An Interfacial Affinity Interaction-Based Method for Detecting HOTAIR lncRNA in Cancer Plasma Samples
Abstract
:1. Introduction
2. Materials and Methods
2.1. Reagents and Materials
2.2. Preparation of RNA from Cell Line and Ovarian Cancer Samples
2.3. Magnetic Isolation and Purification of HOTAIR lncRNA Target
2.4. Electrochemical Detection of HOTAIR lncRNAs
2.5. Quantitative Reverse-Transcription Polymerase Chain Reaction (RT-qPCR)
3. Results and Discussion
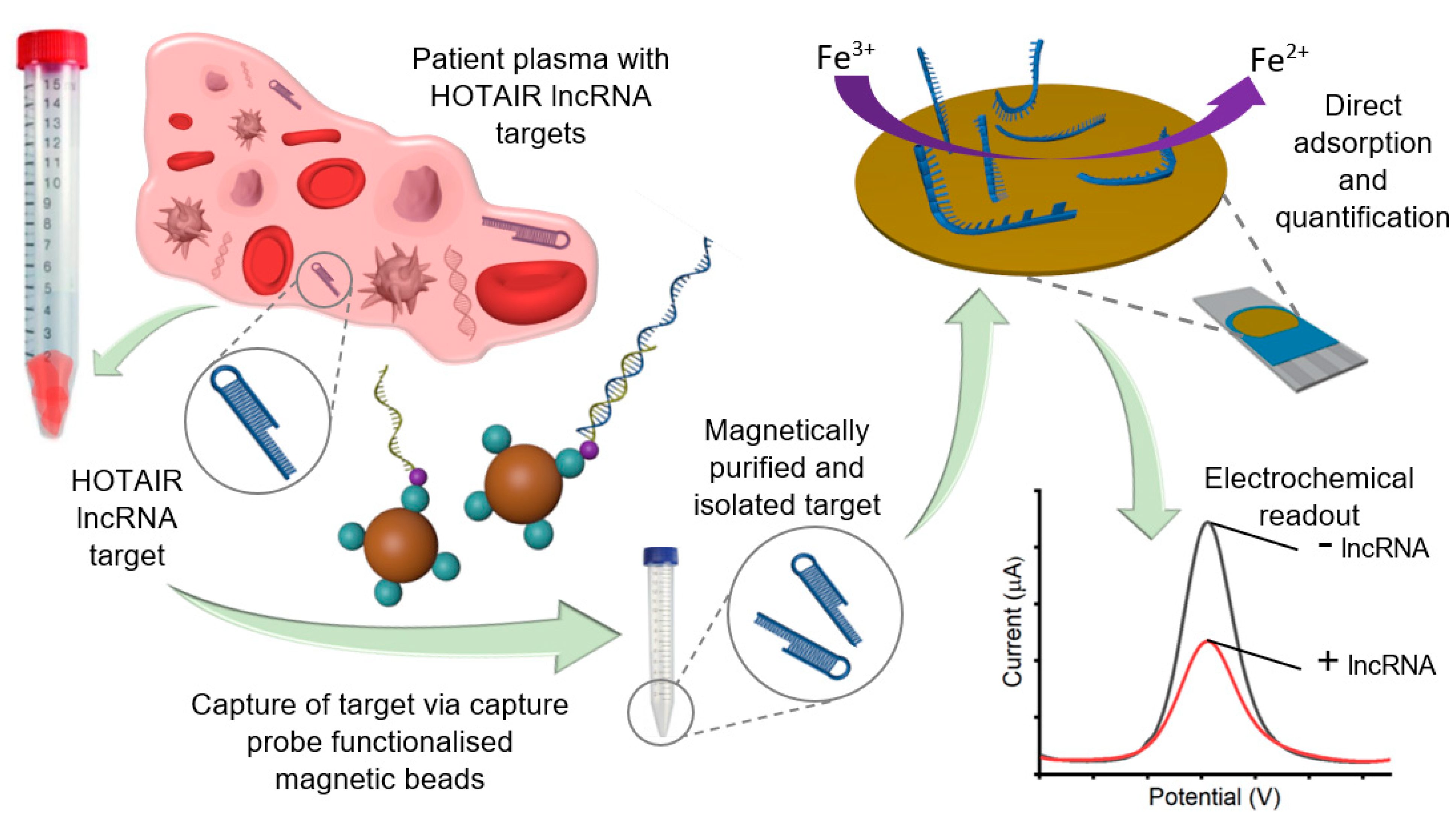
3.1. Assay Principle
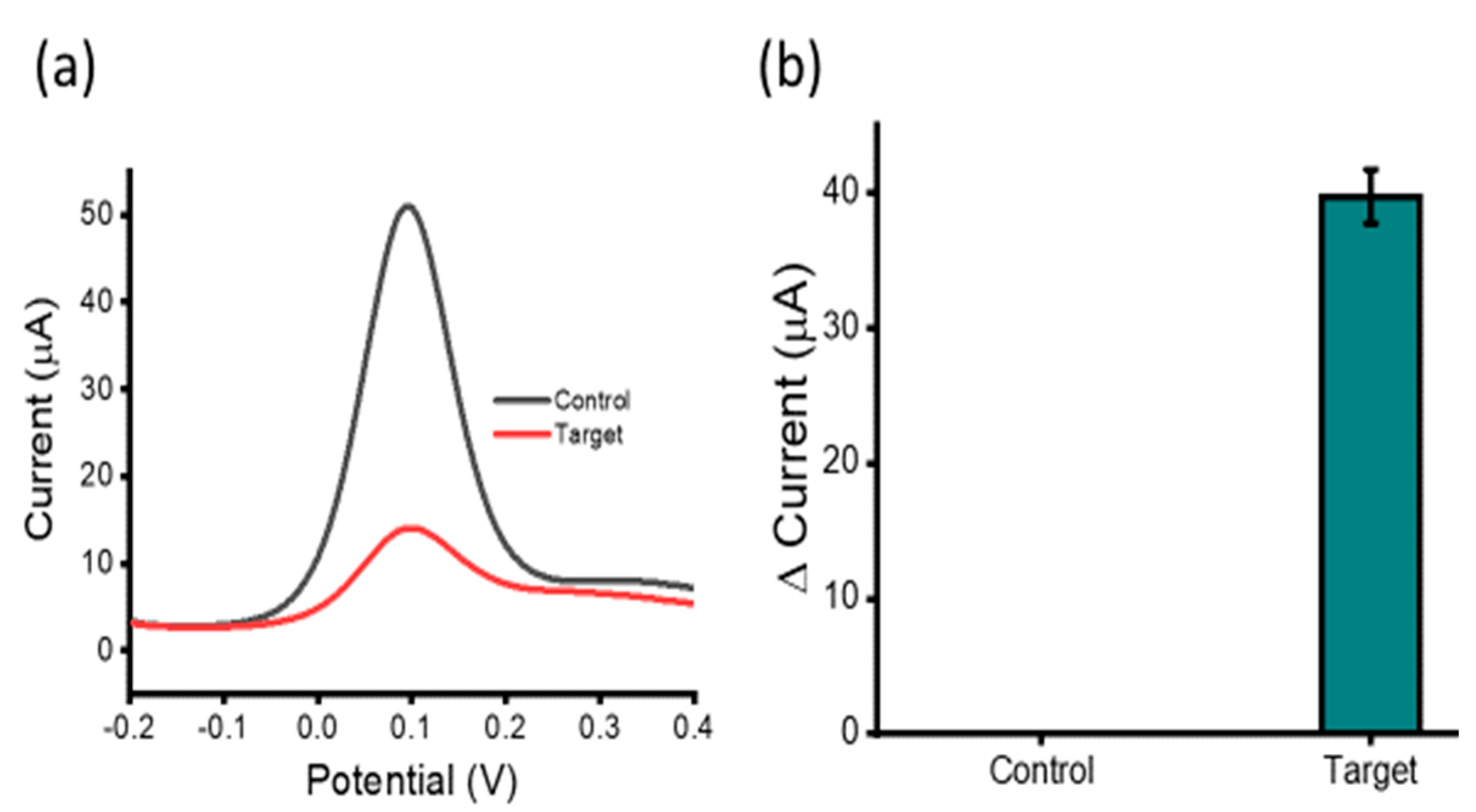
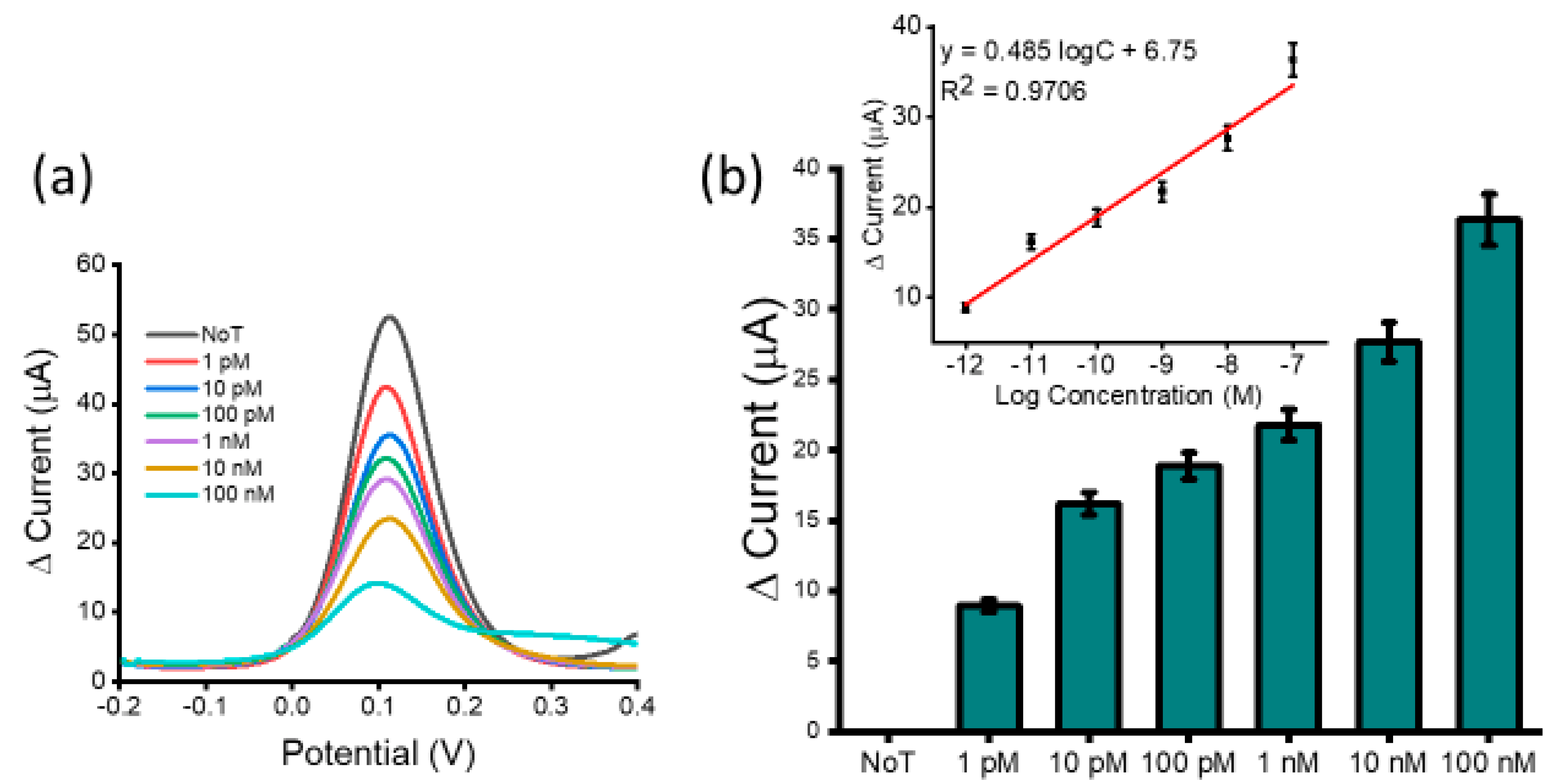
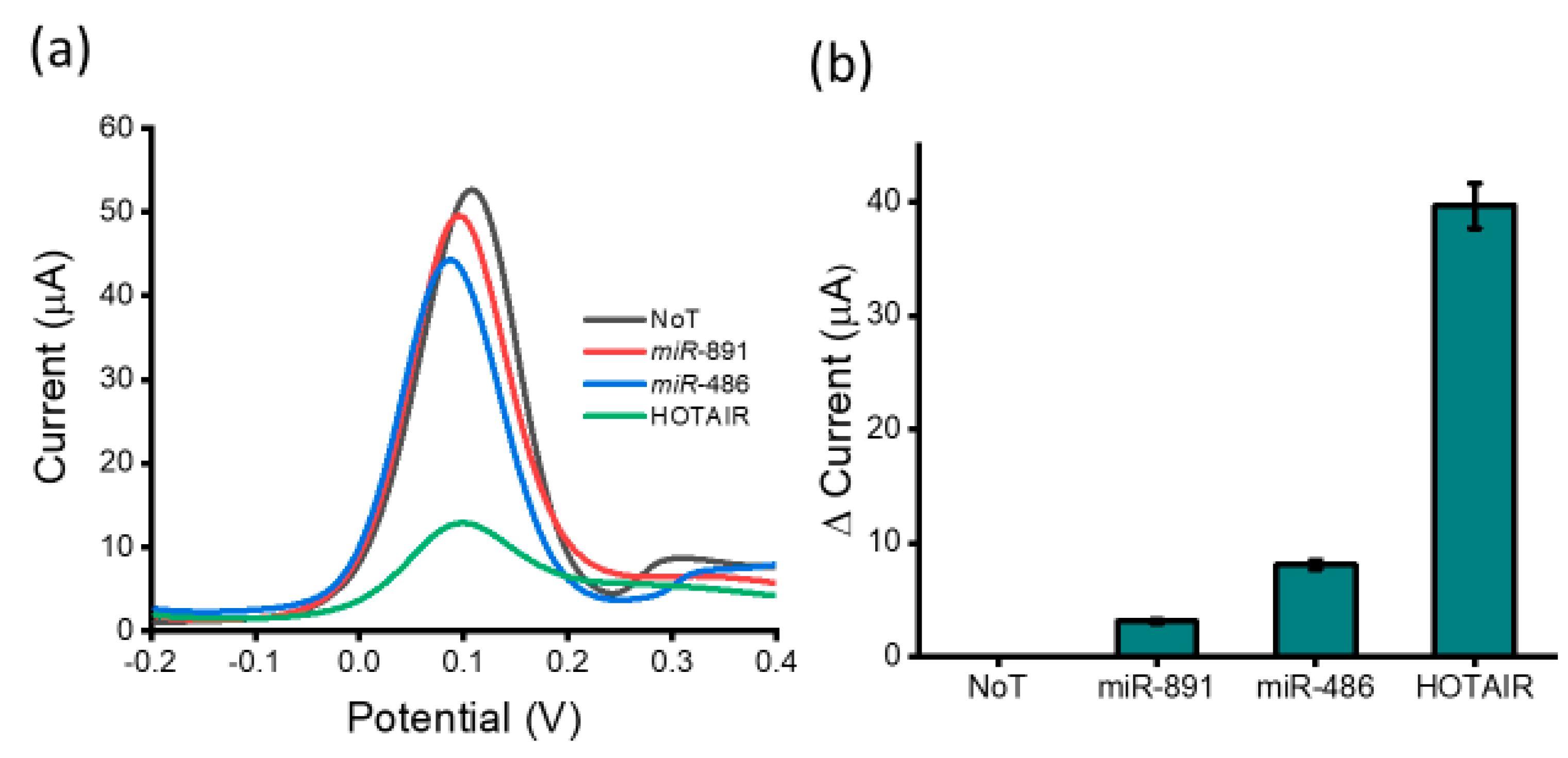
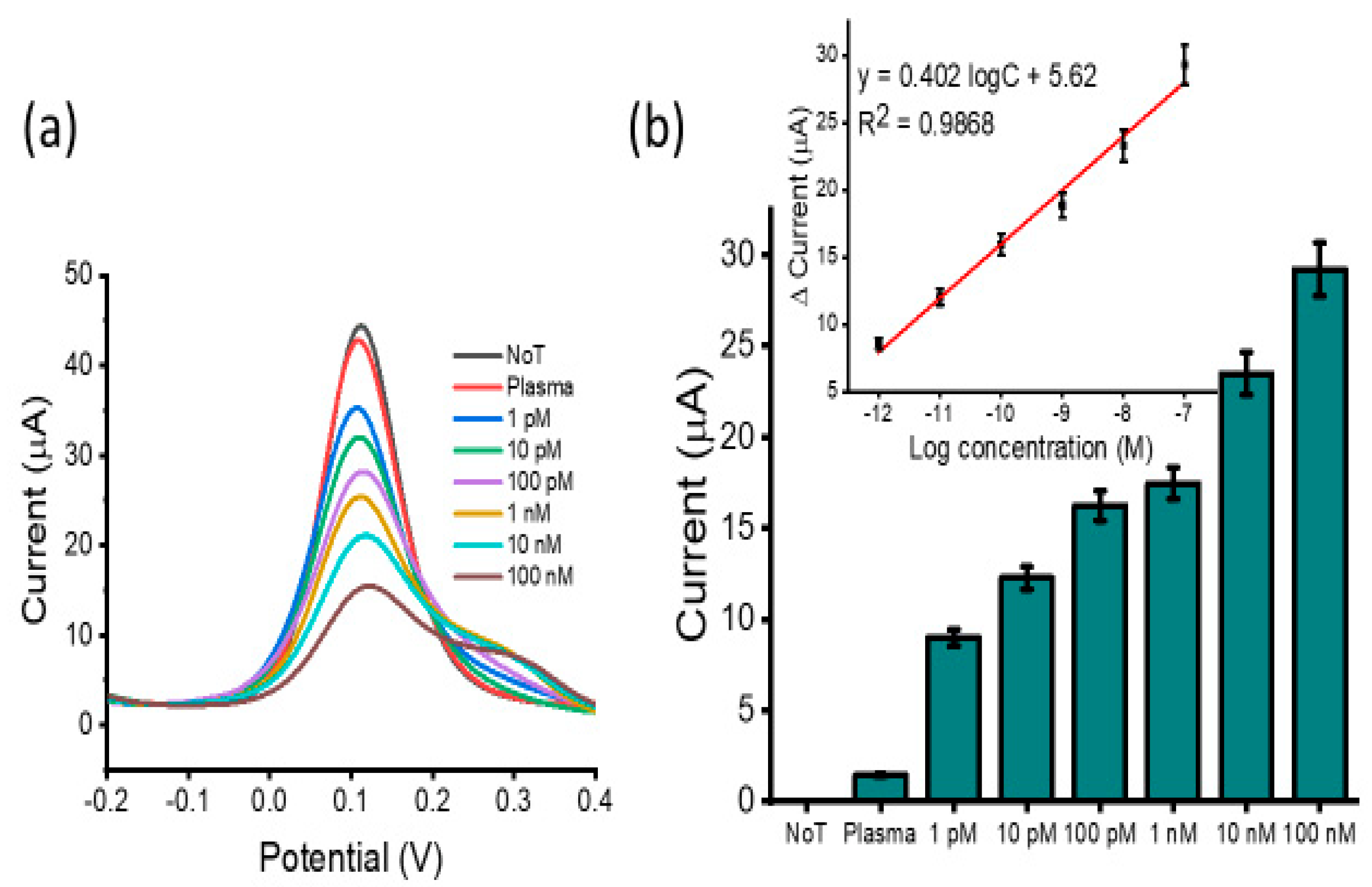
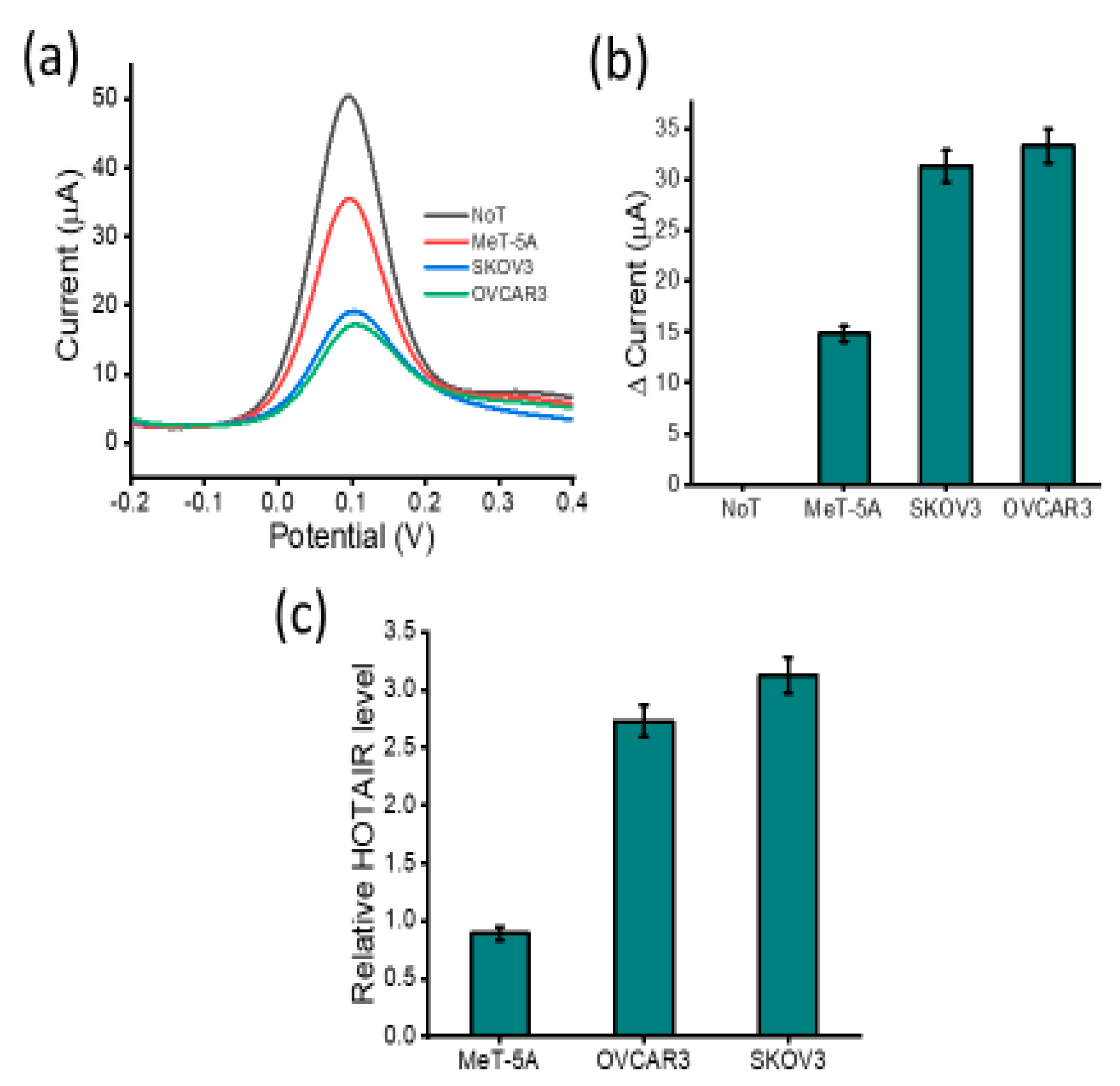
3.2. Analytical Performance of the Assay
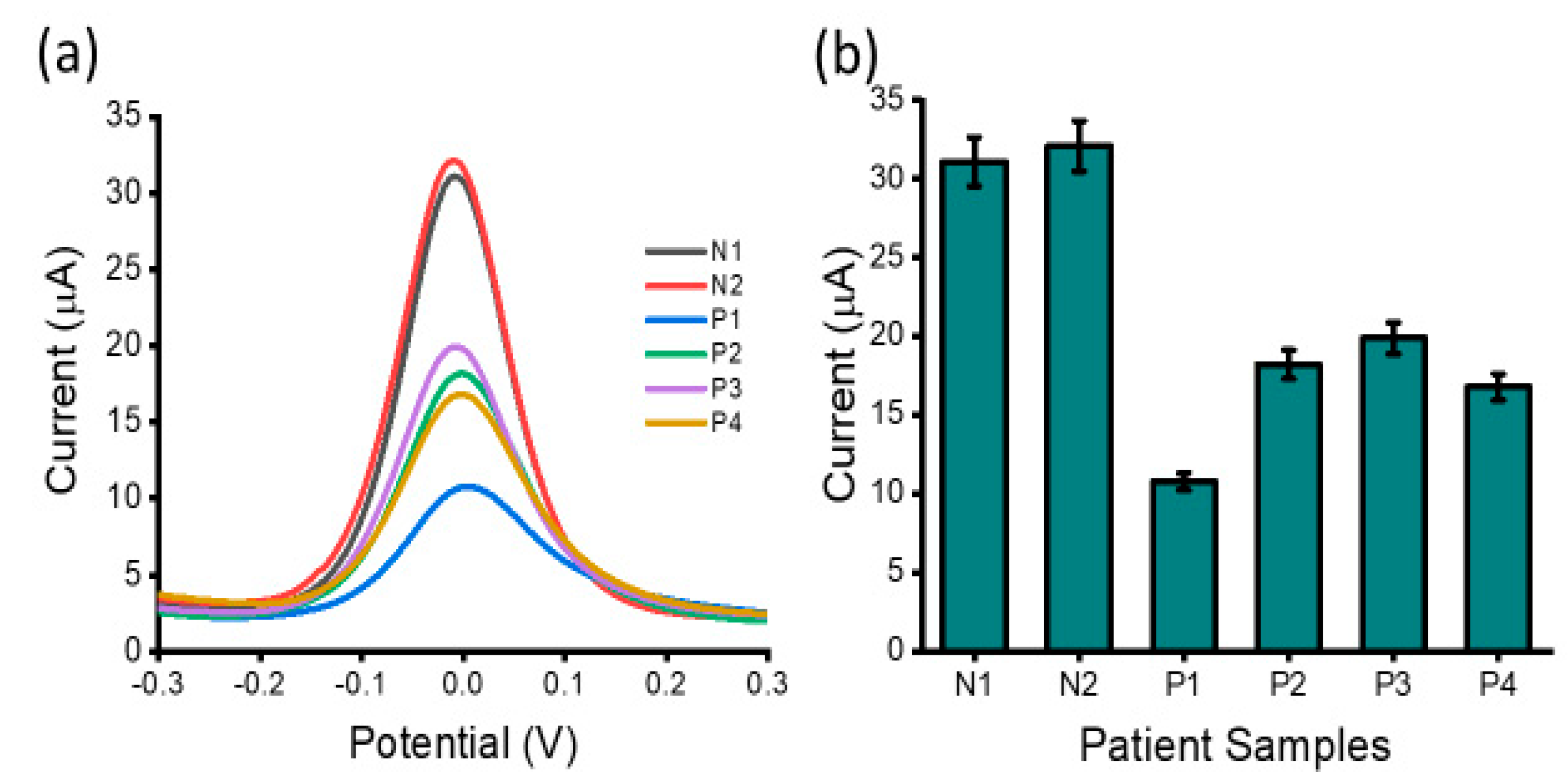
3.3. LncRNA Detection in Ovarian Cancer Patient Samples
3.4. Advantages and Disadvantages of the Assay
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Mercer, T.R.; Dinger, M.E.; Mattick, J.S. Long non-coding RNAs: Insights into functions. Nat. Rev. Genet. 2009, 10, 155–159. [Google Scholar] [CrossRef] [PubMed]
- Yao, R.-W.; Wang, Y.; Chen, L.-L. Cellular functions of long noncoding RNAs. Nat. Cell Biol. 2019, 21, 542–551. [Google Scholar] [CrossRef] [PubMed]
- Bhan, A.; Mandal, S.S. LncRNA HOTAIR: A master regulator of chromatin dynamics and cancer. Biochim. Biophys. Acta Rev. Cancer 2015, 1856, 151–164. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kopp, F.; Mendell, J.T. Functional Classification and Experimental Dissection of Long Noncoding RNAs. Cell 2018, 172, 393–407. [Google Scholar] [CrossRef] [Green Version]
