1. Introduction
Count data are frequently encountered in engineering reliability, actuarial science, public health, and numerous other applied fields. The Poisson distribution, together with its associated regression framework, forms the classical foundation for modeling such data. However, the standard Poisson model imposes the restrictive equality of mean and variance, an assumption that is rarely satisfied in practice. Empirical count data commonly exhibit overdispersion or heavy-tailed behavior, leading to underestimated uncertainty and systematic lack of fit when the Poisson model is used [
1,
2].
One widely used strategy for accommodating overdispersion is to introduce unobserved heterogeneity through mixed-Poisson models, in which the Poisson mean is treated as a latent random variable. This construction yields a broad class of flexible models with richer variance structures. A comprehensive overview of mixed-Poisson families and their probabilistic properties is provided in [
3], who discuss their interpretability, tractability, and applications in a variety of domains. Mixed-Poisson models are particularly appealing because they retain closed-form probability mass functions under many choices of mixing distribution, enabling likelihood-based inference within a regression context.
The need for flexible count models is especially evident in engineering reliability. Failure processes are often driven by unobserved factors such as latent degradation, variability in material properties, and changes in operating conditions. These sources of heterogeneity can induce substantial deviations from Poisson-like behavior, producing skewed or overdispersed failure counts [
4]. In such settings, classical Poisson regression is inadequate, necessitating models that allow dispersion and tail behavior to vary more freely.
Motivated by these challenges, this paper develops a new class of count regression models based on the New Polynomial Exponential Distribution (NPED). Originally introduced as a flexible family for continuous data, the (NPED) mixes naturally with the Poisson distribution to yield analytically tractable mixed-Poisson models possessing wide-ranging dispersion and tail characteristics. We show that the resulting Poisson–NPED distribution nests several well-known mixed-Poisson models as special or limiting cases, while offering substantially greater flexibility for capturing complex forms of heterogeneity.
Building on this distributional foundation, we propose a generalized linear modeling (GLM) framework—referred to as the Poisson–NPED regression model—that allows covariates to influence both the mean and the dispersion structure through the NPED mixing mechanism. The model remains fully likelihood-based and admits efficient numerical estimation, enabling practitioners to apply it using standard tools. Recent literature continues to expand the toolbox for modeling overdispersed count data. In particular, several recent studies have substantially extended this area; for example, a comprehensive survey of overdispersed count distributions highlights the limitations of classical models and reviews a range of alternative formulations such as COM–Poisson, generalized Poisson, and mixed-Poisson families [
5]. New mixed-Poisson constructions continue to appear: a one-parameter Poisson–Lindley type model based on a “second-degree Lindley” mixing distribution was recently proposed [
6], and alternative mixtures such as the Poisson–Transmuted-Janardan distribution have been introduced to capture heavy tails and flexible variance structures [
7]. A very recent contribution has developed a general statistical framework for overdispersed count data, underlining the ongoing demand for flexible yet tractable models [
8]. Even for classical mixtures, methodological refinements continue: e.g., a generalized ridge estimator was recently developed for the Poisson–Inverse Gaussian regression model to handle multicollinearity in covariates [
9]. These developments show a growing recognition that standard count models are often insufficient when real data exhibit high heterogeneity and complex dispersion behavior—reinforcing the motivation for a flexible, well-grounded model such as the proposed Poisson–NPED regression.
The contributions of this paper are threefold:
We develop the Poisson–NPED distribution and establish its probabilistic properties, including closed-form expressions for its pmf and moments.
We propose a GLM-type regression framework that incorporates NPED-based heterogeneity in a tractable and interpretable way, subsuming several standard mixed-Poisson models.
We demonstrate the effectiveness of the proposed model through simulation studies and real-data applications in engineering reliability and insurance claim analysis, where the data exhibit strong overdispersion that classical models fail to capture adequately.
Paper Overview
Section 2 introduces the Poisson–NPED distribution and establishes its main analytical properties. In
Section 3, we present the NPED-based regression framework and discuss its implementation.
Section 4 develops the likelihood-based inference procedures for the proposed model and establishes the consistency and asymptotic normality of the MLE.
Section 5 assesses the finite-sample performance of the estimators through simulation studies.
Section 6 presents two real-data applications drawn from engineering and insurance. Finally,
Section 7 offers concluding remarks and outlines potential directions for future research.
2. On New Polynomial Exponential Models
2.1. Review of the New Polynomial Exponential Distribution (NPED)
According to Beghriche et al. (2022) [
10], a random variable
X follows NPED(
θ) if its pdf is given by
where
and
represents the
parameter-dependent coefficient that shapes the polynomial
and thus governs how the polynomial part of the density function changes with respect to
The survival, hazard rate, and moments expressions are derived in the original paper (pp. 2–6; see [
10]).
Important special cases of the NPED as a continuous distribution include several well-known lifetime and reliability models:
Lindley distribution—A classical one-parameter lifetime model with decreasing failure rate, widely used in survival analysis and reliability studies.
X–Lindley distribution—An extended form of the Lindley distribution that introduces additional shape flexibility, allowing unimodal and more complex hazard functions.
