A Selective Fluorescent l-Lactate Biosensor Based on an l-Lactate-Specific Transcription Regulator and Förster Resonance Energy Transfer
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Bacterial Strains and Culture Conditions
2.3. Expression, Purification, and Characterization of Lactate Utilization Operon Regulator LldR
2.4. Fluorescence-Based Thermal Shift (FTS) Assay
2.5. Construction and Purification of Fluorescent l-Lactate Biosensor FILLac
2.6. Characterization of FILLac In-Virto
2.7. Batch Fermentation for Lactate Production
2.8. Jiaosu and Yogurt Samples Preparation
2.9. Quantification of l-Lactate by HPLC, SBA-40D Bioanalyzer and FILLac10N0C
3. Results and Discussion
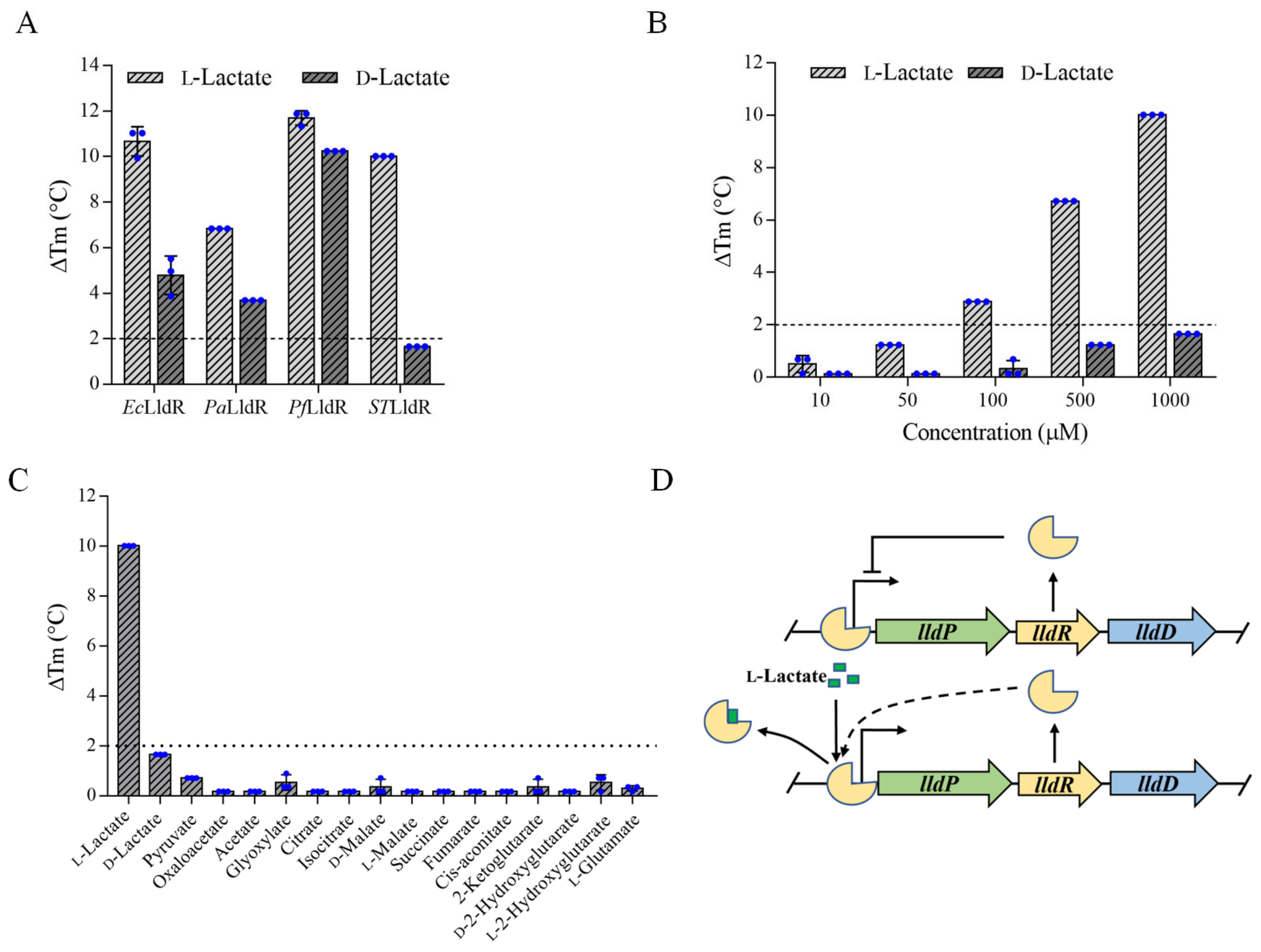
3.1. STLldR as a Specific Recognition Element for l-Lactate
3.2. Design and Optimization of the Fluorescent l-Lactate Biosensor
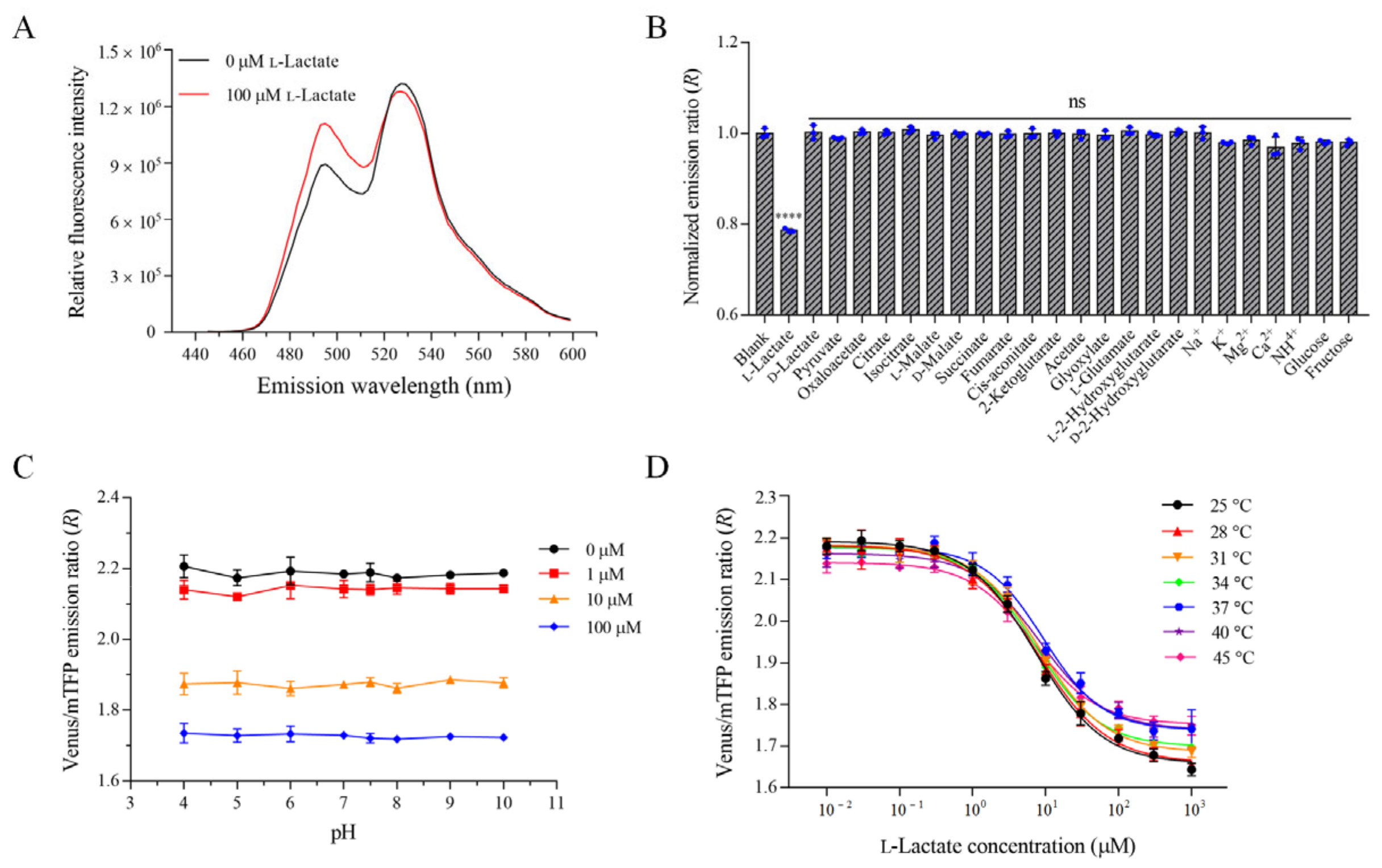
3.3. Characterization of the Optimal l-Lactate Biosensor FILLac10N0C
3.4. Performance of FILLac10N0C in l-Lactate Quantitation
3.5. Quantitation of l-Lactate in Different Fermentation Samples by FILLac10N0C
3.6. Determination of l-Lactate in Food Samples by FILLac10N0C
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hofvendahl, K.; Hahn-Hagerdal, B. Factors affecting the fermentative lactic acid production from renewable resources. Enzyme Microb. Technol. 2000, 26, 87–107. [Google Scholar] [CrossRef] [PubMed]
- Faubert, B.; Li, K.Y.; Cai, L.; Hensley, C.T.; Kim, J.; Zacharias, L.G.; Yang, C.D.; Do, Q.N.; Doucette, S.; Burguete, D.; et al. Lactate metabolism in human lung tumors. Cell 2017, 171, 358. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chrusch, C.; Bands, C.; Bose, D.; Li, X.; Jacobs, H.; Duke, K.; Bautista, E.; Eschun, G.; Light, R.B.; Mink, S.N. Impaired hepatic extraction and increased splanchnic production contribute to lactic acidosis in canine sepsis. Am. J. Respir. Crit. Care Med. 2000, 161, 517–526. [Google Scholar] [CrossRef] [PubMed]
- Kliegel, A.; Losert, H.; Sterz, F.; Holzer, M.; Zeiner, A.; Havel, C.; Laggner, A.N. Serial lactate determinations for prediction of outcome after cardiac arrest. Medicine 2004, 83, 274–279. [Google Scholar] [CrossRef]
