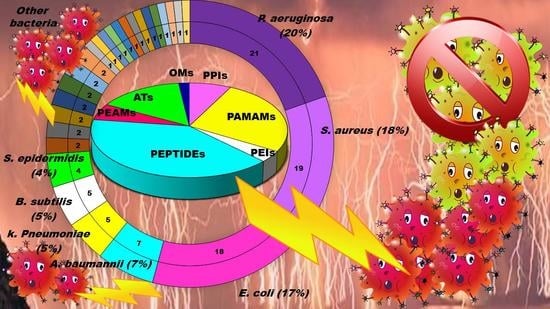
From Nanobiotechnology, Positively Charged Biomimetic Dendrimers as Novel Antibacterial Agents: A Review
Abstract
1. Introduction
2. Biofilm as a Mechanism for Antibiotic Resistance
3. Dendrimers (Ds)
Dendrimers as Antimicrobials
Principal Types of Antimicrobial Dendrimers Developed in the Previous Decade
4. Cationic Antibacterial Dendrimers (CADs)
4.1. PAMAM and PPI Dendrimers (PAMAM Ds and PPI Ds)
4.2. PEI-Based Dendrimers (PEI-Ds)
4.3. Cationic Peptides Dendrimers
4.4. Entirely Polyester-Based Uncharged Scaffolds Peripherally Modified with Cationic Amino Acids
4.5. Ammonium-Terminated Dendrimers
4.5.1. Ammonium-Terminated Phosphorous Dendrimers
4.5.2. Ammonium-Terminated Carbosilane Dendrimers
4.5.3. Ammonium-Terminated Ruthenium and Zinc Encased Phthalocyanines (Pcs) Dendrimers
4.6. Cationic Organometallic Ds
5. Conclusions and Considerations of the Authors
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Schito, A.M.; Alfei, S. Antibacterial activity of non-cytotoxic, amino acid-modified polycationic dendrimers against Pseudomonas aeruginosa and other non-fermenting Gram-negative bacteria. Polymers 2020, 12, 1818. [Google Scholar] [CrossRef] [PubMed]
- Alfei, S.; Schito, A.M. Positively Charged Polymers as Promising Devices against Multidrug Resistant Gram-Negative Bacteria: A Review. Polymers 2020, 12, 1195. [Google Scholar] [CrossRef] [PubMed]
- Mintzer, M.A.; Dane, E.L.; O’Toole, G.A.; Grinstaff, M.W. Exploiting Dendrimer Multivalency To Combat Emerging and Re-Emerging Infectious Diseases. Mol. Pharm. 2012, 9, 342–354. [Google Scholar] [CrossRef] [PubMed]
- Peleg, A.Y.; Hooper, D.C. Hospital-Acquired Infections Due to Gram-Negative Bacteria. N. Engl. J. Med. 2010, 362, 1804–1813. [Google Scholar] [CrossRef]
- WHO. Antibacterial Agents in Clinical Development: An Analysis of the Antibacterial Clinical Development Pipeline; WHO: Geneva, Switzerland, 2019. [Google Scholar]
- WHO. Antibacterial Agents in Preclinical Development: An Open Access Database; WHO: Geneva, Switzerland, 2019. [Google Scholar]
- WHO. Prioritization of Pathogens to Guide Discovery, Research and Development of New Antibiotics for Drug Resistant Bacterial Infections, Including Tuberculosis; WHO: Geneva, Switzerland, 2017. [Google Scholar]
- Jain, A.; Duvvuri, L.S.; Farah, S.; Beyth, N.; Domb, A.J.; Khan, W. Antimicrobial Polymers. Adv. Healthc. Mater. 2014, 3, 1969–1985. [Google Scholar] [CrossRef]
- Alfei, S.; Signorello, M.G.; Schito, A.; Catena, S.; Turrini, F. Reshaped as polyester-based nanoparticles, gallic acid inhibits platelet aggregation, reactive oxygen species production and multi-resistant Gram-positive bacteria with an efficiency never obtained. Nanoscale Adv. 2019, 1, 4148–4157. [Google Scholar] [CrossRef]
- Alfei, S.; Marengo, B.; Domenicotti, C. Polyester-Based Dendrimer Nanoparticles Combined with Etoposide Have an Improved Cytotoxic and Pro-Oxidant Effect on Human Neuroblastoma Cells. Antioxidants 2020, 9, 50. [Google Scholar] [CrossRef]
- Alfei, S.; Catena, S.; Turrini, F. Biodegradable and biocompatible spherical dendrimer nanoparticles with a gallic acid shell and a double-acting strong antioxidant activity as potential device to fight diseases from “oxidative stress.”. Drug Deliv. Transl. Res. 2020, 10, 259–270. [Google Scholar] [CrossRef]
- Alfei, S.; Marengo, B.; Zuccari, G.; Turrini, F.; Domenicotti, C. Dendrimer Nanodevices and Gallic Acid as Novel Strategies to Fight Chemoresistance in Neuroblastoma Cells. Nanomaterials 2020, 10, 1243. [Google Scholar] [CrossRef]
- Castonguay, A.; Ladd, E.; van de Ven, T.G.M.; Kakkar, A. Dendrimers as bactericides. New J. Chem. 2012, 36, 199–204. [Google Scholar] [CrossRef]
- Bahar, A.A.; Liu, Z.; Totsingan, F.; Buitrago, C.; Kallenbach, N.; Ren, D. Synthetic dendrimeric peptide active against biofilm and persister cells of Pseudomonas aeruginosa. Appl. Microbiol. Biotechnol. 2015, 99, 8125–8135. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.; Qu, H.; Ma, M.; Xu, Z.; Xu, P.; Fang, Y.; Xu, T. Polyamidoamine (PAMAM) dendrimers as biocompatible carriers of quinolone antimicrobials: An in vitro study. Eur. J. Med. Chem. 2007, 42, 1032–1038. [Google Scholar] [CrossRef] [PubMed]
- Wronska, N.; Majoral, J.P.; Appelhans, D.; Bryszewska, M.; Lisowska, K. Synergistic effects of anionic/cationic dendrimers and levofloxacin on antibacterial activities. Molecules 2019, 24, 2894. [Google Scholar] [CrossRef] [PubMed]
- Navath, R.S.; Menjoge, A.R.; Dai, H.; Romero, R.; Kannan, S.; Kannan, R.M. Injectable PAMAM Dendrimer–PEG Hydrogels for the Treatment of Genital Infections: Formulation and in Vitro and in Vivo Evaluation. Mol. Pharm. 2011, 8, 1209–1223. [Google Scholar] [CrossRef]
- Stenström, P.; Hjorth, E.; Zhang, Y.; Andrén, O.C.J.; Guette-Marquet, S.; Schultzberg, M.; Malkoch, M. Synthesis and in Vitro Evaluation of Monodisperse Amino-Functional Polyester Dendrimers with Rapid Degradability and Antibacterial Properties. Biomacromolecules 2017, 18, 4323–4330. [Google Scholar] [CrossRef]
- Quadir, M.A.; Haag, R. Biofunctional nanosystems based on dendritic polymers. J. Control. Release 2012, 161, 484–495. [Google Scholar] [CrossRef]
- Dincer, S.; Uslu, F.M.; Delik, A. Antibiotic Resistance in Biofilm. In Bacterial Biofilms; IntechOpen: London, UK, 2020; pp. 1–14. [Google Scholar] [CrossRef]
- Kostakioti, M.; Hadjifrangiskou, M.; Hultgren, S.J. Bacterial biofilms: Development, dispersal, and therapeutic strategies in the dawn of the postantibiotic era. Cold Spring Harb. Perspect. Med. 2013, 3, a010306. [Google Scholar] [CrossRef]
- Klebensberger, J.; Birkenmaier, A.; Geffers, R.; Kjelleberg, S.; Philipp, B. SiaA and SiaD are essential for inducing autoaggregation as a specific response to detergent stress in Pseudomonas aeruginosa. Environ. Microbiol. 2009, 11, 3073–3086. [Google Scholar] [CrossRef]
