Unveiling the Potential Role of Nanozymes in Combating the COVID-19 Outbreak
Abstract
1. Introduction
2. Overview of Previously Used Nanozymes in Various Antiviral Therapies
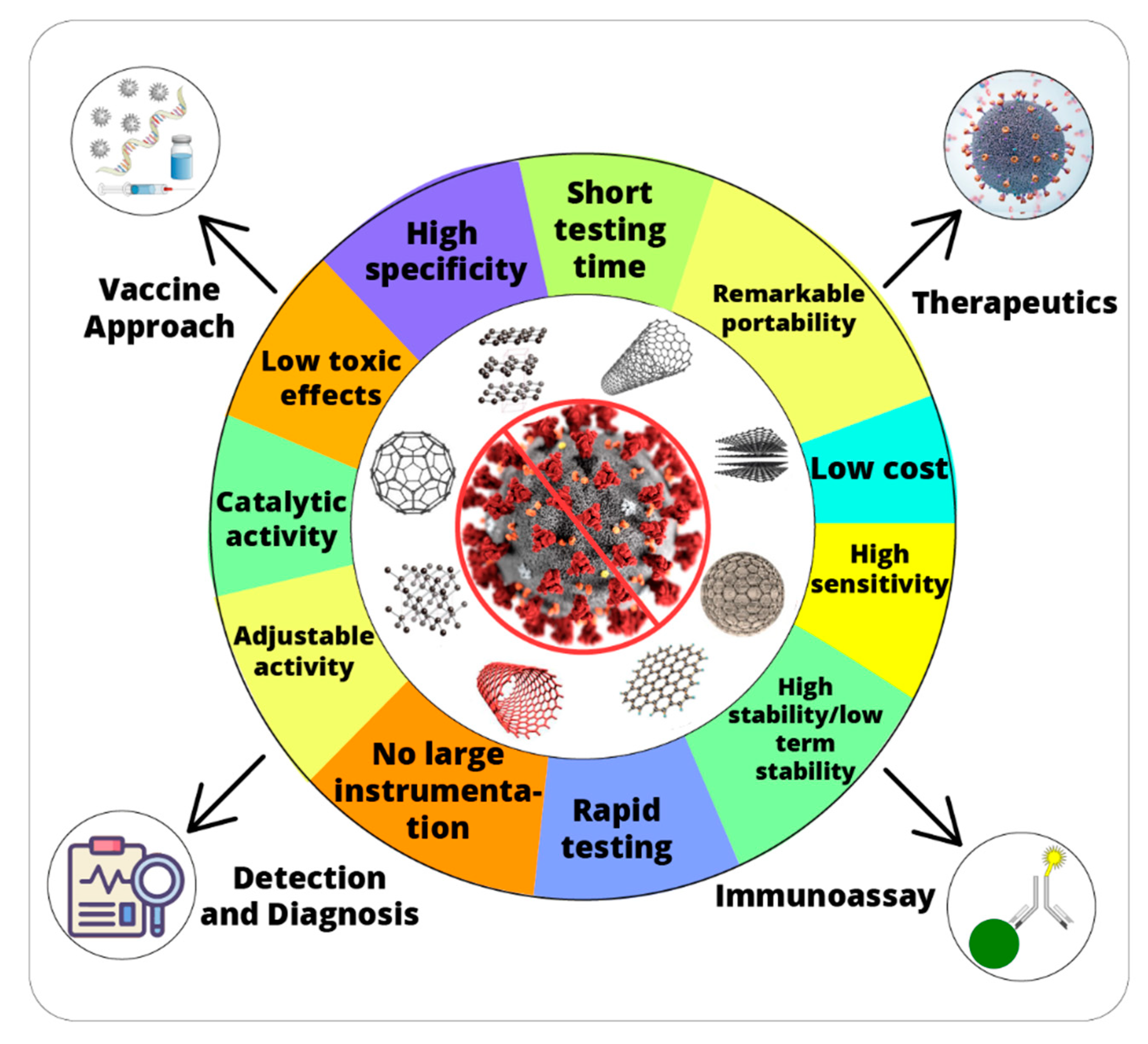
3. Contribution of Nanozymes for Combating Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2)
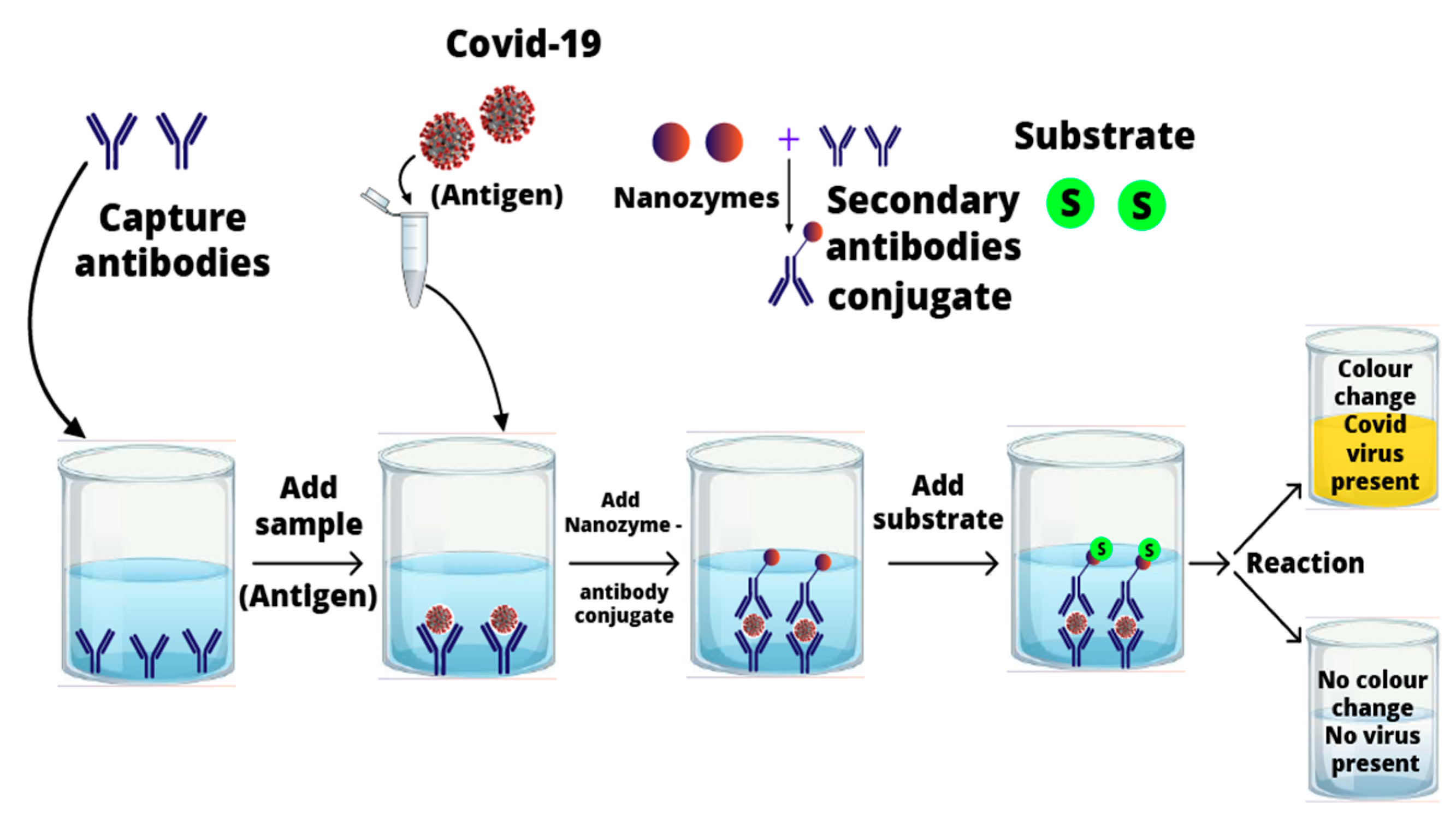
3.1. Nanozymes in the Detection of SARS-CoV-2
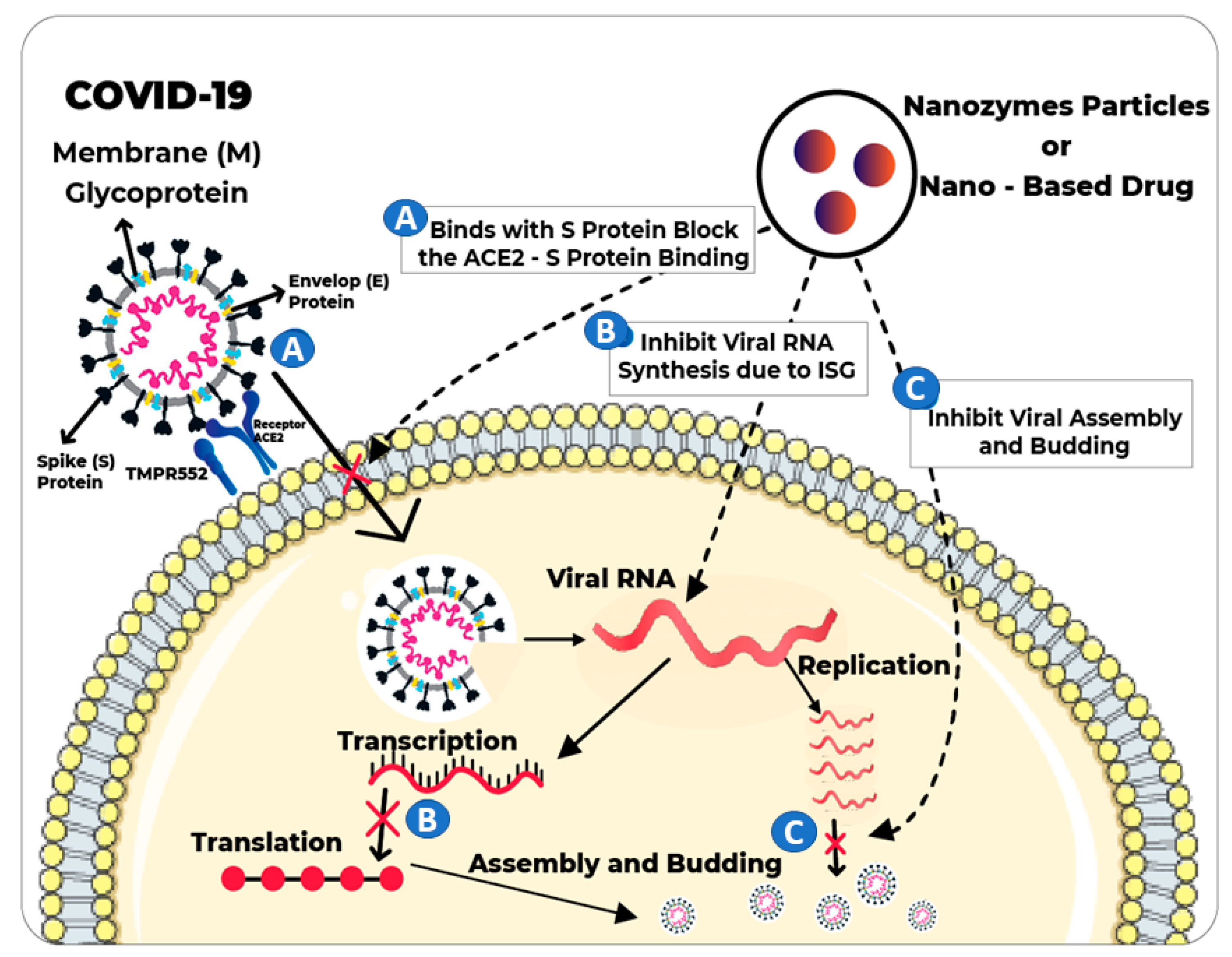
3.2. Nanozymes for Inactivation/Inhibition of SARS-CoV-2
3.2.1. Inhibition of Viral Entry
3.2.2. Blocking the Synthesis of Viral RNA and Viral Assembly
3.2.3. Vaccine Approach
4. Conclusions and Recommendations
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kilbourne, E.D. Influenza pandemics of the 20th century. Emerg. Infect. Dis. 2006, 12, 9–14. [Google Scholar] [CrossRef] [PubMed]
- Taubenberger, J.K. The origin and virulence of the 1918 “Spanish” influenza virus. Proc. Am. Philos. Soc. 2006, 150, 86–112. [Google Scholar] [PubMed]
- Greenberg, M.; Jacobziner, H.; Pakter, J.; Weisl, B.A. Maternal mortality in the epidemic of Asian influenza, New York City, 1957. Am. J. Obstet. Gynecol. 1958, 76, 897–902. [Google Scholar] [CrossRef]
- Rajagopal, S.; Treanor, J. Pandemic (avian) influenza. Semin. Respir. Crit. Care Med. 2007, 28, 159–170. [Google Scholar] [CrossRef] [PubMed]
- Meng, L.; Hua, F.; Bian, Z. Coronavirus Disease 2019 (COVID-19): Emerging and Future Challenges for Dental and Oral Medicine. J. Dent. Res. 2020, 99, 481–487. [Google Scholar] [CrossRef]
- Su Eun Park Epidemiology, virology, and clinical features of severe acute respiratory syndrome-coronavirus-2. Clin. Exp. Pediatr. 2020, 63, 119–124. [CrossRef] [PubMed]
- Harapan, H.; Itoh, N.; Yufika, A.; Winardi, W.; Keam, S.; Te, H.; Megawati, D.; Hayati, Z.; Wagner, A.L.; Mudatsir, M. Coronavirus disease 2019 (COVID-19): A literature review. J. Infect. Public Health 2020, 13, 667–673. [Google Scholar] [CrossRef]
- Li, G.; De Clercq, E. Therapeutic options for the 2019 novel coronavirus (2019-nCoV). Nat. Rev. Drug Discov. 2020, 19, 149–150. [Google Scholar] [CrossRef]
- Forman, R.; Shah, S.; Jeurissen, P.; Jit, M.; Mossialos, E. COVID-19 vaccine challenges: What have we learned so far and what remains to be done? Health Policy 2021, 125, 553–567. [Google Scholar] [CrossRef]
- Ledford, H. How ‘killer’ T cells could boost COVID immunity in face of new variants. Nature 2021, 2–7. [Google Scholar] [CrossRef]
- Samhan, F.A.; Stedtfeld, T.M.; Waseem, H.; Williams, M.R.; Stedtfeld, R.D.; Hashsham, S.A. On-filter direct amplification of Legionella pneumophila for rapid assessment of its abundance and viability. Water Res. 2017, 121. [Google Scholar] [CrossRef] [PubMed]
