Intracellular Imaging with Genetically Encoded RNA-Based Molecular Sensors
Abstract
:1. Introduction
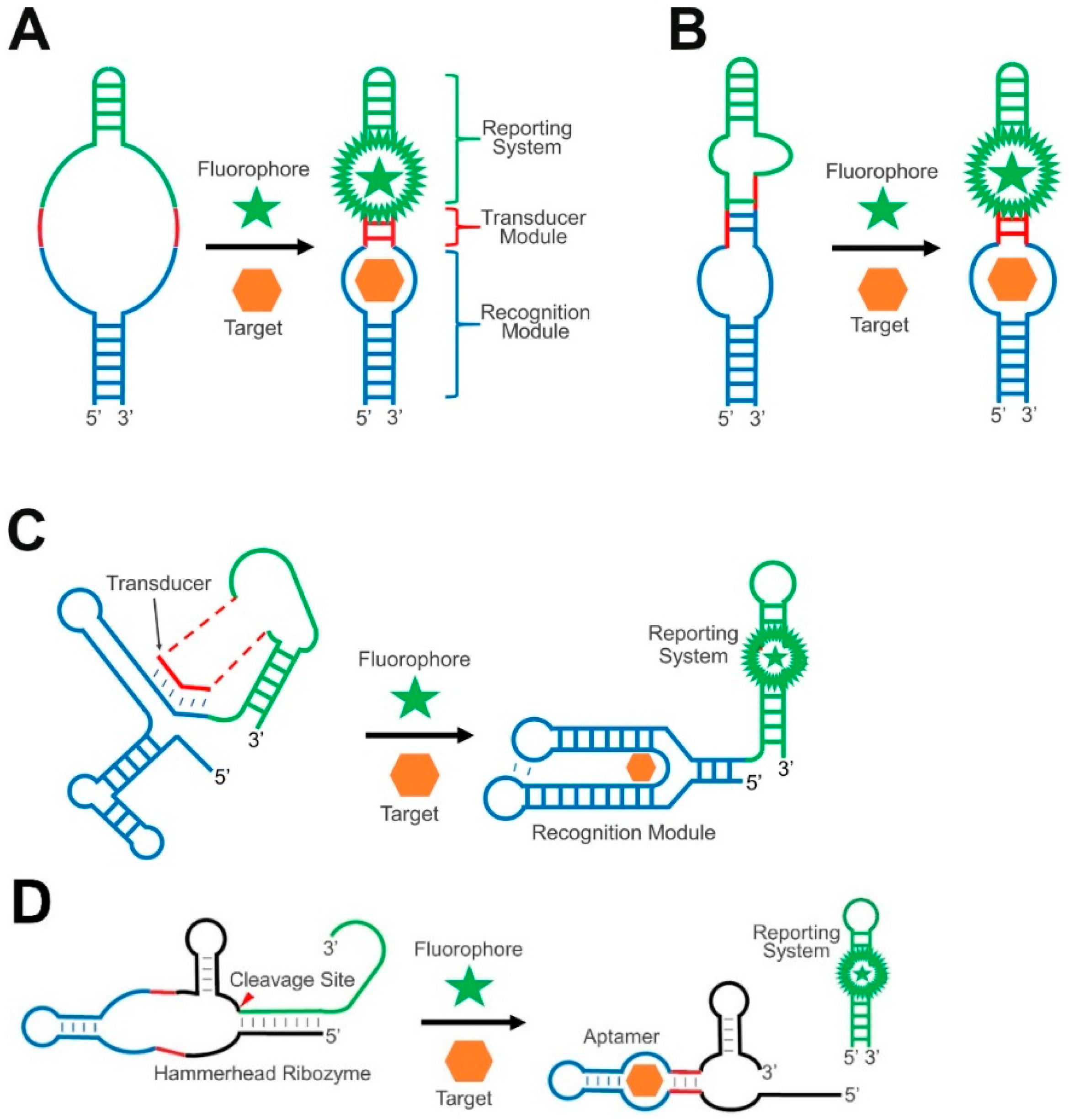
2. Transducer Modules in GERMS
2.1. RNA Duplex Formation or Helix Slipping
2.2. RNA Strand Displacement
2.3. Ribozyme-Based Transducers
3. Reporting Systems in GERMS
3.1. Protein-Based Reporters
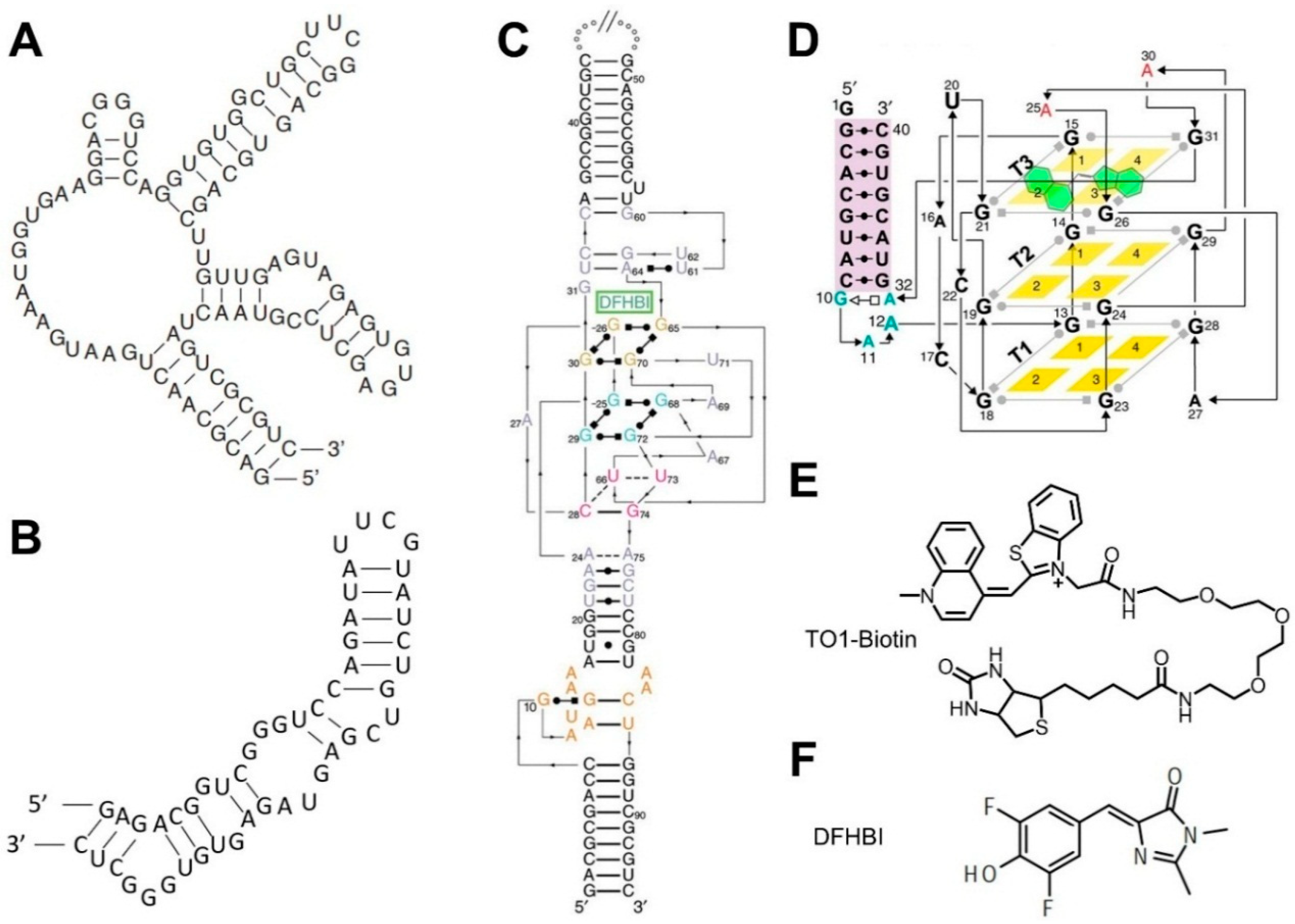
3.2. Fluorogenic RNA Complexes
4. Recognition Modules in GERMS
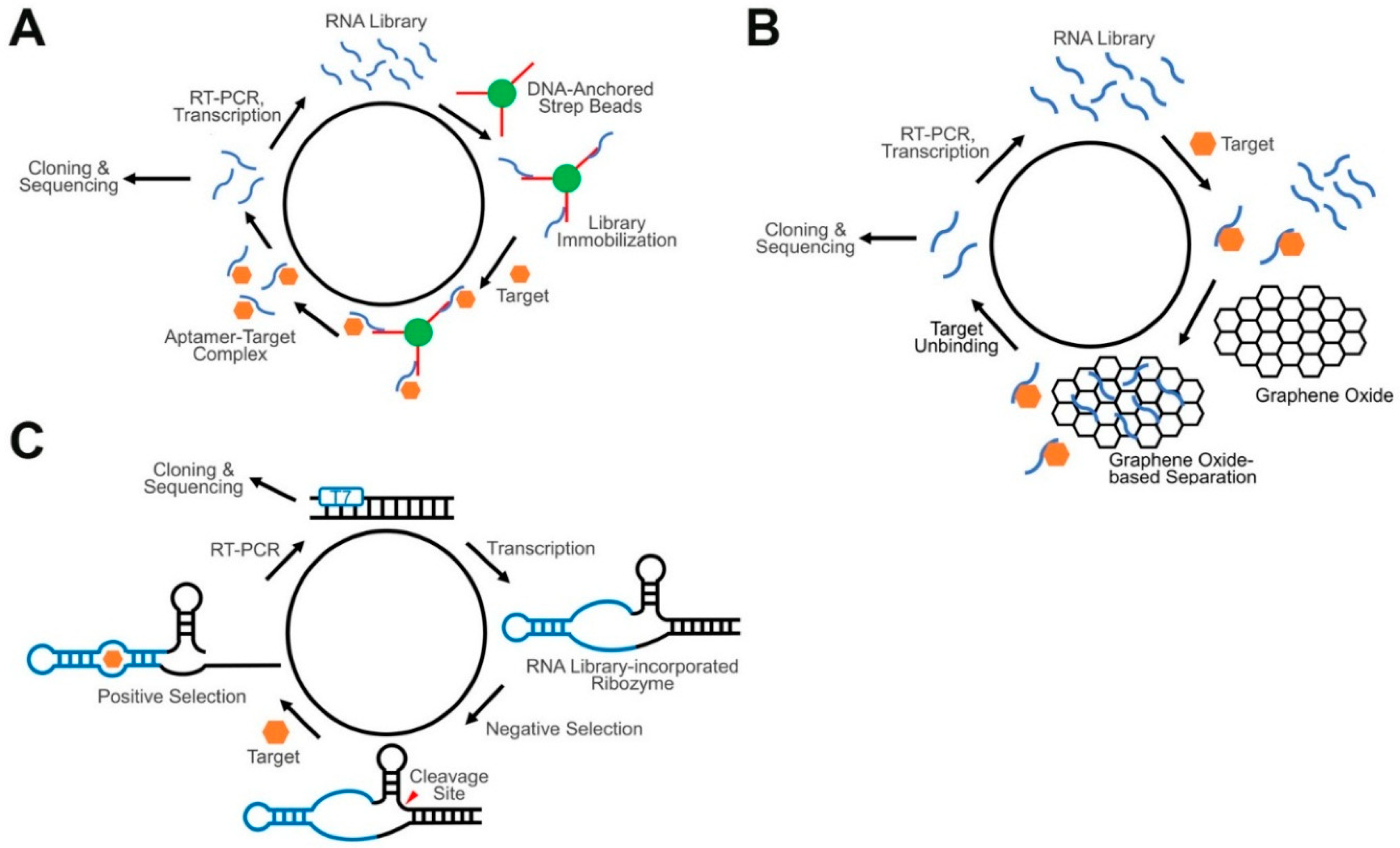
4.1. Aptamers and Conventional SELEX
4.2. Advanced SELEX Approaches for GERMS
4.3. Riboswitch-Based Recognition Modules
4.4. Specific Base Pair Formation
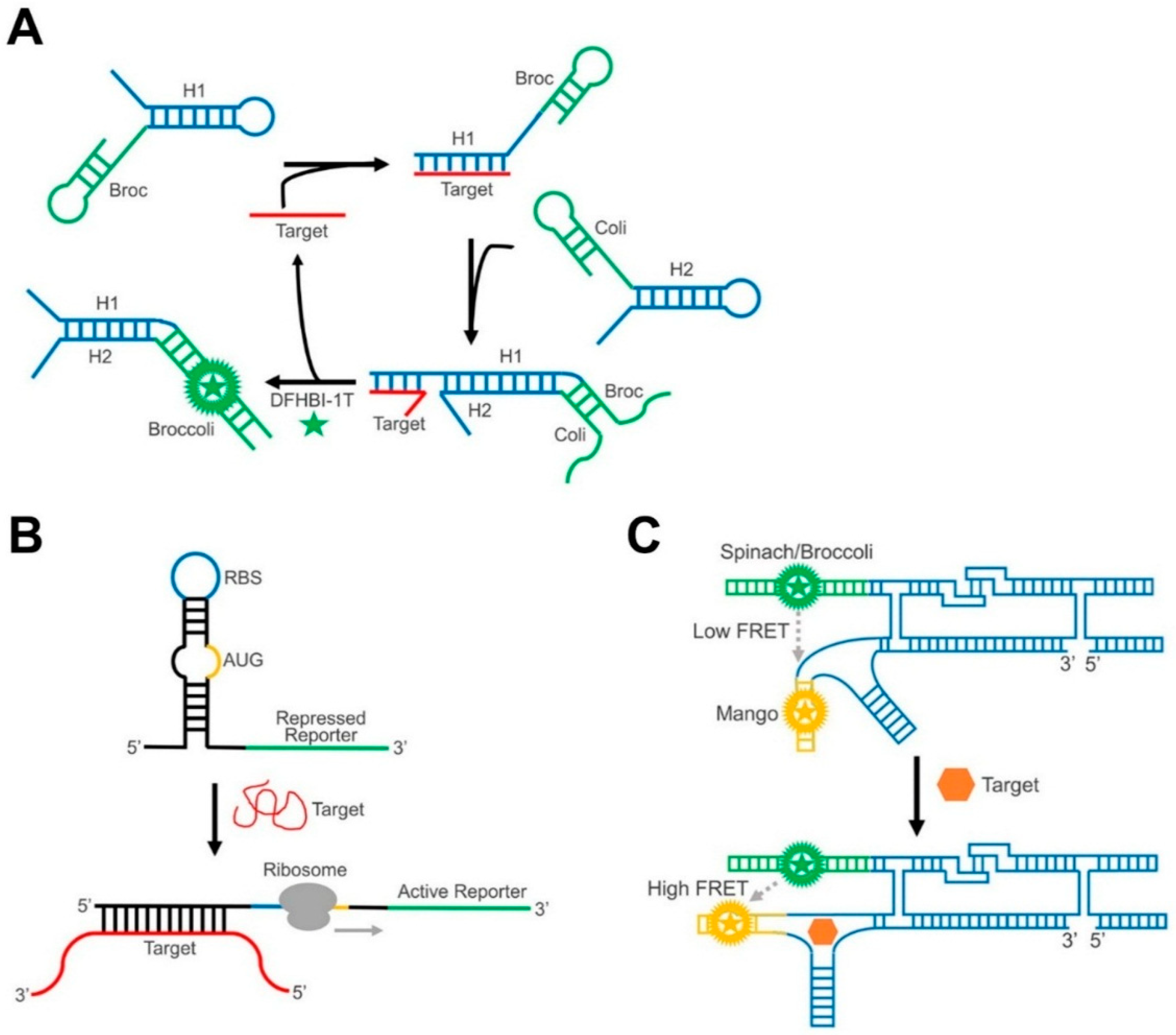
5. Recent Examples of GERMS
6. Conclusions and Outlook
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| GERMS | Genetically Encoded RNA-based Molecular Sensors |
| FRET | Förster Resonance Energy Transfer |
| FACS | Fluorescence-Activated Cell Sorting |
| SELEX | Systematic Evolution of Ligands by EXponential enrichment |
| GO | Graphene Oxide |
| PCR | Polymerase Chain Reaction |
| PAGE | Polyacrylamide Gel Electrophoresis |
| CHARGE | Catalytic Hairpin Assembly RNA circuit that is Genetically Encoded |
| BODIPY | Boron Dipyrromethene |
| FP, GFP, EGFP, BFP, RFP | Fluorescent Protein, Green Fluorescent Protein, Enhanced Green Fluorescent Protein, Blue Fluorescent Protein, Red Fluorescent Protein |
| DFHBI | 3,5-difluoro-4-hydroxybenzylidene Imidazolinone |
| DFHO | 3,5-difluoro-4-hydroxybenzylidene Imidazolinone-2-oxime |
| DNB, SRB, SR-DN | Dinitroaniline-Binding aptamer, Sulforhodamine B, Sulforhodamine-Dinitroaniline |
| TO-Biotin | Thizole Orange-Biotin |
| Co2+, Ag+ | Cobalt ion, Silver ion |
| ATP, ADP, AMP, GMP | Adenosine Triphosphate, Adenosine Diphosphate, Adenosine Monophosphate, Guanosine Monophosphate |
| HIV | Human Immunodeficiency Virus |
| SAH, SAM | S-adenosylhomocysteine, S-adenosylmethionine |
| MTase | Methyltransferase |
| TPP | Thiamine 5’-pyrophosphate |
| MCP | MS2 Coat Protein |
| 5-HTP | 5-hydroxytryptophan |
| L-DOPA | L-3,4-dihydroxyphenylalanine |
References
- Spiller, D.G.; Wood, C.D.; Rand, D.A.; White, M.R.H. Measurement of Single-Cell Dynamics. Nature 2010, 465, 736–745. [Google Scholar] [CrossRef] [PubMed]
- Carlo, D.D.; Lee, L.P. Dynamic Single-Cell Analysis for Quantitative Biology. Anal. Chem. 2006, 78, 7918–7925. [Google Scholar] [CrossRef]
- Jares-Erijman, E.A.; Jovin, T.M. FRET Imaging. Nat. Biotechnol. 2003, 21, 1387–1395. [Google Scholar] [CrossRef]
- Manz, A.; Dittrich, P.S.; Pamme, N.; Iossifidis, D. Bioanalytical Chemistry; IMPERIAL COLLEGE PRESS: London, UK, 2015. [Google Scholar]
- Zhang, J.; Campbell, R.E.; Ting, A.Y.; Tsien, R.Y. Creating New Fluorescent Probes for Cell Biology. Nat. Rev. Mol. Cell Biol. 2002, 3, 906–918. [Google Scholar] [CrossRef] [PubMed]
- Specht, E.A.; Braselmann, E.; Palmer, A.E. A Critical and Comparative Review of Fluorescent Tools for Live-Cell Imaging. Annu. Rev. Physiol. 2017, 79, 93–117. [Google Scholar] [CrossRef]
- Wysocki, L.M.; Lavis, L.D. Advances in the Chemistry of Small Molecule Fluorescent Probes. Curr. Opin. Chem. Biol. 2011, 15, 753–759. [Google Scholar] [CrossRef]
- Yuan, L.; Lin, W.; Zheng, K.; Zhu, S. FRET-Based Small-Molecule Fluorescent Probes: Rational Design and Bioimaging Applications. Acc. Chem. Res. 2013, 46, 1462–1473. [Google Scholar] [CrossRef]
- Fernández-Suárez, M.; Ting, A.Y. Fluorescent Probes for Super-Resolution Imaging in Living Cells. Nat. Rev. Mol. Cell Biol. 2008, 9, 929–943. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, E.; Bértolo, E.; Núñez, C.; Pilla, V.; Santos, H.M.; Fernández-Lodeiro, J.; Fernández-Lodeiro, A.; Djafari, J.; Capelo, J.L.; Lodeiro, C. Green and Red Fluorescent Dyes for Translational Applications in Imaging and Sensing Analytes: A Dual-Color Flag. ChemistryOpen 2018, 7, 3. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Zhou, B.; Ye, R.; Zhu, J.; Bao, X. Synthesis and Evaluation of a New Fluorescein and Rhodamine B-Based Chemosensor for Highly Sensitive and Selective Detection of Cysteine over Other Amino Acids and Its Application in Living Cell Imaging. Sens. Actuat. B-Chem. 2017, 251, 481–489. [Google Scholar] [CrossRef]