- Jamal, M.; Ahmad, W.; Andleeb, S.; Jalil, F.; Imran, M.; Nawaz, M.A.; Hussain, T.; Ali, M.; Rafiq, M.; Kamil, M.A. Bacterial biofilm and associated infections. J. Chi. Med. Assoc. 2018, 81, 7–11. [Google Scholar] [CrossRef]
- Hathroubi, S.; Mekni, M.A.; Domenico, P.; Nguyen, D.; Jacques, M. Biofilms: Microbial Shelters against Antibiotics. Microb. Drug Resistance 2016, 23, 147–156. [Google Scholar] [CrossRef]
- Gebreyohannes, G.; Nyerere, A.; Bii, C.; Sbhatu, D.B. Challenges of intervention, treatment, and antibiotic resistance of biofilm-forming microorganisms. Heliyon 2019, 5, e02192. [Google Scholar] [CrossRef] [PubMed]
- Nadell, C.D.; Drescher, K.; Wingreen, N.S.; Bassler, B.L. Extracellular matrix structure governs invasion resistance in bacterial biofilms. ISME J. 2015, 9, 1700–1709. [Google Scholar] [CrossRef]
- Colvin, K.M.; Gordon, V.D.; Murakami, K.; Borlee, B.R.; Wozniak, D.J.; Wong, G.C.L.; Parsek, M.R. The Pel Polysaccharide Can Serve a Structural and Protective Role in the Biofilm Matrix of Pseudomonas aeruginosa. PLOS Pathog. 2011, 7, e1001264. [Google Scholar] [CrossRef] [PubMed]
- Anderl, J.N.; Franklin, M.J.; Stewart, P.S. Role of antibiotic penetration limitation in Klebsiella pneumoniae biofilm resistance to ampicillin and ciprofloxacin. Antimicrob. Agents Chemother. 2000, 44, 1818–1824. [Google Scholar] [CrossRef]
- Bagge, N.; Hentzer, M.; Andersen, J.B.; Ciofu, O.; Givskov, M.; Høiby, N. Dynamics and spatial distribution of beta-lactamase expression in Pseudomonas aeruginosa biofilms. Antimicrob. Agents Chemother. 2004, 48, 1168–1174. [Google Scholar] [CrossRef] [PubMed]
- Mulcahy, H.; Charron-Mazenod, L.; Lewenza, S. Extracellular DNA Chelates Cations and Induces Antibiotic Resistance in Pseudomonas aeruginosa Biofilms. PLoS Pathog. 2008, 4, e1000213. [Google Scholar] [CrossRef]
- Lewenza, S. Extracellular DNA-induced antimicrobial peptide resistance mechanisms in Pseudomonas aeruginosa. Front. Microbiol. 2013, 4, 21. [Google Scholar] [CrossRef]
- Johnson, L.; Mulcahy, H.; Kanevets, U.; Shi, Y.; Lewenza, S. Surface-Localized Spermidine Protects the Pseudomonas aeruginosa Outer Membrane from Antibiotic Treatment and Oxidative Stress. J. Bacteriol. 2012, 194, 813. [Google Scholar] [CrossRef]
- Keren, I.; Mulcahy, L.R.; Lewis, K. Persistent eradication: Lessons from the world of natural products. In Natural Product Biosynthesis by Microorganisms and Plants, Part C; Hopwood, D.A., Ed.; Springer: Berlin, Germany, 2012; p. 449. [Google Scholar] [CrossRef]
- Dörr, T.; Vulić, M.; Lewis, K. Ciprofloxacin Causes Persister Formation by Inducing the TisB toxin in Escherichia coli. PLoS Biol. 2010, 8, e1000317. [Google Scholar] [CrossRef]
- Nguyen, D.; Joshi-Datar, A.; Lepine, F.; Bauerle, E.; Olakanmi, O.; Beer, K.; McKay, G.; Siehnel, R.; Schafhauser, J.; Wang, Y.; et al. Active Starvation Responses Mediate Antibiotic Tolerance in Biofilms and Nutrient-Limited Bacteria. Science 2011, 334, 982. [Google Scholar] [CrossRef]
- Zheng, Z.; Stewart, P.S. Growth limitation of Staphylococcus epidermidis in biofilms contributes to rifampin tolerance. Biofilms 2004, 1, 31–35. [Google Scholar] [CrossRef][Green Version]
- Pagès, J.-M.; James, C.E.; Winterhalter, M. The porin and the permeating antibiotic: A selective diffusion barrier in Gram-negative bacteria. Nat. Rev. Microbiol. 2008, 6, 893–903. [Google Scholar] [CrossRef]
- Gillis, R.J.; White, K.G.; Choi, K.-H.; Wagner, V.E.; Schweizer, H.P.; Iglewski, B.H. Molecular Basis of Azithromycin-Resistant Pseudomonas aeruginosa Biofilms. Antimicrob. Agents Chemother. 2005, 49, 3858. [Google Scholar] [CrossRef] [PubMed]
- Wilton, M.; Charron-Mazenod, L.; Moore, R.; Lewenza, S. Extracellular DNA Acidifies Biofilms and Induces Aminoglycoside Resistance in Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 2015, 60, 544–553. [Google Scholar] [CrossRef]
- Dalton, T.; Dowd, S.E.; Wolcott, R.D.; Sun, Y.; Watters, C.; Griswold, J.A.; Rumbaugh, K.P. An In Vivo Polymicrobial Biofilm Wound Infection Model to Study Interspecies Interactions. PLoS ONE 2011, 6, e27317. [Google Scholar] [CrossRef]
- Perez, A.C.; Pang, B.; King, L.B.; Tan, L.; Murrah, K.A.; Reimche, J.L.; Wren, J.T.; Richardson, S.H.; Ghandi, U.; Swords, W.E. Residence of Streptococcus pneumoniae and Moraxella catarrhalis within polymicrobial biofilm promotes antibiotic resistance and bacterial persistence in vivo. Pathog. Dis. 2014, 70, 280–288. [Google Scholar] [CrossRef] [PubMed]
- Ryan, R.P.; Fouhy, Y.; Garcia, B.F.; Watt, S.A.; Niehaus, K.; Yang, L.; Tolker-Nielsen, T.; Dow, J.M. Interspecies signalling via the Stenotrophomonas maltophilia diffusible signal factor influences biofilm formation and polymyxin tolerance in Pseudomonas aeruginosa. Mol. Microbiol. 2008, 68, 75–86. [Google Scholar] [CrossRef] [PubMed]
- VanKoten, H.W.; Dlakic, W.M.; Engel, R.; Cloninger, M.J. Synthesis and Biological Activity of Highly Cationic Dendrimer Antibiotics. Mol. Pharm. 2016, 13, 3827–3834. [Google Scholar] [CrossRef]
- Wang, L.; Erasquin, U.J.; Zhao, M.; Ren, L.; Zhang, M.Y.; Cheng, G.J.; Wang, Y.; Cai, C. Stability, antimicrobial activity, and cytotoxicity of poly (amidoamine) dendrimers on titanium substrates. ACS Appl. Mat. Interfaces 2011, 3, 2885–2894. [Google Scholar] [CrossRef]
- Carlmark, A.; Malmström, E.; Malkoch, M. Dendritic architectures based on bis-MPA: Functional polymeric scaffolds for application-driven research. Chem. Soc. Rev. 2013, 42, 5858–5879. [Google Scholar] [CrossRef]
- Khalid, K.; Tan, X.; Zaid, H.F.M.; Tao, Y.; Chew, C.L.; Chu, D.-T.; Lam, M.K.; Ho, Y.-C.; Lim, J.W.; Wei, L.C. Advanced in developmental organic and inorganic nanomaterial: A review. Bioengineered 2020, 11, 328–355. [Google Scholar] [CrossRef] [PubMed]
