Exosomes as Naturally Occurring Vehicles for Delivery of Biopharmaceuticals: Insights from Drug Delivery to Clinical Perspectives
Abstract
1. Introduction
2. Isolation or Purification Methods
2.1. Ultracentrifugation
2.2. Density Gradient Ultracentrifugation
2.3. Ultrafiltration
2.4. Size Exclusion Chromatography (SEC)
2.5. Precipitation Technique
2.6. Immunoisolation
2.7. Microfluidics-Derived Chip Isolation Methods
| Isolation or Purification Methods | Mechanism | Advantages | Limitations | References |
|---|---|---|---|---|
| Ultracentrifugation | Sedimentation coefficient of exosomes and other substances in a sample. | Capable of producing a large number of exosomes with high separation purity. | Poor repeatability, low recovery rate, requires more time, unsuitable for clinical diagnosis. | [65,66,67] |
| Density gradient ultracentrifugation | Separation based on the different densities. | Preserves exosome vesicle integrity, while yield can be maximized for samples that are already pure. | Difficult to scale-up, multi-step procedure. | [11,44,68] |
| Ultrafiltration | Separation based on size and molecular weight. | Faster, requires no special equipment, easy to handle compared to ultracentrifugation. | Deformation and breaking up of large vesicles may occur due to the use of force. | [69,70] |
| Size exclusion liquid chromatography | Utilizes a column packed with porous polymeric beads, which separate the exosomes based on size. | Allows separation of large and small molecules. The structure of the exosomes isolated by this method is not affected by shearing force compared to centrifugation methods. | Requires a long running time, which limits the processing of multiple biological samples. | [71,72] |
| Immunoaffinity capture-based techniques | Based on the specific interactions between immobilized antibodies (ligands) and membrane-bound antigens (receptors) of exosomes. | Suitable for isolation of specific exosomes, high possibility of subtyping, high-purity isolation. | High reagent cost, low capacity and yield, cannot be used for the separation of exosomes at a large scale. Requires non-physiological salt and pH conditions. | [66,73] |
| Precipitation technique | Change in the dispersibility or solubility of exosome vesicles using water excluding polymers. | Easy to use, requires no specialized equipment, and scalable sample capacity. | Requires long running times and preand postseparation cleanup. Co-precipitation of non-exosome contaminants within the sample. | [74,75,76] |
| Microfluidic technologies | Immunoaffinity, sieving, and trapping of exosomes on porous structure. | Quantitative technique that allows high-throughput analysis of exosome contents with high sensitivity. | Low sample capacity. Lack of standardization, method validation, and large-scale tests on clinical samples. | [70,77] |
3. Exosomes in Drug Delivery
3.1. Protein and Peptide Delivery:
3.2. Exosomes in Gene Delivery
3.3. Exosomes in Delivery of Other Therapeutic Compounds
| Cargo Type | Origin of Exosomes | Disease Type | Isolation or Purification Method | Drug Loading Method | Outcome | Reference | |
|---|---|---|---|---|---|---|---|
| Proteins | Signal regulatory protein α | Human embryonic kidney293T cells | Cancer | Centrifugation | Transfection | Enhanced phagocytosis of tumor cells | [82] |
| Survivin-T34A | Melanoma cell lines | Pancreatic cancer | Centrifugation | NA | Apoptotic death of cells | [128] | |
| Antiepidermal growth factor receptor | Mouse neuroblastoma | Epidermoid carcinoma | Ultrafiltration/size exclusion liquid chromatography | NA | Target specificity | [85] | |
| 20S proteasome | Mesenchymal stem cells | Mouse myocardium | Tangential flow filtration | NA | Reduction in myocardial infraction | [129] | |
| Genetic substances | miRNA | Glioblastoma cells | Glioblastoma tumor | Differential centrifugation | Transfection | Providing diagnostic information | [106] |
| miRNA | Human cord blood endothelial colony-forming cells | Ischemic kidney injury | Centrifugation | Transfection | Protected kidney function and reduced kidney injury | [130] | |
| Spherical nucleic acids | PC-3 cells | Prostate cancer | Centrifugation | Naturally encased | 3000-fold-enhanced knockdown of miR-21 | [131] | |
| siRNA | Human embryonic kidneycells (HEK293) | Breast cancer | Sequential centrifugation | Electroporation | TPD52 gene expression was downregulated up to 70% compared with non-targeted exosomes | [113] | |
| Small molecules | Paclitaxel | Prostate cancer cell lines (PC-3 and LNCaP) | Autologous prostate cancer | Differential centrifugation | Co-incubation | Enhanced drug cytotoxicity to cancer cells | [132] |
| Doxorubicin | Immature mouse dendritic cells transfected with the vector-expressing iRGD-Lamp2b fusion proteins | Breast cancer | Centrifugation and ultrafiltration | Electroporation | Specific drug delivery to the tumor site andinhibited tumor growth | [116] | |
| Curcumin | Tumor cells (GL26-Luc, BV2, 3T3L1, 4T1, CT26, A20, and EL-4) | Brain tumor and autoimmune encephalitis | Sucrose gradient centrifugation | Direct mixing | Inhibited brain inflammation and delayed brain tumor growth | [121] | |
| Dopamine | Kunming mouse blood | Parkinson’s disease | Ultracentrifugation | Co-incubation | Enhanced therapeutic effect due to brain specific drug delivery | [133] | |
4. Exosomes Drug Loading Techniques
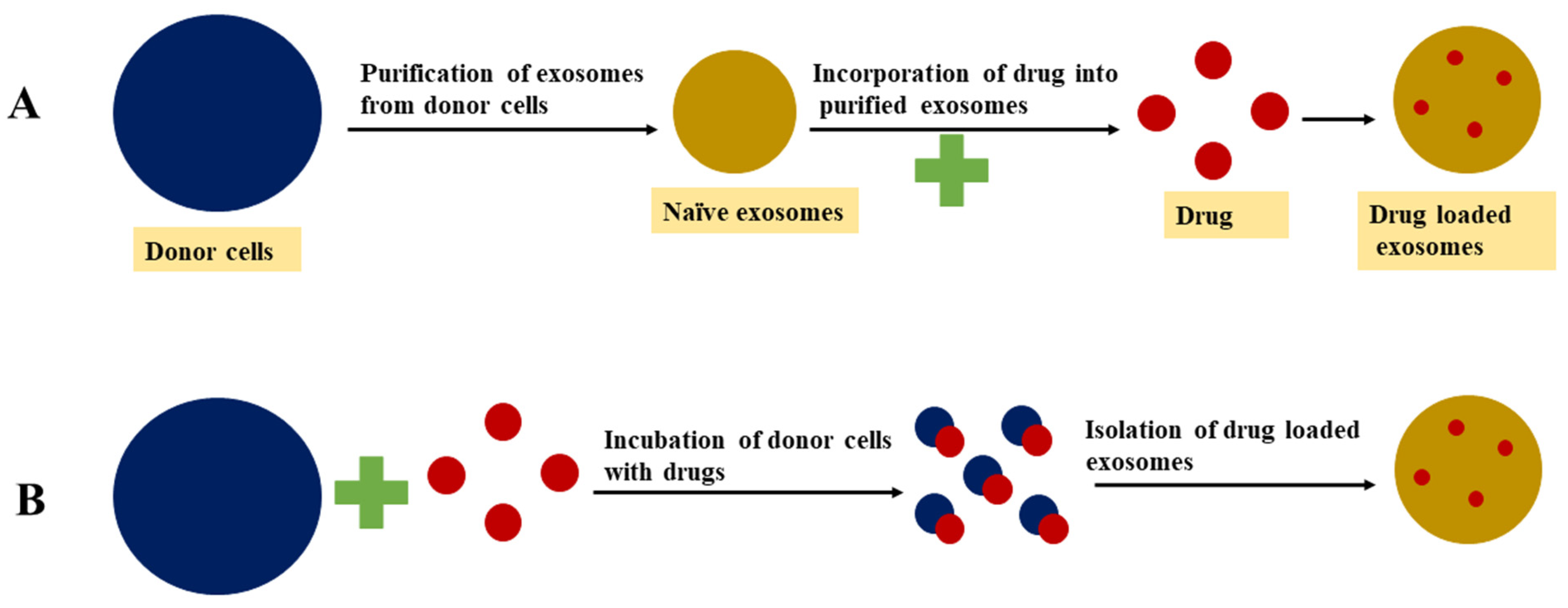
4.1. Passive Loading Approach
4.1.1. Incubation of Drugs with Exosomes
4.1.2. Incubation of Drugs with Donor Cells
4.2. Active Drug Loading Approaches
4.2.1. Sonication
4.2.2. Extrusion
4.2.3. Freeze–Thaw Cycles
4.2.4. Electroporation
4.2.5. Incubation with Membrane Permeabilizers
| Drug Loading Approach | Mechanism | Advantages | Disadvantages | |
|---|---|---|---|---|
| Passive loading | Incubation of exosomes and free drugs. | Diffusion of cargo into a cell or exosomal membrane. | Simple operation. Does not compromise the membrane integrity. | Loading efficiency. Drugs may cause cytotoxicity to the donor cells. |
| Incubation of the donor cells with free drugs. | ||||
| Active loading | Sonication | Creation of micropores for diffusion by mechanical shear force. | Higher loading capacity than the simple incubation method. | Sonication-induced membrane damage is a roadblock for large scale application. Influence on exosome integrity and cargo aggregation. |
| Extrusion | Membrane recombination. | High cargo loading efficiency. Repeated extrusion provides a homogeneous blend of exosomes with cargoes. | Recombination of exosomal surface structure may compromise the immune-privileged status of exosomes, making exosomes visible to immune cells such as mononuclear phagocytes. | |
| Freeze–thaw cycles | Membrane fusion. | Simple and effective strategy to load various cargoes (drugs, proteins, and peptides) into exosomes directly. | Repeated freeze–thaw may cause protein degeneration and exosome aggregation. Drug loading efficiency is lower than sonication and extrusion methods. | |
| Electroporation | Creation of micropores for diffusion by the electric field. | High loading efficiency | The loading efficiency and aggregation of cargoes are major limitations. | |
| Incubation with membrane permeabilizers | Dissolves membrane molecules (cholesterol), create pores on the exosomal surface. | Higher loading capacity as compared with the simple incubation method | Saponin is hemolytically active in vivo, limiting the concentration (toxicity) of saponin used for drug loading. Extra purification process may be required to remove saponin. | |
5. Exosomes Administration Routes
5.1. Intravenous Administration
5.2. Intratumoral Injection
5.3. Intraperitoneal Route
5.4. Oral Administration
5.5. Intranasal Administration
| Routes of Administration | Targeted Disease | Advantages |
|---|---|---|
| Intravenous | Stroke, Parkinson’s disease, traumatic brain injury, acute kidney injury, antitumor therapies (prostate and breast cancer). | Most common route for systemic administration of exosomes. |
| Intraperitoneal | Bronchopulmonary dysplasia Autoimmune type 1 diabetes | Allows the loading of larger EV doses. |
| Oral | Facilitates resolution of colitis, arthritis | Convenient administration route for patients. |
| Intranasal | Brain parenchyma Brain cancer Encephalitis (inflammation of the brain), Parkinson’s disease therapy | Suitable for EV delivery into the brain, surpassing the blood brain barrier. |
| Intratumoral | Glioblastoma multiforme antitumor therapies | More effective strategy for antitumor therapies due to higher EV retention in tumors. |
6. Characterization Techniques
6.1. Imaging
6.1.1. AFM
6.1.2. TEM
6.1.3. SEM
6.2. Dynamic Light Scattering (DLS)
6.3. Flow Cytometry
6.4. Nanoparticle Tracking Analysis (NTA)
6.5. Tunable Resistance Pulse Sensing (TRPS)
6.6. Protein Characterization
6.6.1. ELISA
6.6.2. Western Blotting
| Identification or Quantification Methods | Purpose | Advantages | Limitations |
|---|---|---|---|
| Dynamic light scattering | Exosomes size distribution. | The lower measurement limit is 10 nm, suitable for the determination of monodisperse systems. Sample preservation for downstream analysis and requires no sample preparation. | Difficult to distinguish contaminated proteins with exosomes, not suitable for measuring complex exosome samples with large size ranges. Inaccurate with polydispersed and heterogeneous samples. |