- Schmitt, A.M.; Chang, H.Y. Long Noncoding RNAs in Cancer Pathways. Cancer Cell 2016, 29, 452–463. [Google Scholar] [CrossRef] [Green Version]
- Xue, X.; Yang, Y.A.; Zhang, A.; Fong, K.; Kim, J.; Song, B.; Li, S.; Zhao, J.C.; Yu, J. LncRNA HOTAIR enhances ER signaling and confers tamoxifen resistance in breast cancer. Oncogene 2016, 35, 2746–2755. [Google Scholar] [CrossRef] [Green Version]
- Hajjari, M.; Salavaty, A. HOTAIR: An oncogenic long non-coding RNA in different cancers. Cancer Biol. Med. 2015, 12, 1. [Google Scholar]
- Chen, J.; Deng, B.; Wu, P.; Li, F.; Li, X.F.; Le, X.C.; Zhang, H.; Hou, X. Amplified binding-induced homogeneous assay through catalytic cycling of analyte for ultrasensitive protein detection. Chem. Commun. 2016, 52, 1816–1819. [Google Scholar] [CrossRef]
- Lee, T.-L.; Xiao, A.; Rennert, O.M. Identification of novel long noncoding RNA transcripts in male germ cells. Methods Mol. Biol. 2012, 825, 105–114. [Google Scholar] [CrossRef] [Green Version]
- Lee, C.; Kikyo, N. Strategies to identify long noncoding RNAs involved in gene regulation. Cell Biosci. 2012, 2, 37. [Google Scholar] [CrossRef] [Green Version]