- Loudet, A.; Burgess, K. BODIPY Dyes and Their Derivatives: Syntheses and Spectroscopic Properties. Chem. Rev. 2007, 107, 4891–4932. [Google Scholar] [CrossRef]
- Luo, S.; Zhang, E.; Su, Y.; Cheng, T.; Shi, C. A Review of NIR Dyes in Cancer Targeting and Imaging. Biomaterials 2011, 32, 7127–7138. [Google Scholar] [CrossRef]
- Ueno, T.; Nagano, T. Fluorescent Probes for Sensing and Imaging. Nat. Methods 2011, 8, 642–645. [Google Scholar] [CrossRef]
- Van de Linde, S.; Aufmkolk, S.; Franke, C.; Holm, T.; Klein, T.; Löschberger, A.; Proppert, S.; Wolter, S.; Sauer, M. Investigating Cellular Structures at the Nanoscale with Organic Fluorophores. Chem. Biol. 2013, 20, 8–18. [Google Scholar] [CrossRef]
- Zhu, H.; Fan, J.; Du, J.; Peng, X. Fluorescent Probes for Sensing and Imaging within Specific Cellular Organelles. Acc. Chem. Res. 2016, 49, 2115–2126. [Google Scholar] [CrossRef]
- Alford, R.; Simpson, H.M.; Duberman, J.; Hill, G.C.; Ogawa, M.; Regino, C.; Kobayashi, H.; Choyke, P.L. Toxicity of Organic Fluorophores Used in Molecular Imaging: Literature Review. Mol. Imaging 2009, 8, 7290.2009.00031. [Google Scholar] [CrossRef]
- Crawford, J.M.; Braunwald, N.S. Toxicity in Vital Fluorescence Microscopy: Effect of Dimethylsulfoxide, Rhodamine-123, and DiI-Low Density Lipoprotein on Fibroblast Growth in Vitro. Vitr. Cell. Dev. Biol.-Anim. 1991, 27, 633–638. [Google Scholar] [CrossRef]
- Fei, X.; Gu, Y. Progress in Modifications and Applications of Fluorescent Dye Probe. Prog. Nat. Sci. 2009, 19, 1–7. [Google Scholar] [CrossRef]
- Heim, R.; Prasher, D.C.; Tsien, R.Y. Wavelength Mutations and Posttranslational Autoxidation of Green Fluorescent Protein. Proc. Natl. Acad. Sci. USA 1994, 91, 12501–12504. [Google Scholar] [CrossRef]
- Morise, H.; Shimomura, O.; Johnson, F.H.; Winant, J. Intermolecular Energy Transfer in the Bioluminescent System of Aequorea. Biochemistry 1974, 13, 2656–2662. [Google Scholar] [CrossRef]
- Chudakov, D.M.; Matz, M.V.; Lukyanov, S.; Lukyanov, K.A. Fluorescent Proteins and Their Applications in Imaging Living Cells and Tissues. Physiol. Rev. 2010, 90, 1103–1163. [Google Scholar] [CrossRef]
- Day, R.N.; Davidson, M.W. The Fluorescent Protein Palette: Tools for Cellular Imaging. Chem. Soc. Rev. 2009, 38, 2887. [Google Scholar] [CrossRef]
- Kremers, G.-J.; Gilbert, S.G.; Cranfill, P.J.; Davidson, M.W.; Piston, D.W. Fluorescent Proteins at a Glance. J. Cell Sci. 2011, 124, 157–160. [Google Scholar] [CrossRef]
- Davidson, M.W.; Campbell, R.E. Engineered Fluorescent Proteins: Innovations and Applications. Nat. Methods 2009, 6, 713–717. [Google Scholar] [CrossRef]
- Shaner, N.C.; Patterson, G.H.; Davidson, M.W. Advances in Fluorescent Protein Technology. J. Cell Sci. 2007, 120, 4247–4260. [Google Scholar] [CrossRef]
- Okumoto, S.; Jones, A.; Frommer, W.B. Quantitative Imaging with Fluorescent Biosensors. Annu. Rev. Plant Biol. 2012, 63, 663–706. [Google Scholar] [CrossRef]
- Piston, D.W.; Kremers, G.-J. Fluorescent Protein FRET: The Good, the Bad and the Ugly. Trends Biochem. Sci. 2007, 32, 407–414. [Google Scholar] [CrossRef]
- Paige, J.S.; Nguyen-Duc, T.; Song, W.; Jaffrey, S.R. Fluorescence Imaging of Cellular Metabolites with RNA. Science 2012, 335, 1194. [Google Scholar] [CrossRef]
- Song, W.; Strack, R.L.; Jaffrey, S.R. Imaging Bacterial Protein Expression Using Genetically Encoded RNA Sensors. Nat. Methods 2013, 10, 873–875. [Google Scholar] [CrossRef]
- Song, W.; Strack, R.L.; Svensen, N.; Jaffrey, S.R. Plug-and-Play Fluorophores Extend the Spectral Properties of Spinach. J. Am. Chem. Soc. 2014, 136, 1198–1201. [Google Scholar] [CrossRef]
- You, M.; Litke, J.L.; Jaffrey, S.R. Imaging Metabolite Dynamics in Living Cells Using a Spinach-Based Riboswitch. Proc. Natl. Acad. Sci. USA 2015, 112, E2756–E2765. [Google Scholar] [CrossRef]
- Song, K.-M.; Lee, S.; Ban, C. Aptamers and Their Biological Applications. Sensors 2012, 12, 612–631. [Google Scholar] [CrossRef]