| Nanoparticle tracking analysis Technology | Measurement of size and concentration of exosomes. | Higher resolution than flow cytometer, exosomes can be observed in real time with faster detection speed. | Detection threshold and camera levels will affect the quantification of exosomes. |
| Atomic force microscopy | Detection of exosomal morphology. | Require small sample amount, no sample fixation or staining. | Sample dehydration on mica surfaces may lead to modifications of the size and morphology of exosomes. |
| SEM and TEM | Detection of exosomal morphology. | SEM can be used to directly observe the surface structure, whereas TEM can be used to observe the internal structure of exosomes and provide information about particle size distribution. | SEM resolution is lower than TEM, high requirements in terms of sample preparation make TEM not suitable for rapid measurement of a large number of samples. |
| Flow cytometry | Detection of biomarkers of exosomes. | Qualitative and quantitative characterization of exosomes. | Detection limit is 400 nm, identification of multiple vesicles as a single event is possible, the particle size of exosomes cannot be measured, detection of proteins or antibody aggregates limits its application. |
| ELISA | Exosome protein quantification. | Suitable for high-throughput analysis and rapid detection with high specificity, can be used to analyze the marker proteins quantitatively and qualitatively. | Time-consuming, possible detection of non exosomal marker proteins, complicated operation with less repeatability. |
| Western blot | Exosome marker protein quantification. | Easy to analyze exosomes from cell culture media, the classic method for qualitative and quantitative analysis of marker proteins. | The detection of exosomal marker proteins varies depending on the type of parental cell, meaning this technique not suitable for the detection of exosomal marker proteins in biological fluid. Provides non-specific information on exosome concentration and size or distribution. |
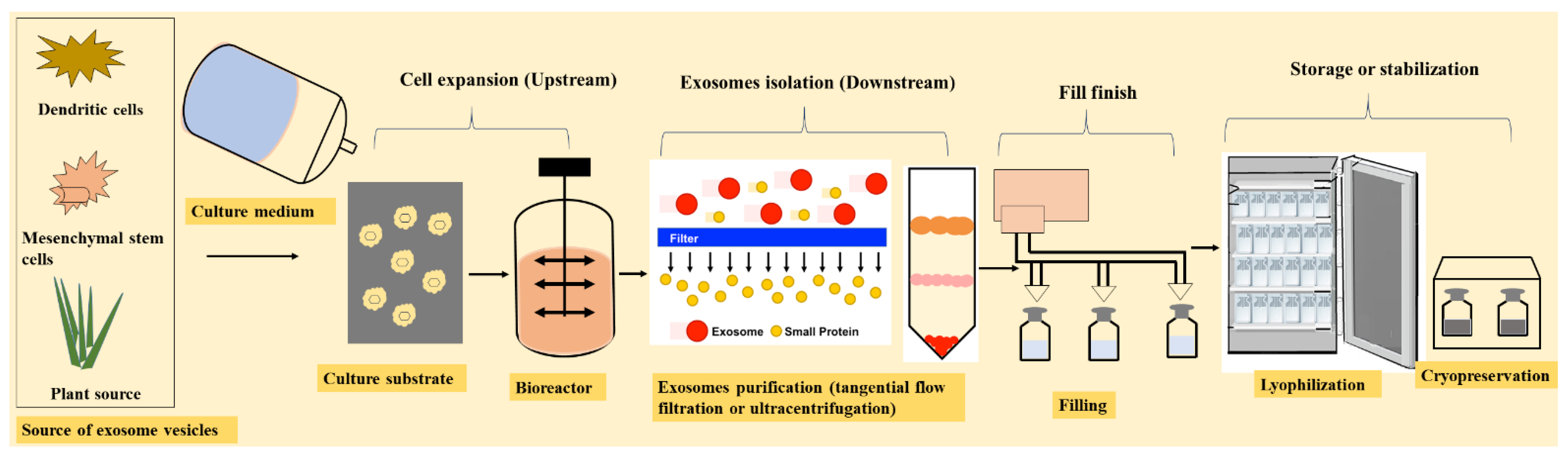
7. Manufacturing of Exosomes
7.1. Upstream Processing
7.2. Downstream Processing
7.3. Fill Finish
8. Stability and Stabilization Methods
9. Scalability Challenges
10. Exosomes in Clinical Trials
| Study Title | NCT Number | Conditions | Phase | Outcome Measures | Source of Exosomes |
|---|---|---|---|---|---|
| Allogenic mesenchymal stem cell derived exosome in patients with acute ischemic stroke. | NCT03384433 | Cerebrovascular disorders | Phase 1/2 | Incidence of treatment-emergent adverse events (deteriorating stroke, stroke recurrences, brain edema, seizures, hemorrhagic transformation). The degree of disability in stroke patients. | Mesenchymal stem cell |
| Evaluation of adipose derived stem cells exosomes in treatment of periodontitis. | NCT04270006 | Periodontitis | Early phase 1 | Changes in gingival inflammation, bone levels, probing depth. | Adipose-derived stem cells |
| Study investigating the ability of plant exosomes to deliver curcumin to normal and colon cancer tissue. | NCT01294072 | Colon cancer | Phase 1 | The concentrations of curcumin in normal and cancerous tissue, safety and tolerability of curcumin, and immune system response to curcumin. | Plants (fruit) |
| Effect of plasma derived exosomes on cutaneous wound healing. | NCT02565264 | Ulcers | Early phase 1 | Ulcer size, the pain of cutaneous wounds. | Plasma |
| Trial of a vaccination with tumor antigen-loaded dendritic cell-derived exosomes. | NCT01159288 | Non-small cell lung cancer | Phase 2 | Progression-free survival. | Dendritic cells |
| Edible plant exosome ability to prevent oral mucositis associated with chemoradiation treatment of head and neck cancer. | NCT01668849 | Head and neck cancer and oral mucositis | Phase 1 | Pain caused by oral mucositis, levels of immune biomarkers in blood and mucosal tissue. | Plants (grape) |
| Effect of microvesicles and exosomes therapy on β-cell mass in type I diabetes mellitus. | NCT02138331 | Diabetes mellitus type 1 | Phase 2 and phase 3 | Total daily insulin dose, pancreatic β-cell mass, and hemoglobin A1c. | Mesenchymal stem cells |
| Exosomes and Immunotherapy in Non-Hodgkin B-cell lymphomas. | NCT03985696 | Lymphoma, B-cell, aggressive non-Hodgkin | Not applicable | Quantification of CD20 and PDL-1 in exosomes purified from cell cultures of diffuse large B-cell lymphoma (DLBCL) human cell lines and evaluation of whether peripheral exosomes can be used as novel diagnostic biomarkers in DLBCL. | Tumor B cells |
| Exosomes in treating participants with metastatic pancreas cancer with KrasG12D mutation. | NCT03608631 | Metastatic pancreatic adenocarcinoma Pancreatic ductal adenocarcinoma Stage IV pancreatic cancer AJCC v8 | Phase 1 | Overall survival, progression-free survival, minimal residual disease rate in high-risk patients, and maximum tolerated dose determined by dose-limiting toxicity. | Mesenchymal stromal cells |
| A safety study of IV stem cell-derived extracellular Vesicles (UNEX-42) in preterm neonates at high risk for BPD. | NCT03857841 | Bronchopulmonary dysplasia (BPD) | Phase 1 | Safety and tolerability, incidence, and severity of BPD at 36 weeks postmenstrual age and incidence of death at 36 weeks postmenstrual age. | Bone marrow mesenchymal stem cells |
11. Regulatory Challenges
12. Future Prospects and Concluding Remarks
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Elsharkasy, O.M.; Nordin, J.Z.; Hagey, D.W.; de Jong, O.G.; Schiffelers, R.M.; Andaloussi, S.E.; Vader, P. Extracellular Vesicles as Drug Delivery Systems: Why and How? Adv. Drug Deliv. Rev. 2020, 159, 332–343. [Google Scholar] [CrossRef]
- Svenson, S. Clinical Translation of Nanomedicines. Curr. Opin. Solid State Mater. Sci. 2012, 16, 287–294. [Google Scholar] [CrossRef]
- Van der Meel, R.; Fens, M.H.A.M.; Vader, P.; van Solinge, W.W.; Eniola-Adefeso, O.; Schiffelers, R.M. Extracellular Vesicles as Drug Delivery Systems: Lessons from the Liposome Field. J. Control. Release 2014, 195, 72–85. [Google Scholar] [CrossRef]
- Bulbake, U.; Doppalapudi, S.; Kommineni, N.; Khan, W. Liposomal Formulations in Clinical Use: An Updated Review. Pharmaceutics 2017, 9, 12. [Google Scholar] [CrossRef]
- Balachandran, B.; Yuana, Y. Extracellular Vesicles-Based Drug Delivery System for Cancer Treatment. Cogent Med. 2019, 6, 1635806. [Google Scholar] [CrossRef]
- Sercombe, L.; Veerati, T.; Moheimani, F.; Wu, S.Y.; Sood, A.K.; Hua, S. Advances and Challenges of Liposome Assisted Drug Delivery. Front. Pharmacol. 2015, 6, 286. [Google Scholar] [CrossRef]
- Shao, J.; Zaro, J.; Shen, Y. Advances in Exosome-Based Drug Delivery and Tumor Targeting: From Tissue Distribution to Intracellular Fate. Int. J. Nanomed. 2020, 15, 9355–9371. [Google Scholar] [CrossRef] [PubMed]
- Kooijmans, S.A.A.; Vader, P.; van Dommelen, S.M.; van Solinge, W.W.; Schiffelers, R.M. Exosome Mimetics: A Novel Class of Drug Delivery Systems. Int. J. Nanomed. 2012, 7, 1525–1541. [Google Scholar] [CrossRef]
- Schiller, M.; Bekeredjian-Ding, I.; Heyder, P.; Blank, N.; Ho, A.D.; Lorenz, H.-M. Autoantigens Are Translocated into Small Apoptotic Bodies during Early Stages of Apoptosis. Cell Death Differ. 2008, 15, 183–191. [Google Scholar] [CrossRef]
- Gregory, C.D.; Pound, J.D. Microenvironmental Influences of Apoptosis In Vivo and In Vitro. Apoptosis 2010, 15, 1029–1049. [Google Scholar] [CrossRef]
- Wang, J.; Chen, D.; Ho, E.A. Challenges in the Development and Establishment of Exosome-Based Drug Delivery Systems. J. Control. Release 2021, 329, 894–906. [Google Scholar] [CrossRef]
- Ha, D.; Yang, N.; Nadithe, V. Exosomes as Therapeutic Drug Carriers and Delivery Vehicles across Biological Membranes: Current Perspectives and Future Challenges. Acta Pharm. Sin. B 2016, 6, 287–296. [Google Scholar] [CrossRef]
- Bunggulawa, E.J.; Wang, W.; Yin, T.; Wang, N.; Durkan, C.; Wang, Y.; Wang, G. Recent Advancements in the Use of Exosomes as Drug Delivery Systems. J. Nanobiotechnology 2018, 16, 81. [Google Scholar] [CrossRef]
- Gebeyehu, A.; Kommineni, N.; Meckes, D.; Sachdeva, M. Exosome Vehicles as Nano-Drug Delivery Materials for Chemotherapeutic Drugs. CRT 2021. [Google Scholar] [CrossRef]
- Mittelbrunn, M.; Sánchez-Madrid, F. Intercellular Communication: Diverse Structures for Exchange of Genetic Information. Nat. Rev. Mol. Cell Biol. 2012, 13, 328–335. [Google Scholar] [CrossRef] [PubMed]
- Heijnen, H.F.; Schiel, A.E.; Fijnheer, R.; Geuze, H.J.; Sixma, J.J. Activated Platelets Release Two Types of Membrane Vesicles: Microvesicles by Surface Shedding and Exosomes Derived from Exocytosis of Multivesicular Bodies and Alpha-Granules. Blood 1999, 94, 3791–3799. [Google Scholar] [CrossRef]
- Savina, A.; Furlán, M.; Vidal, M.; Colombo, M.I. Exosome Release Is Regulated by a Calcium-Dependent Mechanism in K562 Cells. J. Biol. Chem. 2003, 278, 20083–20090. [Google Scholar] [CrossRef]
- Hung, W.-T.; Hong, X.; Christenson, L.K.; McGinnis, L.K. Extracellular Vesicles from Bovine Follicular Fluid Support Cumulus Expansion. Biol. Reprod. 2015, 93, 117. [Google Scholar] [CrossRef]
- Machtinger, R.; Laurent, L.C.; Baccarelli, A.A. Extracellular Vesicles: Roles in Gamete Maturation, Fertilization and Embryo Implantation. Hum. Reprod. Update 2016, 22, 182–193. [Google Scholar] [CrossRef]