- Freeman, E.C.; Weiland, L.M.; Meng, W.S. Modeling the proton sponge hypothesis: Examining proton sponge effectiveness for enhancing intracellular gene delivery through multiscale modeling. J. Biomater. Sci. Polym. Ed. 2013, 24, 398–416. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Lopina, S.T. Penicillin V-conjugated PEG-PAMAM star polymers. J. Biomater. Sci. Polym. Ed. 2003, 14, 1043–1056. [Google Scholar] [CrossRef]
- Xue, X.; Chen, X.; Mao, X.; Hou, Z.; Zhou, Y.; Bai, H.; Meng, J.; Da, F.; Sang, G.; Wang, Y.; et al. Amino-Terminated Generation 2 Poly(amidoamine) Dendrimer as a Potential Broad-Spectrum, Nonresistance-Inducing Antibacterial Agent. AAPS J. 2013, 15, 132–142. [Google Scholar] [CrossRef] [PubMed]
- Gholami, M.; Mohammadi, R.; Arzanlou, M.; Akbari Dourbash, F.; Kouhsari, E.; Majidi, G.; Mohseni, S.M.; Nazari, S. In vitro antibacterial activity of poly (amidoamine)-G7 dendrimer. BMC Infect. Dis. 2017, 17, 395. [Google Scholar] [CrossRef]
- Chen, C.Z.; Tan, N.C.B.; Cooper, S.L. Incorporation of dimethyldodecylammonium chloride functionalities onto poly(propylene imine) dendrimers significantly enhances their antimicrobial properties. Chem. Commun. 1999, 1585–1586. [Google Scholar] [CrossRef]
- Chen, C.Z.; Beck-Tan, N.C.; Dhurjati, P.; van Dyk, T.K.; LaRossa, R.A.; Cooper, S.L. Quaternary ammonium functionalized poly(propylene imine) dendrimers as effective antimicrobials: Structure-activity studies. Biomacromolecules 2000, 1, 473–480. [Google Scholar] [CrossRef]
- Kannan, R.; Prabakaran, P.; Basu, R.; Pindi, C.; Senapati, S.; Muthuvijayan, V.; Prasad, E. Mechanistic Study on the Antibacterial Activity of Self-Assembled Poly(aryl ether)-Based Amphiphilic Dendrimers. ACS Appl. Bio Mater. 2019, 2, 3212–3224. [Google Scholar] [CrossRef]
- Karthikeyan, R.; Kumar, P.V.; Koushik, O.S. Dendrimeric Biocides—A Tool for Effective Antimicrobial Therapy. J. Nanomed. Nanotechnol. 2016, 7, 2. [Google Scholar] [CrossRef]
- Avval, M.M.; Murthy, S.V.; Shashikanth, S. Synthesis and antimicrobial activity evaluation of poly ethylene imine (PEI) dendrimer modified with 1,2,4-oxadiazole derivatives. Int. J. Chem. Pharm. Sci. 2014, 2, 678–683. [Google Scholar]
- Gibney, K.A.; Sovadinova, I.; Lopez, A.I.; Urban, M.; Ridgway, Z.; Caputo, G.A.; Kuroda, K. Poly(ethylene imine)s as antimicrobial agents with selective activity. Macromol. Biosci. 2012, 12, 1279–1289. [Google Scholar] [CrossRef] [PubMed]
- Kalomiraki, M.; Thermos, K.; Chaniotakis, N.A. Dendrimers as tunable vectors of drug delivery systems and biomedical and ocular applications. Int. J. Nanomed. 2015, 11, 1–12. [Google Scholar] [CrossRef]
- Young, A.W.; Liu, Z.; Zhou, C.; Totsingan, F.; Jiwrajka, N.; Shi, Z.; Kallenbach, N.R. Structure and antimicrobial properties of multivalent short peptides. Med. Chem. Comm. 2011, 2, 308–314. [Google Scholar] [CrossRef]
- Milowska, K.; Rybczyńska, A.; Mosiolek, J.; Durdyn, J.; Szewczyk, E.M.; Katir, N.; Brahmi, Y.; Majoral, J.-P.; Bousmina, M.; Bryszewska, M.; et al. Biological Activity of Mesoporous Dendrimer-Coated Titanium Dioxide: Insight on the Role of the Surface–Interface Composition and the Framework Crystallinity. ACS Appl. Mater. Interfaces 2015, 7, 19994–20003. [Google Scholar] [CrossRef]
- Ortega, P.; Cobaleda, B.M.; Hernández-Ros, J.M.; Fuentes-Paniagua, E.; Sánchez-Nieves, J.; Tarazona, M.P.; Copa-Patiño, J.; Soliveri, J.; de la Mata, F.J.; Gómez, R. Hyperbranched polymers versus dendrimers containing a carbosilane framework and terminal ammonium groups as antimicrobial agents. Org. Biomol. Chem. 2011, 9, 5238–5248. [Google Scholar] [CrossRef]
- Pakrudheen, I.; Banu, A.N.; Murugan, E. Cationic amphiphilic dendrimers with tunable hydrophobicity show in vitro activity. Environ. Chem. Lett. 2018, 16, 1513–1519. [Google Scholar] [CrossRef]
- Neelgund, G.M.; Oki, A.; Luo, Z. Antimicrobial activity of CdS and Ag2S quantum dots immobilized on poly(amidoamine) grafted carbon nanotubes. Colloids Surf. B Biointerfaces 2012, 100, 215–221. [Google Scholar] [CrossRef]
- Felczak, A.; Wronska, N.; Janaszewska, A.; Klajnert, B.; Bryszewska, M.; Appelhans, D.; Voit, B.; Rozalska, S.; Lisowska, K. Antimicrobial activity of poly(propylene imine) dendrimers. New J. Chem. 2012, 36, 2215–2222. [Google Scholar] [CrossRef]
- Sun, B.; Slomberg, D.L.; Chudasama, S.L.; Lu, Y.; Schoenfisch, M.H. Nitric oxide-releasing dendrimers as antibacterial agents. Biomacromolecules 2012, 13, 3343–3354. [Google Scholar] [CrossRef]
- Lu, Y.; Slomberg, D.L.; Shah, A.; Schoenfisch, M.H. Nitric oxide-releasing amphiphilic poly (amidoamine) (PAMAM) dendrimers as antibacterial agents. Biomacromolecules 2013, 14, 3589–3598. [Google Scholar] [CrossRef]
- Worley, B.V.; Slomberg, D.L.; Schoenfisch, M.H. Nitric oxide-releasing quaternary ammonium-modified poly (amidoamine) dendrimers as dual action antibacterial agents. Bioconj. Chem. 2014, 25, 918–927. [Google Scholar] [CrossRef] [PubMed]
- Backlund, C.J.; Sergesketter, A.R.; Offenbacher, S.; Schoenfisch, M.H. Antibacterial efficacy of exogenous nitric oxide on periodontal pathogens. J. Dent. Res. 2014, 93, 1089–1094. [Google Scholar] [CrossRef]
- Backlund, C.J.; Worley, B.V.; Schoenfisch, M.H. Anti-biofilm action of nitric oxide-releasing alkyl-modified poly-(amidoamine) dendrimers against Streptococcus mutans. Acta Biomater. 2016, 29, 198–205. [Google Scholar] [CrossRef] [PubMed]
- Worley, B.V.; Schilly, K.M.; Schoenfisch, M.H. Anti-biofilm efficacy of dual-action nitric oxide-releasing alkyl chain modified poly (amidoamine) dendrimers. Mol. Pharm. 2015, 12, 1573–1583. [Google Scholar] [CrossRef] [PubMed]
- Ciepluch, K.; Maciejewska, B.; Gałczyńska, K.; Kuc-Ciepluch, D.; Bryszewska, M.; Appelhans, D.; Drulis-Kawa, Z.; Arabski, M. The influence of cationic dendrimers on antibacterial activity of phage endolysin against P. aeruginosa cells. Biorgan. Chem. 2019, 91, 103121. [Google Scholar] [CrossRef]