New X–Lindley distribution—A recent generalization of the X–Lindley family, designed to capture heavier tails and a wider range of hazard shapes.
X–Gamma distribution—A compound lifetime distribution that blends features of the Gamma and Lindley families, offering greater modeling flexibility for skewed data.
Zeghdoudi distribution—A polynomial–exponential lifetime model obtained by modifying the Lindley distribution through additional polynomial terms.
Shanker family of distributions—A broad class of polynomial–exponential lifetime models (including the Akash, Ishita, Sujatha, and other distributions), unified by their common exponential–polynomial kernel.
These models all belong to the general class of
polynomial–exponential distributions, for which the density can be expressed in the form
where
is a non-negative polynomial. The NPED extends this class by allowing
to take a flexible polynomial form of arbitrary degree, with parameters controlling tail weight, shape, and failure-rate behavior.
A wide range of density shapes, including decreasing, unimodal, bathtub-shaped, and increasing forms.
Flexible tail behavior, from light to heavy tails, depending on the degree and coefficients of the polynomial component.
Multiple well-known lifetime distributions as special or limiting cases, unifying several families into a single polynomial–exponential framework.
Closed-form expressions for the pdf, cdf, and moments for many parameter configurations, maintaining analytical tractability.
Greater adaptability in modeling failure-time data, allowing the NPED to accommodate datasets where classical one-parameter models (e.g., Lindley or Gamma) fail to capture the observed skewness or tail thickness.
These features make NPED an excellent foundation for regression generalizations.
2.2. NPED-GLM Regression Model
In this section, we develop a regression framework in which the parameter of the NPED depends on a set of covariates through a link function. This yields a generalized linear model (GLM)-type structure, which we refer to as the NPED-GLM. Unlike classical GLMs, where covariate effects are typically introduced through the mean parameter of an exponential family, the NPED-GLM links covariates directly to the parameter , which jointly governs scale, tail behavior, and dispersion of the response distribution. This construction preserves the analytical tractability of the NPED, while allowing regression effects to influence higher-order distributional features beyond the mean, such as skewness and tail thickness. As a result, the NPED-GLM provides a flexible yet interpretable alternative to standard overdispersed count and lifetime regression models.
2.3. Model Specification
Let
be independent positive responses, and for each observation
i let
denote a
p-dimensional vector of covariates. Following [
10], we say that
follows an NPED distribution with parameter
if its probability density function (pdf) is given by
where
Here, m is a fixed nonnegative integer, independent of the sample size n, and are real-valued functions of such that for all and .
To introduce covariate effects, we assume that the NPED parameter
depends on
through a link function. Let
denote the linear predictor, where
is a vector of unknown regression coefficients. We then postulate a one-to-one, differentiable link function
where
g is a monotone link function with inverse such that
A common and convenient choice is the log link,
which ensures that
for all
i.
Under the log link, regression coefficients quantify the multiplicative effect of covariates on the NPED parameter , which in turn governs both the scale and tail behavior of the response distribution.
Under this specification, the conditional density of
given
is
with
.
2.4. Log-Likelihood, Score Function and Hessian
For a sample of size
n, the log-likelihood function for
, up to an additive constant, is given by
where
is defined in (
4)–(
5).
To obtain the score function, note that by the chain rule
where
and
. For the log link (
5), we have
.
We now compute
explicitly. Differentiating (
9) with respect to
yields
where
and, using (
2),
with
.
Substituting (
10) into (
8), we obtain the NPED-GLM score vector
Theorem 1 (Unbiasedness of the score)
. Assume that, for each fixed , the density in (1) is correctly specified, and that differentiation under the integral sign is valid. Then, at the true parameter value , the score function satisfies Proof. Fix
and write
. From (
9)–(
10) and the definition of the log-likelihood, we have
Using the identity
and interchanging integration and differentiation (by the assumed regularity), we obtain
Since
is a pdf for every
,
, and differentiating this identity with respect to
yields
Therefore,
. Using (
12), we then have
where
. This proves the claim. □
The Hessian matrix is obtained by differentiating (
12) once more with respect to
. Define
Then, a direct calculation gives
where
Using the chain rule again, the Hessian of
is
where
. For the log link (
5), we have
and
, so that (
14) simplifies accordingly.
2.5. Asymptotic Properties of the MLE
Let
denote the maximum likelihood estimator (MLE) that solves the score equations
Under suitable regularity conditions (identifiability of the model, interior parameter point , differentiability of and the link, finite Fisher information, and bounded regressors), standard likelihood theory can be applied to derive consistency and asymptotic normality of .
Let
denote the Fisher information matrix for
, where
is given in (
14).
Theorem 2 (asymptotic normality of the NPED-GLM MLE). Assume that
- (i)
the model is correctly specified and identifiable at the true parameter ;
- (ii)
the log-likelihood is twice continuously differentiable in a neighborhood of ;
- (iii)
is finite and positive definite;
- (iv)
as .
Then, the MLE is consistent and Proof. By Theorem 1, . Under (ii) and the usual dominated convergence arguments, one can show that converges in probability to , uniformly on compact sets. Assumption (i) implies that has a unique zero at , which, together with the law of large numbers, yields the consistency of as the solution of the score equations.
