SERS-Based Aptasensor for Rapid Quantitative Detection of SARS-CoV-2
Abstract
1. Introduction
2. Materials and Methods
2.1. Procedure of AgNPs Synthesis
2.2. Viruses
2.3. Aptamers and Their Assembly
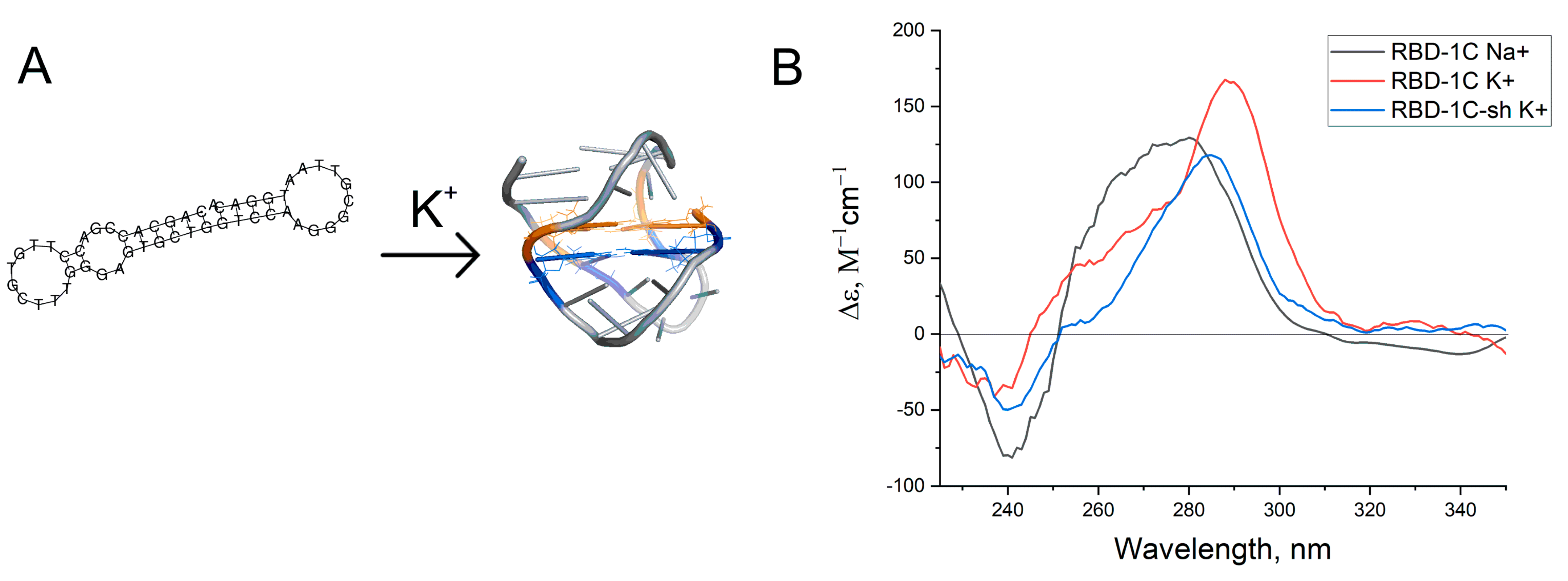
2.4. Circular Dichroism and UV-Spectroscopy
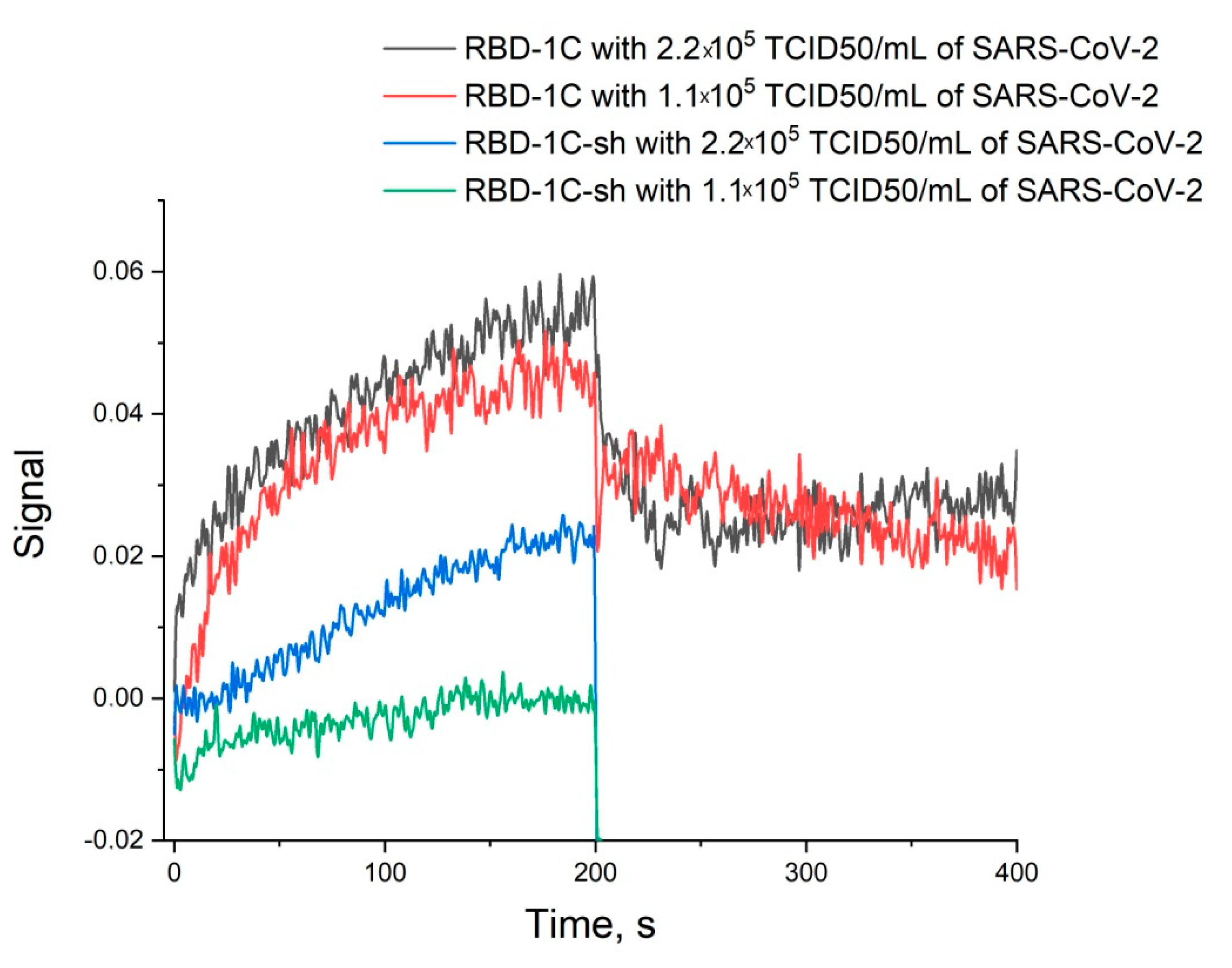
2.5. Determination of Aptamer Affinity to S-Protein and Binding to SARS-CoV-2 Virus
- (1)
- baseline in the buffer during 30 s;
- (2)
- association stage in 80–1200nM RBD of S-protein (from HyTest, Turku, Finland) solution during 200 s;
- (3)
- dissociation stage in the buffer during 300 s.
- (1)
- An amount of 0.1 M NaH2PO4 pH = 6.2 for 2 min.
- (2)
- The activation of -NH2 groups of APTES by EDC (50 mg/mL) and NHS (50 mg/mL) for 20 min.
- (3)
- An amount of 50 mM MES pH~5.0 for 5 min.
- (4)
- A covalent conjugation of streptavidin to the activated -NH2 groups. Streptavidin (40 µg/mL) in 50 mM MES pH~5.0 for 30 min.
- (5)
- Washing with 50 mM MES pH~5.0 for 2 min.
- (6)
- Washing with 5 mM glycine in 25 mM HCl (pH~2) for 1–2 min.
- (7)
- Washing with 10 mM Tris-HCl (pH = 7.4), 140 mM NaCl, 10 mM KCl for 5 min.
- (8)
- Loading of biotinylated aptamer to SARS-CoV-2 (RBD-1C) for 15–20 min.