- Zhan, J.; Wang, L.; Liu, S.; Chen, J.; Ren, L.; Wang, Y. Antimicrobial Hyaluronic Acid/Poly(amidoamine) Dendrimer Multilayer on Poly(3-hydroxybutyrate-co-4-hydroxybutyrate) Prepared by a Layer-by-Layer Self-Assembly Method. ACS Appl. Mater. Interfaces 2015, 7, 13876–13881. [Google Scholar] [CrossRef]
- Klaykruayat, B.; Siralertmukul, K.; Srikulkit, K. Chemical modification of chitosan with cationic hyperbranched dendritic polyamidoamine and its antimicrobial activity on cotton fabric. Carbohydr. Polym. 2010, 80, 197–207. [Google Scholar] [CrossRef]
- Niederhafner, P.; Bednárová, L.; Buděšínský, M.; Šafařík, M.; Ehala, S.; Ježek, J.; Borovičková, L.; Fučík, V.; Čeřovský, V.; Slaninová, J. Melectin MAPs: The influence of dendrimerization on antimicrobial and hemolytic activity. Amino Acids 2010, 39, 1553–1561. [Google Scholar] [CrossRef]
- Pires, J.; Siriwardena, T.N.; Stach, M.; Tinguely, R.; Kasraian, S.; Luzzaro, F.; Leib, S.L.; Darbre, T.; Reymond, J.-L.; Endimiani, A. In Vitro activity of the novel antimicrobial peptide dendrimer G3KL against multidrug-resistant Acinetobacter baumannii and Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 2015, 59, 7915–7918. [Google Scholar] [CrossRef]
- Polcyn, P.; Zielinska, P.; Zimnicka, M.; Troć, A.; Kalicki, P.; Solecka, J.; Laskowska, A.; Urbanczyk-Lipkowska, Z. Novel antimicrobial peptide dendrimers with amphiphilic surface and their interactions with phospholipids—Insights from mass spectrometry. Molecules 2013, 18, 7120–7144. [Google Scholar] [CrossRef]
- Stach, M.; Maillard, N.; Kadam, R.U.; Kalbermatter, D.; Meury, M.; Page, M.G.P.; Fotiadis, D.; Darbre, T.; Reymond, J.-L. Membrane disrupting antimicrobial peptide dendrimers with multiple amino termini. Med. Chem. Commun. 2012, 3, 86–89. [Google Scholar] [CrossRef]
- Scorciapino, M.A.; Pirri, G.; Vargiu, A.V.; Ruggerone, P.; Giuliani, A.; Casu, M.; Buerck, J.; Wadhwani, P.; Ulrich, A.S.; Rinaldi, A.C. A novel dendrimeric peptide with antimicrobial properties: Structure-function analysis of SB056. Biophys. J. 2012, 102, 1039–1048. [Google Scholar] [CrossRef] [PubMed]
- Batoni, G.; Casu, M.; Giuliani, A.; Luca, V.; Maisetta, G.; Mangoni, M.L.; Manzo, G.; Pintus, M.; Pirri, G.; Rinaldi, A.C.; et al. Rational modification of a dendrimeric peptide with antimicrobial activity: Consequences on membrane-binding and biological properties. Amino Acids 2016, 48, 887–900. [Google Scholar] [CrossRef] [PubMed]
- Serra, I.; Casu, M.; Ceccarelli, M.; Gameiro, P.; Rinaldi, A.C.; Scorciapino, M.A. Effects of amphipathic profile regularization on structural order and interaction with membrane models of two highly cationic branched peptides with β-sheet propensity. Peptides 2018, 105, 28–36. [Google Scholar] [CrossRef] [PubMed]
- Stach, M.; Siriwardena, T.N.; Köhler, T.; Van Delden, C.; Darbre, T.; Reymond, J.-L. Combining topology and sequence design for the discovery of potent antimicrobial peptide dendrimers against multidrug-resistant pseudomonas aeruginosa. Angew. Chem. 2014, 53, 12827–12831. [Google Scholar] [CrossRef] [PubMed]
- Siriwardena, T.N.; Capecchi, A.; Gan, B.-H.; Jin, X.; He, R.; Wei, D.; Ma, L.; Köhler, T.; van Delden, C.; Javor, S.; et al. Optimizing Antimicrobial Peptide Dendrimers in Chemical Space. Angew. Chem. 2018, 57, 8483–8487. [Google Scholar] [CrossRef]
- Siriwardena, T.N.; Lüscher, A.; Köhler, T.; van Delden, C.; Javor, S.; Reymond, J.-L. Antimicrobial Peptide Dendrimer Chimera. Helv. Chim. Acta 2019, 102. [Google Scholar] [CrossRef]
- Chen, X.; Zhang, M.; Zhou, C.; Kallenbach, N.R.; Ren, D. Control of Bacterial Persister Cells by Trp/Arg-Containing Antimicrobial Peptides. Appl. Environ. Microbiol. 2011, 77, 4878. [Google Scholar] [CrossRef]
- Kadam, R.U.; Bergmann, M.; Garg, D.; Gabrieli, G.; Stocker, A.; Darbre, T.; Reymond, J.-L. Structure-Based Optimization of the Terminal Tripeptide in Glycopeptide Dendrimer Inhibitors of Pseudomonas aeruginosa Biofilms Targeting LecA. Chem. Eur. J. 2013, 19, 17054–17063. [Google Scholar] [CrossRef]
- Johansson, E.M.; Crusz, S.A.; Kolomiets, E.; Buts, L.; Kadam, R.U.; Cacciarini, M.; Bartels, K.M.; Diggle, S.P.; Camara, M.; Williams, P.; et al. Inhibition and dispersion of Pseudomonas aeruginosa biofilms by glycopeptide dendrimers targeting the fucose-specific lectin LecB. Chem. Biol. 2008, 15, 1249–1257. [Google Scholar] [CrossRef]
- Kadam, R.U.; Bergmann, M.; Hurley, M.; Garg, D.; Cacciarini, M.; Swiderska, M.A.; Nativi, C.; Sattler, M.; Smyth, A.R.; Williams, P.; et al. A Glycopeptide Dendrimer Inhibitor of the Galactose-Specific Lectin LecA and of Pseudomonas aeruginosa Biofilms. Angew. Chem. Int. Ed. 2011, 50, 10631–10635. [Google Scholar] [CrossRef]
- Kolomiets, E.; Swiderska, M.A.; Kadam, R.U.; Johansson, E.M.V.; Jaeger, K.-E.; Darbre, T.; Reymond, J.-L. Glycopeptide Dendrimers with High Affinity for the Fucose-Binding Lectin LecB from Pseudomonas aeruginosa. ChemMedChem 2009, 4, 562–569. [Google Scholar] [CrossRef] [PubMed]
- Johansson, E.M.V.; Kadam, R.U.; Rispoli, G.; Crusz, S.A.; Bartels, K.-M.; Diggle, S.P.; Cámara, M.; Williams, P.; Jaeger, K.-E.; Darbre, T.; et al. Inhibition of Pseudomonas aeruginosa biofilms with a glycopeptide dendrimer containing D-amino acids. Med. Chem. Commun. 2011, 2, 418–420. [Google Scholar] [CrossRef]
- Visini, R.; Jin, X.; Bergmann, M.; Michaud, G.; Pertici, F.; Fu, O.; Pukin, A.; Branson, T.R.; Thies-Weesie, D.M.E.; Kemmink, J.; et al. Structural Insight into Multivalent Galactoside Binding to Pseudomonas aeruginosa Lectin LecA. ACS Chem. Biol. 2015, 10, 2455–2462. [Google Scholar] [CrossRef] [PubMed]
- Bergmann, M.; Michaud, G.; Visini, R.; Jin, X.; Gillon, E.; Stocker, A.; Imberty, A.; Darbre, T.; Reymond, J.-L. Multivalency effects on Pseudomonas aeruginosa biofilm inhibition and dispersal by glycopeptide dendrimers targeting lectin LecA. Org. Biomol. Chem. 2016, 14, 138–148. [Google Scholar] [CrossRef]