- Lakhin, A.V.; Tarantul, V.Z.; Gening, L.V. Aptamers: Problems, Solutions and Prospects. Acta Nat. 2013, 5, 34–43. [Google Scholar]
- Paige, J.S.; Wu, K.Y.; Jaffrey, S.R. RNA Mimics of Green Fluorescent Protein. Science 2011, 333, 642–646. [Google Scholar] [CrossRef]
- Iii, R.J.T.; Truong, L.; Ferré-D’amaré, A.R. Structural Principles of Fluorescent RNA Aptamers. Trends Pharmacol. Sci. 2017, 38, 928–939. [Google Scholar] [CrossRef]
- Ouellet, J. RNA Fluorescence with Light-Up Aptamers. Front. Chem. 2016, 4, 29. [Google Scholar] [CrossRef]
- Kellenberger, C.A.; Wilson, S.C.; Sales-Lee, J.; Hammond, M.C. RNA-Based Fluorescent Biosensors for Live Cell Imaging of Second Messengers Cyclic Di-GMP and Cyclic AMP-GMP. J. Am. Chem. Soc. 2013, 135, 4906–4909. [Google Scholar] [CrossRef]
- Kellenberger, C.A.; Chen, C.; Whiteley, A.T.; Portnoy, D.A.; Hammond, M.C. RNA-Based Fluorescent Biosensors for Live Cell Imaging of Second Messenger Cyclic Di-AMP. J. Am. Chem. Soc. 2015, 137, 6432–6435. [Google Scholar] [CrossRef]
- Cromie, M.J.; Shi, Y.; Latifi, T.; Groisman, E.A. An RNA Sensor for Intracellular Mg2+. Cell 2006, 125, 71–84. [Google Scholar] [CrossRef]
- Bose, D.; Su, Y.; Marcus, A.; Raulet, D.H.; Hammond, M.C. An RNA-Based Fluorescent Biosensor for High-Throughput Analysis of the CGAS-CGAMP-STING Pathway. Cell Chem. Biol. 2016, 23, 1539–1549. [Google Scholar] [CrossRef]
- Win, M.N.; Smolke, C.D. A Modular and Extensible RNA-Based Gene-Regulatory Platform for Engineering Cellular Function. Proc. Natl. Acad. Sci. USA 2007, 104, 14283–14288. [Google Scholar] [CrossRef]
- Wang, R.E.; Zhang, Y.; Cai, J.; Cai, W.; Gao, T. Aptamer-Based Fluorescent Biosensors. Curr. Med. Chem. 2011, 18, 4175–4184. [Google Scholar] [CrossRef]
- Strack, R.L.; Jaffrey, S.R. New Approaches for Sensing Metabolites and Proteins in Live Cells Using RNA. Curr. Opin. Chem. Biol. 2013, 17, 651–655. [Google Scholar] [CrossRef]
- Kellenberger, C.A.; Hammond, M.C. In Vitro Analysis of Riboswitch–Spinach Aptamer Fusions as Metabolite-Sensing Fluorescent Biosensors. Methods Enzymol. 2015, 550, 147–172. [Google Scholar] [CrossRef]
- Jepsen, M.D.E.E.; Sparvath, S.M.; Nielsen, T.B.; Langvad, A.H.; Grossi, G.; Gothelf, K.V.; Andersen, E.S. Development of a Genetically Encodable FRET System Using Fluorescent RNA Aptamers. Nat. Commun. 2018, 9, 1–10. [Google Scholar] [CrossRef]
- Zhang, C.; Wei, Z.H.; Ye, B.C. Imaging and Tracing of Intracellular Metabolites Utilizing Genetically Encoded Fluorescent Biosensors. Biotechnol. J. 2013, 8, 1280–1291. [Google Scholar] [CrossRef]
- Chakraborty, K.; Veetil, A.T.; Jaffrey, S.R.; Krishnan, Y. Nucleic Acid-Based Nanodevices in Biological Imaging. Annu. Rev. Biochem. 2016, 85, 349–373. [Google Scholar] [CrossRef]
- Feagin, T.A.; Maganzini, N.; Soh, H.T. Strategies for Creating Structure-Switching Aptamers. ACS Sens. 2018, 3, 1611–1615. [Google Scholar] [CrossRef]
- Hall, B.; Hesselberth, J.R.; Ellington, A.D. Computational Selection of Nucleic Acid Biosensors via a Slip Structure Model. Biosens. Bioelectron. 2007, 22, 1939–1947. [Google Scholar] [CrossRef]
- Winkler, W.; Nahvi, A.; Breaker, R.R. Thiamine Derivatives Bind Messenger RNAs Directly to Regulate Bacterial Gene Expression. Nature 2002, 419, 952–956. [Google Scholar] [CrossRef]
- Espah Borujeni, A.; Mishler, D.M.; Wang, J.; Huso, W.; Salis, H.M. Automated Physics-Based Design of Synthetic Riboswitches from Diverse RNA Aptamers. Nucleic Acids Res. 2016, 44, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Chappell, J.; Westbrook, A.; Verosloff, M.; Lucks, J.B. Computational Design of Small Transcription Activating RNAs for Versatile and Dynamic Gene Regulation. Nat. Commun. 2017, 8, 1051. [Google Scholar] [CrossRef] [PubMed]
- Ausländer, S.; Fuchs, D.; Hürlemann, S.; Ausländer, D.; Fussenegger, M. Engineering a Ribozyme Cleavage-Induced Split Fluorescent Aptamer Complementation Assay. Nucleic Acids Res. 2016, 44, e94. [Google Scholar] [CrossRef] [PubMed]