- Girum, T.; Lentiro, K.; Geremew, M.; Migora, B.; Shewamare, S. Global strategies and effectiveness for COVID-19 prevention through contact tracing, screening, quarantine, and isolation: A systematic review. Trop. Med. Health 2020, 48, 1–15. [Google Scholar] [CrossRef]
- Chakravarty, M.; Vora, A. Nanotechnology-based antiviral therapeutics. Drug Deliv. Transl. Res. 2020, 1–40. [Google Scholar] [CrossRef] [PubMed]
- Hou, T.; Zeng, W.; Yang, M.; Chen, W.; Ren, L.; Ai, J.; Wu, J.; Liao, Y.; Gou, X.; Li, Y.; et al. Development and evaluation of a rapid CRISPR-based diagnostic for COVID-19. PLoS Pathog. 2020, 16, e1008705. [Google Scholar] [CrossRef] [PubMed]
- Waseem, H.; Saleem ur Rehman, H.; Ali, J.; Iqbal, M.J.; Ali, M.I. Antibiotics and Antimicrobial Resistance Genes in the Environment; Elsevier: Amsterdam, The Netherlands, 2020; pp. 206–222. [Google Scholar]
- Munir, A.; Waseem, H.; Williams, M.R.; Stedtfeld, R.D.; Gulari, E.; Tiedje, J.M.; Hashsham, S.A. Modeling hybridization kinetics of gene probes in a DNA biochip using FEMLAB. Microarrays 2017, 6, 9. [Google Scholar] [CrossRef]
- Waseem, H.; Jameel, S.; Ali, J.; Ur Rehman, H.S.; Tauseef, I.; Farooq, U.; Jamal, A.; Ali, M.I. Contributions and challenges of high throughput qPCR for determining antimicrobial resistance in the environment: A critical review. Molecules 2019, 24, 163. [Google Scholar] [CrossRef]
- Yu, C.Y.; Chan, K.G.; Yean, C.Y.; Ang, G.Y. Nucleic Acid-Based Diagnostic Tests for the Detection SARS-CoV-2: An Update. Diagnostics 2021, 11, 53. [Google Scholar] [CrossRef]
- De Gasparo, R.; Pedotti, M.; Simonelli, L.; Nickl, P.; Muecksch, F.; Cassaniti, I.; Percivalle, E.; Lorenzi, J.C.C.; Mazzola, F.; Magrì, D.; et al. Bispecific IgG neutralizes SARS-CoV-2 variants and prevents escape in mice. Nature 2021. [Google Scholar] [CrossRef]
- Ayove, T.; Houniei, W.; Wangnapi, R.; Bieb, S.V.; Kazadi, W.; Luke, L.-N.; Manineng, C.; Moses, P.; Paru, R.; Esfandiari, J.; et al. Sensitivity and specificity of a rapid point-of-care test for active yaws: A comparative study. Lancet Glob. Heal. 2014, 2, e415–e421. [Google Scholar] [CrossRef]
- Waseem, H.; Williams, M.R.; Stedtfeld, T.; Chai, B.; Stedtfeld, R.D.; Cole, J.R.; Tiedje, J.M.; Hashsham, S.A. Virulence factor activity relationships (VFARs): A bioinformatics perspective. Environ. Sci. Process. Impacts 2017, 19, 247–260. [Google Scholar] [CrossRef]
- Waseem, H.; Ali, J.; Syed, J.H.; Jones, K.C. Establishing the relationship between molecular biomarkers and biotransformation rates: Extension of knowledge for dechlorination of polychlorinated dibenzo-p-dioxins and furans (PCDD/Fs). Environ. Pollut. 2020, 263, 114676. [Google Scholar] [CrossRef]
- Bilal, M.; Barceló, D.; Iqbal, H.M.N. Nanostructured materials for harnessing the power of horseradish peroxidase for tailored environmental applications. Sci. Total Env. 2020, 749, 142360. [Google Scholar] [CrossRef] [PubMed]
