Effects of Doxorubicin Delivery by Nitrogen-Doped Graphene Quantum Dots on Cancer Cell Growth: Experimental Study and Mathematical Modeling
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Culture
2.2. Cell Growth Assay
2.3. MTT Cell Viability Assay
2.4. Synthesis of DOX-N-GQDs
2.5. Microscopy
2.6. Image Analysis
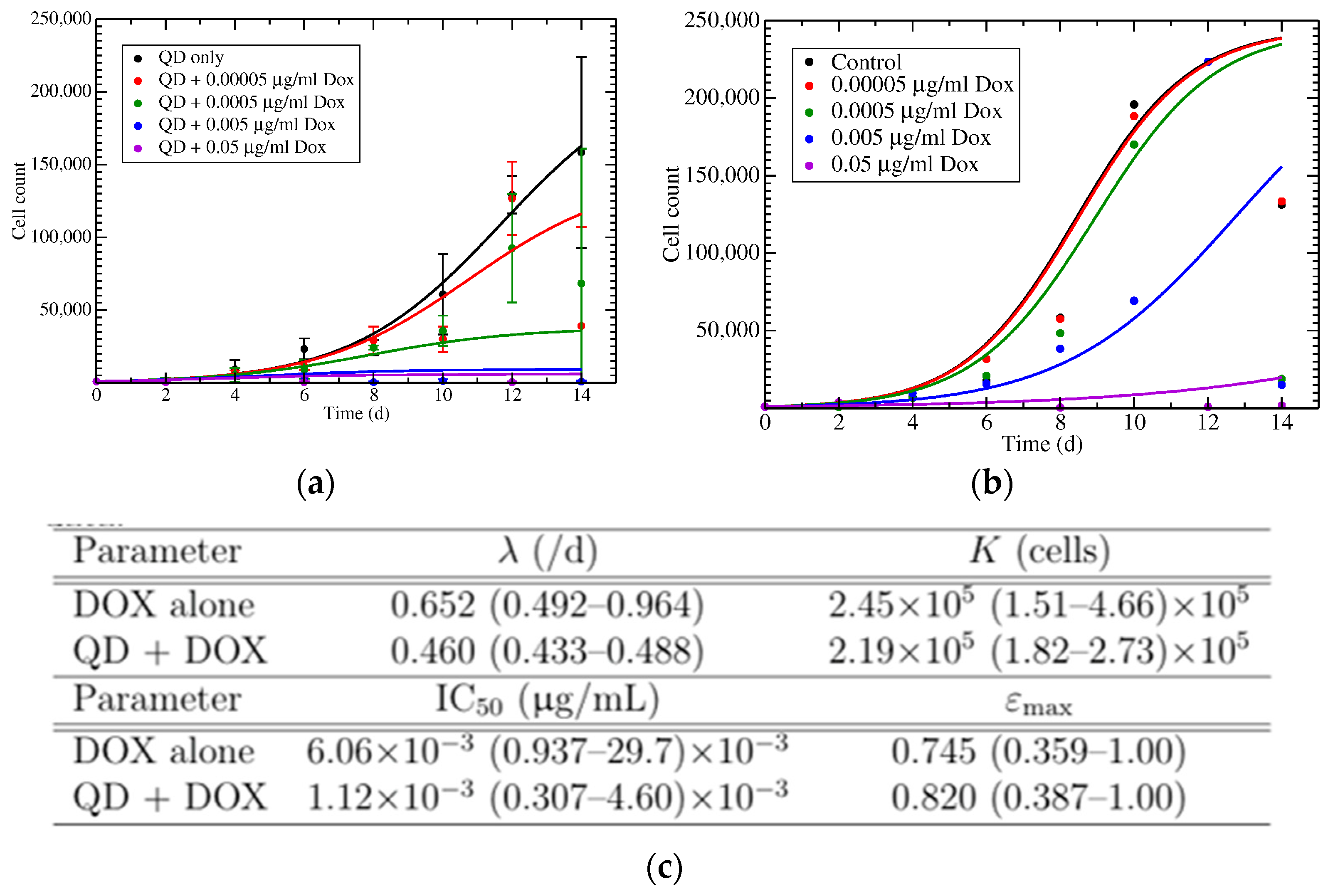
2.7. Mathematical Modeling
3. Results
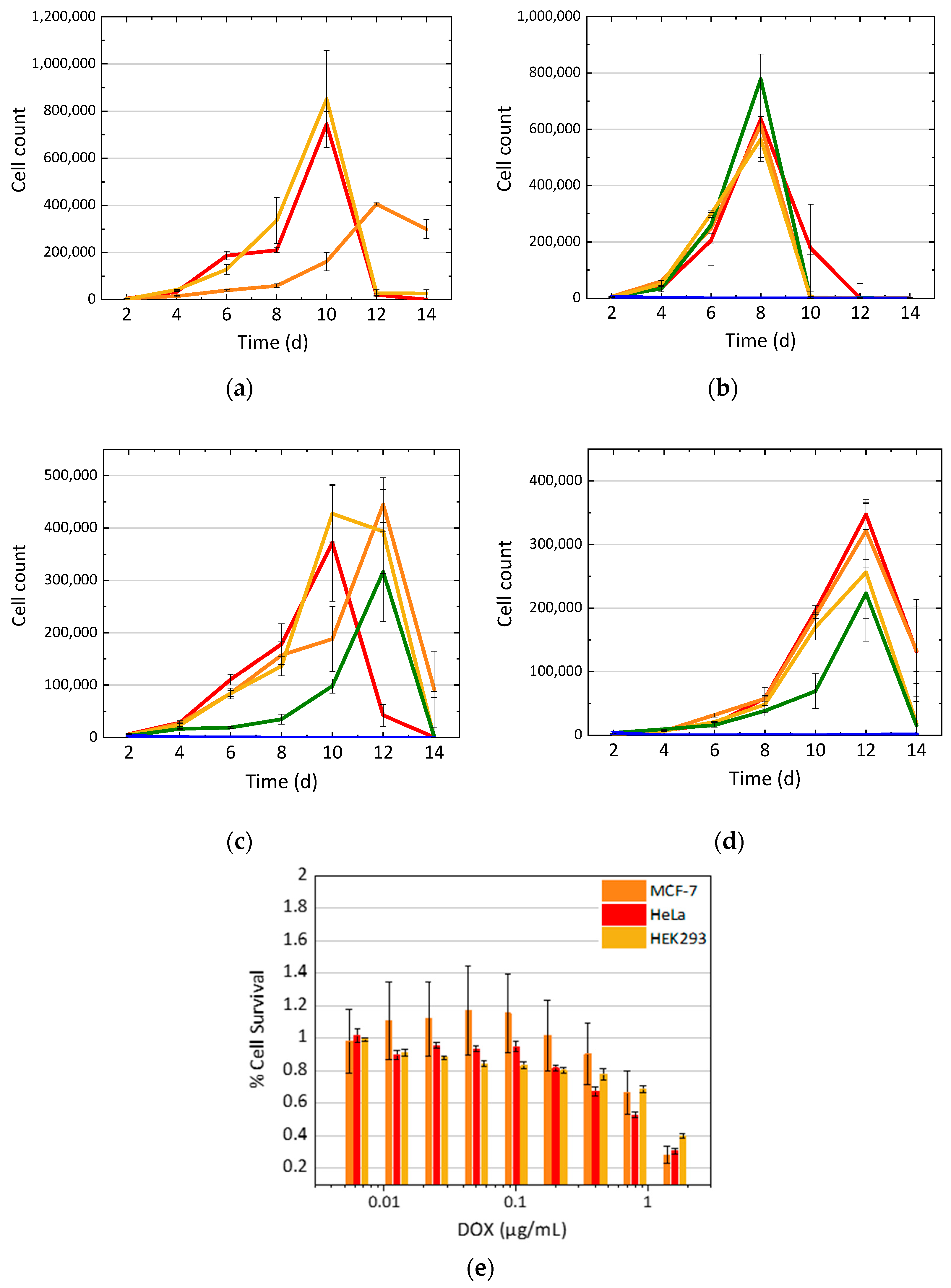
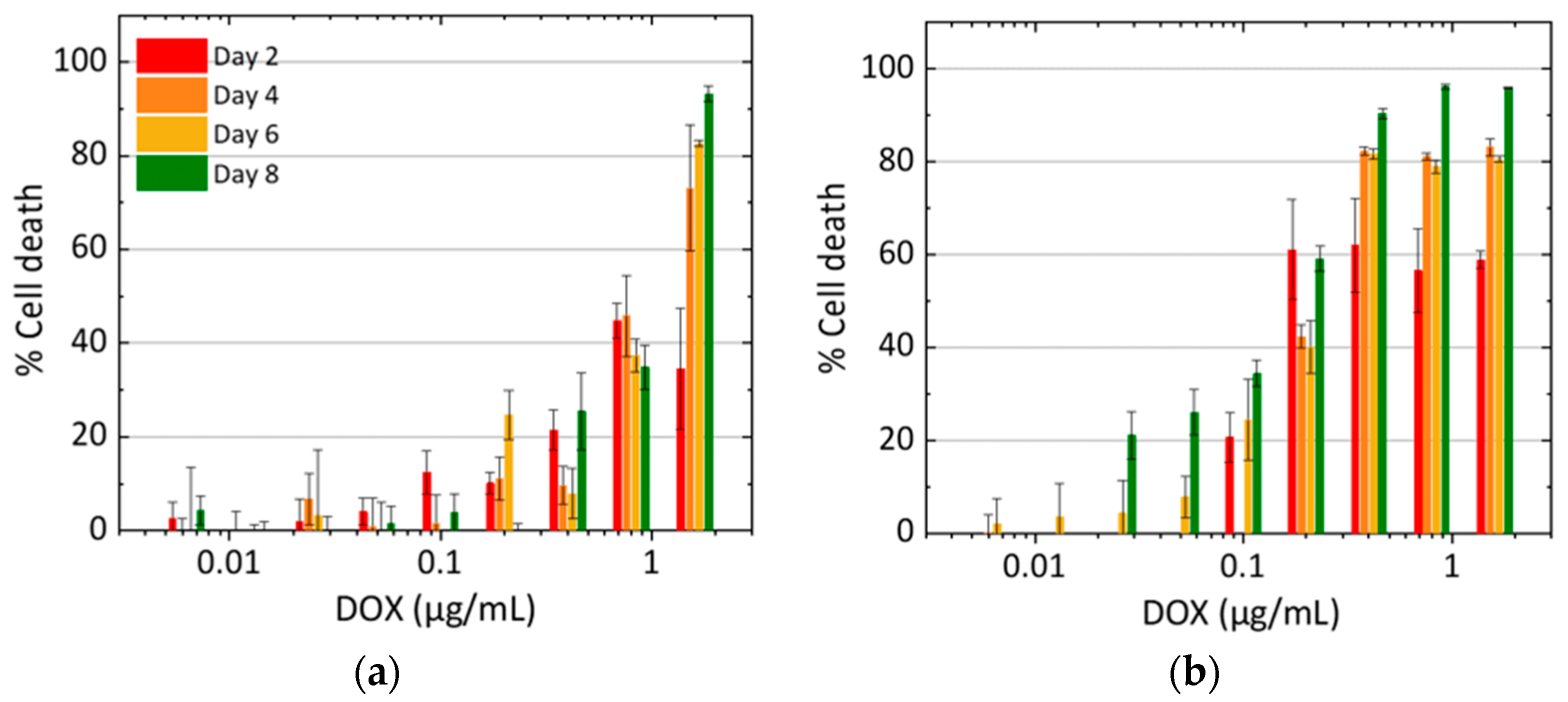
3.1. Cell Growth with DOX Treatment
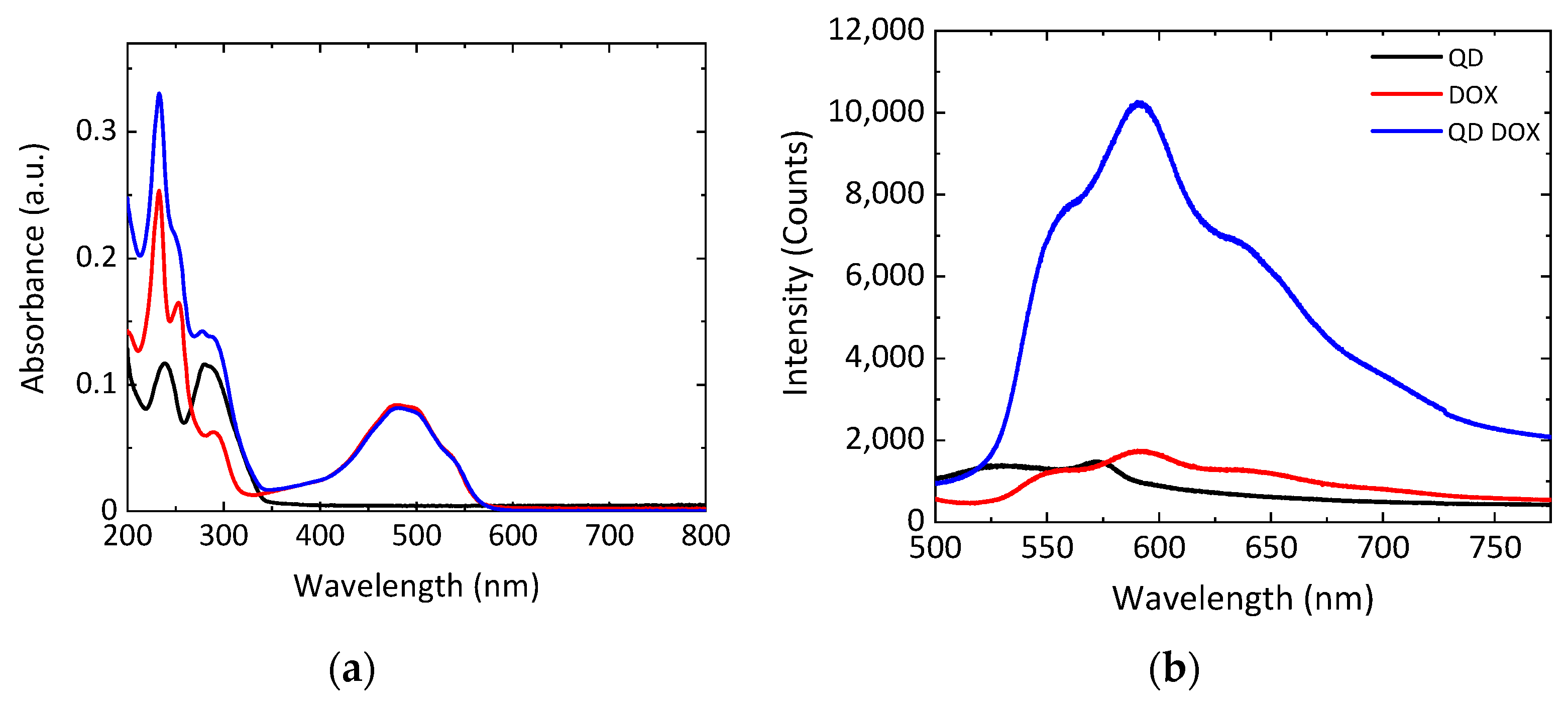
3.2. Characterization of DOX-N-GQDs
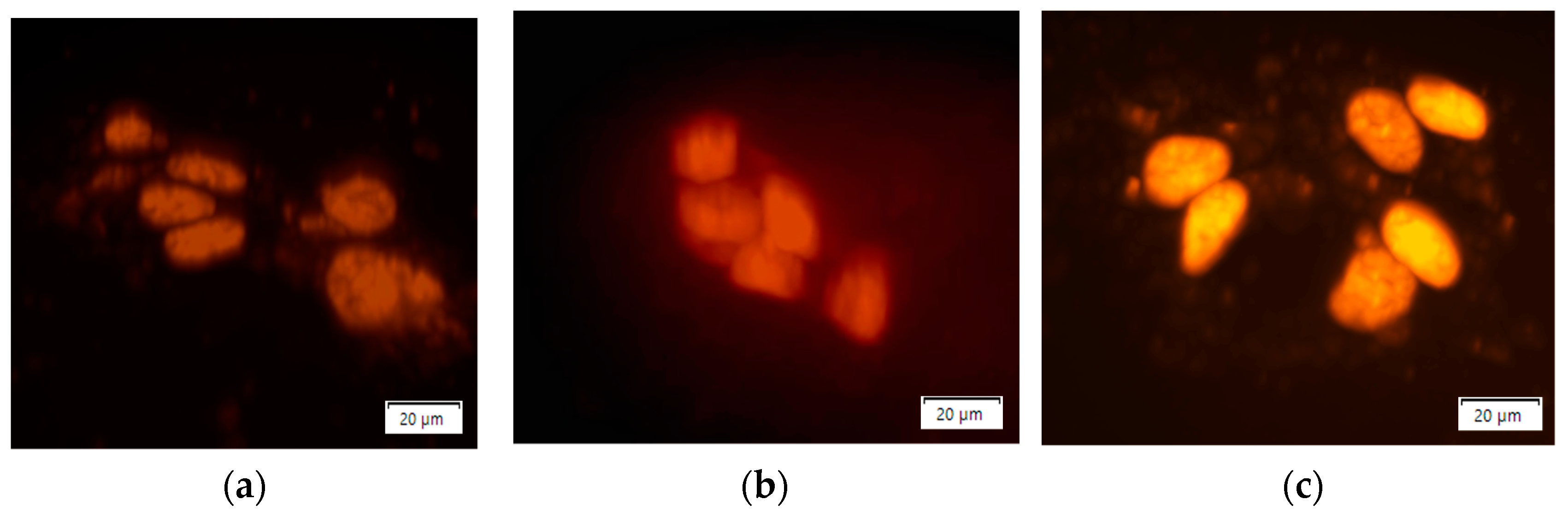
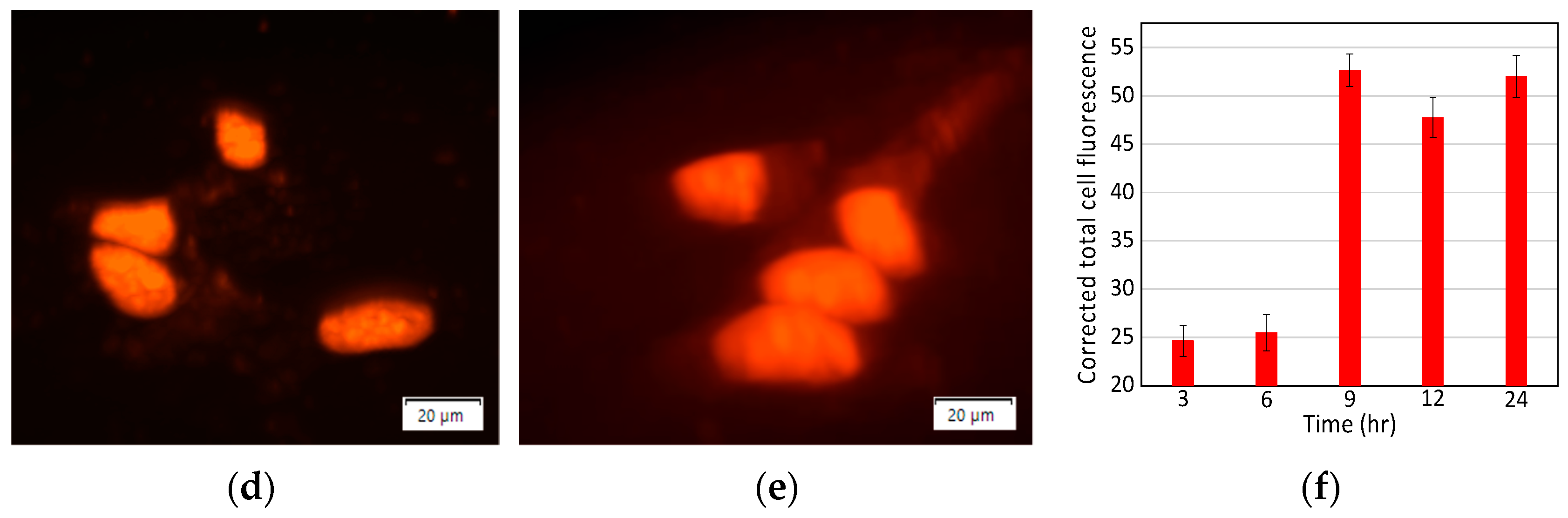
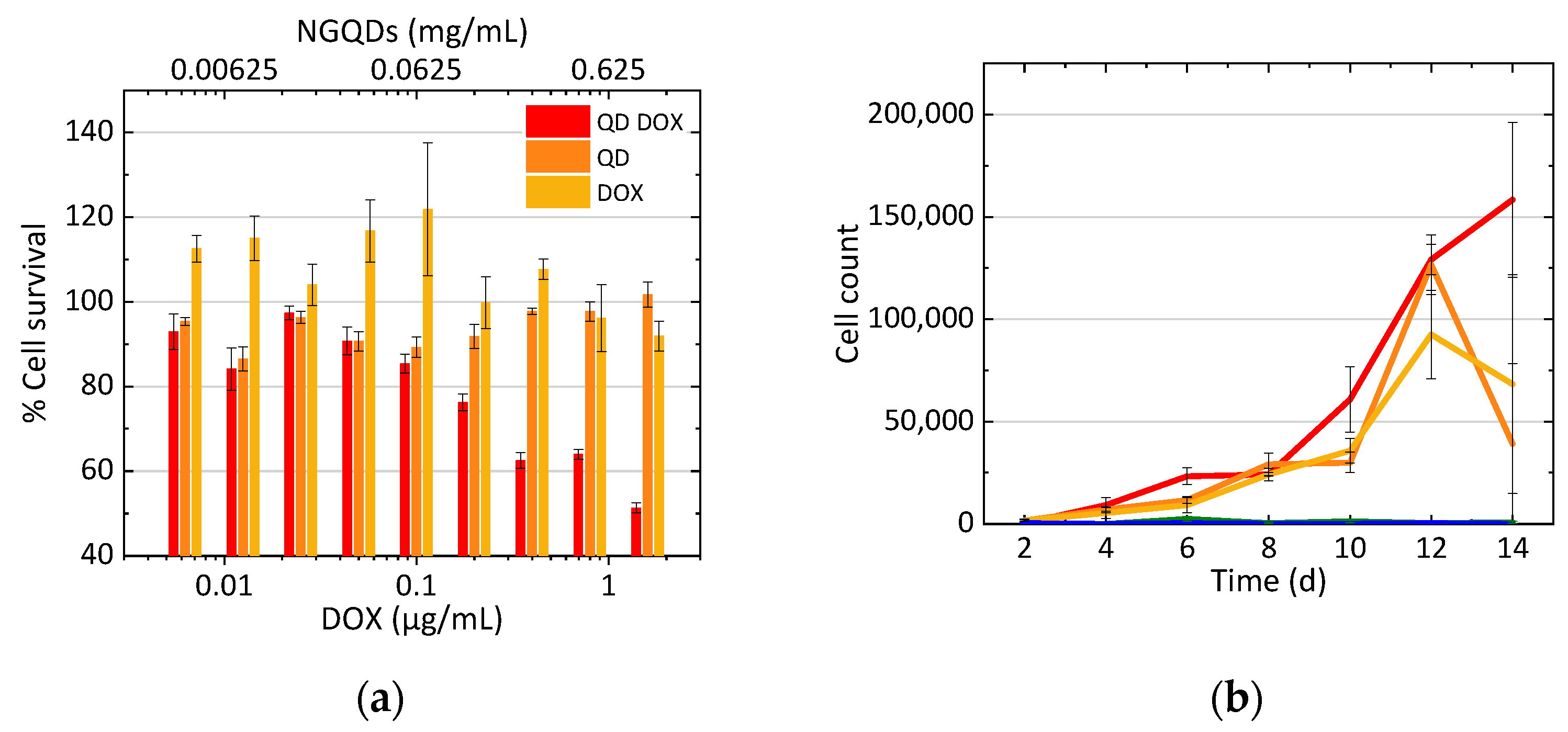
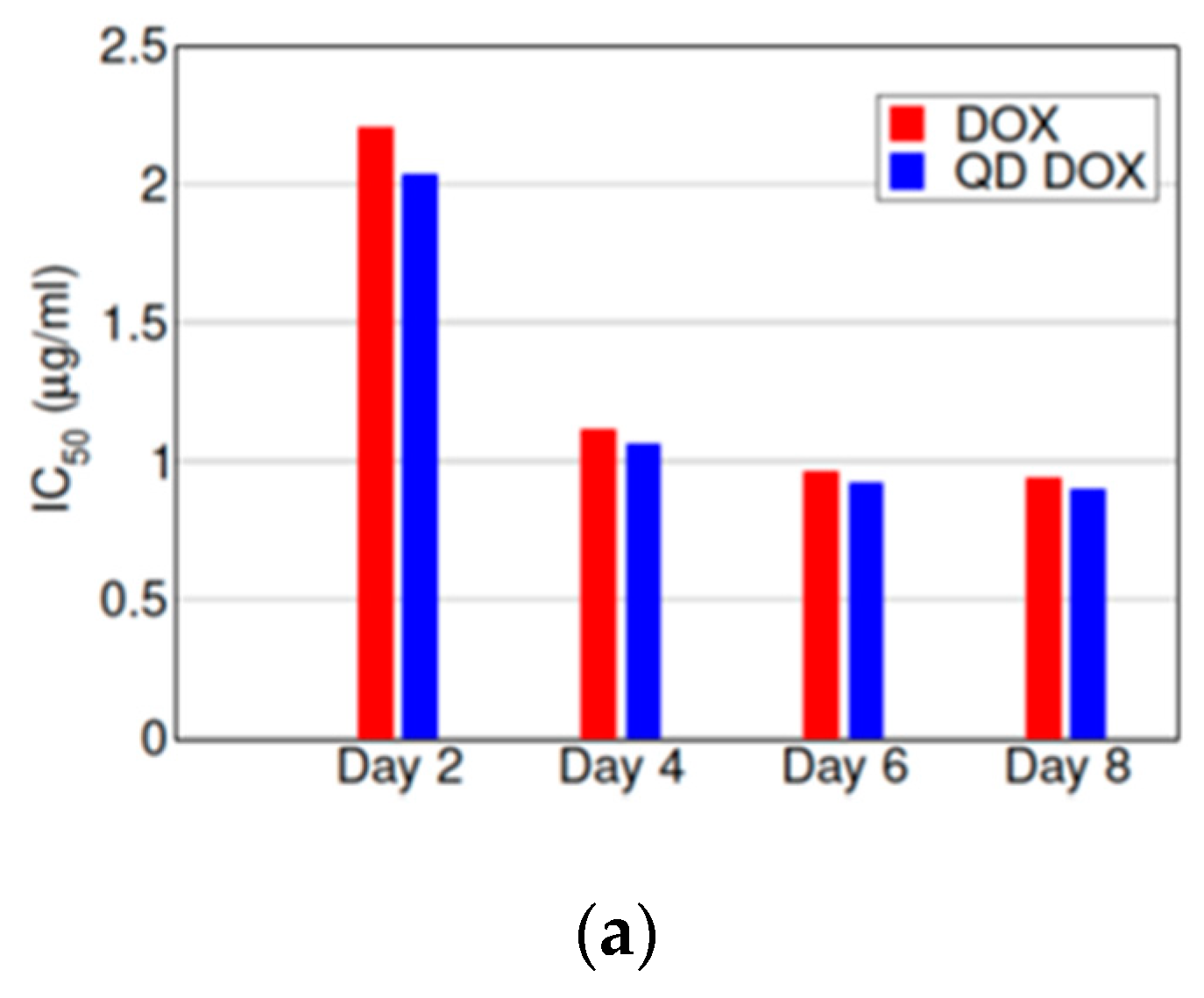
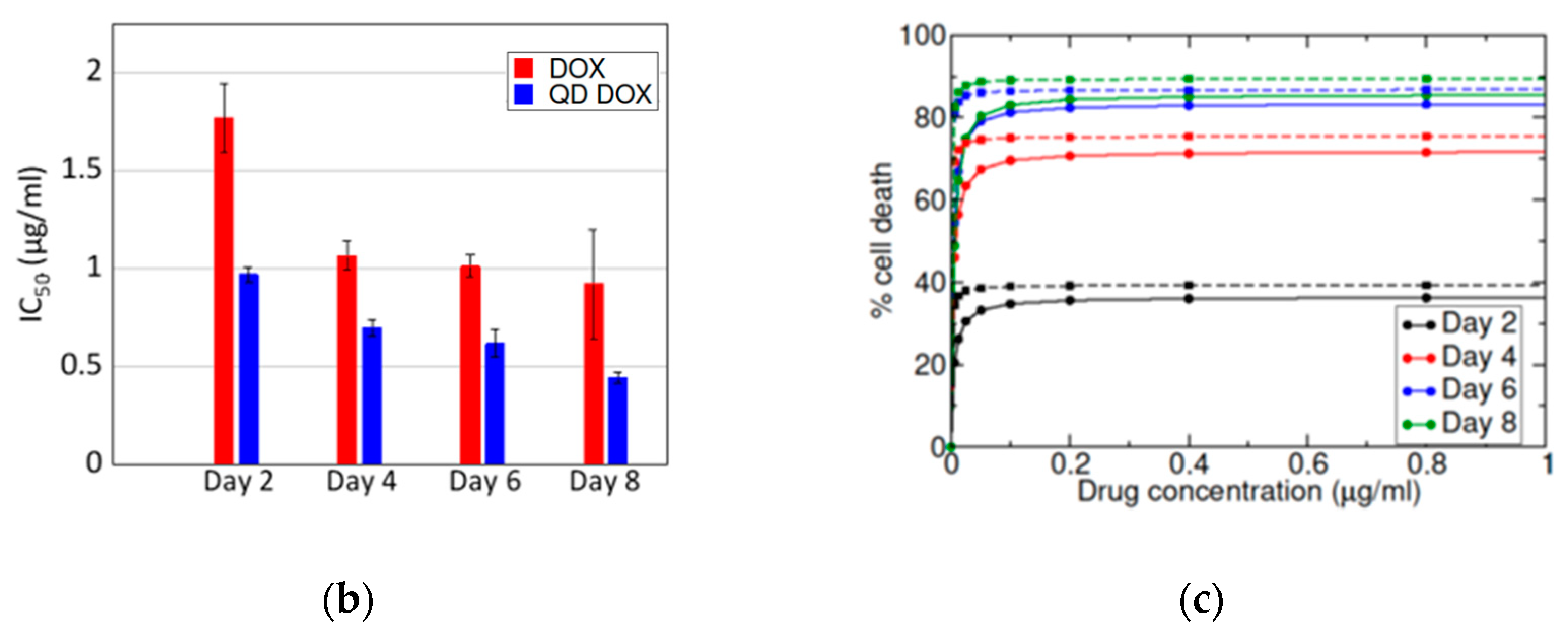
3.3. Treatment of Cells with N-GQD-DOX Complex
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global Cancer Statistics 2018: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. Ca-A Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R. Hallmarks of Cancer: The Next Generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef] [PubMed]
- Siddik, Z.H. Mechanisms of Action of Cancer Chemotherapeutic Agents: DNA-Interactive Alkylating Agents and Antitumour Platinum-Based Drugs. In The Cancer Handbook, 1st ed.; Alison, M.R., Ed.; John Wily & Sons, Ltd.: Chichester, UK, 2005; p. 1. [Google Scholar]
- Thorn, C.F.; Oshiro, C.; Marsh, S.; Hernandez-Boussard, T.; McLeod, H.; Klein, T.E.; Altman, R.B. Doxorubicin pathways: Pharmacodynamics and adverse effects. Pharm. Genom. 2011, 21, 440–446. [Google Scholar] [CrossRef] [PubMed]
- Riccio, A.A.; Schellenberg, M.J.; Williams, R.S. Molecular mechanisms of topoisomerase 2 DNA–protein crosslink resolution. Cell. Mol. Life Sci. 2020, 77, 81–91. [Google Scholar] [CrossRef] [PubMed]