- Felletti, M.; Stifel, J.; Wurmthaler, L.A.; Geiger, S.; Hartig, J.S. Twister Ribozymes as Highly Versatile Expression Platforms for Artificial Riboswitches. Nat. Commun. 2016, 7, 12834. [Google Scholar] [CrossRef] [PubMed]
- Furukawa, K.; Gu, H.; Breaker, R.R. In Vitro Selection of Allosteric Ribozymes That Sense the Bacterial Second Messenger C-Di-GMP. Methods Mol. Biol. 2014, 1111, 209–220. [Google Scholar] [CrossRef] [PubMed]
- Murray, J.B.; Terwey, D.P.; Maloney, L.; Karpeisky, A.; Usman, N.; Beigelman, L.; Scott, W.G. The Structural Basis of Hammerhead Ribozyme Self-Cleavage. Cell 1998, 92, 665–673. [Google Scholar] [CrossRef]
- Lee, E.R.; Baker, J.L.; Weinberg, Z.; Sudarsan, N.; Breaker, R.R. An Allosteric Self-Splicing Ribozyme Triggered by a Bacterial Second Messenger. Science 2010, 329, 845–848. [Google Scholar] [CrossRef]
- Roth, A.; Weinberg, Z.; Chen, A.G.Y.; Kim, P.B.; Ames, T.D.; Breaker, R.R. A Widespread Self-Cleaving Ribozyme Class Is Revealed by Bioinformatics. Nat. Chem. Biol. 2014, 10, 56–60. [Google Scholar] [CrossRef]
- Weinberg, Z.; Kim, P.B.; Chen, T.H.; Li, S.; Harris, K.A.; Lünse, C.E.; Breaker, R.R. New Classes of Self-Cleaving Ribozymes Revealed by Comparative Genomics Analysis. Nat. Chem. Biol. 2015, 11, 606–610. [Google Scholar] [CrossRef]
- Lilley, D.M.J. The Varkud Satellite Ribozyme. RNA 2004, 10, 151–158. [Google Scholar] [CrossRef]
- Fedor, M.J. Structure and Function of the Hairpin Ribozyme. J. Mol. Biol. 2000, 297, 269–291. [Google Scholar] [CrossRef] [PubMed]
- Ferré-D’Amaré, A.R.; Scott, W.G. Small Self-Cleaving Ribozymes. Cold Spring Harb. Perspect. Biol. 2010, 2, a003574. [Google Scholar] [CrossRef] [PubMed]
- Vesuna, F.; Winnard, P.; Raman, V. Enhanced Green Fluorescent Protein as an Alternative Control Reporter to Renilla Luciferase. Anal. Biochem. 2005, 342, 345–347. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.S.; Jones, K.A.; McCutcheon, D.C.; Prescher, J.A. Pyridone Luciferins and Mutant Luciferases for Bioluminescence Imaging. ChemBioChem 2018, 19, 470–477. [Google Scholar] [CrossRef] [PubMed]
- Culler, S.J.; Hoff, K.G.; Smolke, C.D. Reprogramming Cellular Behavior with RNA Controllers Responsive to Endogenous Proteins. Science 2010, 330, 1251–1255. [Google Scholar] [CrossRef] [PubMed]
- Yoshimatsu, T.; Nagawa, F. Control of Gene Expression by Artificial Introns in Saccharomyces Cerevisiae. Science 1989, 244, 1346–1348. [Google Scholar] [CrossRef] [PubMed]
- Swinburne, I.A.; Miguez, D.G.; Landgraf, D.; Silver, P.A. Intron Length Increases Oscillatory Periods of Gene Expression in Animal Cells. Genes Dev. 2008, 22, 2342–2346. [Google Scholar] [CrossRef]
- Carothers, J.M.; Goler, J.A.; Juminaga, D.; Keasling, J.D. Model-Driven Engineering of RNA Devices to Quantitatively Program Gene Expression. Science 2011, 334, 1716–1719. [Google Scholar] [CrossRef]
- Purnick, P.E.M.; Weiss, R. The second wave of synthetic biology: From modules to systems. Nat. Rev. Mol. Cell Biol. 2009, 10, 410–422. [Google Scholar] [CrossRef]
- Ausländer, S.; Ketzer, P.; Hartig, J.S. A Ligand-Dependent Hammerhead Ribozyme Switch for Controlling Mammalian Gene Expression. Mol. Biosyst. 2010, 6, 807. [Google Scholar] [CrossRef]
- Qi, L.; Haurwitz, R.E.; Shao, W.; Doudna, J.A.; Arkin, A.P. RNA Processing Enables Predictable Programming of Gene Expression. Nat. Biotechnol. 2012, 30, 1002–1006. [Google Scholar] [CrossRef] [PubMed]
- Salmena, L.; Poliseno, L.; Tay, Y.; Kats, L.; Pandolfi, P.P. A CeRNA Hypothesis: The Rosetta Stone of a Hidden RNA Language? Cell 2011, 146, 353–358. [Google Scholar] [CrossRef] [PubMed]
- Rackham, O.; Chin, J.W. A Network of Orthogonal Ribosome·mRNA Pairs. Nat. Chem. Biol. 2005, 1, 159–166. [Google Scholar] [CrossRef] [PubMed]
- Anderson, J.C.; Voigt, C.A.; Arkin, A.P. Environmental Signal Integration by a Modular AND Gate. Mol. Syst. Biol. 2007, 3, 133. [Google Scholar] [CrossRef] [PubMed]
- Venkatesan, A.; Dasgupta, A. Novel Fluorescence-Based Screen to Identify Small Synthetic Internal Ribosome Entry Site Elements. Mol. Cell Biol. 2001, 21, 2826–2837. [Google Scholar] [CrossRef] [PubMed]
