Reagentless Vis-NIR Spectroscopy Point-of-Care for Feline Total White Blood Cell Counts
Abstract
1. Introduction
1.1. White Blood Cells and Blood Spectroscopy
1.2. Data Augmentation
2. Methods
2.1. Hemogram Analysis
2.2. Spectroscopy
2.3. Spectral Data Augmentation
2.4. Benchmarking
- Partial least squares regression (PLS): PLS maximizes the covariance between the spectra matrix and the blood hemogram matrix , by determining the eigenvectors of [46,47]. The method forces the latent structures of spectra and hemogram ( and ) to be equal (NIPALS or SVD algorithm), for the correspondent basis and [46], where and . It further deflates sequential orthogonal eigenvectors of the remaining information in . The number of latent variables (LV) is optimized by the cross-validation of the minimal predicted sum of squares. PLS uses an oblique projection to determine the coefficients in , which, albeit it linear combination of LVs, can model spectral information non-linearities [47,48].
- Local PLS (LocPLS): breaks down the complexity of the feature space by KNN classification based on the similarity of spectral features given by the Euclidean distance between sample coordinates. An ensemble of PLS models is generated for each cluster. The dimensions of the feature space and number of the cluster was optimized by cross-validation/hold-out samples [49].
- Self-learning Artificial Intelligence (SLAI): uses a two-step approach that (a) features space optimization for providing equivalence between the spectral information () and blood hemogram () latent spaces; and (b) performs a local covariance mode (CovM) search for unscrambling groups of samples that have the same type of interferences, providing a direct quantification between spectra and nutrients with a single dimension (or eigenvector) exhibiting stable co-variance () [1].
2.5. Total Error and Bias Analysis
3. Results and Discussion
3.1. WBC Blood Spectroscopy
3.2. Significance and Representability
3.3. WBC Quantification
3.4. Bias–Variance Analysis
3.5. CovMs Interpretation
Towards Information Specificity
- Using independent spectral information allows us to diagnose natural correlations, which are highly common in complex biological information. A natural correlation between WBCs and RBCs can lead to dataset specificity but not WBC specificity; that is, WBCs are inferred by the intrinsic correlation of WBCs and RBCs. The isolation of WBC information provides a cause–effect model, preserving both information equivalence and statistical significance;
- The isolation of independent non-dominant information minimizes interference and maximizes WBC information. In this way, the sensitivity of WBC detection and quantification is increased (e.g., 85% at 7 × cell/L, Figure 4c).
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| ASVCP | American Society of Veterinary Clinical Pathology |
| BBL | Beer–Lambert Law |
| Bil | Bilirubin |
| BT | Blind Testing |
| CovM | Covariance Mode |
| CV | Cross-validation |
| EDTA | Ethylenediaminetetraacetic Acid |
| Hgb | Hemoglobin |
| HO | Hold out samples |
| HTC | Hematocrit |
| IoT | Internet of Things |
| LocPLS | Local Partial Least Squares |
| LV | Latent Variables |
| MAPE | Mean Absolute Percentage Error |
| PCV | Packed Cell Volume |
| PLS | Partial Least Squares |
| PLT | Platelets |
| POC | Point-of-care |
| R | Pearson Correlation |
| RBCs | Red Blood Cells |
| RI | Reference Interval |
| ROI | Region of Interest |
| RWD | Real World Data |
| SLAI | Self-learning Artificial Intelligence |
| SSD | Synthetic Spectroscopy Data |
| TAE | Total Allowable Error |
| TE | Total Error |
| TV | Target Values |
| UV-Vis | Ultraviolet–visible |
| Vis-SWNIR | Visible Short-Wave Infrared |
| WBCs | White Blood Cells |
References
- Barroso, T.G.; Ribeiro, L.; Gregório, H.; Santos, F.; Martins, R.C. Point-of-care Vis-SWNIR spectroscopy towards reagent-less hemogram analysis. Sens. Actuators B Chem. 2021, 343, 130138. [Google Scholar] [CrossRef]
- Barroso, T.G.; Ribeiro, L.; Gregório, H.; Monteiro-Silva, F.; Neves dos Santos, F.; Martins, R.C. Point-of-Care Using Vis-NIR Spectroscopy for White Blood Cell Count Analysis. Chemosensors 2022, 10, 460. [Google Scholar] [CrossRef]