- Wallace, K.; Sardão, V.; Oliveira, P. Mitochondrial Determinants of Doxorubicin-Induced Cardiomyopathy. Circ. Res. 2020, 126, 926–941. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Rodriguez, R.; Campbell, E.; Naumov, A. Multifunctional Graphene Oxide/Iron Oxide Nanoparticles for Magnetic Targeted Drug Delivery Dual Magnetic Resonance/Fluorescence Imaging and Cancer Sensing. PLoS ONE 2019, 14, e0217072. [Google Scholar] [CrossRef] [PubMed]
- Kersting, D.; Fasbender, S.; Pilch, R.; Kurth, J.; Franken, A.; Ludescher, M.; Naskou, J.; Hallenberger, A.; von Gall, C.; Mohr, C.J.; et al. From in vitro to ex vivo: Subcellular localization and uptake of graphene quantum dots into solid tumors. Nanotechnology 2019, 30, 395101. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Tan, X.; Pang, X.; Liu, L.; Tan, F.; Li, N. MoS2 Quantum Dot@Polyaniline Inorganic–Organic Nanohybrids for in Vivo Dual-Modal Imaging Guided Synergistic Photothermal/Radiation Therapy. ACS Appl. Mater. Interfaces 2016, 8, 24331–24338. [Google Scholar] [CrossRef]
- Tamarov, K.; Xu, W.; Osminkina, L.; Zinovyev, S.; Soininen, P.; Kudryavtsev, A.; Gongalsky, M.; Gaydarova, A.; Närvänen, A.; Timoshenko, V.; et al. Temperature Responsive Porous Silicon Nanoparticles for Cancer Therapy–Spatiotemporal Triggering through Infrared and Radiofrequency Electromagnetic Heating. J. Control. Release 2016, 241, 220–228. [Google Scholar] [CrossRef]
- Raghav, R.; Srivastava, S. Core–shell Gold–silver Nanoparticles Based Impedimetric Immunosensor for Cancer Antigen CA125. Sens. Actuators B Chem. 2015, 220, 557–564. [Google Scholar] [CrossRef]
- Miyabe, H.; Hyodo, M.; Nakamura, T.; Sato, Y.; Hayakawa, Y.; Harashima, H. A New Adjuvant Delivery System ‘cyclic Di-GMP/YSK05 Liposome’ for Cancer Immunotherapy. J. Control. Release 2014, 184, 20–27. [Google Scholar] [CrossRef] [PubMed]
