Chromophoric Dissolved Organic Matter as a Tracer of Fecal Contamination for Bathing Water Quality Monitoring in the Northern Tyrrhenian Sea (Latium, Italy)
Abstract
1. Introduction
2. Materials and Methods
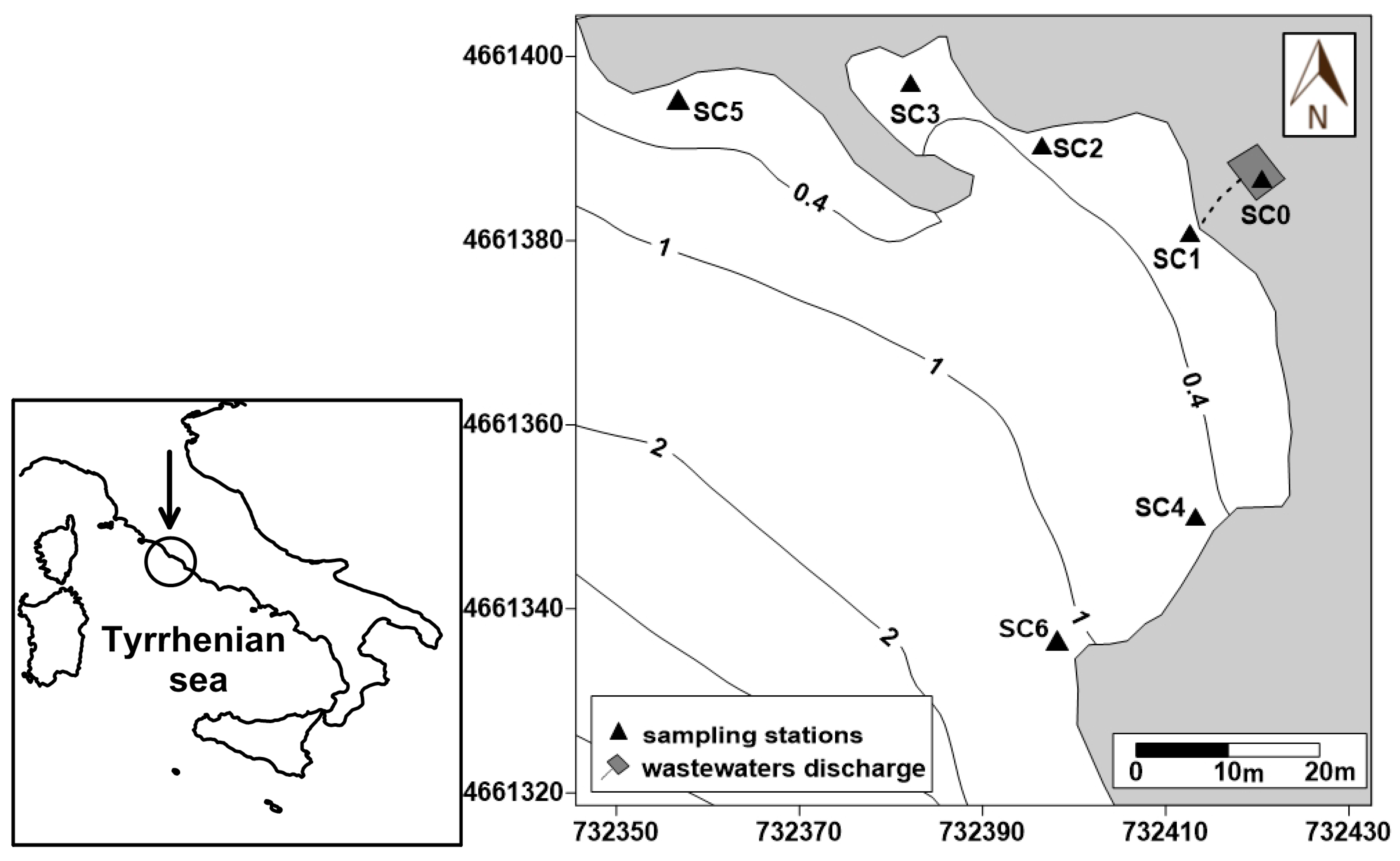
2.1. Study Area and Samplings
2.2. Laboratory Analyses
2.2.1. Chromophoric Dissolved Organic Matter (CDOM) Absorption and Fluorescence Measurements
2.2.2. Microbiological Counts and Extracellular Enzymatic Activities
2.2.3. Statistical Analysis
3. Results
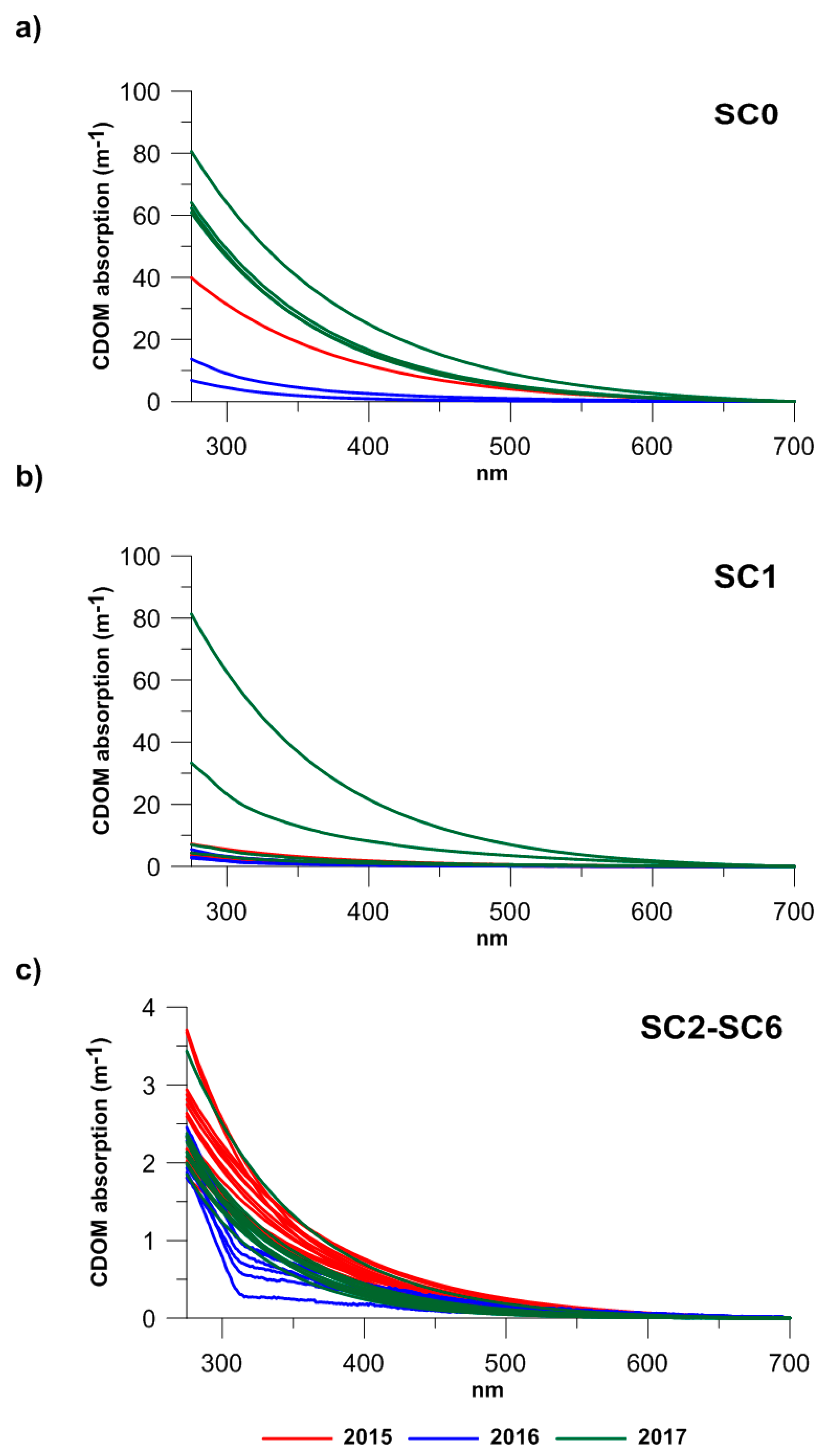
3.1. CDOM Absorption
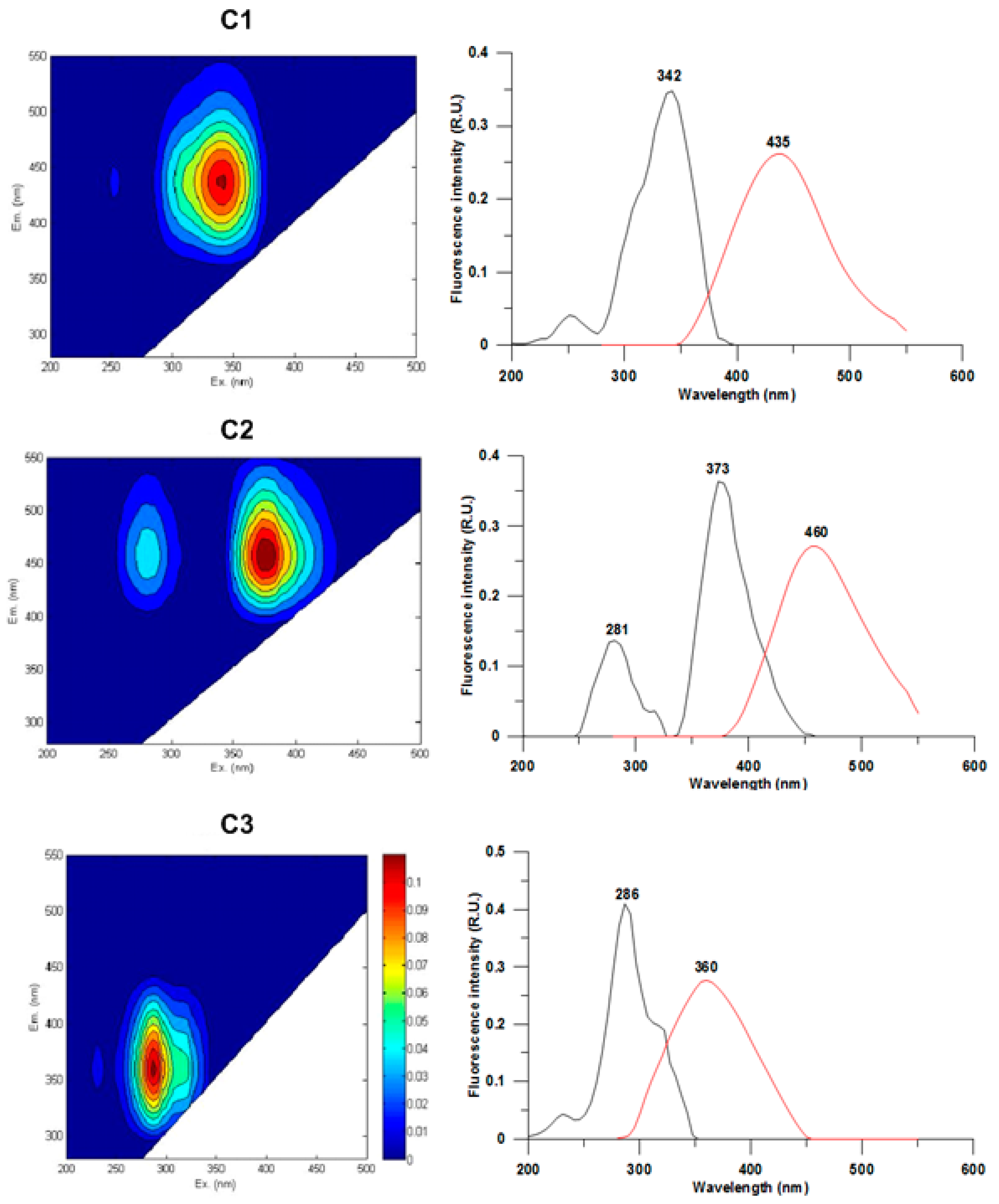
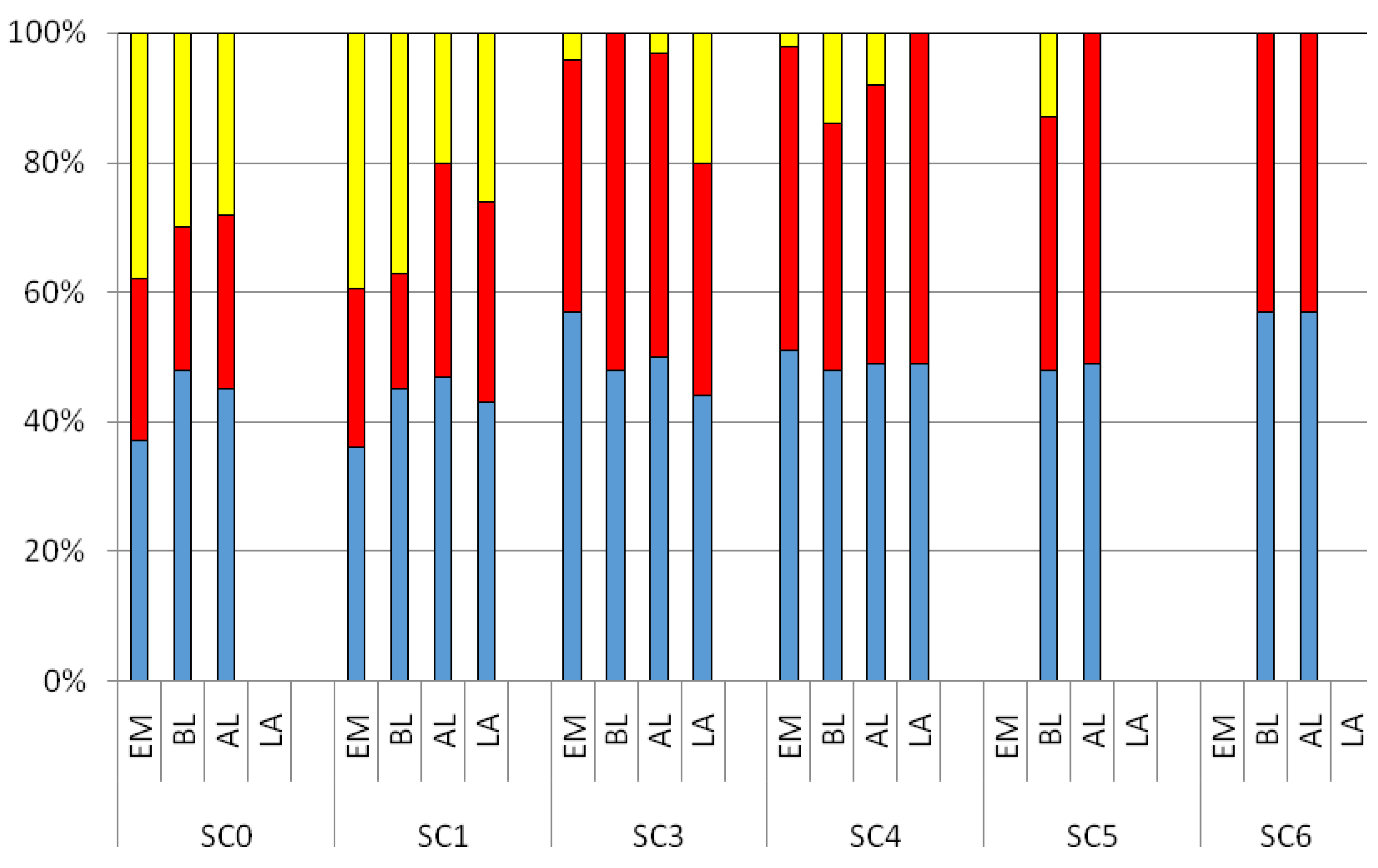
3.2. CDOM Fluorescence
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Holm, S.E.; Windsor, J.G., Jr. Exposure assessment of sewage treatment plant effluent by a selected chemical marker method. Arch. Environ. Contam. Toxicol. 1990, 19, 674–679. [Google Scholar] [CrossRef] [PubMed]
- Chambers, P.A.; Allard, M.; Walker, S.L.; Marsalek, J.; Lawrence, J.; Servos, M.; Busnarda, J.; Munger, K.S.; Jefferson, C.; Kent, R.A.; et al. Impacts of municipal wastewater effluents on Canadian waters: A review. Water Qual. Res. J. Can. 1997, 32, 659–714. [Google Scholar] [CrossRef]
- Singh, K.P.; Mohan, D.; Sinha, S.; Dalwani, R. Impact assessment of treated/untreated wastewater toxicants discharged by sewage treatment plants on health, agricultural, and environmental quality in the wastewater disposal area. Chemosphere 2004, 55, 227–255. [Google Scholar] [CrossRef] [PubMed]
- Brown, C.J.; Knight, B.W.; McMaster, M.E.; Munkittrick, K.R.; Oakes, K.D.; Tetreault, G.R.; Servos, M.R. The effects of tertiary treated municipal wastewater on fish communities of a small river tributary in southern Ontario, Canada. Environ. Pollut. 2011, 159, 1923–1931. [Google Scholar] [CrossRef]
- Tedetti, M.; Longhitano, R.; Garcia, N.; Guigue, C.; Ferretto, N.; Goutx, M. Fluorescence properties of dissolved organic matter in coastal Mediterranean waters influenced by a municipal sewage effluent (Bay of Marseilles, France). Environ. Chem. 2012, 9, 438–449. [Google Scholar] [CrossRef]