- (9)
- Washing with 10 mM Tris-HCl (pH = 7.4), 140 mM NaCl, 10 mM KCl for 5 min.
- (9a)
- Interaction with cell medium for 3 min.
- (9b)
- Interaction with SARS-CoV-2 virus with a titer of 0.22·106 TCID50/mL or 0.11·106 TCID50/mL for 3 min.
- (10)
- Dissociation in 10 mM Tris-HCl (pH = 7.4), 140 mM NaCl, 10 mM KCl for 5 min.
2.6. SERS Measurements
3. Results
3.1. Characterization of Silver Colloidal Nanoparticles
3.2. Structural Characterization of Aptamer RBD-1C
3.3. Affinity of RBD-1C and Its Shortened Version to RBD of S-Protein and the Whole Virus
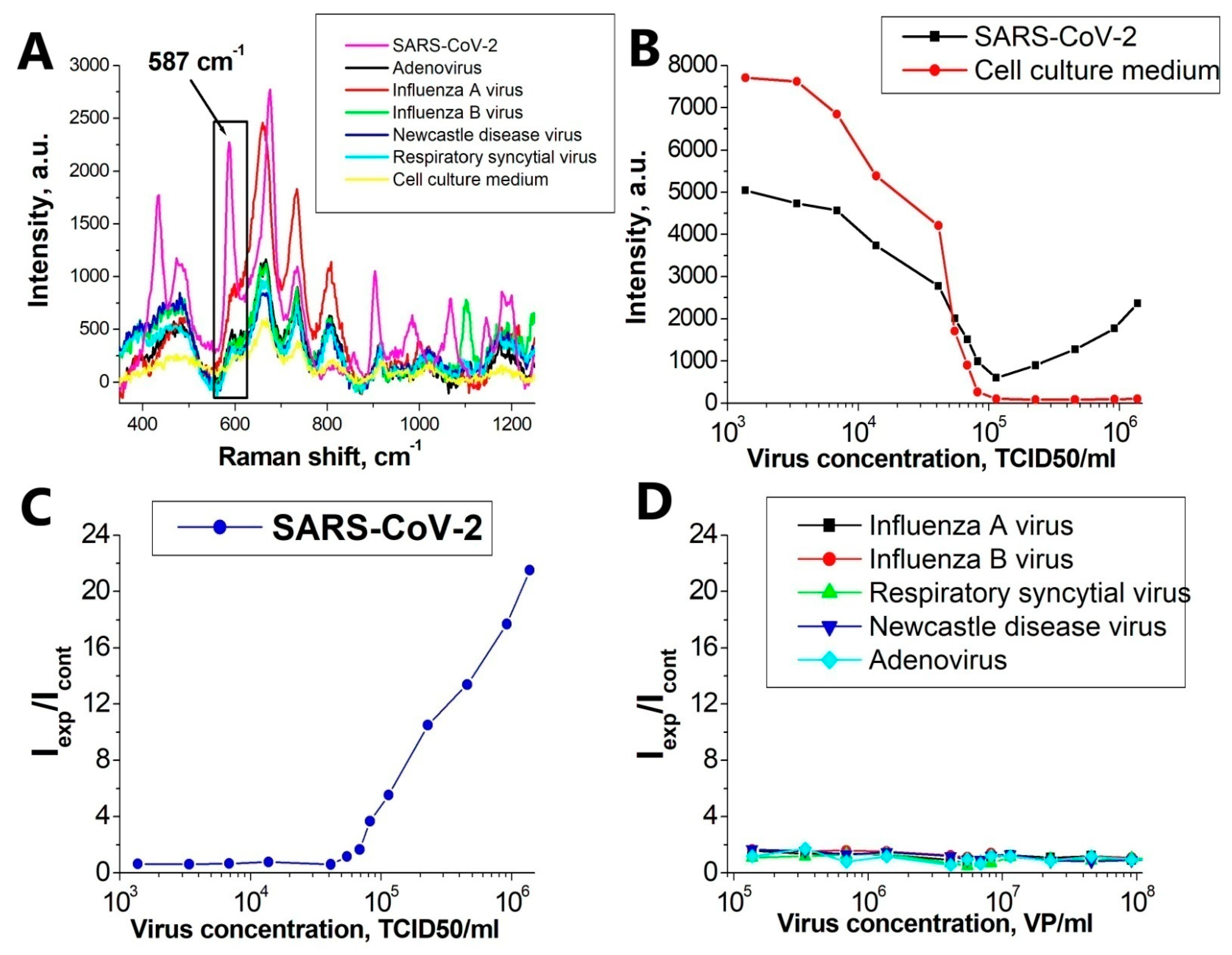
3.4. Specificity of SERS Aptasensor
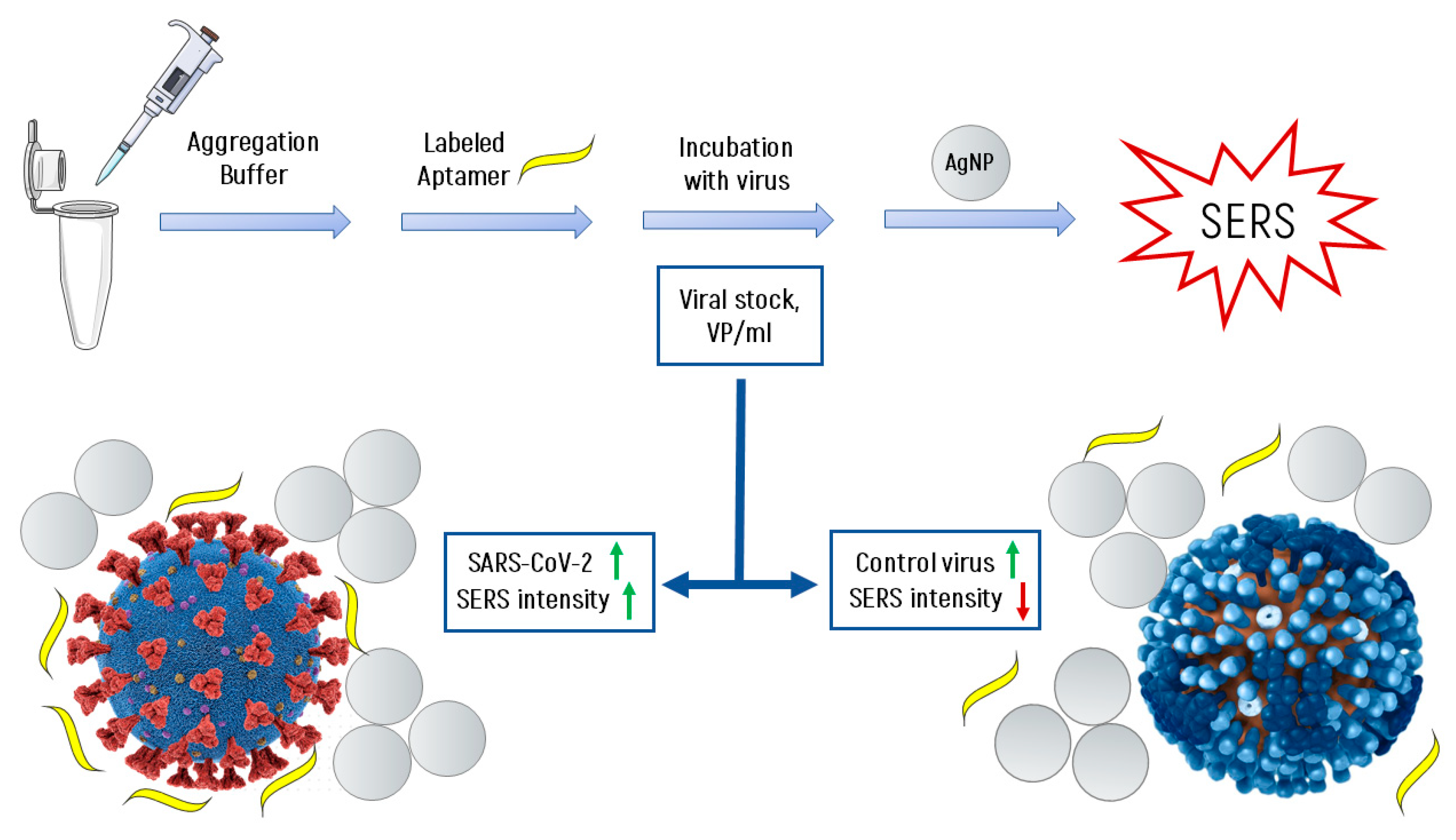
- ❖
- It is known [31,32] that at high ionic strength, silver nanoparticles interact well with the surface proteins of the virus, forming aggregates on the virus particles. With an increase in the concentration of proteins, the aggregates of silver nanoparticles create inhomogeneities of the electromagnetic field with a locally high density («hot spots») near their surface.