- Patton, A.L.; McCallie, B.; Parks, J.C.; Schoolcraft, W.B.; Katz-Jaffe, M. Exosome Bound MicroRNAs Transcriptionally Regulate Embryo-Endometrial Dialogue Impacting Implantation Potential for AMA Patients. Fertil. Steril. 2015, 104, e308. [Google Scholar] [CrossRef]
- Matsuno, Y.; Onuma, A.; Fujioka, Y.A.; Yasuhara, K.; Fujii, W.; Naito, K.; Sugiura, K. Effects of Exosome-like Vesicles on Cumulus Expansion in Pigs In Vitro. J. Reprod. Dev. 2017, 63, 51–58. [Google Scholar] [CrossRef]
- Qu, P.; Qing, S.; Liu, R.; Qin, H.; Wang, W.; Qiao, F.; Ge, H.; Liu, J.; Zhang, Y.; Cui, W.; et al. Effects of Embryo-Derived Exosomes on the Development of Bovine Cloned Embryos. PLoS ONE 2017, 12, e0174535. [Google Scholar] [CrossRef]
- Mobarak, H.; Heidarpour, M.; Lolicato, F.; Nouri, M.; Rahbarghazi, R.; Mahdipour, M. Physiological Impact of Extracellular Vesicles on Female Reproductive System; Highlights to Possible Restorative Effects on Female Age-Related Fertility. Biofactors 2019, 45, 293–303. [Google Scholar] [CrossRef]
- Tsochandaridis, M.; Nasca, L.; Toga, C.; Levy-Mozziconacci, A. Circulating MicroRNAs as Clinical Biomarkers in the Predictions of Pregnancy Complications. Biomed. Res. Int. 2015, 2015, 294954. [Google Scholar] [CrossRef]
- 25 Koh, Y.Q.; Peiris, H.N.; Vaswani, K.; Reed, S.; Rice, G.E.; Salomon, C.; Mitchell, M.D. Characterization of Exosomal Release in Bovine Endometrial Intercaruncular Stromal Cells. Reprod. Biol. Endocrinol. 2016, 14, 78. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, R.; Frenette, G.; Girouard, J. Epididymosomes Are Involved in the Acquisition of New Sperm Proteins during Epididymal Transit. Asian J. Androl. 2007, 9, 483–491. [Google Scholar] [CrossRef]
- Andrews, R.E.; Galileo, D.S.; Martin-DeLeon, P.A. Plasma Membrane Ca2+-ATPase 4: Interaction with Constitutive Nitric Oxide Synthases in Human Sperm and Prostasomes Which Carry Ca2+/CaM-Dependent Serine Kinase. Mol. Hum. Reprod. 2015, 21, 832–843. [Google Scholar] [CrossRef]
- Choy, K.H.K.; Chan, S.Y.; Lam, W.; Jin, J.; Zheng, T.; Yu, S.S.; Wang, W.; Li, L.; Xie, G.; Yim, H.C.H.; et al. The Repertoire of Testicular Extracellular Vesicles Cargoes and Their Involvement in Inter-Compartmental Communication Required for Spermatogenesis. bioRxiv 2021. [Google Scholar] [CrossRef]
- Baskaran, S.; Panner Selvam, M.K.; Agarwal, A. Exosomes of Male Reproduction. Adv. Clin. Chem. 2020, 95, 149–163. [Google Scholar] [CrossRef]
- Zhou, Y.; Tian, T.; Zhu, Y.; Jaffar Ali, D.; Hu, F.; Qi, Y.; Sun, B.; Xiao, Z. Exosomes Transfer Among Different Species Cells and Mediating MiRNAs Delivery. J. Cell. Biochem. 2017, 118, 4267–4274. [Google Scholar] [CrossRef]
- Baj-Krzyworzeka, M.; Majka, M.; Pratico, D.; Ratajczak, J.; Vilaire, G.; Kijowski, J.; Reca, R.; Janowska-Wieczorek, A.; Ratajczak, M.Z. Platelet-Derived Microparticles Stimulate Proliferation, Survival, Adhesion, and Chemotaxis of Hematopoietic Cells. Exp. Hematol. 2002, 30, 450–459. [Google Scholar] [CrossRef]
- Al-Nedawi, K.; Meehan, B.; Kerbel, R.S.; Allison, A.C.; Rak, J. Endothelial Expression of Autocrine VEGF upon the Uptake of Tumor-Derived Microvesicles Containing Oncogenic EGFR. Proc. Natl. Acad. Sci. USA 2009, 106, 3794–3799. [Google Scholar] [CrossRef]
- Tian, T.; Wang, Y.; Wang, H.; Zhu, Z.; Xiao, Z. Visualizing of the Cellular Uptake and Intracellular Trafficking of Exosomes by Live-Cell Microscopy. J. Cell. Biochem. 2010, 111, 488–496. [Google Scholar] [CrossRef]
- Mayer, A. Intracellular Membrane Fusion: SNAREs Only? Curr. Opin. Cell Biol. 1999, 11, 447–452. [Google Scholar] [CrossRef]
- Simpson, R.J.; Jensen, S.S.; Lim, J.W.E. Proteomic Profiling of Exosomes: Current Perspectives. Proteomics 2008, 8, 4083–4099. [Google Scholar] [CrossRef]
- Valadi, H.; Ekström, K.; Bossios, A.; Sjöstrand, M.; Lee, J.J.; Lötvall, J.O. Exosome-Mediated Transfer of MRNAs and MicroRNAs Is a Novel Mechanism of Genetic Exchange between Cells. Nat. Cell Biol. 2007, 9, 654–659. [Google Scholar] [CrossRef]
- Yeo, R.W.Y.; Lai, R.C.; Zhang, B.; Tan, S.S.; Yin, Y.; Teh, B.J.; Lim, S.K. Mesenchymal Stem Cell: An Efficient Mass Producer of Exosomes for Drug Delivery. Adv. Drug Deliv. Rev. 2013, 65, 336–341. [Google Scholar] [CrossRef] [PubMed]
- Cabeza, L.; Perazzoli, G.; Peña, M.; Cepero, A.; Luque, C.; Melguizo, C.; Prados, J. Cancer Therapy Based on Extracellular Vesicles as Drug Delivery Vehicles. J. Control. Release 2020, 327, 296–315. [Google Scholar] [CrossRef]
- Armstrong, J.P.K.; Stevens, M.M. Strategic Design of Extracellular Vesicle Drug Delivery Systems. Adv. Drug Deliv. Rev. 2018, 130, 12–16. [Google Scholar] [CrossRef]
- Kwon, S.; Shin, S.; Do, M.; Oh, B.H.; Song, Y.; Bui, V.D.; Lee, E.S.; Jo, D.-G.; Cho, Y.W.; Kim, D.-H.; et al. Engineering Approaches for Effective Therapeutic Applications Based on Extracellular Vesicles. J. Control. Release 2021, 330, 15–30. [Google Scholar] [CrossRef]
- Liao, W.; Du, Y.; Zhang, C.; Pan, F.; Yao, Y.; Zhang, T.; Peng, Q. Exosomes: The next Generation of Endogenous Nanomaterials for Advanced Drug Delivery and Therapy. Acta Biomater. 2019, 86, 1–14. [Google Scholar] [CrossRef]
- Théry, C.; Amigorena, S.; Raposo, G.; Clayton, A. Isolation and Characterization of Exosomes from Cell Culture Supernatants and Biological Fluids. Curr. Protoc. Cell Biol. 2006, 30, 3.22.1–3.22.29. [Google Scholar] [CrossRef]
- Greening, D.W.; Xu, R.; Ji, H.; Tauro, B.J.; Simpson, R.J. A Protocol for Exosome Isolation and Characterization: Evaluation of Ultracentrifugation, Density-Gradient Separation, and Immunoaffinity Capture Methods. Methods Mol. Biol. 2015, 1295, 179–209. [Google Scholar] [CrossRef]
- Lobb, R.J.; Becker, M.; Wen, S.W.; Wong, C.S.F.; Wiegmans, A.P.; Leimgruber, A.; Möller, A. Optimized Exosome Isolation Protocol for Cell Culture Supernatant and Human Plasma. J. Extracell. Vesicles 2015, 4, 27031. [Google Scholar] [CrossRef]
- Li, K.; Wong, D.K.; Hong, K.Y.; Raffai, R.L. Cushioned-Density Gradient Ultracentrifugation (C-DGUC): A Refined and High Performance Method for the Isolation, Characterization, and Use of Exosomes. Methods Mol. Biol. 2018, 1740, 69–83. [Google Scholar] [CrossRef]
- Taylor, D.D.; Shah, S. Methods of Isolating Extracellular Vesicles Impact Down-Stream Analyses of Their Cargoes. Methods 2015, 87, 3–10. [Google Scholar] [CrossRef]
- Sódar, B.W.; Kittel, Á.; Pálóczi, K.; Vukman, K.V.; Osteikoetxea, X.; Szabó-Taylor, K.; Németh, A.; Sperlágh, B.; Baranyai, T.; Giricz, Z.; et al. Low-Density Lipoprotein Mimics Blood Plasma-Derived Exosomes and Microvesicles during Isolation and Detection. Sci. Rep. 2016, 6, 24316. [Google Scholar] [CrossRef]
- Onódi, Z.; Pelyhe, C.; Terézia Nagy, C.; Brenner, G.B.; Almási, L.; Kittel, Á.; Manček-Keber, M.; Ferdinandy, P.; Buzás, E.I.; Giricz, Z. Isolation of High-Purity Extracellular Vesicles by the Combination of Iodixanol Density Gradient Ultracentrifugation and Bind-Elute Chromatography from Blood Plasma. Front. Physiol. 2018, 9, 1479. [Google Scholar] [CrossRef]
- Heinemann, M.L.; Ilmer, M.; Silva, L.P.; Hawke, D.H.; Recio, A.; Vorontsova, M.A.; Alt, E.; Vykoukal, J. Benchtop Isolation and Characterization of Functional Exosomes by Sequential Filtration. J. Chromatogr. A 2014, 1371, 125–135. [Google Scholar] [CrossRef]
- Busatto, S.; Vilanilam, G.; Ticer, T.; Lin, W.-L.; Dickson, D.W.; Shapiro, S.; Bergese, P.; Wolfram, J. Tangential Flow Filtration for Highly Efficient Concentration of Extracellular Vesicles from Large Volumes of Fluid. Cells 2018, 7, 273. [Google Scholar] [CrossRef]
- Yang, D.; Zhang, W.; Zhang, H.; Zhang, F.; Chen, L.; Ma, L.; Larcher, L.M.; Chen, S.; Liu, N.; Zhao, Q.; et al. Progress, Opportunity, and Perspective on Exosome Isolation—Efforts for Efficient Exosome-Based Theranostics. Theranostics 2020, 10, 3684–3707. [Google Scholar] [CrossRef]
- Wang, Y.; Qin, X.; Zhu, X.; Chen, W.; Zhang, J.; Chen, W. Oral Cancer-Derived Exosomal NAP1 Enhances Cytotoxicity of Natural Killer Cells via the IRF-3 Pathway. Oral Oncol. 2018, 76, 34–41. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Huang, W.; Wani, M.; Yu, X.; Ashraf, M. Ischemic Preconditioning Potentiates the Protective Effect of Stem Cells through Secretion of Exosomes by Targeting Mecp2 via MiR-22. PLoS ONE 2014, 9, e88685. [Google Scholar] [CrossRef]
- Yang, X.-X.; Sun, C.; Wang, L.; Guo, X.-L. New Insight into Isolation, Identification Techniques and Medical Applications of Exosomes. J. Control. Release 2019, 308, 119–129. [Google Scholar] [CrossRef] [PubMed]
- Zeringer, E.; Barta, T.; Li, M.; Vlassov, A.V. Strategies for Isolation of Exosomes. Cold Spring Harb Protoc. 2015, 2015, 319–323. [Google Scholar] [CrossRef] [PubMed]
- Kanchi Ravi, R.; Khosroheidari, M.; DiStefano, J.K. A Modified Precipitation Method to Isolate Urinary Exosomes. J. Vis. Exp. 2015, 51158. [Google Scholar] [CrossRef]
- Deregibus, M.C.; Figliolini, F.; D’Antico, S.; Manzini, P.M.; Pasquino, C.; De Lena, M.; Tetta, C.; Brizzi, M.F.; Camussi, G. Charge-Based Precipitation of Extracellular Vesicles. Int. J. Mol. Med. 2016, 38, 1359–1366. [Google Scholar] [CrossRef]
- Clayton, A.; Court, J.; Navabi, H.; Adams, M.; Mason, M.D.; Hobot, J.A.; Newman, G.R.; Jasani, B. Analysis of Antigen Presenting Cell Derived Exosomes, Based on Immuno-Magnetic Isolation and Flow Cytometry. J. Immunol. Methods 2001, 247, 163–174. [Google Scholar] [CrossRef]