- Pasquini, C. Near infrared spectroscopy: A mature analytical technique with new perspectives—A review. Anal. Chim. Acta 2018, 1026, 8–36. [Google Scholar] [CrossRef] [PubMed]
- Olinger, J.M.; Griffiths, P.R. Quantitative effects of an absorbing matrix on near-infrared diffuse reflectance spectra. Anal. Chem. 1988, 60, 2427–2435. [Google Scholar] [CrossRef]
- Sparén, A.; Hartman, M.; Fransson, M.; Johansson, J.; Svensson, O. Matrix Effects in Quantitative Assessment of Pharmaceutical Tablets Using Transmission Raman and Near-Infrared (NIR) Spectroscopy. Appl. Spectrosc. 2015, 69, 580–589. [Google Scholar] [CrossRef] [PubMed]
- Nishat, S.; Jafry, A.T.; Martinez, A.W.; Awan, F.R. Paper-based microfluidics: Simplified fabrication and assay methods. Sens. Actuators B Chem. 2021, 336, 129681. [Google Scholar] [CrossRef]
- Jiang, K.; Wu, J.; Qiu, Y.; Go, Y.Y.; Ban, K.; Park, H.J.; Lee, J.H. Plasmonic colorimetric PCR for Rapid molecular diagnostic assays. Sens. Actuators B Chem. 2021, 337, 129762. [Google Scholar] [CrossRef]
- Lewińska, I.; Speichert, M.; Granica, M.; Tymecki, Ł. Colorimetric point-of-care paper-based sensors for urinary creatinine with smartphone readout. Sens. Actuators B Chem. 2021, 340, 129915. [Google Scholar] [CrossRef]
- Zhou, L.; Vestri, A.; Marchesano, V.; Rippa, M.; Sagnelli, D.; Picazio, G.; Fusco, G.; Han, J.; Zhou, J.; Petti, L. The Label-Free Detection and Identification of SARS-CoV-2 Using Surface-Enhanced Raman Spectroscopy and Principal Component Analysis. Biosensors 2023, 13, 1014. [Google Scholar] [CrossRef]
- Popov, E.; Polishchuk, A.; Kovalev, A.; Vitkin, V. Raman Spectroscopy for Urea Breath Test. Biosensors 2023, 13, 609. [Google Scholar] [CrossRef]
- Monteiro-Silva, F.; Jorge, P.A.S.; Martins, R.C. Optical Sensing of Nitrogen, Phosphorus and Potassium: A Spectrophotometrical Approach toward Smart Nutrient Deployment. Chemosensors 2019, 7, 51. [Google Scholar] [CrossRef]
- Martins, R.C.; Barroso, T.G.; Jorge, P.; Cunha, M.; Santos, F. Unscrambling spectral interference and matrix effects in Vitis vinifera Vis-NIR spectroscopy: Towards analytical grade ‘in vivo’ sugars and acids quantification. Comput. Electron. Agric. 2022, 194, 106710. [Google Scholar] [CrossRef]
- Lin, K.; Cheng, D.L.P.; Huang, Z. Optical diagnosis of laryngeal cancer using high wavenumber Raman spectroscopy. Biosens. Bioelectron. 2012, 35, 213–217. [Google Scholar] [CrossRef]
- Coelho, B.F.O.; Nunes, S.L.P.; de França, C.A.; dos Santos Costa, D.; do Carmo, R.F.; Prates, R.M.; Filho, E.F.S.; Ramos, R.P. On the feasibility of Vis–NIR spectroscopy and machine learning for real time SARS-CoV-2 detection. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2024, 308, 123735. [Google Scholar] [CrossRef] [PubMed]
- Burns, D.; Ciurczak, E. Handbook of Near Infrared Analysis, 2nd ed.; Marcel Dekker, Inc.: New York, NY, USA, 2001. [Google Scholar]
- Barroso, T.G.; Martins, R.C.; Fernandes, E.; Cardoso, S.; Rivas, J.; Freitas, P.P. Detection of BCG bacteria using a magnetoresistive biosensor: A step towards a fully electronic platform for tuberculosis point-of-care detection. Biosens. Bioelectron. 2018, 100, 259–265. [Google Scholar] [CrossRef] [PubMed]
- Tvedten, H.; Raskin, R.E. 4—Leukocyte Disorders. In Small Animal Clinical Diagnosis by Laboratory Methods, 5th ed.; Willard, M.D., Tvedten, H., Eds.; W.B. Saunders: Saint Louis, MO, USA, 2012; pp. 63–91. [Google Scholar] [CrossRef]
- Burton, A.G.; Jandrey, K.E. Leukocytosis and Leukopenia. In Textbook of Small Animal Emergency Medicine; John Wiley & Sons, Ltd.: Hoboken, NJ, USA, 2018; Chapter 64; pp. 405–412. [Google Scholar] [CrossRef]
- Philo, J.; Adams, M.; Schuster, T.M. Association-dependent absorption spectra of oxyhemoglobin A and its subunits. J. Biol. Chem. 1981, 256, 7917–7924. [Google Scholar] [CrossRef]
- Tan, H.; Liao, S.; Pan, T.; Zhang, J.; Chen, J. Rapid and simultaneous analysis of direct and indirect bilirubin indicators in serum through reagent-free visible-near-infrared spectroscopy combined with chemometrics. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2020, 233, 118215. [Google Scholar] [CrossRef]