- Wang, Z.; Gerstein, M.; Snyder, M. RNA-Seq: A revolutionary tool for transcriptomics. Nat. Rev. Genet. 2009, 10, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Koshiol, J.; Wang, E.; Zhao, Y.D.; Marincola, F.; Landi, M.T. Strengths and Limitations of Laboratory Procedures for MicroRNA Detection. Cancer Epidemiol. Prev. Biomark. 2010, 19, 907–911. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Islam, M.N.; Masud, M.K.; Haque, M.H.; Al Hossain, M.S.; Yamauchi, Y.; Nguyen, N.-T.; Shiddiky, M.J.A. RNA Biomarkers: Diagnostic and Prognostic Potentials and Recent Developments of Electrochemical Biosensors. Small Methods 2017, 1, 1700131. [Google Scholar] [CrossRef]
- Soda, N.; Umer, M.; Kashaninejad, N.; Kasetsirikul, S.; Kline, R.; Salomon, C.; Nam-Trung, N.; Shiddiky, M.J.A. PCR-Free Detection of Long Non-Coding HOTAIR RNA in Ovarian Cancer Cell Lines and Plasma Samples. Cancers 2020, 12, 2233. [Google Scholar] [CrossRef] [PubMed]
- Islam, M.N.; Moriam, S.; Umer, M.; Phan, H.-P.; Salomon, C.; Kline, R.; Nguyen, N.-T.; Shiddiky, M.J.A. Naked-eye and electrochemical detection of isothermally amplified HOTAIR long non-coding RNA. Analyst 2018, 143, 3021–3028. [Google Scholar] [CrossRef]
- Soda, N.; Rehm, B.H.A.; Sonar, P.; Nguyen, N.-T.; Shiddiky, M.J.A. Advanced liquid biopsy technologies for circulating biomarker detection. J. Mater. Chem. B 2019, 7, 6670–6704. [Google Scholar] [CrossRef]
- Ahmed, M.; Carrascosa, L.G.; Sina, A.A.I.; Zarate, E.M.; Korbie, D.; Ru, K.-L.; Shiddiky, M.J.; Mainwaring, P.; Trau, M. Detection of aberrant protein phosphorylation in cancer using direct gold-protein affinity interactions. Biosens. Bioelectron. 2017, 91, 8–14. [Google Scholar] [CrossRef]