- Shin, H.Y.; Park, T.J.; Kim, M. Il Recent Research Trends and Future Prospects in Nanozymes. J. Nanomater. 2015, 2015, 756278. [Google Scholar] [CrossRef]
- Poon, L.L.M.; Chan, K.H.; Wong, O.K.; Yam, W.C.; Yuen, K.Y.; Guan, Y.; Lo, Y.M.D.; Peiris, J.S.M. Early diagnosis of SARS Coronavirus infection by real time RT-PCR. J. Clin. Virol. 2003, 28, 233–238. [Google Scholar] [CrossRef] [PubMed]
- Vogels, C.B.F.; Brackney, D.E.; Wang, J.; Kalinich, C.C.; Ott, I.M.; Kudo, E.; Lu, P.; Venkataraman, A.; Tokuyama, M.; Moore, A.J.; et al. SalivaDirect: Simple and sensitive molecular diagnostic test for SARS-CoV-2 surveillance. medRxiv 2020. [Google Scholar] [CrossRef]
- Weissleder, R.; Lee, H.; Ko, J.; Pittet, M.J. COVID-19 diagnostics in context. Sci. Transl. Med. 2020, 12. [Google Scholar] [CrossRef]
- Suo, T.; Liu, X.; Feng, J.; Guo, M.; Hu, W.; Guo, D.; Ullah, H.; Yang, Y.; Zhang, Q.; Wang, X.; et al. ddPCR: A more accurate tool for SARS-CoV-2 detection in low viral load specimens. Emerg. Microbes Infect. 2020, 9, 1259–1268. [Google Scholar] [CrossRef]
- Dao Thi, V.L.; Herbst, K.; Boerner, K.; Meurer, M.; Kremer, L.P.M.; Kirrmaier, D.; Freistaedter, A.; Papagiannidis, D.; Galmozzi, C.; Stanifer, M.L.; et al. A colorimetric RT-LAMP assay and LAMP-sequencing for detecting SARS-CoV-2 RNA in clinical samples. Sci. Transl. Med. 2020, 12, eabc7075. [Google Scholar] [CrossRef]
- Flynn, M.J.; Snitser, O.; Flynn, J.; Green, S.; Yelin, I.; Szwarcwort-Cohen, M.; Kishony, R.; Elowitz, M.B. A simple direct RT-LAMP SARS-CoV-2 saliva diagnostic. medRxiv 2020. [Google Scholar] [CrossRef]
- Xia, S.; Chen, X. Single-copy sensitive, field-deployable, and simultaneous dual-gene detection of SARS-CoV-2 RNA via modified RT-RPA. Cell Discov. 2020, 6, 37. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Yu, L.; Liu, C.; Ye, S.; Chen, W.; Li, D.; Huang, W. One-tube SARS-CoV-2 detection platform based on RT-RPA and CRISPR/Cas12a. J. Transl. Med. 2021, 19, 1–10. [Google Scholar] [CrossRef]
- El Wahed, A.A.; Patel, P.; Maier, M.; Pietsch, C.; Rüster, D.; Böhlken-Fascher, S.; Kissenkötter, J.; Behrmann, O.; Frimpong, M.; Diagne, M.M.; et al. Suitcase Lab for Rapid Detection of SARS-CoV-2 Based on Recombinase Polymerase Amplification Assay. Anal. Chem. 2021, 93, 2627–2634. [Google Scholar] [CrossRef] [PubMed]
- Daher, R.K.; Stewart, G.; Boissinot, M.; Bergeron, M.G. Recombinase polymerase amplification for diagnostic applications. Clin. Chem. 2016, 62, 947–958. [Google Scholar] [CrossRef]
- Chaibun, T.; Puenpa, J.; Ngamdee, T.; Boonapatcharoen, N.; Athamanolap, P.; O’Mullane, A.P.; Vongpunsawad, S.; Poovorawan, Y.; Lee, S.Y.; Lertanantawong, B. Rapid electrochemical detection of coronavirus SARS-CoV-2. Nat. Commun. 2021, 12, 802. [Google Scholar] [CrossRef]