- Bonamano, S.; Piermattei, V.; Madonia, A.; Paladini de Mendoza, F.; Pierattini, A.; Martellucci, R.; Stefanì, C.; Zappalà, G.; Caruso, G.; Marcelli, M. The Civitavecchia Coastal Environment Monitoring System (C-CEMS): A new tool to analyse the conflicts between coastal pressures and sensitivity areas. Ocean Sci. 2016, 12, 87–100. [Google Scholar] [CrossRef]
- Odonkor, S.T.; Ampofo, J.K. Escherichia coli as an indicator of bacteriological quality of water: An overview. Microbiol. Res. 2013, 4, e2. [Google Scholar] [CrossRef]
- TEPatCotE Union. Directive 2006/7/EC of the European Parliament and of the Council of 15 February 2006 concerning the management of bathing water quality and repealing Directive 76/160/EEC. Off. J. Eur. Union 2006, 64, 37–51. [Google Scholar]
- Caruso, G.; La Ferla, R.; Azzaro, M.; Zoppini, A.; Marino, G.; Petochi, T.; Corinaldesi, C.; Leonardi, M.; Zaccone, R.; Fonda Umani, S.; et al. Microbial assemblages for environmental quality assessment: Knowledge, gaps and usefulness in the European Marine Strategy Framework Directive. Crit. Rev. Microbiol. 2016, 42, 883–904. [Google Scholar] [CrossRef]
- Sorensen, J.P.R.; Baker, A.; Cumberland, S.A.; Lapworth, D.J.; MacDonald, A.M.; Pedley, S.; Taylor, R.G.; Ward, J.S.T. Real-time detection of faecally contaminated drinking water with tryptophan-like fluorescence: Defining threshold values. Sci. Total Environ. 2018, 622–623, 1250–1257. [Google Scholar] [CrossRef]
- Zaccone, R.; Crisafi, E.; Caruso, G. Evaluation of fecal pollution in coastal Italian waters by immunofluorescence. Aquat. Microb. Ecol. 1995, 9, 79–85. [Google Scholar] [CrossRef][Green Version]
- Caruso, G.; Crisafi, E.; Mancuso, M. Development of an enzyme assay for rapid assessment of Escherichia coli in seawaters. J. Appl. Microbiol. 2002, 93, 548–556. [Google Scholar] [CrossRef] [PubMed]
- Caruso, G.; Manucso, M.; Crisafi, E. Combined fluorescent antibody assay and viability staining for the assessment of the physiological states of Escherichia coli in seawaters. J. Appl. Microbiol. 2003, 95, 225–233. [Google Scholar] [CrossRef] [PubMed]