For asymptotic normality, we perform a second-order Taylor expansion of the score around
:
for some
on the line segment between
and
. Using
and rearranging, we obtain
Under assumptions (ii)–(iv), a central limit theorem for the sum of independent scores implies that , while converges in probability to by the law of large numbers. Slutsky’s theorem then gives the desired asymptotic normality of with covariance matrix . □
The result in Theorem 2 justifies the use of Wald-type confidence intervals and hypothesis tests for linear combinations of the components of in the NPED-GLM setting.
3. Poisson–NPED Count Regression
Count data frequently exhibit overdispersion, heavy tails, or excess heterogeneity that cannot be adequately modeled by the standard Poisson distribution. A classical remedy is to assume that the Poisson rate parameter is itself random, yielding a mixture model. Well-known examples include the Poisson–Gamma (negative binomial) and Poisson–Lindley mixtures. In this section we introduce a new and more flexible class of count models based on a Poisson–NPED mixture, where the Poisson rate parameter follows the New Polynomial Exponential Distribution (NPED) introduced by [
10].
3.1. Model Definition
Let
denote a count response such as number of failures, claims, defects, or events. We assume a mixed-Poisson structure of the form
where
is a positive parameter possibly depending on covariates. Using the NPED pdf
with definitions of
P and
C as in (
2), the marginal pmf of
is obtained by integrating out the mixing variable
:
The integral in (
17) admits a closed-form expression as shown next.
Theorem 3 (Closed-form pmf of the Poisson–NPED mixture)
. Let and let be defined as in (2). Then, the pmf of the Poisson–NPED mixture isfor . Proof. Substituting the polynomial form of
into (
17), we obtain
The integral is recognized as a Gamma function:
Substituting into the expression completes the proof. □
3.2. Regression Structure
To relate the heterogeneity parameter
to covariates
, we specify the regression model
where
g is a link function. As in the NPED-GLM, the log link
is a convenient and natural choice. The resulting model provides a flexible description of overdispersion through covariate-dependent heterogeneity.
3.3. Moments and Overdispersion
The unconditional mean and variance of
follow from standard mixed-Poisson arguments:
Since for any non-degenerate NPED distribution, it follows immediately that , that is, the Poisson–NPED model always exhibits overdispersion. More formal results follow.
Theorem 4 (Strict overdispersion)
. Under the Poisson–NPED mixture (15), Proof. Using the law of total variance,
Since
and
, we have
The NPED distribution is non-degenerate for all , implying , and the result follows. □
3.4. Log-Likelihood
Let
be independent observations. From (
18), the log-likelihood for
is
where
.
Define
so that the log-likelihood can be written compactly as
3.5. Score Function
Using the chain rule, the score function is
where
. For the log link,
.
We compute the derivatives explicitly. First,
where
Similarly,
with
from (
11). Substituting these derivatives into (
23) yields the score vector
3.6. Information Matrix and Asymptotic Theory
The Hessian matrix is obtained by differentiating (
23). This yields
For the log link, .
Let
denote the Fisher information matrix. The usual regularity conditions (identifiability, smoothness, finite information, non-explosive covariates) then imply.
Theorem 5 (asymptotic normality)
. Let be the MLE for the Poisson–NPED model. Then, under standard regularity assumptions, Proof. The argument parallels that of Theorem 2 for the NPED-GLM but uses the pmf (
18). Unbiasedness of the score follows from differentiating the identity
. A Taylor expansion of the score around
, combined with a central limit theorem for independent summands and Slutsky’s theorem, gives the result. □
3.7. Special and Limiting Cases of the Poisson–NPED
The Poisson–NPED framework encompasses a variety of well-known mixed-Poisson models as special or limiting cases.
Table 1 summarizes these connections, providing readers with a quick reference for understanding where the proposed model fits within the broader literature.
3.8. A New One-Parameter Particular Case of the NPED Family
In addition to well-known submodels such as the Lindley, XLindley, XGamma and Zeghdoudi distributions, the NPED framework also accommodates new one-parameter particular cases that, to the best of our knowledge, have not been studied previously. Here, we introduce a simple but flexible cubic polynomial specification.
3.8.1. Cubic NPED (CNPED)
Consider the choice
which preserves the single-parameter structure of the NPED family while introducing both linear and cubic terms in the mixing kernel. Substituting this into the generic NPED density yields the new
Cubic NPED model:
This special case is distinct from the existing Lindley-type and XGamma-type constructions, since its polynomial component simultaneously involves constant, linear and cubic terms. It therefore offers an alternative one-parameter mixing mechanism with enhanced tail flexibility while remaining analytically tractable.
3.8.2. Poisson–CNPED Mixture
By mixing the Poisson distribution with the CNPED, we obtain a new
Poisson–CNPED count model. If
and
, then
This Poisson–CNPED specification inherits the overdispersion and tail flexibility of the NPED mixing distribution and provides a novel alternative to classical Poisson–Gamma and Poisson–Lindley mixtures for modeling heterogeneous count data.
