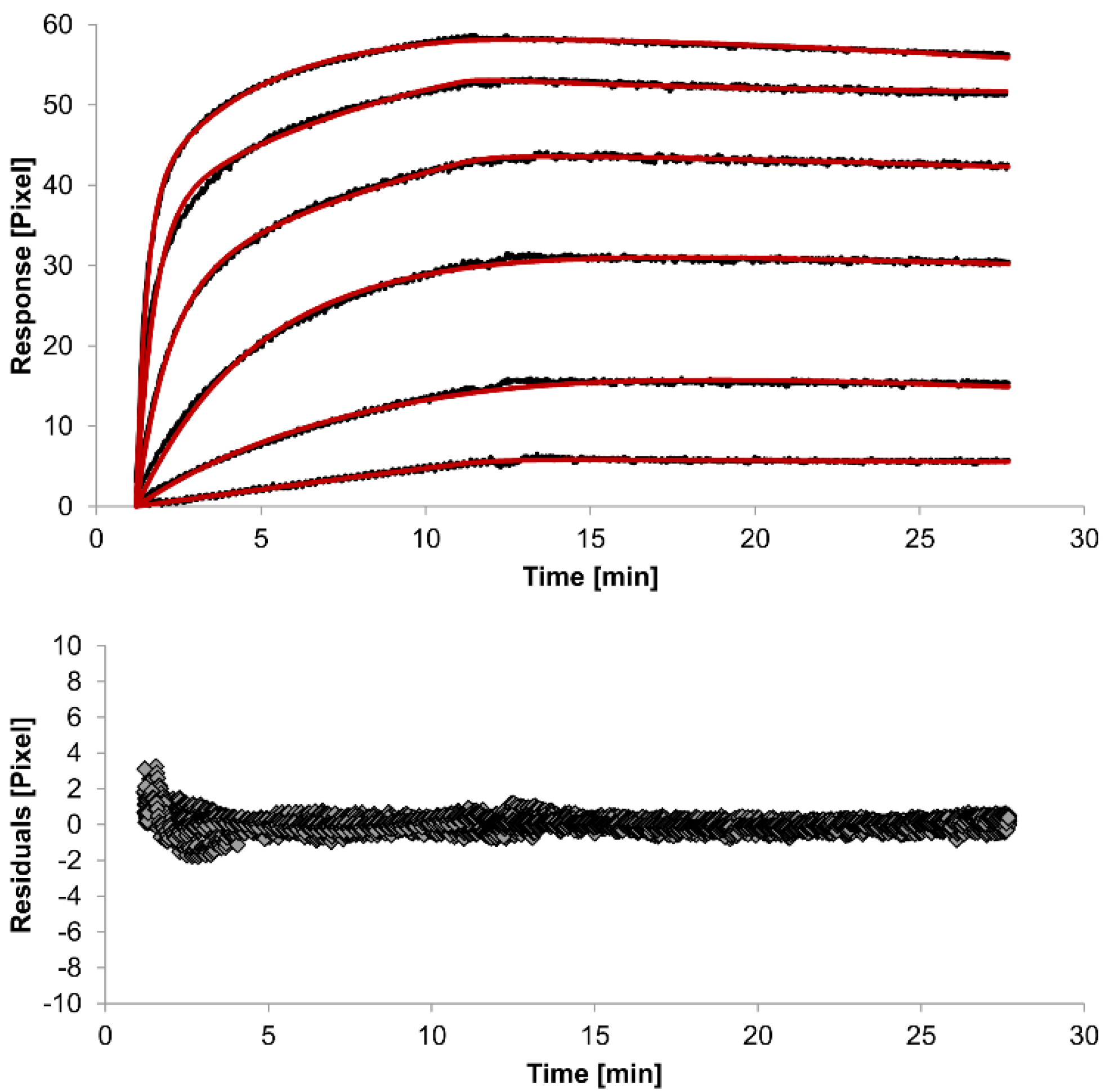
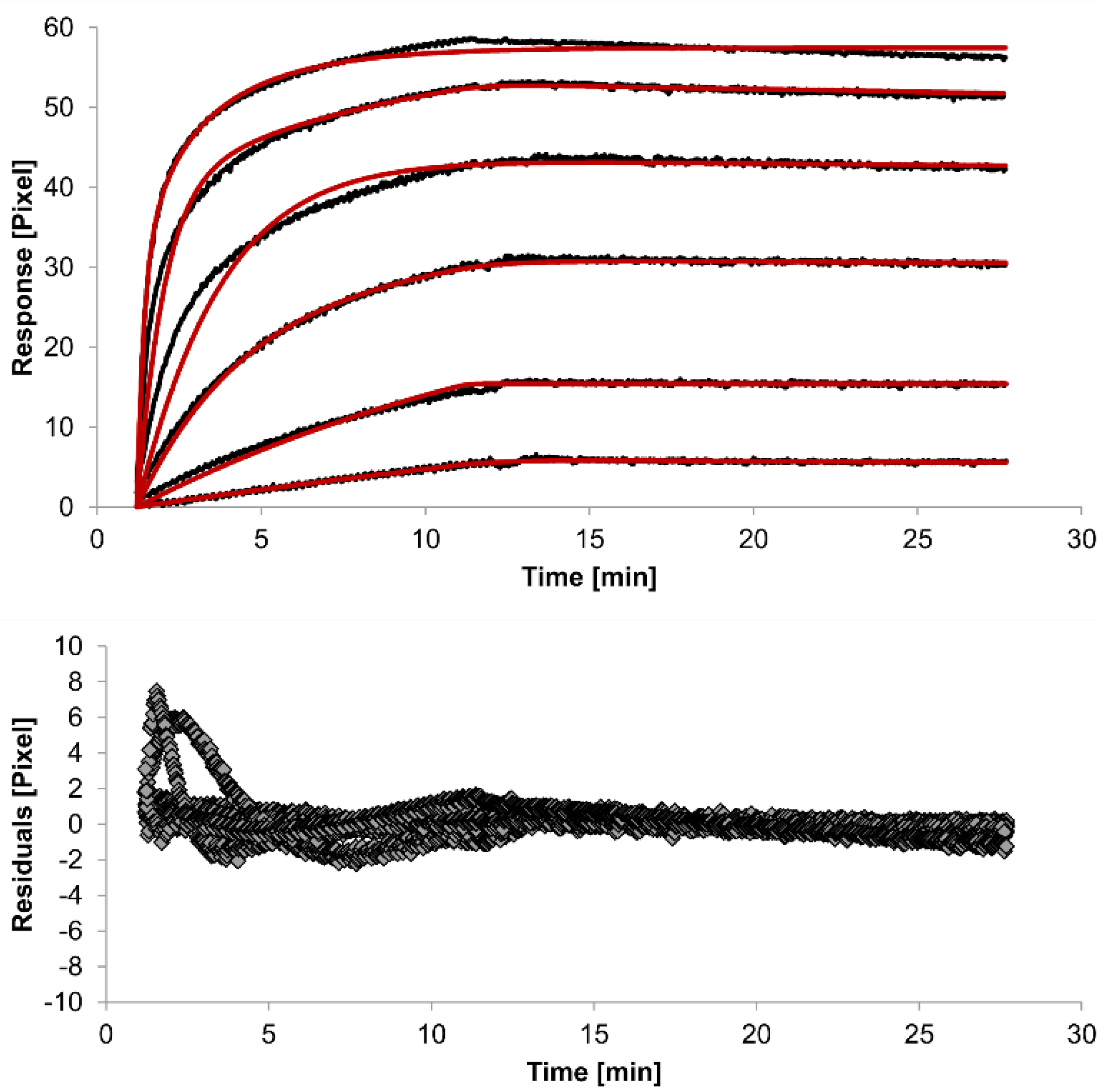
| Interaction Model | ka1 [M−1s−1] | kd1 [s−1] | ka2 [M−1s−1] | kd2 [s−1] | χ2 [Pixel2] |
|---|---|---|---|---|---|
| 1:1 binding | 4 (± 1) ×104 | 2.1 (± 0.2) ×10−3 | / | / | 1.8 |
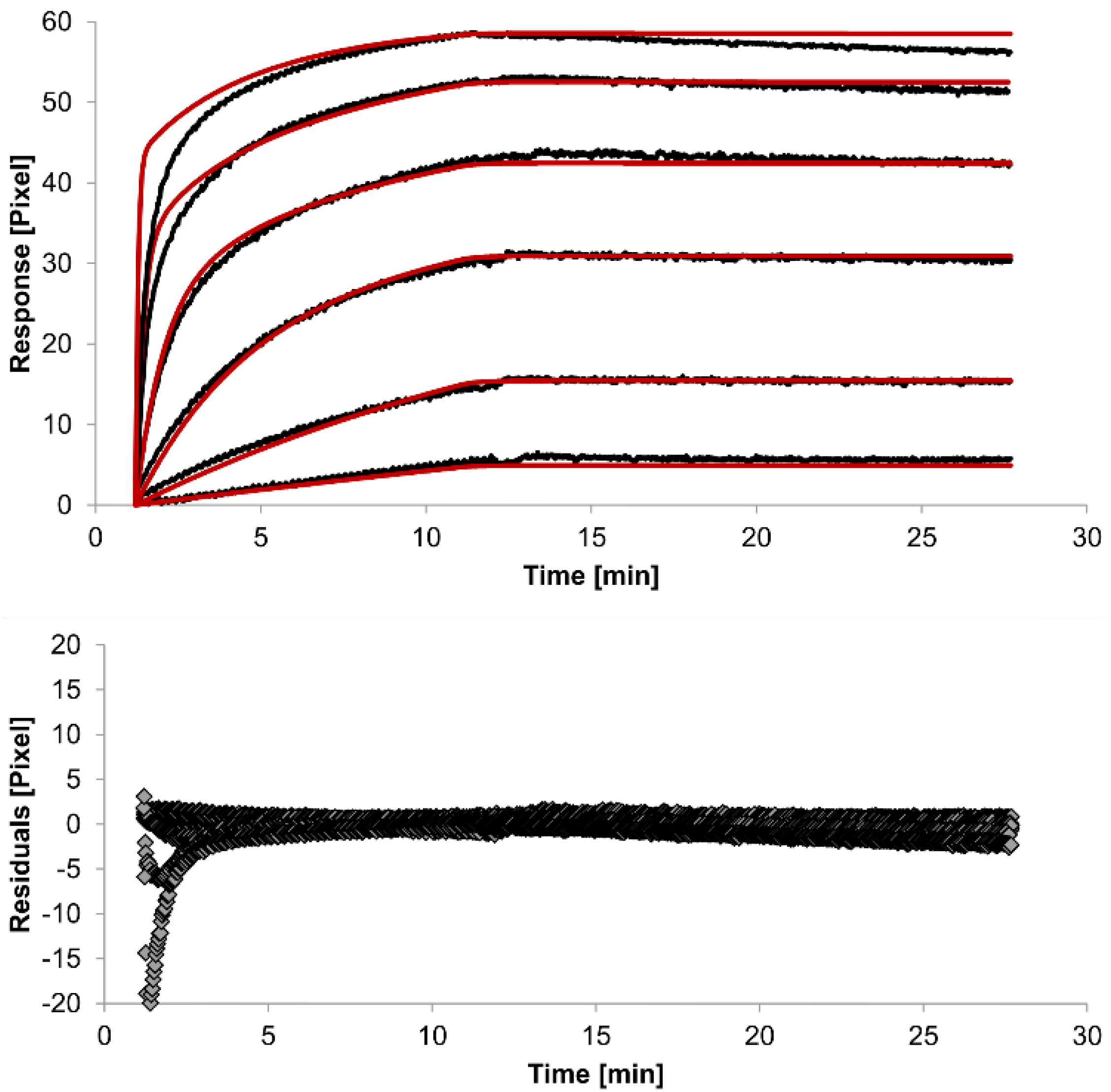
| heterogeneous analyte | 2.3 (± 0.9) ×104 | 1.2 (± 0.4) ×10−2 | 5 (± 2) ×104 | 7 (± 4) ×10−4 | 0.1 |
| bivalent analyte | 3.1 (± 0.7) ×104 | 1.8 (± 0.7) ×10−4 | 1.2 (± 0.7) ×106 [Pixel−1s−1] | 9 (± 5) ×10−3 | 0.8 |
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Liedberg, B.; Nylander, C.; Lunström, I. Surface plasmon resonance for gas detection and biosensing. Sens. Actuator 1983, 4, 299–304. [Google Scholar] [CrossRef]
- Fan, X.; White, I.M.; Shopova, S.I.; Zhu, H.; Suter, J.D.; Sun, Y. Sensitive optical biosensors for unlabeled targets: A review. Anal. Chim. Acta 2008, 620, 8–26. [Google Scholar] [CrossRef] [PubMed]
- Homola, J. Surface plasmon resonance sensors for detection of chemical and biological species. Chem. Rev. 2008, 108, 462–493. [Google Scholar] [CrossRef] [PubMed]
- Kretschmann, E.; Raether, H. Radiative decay of nonradiative surface plasmons excited by light. Z. Naturforsch. A 1968, 23, 2135–2136. [Google Scholar]
- Shankaran, D.R.; Gobi, K.V.; Miura, N. Recent advancements in surface plasmon resonance immunosensors for detection of small molecules of biomedical, food and environmental interest. Sens. Actuator B Chem. 2007, 121, 158–177. [Google Scholar] [CrossRef]