- Michaud, G.; Visini, R.; Bergmann, M.; Salerno, G.; Bosco, R.; Gillon, E.; Richichi, B.; Nativi, C.; Imberty, A.; Stocker, A.; et al. Overcoming antibiotic resistance in Pseudomonas aeruginosa biofilms using glycopeptide dendrimers. Chem. Sci. 2016, 7, 166–182. [Google Scholar] [CrossRef]
- Alfei, S.; Castellaro, S. Synthesis and Characterization of Polyester-Based Dendrimers Containing Peripheral Arginine or Mixed Amino Acids as Potential Vectors for Gene and Drug Delivery. Macromol. Res. 2017, 25, 1172–1186. [Google Scholar] [CrossRef]
- Alfei, S.; Castellaro, S.; Taptue, G.B. Synthesis and NMR characterization of dendrimers based on 2, 2-bis-(hydroxymethyl)-propanoic acid (bis-HMPA) containing peripheral amino acid residues for gene transfection. Org. Commun. 2017, 10, 144–177. [Google Scholar] [CrossRef]
- Alfei, S.; Catena, S. Synthesis and characterization of versatile amphiphilic dendrimers peripherally decorated with positive charged amino acids. Polym. Int. 2018, 67, 1572–1584. [Google Scholar] [CrossRef]
- Alfei, S.; Catena, S. Synthesis and characterization of fourth generation polyester-based dendrimers with cationic amino acids-modified crown as promising water soluble biomedical devices. Polym. Adv. Technol. 2018, 29, 2735–2749. [Google Scholar] [CrossRef]
- Chen, A.; Karanastasis, A.; Casey, K.R.; Necelis, M.; Carone, B.R.; Caputo, G.A.; Palermo, E.F. Cationic Molecular Umbrellas as Antibacterial Agents with Remarkable Cell-Type Selectivity. ACS Appl. Mater. Interfaces 2020, 12, 21270–21282. [Google Scholar] [CrossRef]
- Xu, L.; He, C.; Hui, L.; Xie, Y.; Li, J.-M.; He, W.-D.; Yang, L. Bactericidal Dendritic Polycation Cloaked with Stealth Material via Lipase-Sensitive Intersegment Acquires Neutral Surface Charge without Losing Membrane-Disruptive Activity. ACS Appl. Mater. Interfaces 2015, 7, 27602–27607. [Google Scholar] [CrossRef] [PubMed]
- Leire, E.; Amaral, S.P.; Louzao, I.; Winzer, K.; Alexander, C.; Fernandez-Megia, E.; Fernandez-Trillo, F. Dendrimer mediated clustering of bacteria: Improved aggregation and evaluation of bacterial response and viability. Biomater. Sci. 2016, 4, 998–1006. [Google Scholar] [CrossRef]
- Ladd, E.; Sheikhi, A.; Li, N.; Van de Ven, T.G.; Kakkar, A. Design and Synthesis of Dendrimers with Facile Surface Group Functionalization, and an Evaluation of Their Bactericidal Efficacy. Molecules 2017, 22, 868. [Google Scholar] [CrossRef]
- Fernandez, J.; Acosta, G.; Pulido, D.; Malý, M.; Copa-Patiño, J.L.; Soliveri, J.; Royo, M.; Gómez, R.; Albericio, F.; Ortega, P.; et al. Carbosilane Dendron–Peptide Nanoconjugates as Antimicrobial Agents. Mol. Pharm. 2019, 16, 2661–2674. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-González, R.; Setaro, F.; Gulías, Ò.; Agut, M.; Hahn, U.; Torres, T.; Nonell, S. Cationic phthalocyanine dendrimers as potential antimicrobial photosensitisers. Org. Biomol. Chem. 2017, 15, 9008–9017. [Google Scholar] [CrossRef] [PubMed]
- Abd-El-Aziz, A.S.; Agatemor, C.; Etkin, N.; Overy, D.P.; Lanteigne, M.; McQuillan, K.; Kerr, R.G. Antimicrobial Organometallic Dendrimers with Tunable Activity against Multidrug-Resistant Bacteria. Biomacromolecules 2015, 16, 3694–3703. [Google Scholar] [CrossRef] [PubMed]
| Reasons for Failure of Antibiotics | Biofilm Function | Factors | Bacteria | Inactivated Antibiotics | Ref. |
|---|---|---|---|---|---|
| Hampered antibiotic penetration | Anti-spread barrier | EPS | P. aeruginosa (exopolysaccharides) | Cationic antibiotics aminoglycosides | [26,27] |
| Presence of antibiotic-degrading enzymes | To provide β-lactamases (β-LS) | ↑ β-LS | K. pneumoniae | Ampicillin | [28] |
| P. aeruginosa | Imipenem Ceftazidime | [29] | |||
| Increased biofilm resistance | To provide eDNA | ↑ eDNA ↓ Mg2+ | P. aeruginosa (Spermidine) Salmonella enterica | Cationic Peptides Aminoglycosides | [30,31,32] |
| Presence of persistent cells | To cause gradients in nutrients and oxygen concentration To promote differentiation in cell growth | Endogenous stress TA 1-systems | P. aeruginosa E. coli | Rifampicin Aminoglycosides | [33] |
| Presence of dormant cells | ↓ Functions ↓ Energy ↓ Biosynthesis | E. coli | Fluoroquinolones | [34] | |
| ↑ Resistance to stress | To cause adaptive stress responses by heterogeneity | Changes in component/processes target of antibiotics | P. aeruginosa | Ofloxacin Gentamicin Meropenem Colistin | [35] |
| E. coli K-12 | Ofloxacin | [36] | |||
| ↑ Exporting membrane proteins | To up-regulate the production of some efflux pumps | ↑ Efflux pumps QS | E. coli Enterobacter aerogenes K. pneumonia | Multi-drugs | [37] |
| P. aeruginosa | Azithromycin | [38] | |||
| Genetic diversity | To act as reservoir of genetic diversity by promoting plasmids transfer | Horizontal gene transfer (HGT) eDNA QS | P. aeruginosa | Aminoglycosides | [39] |
| Dendrimer Structure | Name | Structural Features |
|---|---|---|
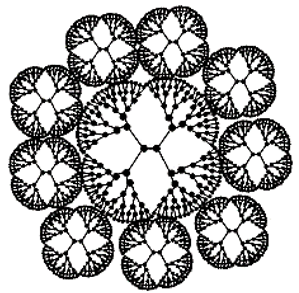 | Tecto Ds | Central dendrimer with multiple peripheral Ds able of differentiate activities |
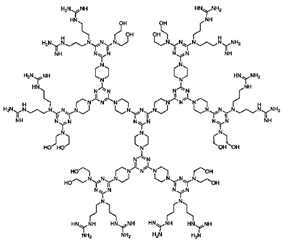 | Triazine Ds | Triazine core with repeated triazine units |
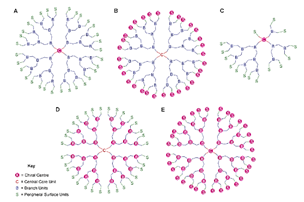 | Chiral Ds | Stereogenic centers within the dendritic structure |
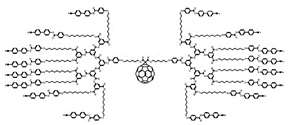 | Liquid crystalline Ds | Block molecules with dendritic core and mesogenic terminal groups |
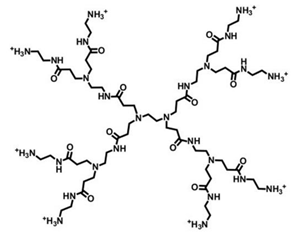 | PAMAM Ds | Amidoamine core and amidoamine repeated units |
 | PPI Ds | Propylene diamine core and similar repeated units |
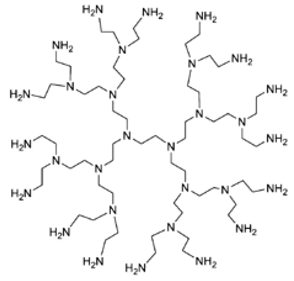 | Poly(ethyleneimine) (PEI-Ds) | Ethylene diamine core and similar repeated units |
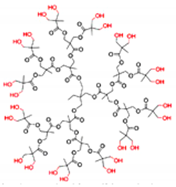 | Polyester-based Ds | AB2 monomer repeated units linked by ester-type hydrolysable bonds |
