Interactive Visual Analysis of Mass Spectrometry Imaging Data Using Linear and Non-Linear Embeddings
Abstract
1. Introduction
- the detection of image regions based on both all relevant information contained in the spectra and the spatial distribution in the imaging space and
- the analysis of the detected image regions to compare different regions with respect to which compounds are decisive for forming the regions and contrasting them against other regions.
2. Background
3. Requirement Analysis
- (T1)
- Which image regions form homogeneous areas with respect to the entire spectral information?
- (T2)
- Which peaks are most descriptive to distinguish image regions?
- (T3)
- What are the distributions of descriptive peaks within an image region?
- (T4)
- How do the spectra/descriptive peaks compare for different image regions?
4. Related Work
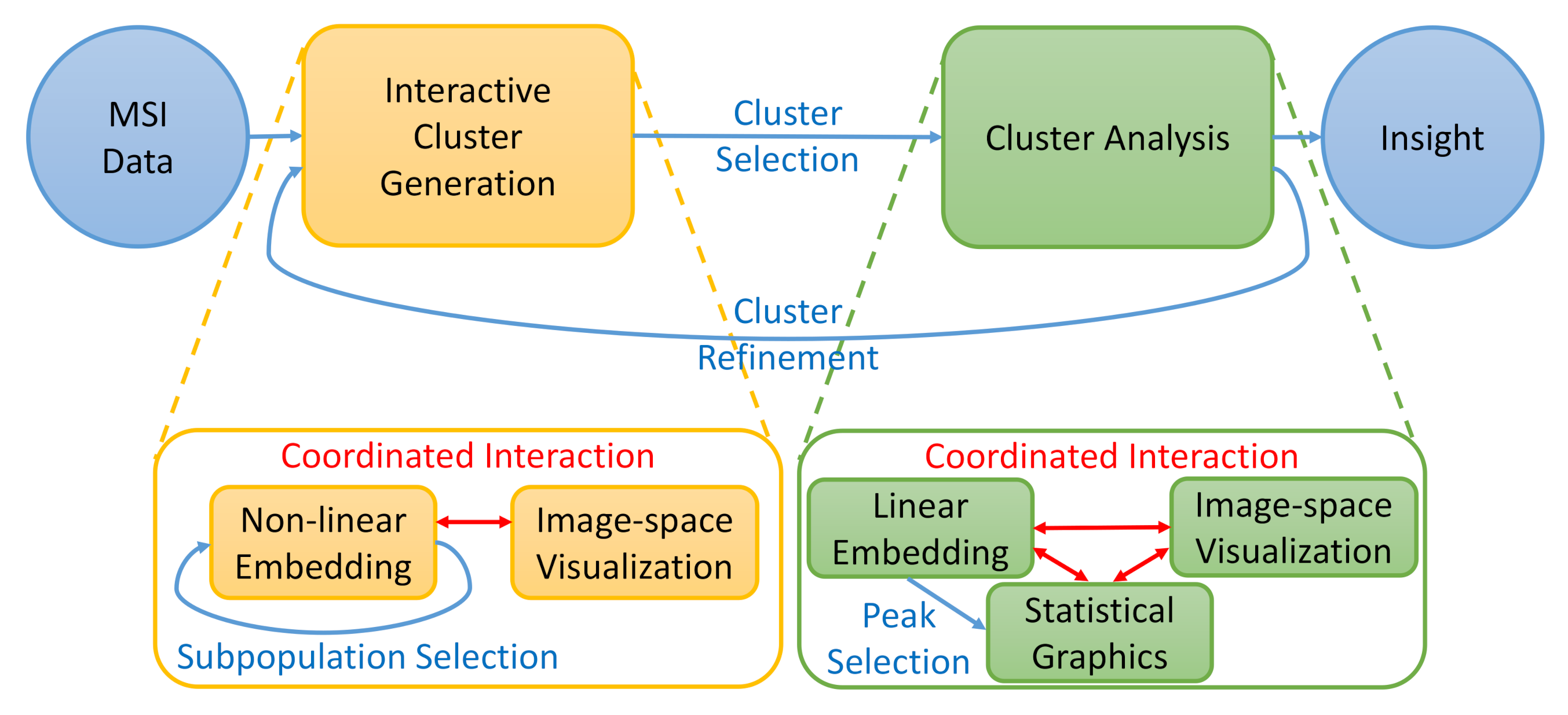
5. Methodology
5.1. Interactive Cluster Generation
5.2. Cluster Analysis
6. Results
7. Discussion and User Feedback
8. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Yuste, R. Fluorescence microscopy today. Nat. Methods 2005, 2, 902. [Google Scholar] [CrossRef] [PubMed]
- Buchberger, A.R.; DeLaney, K.; Johnson, J.; Li, L. Mass Spectrometry Imaging: A Review of Emerging Advancements and Future Insights. Anal. Chem. 2018, 90, 240–265. [Google Scholar] [CrossRef] [PubMed]
- Cole, L.M. (Ed.) Imaging Mass Spectrometry; Springer: New York, NY, USA, 2017. [Google Scholar] [CrossRef]
- Ifa, D.R.; Wu, C.; Ouyang, Z.; Cooks, R.G. Desorption electrospray ionization and other ambient ionization methods: Current progress and preview. Analyst 2010, 135, 669–681. [Google Scholar] [CrossRef] [PubMed]
- Aichler, M.; Walch, A. MALDI Imaging mass spectrometry: Current frontiers and perspectives in pathology research and practice. Lab. Investig. 2015, 95, 422–431. [Google Scholar] [CrossRef]