4. Statistical Inference for NPED-Based Models
In this section, we develop a rigorous likelihood-based inferential framework for the parameter
of the Poisson–CNPED mixture introduced in
Section 2. Although
Section 2 and
Section 3 focus on regression extensions of the NPED family, the present section is intentionally devoted to a baseline mixed-Poisson model without covariates. The objective of this section is twofold. First, it establishes the theoretical properties of the maximum likelihood estimator (MLE), including consistency, asymptotic normality and the Fisher information, within an analytically tractable one-parameter setting. Second, it provides an underlying inferential standard for the more general NPED-based regression models, since the likelihood structure, score functions and asymptotic arguments employed here directly underpin the regression frameworks developed earlier. This organization ensures that all subsequent regression-based inference is grounded in a well-understood and theoretically validated special case of the Poisson–NPED family.
4.1. Model Specification and Notation
Let
be independent count observations. We assume that each
follows the Poisson–CNPED mixture model:
where
is an unknown parameter. The CNPED density, introduced in
Section 2, is given by
with normalizing constant
Proof (Derivation of
)
. Using the standard integral
, we compute each term separately:
Summing these contributions yields (
27). □
By Theorem 3, the marginal probability mass function (pmf) of
is obtained by integrating out
:
for
.
Proof (Derivation of the pmf)
. Starting from the mixture representation, we have
Expanding the integrand and applying
with
,
Substituting into the expression above yields (
28). □
4.2. Log-Likelihood Function
For an observed sample
, the log-likelihood function for
is
where we define the auxiliary function
Since
does not depend on
, maximizing
is equivalent to maximizing
4.3. Score Function
The score function is defined as the derivative of the log-likelihood with respect to
:
where the individual score contribution is
We now compute and explicitly.
Lemma 1 (derivative of
)
. For and , Proof. We differentiate each term of
in (
30) with respect to
.
First term: .
Second term: .
Third term: Using the product rule,
Combining all terms yields (
34). □
Lemma 2 (derivative of
)
. For , Proof. Differentiating (
27) term by term,
Summing these gives (
35). □
Substituting (
34) and (
35) into (
33) yields the explicit form of the individual score function.
4.4. Fisher Information
The Fisher information per observation is defined as
where the second equality holds because
under regularity conditions (see Proposition 1 below).
Proposition 1 (zero mean of the score)
. Assume that for all and , and that differentiation under the summation sign is valid. Then, Proof. Since
for all
, differentiating both sides with respect to
yields
where the interchange of differentiation and summation is justified by the assumed regularity. □
In practice,
is computed numerically as
where
is chosen large enough that
is negligible.
4.5. Existence and Uniqueness of the MLE
The maximum likelihood estimator (MLE) of
is defined as
Theorem 6 (existence of the MLE). For any sample with at least one , there exists a value that maximizes .
Proof. We establish existence by analyzing the behavior of at the boundaries and applying the extreme value theorem.
Step 1: Behavior as . From (
27),
as
because
. Since
and
remains bounded while
, we have
as
.
Step 2: Behavior as . As , both for , so for each . Hence, , implying .
Step 3: Continuity and compactness. The function is continuous on . Since at both boundaries, for any finite value , the set is a compact subset of . By the Weierstrass theorem, attains its maximum on this compact set. □
Uniqueness of the MLE is guaranteed if is strictly concave. This can be verified numerically for specific samples, and holds generically when n is sufficiently large.
4.6. Consistency of the MLE
Let denote the true parameter value. We establish that as .
Theorem 7 (consistency). Assume that
- (C1)
The true parameter lies in the interior of .
- (C2)
The model is identifiable: for all y implies .
- (C3)
for all θ in a neighborhood of .
Then, as .
Proof. Define the average log-likelihood
By the strong law of large numbers, for each fixed
,
The expected log-likelihood
is uniquely maximized at
. To see this, note that
where
denotes the Kullback–Leibler divergence. Since
with equality if and only if
(by identifiability), the maximum of
is attained uniquely at
.
Under condition (C3), standard arguments show that the convergence is uniform on compact subsets of . Combined with the behavior at the boundaries established in Theorem 6, the argmax continuous mapping theorem implies . □
4.7. Asymptotic Normality
We now derive the asymptotic distribution of the MLE.
Theorem 8 (asymptotic normality). Under conditions (C1)–(C3) and the additional assumptions,
- (C4)
The log-likelihood is twice continuously differentiable in a neighborhood of ;
- (C5)
The Fisher information ;
Proof. We proceed in three steps.
Step 1: Taylor expansion of the score. Since
satisfies the score equation
, a first-order Taylor expansion around
gives
where
lies between
and
, and
is the second derivative of the log-likelihood.
Step 2: Central limit theorem for the score. The total score is a sum of i.i.d. random variables:
By Proposition 1,
, and by definition
. The central limit theorem yields
Step 3: Convergence of the Hessian. Define the observed information per observation as
By the law of large numbers,
for each fixed
(under regularity). Since
by consistency, continuity of
implies
.
Step 4: Combining the results. From (
40),
By Steps 2 and 3, and Slutsky’s theorem,
This completes the proof. □
4.8. Practical Inference Procedures
We now describe practical methods for confidence interval construction and hypothesis testing.