- Chabot-Richards, D.S.; George, T.I. White Blood Cell Counts: Reference Methodology. Clin. Lab. Med. 2015, 35, 11–24. [Google Scholar] [CrossRef]
- Rishniw, M.; Pion, P.D. Evaluation of performance of veterinary in-clinic hematology analyzers. Vet. Clin. Pathol. 2016, 45, 604–614. [Google Scholar] [CrossRef]
- Riond, B.; Waßmuth, A.K.; Hartnack, S.; Hofmann-Lehmann, R.; Lutz, H. Study on the kinetics and influence of feline platelet aggregation and deaggregation. BMC Vet. Res. 2015, 11, 276. [Google Scholar] [CrossRef]
- Gulati, G.; Uppal, G.; Gong, J. Unreliable Automated Complete Blood Count Results: Causes, Recognition, and Resolution. Ann. Lab. Med. 2022, 42, 515–530. [Google Scholar] [CrossRef] [PubMed]
- Nabity, M.B.; Harr, K.E.; Camus, M.S.; Flatland, B.; Vap, L. ASVCP guidelines: Allowable total error hematology. Vet. Clin. Pathol. 2018, 47, 9–21. [Google Scholar] [CrossRef] [PubMed]
- Harvey, J.W. The feline blood film: 2. Leukocyte and platelet morphology. J. Feline Med. Surg. 2017, 19, 747–757. [Google Scholar] [CrossRef] [PubMed]
- Meredith, A.M.; Bienzle, D.; Beeler-Marfisi, J. What is your diagnosis? A feline blood smear. Vet. Clin. Pathol. 2019, 48, 128–130. [Google Scholar] [CrossRef] [PubMed]
- Terent’yeva, Y.G.; Yashchuk, V.M.; Zaika, L.A.; Snitserova, O.M.; Losytsky, M.Y. The manifestation of optical centers in UV–Vis absorption and luminescence spectra of white blood human cells. Methods Appl. Fluoresc. 2016, 4, 044010. [Google Scholar] [CrossRef]
- Ramesh, J.; Kapelushnik, J.; Mordehai, J.; Moser, A.; Huleihel, M.; Erukhimovitch, V.; Levi, C.; Mordechai, S. Novel methodology for the follow-up of acute lymphoblastic leukemia using FTIR microspectroscopy. J. Biochem. Biophys. Methods 2002, 51, 251–261. [Google Scholar] [CrossRef]
- Ramesh, J.; Huleihel, M.; Mordehai, J.; Moser, A.; Erukhimovich, V.; Levi, C.; Kapelushnik, J.; Mordechai, S. Preliminary results of evaluation of progress in chemotherapy for childhood leukemia patients employing Fourier-transform infrared microspectroscopy and cluster analysis. J. Lab. Clin. Med. 2003, 141, 385–394. [Google Scholar] [CrossRef]
- Liu, W.; Howarth, M.; Greytak, A.B.; Zheng, Y.; Nocera, D.G.; Ting, A.Y.; Bawendi, M.G. Compact biocompatible quantum dots functionalized for cellular imaging. J. Am. Chem. Soc. 2007, 130, 1274–1284. [Google Scholar] [CrossRef]
- Managò, S.; Valente, C.; Mirabelli, P.; Circolo, D.; Basile, F.; Corda, D.; De Luca, A.C. A reliable Raman-spectroscopy-based approach for diagnosis, classification and follow-up of B-cell acute lymphoblastic leukemia. Sci. Rep. 2016, 6, 24821. [Google Scholar] [CrossRef]
- Lima, A.M.F.; Silva, J.D.P.d.; Daniel, C.R. Diagnosis, classification and monitoring of leukemia based on Raman spectroscopy. Res. Soc. Dev. 2021, 10, e67101421657. [Google Scholar] [CrossRef]
- Sahu, R.K.; Zelig, U.; Huleihel, M.; Brosh, N.; Talyshinsky, M.; Ben-Harosh, M.; Mordechai, S.; Kapelushnik, J. Continuous monitoring of WBC (biochemistry) in an adult leukemia patient using advanced FTIR-spectroscopy. Leuk. Res. 2006, 30, 687–693. [Google Scholar] [CrossRef] [PubMed]
- Zelig, U.; Mordechai, S.; Shubinsky, G.; Sahu, R.K.; Huleihel, M.; Leibovitz, E.; Nathan, I.; Kapelushnik, J. Pre-screening and follow-up of childhood acute leukemia using biochemical infrared analysis of peripheral blood mononuclear cells. Biochim. Biophys. Acta (BBA)-Gen. Subj. 2011, 1810, 827–835. [Google Scholar] [CrossRef] [PubMed]
- Chaber, R.; Kowal, A.; Jakubczyk, P.; Arthur, C.; Łach, K.; Wojnarowska-Nowak, R.; Kusz, K.; Zawlik, I.; Paszek, S.; Cebulski, J. A Preliminary Study of FTIR Spectroscopy as a Potential Non-Invasive Screening Tool for Pediatric Precursor B Lymphoblastic Leukemia. Molecules 2021, 26, 1174. [Google Scholar] [CrossRef] [PubMed]
- Kochan, K.; Bedolla, D.E.; Perez-Guaita, D.; Adegoke, J.A.; Veettil, T.C.P.; Martin, M.; Roy, S.; Pebotuwa, S.; Heraud, P.; Wood, B.R. Infrared Spectroscopy of Blood. Appl. Spectrosc. 2021, 75, 611–646. [Google Scholar] [CrossRef] [PubMed]
- Agbaria, A.H.; Beck, G.; Lapidot, I.; Rich, D.H.; Kapelushnik, J.; Mordechai, S.; Salman, A.; Huleihel, M. Diagnosis of inaccessible infections using infrared microscopy of white blood cells and machine learning algorithms. Analyst 2020, 145, 6955–6967. [Google Scholar] [CrossRef]