- Kompauer, M.; Heiles, S.; Spengler, B. Atmospheric pressure MALDI mass spectrometry imaging of tissues and cells at 1.4-mm lateral resolution. Nat. Methods 2016, 14, 90–96. [Google Scholar] [CrossRef]
- Bouschen, W.; Schulz, O.; Eikel, D.; Spengler, B. Matrix vapor deposition/recrystallization and dedicated spray preparation for high-resolution scanning microprobe matrix-assisted laser desorption/ionization imaging mass spectrometry (SMALDI-MS) of tissue and single cells. Rapid Commun. Mass Spectrom. 2010, 24, 355–364. [Google Scholar] [CrossRef]
- Dreisewerd, K. The Desorption Process in MALDI. Chem. Rev. 2003, 103, 395–426. [Google Scholar] [CrossRef]
- Soltwisch, J.; Kettling, H.; Vens-Cappell, S.; Wiegelmann, M.; Muthing, J.; Dreisewerd, K. Mass spectrometry imaging with laser-induced postionization. Science 2015, 348, 211–215. [Google Scholar] [CrossRef]
- Ellis, S.R.; Bruinen, A.L.; Heeren, R.M.A. A critical evaluation of the current state-of-the-art in quantitative imaging mass spectrometry. Anal. Bioanal. Chem. 2014, 406, 1275–1289. [Google Scholar] [CrossRef]
- Ràfols, P.; Vilalta, D.; Brezmes, J.; Cañellas, N.; del Castillo, E.; Yanes, O.; Ramírez, N.; Correig, X. Signal preprocessing, multivariate analysis and software tools for MA(LDI)-TOF mass spectrometry imaging for biological applications. Mass Spectrom. Rev. 2016, 37, 281–306. [Google Scholar] [CrossRef]
- Kriegsmann, J.; Kriegsmann, M.; Casadonte, R. MALDI TOF imaging mass spectrometry in clinical pathology: A valuable tool for cancer diagnostics (Review). Int. J. Oncol. 2014, 46, 893–906. [Google Scholar] [CrossRef] [PubMed]
- Stoeckli, M.; Staab, D.; Schweitzer, A. Compound and metabolite distribution measured by MALDI mass spectrometric imaging in whole-body tissue sections. Int. J. Mass Spectrom. 2007, 260, 195–202. [Google Scholar] [CrossRef]
- WATERS. The Science of What’s Possible. Available online: http://www.waters.com/waters/en_GB/SYNAPT-G2-Si-High-Definition-Mass-Spectrometry/nav.htm?cid=134740622&locale=en_GB (accessed on 30 November 2020).