- Salis, H.M.; Mirsky, E.A.; Voigt, C.A. Automated Design of Synthetic Ribosome Binding Sites to Control Protein Expression. Nat. Biotechnol. 2009, 27, 946–950. [Google Scholar] [CrossRef] [PubMed]
- Ketterer, S.; Fuchs, D.; Weber, W.; Meier, M. Systematic Reconstruction of Binding and Stability Landscapes of the Fluorogenic Aptamer Spinach. Nucleic Acids Res. 2015, 43, 9564–9572. [Google Scholar] [CrossRef]
- Filonov, G.S.; Moon, J.D.; Svensen, N.; Jaffrey, S.R. Broccoli: Rapid Selection of an RNA Mimic of Green Fluorescent Protein by Fluorescence-Based Selection and Directed Evolution. J. Am. Chem. Soc. 2014, 136, 16299–16308. [Google Scholar] [CrossRef]
- Strack, R.L.; Disney, M.D.; Jaffrey, S.R. A Superfolding Spinach2 Reveals the Dynamic Nature of Trinucleotide Repeat–containing RNA. Nat. Methods 2013, 10, 1219–1224. [Google Scholar] [CrossRef]
- Okuda, M.; Fourmy, D.; Yoshizawa, S. Use of Baby Spinach and Broccoli for Imaging of Structured Cellular RNAs. Nucleic Acids Res. 2016, 45, gkw794. [Google Scholar] [CrossRef]
- Alam, K.K.; Tawiah, K.D.; Lichte, M.F.; Porciani, D.; Burke, D.H. A Fluorescent Split Aptamer for Visualizing RNA–RNA Assembly In Vivo. ACS Synth. Biol. 2017, 6, 1710–1721. [Google Scholar] [CrossRef] [PubMed]
- Song, W.; Filonov, G.S.; Kim, H.; Hirsch, M.; Li, X.; Moon, J.D.; Jaffrey, S.R. Imaging RNA Polymerase III Transcription Using a Photostable RNA-Fluorophore Complex. Nat. Chem. Biol. 2017, 13, 1187–1194. [Google Scholar] [CrossRef] [PubMed]
- Dolgosheina, E.V.; Jeng, S.C.Y.; Panchapakesan, S.S.S.; Cojocaru, R.; Chen, P.S.K.; Wilson, P.D.; Hawkins, N.; Wiggins, P.A.; Unrau, P.J. RNA Mango Aptamer-Fluorophore: A Bright, High-Affinity Complex for RNA Labeling and Tracking. ACS Chem. Biol. 2014, 9, 2412–2420. [Google Scholar] [CrossRef] [PubMed]
- Autour, A.; Jeng, S.C.Y.; Cawte, A.D.; Abdolahzadeh, A.; Galli, A.; Panchapakesan, S.S.S.; Rueda, D.; Ryckelynck, M.; Unrau, P.J.; Jeng, S.C.Y.; et al. Fluorogenic RNA Mango Aptamers for Imaging Small Non-Coding RNAs in Mammalian Cells. Nat. Commun. 2018, 9, 656. [Google Scholar] [CrossRef]
- Sunbul, M.; Jäschke, A. Contact-Mediated Quenching for RNA Imaging in Bacteria with a Fluorophore-Binding Aptamer. Angew. Chem. Int. Ed. 2013, 52, 13401–13404. [Google Scholar] [CrossRef] [PubMed]
- Arora, A.; Sunbul, M.; Jäschke, A. Dual-Colour Imaging of RNAs Using Quencher- and Fluorophore-Binding Aptamers. Nucleic Acids Res. 2015, 43, gkv718. [Google Scholar] [CrossRef] [PubMed]
- Warner, K.D.; Chen, M.C.; Song, W.; Strack, R.L.; Thorn, A.; Jaffrey, S.R.; Ferré-D’Amaré, A.R. Structural Basis for Activity of Highly Efficient RNA Mimics of Green Fluorescent Protein. Nat. Struct. Mol. Biol. 2014, 21, 658–663. [Google Scholar] [CrossRef]
- Karunanayake Mudiyanselage, A.P.K.K.; Wu, R.; Leon-Duque, M.A.; Ren, K.; You, M. “Second-Generation” Fluorogenic RNA-Based Sensors. Methods 2019, in press. [Google Scholar] [CrossRef]
- Ellington, A.D.; Szostak, J.W. In Vitro Selection of RNA Molecules that Bind Specific Ligands. Nature 1990, 346, 818–822. [Google Scholar] [CrossRef]
- Tuerk, C.; Gold, L. Systematic Evolution of Ligands by Exponential Enrichment: RNA Ligands to Bacteriophage T4 DNA Polymerase. Science 1990, 249, 505–510. [Google Scholar] [CrossRef]
- Stoltenburg, R.; Reinemann, C.; Strehlitz, B. SELEX—A (r)Evolutionary Method to Generate High-Affinity Nucleic Acid Ligands. Biomol. Eng. 2007, 24, 381–403. [Google Scholar] [CrossRef] [PubMed]
- Mandal, M.; Breaker, R.R. Gene Regulation by Riboswitches. Nat. Rev. Mol. Cell Biol. 2004, 5, 451–463. [Google Scholar] [CrossRef] [PubMed]
- Wieland, M.; Benz, A.; Klauser, B.; Hartig, J.S. Artificial Ribozyme Switches Containing Natural Riboswiteh Aptamer Domains. Angew. Chem. Int. Ed. 2009, 48, 2715–2718. [Google Scholar] [CrossRef] [PubMed]
- Breaker, R.R.; McCown, P.; Corbino, K.; Stav, S.; Sherlock, M. Riboswitch Diversity and Distribution. RNA 2017, rna-061234. [Google Scholar] [CrossRef]
- Aquino-Jarquin, G.; Toscano-Garibay, J.D. RNA Aptamer Evolution: Two Decades of SELEction. Int. J. Mol. Sci. 2011, 12, 9155–9171. [Google Scholar] [CrossRef] [PubMed]