- De Paz, H.D.; Brotons, P.; Muñoz-Almagro, C. Molecular isothermal techniques for combating infectious diseases: Towards low-cost point-of-care diagnostics. Expert Rev. Mol. Diagn. 2014, 14, 827–843. [Google Scholar] [CrossRef] [PubMed]
- Broughton, J.P.; Deng, X.; Yu, G.; Fasching, C.L.; Servellita, V.; Singh, J.; Miao, X.; Streithorst, J.A.; Granados, A.; Sotomayor-Gonzalez, A.; et al. CRISPR–Cas12-based detection of SARS-CoV-2. Nat. Biotechnol. 2020, 38, 870–874. [Google Scholar] [CrossRef] [PubMed]
- Taylor, M. CRISPR-Based COVID Test Is Rapid, Accurate and Costs Less Than $1. Available online: https://www.laboratoryequipment.com/567702-CRISPR-based-COVID-Test-is-Rapid-Accurate-and-Costs-Less-Than-1/ (accessed on 21 March 2021).
- Yip, C.C.; Sridhar, S.; Leung, K.-H.; Ng, A.C.; Chan, K.-H.; Chan, J.F.; Tsang, O.T.; Hung, I.F.; Cheng, V.C.; Yuen, K.-Y.; et al. Development and Evaluation of Novel and Highly Sensitive Single-Tube Nested Real-Time RT-PCR Assays for SARS-CoV-2 Detection. Int. J. Mol. Sci. 2020, 21, 5674. [Google Scholar] [CrossRef]
- Chan, J.F.-W.; Yip, C.C.-Y.; To, K.K.-W.; Tang, T.H.-C.; Wong, S.C.-Y.; Leung, K.-H.; Fung, A.Y.-F.; Ng, A.C.-K.; Zou, Z.; Tsoi, H.-W.; et al. Improved molecular diagnosis of COVID-19 by the novel, highly sensitive and specific COVID-19-RdRp/Hel real-time reverse transcription-PCR assay validated in vitro and with clinical specimens. J. Clin. Microbiol. 2020, 58, 5. [Google Scholar] [CrossRef]
- Brandsma, E.; Verhagen, H.J.M.P.; van de Laar, T.J.W.; Claas, E.C.J.; Cornelissen, M.; van den Akker, E. Rapid, Sensitive, and Specific Severe Acute Respiratory Syndrome Coronavirus 2 Detection: A Multicenter Comparison Between Standard Quantitative Reverse-Transcriptase Polymerase Chain Reaction and CRISPR-Based DETECTR. J. Infect. Dis. 2021, 223, 206–213. [Google Scholar] [CrossRef]
- Mercier-Delarue, S.; Vray, M.; Plantier, J.C.; Maillard, T.; Adjout, Z.; de Olivera, F.; Schnepf, N.; Maylin, S.; Simon, F.; Delaugerre, C. Higher Specificity of Nucleic Acid Sequence-Based Amplification Isothermal Technology than of Real-Time PCR for Quantification of HIV-1 RNA on Dried Blood Spots. J. Clin. Microbiol. 2014, 52, 52–56. [Google Scholar] [CrossRef][Green Version]
- Wu, Q.; Suo, C.; Brown, T.; Wang, T.; Teichmann, S.A.; Bassett, A.R. INSIGHT: A population-scale COVID-19 testing strategy combining point-of-care diagnosis with centralized high-throughput sequencing. Sci. Adv. 2021, 7. [Google Scholar] [CrossRef]
- Kilic, T.; Weissleder, R.; Lee, H. Molecular and Immunological Diagnostic Tests of COVID-19: Current Status and Challenges. iScience 2020, 23, 101406. [Google Scholar] [CrossRef]
- Elisa Test for Covid-19, We Clear 10 Of The Most Common Doubts. Available online: https://lifelength.com/elisa-test-for-covid-19-we-clear-10-of-the-most-common-doubts/ (accessed on 21 March 2021).