- Klinkert, I.; Chughtai, K.; Ellis, S.R.; Heeren, R.M. Methods for full resolution data exploration and visualization for large 2D and 3D mass spectrometry imaging datasets. Int. J. Mass Spectrom. 2014, 362, 40–47. [Google Scholar] [CrossRef]
- Avtonomov, D.M.; Raskind, A.; Nesvizhskii, A.I. BatMass: A Java Software Platform for LC–MS Data Visualization in Proteomics and Metabolomics. J. Proteome Res. 2016, 15, 2500–2509. [Google Scholar] [CrossRef] [PubMed]
- Paschke, C.; Leisner, A.; Hester, A.; Maass, K.; Guenther, S.; Bouschen, W.; Spengler, B. Mirion—A software package for automatic processing of mass spectrometric images. J. Am. Soc. Mass Spectrom. 2013, 24, 1296–1306. [Google Scholar] [CrossRef] [PubMed]
- Martin, R.; Markus, S. BioMap. Available online: https://ms-imaging.org/wp/biomap/ (accessed on 30 November 2020).
- Bokhart, M.T.; Nazari, M.; Garrard, K.P.; Muddiman, D.C. MSiReader v1.0: Evolving Open-Source Mass Spectrometry Imaging Software for Targeted and Untargeted Analyses. J. Am. Soc. Mass Spectrom. 2018, 29, 8–16. [Google Scholar] [CrossRef]
- Hayakawa, E.; Fujimura, Y.; Miura, D. MSIdV: A versatile tool to visualize biological indices from mass spectrometry imaging data. Bioinformatics 2016, 32, 3852–3854. [Google Scholar] [CrossRef][Green Version]
- Wijetunge, C.D.; Saeed, I.; Boughton, B.A.; Spraggins, J.M.; Caprioli, R.M.; Bacic, A.; Roessner, U.; Halgamuge, S.K. EXIMS: An improved data analysis pipeline based on a new peak picking method for EXploring Imaging Mass Spectrometry data. Bioinformatics 2015, 31, 3198–3206. [Google Scholar] [CrossRef][Green Version]
- Albert-Jan, Y.; Joris, B.; Marilou, D.; Marnix, K.; Michel, C.; Roeland, L.; Steven, B.; Taco, W. MS Spectre: Mass Spectrometry Analysis Software. Available online: http://ms-spectre.sourceforge.net/ (accessed on 30 November 2020).
- Bemis, K.D.; Harry, A.; Eberlin, L.S.; Ferreira, C.; van de Ven, S.M.; Mallick, P.; Stolowitz, M.; Vitek, O. Cardinal: An R package for statistical analysis of mass spectrometry-based imaging experiments. Bioinformatics 2015, 31, 2418–2420. [Google Scholar] [CrossRef]
- Goracci, L.; Tortorella, S.; Tiberi, P.; Pellegrino, R.M.; Di Veroli, A.; Valeri, A.; Cruciani, G. Lipostar, a Comprehensive Platform-Neutral Cheminformatics Tool for Lipidomics. Anal. Chem. 2017, 89, 6257–6264. [Google Scholar] [CrossRef]
- Zweigniederlassung Bremen der Bruker Daltonik GmbH, University of Bremen. SCiLS. Available online: https://scils.de/ (accessed on 30 November 2020).
- Abdelmoula, W.M.; Balluff, B.; Englert, S.; Dijkstra, J.; Reinders, M.J.; Walch, A.; McDonnell, L.A.; Lelieveldt, B.P. Data-driven identification of prognostic tumor subpopulations using spatially mapped t-SNE of mass spectrometry imaging data. Proc. Natl. Acad. Sci. USA 2016, 113, 12244–12249. [Google Scholar] [CrossRef] [PubMed]
- Abdelmoula, W.M.; Pezzotti, N.; Höllt, T.; Dijkstra, J.; Vilanova, A.; McDonnell, L.A.; Lelieveldt, B. Interactive Visual Exploration of 3D Mass Spectrometry Imaging Data Using Hierarchical Stochastic Neighbor Embedding Reveals Spatiomolecular Structures at Full Data Resolution. J. Proteome Res. 2018, 17, 1054–1064. [Google Scholar] [CrossRef] [PubMed]
- Nunes, M.; Laruelo, A.; Ken, S.; Laprie, A.; Bühler, K.A. A Survey on Visualizing Magnetic Resonance Spectroscopy Data. In Eurographics Workshop on Visual Computing for Biology and Medicine; Viola, I., Bühler, K., Ropinski, T., Eds.; The Eurographics Association: Geneva, Switzerland, 2014. [Google Scholar] [CrossRef]