- ❖
- Due to significant differences in the number of the reporter molecules near non-specific viral particles and SARS-CoV-2, the SERS signal decreases with increasing concentration of the non-specific virus, and in the case of SARS-CoV-2, it increases.
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Johansson, M.A.; Quandelacy, T.M.; Kada, S.; Prasad, P.V.; Steele, M.; Brooks, J.T.; Slayton, R.B.; Biggerstaff, M.; Butler, J.C. SARS-CoV-2 Transmission from People Without COVID-19 Symptoms. JAMA Netw. Open. 2021, 4, e2035057. [Google Scholar] [CrossRef] [PubMed]
- Ye, F.; Xu, S.; Rong, Z.; Xu, R.; Liu, X.; Deng, P.; Liu, H.; Xu, X. Delivery of infection from asymptomatic carriers of COVID-19 in a familial cluster. Int. J. Infect. Dis. 2020, 94, 133–138. [Google Scholar] [CrossRef]
- Sharma, R.; Agarwal, M.; Gupta, M.; Somendra, S.; Saxena, S. Clinical Characteristics and Differential Clinical Diagnosis of Novel Coronavirus Disease 2019 (COVID-19). Coronavirus Disease 2019 (COVID-19). In Medical Virology: From Pathogenesis to Disease Control; Saxena, S., Ed.; Springer: Singapore, 2020; pp. 55–70. [Google Scholar] [CrossRef]
- Borst, A.; Box, A.T.A.; Fluit, A.C. False-Positive Results and Contamination in Nucleic Acid Amplification Assays: Suggestions for a Prevent and Destroy Strategy. Eur. J. Clin. Microbiol. Infect. Dis. 2004, 23, 289–299. [Google Scholar] [CrossRef] [PubMed]
- Fruehwirth, M.; Rivas, A.; Fitz, A.; Cechinel-Assing-Batista, A.; Silveira, C.; Delai, R. False-negative result in molecular diagnosis of SARS-CoV-2 in samples with amplification inhibitors. J. Bras. Patol. Med. Lab. 2020, 56, 1–8. [Google Scholar] [CrossRef]
- Conklin, S.E.; Martin, K.; Manabe, Y.C.; Schmidt, H.A.; Miller, J.; Keruly, M.; Klock, E.; Kirby, C.S.; Baker, O.R.; Fernandez, R.E.; et al. Evaluation of Serological SARS-CoV-2 Lateral Flow Assays for Rapid Point-of-Care Testing. J. Clin. Microbiol. 2021, 59, e02020-20. [Google Scholar] [CrossRef]
- Kilic, T.; Weissleder, R.; Lee, H. Molecular and Immunological Diagnostic Tests of COVID-19: Current Status and Challenges. iScience 2020, 23, 101406–101424. [Google Scholar] [CrossRef] [PubMed]
- OECD. Testing for COVID-19: How to Best Use the Various Tests? OECD Policy Responses to Coronavirus (COVID-19); OECD Publishing: Paris, France, 2020; Available online: https://www.oecd.org/coronavirus/policy-responses/testing-for-covid-19-how-to-best-use-the-various-tests-c76df201/ (accessed on 24 April 2021).