- Michel, M.; Bouam, A.; Edouard, S.; Fenollar, F.; Di Pinto, F.; Mège, J.L.; Drancourt, M.; Vitte, J. Evaluating ELISA, Immunofluorescence, and Lateral Flow Assay for SARS-CoV-2 Serologic Assays. Front. Microbiol. 2020, 11, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Edouard, S.; Colson, P.; Melenotte, C.; Di Pinto, F.; Thomas, L.; La Scola, B.; Million, M.; Tissot-Dupont, H.; Gautret, P.; Stein, A.; et al. Evaluating the serological status of COVID-19 patients using an indirect immunofluorescent assay, France. Eur. J. Clin. Microbiol. Infect. Dis. 2021, 40, 361–371. [Google Scholar] [CrossRef] [PubMed]
- Pinto, D.; Park, Y.-J.; Beltramello, M.; Walls, A.C.; Tortorici, M.A.; Bianchi, S.; Jaconi, S.; Culap, K.; Zatta, F.; De Marco, A.; et al. Cross-neutralization of SARS-CoV-2 by a human monoclonal SARS-CoV antibody. Nature 2020, 583, 290–295. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Tang, B.; Wang, Y.; Gu, B. Since January 2020 Elsevier has created a COVID-19 resource centre with free information in English and Mandarin on the novel coronavirus COVID- 19. The COVID-19 resource centre is hosted on Elsevier Connect, the company’s public news and information. Clin. Chim. Acta 2020, 510, 35–46. [Google Scholar] [CrossRef] [PubMed]
- Fenollar, F.; Bouam, A.; Ballouche, M.; Fuster, L.; Prudent, E.; Colson, P.; Tissot-Dupont, H.; Million, M.; Drancourt, M.; Raoult, D.; et al. Evaluation of the panbio COVID-19 rapid antigen detection test device for the screening of patients with COVID-19. J. Clin. Microbiol. 2021, 59, 19–21. [Google Scholar] [CrossRef]
- Eshghifar, N.; Busheri, A.; Shrestha, R.; Beqaj, S. Evaluation of analytical performance of seven rapid antigen detection kits for detection of SARS-CoV-2 virus. Int. J. Gen. Med. 2021, 14, 435–440. [Google Scholar] [CrossRef]
- Abbott PanbioTM: COVID-19 Ag Rapid Test Device 41FK10. Available online: https://www.meteka.com/wp-content/uploads/2020/10/Testanleitung_Abbott.pdf (accessed on 15 March 2021).
- Grossberg, A.N.; Koza, L.A.; Ledreux, A.; Prusmack, C.; Krishnamurthy, H.K.; Jayaraman, V.; Granholm, A.C.; Linseman, D.A. A multiplex chemiluminescent immunoassay for serological profiling of COVID-19-positive symptomatic and asymptomatic patients. Nat. Commun. 2021, 12. [Google Scholar] [CrossRef]
- Mendoza, R.; Silver, M.; Zuretti, A.R.; Christian, M.; Das, B.; Norin, A.J.; Borgen, P.; Libien, J.; Bluth, M.H. Correlation of Automated Chemiluminescent Method with Enzyme-Linked Immunosorbent Assay (ELISA) Antibody Titers in Convalescent COVID-19 Plasma Samples: Development of Rapid, Cost-Effective Semi-Quantitative Diagnostic Methods. J. Blood Med. 2021, 12, 157–164. [Google Scholar] [CrossRef]
- He, Q.; Chong, K.H.; Chng, H.H.; Leung, B.; Ling, A.E.; Wei, T.; Chan, S.W.; Ooi, E.E.; Kwang, J. Development of a Western Blot Assay for Detection of Antibodies against Coronavirus Causing Severe Acute Respiratory Syndrome. Clin. Diagn. Lab. Immunol. 2004, 11, 417–422. [Google Scholar] [CrossRef]
- Wu, J.-L.; Tseng, W.-P.; Lin, C.-H.; Lee, T.-F.; Chung, M.-Y.; Huang, C.-H.; Chen, S.-Y.; Hsueh, P.-R.; Chen, S.-C. Four point-of-care lateral flow immunoassays for diagnosis of COVID-19 and for assessing dynamics of antibody responses to SARS-CoV-2. J. Infect. 2020, 81, 435–442. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-moncayo, R.; Cedillo-alcantar, D.F.; Guevara-pantoja, P.E.; Chavez-pineda, O.G.; Hernandez-ortiz, J.A.; Amador-hernandez, J.U.; Rojas-velasco, G.; Sanchez-muñoz, F.; Manzur-sandoval, D.; Patino-lopez, L.D.; et al. A high-throughput multiplexed microfluidic device for COVID-19 serology assays. Lab Chip 2021, 21, 93–104. [Google Scholar] [CrossRef] [PubMed]
- Benzigar, M.R.; Bhattacharjee, R.; Baharfar, M.; Liu, G. Current methods for diagnosis of human coronaviruses: Pros and cons. Anal. Bioanal. Chem. 2021, 413, 2311–2330. [Google Scholar] [CrossRef] [PubMed]
- Pohanka, M. Point-of-Care Diagnoses and Assays Based on Lateral Flow Test. Int. J. Anal. Chem. 2021. [Google Scholar] [CrossRef]
- EUA Authorized Serology Test Performance. Available online: https://www.fda.gov/medical-devices/coronavirus-disease-2019-covid-19-emergency-use-authorizations-medical-devices/eua-authorized-serology-test-performance (accessed on 15 March 2021).
- Liang, Y.; Yan, H.; Huang, L.; Zhao, J.; Wang, H.; Kang, M.; Wan, Z.; Shui, J.; Tang, S. A luciferase immunosorbent assay for quantitative detection of IgG antibodies against SARS-CoV-2 nucleoprotein. J. Virol. Methods 2021, 292, 114141. [Google Scholar] [CrossRef]
- Zhang, C.; Zhou, L.; Liu, H.; Zhang, S.; Tian, Y.; Huo, J.; Zhang, Y.; Wei, B.; Xu, D.; Hu, J.; et al. Establishing a high sensitivity detection method for SARS-CoV-2 IgM / IgG and developing a clinical application of this method. Emerg. Microbes Infect. 2020, 9, 2020–2029. [Google Scholar] [CrossRef]
- Celltrion to Launch Both Antigen and Antibody Testing Kits in the U.S. Available online: https://www.pharmasalmanac.com/articles/celltrion-to-launch-both-antigen-and-antibody-testing-kits-in-the-u.s. (accessed on 15 March 2021).