- Petrenko, A.A.; Jones, B.H.; Dickey, T.D.; LeHaitre, M.; Moore, C. Effects of a sewage plume on the biology, optical characteristics, and particle size distributions of coastal waters. J. Geophys. Res. 1997, 102, 25061. [Google Scholar] [CrossRef]
- Clark, C.D.; O’Connor, A.P.; Foley, D.M.; de Bruyn, W.J. A study of fecal coliform sources at a coastal site using colored dissolved organic matter (CDOM) as a water source tracer. Mar. Pollut. Bull. 2007, 54, 1507–1513. [Google Scholar] [CrossRef]
- Hudson, N.; Baker, A.; Reynolds, D. Fluorescence analysis of dissolved organic matter in natural, waste and polluted waters-a review. River Res. Appl. 2007, 23, 631–649. [Google Scholar] [CrossRef]
- Tedetti, M.; Guigue, C.; Goutx, M. Utilization of a submersible UV fluorometer for monitoring anthropogenic inputs in the Mediterranean coastal waters. Mar. Pollut. Bull. 2010, 60, 350–362. [Google Scholar] [CrossRef]
- Zhuo, J.F.; Guo, W.D.; Deng, X.; Zhang, Z.Y.; Xu, J.; Huang, L.F. Fluorescence excitation-emission matrix spectroscopy of CDOM from Yundang Lagoon and its indication for organic pollution. Spectrosc. Spect. Anal. 2010, 30, 1539–1544. [Google Scholar] [CrossRef]
- Tzortziou, M.; Zeri, C.; Dimitriou, E.; Ding, Y.; Jaffé, R.; Anagnostou, E.; Pitta, E.; Mentzafou, A. Colored dissolved organic matter dynamics and anthropogenic influences in a major transboundary river and its coastal wetland. Limnol. Oceanogr. 2015, 60, 1222–1240. [Google Scholar] [CrossRef]
- Coble, P.G. Marine optical biogeochemistry—The chemistry of ocean color. Chem. Rev. 2007, 107, 402–418. [Google Scholar] [CrossRef]
- Stedmon, C.A.; Markager, S.; Bro, R. Tracing dissolved organic matter in aquatic environments using a new approach to fluorescence spectroscopy. Mar. Chem. 2003, 82, 239–254. [Google Scholar] [CrossRef]
- Murphy, K.R.; Stedmon, C.A.; Waite, T.D.; Ruiz, G.M. Distinguishing between terrestrial and autochthonous organic matter sources in marine environments using fluorescence spectroscopy. Mar. Chem. 2008, 108, 40–58. [Google Scholar] [CrossRef]