- Homola, J. Surface Plasmon Resonance Based Sensors; Springer: Berlin, Germany, 2006. [Google Scholar]
- Henseleit, A.; Schmieder, S.; Bley, T.; Sonntag, F.; Schilling, N.; Quenzel, P.; Danz, N.; Klotzbach, U.; Boschke, E. A compact and rapid aptasensor platform based on surface plasmon resonance. Eng. Life Sci. 2011, 11, 573–579. [Google Scholar] [CrossRef]
- Bernardi, M.; Maggioli, C.; Zaccherini, G. Human Albumin in the Management of Complications of Liver Cirrhosis. In Annual Update in Intensive Care and Emergency Medicine 2012; Vincent, J.-L., Ed.; Annual Update in Intensive Care and Emergency Medicine; Springer Berlin Heidelberg: Heidelberg, Germany, 2012; Volume 2012, pp. 421–430. [Google Scholar]
- Kim, S.-J.; Rhee, H.-W.; Park, H.-J.; Kim, H.-Y.; Kim, H.-S.; Hong, J.-I. Fluorescent probes designed for detecting human serum albumin on the basis of its pseudo-esterase activity. Bioorg. Med. Chem. Lett. 2013, 23, 2093–2097. [Google Scholar] [CrossRef] [PubMed]
- Taverna, M.; Marie, A.-L.; Mira, J.-P.; Guidet, B. Specific antioxidant properties of human serum albumin. Ann. Intensive Care 2013, 3. [Google Scholar] [CrossRef] [PubMed]
- Colmenarejo, G.; Alvarez-Pedraglio, A.; Lavandera, J.L. Cheminformatic models to predict binding affinities to human serum albumin. J. Med. Chem. 2001, 44, 4370–4378. [Google Scholar] [CrossRef] [PubMed]
- Fanali, G.; di Masi, A.; Trezza, V.; Marino, M.; Fasano, M.; Ascenzi, P. Human serum albumin: From bench to bedside. Mol. Aspects Med. 2012, 33, 209–290. [Google Scholar] [CrossRef] [PubMed]