- El-Sayed, I.H.; Huang, X.; El-Sayed, M.A. Surface Plasmon Resonance Scattering and Absorption of Anti- EGFR Antibody Conjugated Gold Nanoparticles in Cancer Diagnostics: Applications in Oral Cancer. Nano Lett. 2005, 5, 829–834. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Salehi, M.; Schütz, M.; Rudi, K.; Schlücker, S. Microspectroscopic SERS Detection of Interleukin-6 with Rationally Designed Gold/Silver Nanoshells. Analyst 2013, 138, 1764–1771. [Google Scholar] [CrossRef] [PubMed]
- Jokerst, J.V.; Cole, A.J.; Sompel, D.; Gambhir, S.S. Gold Nanorods for Ovarian Cancer Detection with Photoacoustic Imaging and Resection Guidance Via Raman Imaging in Living Mice. ACS Nano 2012, 6, 10366–10377. [Google Scholar] [CrossRef]
- Xin, Y.; Yin, M.; Zhao, L.; Meng, F.; Luo, L. Recent progress on nanoparticle-based drug delivery systems for cancer therapy. Cancer Biol. Med. 2017, 14, 228–241. [Google Scholar] [CrossRef]
- Eisbruch, A.; Blick, M.; Lee, J.S.; Sacks, P.G.; Gutterman, J. Analysis of the Epidermal Growth Factor Receptor Gene in Fresh Human Head and Neck Tumors. Cancer Res. 1987, 47, 3603–3605. [Google Scholar]
- Li, X.; Guo, X.; Wang, D.; Wang, Y.; Li, X.; Zhang, X.; Lin, J. Spectral Analysis for Diagnosis of Rectum Cancer using Fluorescence and Raman Spectroscopy of Serum. In Proceedings of the 2005 IEEE Engineering in Medicine and Biology 27th Annual Conference, Shangai, China, 17–18 January 2006; pp. 5449–5452. [Google Scholar]
- Hasan, M.T.; Campbell, E.; Sizova, O.; Lyle, V.; Akkaraju, G.; Kirkpatrick, D.L.; Naumov, A.V. Multi-Drug/Gene NASH Therapy Delivery and Selective Hyperspectral NIR Imaging using Chirality-Sorted Single-Walled Carbon Nanotubes. Cancers 2019, 11, 1175. [Google Scholar] [CrossRef]
- Pantarotto, D.; Singh, R.; McCarthy, D.; Erhardt, M.; Briand, J.; Prato, M.; Kostarelos, K.; Bianco, A. Functionalized Carbon Nanotubes for Plasmid DNA Gene Delivery. Angew. Chem. 2004, 43, 5242–5246. [Google Scholar] [CrossRef]
- Kirkpatrick, D.L.; Weiss, M.; Naumov, A.; Bartholomeusz, G.; Weisman, R.B.; Gliko, O. Carbon Nanotubes: Solution for the Therapeutic Delivery of siRNA? Materials 2012, 5, 278–301. [Google Scholar] [CrossRef]