- Théry, C.; Ostrowski, M.; Segura, E. Membrane Vesicles as Conveyors of Immune Responses. Nat. Rev. Immunol. 2009, 9, 581–593. [Google Scholar] [CrossRef]
- Yáñez-Mó, M.; Barreiro, O.; Gordon-Alonso, M.; Sala-Valdés, M.; Sánchez-Madrid, F. Tetraspanin-Enriched Microdomains: A Functional Unit in Cell Plasma Membranes. Trends Cell Biol. 2009, 19, 434–446. [Google Scholar] [CrossRef]
- Kim, D.; Nishida, H.; An, S.Y.; Shetty, A.K.; Bartosh, T.J.; Prockop, D.J. Chromatographically Isolated CD63+CD81+ Extracellular Vesicles from Mesenchymal Stromal Cells Rescue Cognitive Impairments after TBI. Proc. Natl. Acad. Sci. USA 2016, 113, 170–175. [Google Scholar] [CrossRef]
- Wubbolts, R.; Leckie, R.S.; Veenhuizen, P.T.M.; Schwarzmann, G.; Möbius, W.; Hoernschemeyer, J.; Slot, J.-W.; Geuze, H.J.; Stoorvogel, W. Proteomic and Biochemical Analyses of Human B Cell-Derived Exosomes. Potential Implications for Their Function and Multivesicular Body Formation. J. Biol. Chem. 2003, 278, 10963–10972. [Google Scholar] [CrossRef]
- Chen, C.; Skog, J.; Hsu, C.-H.; Lessard, R.T.; Balaj, L.; Wurdinger, T.; Carter, B.S.; Breakefield, X.O.; Toner, M.; Irimia, D. Microfluidic Isolation and Transcriptome Analysis of Serum Microvesicles. Lab Chip 2010, 10, 505–511. [Google Scholar] [CrossRef]
- Davies, R.T.; Kim, J.; Jang, S.C.; Choi, E.-J.; Gho, Y.S.; Park, J. Microfluidic Filtration System to Isolate Extracellular Vesicles from Blood. Lab Chip 2012, 12, 5202–5210. [Google Scholar] [CrossRef] [PubMed]
- Patil, S.M.; Sawant, S.S.; Kunda, N.K. Exosomes as Drug Delivery Systems: A Brief Overview and Progress Update. Eur. J. Pharm. Biopharm. 2020, 154, 259–269. [Google Scholar] [CrossRef]
- Momen-Heravi, F.; Balaj, L.; Alian, S.; Mantel, P.-Y.; Halleck, A.E.; Trachtenberg, A.J.; Soria, C.E.; Oquin, S.; Bonebreak, C.M.; Saracoglu, E.; et al. Current Methods for the Isolation of Extracellular Vesicles. Biol. Chem. 2013, 394, 1253–1262. [Google Scholar] [CrossRef]
- Li, Y.; Zheng, Q.; Bao, C.; Li, S.; Guo, W.; Zhao, J.; Chen, D.; Gu, J.; He, X.; Huang, S. Circular RNA Is Enriched and Stable in Exosomes: A Promising Biomarker for Cancer Diagnosis. Cell Res. 2015, 25, 981–984. [Google Scholar] [CrossRef]
- Tauro, B.J.; Greening, D.W.; Mathias, R.A.; Ji, H.; Mathivanan, S.; Scott, A.M.; Simpson, R.J. Comparison of Ultracentrifugation, Density Gradient Separation, and Immunoaffinity Capture Methods for Isolating Human Colon Cancer Cell Line LIM1863-Derived Exosomes. Methods 2012, 56, 293–304. [Google Scholar] [CrossRef]
- Boriachek, K.; Islam, M.N.; Möller, A.; Salomon, C.; Nguyen, N.T.; Hossain, M.S.A.; Yamauchi, Y.; Shiddiky, M.J. Biological functions and current advances in isolation and detection strategies for exosome nanovesicles. Small 2018, 14, 1702153. [Google Scholar] [CrossRef]
- Li, P.; Kaslan, M.; Lee, S.H.; Yao, J.; Gao, Z. Progress in Exosome Isolation Techniques. Theranostics 2017, 7, 789–804. [Google Scholar] [CrossRef]
- Yakimchuk, K. Exosomes: Isolation and Characterization Methods and Specific Markers. Mater. Methods 2021. [Google Scholar] [CrossRef]
- Muller, L.; Hong, C.-S.; Stolz, D.B.; Watkins, S.C.; Whiteside, T.L. Isolation of Biologically-Active Exosomes from Human Plasma. J. Immunol. Methods 2014, 411, 55–65. [Google Scholar] [CrossRef]
- Batrakova, E.V.; Kim, M.S. Using Exosomes, Naturally-Equipped Nanocarriers, for Drug Delivery. J. Control. Release 2015, 219, 396–405. [Google Scholar] [CrossRef]
- Yamashita, T.; Takahashi, Y.; Nishikawa, M.; Takakura, Y. Effect of Exosome Isolation Methods on Physicochemical Properties of Exosomes and Clearance of Exosomes from the Blood Circulation. Eur. J. Pharm. Biopharm. 2016, 98, 1–8. [Google Scholar] [CrossRef]
- Chen, B.-Y.; Sung, C.W.-H.; Chen, C.; Cheng, C.-M.; Lin, D.P.-C.; Huang, C.-T.; Hsu, M.-Y. Advances in Exosomes Technology. Clin. Chim. Acta 2019, 493, 14–19. [Google Scholar] [CrossRef]
- Soares Martins, T.; Catita, J.; Martins Rosa, I.; A B da Cruz E Silva, O.; Henriques, A.G. Exosome Isolation from Distinct Biofluids Using Precipitation and Column-Based Approaches. PLoS ONE 2018, 13, e0198820. [Google Scholar] [CrossRef]
- Panagiotara, A.; Markou, A.; Lianidou, E.S.; Patrinos, G.P.; Katsila, T. Exosomes: A Cancer Theranostics Road Map. Public Health Genom. 2017, 20, 116–125. [Google Scholar] [CrossRef] [PubMed]
- Milman, N.; Ginini, L.; Gil, Z. Exosomes and Their Role in Tumorigenesis and Anticancer Drug Resistance. Drug Resist. Updates 2019, 45, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Tan, A.; Rajadas, J.; Seifalian, A.M. Exosomes as Nano-Theranostic Delivery Platforms for Gene Therapy. Adv. Drug Deliv. Rev. 2013, 65, 357–367. [Google Scholar] [CrossRef]
- Graner, M.W.; Alzate, O.; Dechkovskaia, A.M.; Keene, J.D.; Sampson, J.H.; Mitchell, D.A.; Bigner, D.D. Proteomic and Immunologic Analyses of Brain Tumor Exosomes. FASEB J. 2009, 23, 1541–1557. [Google Scholar] [CrossRef]
- Van Dommelen, S.M.; Vader, P.; Lakhal, S.; Kooijmans, S.A.A.; van Solinge, W.W.; Wood, M.J.A.; Schiffelers, R.M. Microvesicles and Exosomes: Opportunities for Cell-Derived Membrane Vesicles in Drug Delivery. J. Control. Release 2012, 161, 635–644. [Google Scholar] [CrossRef] [PubMed]
- Cho, E.; Nam, G.-H.; Hong, Y.; Kim, Y.K.; Kim, D.-H.; Yang, Y.; Kim, I.-S. Comparison of Exosomes and Ferritin Protein Nanocages for the Delivery of Membrane Protein Therapeutics. J. Control. Release 2018, 279, 326–335. [Google Scholar] [CrossRef] [PubMed]
- Théry, C.; Zitvogel, L.; Amigorena, S. Exosomes: Composition, Biogenesis and Function. Nat. Rev. Immunol. 2002, 2, 569–579. [Google Scholar] [CrossRef]
- Khan, S.; Bennit, H.F.; Wall, N.R. The Emerging Role of Exosomes in Survivin Secretion. Histol. Histopathol. 2015, 30, 43–50. [Google Scholar] [CrossRef]
- Kooijmans, S.A.A.; Aleza, C.G.; Roffler, S.R.; van Solinge, W.W.; Vader, P.; Schiffelers, R.M. Display of GPI-Anchored Anti-EGFR Nanobodies on Extracellular Vesicles Promotes Tumour Cell Targeting. J. Extracell. Vesicles 2016, 5, 31053. [Google Scholar] [CrossRef]
- Peng, H.; Ji, W.; Zhao, R.; Yang, J.; Lu, Z.; Li, Y.; Zhang, X. Exosome: A Significant Nano-Scale Drug Delivery Carrier. J. Mater. Chem. B 2020, 8, 7591–7608. [Google Scholar] [CrossRef] [PubMed]
- Gehrmann, U.; Hiltbrunner, S.; Georgoudaki, A.-M.; Karlsson, M.C.; Näslund, T.I.; Gabrielsson, S. Synergistic Induction of Adaptive Antitumor Immunity by Codelivery of Antigen with α-Galactosylceramide on Exosomes. Cancer Res. 2013, 73, 3865–3876. [Google Scholar] [CrossRef]
- Tian, T.; Zhang, H.-X.; He, C.-P.; Fan, S.; Zhu, Y.-L.; Qi, C.; Huang, N.-P.; Xiao, Z.-D.; Lu, Z.-H.; Tannous, B.A.; et al. Surface Functionalized Exosomes as Targeted Drug Delivery Vehicles for Cerebral Ischemia Therapy. Biomaterials 2018, 150, 137–149. [Google Scholar] [CrossRef]
- De Toro, J.; Herschlik, L.; Waldner, C.; Mongini, C. Emerging Roles of Exosomes in Normal and Pathological Conditions: New Insights for Diagnosis and Therapeutic Applications. Front. Immunol. 2015, 6, 203. [Google Scholar] [CrossRef]
- Anand, P.K. Exosomal Membrane Molecules Are Potent Immune Response Modulators. Commun. Integr. Biol. 2010, 3, 405–408. [Google Scholar] [CrossRef]
- Gehrmann, U.; Näslund, T.I.; Hiltbrunner, S.; Larssen, P.; Gabrielsson, S. Harnessing the Exosome-Induced Immune Response for Cancer Immunotherapy. Semin. Cancer Biol. 2014, 28, 58–67. [Google Scholar] [CrossRef]
- Jesus, S.; Soares, E.; Cruz, M.T.; Borges, O. Exosomes as Adjuvants for the Recombinant Hepatitis B Antigen: First Report. Eur. J. Pharm. Biopharm. 2018, 133, 1–11. [Google Scholar] [CrossRef]
- Liu, Y.; Fu, N.; Su, J.; Wang, X.; Li, X. Rapid Enkephalin Delivery Using Exosomes to Promote Neurons Recovery in Ischemic Stroke by Inhibiting Neuronal P53/Caspase-3. BioMed Res. Int. 2019, 2019, e4273290. [Google Scholar] [CrossRef]
- Barok, M.; Puhka, M.; Vereb, G.; Szollosi, J.; Isola, J.; Joensuu, H. Cancer-Derived Exosomes from HER2-Positive Cancer Cells Carry Trastuzumab-Emtansine into Cancer Cells Leading to Growth Inhibition and Caspase Activation. BMC Cancer 2018, 18, 504. [Google Scholar] [CrossRef]
- Parolini, I.; Federici, C.; Raggi, C.; Lugini, L.; Palleschi, S.; De Milito, A.; Coscia, C.; Iessi, E.; Logozzi, M.; Molinari, A.; et al. Microenvironmental PH Is a Key Factor for Exosome Traffic in Tumor Cells. J. Biol. Chem. 2009, 284, 34211–34222. [Google Scholar] [CrossRef]
- Kim, H.; Yun, N.; Mun, D.; Kang, J.-Y.; Lee, S.-H.; Park, H.; Park, H.; Joung, B. Cardiac-Specific Delivery by Cardiac Tissue-Targeting Peptide-Expressing Exosomes. Biochem. Biophys. Res. Commun. 2018, 499, 803–808. [Google Scholar] [CrossRef]
- Haney, M.J.; Klyachko, N.L.; Zhao, Y.; Gupta, R.; Plotnikova, E.G.; He, Z.; Patel, T.; Piroyan, A.; Sokolsky, M.; Kabanov, A.V.; et al. Exosomes as Drug Delivery Vehicles for Parkinson’s Disease Therapy. J. Control. Release 2015, 207, 18–30. [Google Scholar] [CrossRef]
- Kole, R.; Krainer, A.R.; Altman, S. RNA Therapeutics: Beyond RNA Interference and Antisense Oligonucleotides. Nat. Rev. Drug Discov. 2012, 11, 125–140. [Google Scholar] [CrossRef] [PubMed]
- Guo, P. The Emerging Field of RNA Nanotechnology. Nat. Nanotechnol. 2010, 5, 833–842. [Google Scholar] [CrossRef]
- Bramsen, J.B.; Kjems, J. Development of Therapeutic-Grade Small Interfering RNAs by Chemical Engineering. Front. Genet. 2012, 3, 154. [Google Scholar] [CrossRef] [PubMed]