- Jawad, M.; Molchanov, V.; Linsen, L. Coordinated Image and Featurespace Visualization for Interactive Magnetic Resonance Spectroscopy Imaging Data Analysis. Int. Conf. Inf. Vis. Theory Appl. 2019, 10, 118–128. [Google Scholar]
- Garrison, L.; Vašíček, J.; Grüner, R.; Smit, N.N.; Bruckner, S. SpectraMosaic: An Exploratory Tool for the Interactive Visual Analysis of Magnetic Resonance Spectroscopy Data. In Eurographics Workshop on Visual Computing for Biology and Medicine; Kozlíková, B., Linsen, L., Vázquez, P.P., Lawonn, K., Raidou, R.G., Eds.; The Eurographics Association: Geneva, Switzerland, 2019. [Google Scholar] [CrossRef]
- Murchie, S.L.; Seelos, F.P.; Hash, C.D.; Humm, D.C.; Malaret, E.; McGovern, J.A.; Choo, T.H.; Seelos, K.D.; Buczkowski, D.L.; Morgan, M.F.; et al. Compact Reconnaissance Imaging Spectrometer for Mars investigation and data set from the Mars Reconnaissance Orbiter’s primary science phase. J. Geophys. Res. Planets 2009, 114, E00D07. [Google Scholar] [CrossRef]
- Pelkey, S.M.; Mustard, J.F.; Murchie, S.; Clancy, R.T.; Wolff, M.; Smith, M.; Milliken, R.; Bibring, J.P.; Gendrin, A.; Poulet, F.; et al. CRISM multispectral summary products: Parameterizing mineral diversity on Mars from reflectance. J. Geophys. Res. Planets 2007, 112. [Google Scholar] [CrossRef]
- Blaas, J.; Botha, C.P.; Post, F.H. Interactive Visualization of Multi-Field Medical Data Using Linked Physical and Feature-Space Views. In Proceedings of the 9th Joint Eurographics/IEEE VGTC Conference on Visualization, Norrkoping, Sweden; Eurographics Association: Goslar, Germany, 2007; pp. 123–130. [Google Scholar]
- Linsen, L.; Long, T.V.; Rosenthal, P. Linking Multidimensional Feature Space Cluster Visualization to Multifield Surface Extraction. IEEE Comput. Graph. Appl. 2009, 29, 85–89. [Google Scholar] [CrossRef]
- He, X.; Tao, Y.; Wang, Q.; Lin, H. Multivariate Spatial Data Visualization: A Survey. J. Vis. 2019, 22, 897–912. [Google Scholar] [CrossRef]
- van der Maaten, L.; Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 2008, 9, 2579–2605. [Google Scholar]
- Pezzotti, N.; Lelieveldt, B.P.F.; van der Maaten, L.; Höllt, T.; Eisemann, E.; Vilanova, A. Approximated and User Steerable tSNE for Progressive Visual Analytics. IEEE Trans. Vis. Comput. Graph. 2017, 23, 1739–1752. [Google Scholar] [CrossRef]
- Rubio-Sánchez, M.; Sanchez, A. Axis Calibration for Improving Data Attribute Estimation in Star Coordinates Plots. Vis. Comput. Graph. IEEE Trans. 2014, 20, 2013–2022. [Google Scholar] [CrossRef]
- Elmqvist, N.; Dragicevic, P.; Fekete, J. Rolling the Dice: Multidimensional Visual Exploration using Scatterplot Matrix Navigation. IEEE Trans. Vis. Comput. Graph. 2008, 14, 1539–1548. [Google Scholar] [CrossRef] [PubMed]
- Inselberg, A. The plane with parallel coordinates. Vis. Comput. 1985, 1, 69–91. [Google Scholar] [CrossRef]
- Cox, T.F.; Cox, M.A.A. Multidimensional Scaling; Chapman and Hall: London, UK, 1994. [Google Scholar]
- van der Maaten, L. Accelerating t-SNE using tree-based algorithms. J. Mach. Learn. Res. 2014, 15, 3221–3245. [Google Scholar]
- Peano, G. Sur une courbe, qui remplit toute une aire plane. Math. Ann. 1890, 36, 157–160. [Google Scholar] [CrossRef]
- Sierpínski, W. Sur une nouvelle courbe continue qui remplit toute une aire plane. Bull. Acad. Sci. Crac. 1912, 462–478. [Google Scholar]
- Hilbert, D. Über die stetige Abbildung einer Linie auf ein Flächenstück. In Dritter Band: Analysis· Grundlagen der Mathematik· Physik Verschiedenes; Springer: Berlin/Heidelberg, Germany, 1935; pp. 1–2. [Google Scholar]