- Guo, W.; Yang, L.; Hong, H.; Stedmon, C.A.; Wang, F.; Xu, J.; Xie, Y. Assessing the dynamics of chromophoric dissolved organic matter in a subtropical estuary using parallel factor analysis. Mar. Chem. 2011, 124, 125–133. [Google Scholar] [CrossRef]
- Jørgensen, L.; Stedmon, C.A.; Kragh, T.; Markager, S.; Middelboe, M.; Søndergaard, M. Global trends in the fluorescence characteristics and distribution of marine dissolved organic matter. Mar. Chem. 2011, 126, 139–148. [Google Scholar] [CrossRef]
- Dainard, P.G.; Guéguen, C. Distribution of PARAFAC modeled CDOM components in the North Pacific ocean, Bering, Chukchi and Beaufort seas. Mar. Chem. 2013, 157, 216–223. [Google Scholar] [CrossRef]
- Yamashita, Y.; Boyer, J.N.; Jaffé, R. Evaluating the distribution of terrestrial dissolved organic matter in a complex coastal ecosystem using fluorescence spectroscopy. Cont. Shelf Res. 2013, 66, 136–144. [Google Scholar] [CrossRef]
- Galletti, Y.; Gonnelli, M.; Brogi, S.R.; Vestri, S.; Santinelli, C. DOM dynamics in open waters of the Mediterranean Sea: New insights from optical properties. Deep Sea Res. Pt. I 2019, 144, 95–114. [Google Scholar] [CrossRef]
- Stedmon, C.A.; Seredyńska-Sobecka, B.; Boe-Hansen, R.; Le Tallec, N.; Waul, C.K.; Arvin, E. A potential approach for monitoring drinking water quality from groundwater systems using organic matter fluorescence as an early warning for contamination events. Water Res. 2011, 45, 6030–6038. [Google Scholar] [CrossRef]
- Nowicki, S.; Lapworth, D.J.; Ward, J.S.; Thomson, P.; Charles, K. Tryptophan-like fluorescence as a measure of microbial contamination risk in groundwater. Sci. Total Environ. 2019, 646, 782–791. [Google Scholar] [CrossRef]
- Henderson, R.K.; Baker, A.; Murphy, K.R.; Hambly, A.; Stuetz, R.M.; Khan, S.J. Fluorescence as a potential monitoring tool for recycled water systems: A review. Water Res. 2009, 43, 863–881. [Google Scholar] [CrossRef]
- Hambly, A.C.; Henderson, R.K.; Storey, M.V.; Baker, A.; Stuetz, R.M.; Khan, S.J. Fluorescence monitoring at a recycled water treatment plant and associated dual distribution system-implications for cross-connection detection. Water Res. 2010, 44, 5323–5333. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.; Song, Y.; Gao, H.; Liu, L.; Yao, L.; Peng, J. Applying fluorescence spectroscopy and multivariable analysis to characterize structural composition of dissolved organic matter and its correlation with water quality in an urban river. Environ. Earth Sci. 2015, 73, 5163–5171. [Google Scholar] [CrossRef]