- Seow, Y.; Wood, M.J. Biological Gene Delivery Vehicles: Beyond Viral Vectors. Mol. Ther. 2009, 17, 767–777. [Google Scholar] [CrossRef]
- Foged, C. SiRNA Delivery with Lipid-Based Systems: Promises and Pitfalls. Curr. Top. Med. Chem. 2012, 12, 97–107. [Google Scholar] [CrossRef] [PubMed]
- El Andaloussi, S.; Mäger, I.; Breakefield, X.O.; Wood, M.J.A. Extracellular Vesicles: Biology and Emerging Therapeutic Opportunities. Nat. Rev. Drug Discov. 2013, 12, 347–357. [Google Scholar] [CrossRef]
- Wahlgren, J.; Karlson, T.D.L.; Brisslert, M.; Vaziri Sani, F.; Telemo, E.; Sunnerhagen, P.; Valadi, H. Plasma Exosomes Can Deliver Exogenous Short Interfering RNA to Monocytes and Lymphocytes. Nucleic Acids Res. 2012, 40, e130. [Google Scholar] [CrossRef] [PubMed]
- Shtam, T.A.; Kovalev, R.A.; Varfolomeeva, E.Y.; Makarov, E.M.; Kil, Y.V.; Filatov, M.V. Exosomes Are Natural Carriers of Exogenous SiRNA to Human Cells In Vitro. Cell Commun. Signal. 2013, 11, 88. [Google Scholar] [CrossRef] [PubMed]
- Skog, J.; Würdinger, T.; van Rijn, S.; Meijer, D.H.; Gainche, L.; Sena-Esteves, M.; Curry, W.T.; Carter, B.S.; Krichevsky, A.M.; Breakefield, X.O. Glioblastoma Microvesicles Transport RNA and Proteins that Promote Tumour Growth and Provide Diagnostic Biomarkers. Nat. Cell Biol. 2008, 10, 1470–1476. [Google Scholar] [CrossRef] [PubMed]
- Alvarez-Erviti, L.; Seow, Y.; Yin, H.; Betts, C.; Lakhal, S.; Wood, M.J.A. Delivery of SiRNA to the Mouse Brain by Systemic Injection of Targeted Exosomes. Nat. Biotechnol. 2011, 29, 341–345. [Google Scholar] [CrossRef] [PubMed]
- Banizs, A.B.; Huang, T.; Dryden, K.; Berr, S.S.; Stone, J.R.; Nakamoto, R.K.; Shi, W.; He, J. In Vitro Evaluation of Endothelial Exosomes as Carriers for Small Interfering Ribonucleic Acid Delivery. Int. J. Nanomed. 2014, 9, 4223–4230. [Google Scholar] [CrossRef]
- Zheng, M.; Huang, M.; Ma, X.; Chen, H.; Gao, X. Harnessing Exosomes for the Development of Brain Drug Delivery Systems. Bioconjugate Chem. 2019, 30, 994–1005. [Google Scholar] [CrossRef]
- Saeedi, S.; Israel, S.; Nagy, C.; Turecki, G. The Emerging Role of Exosomes in Mental Disorders. Transl. Psychiatry 2019, 9, 1–11. [Google Scholar] [CrossRef]
- Lamichhane, T.N.; Raiker, R.S.; Jay, S.M. Exogenous DNA Loading into Extracellular Vesicles via Electroporation Is Size-Dependent and Enables Limited Gene Delivery. Mol. Pharm. 2015, 12, 3650–3657. [Google Scholar] [CrossRef]
- Faruqu, F.N.; Xu, L.; Al-Jamal, K.T. Preparation of Exosomes for SiRNA Delivery to Cancer Cells. J. Vis. Exp. 2018. [Google Scholar] [CrossRef]
- Limoni, S.K.; Moghadam, M.F.; Moazzeni, S.M.; Gomari, H.; Salimi, F. Engineered Exosomes for Targeted Transfer of SiRNA to HER2 Positive Breast Cancer Cells. Appl. Biochem. Biotechnol. 2019, 187, 352–364. [Google Scholar] [CrossRef]
- Zhang, D.; Lee, H.; Wang, X.; Rai, A.; Groot, M.; Jin, Y. Exosome-Mediated Small RNA Delivery: A Novel Therapeutic Approach for Inflammatory Lung Responses. Mol. Ther. 2018, 26, 2119–2130. [Google Scholar] [CrossRef]
- Melzer, C.; Rehn, V.; Yang, Y.; Bähre, H.; von der Ohe, J.; Hass, R. Taxol-Loaded MSC-Derived Exosomes Provide a Therapeutic Vehicle to Target Metastatic Breast Cancer and Other Carcinoma Cells. Cancers 2019, 11, 798. [Google Scholar] [CrossRef]
- Tian, Y.; Li, S.; Song, J.; Ji, T.; Zhu, M.; Anderson, G.J.; Wei, J.; Nie, G. A Doxorubicin Delivery Platform Using Engineered Natural Membrane Vesicle Exosomes for Targeted Tumor Therapy. Biomaterials 2014, 35, 2383–2390. [Google Scholar] [CrossRef]
- Schindler, C.; Collinson, A.; Matthews, C.; Pointon, A.; Jenkinson, L.; Minter, R.R.; Vaughan, T.J.; Tigue, N.J. Exosomal Delivery of Doxorubicin Enables Rapid Cell Entry and Enhanced In Vitro Potency. PLoS ONE 2019, 14, e0214545. [Google Scholar] [CrossRef]
- Mulcahy, L.A.; Pink, R.C.; Carter, D.R.F. Routes and Mechanisms of Extracellular Vesicle Uptake. J. Extracell. Vesicles 2014, 3. [Google Scholar] [CrossRef]
- Finn, O.J. Immuno-Oncology: Understanding the Function and Dysfunction of the Immune System in Cancer. Ann. Oncol. 2012, 23, viii6-9. [Google Scholar] [CrossRef]
- Kim, M.S.; Haney, M.J.; Zhao, Y.; Mahajan, V.; Deygen, I.; Klyachko, N.L.; Inskoe, E.; Piroyan, A.; Sokolsky, M.; Okolie, O.; et al. Development of Exosome-Encapsulated Paclitaxel to Overcome MDR in Cancer Cells. Nanomedicine 2016, 12, 655–664. [Google Scholar] [CrossRef]
- Sun, D.; Zhuang, X.; Xiang, X.; Liu, Y.; Zhang, S.; Liu, C.; Barnes, S.; Grizzle, W.; Miller, D.; Zhang, H.-G. A Novel Nanoparticle Drug Delivery System: The Anti-Inflammatory Activity of Curcumin Is Enhanced When Encapsulated in Exosomes. Mol. Ther. 2010, 18, 1606–1614. [Google Scholar] [CrossRef]
- Zhuang, X.; Xiang, X.; Grizzle, W.; Sun, D.; Zhang, S.; Axtell, R.C.; Ju, S.; Mu, J.; Zhang, L.; Steinman, L.; et al. Treatment of Brain Inflammatory Diseases by Delivering Exosome Encapsulated Anti-Inflammatory Drugs from the Nasal Region to the Brain. Mol. Ther. 2011, 19, 1769–1779. [Google Scholar] [CrossRef]
- Canfrán-Duque, A.; Pastor, O.; Quintana-Portillo, R.; Lerma, M.; de la Peña, G.; Martín-Hidalgo, A.; Fernández-Hernando, C.; Lasunción, M.A.; Busto, R. Curcumin Promotes Exosomes/Microvesicles Secretion That Attenuates Lysosomal Cholesterol Traffic Impairment. Mol. Nutr. Food Res. 2014, 58, 687–697. [Google Scholar] [CrossRef]
- Kalani, A.; Kamat, P.K.; Chaturvedi, P.; Tyagi, S.C.; Tyagi, N. Curcumin-Primed Exosomes Mitigate Endothelial Cell Dysfunction during Hyperhomocysteinemia. Life Sci. 2014, 107, 1–7. [Google Scholar] [CrossRef]
- Kim, S.M.; Yang, Y.; Oh, S.J.; Hong, Y.; Seo, M.; Jang, M. Cancer-Derived Exosomes as a Delivery Platform of CRISPR/Cas9 Confer Cancer Cell Tropism-Dependent Targeting. J. Control. Release 2017, 266, 8–16. [Google Scholar] [CrossRef]
- Hou, S.; Niu, X.; Li, L.; Zhou, J.; Qian, Z.; Yao, D.; Yang, F.; Ma, P.X.; Fan, Y. Simultaneous Nano- and Microscale Structural Control of Injectable Hydrogels via the Assembly of Nanofibrous Protein Microparticles for Tissue Regeneration. Biomaterials 2019, 223, 119458. [Google Scholar] [CrossRef]
- Pardridge, W.M. Drug Transport across the Blood-Brain Barrier. J. Cereb. Blood Flow Metab. 2012, 32, 1959–1972. [Google Scholar] [CrossRef] [PubMed]
- Aspe, J.R.; Diaz Osterman, C.J.; Jutzy, J.M.S.; Deshields, S.; Whang, S.; Wall, N.R. Enhancement of Gemcitabine Sensitivity in Pancreatic Adenocarcinoma by Novel Exosome-Mediated Delivery of the Survivin-T34A Mutant. J. Extracell. Vesicles 2014, 3. [Google Scholar] [CrossRef] [PubMed]
- Lai, R.C.; Tan, S.S.; Teh, B.J.; Sze, S.K.; Arslan, F.; de Kleijn, D.P.; Choo, A.; Lim, S.K. Proteolytic Potential of the MSC Exosome Proteome: Implications for an Exosome-Mediated Delivery of Therapeutic Proteasome. Int. J. Proteomics 2012, 2012, 971907. [Google Scholar] [CrossRef] [PubMed]
- Viñas, J.L.; Burger, D.; Zimpelmann, J.; Haneef, R.; Knoll, W.; Campbell, P.; Gutsol, A.; Carter, A.; Allan, D.S.; Burns, K.D. Transfer of MicroRNA-486-5p from Human Endothelial Colony Forming Cell-Derived Exosomes Reduces Ischemic Kidney Injury. Kidney Int. 2016, 90, 1238–1250. [Google Scholar] [CrossRef]
- Alhasan, A.H.; Patel, P.C.; Choi, C.H.J.; Mirkin, C.A. Exosome Encased Spherical Nucleic Acid Gold Nanoparticle Conjugates as Potent MicroRNA Regulation Agents. Small 2014, 10, 186–192. [Google Scholar] [CrossRef]
- Saari, H.; Lázaro-Ibáñez, E.; Viitala, T.; Vuorimaa-Laukkanen, E.; Siljander, P.; Yliperttula, M. Microvesicle- and Exosome-Mediated Drug Delivery Enhances the Cytotoxicity of Paclitaxel in Autologous Prostate Cancer Cells. J. Control. Release 2015, 220, 727–737. [Google Scholar] [CrossRef]
- Qu, M.; Lin, Q.; Huang, L.; Fu, Y.; Wang, L.; He, S.; Fu, Y.; Yang, S.; Zhang, Z.; Zhang, L.; et al. Dopamine-Loaded Blood Exosomes Targeted to Brain for Better Treatment of Parkinson’s Disease. J. Control. Release 2018, 287, 156–166. [Google Scholar] [CrossRef]
- Vader, P.; Mol, E.A.; Pasterkamp, G.; Schiffelers, R.M. Extracellular Vesicles for Drug Delivery. Adv. Drug Deliv. Rev. 2016, 106, 148–156. [Google Scholar] [CrossRef]
- Akuma, P.; Okagu, O.D.; Udenigwe, C.C. Naturally Occurring Exosome Vesicles as Potential Delivery Vehicle for Bioactive Compounds. Front. Sustain. Food Syst. 2019. [Google Scholar] [CrossRef]
- Fuhrmann, G.; Serio, A.; Mazo, M.; Nair, R.; Stevens, M.M. Active Loading into Extracellular Vesicles Significantly Improves the Cellular Uptake and Photodynamic Effect of Porphyrins. J. Control. Release 2015, 205, 35–44. [Google Scholar] [CrossRef]
- Luan, X.; Sansanaphongpricha, K.; Myers, I.; Chen, H.; Yuan, H.; Sun, D. Engineering Exosomes as Refined Biological Nanoplatforms for Drug Delivery. Acta Pharmacol. Sin. 2017, 38, 754–763. [Google Scholar] [CrossRef]
- Vashisht, M.; Rani, P.; Onteru, S.K.; Singh, D. Curcumin Encapsulated in Milk Exosomes Resists Human Digestion and Possesses Enhanced Intestinal Permeability In Vitro. Appl. Biochem. Biotechnol. 2017, 183, 993–1007. [Google Scholar] [CrossRef]
- Pascucci, L.; Coccè, V.; Bonomi, A.; Ami, D.; Ceccarelli, P.; Ciusani, E.; Viganò, L.; Locatelli, A.; Sisto, F.; Doglia, S.M.; et al. Paclitaxel Is Incorporated by Mesenchymal Stromal Cells and Released in Exosomes That Inhibit In Vitro Tumor Growth: A New Approach for Drug Delivery. J. Control. Release 2014, 192, 262–270. [Google Scholar] [CrossRef]