- Wrzesinski, J.; Ciesiolka, J. Characterization of Structure and Metal Ions Specificity of Co2+-Binding RNA Aptamers. Biochemistry 2005, 44, 6257–6268. [Google Scholar] [CrossRef]
- Yarus, M. A Specific Amino Acid Binding Site Composed of RNA. Science 1988, 240, 1751–1758. [Google Scholar] [CrossRef] [PubMed]
- Sassanfar, M.; Szostak, J.W. An RNA Motif that Binds ATP. Nature 1993, 364, 550–553. [Google Scholar] [CrossRef] [PubMed]
- Mehlhorn, A.; Rahimi, P.; Joseph, Y. Aptamer-Based Biosensors for Antibiotic Detection: A Review. Biosensors 2018, 8, 54. [Google Scholar] [CrossRef] [PubMed]
- Lorsch, J.R.; Szostak, J.W. In Vitro Selection of RNA Aptamers Specific for Cyanocobalamin. Biochemistry 1994, 33, 973–982. [Google Scholar] [CrossRef]
- Bouhedda, F.; Autour, A.; Ryckelynck, M. Light-up RNA Aptamers and Their Cognate Fluorogens: From Their Development to Their Applications. Int. J. Mol. Sci. 2018, 19, 44. [Google Scholar] [CrossRef] [PubMed]
- White, R.; Rusconi, C.; Scardino, E.; Wolberg, A.; Lawson, J.; Hoffman, M.; Sullenger, B. Generation of Species Cross-Reactive Aptamers Using “Toggle” SELEX. Mol. Ther. 2001, 4, 567–573. [Google Scholar] [CrossRef]
- Mondragón, E.; Maher, L.J., III. Anti-Transcription Factor RNA Aptamers as Potential Therapeutics. Nucleic Acid Ther. 2016, 26, 29–43. [Google Scholar] [CrossRef] [PubMed]
- Tuerk, C.; MacDougal-Waugh, S. In Vitro Evolution of Functional Nucleic Acids: High-Affinity RNA Ligands of HIV-1 Proteins. Gene 1993, 137, 33–39. [Google Scholar] [CrossRef]
- Davydova, A.; Vorobjeva, M.; Pyshnyi, D.; Altman, S.; Vlassov, V.; Venyaminova, A. Aptamers against Pathogenic Microorganisms. Crit. Rev. Microbiol. 2016, 42, 847–865. [Google Scholar] [CrossRef] [PubMed]
- Stoltenburg, R.; Nikolaus, N.; Strehlitz, B. Capture-SELEX: Selection of DNA Aptamers for Aminoglycoside Antibiotics. J. Anal. Methods Chem. 2012, 2012, 415697. [Google Scholar] [CrossRef] [PubMed]
- Kruger, K.; Grabowski, P.J.; Zaug, A.J.; Sands, J.; Gottschling, D.E.; Cech, T.R. Self-Splicing RNA: Autoexcision and Autocyclization of the Ribosomal RNA Intervening Sequence of Tetrahymena. Cell 1982, 31, 147–157. [Google Scholar] [CrossRef]
- Usman, N.; Beigelman, L.; McSwiggen, J.A. Hammerhead ribozyme engineering. Curr. Opin. Struct. Biol. 1996, 6, 527–533. [Google Scholar] [CrossRef]
- Park, J.-W.; Tatavarty, R.; Kim, D.W.; Jung, H.-T.; Gu, M.B. Immobilization-Free Screening of Aptamers Assisted by Graphene Oxide. Chem. Commun. 2012, 48, 2071–2073. [Google Scholar] [CrossRef]
- Nguyen, V.-T.; Kwon, Y.S.; Kim, J.H.; Gu, M.B. Multiple GO-SELEX for Efficient Screening of Flexible Aptamers. Chem. Commun. 2014, 50, 10513–10516. [Google Scholar] [CrossRef] [PubMed]
- Garst, A.; Edwards, A.L.; Batey, R.T. Riboswitches: Structures and Mechanisms. Cold Spring Harb. Perspect. Biol. 2011, 3, a003533. [Google Scholar] [CrossRef] [PubMed]
- Serganov, A.; Patel, D.J. Metabolite Recognition Principles and Molecular Mechanisms Underlying Riboswitch Function. Annu. Rev. Biophys. 2012, 41, 343–370. [Google Scholar] [CrossRef] [PubMed]
- Hammann, C.; Westhof, E. Searching Genomes for Ribozymes and Riboswitches. Genome Biol. 2007, 8, 210. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.X.; Lee, E.R.; Morales, D.R.; Lim, J.; Breaker, R.R. Riboswitches That Sense S-Adenosylhomocysteine and Activate Genes Involved in Coenzyme Recycling. Mol. Cell 2008, 29, 691–702. [Google Scholar] [CrossRef] [PubMed]
- Su, Y.; Hickey, S.F.; Keyser, S.G.L.; Hammond, M.C. In Vitro and In Vivo Enzyme Activity Screening via RNA-Based Fluorescent Biosensors for S-Adenosyl-l-Homocysteine (SAH). J. Am. Chem. Soc. 2016, 138, 7040–7047. [Google Scholar] [CrossRef] [PubMed]
- Yu, Q.; Shi, J.; Mudiyanselage, A.P.K.K.K.; Wu, R.; Zhao, B.; Zhou, M.; You, M. Genetically Encoded RNA-Based Sensors for Intracellular Imaging of Silver Ions. Chem. Commun. 2019, 55, 707–710. [Google Scholar] [CrossRef] [PubMed]