- Javinsky, E. Chapter 25—Hematology and Immune-Related Disorders. In The Cat; Little, S.E., Ed.; W.B. Saunders: Saint Louis, MO, USA, 2012; pp. 643–703. [Google Scholar] [CrossRef]
- Silva, F.; Queirós, C.; Pinho, T.; Boaventura, J.; Santos, F.; Barroso, T.; Pereira, M.; Cunha, M.; Martins, R. Reagent-less spectroscopy towards NPK sensing for hydroponics nutrient solutions. Sens. Actuators B Chem. 2023, 395, 134442. [Google Scholar] [CrossRef]
- Nalepa, J.; Marcinkiewicz, M.; Kawulok, M. Data Augmentation for Brain-Tumor Segmentation: A Review. Front. Comput. Neurosci. 2019, 13. [Google Scholar] [CrossRef]
- Shorten, C.; Khoshgoftaar, T. A survey on Image Data Augmentation for Deep Learning. J. Big Data 2019, 6, 60. [Google Scholar] [CrossRef]
- Brown, M.; Wittwer, C. Flow Cytometry: Principles and Clinical Applications in Hematology. Clin. Chem. 2000, 46, 1221–1229. [Google Scholar] [CrossRef]
- INESCTEC. AgIoT—Modular Solution and Open Source IoT Solution for Agrofood Domain; INESCTEC: Porto, Portugal, 2020. [Google Scholar]
- Martens, H.; Stark, E. Extended multiplicative signal correction and spectral interference subtraction: New preprocessing methods for near infrared spectroscopy. J. Pharm. Biomed. Anal. 1991, 9, 625–635. [Google Scholar] [CrossRef]
- Geladi, P.; Kowalsky, B. Partial least squares regression: A tuturial. Anal. Acta 1988, 185, 1–17. [Google Scholar] [CrossRef]
- Ergon, R. Re-interpretation of NIPALS results solves PLSR inconsistency problem. J. Chemom. 2009, 23, 72–75. [Google Scholar] [CrossRef]
- Phatak, A.; Jong, S. The geometry of partial least squares. J. Chemom. 1997, 11, 311–338. [Google Scholar] [CrossRef]
- Shen, G.; Lesnoff, M.; Baeten, V.; Dardenne, P.; Davrieux, F.; Ceballos, H.; Belalcazar, J.; Dufour, D.; Yang, Z.; Han, L.; et al. Local partial least squares based on global PLS scores. J. Chemom. 2019, 33, e3117. [Google Scholar] [CrossRef]
- R-Core-Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2020; Available online: http://www.r-project.org/ (accessed on 19 December 2023).
- Harr, K.E.; Flatland, B.; Nabity, M.B.; Freeman, K.P. ASVCP Guidelines—Allowable Total Error: Biochemistry; American Society for Veterinary Clinical Pathology: Madison, WI, USA, 2013. [Google Scholar]
- Meng, F.; Alayash, A.I. Determination of extinction coefficients of human hemoglobin in various redox states. Anal. Biochem. 2017, 521, 11–19. [Google Scholar] [CrossRef]
- Cook, A.M.; Moritz, A.; Freeman, K.P.; Bauer, N. Quality requirements for veterinary hematology analyzers in small animals—A survey about veterinary experts’ requirements and objective evaluation of analyzer performance based on a meta-analysis of method validation studies: Bench top hematology analyzer. Vet. Clin. Pathol. 2016, 45, 466–476. [Google Scholar] [CrossRef]
 ) low WBCs, (
) low WBCs, ( ) high WBC examples, and (
) high WBC examples, and ( ) RBC Hgb major inteference bands (539–576 nm); (b) PCA scores of hemogram counts; and (c) Cell packing, scattering and WBC absorbance effects and impact on observed spectra; (d) PLS scores of blood spectra—translating maximum covariance to WBCs, where • SSD blood samples, • RWD blood samples, • low WBCs and • high WBCs and → hemogram PCA loadings and main gradient direction in PLS scores space.
) RBC Hgb major inteference bands (539–576 nm); (b) PCA scores of hemogram counts; and (c) Cell packing, scattering and WBC absorbance effects and impact on observed spectra; (d) PLS scores of blood spectra—translating maximum covariance to WBCs, where • SSD blood samples, • RWD blood samples, • low WBCs and • high WBCs and → hemogram PCA loadings and main gradient direction in PLS scores space.
 ) low WBCs, (
) low WBCs, ( ) high WBC examples, and (
) high WBC examples, and ( ) RBC Hgb major inteference bands (539–576 nm); (b) PCA scores of hemogram counts; and (c) Cell packing, scattering and WBC absorbance effects and impact on observed spectra; (d) PLS scores of blood spectra—translating maximum covariance to WBCs, where • SSD blood samples, • RWD blood samples, • low WBCs and • high WBCs and → hemogram PCA loadings and main gradient direction in PLS scores space.