- Orooji, Y.; Sohrabi, H.; Hemmat, N.; Oroojalian, F.; Baradaran, B.; Mokhtarzadeh, A.; Mohaghegh, M.; Karimi-Maleh, H. An Overview on SARS-CoV-2 (COVID-19) and Other Human Coronaviruses and Their Detection Capability via Amplification Assay, Chemical Sensing, Biosensing, Immunosensing, and Clinical Assays. Nano-Micro Lett. 2021, 13, 18. [Google Scholar] [CrossRef]
- Li, M.; Wu, J.; Ma, M.; Feng, Z.; Mi, Z.; Rong, P.; Liu, D. Alkyne- and nitrile-anchored gold nanoparticles for multiplex SERS imaging of biomarkers in cancer cells and tissues. Nanotheranostics 2019, 3, 113–119. [Google Scholar] [CrossRef]
- Nishi, K.; Isobe, S.; Zhu, Y.; Kiyama, R. Fluorescence-based bioassays for the detection and evaluation of food materials. Sensors 2015, 15, 25831–25867. [Google Scholar] [CrossRef]
- Jiang, L.; Qian, J.; Cai, F.; He, S. Raman reporter-coated gold nanorods and their applications in multimodal optical imaging of cancer cells. Anal Bioanal Chem. 2011, 400, 2793–2800. [Google Scholar] [CrossRef]
- Mugo, S.M.; Zhang, Q. Nano-Sized Structured Platforms for Facile Solid-Phase Nanoextraction for Molecular Capture and (Bio)Chemical Analysis. In Micro and Nano Technologies, Nanomaterials Design for Sensing Applications; Zenkina, O.V., Ed.; Elsevier: Amsterdam, The Netherlands, 2019; pp. 153–195. [Google Scholar] [CrossRef]
- Egan, J.G.; Drossis, N.; Ebralidze, I.I.; Fruehwald, H.M.; Laschuk, N.O.; Poisson, J.; de Haan, H.W.; Zenkina, O.V. Hemoglobin-driven iron-directed assembly of gold nanoparticles. RSC Adv. 2018, 8, 15675–15686. [Google Scholar] [CrossRef]
- Kim, S.M.; Kim, J.; Noh, S.; Sohn, H.; Lee, T. Recent Development of Aptasensor for Influenza Virus Detection. BioChip J. 2020, 14, 327–339. [Google Scholar] [CrossRef] [PubMed]
- Negri, P.; Chen, G.; Kage, A.; Nitsche, A.; Naumann, D.; Xu, B.; Dluhy, R.A. Direct Optical Detection of Viral Nucleoprotein Binding to an Anti-Influenza Aptamer. Anal. Chem. 2012, 84, 5501–5508. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Park, S.-G.; Choi, N.; Moon, J.-I.; Dang, H.; Das, A.; Lee, S.; Kim, D.-G.; Chen, L.; Choo, J. SERS imaging-based aptasensor for ultrasensitive and reproducible detection of influenza virus A. Biosens. Bioelectron. 2020, 167, 112496. [Google Scholar] [CrossRef] [PubMed]
- Stanborough, T.; Given, F.M.; Koch, B.; Sheen, C.R.; Stowers-Hull, A.B.; Waterland, M.R.; Crittenden, D.L. Optical Detection of CoV-SARS-2 Viral Proteins to Sub-Picomolar Concentrations. ACS Omega 2021, 6, 6404–6413. [Google Scholar] [CrossRef]
- Yang, Y.; Peng, Y.; Lin, C.; Long, L.; Hu, J.; He, J.; Zeng, H.; Huang, Z.; Li, Z.Y.; Tanemura, M.; et al. Human ACE2-Functionalized Gold “Virus-Trap” Nanostructures for Accurate Capture of SARS-CoV-2 and Single-Virus SERS Detection. Nanomicro Lett. 2021, 13, 109. [Google Scholar] [CrossRef]
- Kukushkin, V.I.; Ivanov, N.M.; Novoseltseva, A.A.; Gambaryan, A.S.; Yaminsky, I.V.; Kopylov, A.M.; Zavyalova, E.G. Highly sensitive detection of influenza virus with SERS aptasensor. PLoS ONE. 2019, 14, e0216247. [Google Scholar] [CrossRef]
- Gribanyov, D.; Zhdanov, G.; Olenin, A.; Lisichkin, G.; Gambaryan, A.; Kukushkin, V.; Zavyalova, E. SERS-Based Colloidal Aptasensors for Quantitative Determination of Influenza Virus. Int. J. Mol. Sci. 2021, 22, 1842. [Google Scholar] [CrossRef]
- Song, Y.; Song, J.; Wei, X.; Huang, M.; Sun, M.; Zhu, L.; Yang, C. Discovery of Aptamers Targeting Receptor-Binding Domain of the SARS-CoV-2 Spike Glycoprotein. Anal. Chem. 2020, 92, 9895–9900. [Google Scholar] [CrossRef]
- Leopold, A.N.; Lendl, B. A New Method for Fast Preparation of Highly Surface-Enhanced Raman Scattering (SERS) Active Silver Colloids at Room Temperature by Reduction of Silver Nitrate with Hydroxylamine Hydrochloride. J. Phys. Chem. B. 2003, 107, 5723–5727. [Google Scholar] [CrossRef]
- Reed, L.J.; Muench, H. A Simple Method of Estimating Fifty Per Cent Endpoint. Am. J. Epidemiol. 1938, 27, 493–497. [Google Scholar] [CrossRef]
- SPR-Pages. Available online: https://www.sprpages.nl/ (accessed on 15 March 2021).