- Bieroza, M.; Baker, A.; Bridgeman, J. Relating freshwater organic matter fluorescence to organic carbon removal efficiency in drinking water treatment. Sci. Total Environ. 2009, 407, 1765–1774. [Google Scholar] [CrossRef] [PubMed]
- Baghoth, S.A.; Sharma, S.K.; Amy, G.L. Tracking natural organic matter (NOM) in a drinking water treatment plant using fluorescence excitation–emission matrices and PARAFAC. Water Res. 2011, 45, 797–809. [Google Scholar] [CrossRef]
- Zhou, Y.; Jeppesen, E.; Zhang, Y.; Shi, K.; Liu, X.; Zhu, G. Dissolved organic matter fluorescence at wavelength 275/342 nm as a key indicator for detection of point-source contamination in a large Chinese drinking water lake. Chemosphere 2016, 144, 503–509. [Google Scholar] [CrossRef]
- Baker, A. Fluorescence excitation-emission matrix characterization of some sewage-impacted rivers. Environ. Sci. Technol. 2001, 35, 948–953. [Google Scholar] [CrossRef]
- Baker, A.; Spencer, R.G. Characterization of dissolved organic matter from source to sea using fluorescence and absorbance spectroscopy. Sci. Total Environ. 2004, 333, 217–232. [Google Scholar] [CrossRef]
- Cyr, F.; Tedetti, M.; Besson, F.; Beguery, L.; Doglioli, A.M.; Petrenko, A.A.; Goutx, M. A new glider-compatible optical sensor for dissolved organic matter measurements: Test case from the NW Mediterranean Sea. Front. Mar. Sci. 2017, 4, 89. [Google Scholar] [CrossRef]
- Zoppini, A.; Ademollo, N.; Bensi, M.; Berto, D.; Bongiorni, L.; Campanelli, A.; Casentini, B.; Patrolecco, L.; Amalfitano, S. Impact of a river flood on marine water quality and planktonic microbial communities. Estuar. Coast. Shelf Sci. 2019, 224, 62–72. [Google Scholar] [CrossRef]
- Seritti, A.; Russo, D.; Nannicini, L.; Del Vecchio, R. DOC, absorption and fluorescence properties of estuarine and coastal waters of the Northern Tyrrhenian Sea. Chem. Speciat. Bioavailab. 1998, 10, 95–106. [Google Scholar] [CrossRef]