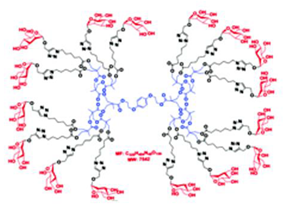 | Glyco Ds | Ds containing saccharide units |
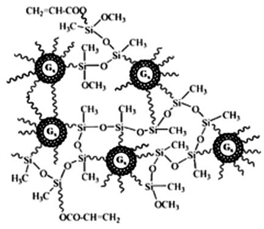 | Polyamidoamine-organosilicon (PAMAMOS) Ds | PAMAM Ds containing silicon organic groups |
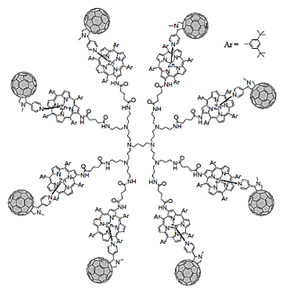 | Fulleropyrrolidine Ds | Ds containing fullero pyrrolidine units |
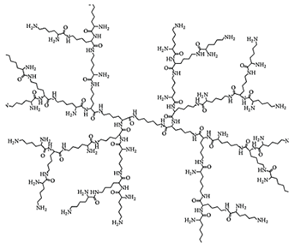 | Poly(lysine) Ds | Ds containing N-lysine repeated units |
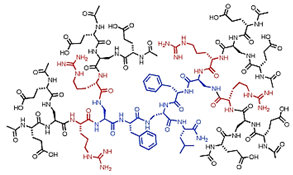 | Peptide Ds | Dendrimer containing amino acids combined by peptide bonds |
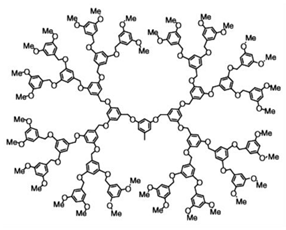 | Polyether Ds | Ds with repeated units linked by ether-type bonds |
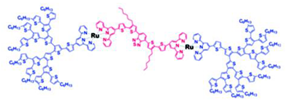 | Metal Ds | Dendrimer with incorporated metal atoms |
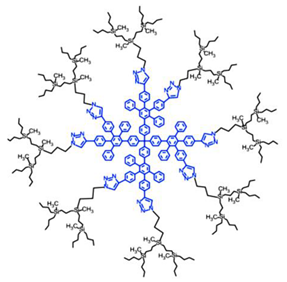 | Hybrid Ds | Ds encompassing more than one type of dendrimer structure |
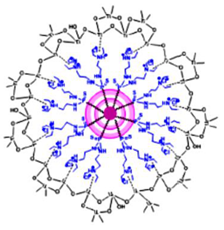 | Ammonium-terminated phosphorous hybrid Ds | Hexachlorocyclo triphosphazene core with hydroxybenzyl phosphor hydrazone repeated units |
 | Mesogenic Ds | Ds containing parts responsible for the formation of a mesophase and liquid crystals |
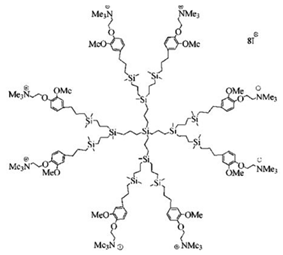 | Carbosilane Ds | Ds containing silicon atoms |
| Dendrimer Class | Mechanism of Action Proposed | Structure Characteristics | Advantages | Target Bacterial Species Toxins | Range of Activity 2,3,4 |
|---|---|---|---|---|---|
| Glyco Ds | Interference with adhesion of toxins to eukaryotic cells (ECs) | 1/2 G 1 PPI Ds 1 G 1 PAMAM Ds | ↑ Interactions with carbohydrate-targets due to dendrimer multivalence | Vibrio cholerae toxin B E. coli enterotoxin | ↑ 1000-times 2,3 |
| ↓ adherence 2,4 | |||||
| 3 G 3,5-di-2-PAE(BA) 5 | Vibrio cholerae toxin B | ↑ 380 × 103-times 2,3 | |||
| Interference with adhesion of bacteria to ECs | 8-valent galabiose 1 G 1 PAMAM Ds 3,5-di-2-PAE(BA) 5 | Streptococcus suis | MIC * 0.3 nM | ||
| Interference with adhesion of bacteria to human erythrocytes | 12-valent mannose 3G 1 PAMAM Ds | Type I fimbriated E. coli | MIC * 19 μM ↑ 400-times 2,4 | ||
| Interference with adhesion of bacteria to ECs | 4-valent α-C-fucosyl-Ds | ↑ Interactions with carbohydrate−targets due to dendrimer multivalence No cytotoxicity ↑ Stability | P. aeruginosa | IC50 § 0.14 μM | |
| Biofilm (P. aeruginosa) | IC50 § 10 μM | ||||
| Cationic Ds | Electrostatic interactions Bacterial membranes disruption | Quaternized PPI Ds | ↑ Positive charge density due to dendrimer multivalence | Recombinant E. coli (TV 1048) | EC50 # 0.14 μM ↑ 100-times 2,3 |
| 1/2 G 1 ammonium-carbosilane Ds | Staphylococcus aureus | MIC * 8 mg/L (2G) 1 mg/mL (1G) | |||
| E. coli | MIC * 64 mg/L (2G) 4 mg/L (1G) | ||||
| 1-3 G 1 ammonium N,N-dimethyl-N′-allyl N′-ethylethylene diamine hydride-terminated carbosilane Ds | ↑ Positive charge density due to dendrimer multivalence Water-soluble | E. coli | MBC $ 1.65 mg/L (3G) 1.70 mg/L (2G) 3.65 mg/L (1G) | ||
| S. aureus | MBC $ 0.82 mg/L (3G) 0.85 mg/L (2G) 1.82 mg/L (1G) | ||||
| 4G 1 nadifloxacin-loaded PAMAM Ds | ↑ Positive charge density due to dendrimer multivalence Antibiotics solubilization by encapsulation | E. coli | As nadifloxacin | ||
| 4G 1 prulifloxacin-loaded PAMAM Ds | ↑ 2-times prulifloxacin | ||||
| PEG/not PEG 5G 1 PAMAM Ds coatings | ↑ Positive charge density due to dendrimer multivalence ↓ Cytotoxicity Prevention of implant-associated infections | P. aeruginosa (ATCC 19660) | EC50 # 1.5 μg/mL | ||
| P. aeruginosa (clinical) | EC50 # 0.9 μg/mL | ||||
| 6% PEG 3G 1 PAMAM Ds coatings | ↑ Positive charge density due to dendrimer multivalence ↓↓↓ Cytotoxicity | P. aeruginosa | MIC * 25 μg/mL | ||
| Hydroxyl-terminated 4G 1 PAMAM Ds | ↑ Positive charge density due to dendrimer multivalence | E. coli | Inhibition of bacteria growth and ascending in uterus 6 | ||
| Amoxicillin-loaded cross-linked 4G 1 PEG-PAMAM Ds | ↑ Positive charge density due to dendrimer multivalence Injectable hydrogels ↑ 72 h residence time No cytotoxicity 240 h sustained drug release 7 | ||||
| Triclosan-loaded 4G 1 PPO 8/PAMAM/PAA 9 layer-by-layer device | ↑ Positive charge density due to dendrimer multivalence ↑ Drug loading than surfactants 20 days sustained drug release 7 | S. aureus | Inhibition of bacteria growth | ||
| PPO 8 triamine core PAMAM Ds | Positive charge density due to dendrimer multivalence | E. coli S. aureus K. pneumoniae B. cereus M. lutens P. vulgaris M. Smegmatis L. monocytogenes P. aeruginosa | MIC * (μg/mL) 3.1–12.5 1.6–6.3 6.1–12.5 6.3–25.0 1.6–12.5 1.6–6.3 12.5 6.3–12.5 6.3–25.0 | ||
| Anionic Ds | Imitating detergent activity | Amphiphilic Ds | ↑ Negative charge density due to dendrimer multivalence Significant selectivity | Bacillus subtilis | EC50 # 41 μM |
| PPO 8 Triamine/anionic PAMAM Ds | ↑ Negative charge density due to dendrimer multivalence Significant selectivity | E. coli S. aureus K. pneumoniae B. cereus M. luteus P. vulgaris M. Smegmatis L. monocytogenes P. aeruginosa | MIC * (μg/mL) 12.5–25.0 12.5 12.5–25.0 12.5–25.0 6.3–12.5 6.3–12.5 12.5–25.0 6.3–12.5 12.5–25.0 | ||
| Peptide-based Ds | Mimicking membrane-active antimicrobial peptides [2] | 2, 4, 8-valent polylysine-Ds with Arg-Leu-Tyr-Arg or Arg-Leu-Tyr-Arg-Lys-Val-Tyr-Gly sequences on surface [D2,4,8(R4), D2,4,8(R8)] | ↑ Positive charge density due to dendrimer multivalence Significant selectivity ↓ Hemolytic activity | E. coli P. aeruginosa P. vulgaris K. oxytoca S. aureus M. luteus E. faecalis | MIC * (μM) D8(R8)/D8(R4) 0.3/0.5–0.7 0.5/0.3–0.9 0.5/0.8–1.3 0.5/0.4–0.8 0.4/0.5–0.6 0.4/0.5–0.7 0.4/0.8–1.3 |
| 4-valent Polylysine-D core with Gln-Lys-Lys-Ile-Arg-Val-Arg-Leu-Ser-Ala sequences on surface | ↑ Positive charge density due to dendrimer multivalence Significant selectivity ↓ Hemolytic activity ↓ Cytotoxicity Good stability in plasma | E. coli K. pneumoniae K. oxytoca E. aerogenes E. doacae P. mirabilis A. baumannii Citrobacter freundii B. cepacia S. aureus P. aeruginosa | MIC * (μg/mL) 10 8 4–16 64 8 4 64→128 16–32 16 64 128 4–8 | ||