- Kramberger, P.; Ciringer, M.; Štrancar, A.; Peterka, M. Evaluation of nanoparticle tracking analysis for total virus particle determination. Virol. J. 2012, 9, 265. [Google Scholar] [CrossRef]
- The ViennaRNA Web Services. Available online: http://rna.tbi.univie.ac.at/ (accessed on 20 April 2021).
- Mergny, J.-L.; Lacroix, L. Analysis of Thermal Melting Curves. In Oligonucleotides; Mary Ann Liebert, Inc.: New Rochelle, NY, USA, 2003; Volume 13, pp. 515–537. [Google Scholar] [CrossRef]
- Kypr, J.; Kejnovska, I.; Renciuk, D.; Vorlickova, M. Circular dichroism and conformational polymorphism of DNA. Nucl. Acids Res. 2009, 37, 1713–1725. [Google Scholar] [CrossRef]
- Villar-Guerra, R.; Trent, J.O.; Chaires, J.B. G-Quadruplex Secondary Structure Obtained from Circular Dichroism Spectroscopy. Angew. Chem. Int. Ed. 2018, 57, 7171–7175. [Google Scholar] [CrossRef]
- Elechiguerra, J.L.; Burt, J.L.; Morones, J.R.; Camacho-Bragado, A.; Gao, X.; Lara, H.H.; Yacaman, M.J. Interaction of silver nanoparticles with HIV-1. J. Nanobiotechnol. 2005, 3, 6. [Google Scholar] [CrossRef]
- Sepunarua, L.; Plowmana, B.J.; Sokolova, S.V.; Young, N.P.; Comptona, R.G. Rapid Electrochemical Detection of Single Influenza Viruses Tagged with Silver Nanoparticles. Chem. Sci. 2016, 7, 3892–3899. [Google Scholar] [CrossRef]
- Betz, J.F.; Yu, W.W.; Cheng, Y.; White, I.M.; Rubloff, G.W. Simple SERS substrates: Powerful, portable, and full of potential. Phys. Chem. Chem. Phys. 2014, 16, 2224–2239. [Google Scholar] [CrossRef]
- Kukushkin, V.I.; Astrakhantseva, A.S.; Morozova, E.N. Influence of the Morphology of Metal Nanoparticles Deposited on Surfaces of Silicon Oxide on the Optical Properties of SERS Substrates. Bull. Russ. Acad. Sci. Phys. 2021, 85, 133–140. [Google Scholar] [CrossRef]
- Kukushkin, V.I.; Grishina, Y.V.; Solov’ev, V.V.; Kukushkin, I.V. Size plasmon–polariton resonance and its contribution to the giant enhancement of the Raman scattering. JETP Lett. 2017, 105, 677–681. [Google Scholar] [CrossRef]
- Jesus Gonçalves Rubira, R.; Alessio Camacho, S.; Silva Martin, C.; Mejía-Salazar, J.R.; Reyes Gómez, F.; Rosa da Silva, R.; Novais de Oliveira, O., Jr.; Alessio, P.; José Leopoldo Constantino, C. Designing Silver Nanoparticles for Detecting Levodopa (3,4-Dihydroxyphenylalanine, L-Dopa) Using Surface-Enhanced Raman Scattering (SERS). Sensors 2020, 20, 15. [Google Scholar] [CrossRef]
- Ma, X.; Xumin, X.X.; Xia, Y.; Wang, Z. SERS Aptasensor for Salmonella typhimurium Detection based on Spiny Gold Nanoparticles. Food Control 2018, 84, 232–237. [Google Scholar] [CrossRef]
- Duana, N.; Shen, M.; Qi, S.; Wang, W.; Wu, S.; Wang, Z. A SERS aptasensor for simultaneous multiple pathogens detection using gold decorated PDMS substrate. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2020, 230, 118103. [Google Scholar] [CrossRef]