- Kushwaha, S.K.S.; Ghoshal, S.; Rai, A.K.; Singh, S. Carbon nanotubes as a novel drug delivery system for anticancer therapy: A review. Braz. J. Pharm. Sci. 2013, 49, 629–643. [Google Scholar] [CrossRef]
- Lin, C.; Yang, H.; Sanchez, S.R.; Mao, W.; Pang, L.; Beckingham, K.M.; Bast, R.C.; Weisman, R.B. In Vivo Optical Detection and Spectral Triangulation of Carbon Nanotubes. ACS Appl. Mater. Interfaces 2017, 9, 41680–41690. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Y.; Zhang, J.; An, L.; Cao, Q.; Deng, Y.; Liang, G. Oligomeric Nanoparticles Functionalized with NIR-Emitting CdTe/CdS QDs and Folate for Tumor-Targeted Imaging. Biomaterials 2014, 35, 7881–7886. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.; Zhang, S. Creating High Yield Water Soluble Luminescent Graphene Quantum Dots Via Exfoliating and Disintegrating Carbon Nanotubes and Graphite Flakes. Chem. Commun. 2012, 48, 10177–10179. [Google Scholar] [CrossRef]
- Zhao, M.; Zhu, B. The Research and Applications of Quantum Dots as Nano-Carriers for Targeted Drug Delivery and Cancer Therapy. Nanoscale Res. Lett. 2016, 11, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Campbell, E.; Hasan, M.T.; Gonzalez Rodriguez, R.; Akkaraju, G.R.; Naumov, A.V. Doped Graphene Quantum Dots for Intracellular Multicolor Imaging and Cancer Detection. ACS Biomater. Sci. Eng. 2019, 5, 4671–4682. [Google Scholar] [CrossRef]
- Al Dine, E.J.; Marchal, S.; Schneider, R.; Hamie, B.; Ghanbaja, J.; Roques-Carmes, T.; Hamieh, T.; Toufaily, J.; Gaffet, E.; Alem, H. A Facile Approach for Doxorubicine Delivery in Cancer Cells by Responsive and Fluorescent Core/Shell Quantum Dots. Bioconjugate Chem. 2018, 29, 2248–2256. [Google Scholar] [CrossRef]
- Yum, K.; Na, S.; Xiang, Y.; Wang, N.; Yu, M. Mechanochemical Delivery and Dynamic Tracking of Fluorescent Quantum Dots in the Cytoplasm and Nucleus of Living Cells. Nano Lett. 2009, 9, 2193–2198. [Google Scholar] [CrossRef]