| 4-valent Lysine-D core with Trp-Arg on the surface | ↑ Positive charge density due to dendrimer multivalence Significant selectivity ↓ Hemolytic activity ↓ Cytotoxicity ↓ Resistance development Capable of synergistic action | E. coli res. | MIC50 * (μg/mL) 4.5 ↓ 33.5% planktonic ↓ 93.5% biofilm | ||
| S. aureus res. | MIC50 * (μg/mL) 16.0 |
| PAMAM/PPI Ds Generation | Number of NH2 Peripheral Groups |
|---|---|
| 0 | 4 |
| 1 | 8 |
| 2 | 16 |
| 3 | 32 |
| 4 | 64 |
| 5 | 128 |
| 6 | 256 |
| 7 | 512 |
| Dendrimer Structure | Proposed Mechanism | HC50 (μg/mL) | Cytotoxicity IC50 (μg/mL) | Target Bacteria | Activity (μg/mL) |
|---|---|---|---|---|---|
| LVFX (0.1 μg/mL) co-administered Highly maltose-modified 3G PPIs (PPI-G3-DS-Mal) [16] | Electrostatic interactions OM/CM damage OM/CM disruption | N.D. | Cell viability% (100 μM) B14 80 HepG 100 N2a 100 BRL-3A 100 | E. coli | MBC80 (μM) 10 |
| C16-DABCO-loaded mannose-terminated 4G PAMAM Ds [43] | Electrostatic interactions OM/CM damage OM/CM disruption | MHC (μM) 0.6 | A549 human lung carcinoma cells Observed at 1.1 μM | S. aureus S. aureus (biofilm) B. cereus P. aeruginosa E. coli | MIC (μM) 0.13 33.3 0.13 2.0 1.1 |
| G7 PAMAM-D [50] | Electrostatic binding Non-specific OM/CM disruption | N.D. | Cells viability (%) 55 HCT116 57 NIH 3 T3 | P. aeruginosa E. coli A. baumannii S. dysenteriae K. pneumoniae P. mirabilis S. aureus B. subtilis | MIC/MBC 4–8/128–256 4–8/128–256 4–8/128–256 1–2/64–128 4–8/128–256 1–2/64–128 4–8/128–256 2–4/64–128 |
| Self-assembly poly(aryl ether)-PAMAM-based amphiphilic Ds [53] | OM/CM disruption assessed by fluorescent assays OM/CM depolarization | N.D. | IC75 250 (48 h) 2 | E. coli S. aureus | MIC 62 31 |
E. coli S. aureus | MBC 125 31 | ||||
| CdS/Ag2S (QDs)-loaded PAMAM Ds/ MWCNTs [62] | OM/CM damage 5 CM disruption 5 DNA damage by CdS/Ag2S penetration | N.D. | N.D. | E. coli P. aeruginosa S. aureus | GR 1% CdS (20 μg/mL) 87.2 68.9 46.7 |
E. coli P. aeruginosa S. aureus | GR 1% Ag2S (20 μg/mL) 97.8 78.5 55.7 | ||||
| G2-G5 PPI-PO-Ds G2-G4 PPI-PEG-Ds G2-G5 PPI-SO-Ds G3-G5 PPI-PO-NO-Ds G2-G5 PPI-PEG-NO-Ds G2-G5 PPI-SO-NO-Ds G2-G5 PPI-NH3+-Ds [64] | General OM/CM disruption action Additional for NO-Ds: Oxidative and nitrosative stresses Reactive NO byproducts [(N2O3), peroxynitrite (ONOO−)] Membrane destruction via Peroxynitrite-induced lipid peroxidation Protein S-nitrosation DNA deamination | Data in the text | N.D. | P. aeruginosa | MBC (μM) 5 2, 1 3 5 4, 0.5 3 |
| S. aureus | MBC (μM) 2.5 5, 0.5 3 0.5 4, 0.5 3 | ||||
| S. aureus MRSA | 5 2, 2.5 3 0.5 4, 0.25 3 | ||||
| G1 PAMAM-ED-NO-D G1 PAMAM-PE3/7-NO-D G1 PAMAM-PE5/5-NO-D G1 PAMAM-PE7/3-NO-D G1 PAMAM-PO-NO-D G3 PAMAM-PE7/3-NO-D [65] | General OM/CM disruption action Oxidative and nitrosative stresses Reactive NO byproducts [(N2O3), peroxynitrite (ONOO−)] Membrane destruction via Peroxynitrite-induced lipid peroxidation Protein S-nitrosation DNA deamination | N.D. | Cells 6 viability at MBEC values 1st G ED 35% 1st G PE3/7 30% 1st G PE5/5 100% 1st G PE7/3 105% 1st G PO 55% 3rd G PE7/3 90% | P. aeruginosa | MBC 5 |
| P. aeruginosa Biofilm | MBIC 15 | ||||
| G1-G4 NO-releasing-alkyl(QA)-PAMAM Ds [66] | General OM/CM disruption action Oxidative and nitrosative stresses Reactive NO byproducts [(N2O3), peroxynitrite (ONOO−)] Membrane destruction via Peroxynitrite-induced lipid peroxidation Protein S-nitrosation DNA deamination | N.D. | Cells 6 at MBC values 80–110% | P. aeruginosa | MBC 10 |
S. aureus | MBC 10 | ||||
| 1G NO-releasing octyl- and dodecyl-modified PAMAM Ds [68] | Electrostatic dendrimer-bacteria interactions OM/CM damage Fast NO-release kinetics from proton-labile N-diazeniumdiolate NO donors | N.D. | MBC 1000–2000 7 15–20% | S. mutans S. mutans biofilm | MBC (pH = 6.4) 15 MBIC 1000 |
| Dendrimer Structure | Proposed Mechanism | HC50 (μg/mL) | Cytotoxicity IC50 (μg/mL) | Target Bacteria | Activity (μg/mL) |
|---|---|---|---|---|---|
| PEI-D 4[N[(Ts)(2-(methyl)-5-aryl-1,3,4 oxadiazole)]] [55] | OM/CM damage OM/CM disruption by PEI fraction Electron donating action of heterocycle groups | N.D. | N.D. | B. subtilis S. aureus S. epidermidis E. coli X. campestris S. typhi P. aeruginosa | MIC 12.5 12.4 12.5 12.5 12.5 5 12.5 |
| Unmodified b-PEI-D [56] | OM/CM damage OM/CM disruption | >4000 | 27→4000 (1 h) 1 7–2305 (24 h) 1 | E. coli S. aureus | MIC 250 16 |
| Dendrimer Structure | Proposed Mechanism | HC50 (μg/mL) | Cytotoxicity IC50 (μg/mL) | Target Bacteria | Activity (μg/mL) |
|---|---|---|---|---|---|
| Cationic peptide dendrimer RW4D [58] | OM/CM damage OM/CM disruption | 1962 | N.D. | E. coli A. baumannii S. aureus | IC50 3.9 15 42 |
| G2-G4 peptide Ds [73] | OM/CM damage OM/CM disruption | μM <6.3 >200 | N.D. | B. subtilis S. aureus E. coli P. aeruginosa | MIC (μM) 0.37 2.5 0.45 13.8 |
| G3 peptide-D (3GKL) [74] | OM/CM damage OM/CM disruption | 840 | N.D. | A. baumannii P. aeruginosa A. baumannii P. aeruginosa | MIC50/90 8/8 8/8 MBC50/90 8/8 8/8 |
| Amphiphilic peptide Ds with Ornitine core [75] | OM/CM damage OM/CM disruption | μM 100 (0–70%) | S. aureus1 S. aureus2 E. coli P. aeruginosa | MIC (μM) 0.93 0.46 1.85 7.8 | |
| Peptide Ds [76] | Electrostatic binding Insertion into lipid bilayer OM/CM damage OM/CM disruption | MHC 20→10,000 | B. subtilis E. coli P. aeruginosa S. aureus ATCC 25923 S. aureus 887 S. epidermidis ATCC 14990 S. epidermidis J147 En. faecium ATCC 19434 En. faecium Van B E38-10 E. coli HB101 (PAT266) E. coli DC2 | MIC (H1, bH1) # 2 3.9 18 MIC (H1, bH1) # 24 12 24 24 2 8 32 32 | |