) RBC Hgb major inteference bands (539–576 nm); (b) PCA scores of hemogram counts; and (c) Cell packing, scattering and WBC absorbance effects and impact on observed spectra; (d) PLS scores of blood spectra—translating maximum covariance to WBCs, where • SSD blood samples, • RWD blood samples, • low WBCs and • high WBCs and → hemogram PCA loadings and main gradient direction in PLS scores space.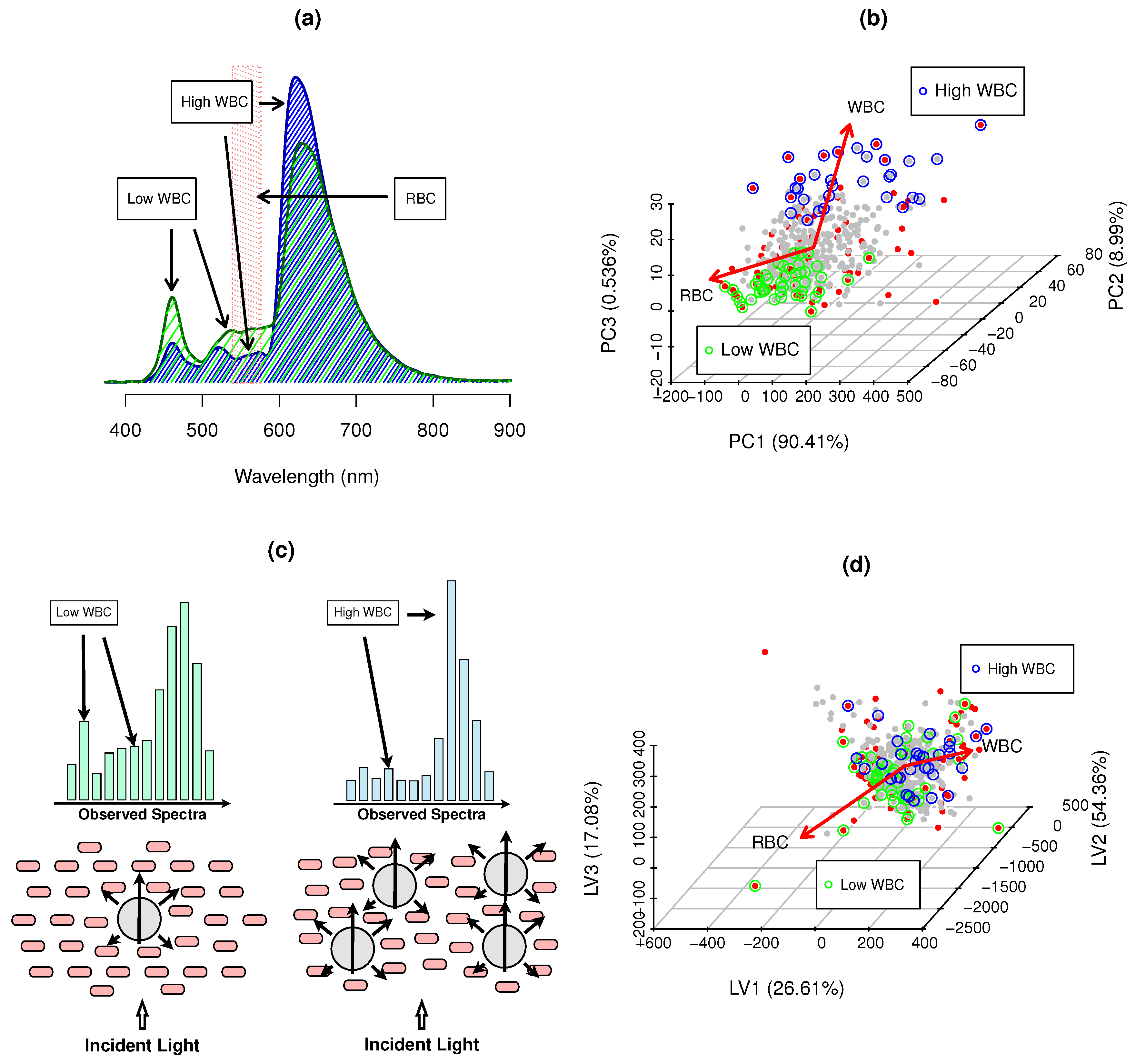
 ) represents the WBC reference interval for cats (5.5–19.5 × cells/L) and red shaded rectangle (
) represents the WBC reference interval for cats (5.5–19.5 × cells/L) and red shaded rectangle ( ) represents the ASVCP total allowable error tolerance for high WBC diagnosis.
) represents the ASVCP total allowable error tolerance for high WBC diagnosis.
 ) represents the WBC reference interval for cats (5.5–19.5 × cells/L) and red shaded rectangle (
) represents the WBC reference interval for cats (5.5–19.5 × cells/L) and red shaded rectangle ( ) represents the ASVCP total allowable error tolerance for high WBC diagnosis.