- Vignudelli, S.; Santinelli, C.; Murru, E.; Nannicini, L.; Seritti, A. Distributions of dissolved organic carbon (DOC) and chromophoric dissolved organic matter (CDOM) in coastal waters of the northern Tyrrhenian Sea (Italy). Estuar. Coast. Shelf Sci. 2004, 60, 133–149. [Google Scholar] [CrossRef]
- Para, J.; Coble, P.G.; Charrière, B.; Tedetti, M.; Fontana, C.; Sempéré, R. Fluorescence and absorption properties of chromophoric dissolved organic matter (CDOM) in coastal surface waters of the northwestern Mediterranean Sea, influence of the Rhône River. Biogeosciences 2010, 7, 4083. [Google Scholar] [CrossRef]
- Gonnelli, M.; Vestri, S.; Santinelli, C. Chromophoric dissolved organic matter and microbial enzymatic activity. Biophys. Chem. 2013, 182, 79–85. [Google Scholar] [CrossRef]
- Retelletti Brogi, S.; Gonnelli, M.; Vestri, S.; Santinelli, C. Biophysical processes affecting DOM dynamics at the Arno river mouth (Tyrrhenian Sea). Biophys. Chem. 2015, 197, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Coble, P. Characterization of marine and terrestrial DOM in seawater using excitation-emission matrix spectroscopy. Mar. Chem. 1996, 51, 325–346. [Google Scholar] [CrossRef]
- Fellman, J.B.; Hood, E.; Spencer, R.G.M. Fluorescence spectroscopy opens new windows into dissolved organic matter dynamics in freshwater ecosystems: A review. Limnol. Oceanogr. 2010, 55, 2452–2462. [Google Scholar] [CrossRef]
- Reynolds, D.M. The differentiation of biodegradable and non-biodegradable dissolved organic matter in wastewaters using fluorescence spectroscopy. J. Chem. Technol. Biotechnol. 2002, 77, 965–972. [Google Scholar] [CrossRef]
- Hudson, N.; Baker, A.; Ward, D.; Reynolds, D.M.; Brunsdon, C.; Carliell-Marquet, C.; Browning, S. Can fluorescence spectrometry be used as a surrogate for the biochemical oxygen demand (BOD) test in water quality assessment? An example from South West England. Sci. Total Environ. 2008, 391, 149–158. [Google Scholar] [CrossRef] [PubMed]