- Chen, T.; Guo, J.; Yang, M.; Zhu, X.; Cao, X. Chemokine-Containing Exosomes Are Released from Heat-Stressed Tumor Cells via Lipid Raft-Dependent Pathway and Act as Efficient Tumor Vaccine. J. Immunol. 2011, 186, 2219–2228. [Google Scholar] [CrossRef]
- Wang, L.-Y.; Shi, X.-Y.; Yang, C.-S.; Huang, D.-M. Versatile RBC-Derived Vesicles as Nanoparticle Vector of Photosensitizers for Photodynamic Therapy. Nanoscale 2013, 5, 416–421. [Google Scholar] [CrossRef]
- Lamichhane, T.N.; Jeyaram, A.; Patel, D.B.; Parajuli, B.; Livingston, N.K.; Arumugasaamy, N.; Schardt, J.S.; Jay, S.M. Oncogene Knockdown via Active Loading of Small RNAs into Extracellular Vesicles by Sonication. Cell. Mol. Bioeng. 2016, 9, 315–324. [Google Scholar] [CrossRef] [PubMed]
- Arun, B.; Narendar, D.; Veerabrahma, K. Development of Olmesartan Medoxomil Lipid-Based Nanoparticles and Nanosuspension: Preparation, Characterization and Comparative Pharmacokinetic Evaluation. Artif. Cells Nanomed. Biotechnol. 2018, 46, 126–137. [Google Scholar] [CrossRef]
- Sato, Y.T.; Umezaki, K.; Sawada, S.; Mukai, S.; Sasaki, Y.; Harada, N.; Shiku, H.; Akiyoshi, K. Engineering Hybrid Exosomes by Membrane Fusion with Liposomes. Sci. Rep. 2016, 6, 21933. [Google Scholar] [CrossRef] [PubMed]
- Johnsen, K.B.; Gudbergsson, J.M.; Skov, M.N.; Pilgaard, L.; Moos, T.; Duroux, M. A Comprehensive Overview of Exosomes as Drug Delivery Vehicles—Endogenous Nanocarriers for Targeted Cancer Therapy. Biochim. Biophys. Acta 2014, 1846, 75–87. [Google Scholar] [CrossRef]
- Podolak, I.; Galanty, A.; Sobolewska, D. Saponins as Cytotoxic Agents: A Review. Phytochem. Rev. 2010, 9, 425–474. [Google Scholar] [CrossRef]
- Reshke, R.; Taylor, J.A.; Savard, A.; Guo, H.; Rhym, L.H.; Kowalski, P.S.; Trung, M.T.; Campbell, C.; Little, W.; Anderson, D.G.; et al. Reduction of the Therapeutic Dose of Silencing RNA by Packaging It in Extracellular Vesicles via a Pre-MicroRNA Backbone. Nat. Biomed. Eng. 2020, 4, 52–68. [Google Scholar] [CrossRef]
- Haney, M.J.; Zhao, Y.; Jin, Y.S.; Li, S.M.; Bago, J.R.; Klyachko, N.L.; Kabanov, A.V.; Batrakova, E.V. Macrophage-Derived Extracellular Vesicles as Drug Delivery Systems for Triple Negative Breast Cancer (TNBC) Therapy. J. Neuroimmune Pharmacol. 2020, 15, 487–500. [Google Scholar] [CrossRef] [PubMed]
- Fu, S.; Wang, Y.; Xia, X.; Zheng, J.C. Exosome Engineering: Current Progress in Cargo Loading and Targeted Delivery. NanoImpact 2020, 20, 100261. [Google Scholar] [CrossRef]
- Xu, M.; Yang, Q.; Sun, X.; Wang, Y. Recent Advancements in the Loading and Modification of Therapeutic Exosomes. Front. Bioeng. Biotechnol. 2020, 8, 586130. [Google Scholar] [CrossRef]
- Meng, W.; He, C.; Hao, Y.; Wang, L.; Li, L.; Zhu, G. Prospects and Challenges of Extracellular Vesicle-Based Drug Delivery System: Considering Cell Source. Drug Deliv. 2020, 27, 585–598. [Google Scholar] [CrossRef] [PubMed]
- Antimisiaris, S.G.; Mourtas, S.; Marazioti, A. Exosomes and Exosome-Inspired Vesicles for Targeted Drug Delivery. Pharmaceutics 2018, 10, 218. [Google Scholar] [CrossRef]
- Jafari, D.; Shajari, S.; Jafari, R.; Mardi, N.; Gomari, H.; Ganji, F.; Forouzandeh Moghadam, M.; Samadikuchaksaraei, A. Designer Exosomes: A New Platform for Biotechnology Therapeutics. BioDrugs 2020, 34, 567–586. [Google Scholar] [CrossRef]
- El-Andaloussi, S.; Lee, Y.; Lakhal-Littleton, S.; Li, J.; Seow, Y.; Gardiner, C.; Alvarez-Erviti, L.; Sargent, I.L.; Wood, M.J.A. Exosome-Mediated Delivery of SiRNA In Vitro and In Vivo. Nat. Protoc. 2012, 7, 2112–2126. [Google Scholar] [CrossRef]
- Takahashi, Y.; Nishikawa, M.; Shinotsuka, H.; Matsui, Y.; Ohara, S.; Imai, T.; Takakura, Y. Visualization and In Vivo Tracking of the Exosomes of Murine Melanoma B16-BL6 Cells in Mice after Intravenous Injection. J. Biotechnol. 2013, 165, 77–84. [Google Scholar] [CrossRef]
- Peinado, H.; Alečković, M.; Lavotshkin, S.; Matei, I.; Costa-Silva, B.; Moreno-Bueno, G.; Hergueta-Redondo, M.; Williams, C.; García-Santos, G.; Ghajar, C.; et al. Melanoma Exosomes Educate Bone Marrow Progenitor Cells toward a Pro-Metastatic Phenotype through MET. Nat. Med. 2012, 18, 883–891. [Google Scholar] [CrossRef]
- Imai, T.; Takahashi, Y.; Nishikawa, M.; Kato, K.; Morishita, M.; Yamashita, T.; Matsumoto, A.; Charoenviriyakul, C.; Takakura, Y. Macrophage-Dependent Clearance of Systemically Administered B16BL6-Derived Exosomes from the Blood Circulation in Mice. J. Extracell. Vesicles 2015, 4, 26238. [Google Scholar] [CrossRef]
- Morishita, M.; Takahashi, Y.; Nishikawa, M.; Sano, K.; Kato, K.; Yamashita, T.; Imai, T.; Saji, H.; Takakura, Y. Quantitative Analysis of Tissue Distribution of the B16BL6-Derived Exosomes Using a Streptavidin-Lactadherin Fusion Protein and Iodine-125-Labeled Biotin Derivative after Intravenous Injection in Mice. J. Pharm. Sci. 2015, 104, 705–713. [Google Scholar] [CrossRef]
- Kosaka, N.; Iguchi, H.; Yoshioka, Y.; Hagiwara, K.; Takeshita, F.; Ochiya, T. Competitive Interactions of Cancer Cells and Normal Cells via Secretory MicroRNAs. J. Biol. Chem. 2012, 287, 1397–1405. [Google Scholar] [CrossRef]
- Mizrak, A.; Bolukbasi, M.F.; Ozdener, G.B.; Brenner, G.J.; Madlener, S.; Erkan, E.P.; Ströbel, T.; Breakefield, X.O.; Saydam, O. Genetically Engineered Microvesicles Carrying Suicide MRNA/Protein Inhibit Schwannoma Tumor Growth. Mol. Ther. 2013, 21, 101–108. [Google Scholar] [CrossRef]
- Rezaie, J.; Ajezi, S.; Avci, Ç.B.; Karimipour, M.; Geranmayeh, M.H.; Nourazarian, A.; Sokullu, E.; Rezabakhsh, A.; Rahbarghazi, R. Exosomes and Their Application in Biomedical Field: Difficulties and Advantages. Mol. Neurobiol. 2018, 55, 3372–3393. [Google Scholar] [CrossRef]
- Agrawal, A.K.; Aqil, F.; Jeyabalan, J.; Spencer, W.A.; Beck, J.; Gachuki, B.W.; Alhakeem, S.S.; Oben, K.; Munagala, R.; Bondada, S.; et al. Milk-Derived Exosomes for Oral Delivery of Paclitaxel. Nanomedicine 2017, 13, 1627–1636. [Google Scholar] [CrossRef] [PubMed]
- Aqil, F.; Munagala, R.; Jeyabalan, J.; Agrawal, A.K.; Gupta, R. Exosomes for the Enhanced Tissue Bioavailability and Efficacy of Curcumin. AAPS J. 2017, 19, 1691–1702. [Google Scholar] [CrossRef]
- Braun, R.K.; Chetty, C.; Balasubramaniam, V.; Centanni, R.; Haraldsdottir, K.; Hematti, P.; Eldridge, M.W. Intraperitoneal Injection of MSC-Derived Exosomes Prevent Experimental Bronchopulmonary Dysplasia. Biochem. Biophys. Res. Commun. 2018, 503, 2653–2658. [Google Scholar] [CrossRef]
- Yu, M.; Shi, J.; Sheng, M. Exosomes: The New Mediator of Peritoneal Membrane Function. KBR 2018, 43, 1010–1022. [Google Scholar] [CrossRef]
- Ortega, A.; Martinez-Arroyo, O.; Forner, M.J.; Cortes, R. Exosomes as Drug Delivery Systems: Endogenous Nanovehicles for Treatment of Systemic Lupus Erythematosus. Pharmaceutics 2020, 13, 3. [Google Scholar] [CrossRef]
- Ju, S.; Mu, J.; Dokland, T.; Zhuang, X.; Wang, Q.; Jiang, H.; Xiang, X.; Deng, Z.-B.; Wang, B.; Zhang, L.; et al. Grape Exosome-like Nanoparticles Induce Intestinal Stem Cells and Protect Mice From DSS-Induced Colitis. Mol. Ther. 2013, 21, 1345–1357. [Google Scholar] [CrossRef]
- Pinheiro, A.; Silva, A.M.; Teixeira, J.H.; Gonçalves, R.M.; Almeida, M.I.; Barbosa, M.A.; Santos, S.G. Extracellular Vesicles: Intelligent Delivery Strategies for Therapeutic Applications. J. Control. Release 2018, 289, 56–69. [Google Scholar] [CrossRef]
- Paolini, L.; Zendrini, A.; Di Noto, G.; Busatto, S.; Lottini, E.; Radeghieri, A.; Dossi, A.; Caneschi, A.; Ricotta, D.; Bergese, P. Residual Matrix from Different Separation Techniques Impacts Exosome Biological Activity. Sci. Rep. 2016, 6, 23550. [Google Scholar] [CrossRef]
- Parisse, P.; Rago, I.; Ulloa Severino, L.; Perissinotto, F.; Ambrosetti, E.; Paoletti, P.; Ricci, M.; Beltrami, A.P.; Cesselli, D.; Casalis, L. Atomic Force Microscopy Analysis of Extracellular Vesicles. Eur. Biophys. J. 2017, 46, 813–820. [Google Scholar] [CrossRef]
- Gurunathan, S.; Kang, M.-H.; Jeyaraj, M.; Qasim, M.; Kim, J.-H. Review of the Isolation, Characterization, Biological Function, and Multifarious Therapeutic Approaches of Exosomes. Cells 2019, 8, 307. [Google Scholar] [CrossRef]
- Wu, Y.; Deng, W.; Ii, D.J.K. Exosomes: Improved Methods to Characterize Their Morphology, RNA Content, and Surface Protein Biomarkers. Analyst 2015, 140, 6631–6642. [Google Scholar] [CrossRef]
- Jung, M.K.; Mun, J.Y. Sample Preparation and Imaging of Exosomes by Transmission Electron Microscopy. J. Vis. Exp. 2018. [Google Scholar] [CrossRef]
- Sokolova, V.; Ludwig, A.-K.; Hornung, S.; Rotan, O.; Horn, P.A.; Epple, M.; Giebel, B. Characterisation of Exosomes Derived from Human Cells by Nanoparticle Tracking Analysis and Scanning Electron Microscopy. Colloids Surf. B Biointerfaces 2011, 87, 146–150. [Google Scholar] [CrossRef]
- Brinton, L.T.; Sloane, H.S.; Kester, M.; Kelly, K.A. Formation and Role of Exosomes in Cancer. Cell. Mol. Life Sci. 2015, 72, 659–671. [Google Scholar] [CrossRef]
- Butreddy, A.; Janga, K.Y.; Ajjarapu, S.; Sarabu, S.; Dudhipala, N. Instability of Therapeutic Proteins—An Overview of Stresses, Stabilization Mechanisms and Analytical Techniques Involved in Lyophilized Proteins. Int. J. Biol. Macromol. 2021, 167, 309–325. [Google Scholar] [CrossRef] [PubMed]
- Hoo, C.M.; Starostin, N.; West, P.; Mecartney, M.L. A Comparison of Atomic Force Microscopy (AFM) and Dynamic Light Scattering (DLS) Methods to Characterize Nanoparticle Size Distributions. J. Nanoparticle Res. 2008, 10, 89–96. [Google Scholar] [CrossRef]