- Chen, M.; He, Y.; Chen, X.; Wang, J. Quantum-Dot-Conjugated Graphene as a Probe for Simultaneous Cancer-Targeted Fluorescent Imaging, Tracking, and Monitoring Drug Delivery. Bioconjugate Chem. 2013, 24, 387–397. [Google Scholar] [CrossRef]
- Wang, C.; Wu, C.; Zhou, X.; Han, T.; Xin, X.; Wu, J.; Zhang, J.; Guo, S. Enhancing Cell Nucleus Accumulation and DNA Cleavage Activity of Anti-Cancer Drug Via Graphene Quantum Dots. Sci. Rep. 2013, 3, 2852. [Google Scholar] [CrossRef]
- Javanbakht, S.; Namazi, H. Doxorubicin Loaded Carboxymethyl Cellulose/Graphene Quantum Dot Nanocomposite Hydrogel Films as a Potential Anticancer Drug Delivery System. Mater. Sci. Eng. C 2018, 87, 50–59. [Google Scholar] [CrossRef]
- Wang, X.; Sun, X.; Lao, J.; He, H.; Cheng, T.; Wang, M.; Wang, S.; Huang, F. Multifunctional Graphene Quantum Dots for Simultaneous Targeted Cellular Imaging and Drug Delivery. Colloids Surf. B Biointerfaces 2014, 122, 638–644. [Google Scholar] [CrossRef] [PubMed]
- Barbolosi, D.; Ciccolini, J.; Lacarelle, B.; Barlési, F.; André, N. Computational Oncology—Mathematical Modelling of Drug Regimens for Precision Medicine. Nat. Rev. Clin. Oncol. 2016, 13, 242–254. [Google Scholar] [CrossRef] [PubMed]
- Lai, X.; Geier, O.M.; Fleischer, T.; Garred, Ø.; Borgen, E.; Funke, S.W.; Kumar, S.; Rognes, M.E.; Seierstad, T.; Børresen-Dale, A.; et al. Toward Personalized Computer Simulation of Breast Cancer Treatment: A Multiscale Pharmacokinetic and Pharmacodynamic Model Informed by Multitype Patient Data. Cancer Res. 2019, 79, 4293–4304. [Google Scholar] [CrossRef] [PubMed]
- Karolak, A.; Markov, D.A.; McCawley, L.S.; Rejniak, K.A. Towards Personalized Computational Oncology: From Spatial Models of Tumour Spheroids, to Organoids, to Tissues. J. R. Soc. Interface 2015, 15, 20170703. [Google Scholar] [CrossRef] [PubMed]