- Murphy, N.D.; Kodakat, S.K.; Wendon, J.A.; Jooste, C.A.; Muiesan, P.; Rela, M.; Heaton, N.D. Liver and intestinal lactate metabolism in patients with acute hepatic failure undergoing liver transplantation. Crit. Care Med. 2001, 29, 2111–2118. [Google Scholar] [CrossRef]
- Gamella, M.; Campuzano, S.; Conzuelo, F.; Curiel, J.A.; Munoz, R.; Reviejo, A.J.; Pingarron, J.M. Integrated multienzyme electrochemical biosensors for monitoring malolactic fermentation in wines. Talanta 2010, 81, 925–933. [Google Scholar] [CrossRef] [Green Version]
- Pundir, C.S.; Narwal, V.; Batra, B. Determination of lactic acid with special emphasis on biosensing methods: A review. Biosens. Bioelectron. 2016, 86, 777–790. [Google Scholar] [CrossRef]
- Castro-Aguirre, E.; Iniguez-Franco, F.; Samsudin, H.; Fang, X.; Auras, R. Poly (lactic acid)-Mass production, processing, industrial applications, and end of life. Adv. Drug Deliver. Rev. 2016, 107, 333–366. [Google Scholar] [CrossRef] [Green Version]
- Rawoof, S.; Kumar, P.S.; Vo, D.; Devaraj, K.; Mani, Y.; Devaraj, T.; Subramanian, S. Production of optically pure lactic acid by microbial fermentation: A review. Environ. Chem. Lett. 2021, 19, 539–556. [Google Scholar] [CrossRef]
- Yuzer, E.; Dogan, V.; Kilic, V.; Sen, M. Smartphone embedded deep learning approach for highly accurate and automated colorimetric lactate analysis in sweat. Sensor. Actuat. B-Chem. 2022, 371, 132489. [Google Scholar] [CrossRef]
- Promphet, N.; Rattanawaleedirojn, P.; Siralertmukul, K.; Soatthiyanon, N.; Potiyaraj, P.; Thanawattano, C.; Hinestroza, J.P.; Rodthongkum, N. Non-invasive textile based colorimetric sensor for the simultaneous detection of sweat pH and lactate. Talanta 2019, 192, 424–430. [Google Scholar] [CrossRef] [PubMed]
- Dias, A.; Silva, R.; Arruda, M. A sequential injection system for indirect spectrophotometric determination of lactic acid in yogurt and fermented mash samples. Microchem. J. 2010, 96, 151–156. [Google Scholar] [CrossRef]
- Zhang, X.M.; Ding, S.S.; Cao, S.M.; Zhu, A.W.; Shi, G.Y. Functional surface engineering of quantum dot hydrogels for selective fluorescence imaging of extracellular lactate release. Biosens. Bioelectron. 2016, 80, 315–322. [Google Scholar] [CrossRef]
- Matoori, S.; Mooney, D.J. Development of a liposomal near-infrared fluorescence lactate assay for human blood. Biomaterials 2022, 283, 121475. [Google Scholar] [CrossRef] [PubMed]