- Gercel-Taylor, C.; Atay, S.; Tullis, R.H.; Kesimer, M.; Taylor, D.D. Nanoparticle Analysis of Circulating Cell-Derived Vesicles in Ovarian Cancer Patients. Anal. Biochem. 2012, 428, 44–53. [Google Scholar] [CrossRef] [PubMed]
- Pospichalova, V.; Svoboda, J.; Dave, Z.; Kotrbova, A.; Kaiser, K.; Klemova, D.; Ilkovics, L.; Hampl, A.; Crha, I.; Jandakova, E.; et al. Simplified Protocol for Flow Cytometry Analysis of Fluorescently Labeled Exosomes and Microvesicles Using Dedicated Flow Cytometer. J. Extracell. Vesicles 2015, 4, 25530. [Google Scholar] [CrossRef]
- Melo, S.A.; Luecke, L.B.; Kahlert, C.; Fernandez, A.F.; Gammon, S.T.; Kaye, J.; LeBleu, V.S.; Mittendorf, E.A.; Weitz, J.; Rahbari, N.; et al. Glypican-1 Identifies Cancer Exosomes and Detects Early Pancreatic Cancer. Nature 2015, 523, 177–182. [Google Scholar] [CrossRef] [PubMed]
- Van der Pol, E.; Coumans, F.A.W.; Grootemaat, A.E.; Gardiner, C.; Sargent, I.L.; Harrison, P.; Sturk, A.; van Leeuwen, T.G.; Nieuwland, R. Particle Size Distribution of Exosomes and Microvesicles Determined by Transmission Electron Microscopy, Flow Cytometry, Nanoparticle Tracking Analysis, and Resistive Pulse Sensing. J. Thromb. Haemost. 2014, 12, 1182–1192. [Google Scholar] [CrossRef]
- Thane, K.E.; Davis, A.M.; Hoffman, A.M. Improved Methods for Fluorescent Labeling and Detection of Single Extracellular Vesicles Using Nanoparticle Tracking Analysis. Sci. Rep. 2019, 9, 12295. [Google Scholar] [CrossRef]
- Vestad, B.; Llorente, A.; Neurauter, A.; Phuyal, S.; Kierulf, B.; Kierulf, P.; Skotland, T.; Sandvig, K.; Haug, K.B.F.; Øvstebø, R. Size and Concentration Analyses of Extracellular Vesicles by Nanoparticle Tracking Analysis: A Variation Study. J. Extracell. Vesicles 2017, 6, 1344087. [Google Scholar] [CrossRef]
- Szatanek, R.; Baj-Krzyworzeka, M.; Zimoch, J.; Lekka, M.; Siedlar, M.; Baran, J. The Methods of Choice for Extracellular Vesicles (EVs) Characterization. Int. J. Mol. Sci. 2017, 18, 1153. [Google Scholar] [CrossRef]
- Anderson, W.; Lane, R.; Korbie, D.; Trau, M. Observations of Tunable Resistive Pulse Sensing for Exosome Analysis: Improving System Sensitivity and Stability. Langmuir 2015, 31, 6577–6587. [Google Scholar] [CrossRef] [PubMed]
- Théry, C.; Witwer, K.W.; Aikawa, E.; Alcaraz, M.J.; Anderson, J.D.; Andriantsitohaina, R.; Antoniou, A.; Arab, T.; Archer, F.; Atkin-Smith, G.K.; et al. Minimal Information for Studies of Extracellular Vesicles 2018 (MISEV2018): A Position Statement of the International Society for Extracellular Vesicles and Update of the MISEV2014 Guidelines. J. Extracell. Vesicles 2018, 7, 1535750. [Google Scholar] [CrossRef]
- Rissin, D.M.; Kan, C.W.; Campbell, T.G.; Howes, S.C.; Fournier, D.R.; Song, L.; Piech, T.; Patel, P.P.; Chang, L.; Rivnak, A.J.; et al. Single-Molecule Enzyme-Linked Immunosorbent Assay Detects Serum Proteins at Subfemtomolar Concentrations. Nat. Biotechnol. 2010, 28, 595–599. [Google Scholar] [CrossRef]
- Shao, H.; Im, H.; Castro, C.M.; Breakefield, X.; Weissleder, R.; Lee, H. New Technologies for Analysis of Extracellular Vesicles. Chem. Rev. 2018, 118, 1917–1950. [Google Scholar] [CrossRef] [PubMed]
- Coumans, F.A.W.; Gool, E.L.; Nieuwland, R. Bulk Immunoassays for Analysis of Extracellular Vesicles. Platelets 2017, 28, 242–248. [Google Scholar] [CrossRef]
- Pisitkun, T.; Shen, R.-F.; Knepper, M.A. Identification and Proteomic Profiling of Exosomes in Human Urine. Proc. Natl. Acad. Sci. USA 2004, 101, 13368–13373. [Google Scholar] [CrossRef]
- Maas, S.L.N.; de Vrij, J.; van der Vlist, E.J.; Geragousian, B.; van Bloois, L.; Mastrobattista, E.; Schiffelers, R.M.; Wauben, M.H.M.; Broekman, M.L.D.; Nolte-’t Hoen, E.N.M. Possibilities and Limitations of Current Technologies for Quantification of Biological Extracellular Vesicles and Synthetic Mimics. J. Control. Release 2015, 200, 87–96. [Google Scholar] [CrossRef]
- Zhang, Y.; Bi, J.; Huang, J.; Tang, Y.; Du, S.; Li, P. Exosome: A Review of Its Classification, Isolation Techniques, Storage, Diagnostic and Targeted Therapy Applications. Int. J. Nanomed. 2020, 15, 6917–6934. [Google Scholar] [CrossRef]
- Chiriacò, M.S.; Bianco, M.; Nigro, A.; Primiceri, E.; Ferrara, F.; Romano, A.; Quattrini, A.; Furlan, R.; Arima, V.; Maruccio, G. Lab-on-Chip for Exosomes and Microvesicles Detection and Characterization. Sensors 2018, 18, 3175. [Google Scholar] [CrossRef]
- Zhang, Y.; Liu, Y.; Liu, H.; Tang, W.H. Exosomes: Biogenesis, Biologic Function and Clinical Potential. Cell Biosci. 2019, 9, 19. [Google Scholar] [CrossRef]
- Chuo, S.T.-Y.; Chien, J.C.-Y.; Lai, C.P.-K. Imaging Extracellular Vesicles: Current and Emerging Methods. J. Biomed. Sci. 2018, 25, 91. [Google Scholar] [CrossRef] [PubMed]
- Dominici, M.; Le Blanc, K.; Mueller, I.; Slaper-Cortenbach, I.; Marini, F.; Krause, D.; Deans, R.; Keating, A.; Prockop, D.; Horwitz, E. Minimal Criteria for Defining Multipotent Mesenchymal Stromal Cells. The International Society for Cellular Therapy Position Statement. Cytotherapy 2006, 8, 315–317. [Google Scholar] [CrossRef]
- Chen, Y.-S.; Lin, E.-Y.; Chiou, T.-W.; Harn, H.-J. Exosomes in Clinical Trial and Their Production in Compliance with Good Manufacturing Practice. Available online: https://www.tcmjmed.com/article.asp?issn=1016-3190;year=2020;volume=32;issue=2;spage=113;epage=120;aulast=Chen (accessed on 3 April 2021).
- Soekmadji, C.; Hill, A.F.; Wauben, M.H.; Buzás, E.I.; Di Vizio, D.; Gardiner, C.; Lötvall, J.; Sahoo, S.; Witwer, K.W. Towards Mechanisms and Standardization in Extracellular Vesicle and Extracellular RNA Studies: Results of a Worldwide Survey. J. Extracell. Vesicles 2018, 7, 1535745. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Lee, Y.; Johansson, H.J.; Mäger, I.; Vader, P.; Nordin, J.Z.; Wiklander, O.P.B.; Lehtiö, J.; Wood, M.J.A.; Andaloussi, S.E. Serum-Free Culture Alters the Quantity and Protein Composition of Neuroblastoma-Derived Extracellular Vesicles. J. Extracell. Vesicles 2015, 4, 26883. [Google Scholar] [CrossRef] [PubMed]
- Andriolo, G.; Provasi, E.; Lo Cicero, V.; Brambilla, A.; Soncin, S.; Torre, T.; Milano, G.; Biemmi, V.; Vassalli, G.; Turchetto, L.; et al. Exosomes From Human Cardiac Progenitor Cells for Therapeutic Applications: Development of a GMP-Grade Manufacturing Method. Front. Physiol. 2018, 9, 1169. [Google Scholar] [CrossRef]
- Pachler, K.; Lener, T.; Streif, D.; Dunai, Z.A.; Desgeorges, A.; Feichtner, M.; Öller, M.; Schallmoser, K.; Rohde, E.; Gimona, M. A Good Manufacturing Practice-Grade Standard Protocol for Exclusively Human Mesenchymal Stromal Cell-Derived Extracellular Vesicles. Cytotherapy 2017, 19, 458–472. [Google Scholar] [CrossRef]
- Doyle, L.M.; Wang, M.Z. Overview of extracellular vesicles, their origin, composition, purpose, and methods for exosome isolation and analysis. Cells 2019, 8, 727. [Google Scholar] [CrossRef] [PubMed]
- Adlerz, K.; Patel, D.; Rowley, J.; Ng, K.; Ahsan, T. Strategies for Scalable Manufacturing and Translation of MSC-Derived Extracellular Vesicles. Stem Cell Research 2020, 48, 101978. [Google Scholar] [CrossRef]
- Colao, I.L.; Corteling, R.; Bracewell, D.; Wall, I. Manufacturing Exosomes: A Promising Therapeutic Platform. Trends Mol. Med. 2018, 24, 242–256. [Google Scholar] [CrossRef] [PubMed]
- Bari, E.; Perteghella, S.; Catenacci, L.; Sorlini, M.; Croce, S.; Mantelli, M.; Avanzini, M.A.; Sorrenti, M.; Torre, M.L. Freeze-Dried and GMP-Compliant Pharmaceuticals Containing Exosomes for Acellular Mesenchymal Stromal Cell Immunomodulant Therapy. Nanomedicine 2019, 14, 753–765. [Google Scholar] [CrossRef]
- Wen, Y.-T.; Chang, Y.-C.; Lin, L.-C.; Liao, P.-C. Collection of In Vivo-like Liver Cell Secretome with Alternative Sample Enrichment Method Using a Hollow Fiber Bioreactor Culture System Combined with Tangential Flow Filtration for Secretomics Analysis. Anal. Chim. Acta 2011, 684, 72–79. [Google Scholar] [CrossRef]
- Morenweiser, R. Downstream Processing of Viral Vectors and Vaccines. Gene Ther. 2005, 12, S103–S110. [Google Scholar] [CrossRef]
- Nordin, J.Z.; Lee, Y.; Vader, P.; Mäger, I.; Johansson, H.J.; Heusermann, W.; Wiklander, O.P.B.; Hällbrink, M.; Seow, Y.; Bultema, J.J.; et al. Ultrafiltration with Size-Exclusion Liquid Chromatography for High Yield Isolation of Extracellular Vesicles Preserving Intact Biophysical and Functional Properties. Nanomedicine 2015, 11, 879–883. [Google Scholar] [CrossRef]
- Gardiner, C.; Di Vizio, D.; Sahoo, S.; Théry, C.; Witwer, K.W.; Wauben, M.; Hill, A.F. Techniques Used for the Isolation and Characterization of Extracellular Vesicles: Results of a Worldwide Survey. J. Extracell. Vesicles 2016, 5, 32945. [Google Scholar] [CrossRef]
- Gimona, M.; Pachler, K.; Laner-Plamberger, S.; Schallmoser, K.; Rohde, E. Manufacturing of Human Extracellular Vesicle-Based Therapeutics for Clinical Use. Int. J. Mol. Sci. 2017, 18, 1190. [Google Scholar] [CrossRef]
- Witwer, K.W.; Buzás, E.I.; Bemis, L.T.; Bora, A.; Lässer, C.; Lötvall, J.; Nolte-’t Hoen, E.N.; Piper, M.G.; Sivaraman, S.; Skog, J.; et al. Standardization of Sample Collection, Isolation and Analysis Methods in Extracellular Vesicle Research. J. Extracell. Vesicles 2013, 2. [Google Scholar] [CrossRef]
- Jeyaram, A.; Jay, S.M. Preservation and Storage Stability of Extracellular Vesicles for Therapeutic Applications. AAPS J. 2017, 20, 1. [Google Scholar] [CrossRef]