- Bregenzer, M.E.; Horst, E.N.; Mehta, P.; Novak, C.M.; Raghavan, S.; Snyder, C.S.; Mehta, G. Integrated Cancer Tissue Engineering Models for Precision Medicine. PLoS ONE 2019, 14, e0216564. [Google Scholar] [CrossRef]
- Yankeelov, T.E.; An, G.; Saut, O.; Luebeck, G.; Popel, A.S.; Ribba, B.; Vicini, P.; Zhou, X.; Weis, J.A.; Ye, K.; et al. Multi-Scale Modeling in Clinical Oncology: Opportunities and Barriers to Success. Ann. Biomed. Eng. 2016, 44, 2626–2641. [Google Scholar] [CrossRef]
- McKenna, M.T.; Weis, J.A.; Quaranta, V.; Yankeelov, T.E. Leveraging Mathematical Modeling to Quantify Pharmacokinetic and Pharmacodynamic Pathways: Equivalent Dose Metric. Front. Physiol. 2019, 10, 616. [Google Scholar] [CrossRef]
- Brocato, T.A.; Coker, E.N.; Durfee, P.N.; Lin, Y.; Townson, J.; Wyckoff, E.F.; Cristini, V.; Brinker, C.J.; Wang, Z. Understanding the Connection between Nanoparticle Uptake and Cancer Treatment Efficacy using Mathematical Modeling. Sci. Rep. 2018, 8, 7538–7841. [Google Scholar] [CrossRef]
- Bodnar, M.; Piotrowska, M.J.; Bogdanska, M.U. Mathematical Analysis of a Generalised Model of Chemotherapy for Low Grade Gliomas. Discret. Contin. Dyn. Syst. 2019, 24, 2149–2167. [Google Scholar] [CrossRef]
- Shariatpanahi, S.P.; Shariatpanahi, S.P.; Madjidzadeh, K.; Hassan, M.; Abedi-Valugerdi, M. Mathematical Modeling of Tumor-Induced Immunosuppression by Myeloid-Derived Suppressor Cells: Implications for Therapeutic Targeting Strategies. J. Theor. Biol. 2018, 442, 1–10. [Google Scholar] [CrossRef]
- Jarrett, A.M.; Shah, A.; Bloom, M.J.; McKenna, M.T.; Hormuth, D.A., II; Yankeelov, T.E.; Sorace, A.G. Experimentally-Driven Mathematical Modeling to Improve Combination Targeted and Cytotoxic Therapy for HER2+breast Cancer. Sci. Rep. 2019, 9, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Murphy, H.; Jaafari, H.; Dobrovolny, H.M. Differences in Predictions of ODE Models of Tumor Growth: A Cautionary Example. BMC Cancer 2016, 16, 163. [Google Scholar] [CrossRef] [PubMed]