- Kevadiya, B.D.; Machhi, J.; Herskovitz, J.; Oleynikov, M.D.; Blomberg, W.R.; Bajwa, N.; Soni, D.; Das, S.; Hasan, M.; Patel, M.; et al. Diagnostics for SARS-CoV-2 infections. Nat. Mater. 2021, 20, 593–605. [Google Scholar] [CrossRef]
- Imai, K.; Tabata, S.; Ikeda, M.; Noguchi, S.; Kitagawa, Y.; Matuoka, M.; Miyoshi, K.; Tarumoto, N.; Sakai, J.; Ito, T.; et al. Clinical evaluation of an immunochromatographic IgM/IgG antibody assay and chest computed tomography for the diagnosis of COVID-19. J. Clin. Virol. 2020, 128, 104393. [Google Scholar] [CrossRef]
- Ali, J.; Wang, L.; Waseem, H.; Djellabi, R.; Oladoja, N.A.; Pan, G. FeS@rGO nanocomposites as electrocatalysts for enhanced chromium removal and clean energy generation by microbial fuel cell. Chem. Eng. J. 2020, 384, 123335. [Google Scholar] [CrossRef]
- Ali, J.; Ali, N.; Wang, L.; Waseem, H.; Pan, G. Revisiting the mechanistic pathways for bacterial mediated synthesis of noble metal nanoparticles. J. Microbiol. Methods 2019, 159, 18–25. [Google Scholar] [CrossRef] [PubMed]
- Ali, J.; Wang, L.; Waseem, H.; Sharif, H.M.A.; Djellabi, R.; Zhang, C.; Pan, G. Bioelectrochemical recovery of silver from wastewater with sustainable power generation and its reuse for biofouling mitigation. J. Clean. Prod. 2019, 235, 1425–1437. [Google Scholar] [CrossRef]
- Djellabi, R.; Ali, J.; Zhao, X.; Saber, A.N.; Yang, B. CuO NPs incorporated into electron-rich TCTA@PVP photoactive polymer for the photocatalytic oxidation of dyes and bacteria inactivation. J. Water Process Eng. 2020, 36, 101238. [Google Scholar] [CrossRef]
- Nguyen, Q.H.; Kim, M. Il Trends in Analytical Chemistry Nanomaterial-mediated paper-based biosensors for colorimetric pathogen detection. Trends Anal. Chem. 2020, 132, 116038. [Google Scholar] [CrossRef]
- Liu, D.; Ju, C.; Han, C.; Shi, R.; Chen, X.; Duan, D. Ultra-sensitive nanozyme-based chemiluminescence paper test for rapid diagnosis of SARS-CoV-2 infection. bioRxiv 2020. [Google Scholar] [CrossRef]
- Duan, D.; Fan, K.; Zhang, D.; Tan, S.; Liang, M.; Liu, Y.; Zhang, J.; Zhang, P.; Liu, W.; Qiu, X.; et al. Biosensors and Bioelectronics Nanozyme-strip for rapid local diagnosis of Ebola. Biosens. Bioelectron. 2015, 74, 134–141. [Google Scholar] [CrossRef] [PubMed]
- Oh, S.; Kim, J.; Tran, V.T.; Lee, D.K.; Ahmed, S.R.; Hong, J.C.; Lee, J.; Park, E.Y.; Lee, J. Magnetic Nanozyme-Linked Immunosorbent Assay for Ultrasensitive Influenza A Virus Detection. ACS Appl. Mater. Interfaces 2018, 10, 12534–12543. [Google Scholar] [CrossRef] [PubMed]
- Long, L.; Cai, R.; Liu, J.; Wu, X. A Novel Nanoprobe Based on Core–Shell Au@ Pt@ Mesoporous SiO2 Nanozyme With Enhanced Activity and Stability for Mumps Virus Diagnosis. Front. Chem. 2020, 8, 1–9. [Google Scholar] [CrossRef]
- Goodacre, R.; Behera, B.K.; Bansal, V. Ultrasensitive Colorimetric Detection of Murine Norovirus Using NanoZyme Aptasensor. Anal. Chem. 2019, 91, 3270–3276. [Google Scholar] [CrossRef]
- Zhao, Y.; Ding, B.; Xiao, X.; Jiang, F.; Wang, M.; Hou, Z.; Xing, B.; Teng, B.; Cheng, Z.; Ma, P.; et al. Virus-Like Fe3O4@ Bi2S3 Nanozymes with Resistance-Free Apoptotic Hyperthermia-Augmented Nanozymitic Activity for Enhanced Synergetic Cancer Therapy. ACS Appl. Mater. Interfaces 2020, 12, 11320–11328. [Google Scholar] [CrossRef]
- Qin, T.; Ma, R.; Yin, Y.; Miao, X.; Chen, S.; Fan, K.; Xi, J.; Liu, Q. Catalytic inactivation of influenza virus by iron oxide nanozyme. Theranostics 2019, 23, 6920–6935. [Google Scholar] [CrossRef] [PubMed]
- Rana, S.; Nazar, U.; Ali, J.; ul Ain Ali, Q.; Ahmad, N.M.; Sarwar, F.; Waseem, H.; Jamil, S.U.U. Improved antifouling potential of polyether sulfone polymeric membrane containing silver nanoparticles: Self-cleaning membranes. Environ. Technol. 2018, 39, 1413–1421. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-hitzky, E.; Darder, M.; Wicklein, B.; Ruiz-garcia, C.; Martín-sampedro, R.; Real, G.; Aranda, P. Nanotechnology Responses to COVID-19. Adv. Healthc. Mater. 2020, 2, 1–26. [Google Scholar] [CrossRef] [PubMed]
- Chauhan, G.; Madou, M.J.; Kalra, S.; Chopra, V.; Ghosh, D.; Martinez-Chapa, S.O. Nanotechnology for COVID-19: Therapeutics and Vaccine Research. ACS Nano 2020, 14, 7760–7782. [Google Scholar] [CrossRef] [PubMed]
- Niu, X.; Cheng, N.; Ruan, X.; Du, D.; Lin, Y. Review—Nanozyme-Based Immunosensors and Immunoassays: Recent Developments and Future Trends Review. J. Electrochem. Soc. 2020, 167, 037508. [Google Scholar] [CrossRef]
- Bu, T.; Huang, Q.; Yan, L.; Huang, L.; Zhang, M.; Yang, Q. SC. Ultra technically-simple and sensitive detection for Salmonella Enteritidis by immunochromatographic assay based on gold growth. Food Control 2017, 84, 536–543. [Google Scholar] [CrossRef]