- Tu, D.; Garzaab, J.T.; Cote, G.L. A SERS aptasensor for sensitive and selective detection of bis(2-ethylhexyl)phthalate. RSC Adv. 2019, 9, 2618–2625. [Google Scholar] [CrossRef]
- Yang, L.; Fu, C.; Wang, H.; Xu, S.; Xu, W. Aptamer-based surface-enhanced Raman scattering (SERS) sensor for thrombin based on supramolecular recognition, oriented assembly, and local field coupling. Anal. Bioanal. Chem. 2017, 409, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Fedotova, Y.V.; Kukushkin, V.I.; Solovyev, V.V.; Kukushkin, I.V. Spoof plasmons enable giant Raman scattering enhancement in Near-Infrared region. Opt. Express 2019, 27, 32578–32586. [Google Scholar] [CrossRef]
- Petti, L.; Capasso, R.; Rippa, M.; Pannico, M.; La Manna, P.; Peluso, G.; Musto, P. A plasmonic nanostructure fabricated by electron beam lithography as a sensitive and highly homogeneous SERS substrate for bio-sensing applications. Vib. Spectrosc. 2016, 82, 22–30. [Google Scholar] [CrossRef]
- Mahmoud, A.Y.; Rusin, C.J.; McDermott, M.T. Gold nanostars as a colloidal substrate for in-solution SERS measurements using a handheld Raman spectrometer. Analyst 2020, 145, 1396–1407. [Google Scholar] [CrossRef] [PubMed]

| Aptamer | ka, M−1s−1 | kd, s−1 | Kd, M |
|---|---|---|---|
| RBD-1C | (1.7 ± 0.8)·105 | (2.17 ± 0.02)·10−3 | (1.3 ± 0.2)·10−8 |
| RBD-1C-sh | (1.7 ± 0.2)·104 | (5.6 ± 0.4)·10−3 | (3.3 ± 0.6)·10−7 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zavyalova, E.; Ambartsumyan, O.; Zhdanov, G.; Gribanyov, D.; Gushchin, V.; Tkachuk, A.; Rudakova, E.; Nikiforova, M.; Kuznetsova, N.; Popova, L.; et al. SERS-Based Aptasensor for Rapid Quantitative Detection of SARS-CoV-2. Nanomaterials 2021, 11, 1394. https://doi.org/10.3390/nano11061394
Zavyalova E, Ambartsumyan O, Zhdanov G, Gribanyov D, Gushchin V, Tkachuk A, Rudakova E, Nikiforova M, Kuznetsova N, Popova L, et al. SERS-Based Aptasensor for Rapid Quantitative Detection of SARS-CoV-2. Nanomaterials. 2021; 11(6):1394. https://doi.org/10.3390/nano11061394
Chicago/Turabian StyleZavyalova, Elena, Oganes Ambartsumyan, Gleb Zhdanov, Dmitry Gribanyov, Vladimir Gushchin, Artem Tkachuk, Elena Rudakova, Maria Nikiforova, Nadezhda Kuznetsova, Liubov Popova, and et al. 2021. "SERS-Based Aptasensor for Rapid Quantitative Detection of SARS-CoV-2" Nanomaterials 11, no. 6: 1394. https://doi.org/10.3390/nano11061394
APA StyleZavyalova, E., Ambartsumyan, O., Zhdanov, G., Gribanyov, D., Gushchin, V., Tkachuk, A., Rudakova, E., Nikiforova, M., Kuznetsova, N., Popova, L., Verdiev, B., Alatyrev, A., Burtseva, E., Ignatieva, A., Iliukhina, A., Dolzhikova, I., Arutyunyan, A., Gambaryan, A., & Kukushkin, V. (2021). SERS-Based Aptasensor for Rapid Quantitative Detection of SARS-CoV-2. Nanomaterials, 11(6), 1394. https://doi.org/10.3390/nano11061394