) represents the ASVCP total allowable error tolerance for high WBC diagnosis.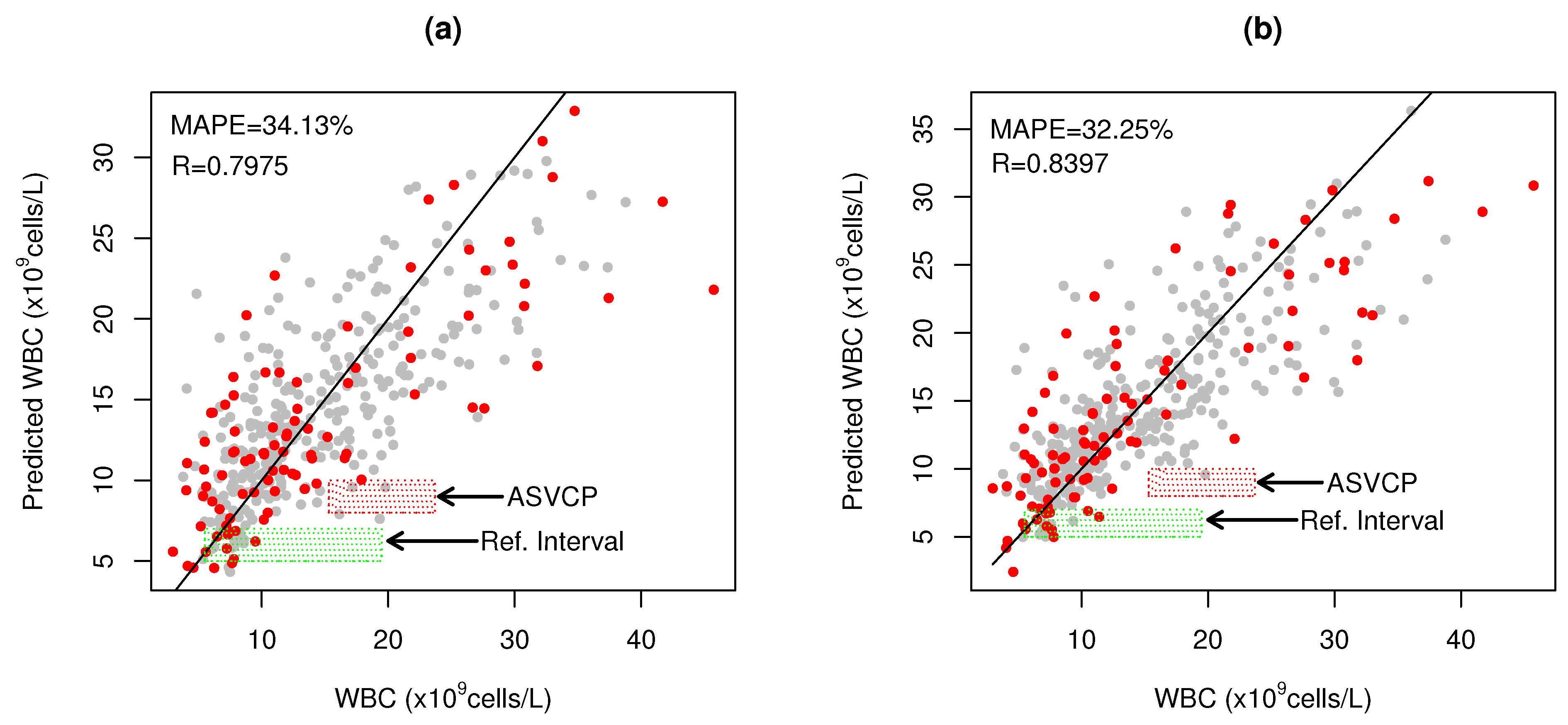
 ) and quantitative (
) and quantitative ( ) results; (b) MAPE benchmarks where (
) results; (b) MAPE benchmarks where ( ) represents the ASVCP TAE for WBCs (21.45%); and (c) POC WBC diagnostic capacity: (
) represents the ASVCP TAE for WBCs (21.45%); and (c) POC WBC diagnostic capacity: ( ) percentage of correct diagnoses within WBC value interval, (– –) accumulated percentage of correct diagnosis, (
) percentage of correct diagnoses within WBC value interval, (– –) accumulated percentage of correct diagnosis, ( ) WBC histogram distibution; and RI for WBC (
) WBC histogram distibution; and RI for WBC ( ) and (
) and ( ) TAE diagnosis error tolerance.
) TAE diagnosis error tolerance.
 ) and quantitative (
) and quantitative ( ) results; (b) MAPE benchmarks where (
) results; (b) MAPE benchmarks where ( ) represents the ASVCP TAE for WBCs (21.45%); and (c) POC WBC diagnostic capacity: (
) represents the ASVCP TAE for WBCs (21.45%); and (c) POC WBC diagnostic capacity: ( ) percentage of correct diagnoses within WBC value interval, (– –) accumulated percentage of correct diagnosis, (
) percentage of correct diagnoses within WBC value interval, (– –) accumulated percentage of correct diagnosis, ( ) WBC histogram distibution; and RI for WBC (
) WBC histogram distibution; and RI for WBC ( ) and (
) and ( ) TAE diagnosis error tolerance.