- Liu, R.; He, L.; Hu, Y.; Luo, Z.; Zhang, J. A serological aptamer-assisted proximity ligation assay for COVID-19 diagnosis and seeking neutralizing aptamers. Chem. Sci. 2020, 11, 12157–12164. [Google Scholar] [CrossRef]
- Yoo, S.M.; Kim, D.; Lee, S.Y. Aptamer-functionalized localized surface plasmon resonance sensor for the multiplexed detection of different bacterial species. Talanta 2015, 132, 112–117. [Google Scholar] [CrossRef]
- Iqbal, H.; Parra, R. The Emergence of Novel-Coronavirus and its Replication Cycle-An Overview. J. Pure Appl. Microbiol. 2020, 14, 13–16. [Google Scholar] [CrossRef]
- Wang, Q.; Zhang, Y.; Wu, L.; Niu, S.; Song, C.; Zhang, Z.; Lu, G.; Qiao, C.; Hu, Y.; Yuen, K.Y.; et al. Structural and Functional Basis of SARS-CoV-2 Entry by Using Human ACE2 ll ll Article Structural and Functional Basis of SARS-CoV-2 Entry by Using Human ACE2. Cell 2020, 181, 894–904. [Google Scholar] [CrossRef]
- Palestino, G.; García-Silva, I.; González-Ortega, O.; Rosales-Mendoza, S. Can nanotechnology help in the fight against COVID-19? Expert Rev. Anti. Infect. Ther. 2020, 18, 849–864. [Google Scholar] [CrossRef] [PubMed]
- Iannazzo, D.; Pistone, A.; Ferro, S.; De Luca, L.; Monforte, A.M.; Romeo, R.; Buemi, M.R.; Pannecouque, C. Graphene Quantum Dots Based Systems as HIV Inhibitors. Bioconjug. Chem. 2018, 29, 3084–3093. [Google Scholar] [CrossRef] [PubMed]
- Medhi, R.; Srinoi, P.; Ngo, N.; Tran, H.; Lee, T.R. Nanoparticle-Based Strategies to Combat COVID-19. ACS Appl. Nano Mater. 2020, 9, 8557–8580. [Google Scholar] [CrossRef]
- Du, T.; Liang, J.; Dong, N.; Lu, J.; Fu, Y.; Fang, L.; Xiao, S.; Han, H. Glutathione-Capped Ag2S Nanoclusters Inhibit Coronavirus Proliferation through Blockage of Viral RNA Synthesis and Budding. ACS Appl. Mater. Interfaces 2018, 10, 4369–4378. [Google Scholar] [CrossRef]
- Sun, H.; Zhou, Y.; Ren, J.; Qu, X. Carbon Nanozymes: Enzymatic Properties, Catalytic Mechanism, and Applications. Angew. Chemie Int. Ed. 2018, 57, 9224–9237. [Google Scholar] [CrossRef] [PubMed]
- Du, T.; Liang, J.; Dong, N.; Liu, L.; Fang, L.; Xiao, S.; Han, H. Carbon dots as inhibitors of virus by activation of type I interferon response. Carbon 2016, 110, 278–285. [Google Scholar] [CrossRef]
- Cagno, V.; Andreozzi, P.; D’Alicarnasso, M.; Jacob Silva, P.; Mueller, M.; Galloux, M.; Le Goffic, R.; Jones, S.T.; Vallino, M.; Hodek, J.; et al. Broad-spectrum non-toxic antiviral nanoparticles with a virucidal inhibition mechanism. Nat. Mater. 2018, 17, 195–203. [Google Scholar] [CrossRef]
- Mackay, F.; Figgett, W.A.; Saulep, D.; Lepage, M.; Hibbs, M.L. B-cell stage and context-dependent requirements for survival signals from BAFF and the B-cell receptor. Immunol. Rev. 2010, 237, 205–225. [Google Scholar] [CrossRef]
- Bettini, E.; Locci, M. SARS-CoV-2 mRNA Vaccines: Immunological mechanism and beyond. Vaccines 2021, 9, 147. [Google Scholar] [CrossRef]
- Jindal, A.; Sarkar, S.; Alam, A. Nanomaterials-Mediated Immunomodulation for Cancer Therapeutics. Front. Chem. 2021, 9, 46. [Google Scholar] [CrossRef]
- Patterson, D.P.; Rynda-Apple, A.; Harmsen, A.L.; Harmsen, A.G.; Douglas, T. Biomimetic antigenic nanoparticles elicit controlled protective immune response to influenza. ACS Nano 2013, 7, 3036–3044. [Google Scholar] [CrossRef] [PubMed]
- Waseem, H.; Williams, M.R.; Jameel, S.; Hashsham, S.A. Antimicrobial Resistance in the Environment. Water Environ. Res. 2018, 90, 865–884. [Google Scholar] [CrossRef] [PubMed]
- Waseem, H.; Ali, J.; Jamal, A.; Ali, M.I. Potential Dissemination of Antimicrobial Resistance from Small Scale Poultry Slaughterhouses in Pakistan. Appl. Ecol. Environ. Res. 2019, 17, 3049–3063. [Google Scholar] [CrossRef]
- Waseem, H.; Williams, M.R.; Stedtfeld, R.D.; Stedtfeld, T.M.; Shanker, R.; Hashsham, S.A. Chapter 8: Organ-on-chip Systems: An Emerging Platform for Toxicity Screening of Chemicals, Pharmaceuticals, and Nanomaterials. Nanotoxicology: Experimental and Computational Perspectives; Dhawan, A., Anderson, D., Shanker, R., Eds.; Royal Society of Chemistry: Cambridge, UK, 2018; pp. 203–231. [Google Scholar]
| Sr.No. | Technique | Sensitivity | Specificity | Limit of Detection | Cost in USD | Processing Time | Sample | Detection Element (Gene/Antibodies) | Reference |
|---|---|---|---|---|---|---|---|---|---|
| Nucleic Acid-Based Techniques | |||||||||
| 1 | RT-qPCR | 80% | 100% | 0.0278 copies/µL | 1.29–4.37 | 2–8 h; >12 h | Nasopharyngeal aspirate | N gene | [25,26,27] |