- Milagres, M.P.; Brandao, S.; Magalhaes, M.A.; Minim, V.; Minim, L.A. Development and validation of the high performance liquid chromatography-ion exclusion method for detection of lactic acid in milk. Food Chem. 2012, 135, 1078–1082. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hsieh, C.L.; Koga, R.; Furusho, A.; Akita, T.; Mita, M.; Ide, T.; Lee, J.A.; Hamase, K. Enantioselective and simultaneous determination of lactate and 3-hydroxybutyrate in human plasma and urine using a narrow-bore online two-dimensional high-performance liquid chromatography system. J. Sep. Sci. 2018, 41, 1298–1306. [Google Scholar] [CrossRef]
- Jackson, T.C.; Zhang, Y.V.; Sime, P.J.; Phipps, R.P.; Kottmann, R.M. Development of an accurate and sensitive method for lactate analysis in exhaled breath condensate by LC MS/MS. J. Chromatogr. B. 2017, 1061, 468–473. [Google Scholar] [CrossRef]
- Zhang, W.; Guo, C.; Jiang, K.Z.; Ying, M.F.; Hu, X. Quantification of lactate from various metabolic pathways and quantification issues of lactate isotopologues and isotopmers. Sci. Rep. 2017, 7, 8489. [Google Scholar] [CrossRef] [Green Version]
- Rattu, G.; Khansili, N.; Maurya, V.K.; Krishna, P.M. Lactate detection sensors for food, clinical and biological applications: A review. Environ. Chem. Lett. 2021, 19, 1135–1152. [Google Scholar] [CrossRef]
- Poletti, F.; Zanfrognini, B.; Favaretto, L.; Quintano, V.; Sun, J.H.; Treossi, E.; Melucci, M.; Palermo, V.; Zanardi, C. Continuous capillary-flow sensing of glucose and lactate in sweat with an electrochemical sensor based on functionalized graphene oxide. Sensor. Actuat. B-Chem. 2021, 344, 130253. [Google Scholar] [CrossRef]
- Zhang, Q.W.; Jiang, D.F.; Xu, C.S.; Ge, Y.C.; Liu, X.H.; Wei, Q.Q.; Huang, L.P.; Ren, X.Q.; Wang, C.D.; Wang, Y. Wearable electrochemical biosensor based on molecularly imprinted Ag nanowires for noninvasive monitoring lactate in human sweat. Sensor. Actuat. B-Chem. 2020, 320, 128325. [Google Scholar] [CrossRef]
- Bott-Neto, J.L.; Martins, T.S.; Buscaglia, L.A.; Santiago, P.; Fernandez, P.S.; Machado, S.; Oliveira, O.N. A portable system for photoelectrochemical detection of lactate on TiO2 nanoparticles and [Ni(salen)] polymeric film. Sensor. Actuat. B-Chem. 2021, 345, 130390. [Google Scholar] [CrossRef]