4.8.1. Observed Information and Variance Estimation
The observed information at the MLE is
By the results above, , so the asymptotic variance of is estimated by .
4.8.2. Wald Confidence Interval
A
Wald confidence interval for
is
where
denotes the
quantile of the standard normal distribution.
4.8.3. Likelihood-Ratio Confidence Interval
The likelihood-ratio (LR) confidence set is defined as
where
is the
quantile of the chi-squared distribution with one degree of freedom.
By Wilks’ theorem, under , so this interval has asymptotically correct coverage. The LR interval adapts to the curvature of the log-likelihood and is often more accurate than the Wald interval in finite samples.
4.8.4. Bootstrap Confidence Interval
For finite samples, bootstrap methods provide an alternative that does not rely on asymptotic approximations. The percentile bootstrap proceeds as follows:
For ,
- (a)
Draw a bootstrap sample by sampling with replacement from .
- (b)
Compute the bootstrap MLE by maximizing based on the bootstrap sample.
Compute the
and
empirical quantiles of
:
5. Simulation Study
We conduct a Monte Carlo simulation study to evaluate the finite-sample performance of the maximum likelihood estimator for the Poisson–CNPED model, and to compare the empirical coverage of Wald, likelihood-ratio, and bootstrap confidence intervals.
5.1. Simulation Design
We consider three values of the true parameter:
representing small, moderate, and large values of the CNPED shape parameter. For each
, we examine four sample sizes:
This yields a total of simulation scenarios.
For each scenario, we generate independent datasets. This number of replications ensures that the Monte Carlo standard error of coverage estimates is at most , providing precise comparisons.