- Carstea, E.M.; Popa, C.L.; Baker, A.; Bridgeman, J. In situ fluorescence measurements of dissolved organic matter: A review. Sci. Total Environ. 2020, 699, 134361. [Google Scholar] [CrossRef]
- Yamashita, Y.; Tanoue, E. Chemical characterization of protein-like fluorophores in DOM in relation to aromatic amino acids. Mar. Chem. 2003, 82, 255–271. [Google Scholar] [CrossRef]
- Determann, S.; Lobbes, J.M.; Reuter, R.; Rullkutter, J. Ultraviolet fluorescence excitation and emission spectroscopy of marine algae and bacteria. Mar. Chem. 1998, 62, 137–156. [Google Scholar] [CrossRef]
- Carlson, C.A. Production and removal processes. In Biogeochemistry of Marine Dissolved Organic Matter; Hansell, D.A., Carlson, C.A., Eds.; Academic Press: San Diego, CA, USA, 2002; pp. 91–152. [Google Scholar]
- Hedges, J.I. Why dissolved organics matter? In Biogeochemistry of Marine Dissolved Organic Matter; Hansell, D.A., Carlson, C.A., Eds.; Academic Press: San Diego, CA, USA, 2002; pp. 1–33. [Google Scholar]
- Jiao, N.; Herndl, G.J.; Hansell, D.A.; Benner, R.; Kattner, G.; Wilhelm, S.W.; Kirchman, D.L.; Weinbauer, M.G.; Luo, T.; Chen, F.; et al. Microbial production of recalcitrant dissolved organic matter: Long-term carbon storage in the global ocean. Nat. Rev. Microbiol. 2010, 8, 593. [Google Scholar] [CrossRef] [PubMed]
- Smith, R.C.; Prezelin, B.B.; Baker, K.E.A.; Bidigare, R.R.; Boucher, N.P.; Coley, T.; Karents, D.; Macintyre, S.; Matlick, H.A.; Menzies, D.; et al. Ozone depletion: Ultraviolet radiation and phytoplankton biology in Antarctic waters. Science 1992, 255, 952–959. [Google Scholar] [CrossRef] [PubMed]
- Herndl, G.J.; Müller-Niklas, G.; Frick, J. Major role of ultraviolet-B in controlling bacterioplankton growth in the surface layer of the ocean. Nature 1993, 361, 717–719. [Google Scholar] [CrossRef]
- Nelson, N.B.; Siegel, D.A. The global distribution and dynamics of chromophoric dissolved organic matter. Annu. Rev. Mar. Sci. 2013, 5, 447–476. [Google Scholar] [CrossRef]
- Hoppe, H.G.; Arnosti, C.; Herndl, G.J. Ecological Significance of Bacterial Enzymes in the Marine Environment. In Enzymes in the Environment: Activity, Ecology and Applications; Burns, R., Dick, R., Eds.; Marcel Dekker: New York, NY, USA, 2002; pp. 73–108. [Google Scholar]