- Ma, Y.S.; Wang, Y.; Liu, Y.J.; Shi, L.J.; Yang, D.Z. Towards a transdermal membrane biosensor for the detection of lactate in body fluids. Sensor. Actuat. B-Chem. 2020, 308, 127645. [Google Scholar] [CrossRef]
- Ma, Y.S.; Wang, Y.; Liu, Y.J.; Shi, L.J.; Yang, D.Z. A cascade-triggered ratiometric fluorescent sensor based on nanocomposite for lactate determination. Sensor. Actuat. B-Chem. 2022, 355, 131295. [Google Scholar] [CrossRef]
- Zhang, X.; Hu, Y.; Yang, X.; Tang, Y.; Han, S.; Kang, A.; Deng, H.; Chi, Y.; Zhu, D.; Lu, Y. FÖrster resonance energy transfer (FRET)-based biosensors for biological applications. Biosens. Bioelectron. 2019, 138, 111314. [Google Scholar] [CrossRef]
- San Martín, A.; Ceballo, S.; Ruminot, I.; Lerchundi, R.; Frommer, W.B.; Barros, L.F. A genetically encoded FRET lactate sensor and its use to detect the Warburg effect in single cancer cells. PLoS ONE 2013, 8, e57712. [Google Scholar] [CrossRef]
- Bekdash, R.; Quejada, J.R.; Ueno, S.; Kawano, F.; Morikawa, K.; Klein, A.D.; Matsumoto, K.; Lee, T.C.; Nakanishi, K.; Chalan, A.; et al. GEM-IL: A highly responsive fluorescent lactate indicator. Cell Rep. Methods 2021, 1, 100092. [Google Scholar] [CrossRef]
- Harada, K.; Chihara, T.; Hayasaka, Y.; Mita, M.; Takizawa, M.; Ishida, K.; Arai, M.; Tsuno, S.; Matsumoto, M.; Ishihara, T.; et al. Green fluorescent protein-based lactate and pyruvate indicators suitable for biochemical assays and live cell imaging. Sci. Rep. 2020, 10, 19562. [Google Scholar] [CrossRef]
- Nasu, Y.; Murphy-Royal, C.; Wen, Y.R.; Haidey, J.N.; Molina, R.S.; Aggarwal, A.; Zhang, S.C.; Kamijo, Y.; Paquet, M.E.; Podgorski, K.; et al. A genetically encoded fluorescent biosensor for extracellular l-lactate. Nat. Commun. 2021, 12, 7058. [Google Scholar] [CrossRef]
- Wellbourne-Wood, J.; Briquet, M.; Alessandri, M.; Binda, F.; Touya, M.; Chatton, J.Y. Evaluation of hydroxycarboxylic acid receptor 1 (HCAR1) as a building block for genetically encoded extracellular lactate biosensors. Biosensors 2022, 12, 143. [Google Scholar] [CrossRef]
- Koveal, D.; Rosen, P.C.; Meyer, D.J.; Diaz-Garcia, C.M.; Wang, Y.C.; Cai, L.H.; Chou, P.J.; Weitz, D.A.; Yellen, G. A high-throughput multiparameter screen for accelerated development and optimization of soluble genetically encoded fluorescent biosensors. Nat. Commun. 2022, 13, 2919. [Google Scholar] [CrossRef] [PubMed]