- Ibn Sina, A.A.; Koo, K.M.; Ahmed, M.; Carrascosa, L.G.; Trau, M. Interfacial Biosensing: Direct Biosensing of Biomolecules at the Bare Metal Interface. In Encyclopedia of Interfacial Chemistry; Wandelt, K., Ed.; Elsevier: Oxford, UK, 2018; pp. 269–277. [Google Scholar] [CrossRef]
- Haque, M.H.; Gopalan, V.; Yadav, S.; Islam, M.N.; Eftekhari, E.; Li, Q.; Carrascosa, L.G.; Nguyen, N.-T.; Lam, A.K.; Shiddiky, M.J.A. Detection of regional DNA methylation using DNA-graphene affinity interactions. Biosens. Bioelectron. 2017, 87, 615–621. [Google Scholar] [CrossRef] [Green Version]
- Lu, C.-H.; Yang, H.-H.; Zhu, C.-L.; Chen, X.; Chen, G.-N. A Graphene Platform for Sensing Biomolecules. Angew. Chem. Int. Ed. 2009, 48, 4785–4787. [Google Scholar] [CrossRef]
- Song, Y.; Qu, K.; Zhao, C.; Ren, J.; Qu, X. Graphene Oxide: Intrinsic Peroxidase Catalytic Activity and Its Application to Glucose Detection. Adv. Mater. 2010, 22, 2206–2210. [Google Scholar] [CrossRef]
- Wu, M.; Kempaiah, R.; Huang, P.-J.J.; Maheshwari, V.; Liu, J. Adsorption and Desorption of DNA on Graphene Oxide Studied by Fluorescently Labeled Oligonucleotides. Langmuir 2011, 27, 2731–2738. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Koo, K.M.; Sina, A.A.I.; Carrascosa, L.G.; Shiddiky, M.J.A.; Trau, M. DNA–bare gold affinity interactions: Mechanism and applications in biosensing. Anal. Methods 2015, 7, 7042–7054. [Google Scholar] [CrossRef]