- Harris, L.A.; Frick, P.L.; Garbett, S.P.; Hardeman, K.N.; Paudel, B.B.; Lopez, C.F.; Quaranta, V.; Tyson, D.R. An unbiased metric of antiproliferative drug effect in vitro. Nat. Methods 2016, 13, 497–500. [Google Scholar] [CrossRef] [PubMed]
- Hosoya, H.; Dobroff, A.S.; Driessen, W.H.P.; Cristini, V.; Brinker, L.M.; Staquicini, F.I.; Cardó-Vila, M.; D’Angelo, S.; Ferrara, F.; Proneth, B.; et al. Integrated nanotechnology platform for tumor-targeted multimodal imaging and therapeutic cargo release. Proc. Natl. Acad. Sci. USA 2016, 113, 1877–1882. [Google Scholar] [CrossRef]
- Pascal, J.; Ashley, C.E.; Wang, Z.; Brocato, T.A.; Butner, J.D.; Carnes, E.C.; Koay, E.J.; Brinker, C.J.; Cristini, V. Mechanistic modeling identifies drug-uptake history as predictor of tumor drug resistance and nano-carrier-mediated response. ACS Nano 2013, 7, 11174–11182. [Google Scholar] [CrossRef]
- Wong, C.; Stylianopoulos, T.; Cui, J.; Martin, J.; Chauhan, V.P.; Jiang, W.; Popovic, Z.; Jain, R.K.; Bawendi, M.G.; Fukumura, D. Multistage nanoparticle delivery system for deep penetration into tumor tissue. Proc. Natl. Acad. Sci. USA 2011, 108, 2426–2431. [Google Scholar] [CrossRef]
- Dogra, P.; Butner, J.D.; Chuang, Y.-L.; Caserta, S.; Goel, S.; Jeffrey Brinker, C.; Cristini, V.; Wang, Z. Mathematical modeling in cancer nanomedicine: A review. Biomed. Microdevices 2019, 21, 40. [Google Scholar] [CrossRef]
- Wang, A.Z.; Langer, R.; Farokhzad, O.C. Nanoparticle delivery of cancer drugs. Annu. Rev. Med. 2012, 63, 185–198. [Google Scholar] [CrossRef]
- Sun, T.; Zhang, Y.S.; Pang, B.; Choon Hyun, D.; Yang, M.; Xia, Y. Engineered nanoparticles for drug delivery in cancer therapy. Angew. Chem. Int. Ed. 2014, 53, 12320–12364. [Google Scholar] [CrossRef]
- Iannazzo, D.; Pistone, A.; Salamò, M.; Galvagno, S.; Romeo, R.; Giofré, S.V.; Branca, C.; Visalli, G.; di Pietro, A. Graphene quantum dots for cancer targeted drug delivery. Int. J. Pharm. 2017, 518, 185–192. [Google Scholar] [CrossRef]