- Xiao, D.; Hu, C.; Xu, X.; Lü, C.; Wang, Q.; Zhang, W.; Gao, C.; Xu, P.; Wang, X.; Ma, C. A d,l-lactate biosensor based on allosteric transcription factor LldR and amplified luminescent proximity homogeneous assay. Biosens. Bioelectron. 2022, 211, 114378. [Google Scholar] [CrossRef] [PubMed]
- Kang, Z.Q.; Zhang, M.M.; Gao, K.Y.; Zhang, W.; Meng, W.S.; Liu, Y.D.; Xiao, D.; Guo, S.T.; Ma, C.Q.; Gao, C.; et al. An l-2-hydroxyglutarate biosensor based on specific transcriptional regulator LhgR. Nat. Commun. 2021, 12, 3619. [Google Scholar] [CrossRef]
- Xia, Y.Z.; Li, K.; Li, J.J.; Wang, T.Q.; Gu, L.C.; Xun, L.Y. T5 exonuclease-dependent assembly offers a low-cost method for efficient cloning and site-directed mutagenesis. Nucleic Acids Res. 2019, 47, e15. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- van Hijum, S.A.; Medema, M.H.; Kuipers, O.P. Mechanisms and evolution of control logic in prokaryotic transcriptional regulation. Microbiol. Mol. Biol. Rev. 2009, 73, 481–509. [Google Scholar] [CrossRef] [Green Version]
- Fernandez-Lopez, R.; Ruiz, R.; de la Cruz, F.; Moncalian, G. Transcription factor-based biosensors enlightened by the analyte. Front. Microbiol. 2015, 6, 648. [Google Scholar] [CrossRef] [Green Version]
- Xiao, D.; Zhang, W.; Guo, X.T.; Liu, Y.D.; Hu, C.X.; Guo, S.T.; Kang, Z.Q.; Xu, X.Z.; Ma, C.Q.; Gao, C.; et al. A d-2-hydroxyglutarate biosensor based on specific transcriptional regulator DhdR. Nat. Commun. 2021, 12, 7108. [Google Scholar] [CrossRef]
- Jiang, T.; Gao, C.; Ma, C.; Xu, P. Microbial lactate utilization: Enzymes, pathogenesis, and regulation. Trends Microbiol. 2014, 22, 589–599. [Google Scholar] [CrossRef]
- Gillis, C.C.; Winter, M.G.; Chanin, R.B.; Zhu, W.H.; Spiga, L.; Winter, S.E. Host-derived metabolites modulate transcription of Salmonella genes involved in l-lactate utilization during gut colonization. Infect. Immun. 2019, 87, e00773–18. [Google Scholar] [CrossRef] [Green Version]
- Augustiniene, E.; Valanciene, E.; Matulis, P.; Syrpas, M.; Jonuskiene, I.; Malys, N. Bioproduction of l- and d-lactic acids: Advances and trends in microbial strain application and engineering. Crit. Rev. Biotechnol. 2022, 42, 342–360. [Google Scholar] [CrossRef]