- Li, H.; Rothberg, L. Colorimetric detection of DNA sequences based on electrostatic interactions with unmodified gold nanoparticles. Proc. Natl. Acad. Sci. USA 2004, 101, 14036–14039. [Google Scholar] [CrossRef] [Green Version]
- Islam, T.; Hasan, M.M.; Awal, A.; Nurunnabi, M.; Ahammad, A.J.S. Metal Nanoparticles for Electrochemical Sensing: Progress and Challenges in the Clinical Transition of Point-of-Care Testing. Molecules 2020, 25, 5787. [Google Scholar] [CrossRef] [PubMed]
- Masud, M.K.; Umer, M.; Hossain, M.S.A.; Yamauchi, Y.; Nguyen, N.T.; Shiddiky, M.J.A. Nanoarchitecture Frameworks for Electrochemical miRNA Detection. Trends Biochem. Sci. 2019, 44, 433–452. [Google Scholar] [CrossRef] [PubMed]
- Sina, A.A.I.; Howell, S.; Carrascosa, L.G.; Rauf, S.; Shiddiky, M.J.A.; Trau, M. eMethylsorb: Electrochemical quantification of DNA methylation at CpG resolution using DNA–gold affinity interactions. Chem. Commun. 2014, 50, 13153–13156. [Google Scholar] [CrossRef] [Green Version]
- Zhang, J.; Wang, L.; Pan, D.; Song, S.; Fan, C. DNA hybridization “turns on” electrocatalysis at gold electrodes. Chem. Commun. 2007, 11, 1154–1156. [Google Scholar] [CrossRef]
- Liu, F.; Xiang, G.; Jiang, D.; Zhang, L.; Chen, X.; Liu, L.; Luo, F.; Li, Y.; Liu, C.; Pu, X. Ultrasensitive strategy based on PtPd nanodendrite/nano-flower-like@ GO signal amplification for the detection of long non-coding RNA. Biosens. Bioelectron. 2015, 74, 214–221. [Google Scholar] [CrossRef]