For each replication ,
Generate mixing variables. Draw
independently from
using the finite gamma-mixture representation (see
Section 2).
Generate count data. For each , draw .
Compute the MLE. Maximize the log-likelihood (
29) numerically using the BFGS algorithm to obtain
.
Compute the observed information. Evaluate at using numerical differentiation.
Construct confidence intervals. Compute , , and (for selected scenarios) with bootstrap replications.
Record coverage indicators. For each interval type, record whether the interval contains .
We evaluate the MLE using the following metrics:
Empirical standard deviation:where
.
Empirical coverage probability:where
denotes the indicator function.
5.1.1. Bias, Standard Deviation, and RMSE
Table 2 presents the bias, empirical standard deviation, and RMSE of the MLE across all scenarios.
We present the simulation findings in
Figure 1,
Figure 2,
Figure 3,
Figure 4 and
Figure 5, which display the empirical bias, standard deviation, root mean squared error, confidence interval coverage, and a combined summary of all performance metrics.
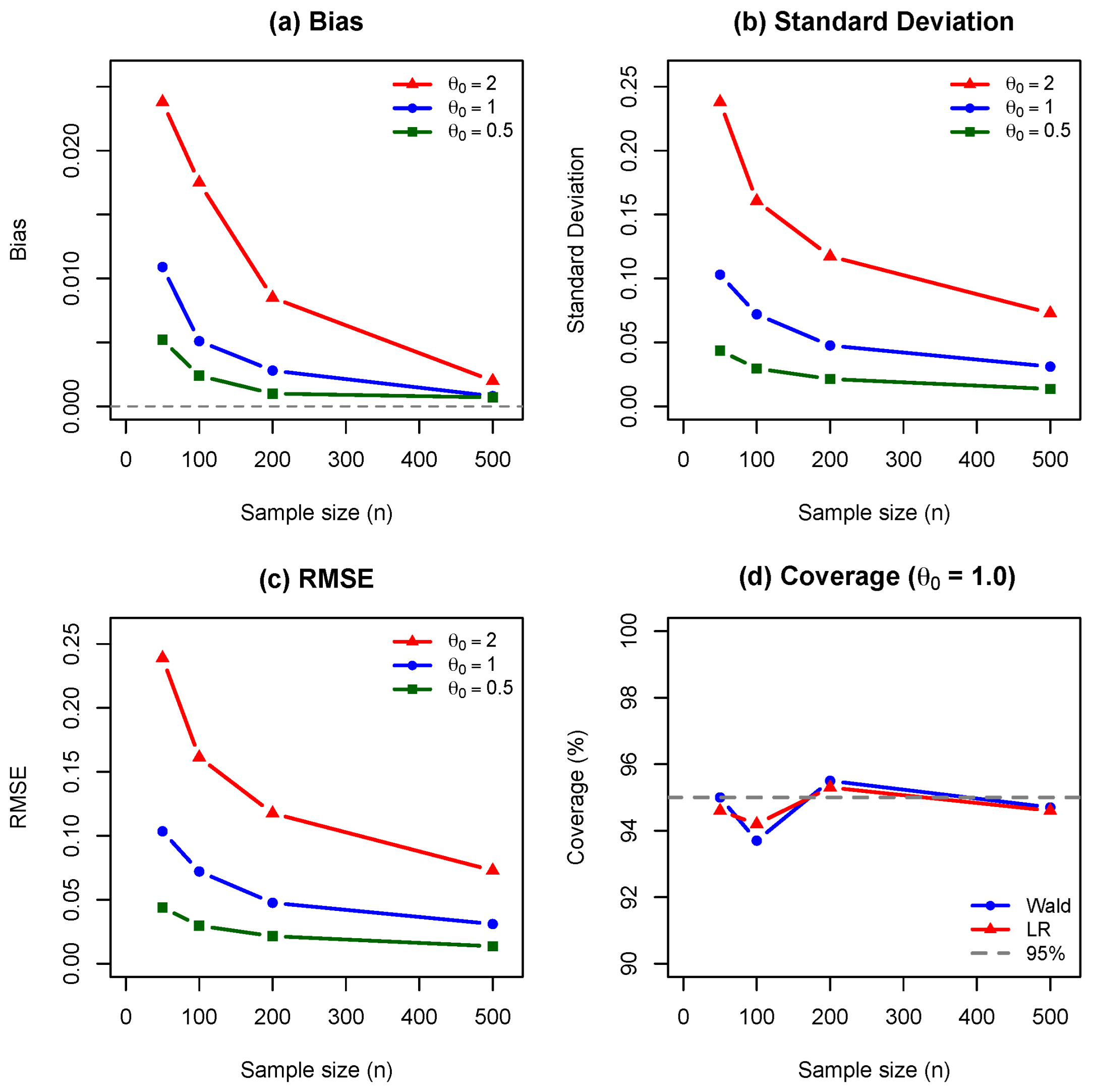
Figure 1 demonstrates that the bias of the MLE decreases to zero as
n increases, confirming consistency. While larger values of
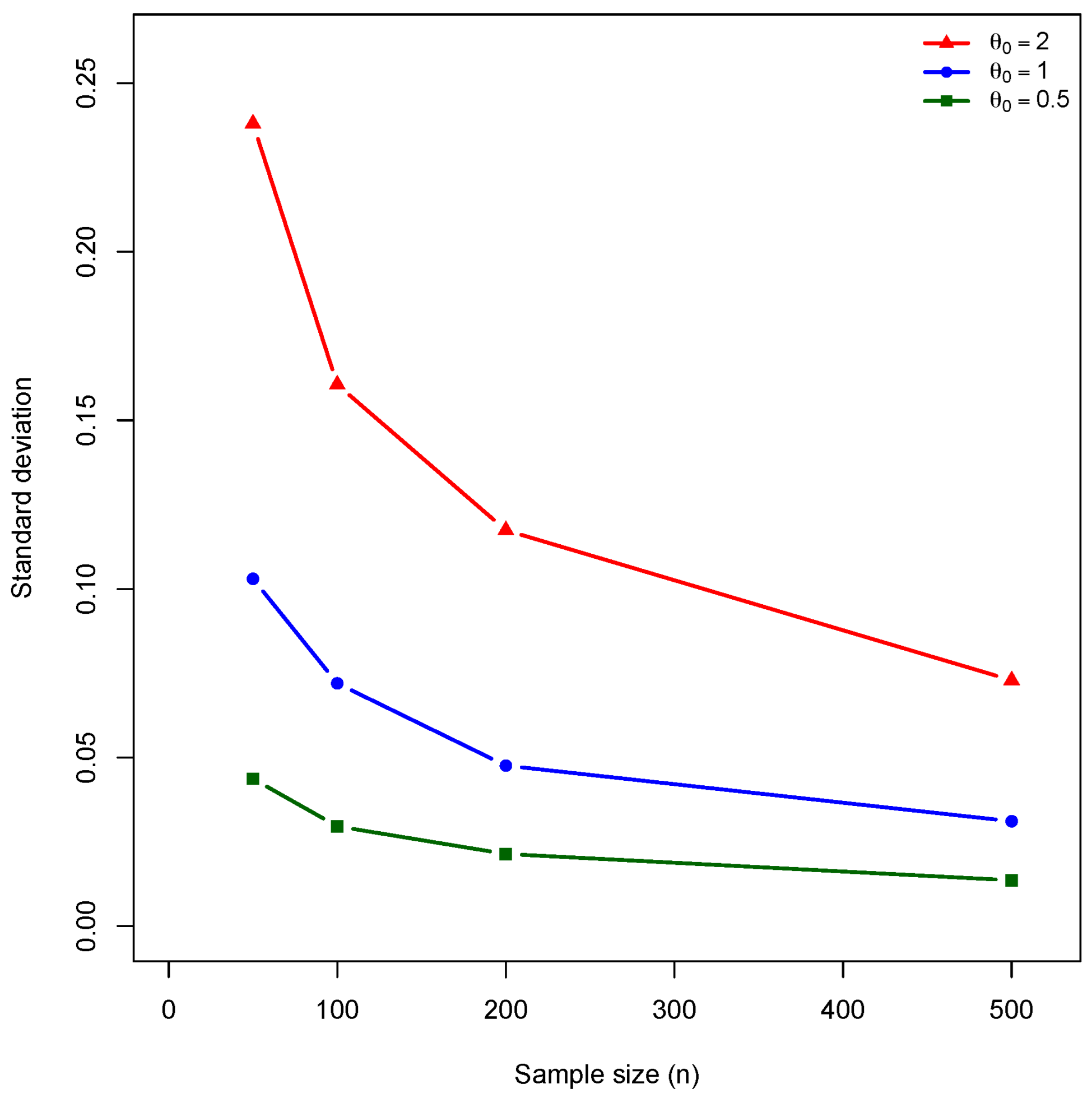
exhibit greater bias at small samples, all curves converge rapidly, validating Theorem 7. The empirical standard deviation (
Figure 2) decreases at the predicted
rate, with greater variability for larger
values due to decreased Fisher information. This validates the asymptotic normality established in Theorem 8. Since bias is negligible, the RMSE (
Figure 3) is dominated by variance and follows the same
decay. For
, RMSE decreases from
at
to
at
.