) TAE diagnosis error tolerance.
 low and
low and  high CovM), (
high CovM), ( ) and (
) and ( ) major and minor RBC/Hgb interference bands; (c,e) high and low CovM sample spectra and corresponding used ROIs to quantify WBCs; (d,f) prediction plot for high and low CovMs.
) major and minor RBC/Hgb interference bands; (c,e) high and low CovM sample spectra and corresponding used ROIs to quantify WBCs; (d,f) prediction plot for high and low CovMs.
 low and
low and  high CovM), (
high CovM), ( ) and (
) and ( ) major and minor RBC/Hgb interference bands; (c,e) high and low CovM sample spectra and corresponding used ROIs to quantify WBCs; (d,f) prediction plot for high and low CovMs.
) major and minor RBC/Hgb interference bands; (c,e) high and low CovM sample spectra and corresponding used ROIs to quantify WBCs; (d,f) prediction plot for high and low CovMs.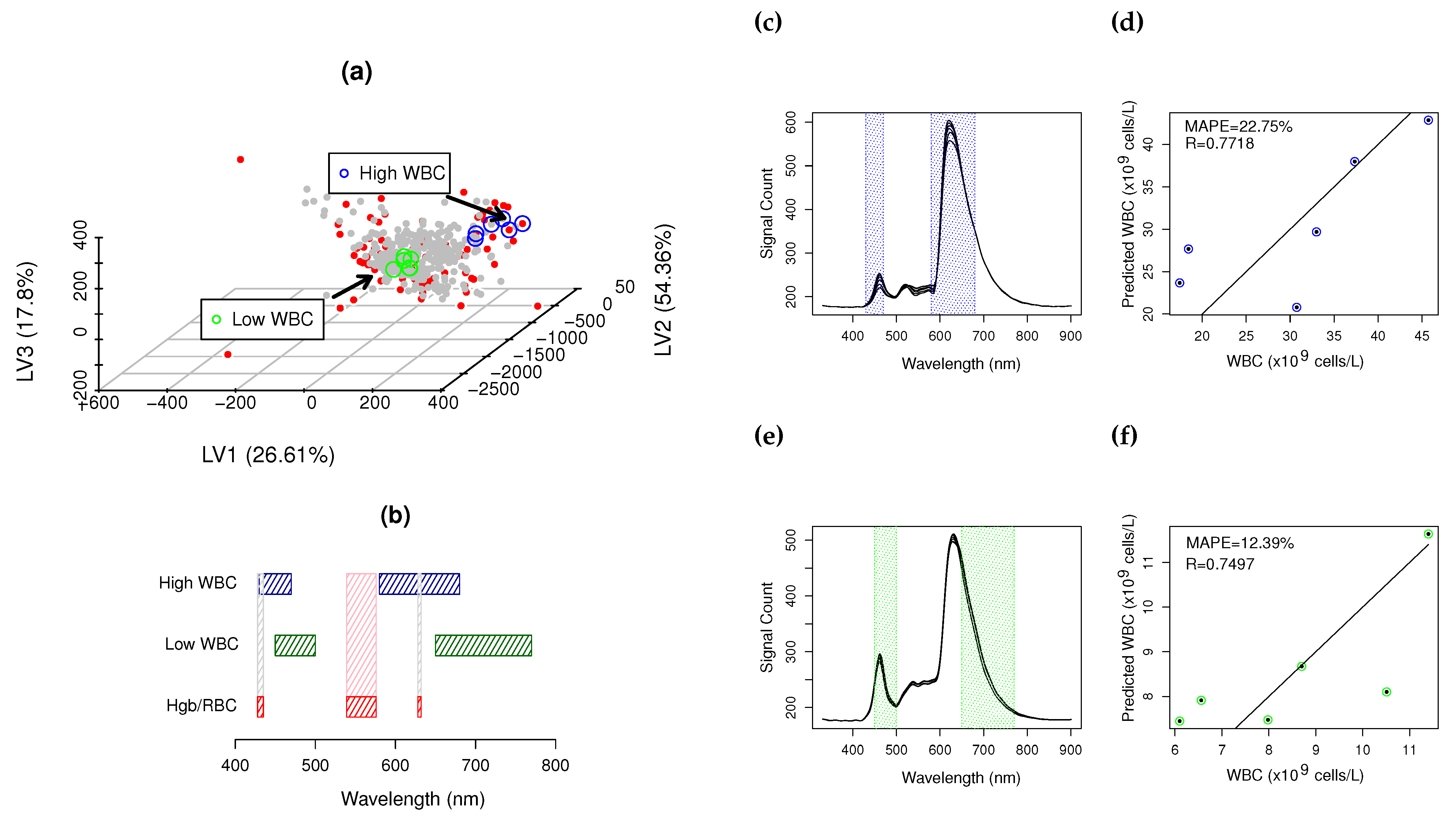
| Real World Data (RWD) | Synthetic Sample Data (SSD) | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Method | nLV | R | R2 | SE | MAPE (%) | p-Value | R | R2 | SE | MAPE (%) | p-Value |
| PLS (1) | 2 | 0.1874 | 0.0350 | 5.498 | 55.45 | 0.001 | – | – | – | – | – |
| PLS (2) | 15 | 0.5775 | 0.3336 | 6.109 | 44.56 | 1.12 × | 0.6802 | 0.4626 | 5.135 | 65.65 | <2 × |
| PLS (3) | 2 | 0.1874 | 0.0350 | 5.498 | 55.45 | 0.001 | 0.2634 | 0.0691 | 9.162 | 44.60 | 0.013 |
| PLS (4) | 13 | 0.5553 | 0.3084 | 7.925 | 57.31 | <2 × | 0.6320 | 0.3995 | 9.736 | 34.18 | <2 × |
| LocPLS (2) | 5 | 0.4546 | 0.2067 | 8.576 | 58.89 | 4.16 × | 0.6840 | 0.4679 | 4.480 | 31.19 | <2 × |
| LocPLS (4) | 7 | 0.7112 | 0.5059 | 6.673 | 46.78 | <2 × | 0.6782 | 0.4599 | 9.188 | 32.16 | <2 × |
| SLAI (2) | 1–2 | 0.7975 | 0.6361 | 5.837 | 34.13 | <2 × | 0.7369 | 0.5460 | 8.492 | 22.35 | <2 × |
| SLAI (4) | 1 | 0.8397 | 0.7051 | 5.192 | 32.25 | <2 × | 0.7723 | 0.5965 | 7.947 | 25.86 | <2 × |
| Real World Data | Synthetic Sample Data | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| % Inside RI | % Outside RI | % Inside RI | % Outside RI | |||||||||