- Liu, F.; Xiang, G.; Zhang, L.; Jiang, D.; Liu, L.; Li, Y.; Liu, C.; Pu, X. A novel label free long non-coding RNA electrochemical biosensor based on green L-cysteine electrodeposition and Au–Rh hollow nanospheres as tags. RSC Adv. 2015, 5, 51990–51999. [Google Scholar] [CrossRef]
- Yin, H.; Zhou, Y.; Yang, Z.; Guo, Y.; Wang, X.; Ai, S.; Zhang, X. Electrochemical immunosensor for N6-methyladenosine RNA modification detection. Sens. Actuators B Chem. 2015, 221, 1–6. [Google Scholar] [CrossRef]
- Rafiee-Pour, H.-A.; Behpour, M.; Keshavarz, M. A novel label-free electrochemical miRNA biosensor using methylene blue as redox indicator: Application to breast cancer biomarker miRNA-21. Biosens. Bioelectron. 2016, 77, 202–207. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Li, Y.; Yang, Y.; Wu, F.; Cao, J.; Bai, L. A polyaniline-reduced graphene oxide nanocomposite as a redox nanoprobe in a voltammetric DNA biosensor for Mycobacterium tuberculosis. Microchim. Acta 2017, 184, 1801–1808. [Google Scholar] [CrossRef]
- Soda, N.; Umer, M.; Kasetsirikul, S.; Salomon, C.; Kline, R.; Nguyen, N.-T.; Rehm, B.H.A.; Shiddiky, M.J.A. An amplification-free method for the detection of HOTAIR long non-coding RNA. Anal. Chim. Acta 2020, 1132, 66–73. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Wu, D.; Tu, S.; Yang, C.; Chen, D.; Xu, Y. A novel biosensor for the ultrasensitive detection of the lncRNA biomarker MALAT1 in non-small cell lung cancer. Sci. Rep. 2021, 11, 3666. [Google Scholar] [CrossRef] [PubMed]
- Soda, N.; Clack, K.; Shiddiky, M.J.A. Recent advances in liquid biopsy technologies for cancer biomarker detection. Sens. Diagn. 2022. [Google Scholar] [CrossRef]