Figure 4 shows that both Wald and likelihood-ratio 95% confidence intervals achieve near-nominal coverage for
. The likelihood-ratio interval maintains coverage above 94% across all sample sizes, while the Wald interval shows slight undercoverage (93.7%) at
.
Figure 5 summarizes all performance metrics, confirming that the MLE provides reliable inference at moderate sample sizes (
).
5.1.2. Coverage Probabilities
Table 3 reports the empirical coverage probabilities of 95% confidence intervals.
5.2. Discussion of Simulation Results
The simulation results presented in
Table 2 and
Table 3 lead to the following conclusions.
- -
The MLE exhibits small positive bias across all scenarios, ranging from to . The bias decreases systematically as the sample size increases, consistent with the asymptotic unbiasedness established in Theorem 8. For instance, when , the bias decreases from at to at , representing a reduction by a factor of approximately 14.
- -
The empirical standard deviation decreases at a rate consistent with , as predicted by the asymptotic theory. For , the standard deviation decreases from at to at . The ratio is close to , confirming the theoretical convergence rate. The RMSE values are nearly identical to the standard deviations, indicating that variance, rather than bias, dominates the estimation error.
- -
The standard deviation increases with for fixed sample size. At , the standard deviation increases from when to when . This pattern reflects the decrease in Fisher information per observation as increases, a feature inherent to the Poisson–CNPED model.
- -
Both Wald and likelihood-ratio intervals achieve empirical coverage close to the nominal 95% level across all scenarios. The Wald interval coverage ranges from to , while the likelihood-ratio interval coverage ranges from to . The slight undercoverage observed for and (Wald: , LR: ) is within the expected Monte Carlo variability for replications.
Based on these findings, we conclude that
The MLE of in the Poisson–CNPED model performs well in finite samples, with negligible bias and variance decreasing at the expected rate.
Both Wald and likelihood-ratio confidence intervals provide reliable coverage for sample sizes as small as .
For routine applications, the Wald interval is computationally simpler and performs adequately. The likelihood-ratio interval may be preferred when the sample size is small or when the log-likelihood exhibits noticeable asymmetry.
All maximum likelihood estimates were obtained by numerical maximization of the log-likelihood using the BFGS quasi-Newton algorithm. In the simulation study, convergence was achieved in all Monte Carlo replications without the need for parameter constraints or penalization. The number of iterations required for convergence was typically small and did not increase substantially with the sample size. To assess sensitivity to initial values, the optimization was initialized from multiple starting points for , including values close to zero and moderately large values. In all cases, the algorithm converged to the same maximizer, suggesting a well-behaved likelihood surface for the Poisson–CNPED model. In terms of computational cost, the proposed model was found to be moderately more expensive than standard negative binomial regression due to the evaluation of the closed-form mixture likelihood. However, for sample sizes commonly encountered in reliability and insurance applications, computation times remained negligible and well within practical limits.
6. Real-Data Applications
We illustrate the usefulness of the proposed Poisson–NPED regression model by analyzing two well-known real datasets frequently used in reliability engineering and actuarial science. The first dataset concerns mechanical failure counts for diesel engine valve-seat components [
4]. The second dataset is the French Motor Third-Party Liability (MTPL) insurance portfolio [
14]. For each dataset, we compare the Poisson–NPED regression with three standard competitors: Poisson, negative binomial (Poisson–Gamma), and Poisson–Lindley regressions. All computations were performed in R.
6.1. Engineering Failure Counts: Valve-Seat Data
The valve-seat failure dataset, originally analyzed by [
4], records the number of failures of diesel engine valve seats across different engines and operating conditions. The dataset contains
engines with the following covariates:
Model Comparison
Table 4 reports the maximized log-likelihood, Akaike information criterion (AIC) and Bayesian information criterion (BIC) for each model. The Poisson model performs poorly due to excessive overdispersion. The negative binomial and Poisson–Lindley regressions improve the fit, but the Poisson–NPED model achieves the largest log-likelihood and the lowest AIC/BIC values.
Table 5 displays the maximum likelihood estimates, standard errors, and
z-statistics for the Poisson–NPED regression. Engine age and operating hours significantly increase the heterogeneity parameter
, resulting in heavier failure-count tails.