- Maroto, R.; Zhao, Y.; Jamaluddin, M.; Popov, V.L.; Wang, H.; Kalubowilage, M.; Zhang, Y.; Luisi, J.; Sun, H.; Culbertson, C.T.; et al. Effects of Storage Temperature on Airway Exosome Integrity for Diagnostic and Functional Analyses. J. Extracell. Vesicles 2017, 6, 1359478. [Google Scholar] [CrossRef]
- Lőrincz, Á.M.; Timár, C.I.; Marosvári, K.A.; Veres, D.S.; Otrokocsi, L.; Kittel, Á.; Ligeti, E. Effect of Storage on Physical and Functional Properties of Extracellular Vesicles Derived from Neutrophilic Granulocytes. J. Extracell. Vesicles 2014, 3, 25465. [Google Scholar] [CrossRef]
- Linares, R.; Tan, S.; Gounou, C.; Arraud, N.; Brisson, A.R. High-Speed Centrifugation Induces Aggregation of Extracellular Vesicles. J. Extracell. Vesicles 2015, 4, 29509. [Google Scholar] [CrossRef] [PubMed]
- Charoenviriyakul, C.; Takahashi, Y.; Nishikawa, M.; Takakura, Y. Preservation of Exosomes at Room Temperature Using Lyophilization. Int. J. Pharm. 2018, 553, 1–7. [Google Scholar] [CrossRef]
- Bahr, M.M.; Amer, M.S.; Abo-El-Sooud, K.; Abdallah, A.N.; El-Tookhy, O.S. Preservation Techniques of Stem Cells Extracellular Vesicles: A Gate for Manufacturing of Clinical Grade Therapeutic Extracellular Vesicles and Long-Term Clinical Trials. Int. J. Vet. Sci. Med. 2020, 8, 1–8. [Google Scholar] [CrossRef]
- Lee, M.; Ban, J.-J.; Im, W.; Kim, M. Influence of Storage Condition on Exosome Recovery. Biotechnol. Bioproc. E 2016, 21, 299–304. [Google Scholar] [CrossRef]
- Yadava, P.; Gibbs, M.; Castro, C.; Hughes, J.A. Effect of Lyophilization and Freeze-Thawing on the Stability of SiRNA-Liposome Complexes. AAPS PharmSciTech 2008, 9, 335–341. [Google Scholar] [CrossRef]
- Butreddy, A.; Dudhipala, N.; Janga, K.Y.; Gaddam, R.P. Lyophilization of Small-Molecule Injectables: An Industry Perspective on Formulation Development, Process Optimization, Scale-Up Challenges, and Drug Product Quality Attributes. AAPS PharmSciTech 2020, 21, 252. [Google Scholar] [CrossRef]
- Bosch, S.; de Beaurepaire, L.; Allard, M.; Mosser, M.; Heichette, C.; Chrétien, D.; Jegou, D.; Bach, J.-M. Trehalose Prevents Aggregation of Exosomes and Cryodamage. Sci. Rep. 2016, 6, 36162. [Google Scholar] [CrossRef]
- Hood, J.L.; Scott, M.J.; Wickline, S.A. Maximizing Exosome Colloidal Stability Following Electroporation. Anal. Biochem. 2014, 448, 41–49. [Google Scholar] [CrossRef]
- Motta, J.P.R.; Paraguassú-Braga, F.H.; Bouzas, L.F.; Porto, L.C. Evaluation of Intracellular and Extracellular Trehalose as a Cryoprotectant of Stem Cells Obtained from Umbilical Cord Blood. Cryobiology 2014, 68, 343–348. [Google Scholar] [CrossRef]
- Wang, Y.; Grainger, D.W. Lyophilized Liposome-Based Parenteral Drug Development: Reviewing Complex Product Design Strategies and Current Regulatory Environments. Adv. Drug Deliv. Rev. 2019, 151–152, 56–71. [Google Scholar] [CrossRef]
- Chen, C.; Han, D.; Cai, C.; Tang, X. An Overview of Liposome Lyophilization and Its Future Potential. J. Control. Release 2010, 142, 299–311. [Google Scholar] [CrossRef]
- Lee, M.K.; Kim, M.Y.; Kim, S.; Lee, J. Cryoprotectants for Freeze Drying of Drug Nano-Suspensions: Effect of Freezing Rate. J. Pharm. Sci. 2009, 98, 4808–4817. [Google Scholar] [CrossRef]
- Akers, J.C.; Ramakrishnan, V.; Yang, I.; Hua, W.; Mao, Y.; Carter, B.S.; Chen, C.C. Optimizing Preservation of Extracellular Vesicular MiRNAs Derived from Clinical Cerebrospinal Fluid. Cancer Biomark. 2016, 17, 125–132. [Google Scholar] [CrossRef]
- Mendt, M.; Kamerkar, S.; Sugimoto, H.; McAndrews, K.M.; Wu, C.-C.; Gagea, M.; Yang, S.; Blanko, E.V.R.; Peng, Q.; Ma, X.; et al. Generation and Testing of Clinical-Grade Exosomes for Pancreatic Cancer. JCI Insight 2018, 3. [Google Scholar] [CrossRef]
- Vishnubhatla, I.; Corteling, R.; Stevanato, L.; Hicks, C.; Sinden, J. The Development of Stem Cell-Derived Exosomes as a Cell-Free Regenerative Medicine. J. Circ. Biomark. 2014, 3, 2. [Google Scholar] [CrossRef]
- Whitford, W.; Guterstam, P. Exosome Manufacturing Status. Future Med. Chem. 2019, 11, 1225–1236. [Google Scholar] [CrossRef] [PubMed]
- Escudier, B.; Dorval, T.; Chaput, N.; André, F.; Caby, M.-P.; Novault, S.; Flament, C.; Leboulaire, C.; Borg, C.; Amigorena, S.; et al. Vaccination of Metastatic Melanoma Patients with Autologous Dendritic Cell (DC) Derived-Exosomes: Results of Thefirst Phase I Clinical Trial. J. Transl. Med. 2005, 3, 10. [Google Scholar] [CrossRef] [PubMed]
- Morse, M.A.; Garst, J.; Osada, T.; Khan, S.; Hobeika, A.; Clay, T.M.; Valente, N.; Shreeniwas, R.; Sutton, M.A.; Delcayre, A.; et al. A Phase I Study of Dexosome Immunotherapy in Patients with Advanced Non-Small Cell Lung Cancer. J. Transl. Med. 2005, 3, 9. [Google Scholar] [CrossRef]
- Besse, B.; Charrier, M.; Lapierre, V.; Dansin, E.; Lantz, O.; Planchard, D.; Le Chevalier, T.; Livartoski, A.; Barlesi, F.; Laplanche, A.; et al. Dendritic Cell-Derived Exosomes as Maintenance Immunotherapy after First Line Chemotherapy in NSCLC. Oncoimmunology 2016, 5, e1071008. [Google Scholar] [CrossRef]
- Pitt, J.M.; André, F.; Amigorena, S.; Soria, J.-C.; Eggermont, A.; Kroemer, G.; Zitvogel, L. Dendritic Cell-Derived Exosomes for Cancer Therapy. J. Clin. Invest. 2016, 126, 1224–1232. [Google Scholar] [CrossRef]
- Tang, K.; Zhang, Y.; Zhang, H.; Xu, P.; Liu, J.; Ma, J.; Lv, M.; Li, D.; Katirai, F.; Shen, G.-X.; et al. Delivery of Chemotherapeutic Drugs in Tumour Cell-Derived Microparticles. Nat. Commun. 2012, 3, 1282. [Google Scholar] [CrossRef]
- Zitvogel, L.; Regnault, A.; Lozier, A.; Wolfers, J.; Flament, C.; Tenza, D.; Ricciardi-Castagnoli, P.; Raposo, G.; Amigorena, S. Eradication of Established Murine Tumors Using a Novel Cell-Free Vaccine: Dendritic Cell-Derived Exosomes. Nat. Med. 1998, 4, 594–600. [Google Scholar] [CrossRef]
- Lamparski, H.G.; Metha-Damani, A.; Yao, J.-Y.; Patel, S.; Hsu, D.-H.; Ruegg, C.; Le Pecq, J.-B. Production and Characterization of Clinical Grade Exosomes Derived from Dendritic Cells. J. Immunol. Methods 2002, 270, 211–226. [Google Scholar] [CrossRef]
- Viaud, S.; Terme, M.; Flament, C.; Taieb, J.; André, F.; Novault, S.; Escudier, B.; Robert, C.; Caillat-Zucman, S.; Tursz, T.; et al. Dendritic Cell-Derived Exosomes Promote Natural Killer Cell Activation and Proliferation: A Role for NKG2D Ligands and IL-15Ralpha. PLoS ONE 2009, 4, e4942. [Google Scholar] [CrossRef]
- Mathivanan, S.; Ji, H.; Simpson, R.J. Exosomes: Extracellular Organelles Important in Intercellular Communication. J. Proteomics 2010, 73, 1907–1920. [Google Scholar] [CrossRef] [PubMed]
- Mendt, M.; Rezvani, K.; Shpall, E. Mesenchymal Stem Cell-Derived Exosomes for Clinical Use. Bone Marrow Transplant. 2019, 54, 789–792. [Google Scholar] [CrossRef]
- Ezquer, F.; Ezquer, M.; Contador, D.; Ricca, M.; Simon, V.; Conget, P. The Antidiabetic Effect of Mesenchymal Stem Cells Is Unrelated to Their Transdifferentiation Potential but to Their Capability to Restore Th1/Th2 Balance and to Modify the Pancreatic Microenvironment. Stem Cells 2012, 30, 1664–1674. [Google Scholar] [CrossRef]
- Zhao, Y.; Jiang, Z.; Zhao, T.; Ye, M.; Hu, C.; Yin, Z.; Li, H.; Zhang, Y.; Diao, Y.; Li, Y.; et al. Reversal of Type 1 Diabetes via Islet β Cell Regeneration Following Immune Modulation by Cord Blood-Derived Multipotent Stem Cells. BMC Med. 2012, 10, 3. [Google Scholar] [CrossRef]
- Jiang, Y.; Zhang, Y.; Zhang, L.; Wang, M.; Zhang, X.; Li, X. Therapeutic Effect of Bone Marrow Mesenchymal Stem Cells on Laser-Induced Retinal Injury in Mice. Int. J. Mol. Sci. 2014, 15, 9372–9385. [Google Scholar] [CrossRef]
- Nikfarjam, S.; Rezaie, J.; Zolbanin, N.M.; Jafari, R. Mesenchymal Stem Cell Derived-Exosomes: A Modern Approach in Translational Medicine. J. Transl. Med. 2020, 18, 449. [Google Scholar] [CrossRef]
- IBI—International Biopharmaceutical Industry. Available online: https://biopharmaceuticalmedia.com/ (accessed on 3 April 2021).
- Aegle Therapeutics. Available online: https://www.aegletherapeutics.com/ (accessed on 4 April 2021).
- Hanna, E.; Rémuzat, C.; Auquier, P.; Toumi, M. Advanced Therapy Medicinal Products: Current and Future Perspectives. J. Mark. Access Health Policy 2016, 4. [Google Scholar] [CrossRef]
- Conlan, R.S.; Pisano, S.; Oliveira, M.I.; Ferrari, M.; Mendes Pinto, I. Exosomes as Reconfigurable Therapeutic Systems. Trends Mol. Med. 2017, 23, 636–650. [Google Scholar] [CrossRef] [PubMed]
- Biopharma Excellence. Available online: https://www.biopharma-excellence.com (accessed on 3 April 2021).
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Butreddy, A.; Kommineni, N.; Dudhipala, N. Exosomes as Naturally Occurring Vehicles for Delivery of Biopharmaceuticals: Insights from Drug Delivery to Clinical Perspectives. Nanomaterials 2021, 11, 1481. https://doi.org/10.3390/nano11061481
Butreddy A, Kommineni N, Dudhipala N. Exosomes as Naturally Occurring Vehicles for Delivery of Biopharmaceuticals: Insights from Drug Delivery to Clinical Perspectives. Nanomaterials. 2021; 11(6):1481. https://doi.org/10.3390/nano11061481
Chicago/Turabian StyleButreddy, Arun, Nagavendra Kommineni, and Narendar Dudhipala. 2021. "Exosomes as Naturally Occurring Vehicles for Delivery of Biopharmaceuticals: Insights from Drug Delivery to Clinical Perspectives" Nanomaterials 11, no. 6: 1481. https://doi.org/10.3390/nano11061481
APA StyleButreddy, A., Kommineni, N., & Dudhipala, N. (2021). Exosomes as Naturally Occurring Vehicles for Delivery of Biopharmaceuticals: Insights from Drug Delivery to Clinical Perspectives. Nanomaterials, 11(6), 1481. https://doi.org/10.3390/nano11061481