- Porter, E.B.; Polaski, J.T.; Morck, M.M.; Batey, R.T. Recurrent RNA Motifs as Scaffolds for Genetically Encodable Small-Molecule Biosensors. Nat. Chem. Biol. 2017, 13, 295–301. [Google Scholar] [CrossRef]
- Kennedy, A.B.; Vowles, J.V.; d’Espaux, L.; Smolke, C.D. Protein-Responsive Ribozyme Switches in Eukaryotic Cells. Nucleic Acids Res. 2014, 42, 12306–12321. [Google Scholar] [CrossRef]
- Klauser, B.; Atanasov, J.; Siewert, L.K.; Hartig, J.S. Ribozyme-Based Aminoglycoside Switches of Gene Expression Engineered by Genetic Selection in S. Cerevisiae. ACS Synth. Biol. 2015, 4, 516–525. [Google Scholar] [CrossRef]
- Win, M.N.; Smolke, C.D. Higher-Order Cellular Information Processing with Synthetic RNA Devices. Science 2008, 322, 456–460. [Google Scholar] [CrossRef]
- Beilstein, K.; Wittmann, A.; Grez, M.; Suess, B. Conditional Control of Mammalian Gene Expression by Tetracycline-Dependent Hammerhead Ribozymes. ACS Synth. Biol. 2015, 4, 526–534. [Google Scholar] [CrossRef] [PubMed]
- Wieland, M.; Hartig, J.S. Improved Aptazyme Design and In Vivo Screening Enable Riboswitching in Bacteria. Angew. Chem. Int. Ed. 2008, 47, 2604–2607. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.C.; Wilson, S.C.; Hammond, M.C. Next-Generation RNA-Based Fluorescent Biosensors Enable Anaerobic Detection of Cyclic Di-GMP. Nucleic Acids Res. 2016, 44, e139. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Zheng, C.; Su, J.; Chen, B.; Fu, Y.; Xie, Y.; Tang, Q.; Chou, S.-H.; He, J. Characterization of a Natural Triple-Tandem c-Di-GMP Riboswitch and Application of the Riboswitch-Based Dual-Fluorescence Reporter. Sci. Rep. 2016, 6, 20871. [Google Scholar] [CrossRef] [PubMed]
- Nomura, Y.; Kumar, D.; Yokobayashi, Y. Synthetic Mammalian Riboswitches Based on Guanine Aptazyme. Chem. Commun. 2012, 48, 7215. [Google Scholar] [CrossRef] [PubMed]
- Wachter, A.; Tunc-Ozdemir, M.; Grove, B.C.; Green, P.J.; Shintani, D.K.; Breaker, R.R. Riboswitch Control of Gene Expression in Plants by Splicing and Alternative 3’ End Processing of MRNAs. Plant Cell 2007, 19, 3437–3450. [Google Scholar] [CrossRef] [PubMed]
- Saragliadis, A.; Hartig, J.S. Ribozyme-Based Transfer RNA Switches for Post-Transcriptional Control of Amino Acid Identity in Protein Synthesis. J. Am. Chem. Soc. 2013, 135, 8222–8226. [Google Scholar] [CrossRef]
- Goldsworthy, V.; LaForce, G.; Abels, S.; Khisamutdinov, E. Fluorogenic RNA Aptamers: A Nano-Platform for Fabrication of Simple and Combinatorial Logic Gates. Nanomaterials 2018, 8, 984. [Google Scholar] [CrossRef]
- Hansen, T.B.; Jensen, T.I.; Clausen, B.H.; Bramsen, J.B.; Finsen, B.; Damgaard, C.K.; Kjems, J. Natural RNA Circles Function as Efficient MicroRNA Sponges. Nature 2013, 495, 384–388. [Google Scholar] [CrossRef]
- Karunanayake Mudiyanselage, A.P.K.K.; Yu, Q.; Leon-Duque, M.A.; Zhao, B.; Wu, R.; You, M. Genetically Encoded Catalytic Hairpin Assembly for Sensitive RNA Imaging in Live Cells. J. Am. Chem. Soc. 2018, 140, 8739–8745. [Google Scholar] [CrossRef]
- You, M.; Litke, J.L.; Wu, R.; Jaffrey, S.R. Detection of Low-Abundance Metabolites in Live Cells Using an RNA Integrator. Cell Chem. Biol. 2019, 26. in press. [Google Scholar] [CrossRef]
- Yin, P.; Choi, H.M.T.; Calvert, C.R.; Pierce, N.A. Programming Biomolecular Self-Assembly Pathways. Nature 2008, 451, 318–322. [Google Scholar] [CrossRef] [PubMed]
- Green, A.A.; Silver, P.A.; Collins, J.J.; Yin, P. Toehold Switches: De-Novo-Designed Regulators of Gene Expression. Cell 2014, 159, 925–939. [Google Scholar] [CrossRef] [PubMed]
- Memczak, S.; Jens, M.; Elefsinioti, A.; Torti, F.; Krueger, J.; Rybak, A.; Maier, L.; Mackowiak, S.D.; Gregersen, L.H.; Munschauer, M.; et al. Circular RNAs Are a Large Class of Animal RNAs with Regulatory Potency. Nature 2013, 495, 333–338. [Google Scholar] [CrossRef]
- Barrett, S.P.; Salzman, J. Circular RNAs: Analysis, Expression and Potential Functions. Development 2016, 143, 1838–1847. [Google Scholar] [CrossRef] [PubMed]
- Holdt, L.M.; Kohlmaier, A.; Teupser, D. Molecular Roles and Function of Circular RNAs in Eukaryotic Cells. Cell. Mol. Life Sci. 2018, 75, 1071–1098. [Google Scholar] [CrossRef] [PubMed]