- Plessas, S.; Bosnea, L.; Psarianos, C.; Koutinas, A.A.; Marchant, R.; Banat, I.M. Lactic acid production by mixed cultures of Kluyveromyces marxianus, Lactobacillus delbrueckii ssp. bulgaricus and Lactobacillus helveticus. Bioresour. Technol. 2008, 99, 5951–5955. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.H.; Harris, H.; McCann, A.; Guo, C.Y.; Argimon, S.; Zhang, W.Y.; Yang, X.W.; Jeffery, I.B.; Cooney, J.C.; Kagawa, T.F.; et al. Expanding the biotechnology potential of lactobacilli through comparative genomics of 213 strains and associated genera. Nat. Commun. 2015, 6, 8322. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dicks, L.; DuPlessis, E.M.; Dellaglio, F.; Lauer, E. Reclassification of Lactobacillus casei subsp. casei ATCC 393 and Lactobacillus rhamnosus ATCC 15820 as Lactobacillus zeae nom. rev., designation of ATCC 334 as the neotype of L. casei subsp. casei, and rejection of the Lactobacillus paracasei. Int. J. Syst. Bacteriol. 1996, 46, 337–340. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Okano, K.; Yoshida, S.; Tanaka, T.; Ogino, C.; Fukuda, H.; Kondo, A. Homo-d-lactic acid fermentation from arabinose by redirection of the phosphoketolase pathway to the pentose phosphate pathway in l-lactate dehydrogenase gene-deficient Lactobacillus plantarum. Appl. Environ. Microbiol. 2009, 75, 5175–5178. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hosnedlova, B.; Sochor, J.; Baron, M.; Bjorklund, G.; Kizek, R. Application of nanotechnology based-biosensors in analysis of wine compounds and control of wine quality and safety: A critical review. Crit. Rev. Food Sci. 2020, 60, 3271–3289. [Google Scholar] [CrossRef] [PubMed]
- Manoj, D.; Shanmugasundaram, S.; Anandharamakrishnan, C. Nanosensing and nanobiosensing: Concepts, methods, and applications for quality evaluation of liquid foods. Food Control 2021, 126, 108017. [Google Scholar] [CrossRef]
- Hui, S.; Ghergurovich, J.M.; Morscher, R.J.; Jang, C.; Teng, X.; Lu, W.Y.; Esparza, L.A.; Reya, T.; Zhan, L.; Guo, J.; et al. Glucose feeds the TCA cycle via circulating lactate. Nature 2017, 551, 115. [Google Scholar] [CrossRef] [Green Version]