- Sugio, S.; Kashima, A.; Mochizuki, S.; Noda, M.; Kobayashi, K. Crystal structure of human serum albumin at 2.5 A resolution. Protein Eng. 1999, 12, 439–446. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.; Sankar, C.; Rajasree, P.H. Human serum albumin nanoparticles for enhanced drug delivery to treat breast cancer: Preparation and In vitro assessment. Int. J. Pharm. Life Sci. 2012, 3, 2055–2063. [Google Scholar]
- Soni, N.; Margarson, M. Albumin. Where are we now? Curr. Anaesth. Crit. Care 2004, 15, 61–68. [Google Scholar] [CrossRef]
- Yang, M.-H.; Jong, S.-B.; Chung, T.-W.; Huang, Y.-F.; Ty, Y.-C. Quartz crystal microbalance in clinical application. In Biosensors for Health, Environment and Biosecurity; Serra, P.A., Ed.; InTech: Rijeka, Croatia, 2011; pp. 257–272. [Google Scholar]
- Vincent, J.-L.; Dubois, M.-J.; Navickis, R.J.; Wilkes, M.M. Hypoalbuminemia in Acute Illness: Is There a Rationale for Intervention? Ann. Surg. 2003, 237, 319–334. [Google Scholar] [PubMed]
- Guzzardi, M.A.; Vozzi, F.; Ahluwalia, A.D. Study of the crosstalk between hepatocytes and endothelial cells using a novel multicompartmental bioreactor: a comparison between connected cultures and cocultures. Tissue Eng. Part. A 2009, 15, 3635–3644. [Google Scholar] [CrossRef] [PubMed]
- Vinci, B.; Cavallone, D.; Vozzi, G.; Mazzei, D.; Domenici, C.; Brunetto, M.; Ahluwalia, A. In vitro liver model using microfabricated scaffolds in a modular bioreactor. Biotechnol. J. 2010, 5, 232–241. [Google Scholar] [CrossRef] [PubMed]