- Hasan, M.T.; Gonzalez-Rodriguez, R.; Ryan, C.; Faerber, N.; Coffer, J.L.; Naumov, A.V. Photo-and Electroluminescence from Nitrogen-Doped and Nitrogen–Sulfur Codoped Graphene Quantum Dots. Adv. Funct. Mater. 2018, 28, 1804337. [Google Scholar] [CrossRef]
- Holford, N.H.; Sheiner, L.B. Understanding the Dose-Effect Relationship: Clinical Application of Pharmacokinetic-Pharmacodynamic Models. Clin. Pharmacokinet. 1981, 6, 429–453. [Google Scholar] [CrossRef] [PubMed]
- Oduola, W.O.; Li, X. Multiscale Tumor Modeling with Drug Pharmacokinetic and Pharmacodynamic Profile using Stochastic Hybrid System. Cancer Inform. 2018, 17, 1176935118790262. [Google Scholar] [CrossRef] [PubMed]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Frieler, M.; Pho, C.; Lee, B.H.; Dobrovolny, H.; Akkaraju, G.R.; Naumov, A.V. Effects of Doxorubicin Delivery by Nitrogen-Doped Graphene Quantum Dots on Cancer Cell Growth: Experimental Study and Mathematical Modeling. Nanomaterials 2021, 11, 140. https://doi.org/10.3390/nano11010140
Frieler M, Pho C, Lee BH, Dobrovolny H, Akkaraju GR, Naumov AV. Effects of Doxorubicin Delivery by Nitrogen-Doped Graphene Quantum Dots on Cancer Cell Growth: Experimental Study and Mathematical Modeling. Nanomaterials. 2021; 11(1):140. https://doi.org/10.3390/nano11010140
Chicago/Turabian StyleFrieler, Madison, Christine Pho, Bong Han Lee, Hana Dobrovolny, Giridhar R. Akkaraju, and Anton V. Naumov. 2021. "Effects of Doxorubicin Delivery by Nitrogen-Doped Graphene Quantum Dots on Cancer Cell Growth: Experimental Study and Mathematical Modeling" Nanomaterials 11, no. 1: 140. https://doi.org/10.3390/nano11010140
APA StyleFrieler, M., Pho, C., Lee, B. H., Dobrovolny, H., Akkaraju, G. R., & Naumov, A. V. (2021). Effects of Doxorubicin Delivery by Nitrogen-Doped Graphene Quantum Dots on Cancer Cell Growth: Experimental Study and Mathematical Modeling. Nanomaterials, 11(1), 140. https://doi.org/10.3390/nano11010140