- Pascucci, V.; Laney, D.E.; Frank, R.J.; Scorzelli, G.; Linsen, L.; Hamann, B.; Gygi, F. Real-time Monitoring of Large Scientific Simulations. In Proceedings of the 2003 ACM Symposium on Applied Computing, Melbourne, FL, USA; ACM: New York, NY, USA, 2003; pp. 194–198. [Google Scholar] [CrossRef]
- Weissenböck, J.; Fröhler, B.; Gröller, E.; Kastner, J.; Heinzl, C. Dynamic Volume Lines: Visual Comparison of 3D Volumes through Space-filling Curves. IEEE Trans. Vis. Comput. Graph. 2019, 25, 1040–1049. [Google Scholar] [CrossRef]
- Holzmüller, D. Efficient neighbor-finding on space-filling curves. arXiv 2017, arXiv:1710.06384. [Google Scholar]
- Skubalska-Rafajłowicz, E. Applications of the space—filling curves with data driven measure—Preserving property. Nonlinear Anal. Theory Methods Appl. 1997, 30, 1305–1310. [Google Scholar] [CrossRef]
- Ye, J. Least Squares Linear Discriminant Analysis. In Proceedings of the 24th International Conference on Machine Learning, Corvalis, OR, USA; ACM: New York, NY, USA, 2007; pp. 1087–1093. [Google Scholar] [CrossRef]
- Molchanov, V.; Linsen, L. Interactive Design of Multidimensional Data Projection Layout. In EuroVis-Short Papers; Elmqvist, N., Hlawitschka, M., Kennedy, J., Eds.; The Eurographics Association: Geneva, Switzerland, 2014. [Google Scholar] [CrossRef]
- Kandogan, E. Star coordinates: A multi-dimensional visualization technique with uniform treatment of dimensions. In Proceedings of the IEEE Information Visualization Symposium, Salt Lake City, UT, USA, 8–13 October 2000; Volume 650, p. 22. [Google Scholar]
- Teoh, S.T.; Ma, K.L. StarClass: Interactive Visual Classification using Star Coordinates. In Proceedings of the 2003 SIAM International Conference on Data Mining, San Francisco, CA, USA, 1–3 May 2003; pp. 178–185. [Google Scholar]
- Chen, K. Optimizing star-coordinate visualization models for effective interactive cluster exploration on big data. Intell. Data Anal. 2014, 18, 117–136. [Google Scholar] [CrossRef]
- Khalid, N.E.A.; Yusoff, M.; Kamaru-Zaman, E.A.; Kamsani, I.I. Multidimensional Data Medical Dataset Using Interactive Visualization Star Coordinate Technique. Procedia Comput. Sci. 2014, 42, 247–254. [Google Scholar] [CrossRef][Green Version]
- Kiyadeh, A.P.H.; Zamiri, A.; Yazdi, H.S.; Ghaemi, H. Discernible visualization of high dimensional data using label information. Appl. Soft Comput. 2015, 27, 474–486. [Google Scholar] [CrossRef]




Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jawad, M.; Soltwisch, J.; Dreisewerd, K.; Linsen, L. Interactive Visual Analysis of Mass Spectrometry Imaging Data Using Linear and Non-Linear Embeddings. Information 2020, 11, 575. https://doi.org/10.3390/info11120575
Jawad M, Soltwisch J, Dreisewerd K, Linsen L. Interactive Visual Analysis of Mass Spectrometry Imaging Data Using Linear and Non-Linear Embeddings. Information. 2020; 11(12):575. https://doi.org/10.3390/info11120575
Chicago/Turabian StyleJawad, Muhammad, Jens Soltwisch, Klaus Dreisewerd, and Lars Linsen. 2020. "Interactive Visual Analysis of Mass Spectrometry Imaging Data Using Linear and Non-Linear Embeddings" Information 11, no. 12: 575. https://doi.org/10.3390/info11120575
APA StyleJawad, M., Soltwisch, J., Dreisewerd, K., & Linsen, L. (2020). Interactive Visual Analysis of Mass Spectrometry Imaging Data Using Linear and Non-Linear Embeddings. Information, 11(12), 575. https://doi.org/10.3390/info11120575