- Wagner, I.; Materne, E.-M.; Brincker, S.; Süßbier, U.; Frädrich, C.; Busek, M.; Sonntag, F.; Sakharov, D.A.; Trushkin, E.V.; Tonevitsky, A.G.; et al. A dynamic multi-organ-chip for long-term cultivation and substance testing proven by 3D human liver and skin tissue co-culture. Lab. Chip 2013, 13, 3538–3547. [Google Scholar] [CrossRef] [PubMed]
- Linke, K.; Schanz, J.; Hansmann, J.; Walles, T.; Brunner, H.; Mertsching, H. Engineered Liver-Like Tissue on a Capillarized Matrix for Applied Research. Tissue Eng. 2007, 13, 2699–2707. [Google Scholar] [CrossRef] [PubMed]
- Rich, R.L.; Myszka, D.G. Grading the commercial optical biosensor literature-Class of 2008: “The Mighty Binders.”. J. Mol. Recognit. 2010, 23, 1–64. [Google Scholar] [CrossRef] [PubMed]
- Drake, A.W.; Myszka, D.G.; Klakamp, S.L. Characterizing high-affinity antigen/antibody complexes by kinetic- and equilibrium-based methods. Anal. Biochem. 2004, 328, 35–43. [Google Scholar] [CrossRef] [PubMed]
- Müller, K.M.; Arndt, K.M.; Plückthun, A. Model and simulation of multivalent binding to fixed ligands. Anal. Biochem. 1998, 261, 149–158. [Google Scholar] [CrossRef] [PubMed]
- Birch, J.R.; Lennox, E.S. Monoclonal Antibodies: Principles and Applications; Wiley-Liss: New York, NY, USA, 1995. [Google Scholar]