| Method | Opt | Des | Accep | Opt | Des | Accep | Opt | Des | Accep | Opt | Des | Accep |
| PLS RWD (1) | 7.81 | 14.06 | 26.56 | 0.00 | 0.00 | 10.00 | na | na | na | na | na | na |
| PLS SSD>RWD (2) | 21.88 | 29.69 | 39.06 | 10.00 | 16.67 | 20.00 | 15.63 | 29.02 | 46.43 | 13.16 | 22.37 | 43.42 |
| PLS RWD>SSD (3) | 7.81 | 14.06 | 26.56 | 0.00 | 0.00 | 10.00 | 16.07 | 25.89 | 34.38 | 1.32 | 2.63 | 9.21 |
| PLS SSD+RWD (4) | 10.94 | 26.56 | 42.19 | 3.33 | 10.00 | 13.33 | 13.84 | 29.91 | 47.77 | 9.21 | 22.37 | 26.32 |
| LocPLS SSD>RWD (2) | 14.06 | 25.00 | 29.69 | 13.33 | 16.67 | 20.00 | 23.21 | 41.52 | 52.68 | 21.05 | 35.53 | 52.63 |
| LocPLS RWD+SSD (4) | 15.63 | 32.81 | 46.88 | 16.67 | 23.33 | 36.67 | 19.20 | 32.14 | 45.98 | 14.47 | 32.89 | 44.74 |
| SLAI SSD>RWD (2) | 20.31 | 34.38 | 50.00 | 13.33 | 30.00 | 43.33 | 23.66 | 38.39 | 54.91 | 17.11 | 26.32 | 43.42 |
| SLAI RWD+SSD (4) | 31.25 | 42.19 | 54.69 | 13.33 | 26.67 | 50.00 | 24.12 | 41.07 | 58.03 | 30.26 | 35.52 | 46.05 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Barroso, T.G.; Queirós, C.; Monteiro-Silva, F.; Santos, F.; Gregório, A.H.; Martins, R.C. Reagentless Vis-NIR Spectroscopy Point-of-Care for Feline Total White Blood Cell Counts. Biosensors 2024, 14, 53. https://doi.org/10.3390/bios14010053
Barroso TG, Queirós C, Monteiro-Silva F, Santos F, Gregório AH, Martins RC. Reagentless Vis-NIR Spectroscopy Point-of-Care for Feline Total White Blood Cell Counts. Biosensors. 2024; 14(1):53. https://doi.org/10.3390/bios14010053
Chicago/Turabian StyleBarroso, Teresa Guerra, Carla Queirós, Filipe Monteiro-Silva, Filipe Santos, António Hugo Gregório, and Rui Costa Martins. 2024. "Reagentless Vis-NIR Spectroscopy Point-of-Care for Feline Total White Blood Cell Counts" Biosensors 14, no. 1: 53. https://doi.org/10.3390/bios14010053
APA StyleBarroso, T. G., Queirós, C., Monteiro-Silva, F., Santos, F., Gregório, A. H., & Martins, R. C. (2024). Reagentless Vis-NIR Spectroscopy Point-of-Care for Feline Total White Blood Cell Counts. Biosensors, 14(1), 53. https://doi.org/10.3390/bios14010053