| 2 | Droplet Digital PCR (ddPCR) | 83–99% | 48–100% | 5 × 103 copies/µL | 100 | 2–4 h | Throat swab samples | ORF1ab and N gene | [27,28] |
| 3 | RT-LAMP | 97.50% | 99.70% | 1 × 102 copies/µL | ∼7 | 1 h | Pharyngeal swab | N gene | [29,30] |
| 4 | Recombinase Polymerase Amplification (RT-RPA) | 65–94% | 77–100% | 0.05 copies/µL | 4.3 | 15–20 min | Swab Samples | N gene, RdRp, E Gene | [31,32,33,34] |
| 5 | RCA-POC | 99% | 99% | 1 copy/µL | 4–10 | <2 h | Nasopharyngeal swab | N gene and S gene | [35,36] |
| 6 | CRISPR | 100% | 100% | 10 copies/µL | 3.50 | 30–40 min | Oropharyngeal swab and Nasopharyngeal swab | E (envelope) and N (nucleoprotein) genes | [37,38] |
| 7 | RdRp/Hel RT-PCR | 100% | 100% | 0.56 copies/µL | NA | <1 h | Respiratory and non-respiratory tract specimens | RdRp/Hel, spike (S), (N), genes | [39,40] |
| 8 | DETECTR (DNA Endonuclease-Targeted CRISPR Trans Reporter) | 95% | 100% | 10 copies/μL | ~0.2 | 40 min | Pharyngeal swab | E/N | [32,41] |
| 9 | Nucleic Acid Sequence-Based Amplification (NASBA) | 89% | 98% | 0.5–5 copies/µL | 3.66 and 12.61 | 10–50 min | Saliva | S gene | [42,43] |
| Serological Based Techniques | |||||||||
| 10 | ELISA | 99.30% | 95–96% | 100 pg/ml | 71.40 | 3–5 h | Blood samples | IgG, IgM | [27,44,45] |
| 11 | IFA (Immunofluorescence Assay) | 64.50% | 96.3–100% | NA | NA | 3 h | Serum samples | IgG, IgM | [46,47] |
| 12 | Neutralization Assay | 76.5–100% | 100% | 0.22 copies/µL | NA | 2–3 days | Serum samples | IgG, IgM | [48,49] |
| 13 | Antigen Detection Assay | 75.50% | 94.90% | 46–750 copies/µL | 5 | 15 min | Nasopharyngeal | Nucleocapsid | [50,51,52] |
| 14 | Chemiluminescence Enzyme Immunoassays (CLIA) | 99.67/90% | 99.77/80% | NA | NA | 45 min | Plasma | SARS-Cov2 RBD | [53,54] |
| 15 | WB (Westren Blotting) | 90.90% | 98.30% | NA | NA | 4 h | Serum | Antinucleocapsid antibody | [27,55,56] |
| 16 | HTP-Microfluidic Device | 95% | 91% | 1.6 ng/mL | NA | 2.5 h | Serum samples | IgG, IgM | [57] |
| 17 | Lateral Flow Assay | 96.7–100% | 97.5–98.8% | 5–20 ng/mL | 27.42 | 15 min | Blood samples | IgG, IgM | [58,59,60] |
| 18 | Luciferase immunosorbent assay | 96.7–100 % | 100% | 0.4–75 pg/μL | NA | <2.5 h | Serum samples | IgG | [61] |
| 19 | MIA (Microsphere Immunoassay) | 90.17% | 99.49% | 0.121 U/L–0.366 U/L | NA | 10–15 min | Serum samples | IgG, IgM, and IgA | [62,63] |
| 20 | Immunochromatographic Assay | 43.20% | 98.0% | 0.18 ng/μL | NA | 10–15 min | Serum samples | IgG, IgM | [64,65] |
| Sr. No. | Nanozymes | Disease/Pathogen | Application/Principle | References |
|---|---|---|---|---|
| 1. | Metal nanozymes (Au-Pt core-shell nanozyme) Peroxidase-mimicking MNP3 | Infectious diseases such as SARS-CoV, MERS-Covid-19. | Paper base biosensors for colorimetric detection. | [70] |
| 2. | Co-Fe@hemin-peroxidase nanozyme | SARS-CoV-2 | Chemiluminescence paper test for rapid and sensitive detection of SARS-CoV-2 antigen. | [71] |
| 3. | Fe3O4 magnetic nanoparticle (Nanozyme probe) | Ebola virus | Detects the glycoprotein of Ebola virus. | [72] |
| 4. | MagLISA | Influenza virus A | Provide detection, reduce spread of influenza virus and provide immediate clinical treatment. | [73] |
| 5. | Au@Pt Mesopotous SiO2 nanozymes | Mumps virus | Diagnosis of mumps-virus, more sensitive compared to conventional immunoassay. | [74] |
| 6. | Nanozyme Aptanser | Human nanovirus (NoV) | Allows ultrasensitive NoV detection rapidly, offering simplicity of use. | [75] |
| 7. | Fe, O4 Bi2, S3 nanozyme | Cancer cell | Induce cancer cell death, Anti-cancerous. | [76] |
| 8. | Iron Oxide nanozyme | Influenza virus | Catalytic inactivation of influenza virus, provide protection from viral transmission and infection. | [77] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ali, J.; Elahi, S.N.; Ali, A.; Waseem, H.; Abid, R.; Mohamed, M.M. Unveiling the Potential Role of Nanozymes in Combating the COVID-19 Outbreak. Nanomaterials 2021, 11, 1328. https://doi.org/10.3390/nano11051328
Ali J, Elahi SN, Ali A, Waseem H, Abid R, Mohamed MM. Unveiling the Potential Role of Nanozymes in Combating the COVID-19 Outbreak. Nanomaterials. 2021; 11(5):1328. https://doi.org/10.3390/nano11051328
Chicago/Turabian StyleAli, Jafar, Saira Naveed Elahi, Asghar Ali, Hassan Waseem, Rameesha Abid, and Mohamed M. Mohamed. 2021. "Unveiling the Potential Role of Nanozymes in Combating the COVID-19 Outbreak" Nanomaterials 11, no. 5: 1328. https://doi.org/10.3390/nano11051328
APA StyleAli, J., Elahi, S. N., Ali, A., Waseem, H., Abid, R., & Mohamed, M. M. (2021). Unveiling the Potential Role of Nanozymes in Combating the COVID-19 Outbreak. Nanomaterials, 11(5), 1328. https://doi.org/10.3390/nano11051328