- Baumann, S.; Grob, P.; Stuart, F.; Pertlik, D.; Ackermann, M.; Suter, M. Indirect immobilization of recombinant proteins to a solid phase using the albumin binding domain of streptococcal protein G and immobilized albumin. J. Immunol. Methods 1998, 221, 95–106. [Google Scholar] [CrossRef] [PubMed]
- Karlsson, R. SPR for molecular interaction analysis: A review of emerging application areas. J. Mol. Recognit. 2004, 17, 151–161. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Henseleit, A.; Pohl, C.; Kaltenbach, H.-M.; Hettwer, K.; Simon, K.; Uhlig, S.; Haustein, N.; Bley, T.; Boschke, E. Kinetic Analyses of Data from a Human Serum Albumin Assay Using the liSPR System. Biosensors 2015, 5, 27-36. https://doi.org/10.3390/bios5010027
Henseleit A, Pohl C, Kaltenbach H-M, Hettwer K, Simon K, Uhlig S, Haustein N, Bley T, Boschke E. Kinetic Analyses of Data from a Human Serum Albumin Assay Using the liSPR System. Biosensors. 2015; 5(1):27-36. https://doi.org/10.3390/bios5010027
Chicago/Turabian StyleHenseleit, Anja, Carolin Pohl, Hans-Michael Kaltenbach, Karina Hettwer, Kirsten Simon, Steffen Uhlig, Natalie Haustein, Thomas Bley, and Elke Boschke. 2015. "Kinetic Analyses of Data from a Human Serum Albumin Assay Using the liSPR System" Biosensors 5, no. 1: 27-36. https://doi.org/10.3390/bios5010027
APA StyleHenseleit, A., Pohl, C., Kaltenbach, H.-M., Hettwer, K., Simon, K., Uhlig, S., Haustein, N., Bley, T., & Boschke, E. (2015). Kinetic Analyses of Data from a Human Serum Albumin Assay Using the liSPR System. Biosensors, 5(1), 27-36. https://doi.org/10.3390/bios5010027