- Monroe, G.R.; van Eerde, A.M.; Tessadori, F.; Duran, K.J.; Savelberg, S.; van Alfen, J.C.; Terhal, P.A.; van der Crabben, S.N.; Lichtenbelt, K.D.; Fuchs, S.A.; et al. Identification of human d-lactate dehydrogenase deficiency. Nat. Commun. 2019, 10, 1477. [Google Scholar] [CrossRef] [Green Version]
- McHale, C.; Keating, E.; O'Donovan, H.; Slattery, E. d-lactic acidosis secondary to short bowel syndrome. Postgrad. Med. J. 2003, 79, 110–112. [Google Scholar] [CrossRef] [Green Version]
- Adeva-Andany, M.; Lopez-Ojen, M.; Funcasta-Calderon, R.; Ameneiros-Rodriguez, E.; Donapetry-Garcia, C.; Vila-Altesor, M.; Rodriguez-Seijas, J. Comprehensive review on lactate metabolism in human health. Mitochondrion 2014, 17, 76–100. [Google Scholar] [CrossRef]
- Engqvist, M.; Drincovich, M.F.; Flugge, U.I.; Maurino, V.G. Two d-2-hydroxy-acid dehydrogenases in Arabidopsis thaliana with catalytic capacities to participate in the last reactions of the methylglyoxal and beta-oxidation pathways. J. Biol. Chem. 2009, 284, 25026–25037. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Becker-Kettern, J.; Paczia, N.; Conrotte, J.F.; Kay, D.P.; Guignard, C.; Jung, P.P.; Linster, C.L. Saccharomyces cerevisiae forms d-2-hydroxyglutarate and couples its degradation to d-lactate formation via a cytosolic transhydrogenase. J. Biol. Chem. 2016, 291, 6036–6058. [Google Scholar] [CrossRef] [PubMed]




Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xu, X.; Xu, R.; Hou, S.; Kang, Z.; Lü, C.; Wang, Q.; Zhang, W.; Wang, X.; Xu, P.; Gao, C.; et al. A Selective Fluorescent l-Lactate Biosensor Based on an l-Lactate-Specific Transcription Regulator and Förster Resonance Energy Transfer. Biosensors 2022, 12, 1111. https://doi.org/10.3390/bios12121111
Xu X, Xu R, Hou S, Kang Z, Lü C, Wang Q, Zhang W, Wang X, Xu P, Gao C, et al. A Selective Fluorescent l-Lactate Biosensor Based on an l-Lactate-Specific Transcription Regulator and Förster Resonance Energy Transfer. Biosensors. 2022; 12(12):1111. https://doi.org/10.3390/bios12121111
Chicago/Turabian StyleXu, Xianzhi, Rong Xu, Shuang Hou, Zhaoqi Kang, Chuanjuan Lü, Qian Wang, Wen Zhang, Xia Wang, Ping Xu, Chao Gao, and et al. 2022. "A Selective Fluorescent l-Lactate Biosensor Based on an l-Lactate-Specific Transcription Regulator and Förster Resonance Energy Transfer" Biosensors 12, no. 12: 1111. https://doi.org/10.3390/bios12121111