- Wesselhoeft, R.A.; Kowalski, P.S.; Anderson, D.G. Engineering Circular RNA for Potent and Stable Translation in Eukaryotic Cells. Nat. Commun. 2018, 9, 2629. [Google Scholar] [CrossRef] [PubMed]
| Aptamer | Fluorophore | KD (nM) | Ex./Em. (nm) | Ɛ (M−1cm−1) a | ɸ b | Brightness c |
|---|---|---|---|---|---|---|
| Spinach [35] | DFHBI | 540 | 469/501 | 24,300 | 0.72 | 100 |
| Spinach2 [31] | DFHBI | 530 | 447/501 | 22,000 | 0.72 | 91 |
| Spinach2 [31] | DFHBI-1T | 560 | 482/505 | 31,000 | 0.94 | 167 |
| Spinach2 [31] | DFHBI-2T | 1300 | 500/523 | 29,000 | 0.12 | 20 |
| Broccoli [79] | DFHBI-1T | 360 | 472/507 | 29,600 | 0.94 | 159 |
| Corn [83] | DFHO | 70 | 505/545 | 29,000 | 0.25 | 41 |
| Mango [84] | TO1-Biotin | 3.2 | 510/535 | 77,500 | 0.14 | 62 |
| Mango [84] | TO3-Biotin | 5.1 | 637/658 | 9300 | N/A d | N/A |
| Mango II [85] | TO1-Biotin | 0.7 | 510/535 | 77,000 | 0.2 | 88 |
| Mango III [85] | TO1-Biotin | 5.6 | 510/535 | 77,000 | 0.56 | 247 |
| Mango IV [85] | TO1-Biotin | 11.1 | 510/535 | 77,000 | 0.42 | 185 |
| DNB [87] | SR-DN | 800 | 572/591 | 50,250 | 0.98 | 282 |
| SRB-2 [86] | SR-DN | 1400 | 579/596 | 85,200 | 0.65 | 317 |
| Target | Recognition Module Source | Transducer Module Type | Reporting System Type | EC50 (μM) | ON/OFF b | Cell System |
|---|---|---|---|---|---|---|
| ADP [29] | SELEX | Duplex Formation | Spinach | 270 | 20 | Bacteria |
| 5-HTP [118] | SELEX | Duplex Formation | Broccoli | N/A c | >5 | Bacteria |
| L-DOPA [118] | SELEX | Duplex Formation | Broccoli | N/A | >5 | Bacteria |
| MS2 coat protein [30] | SELEX | Duplex Formation | Spinach | ~0.6 | 41.7 | Bacteria |
| MS2 coat protein [119] | SELEX | Ribozyme | BFP | N/A | 1.8 | Mammalian |
| Neomycin [120] | SELEX | Ribozyme | β-galactosidase | N/A | 25 | Yeast |
| Streptavidin [30] | SELEX | Duplex Formation | Spinach | <0.2 | 10.3 | Bacteria |
| Tetracycline/Theophylline [121] | SELEX | Ribozyme | EGFP | N/A | N/A | Yeast |
| Tetracycline [122] | SELEX | Ribozyme | Luciferase/EGFP | 35.4 | 4.8 | Mammalian |
| Theophylline [123] | SELEX | Ribozyme | EGFP | ~200 | 10 | Bacteria |
| c-AMP-GMP [38] | Riboswitch | Duplex Formation | Spinach | 4.2 | ~8 (37 °C) | Bacteria |
| c-di-AMP [39] | Riboswitch | Strand Displacement | Spinach2 | 3.4 & 29 | 2.4 & 9.1 | Bacteria |
| c-di-GMP [38] | Riboswitch | Duplex Formation | Spinach | 0.23 | ~6 (37 °C) | Bacteria |
| c-di-GMP [124] | Riboswitch | Duplex Formation | Spinach2 | 0.005–0.4 | ~6 (37 °C) | Bacteria |
| c-di-GMP [125] | Riboswitch | Strand Displacement | TurboRFP | N/A | 38 | Bacteria |
| Guanine [126] | Riboswitch | Ribozyme | EGFP | N/A | 9.6 | Mammalian |
| SAM [29] | Riboswitch | Duplex Formation | Spinach | 120 | 25 | Bacteria |
| TPP [32] | Riboswitch | Strand Displacement | Spinach | 9 | 15.9 | Bacteria |
| TPP [127] | Riboswitch | Strand Displacement | EGFP | N/A | ~5 | Plant |
| TPP [128] | Riboswitch | Ribozyme | tRNA | N/A | 43 | Bacteria |
| N-peptide [129] | Ribonucleoprotein complexes | Ribozyme | EGFP/SEAP | N/A | ~12 | Mammalian |
| RNA [130] | Base Pairing | Ribozyme | EGFP | N/A | ~10 | Bacteria |
| RNA [131] | Base Pairing | Strand Displacement | Split Broccoli | ~0.001 | 2.2 | Bacteria |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sun, Z.; Nguyen, T.; McAuliffe, K.; You, M. Intracellular Imaging with Genetically Encoded RNA-Based Molecular Sensors. Nanomaterials 2019, 9, 233. https://doi.org/10.3390/nano9020233
Sun Z, Nguyen T, McAuliffe K, You M. Intracellular Imaging with Genetically Encoded RNA-Based Molecular Sensors. Nanomaterials. 2019; 9(2):233. https://doi.org/10.3390/nano9020233
Chicago/Turabian StyleSun, Zhining, Tony Nguyen, Kathleen McAuliffe, and Mingxu You. 2019. "Intracellular Imaging with Genetically Encoded RNA-Based Molecular Sensors" Nanomaterials 9, no. 2: 233. https://doi.org/10.3390/nano9020233
APA StyleSun, Z., Nguyen, T., McAuliffe, K., & You, M. (2019). Intracellular Imaging with Genetically Encoded RNA-Based Molecular Sensors. Nanomaterials, 9(2), 233. https://doi.org/10.3390/nano9020233