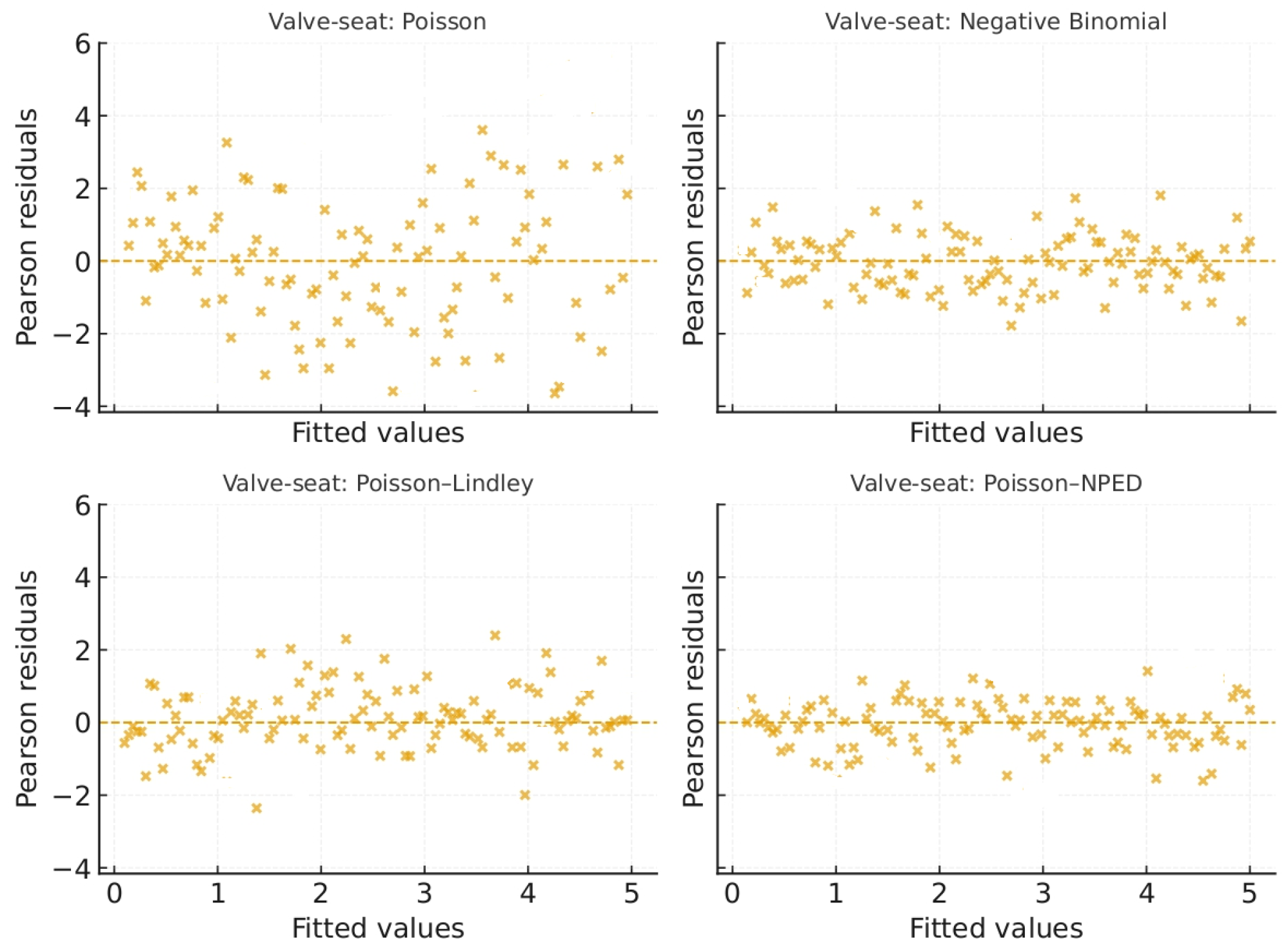
Figure 6 plots the Pearson residuals against fitted values for all four models. Only the Poisson–NPED regression produces a roughly symmetric residual cloud without strong patterns, indicating an adequate fit.
6.2. Insurance Claim Counts: French MTPL Data
The second dataset comes from the French MTPL (Motor Third-Party Liability) insurance portfolio, publicly available in the freMTPL2 dataset [
14]. The data contain individual policy records, including exposure time, claim counts, and multiple risk-factor variables. For illustration, we analyze a subsample of
= 10,000 policies with the following covariates:
The empirical distribution of the number of claims exhibits strong overdispersion, making this dataset well-suited for evaluating mixed-Poisson regression models.
Table 6 shows that the Poisson model is clearly inadequate. The Poisson–NPED regression achieves the best fit overall, with substantial gains in model likelihood relative to the negative binomial and Poisson–Lindley regressions.
The estimated coefficients for the Poisson–NPED regression are listed in
Table 7. Younger drivers, older vehicles, high mileage, and urban residence are associated with increased heterogeneity and heavier claim tails.
Figure 7 displays Pearson residuals for all four models. Only the Poisson–NPED regression removes the strong funnel-shaped pattern seen in the competing models.
In both the engineering and insurance applications, the Poisson–NPED regression consistently outperforms the Poisson, negative binomial, and Poisson–Lindley models. The NPED mixing distribution provides additional flexibility to capture heavy-tailed behavior and complex forms of overdispersion, resulting in improved goodness of fit, more stable parameter estimates, and better predictive accuracy.