- Zhang, Y.; Wang, X.-Y.; Su, X.; Zhang, C.-Y. Ultrasensitive detection of long non-coding RNAs based on duplex-specific nuclease-actuated cyclic enzymatic repairing-mediated signal amplification. Chem. Commun. 2019, 55, 6827–6830. [Google Scholar] [CrossRef] [PubMed]
- Özeş, A.R.; Wang, Y.; Zong, X.; Fang, F.; Pilrose, J.; Nephew, K.P. Therapeutic targeting using tumor specific peptides inhibits long non-coding RNA HOTAIR activity in ovarian and breast cancer. Sci. Rep. 2017, 7, 894. [Google Scholar] [CrossRef] [Green Version]
- Qiu, J.-J.; Lin, Y.-Y.; Ye, L.-C.; Ding, J.-X.; Feng, W.-W.; Jin, H.-Y.; Zhang, Y.; Li, Q.; Hua, K.-Q. Overexpression of long non-coding RNA HOTAIR predicts poor patient prognosis and promotes tumor metastasis in epithelial ovarian cancer. Gynecol. Oncol. 2014, 134, 121–128. [Google Scholar] [CrossRef]
| Oligonucleotides | 5′-Sequences-3′ |
|---|---|
| HOTAIR capture probe (Biotinylated) | ATC AAT TAA TTA GCG CCT CCC AGT CCC/3Bio |
| Synthetic HOTAIR lncRNA target | GGG ACU GGG AGG CGC UAA UUA AUU GAU UCC UUU GGA CUG UAA AAU AUG GCG GCG U |
| miR-891 | UGC AAC GAA CCU GAG CCA CUG A |
| miR-486 | UCC UGU ACU GAG CUG CCC CGA G |
| HOTAIR Forward primer sequence | AAG CAC CTC TAT CTC AGC CG |
| HOTAIR Reverse primer sequence | AGA ACC CTC TGA CAT TTG CCT |
| GAPDH Forward primer sequence | CCG GGA AAC TGT GGC GTG ATG G |
| GAPDH Reverse primer sequence | AGG TGG AGG AGT GGG TGT CGC TGT T |
| Target | Techniques | Linear Range (M) | Detection Limit (M) | References |