| Amphiphilic dimeric peptide-D (SB056) [77] | Electrostatic binding Insertion into lipid bilayer Membranolysis by lipid-induced aggregation | 159 | N.D. | A. baumannii E. cloacae E. coli K. pneumoniae P. aeruginosa S. maltophilia P. mirabilis S. marcescens E. faecalis E. faecium S. epidermidis S. aureus | MIC 4 4 2 8 (MBC) 4 4 8 32 >128 32 8 8 (25.6) 3 32 (51.6) 3 64 (MBC) |
| Dimeric peptide-D (SB056-1) [78] | Electrostatic binding Insertion into lipid bilayer Membranolysis by lipid-induced aggregation | 87 | N.D. | E. coli S. aureus E. coli S. aureus | MIC 16 32 MBC 16 32 |
| G2 KW peptide-D (2D-24) [14] | Electrostatic binding Insertion into lipid bilayer OM/CM damage OM/CM disruption Biofilm and alginate penetration | >1000 | >50 4 | P. aeruginosa (PAO1) P. aeruginosa (PDO300) | MIC GR 5, 46.5 μg/mL 77.5 94% (biofilm) 77.5 94% (biofilm) |
| G3 peptide Ds [80] | Electrostatic binding Non-specific OM/CM damage OM/CM disruption | MHC G3KL 840 DG3KL 680 | N.D. | P. aeruginosa PAO1 E. coli DH5α B. subtilis BR151 P. aeruginosa 6 P. aeruginosa 7 A. baumannii ATCC19606 E. coli W3110 E. aerogenes 13048 | MIC G3KL/DG3KL # 2/4 4/1 3/2 4/4 4/4 8/16 4/2 64/32 |
| G3 peptide-D (T7) [81] | OM/CM damage OM/CM disruption Leakage of cell contents | MHC >2000 | N.D. | P. aeruginosa PAO1 P. aeruginosa 6 E. coli bla 8 E. coli 8 C. freundii bla 8 E. cloacae bla 8 K. pneumoniae 8 K. pneumoniae bla 8 | MIC 4 8 8 8 4 8 16 16 |
| G3 peptide-D (T7) [82] | OM/CM damage OM/CM disruption Leakage of cell contents | MHC 500 | N.D. | P. aeruginosa PAO1 P. aeruginosa 6 E. coli 8 K. pneumoniae 8 K. pneumoniae bla 8 S. aureus 8 | MIC 2 4 8 32 16 32 |
| LecA and LecB ligands glycopeptide Ds (GalAG2, GalBG2, FD2) Heteroglycopeptide-Ds (Het1G2-Het8G2) β-fucoside-D (FucC6G2) Lewisa analogous glycopeptide Ds (LeaxG2) Non-glycosilated peptide Ds (AcG2xK, NG2) [91] | Inactivation of LecA and/or LecB Electrostatic binding OM/CM damage OM/CM disruption | N.D. | N.D. | P. aeruginosa * Biofilm | MBC (μM) 13 (Het7G2) 13 AcG2xK) 2.5 NG2) MBIC (μM) 20 (FD2) 20 (GalAG2) 20 (GalBG2) 13 (Het7G2) 9 (FucC6G2) 30 (LeaxG2) 13 (AcG2xK) 2.5 (NG2) |
| Dendrimer Structure | Proposed Mechanism | HC50 (μg/mL) | Cytotoxicity IC50 (μg/mL) | Target Bacteria | Activity (μg/mL) |
|---|---|---|---|---|---|
| G1-G5 polyester-based alanine Ds NH3 or OH terminated [18] | OM/CM damage OM/CM disruption Leakage of cell contents | N.D. | G1/G2-NH3 NT 203 1,2,3 G3-NH3 NT < 4.3 2,3 <21.3 1 G4-NH3 NT < 0.9 2,3 <21 1 G5-NH3 NT < 1.7 2,3 <42.6 1 | E. coli | MIC G2-NH3 # 203.09 |
| G1-G3 alanine and alkyl (C2-C14)-modified polyester-based dendrons (umbrella molecules) [96] | OM/CM-disrupting action Cationic surfactants-like activity Lytic mechanism | HC50 10 G1/C14 63 G2/C14 5000 G3/C14 | LC50 4 32 μg/mL G3/C14 | E. coli S. aureus P. aeruginosa A. baumannii MRSA E. faecalis | MIC 3.9 (G1/C14) 3.9 (G2/C14) 7.8 (G3/C14) 3.9 (G1/C14) 1.95 (G2/C14) 3.9 (G3/C14) 5.5 (G3/C14) 11.0 (G3/C14) 5.5 (G3/C14) 8.0 (G3/C14) |
| 5G polyester-based cationic Ds (G5K, G5H, G5HK) [1] | Electrostatic binding OM damage Non-lytic bactericidal mechanism | N.D. | Cells viability 4 96.1% | P. aeruginosa (MDR) P. aeruginosa P. aeruginosa ATCC P. putida P. florescens P. straminea A. baumannii A. pittii S. maltophilia | MIC (μM) G5K # 2.1 2.1 2.1 1.0 0.5 1.0 1.1–2.1 2.1 1.1–4.2 |
P. florescens P. straminea A. baumannii 24 A. pittii S. maltophilia 18 | MIC (μM) G5H # 8.3 8.3 8.3 8.3 8.3 | ||||
P. aeruginosa 209 P. aeruginosa 249 P. aeruginosa ATCC P. putida P. florescens P. straminea A. baumannii A. pittii S. maltophilia 18 S. maltophilia 11 S. maltophilia 16 S. maltophilia 19 | MIC (μM) G5HK # 4.2 4.2 8.4 2.1 1.0 1.0 4.2–8.4 2.1 2.1 4.2 8.4 8.4 |
| Dendrimer Structure | Proposed Mechanism | HC50 (μg/mL) | Cytotoxicity IC50 (μg/mL) | Target Bacteria | Activity (μg/mL) |
|---|---|---|---|---|---|
| G2-G3 Ammonium-terminated carbosilane Ds [60] | Membrane impairments and disruption | N.D. | N.D. | E. coli S. aureus E. coli S. aureus | MIC 16 4 MBC 32 8 |
| G2 Ammonium-terminated Ds [97] | Electrostatic binding CM permeabilization | 1024 <10% HC | N.D. | E. coli S. aureus | MBC99.9 3–8 4 |
| G0-G2 Ammonium-terminated Ds [99] | Electrostatic binding Membrane damage Membrane disruption Leakage of cell contents | N.D. | N.D. | E. coli ATCC | MIC μg/mL; μM 0.9; 0.26 (G1) 1–16; 0.12–2 (G2) MBC μg/mL/μM 4–8; 1.1–2.3 (G1) N.D. (G2) |
| G1-G2 ammonium-terminated carbosilane dendrons 3(G1), 4(G2) G1-G2 ammonium-terminated carbosilane dendron nanocomposites 11(3G1), 15(3G2) G1-G2 ammonium-terminated carbosilane dendron nanocomposites 14(4G1), 16(4G2) [100] | Electrostatic binding Membrane damage Membrane disruption Leakage of cell contents | N.D. | N.D. | E. coli | MIC/MBC 16/16 (3) 4/8 (4) 16/16 (11) 16/16 (15) 16/32 (14) 4/4 (16) |
| S. aureus | 128/128 (3) 8/4-8 (4) 64/64 (11) 32/64 (15) 16/16-32 (14) 4-8/8 (16) | ||||
| Ammonium-terminated Ru- and Zn encased Pcs Ds (RuPc, RuPc1, ZnPc, ZnPc1) [101] | Photoinactivating activity Oxidative stress induction by 1O2 | N.D. | N.D. | E. coli | ↓ 6-log10 (μM) ZnPc1 5 1 ZnPc 2.5 1 |
| S. aureus | ZnPc1 1 2 ZnPc 1 2 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Alfei, S.; Schito, A.M. From Nanobiotechnology, Positively Charged Biomimetic Dendrimers as Novel Antibacterial Agents: A Review. Nanomaterials 2020, 10, 2022. https://doi.org/10.3390/nano10102022
Alfei S, Schito AM. From Nanobiotechnology, Positively Charged Biomimetic Dendrimers as Novel Antibacterial Agents: A Review. Nanomaterials. 2020; 10(10):2022. https://doi.org/10.3390/nano10102022
Chicago/Turabian StyleAlfei, Silvana, and Anna Maria Schito. 2020. "From Nanobiotechnology, Positively Charged Biomimetic Dendrimers as Novel Antibacterial Agents: A Review" Nanomaterials 10, no. 10: 2022. https://doi.org/10.3390/nano10102022
APA StyleAlfei, S., & Schito, A. M. (2020). From Nanobiotechnology, Positively Charged Biomimetic Dendrimers as Novel Antibacterial Agents: A Review. Nanomaterials, 10(10), 2022. https://doi.org/10.3390/nano10102022