|---|---|---|---|---|
| DPV | 10 pM–10 nM | 0.247 pM | [29] | |
| DPV | 1.0 pM–10 μM | 0.886 pM | [30] | |
| DPV | 10 pM–10 nM | 2.57 pM | [31] | |
| DPV | 100 fM–500.0 pM | 84.3 fM | [32] | |
| DPV | 100 fM–10 nM | 50 fM | [33] | |
| I-T | 1 fM–1.0 nM | 1 fM | [34] | |
| DPV | 10 fM–10.0 nM | 42.8 fM | [35] | |
| DPV | 1 pM–10 nM | 1 pM | Present work |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Clack, K.; Soda, N.; Kasetsirikul, S.; Kline, R.; Salomon, C.; Shiddiky, M.J.A. An Interfacial Affinity Interaction-Based Method for Detecting HOTAIR lncRNA in Cancer Plasma Samples. Biosensors 2022, 12, 287. https://doi.org/10.3390/bios12050287
Clack K, Soda N, Kasetsirikul S, Kline R, Salomon C, Shiddiky MJA. An Interfacial Affinity Interaction-Based Method for Detecting HOTAIR lncRNA in Cancer Plasma Samples. Biosensors. 2022; 12(5):287. https://doi.org/10.3390/bios12050287
Chicago/Turabian StyleClack, Kimberley, Narshone Soda, Surasak Kasetsirikul, Richard Kline, Carlos Salomon, and Muhammad J. A. Shiddiky. 2022. "An Interfacial Affinity Interaction-Based Method for Detecting HOTAIR lncRNA in Cancer Plasma Samples" Biosensors 12, no. 5: 287. https://doi.org/10.3390/bios12050287
APA StyleClack, K., Soda, N., Kasetsirikul, S., Kline, R., Salomon, C., & Shiddiky, M. J. A. (2022). An Interfacial Affinity Interaction-Based Method for Detecting HOTAIR lncRNA in Cancer Plasma Samples. Biosensors, 12(5), 287. https://doi.org/10.3390/bios12050287