- Caruso, G. Leucine aminopeptidase, b-glucosidase and alkaline phosphatase activity rates and their significance in nutrient cycles in some coastal Mediterranean sites. Mar. Drugs 2010, 8, 916–940. [Google Scholar] [CrossRef]
- Mitchell, B.G.; Kahru, M.; Wieland, J.; Stramska, M. Determination of spectral absorption coefficients of particles, dissolved material and phytoplankton for discrete water samples. In Ocean Optics Protocols for Satellite Ocean Color Sensor Validation, Revision 3; Muller, J.L., Fargion, G.S., Eds.; NASA: Hanover, NH, USA, 2002; Volume 2, Chapter 15; pp. 231–257. [Google Scholar]
- Twardowski, M.S.; Boss, E.; Sullivan, J.M.; Donaghay, P.L. Modeling the spectral shape of absorption by chromophoric dissolved organic matter. Mar. Chem. 2004, 89, 69–88. [Google Scholar] [CrossRef]
- Bricaud, A.; Morel, A.; Prieur, L. Absorption by dissolved organic matter of the sea (yellow substance) in the UV and visible domains 1. Limnol. Oceanogr. 1981, 26, 43–53. [Google Scholar] [CrossRef]
- Helms, J.R.; Stubbins, A.; Ritchie, J.D.; Minor, E.C.; Kieber, D.J.; Mopper, K. Absorption spectral slopes and slope ratios as indicators of molecular weight, source, and photobleaching of chromophoric dissolved organic matter. Limnol. Oceanogr. 2008, 53, 955–969. [Google Scholar] [CrossRef]
- Monticelli, L.S.; Caruso, G.; Decembrini, F.; Caroppo, C.; Fiesoletti, F. Role of prokaryotic biomasses and activities in carbon and phosphorus cycles at a coastal, thermohaline front and in offshore waters (Gulf of Manfredonia, Southern Adriatic Sea). Microb. Ecol. 2014, 67, 501–519. [Google Scholar] [CrossRef]
- Hammer, Ø.; Harper, D.A.T.; Ryan, P.D. PAST: PAleontological STatistics software package for education and data analysis. Paleontol. Electron. 2001, 4, 1–9. Available online: http://palaeo-electronica.org/2001_1/past/issue1_01.htm (accessed on 1 October 2019).
- Bro, R. PARAFAC. Tutorial and applications. Chemometr. Intell. Lab. 1997, 38, 149–172. [Google Scholar] [CrossRef]
- Stedmon, C.A.; Bro, R. Characterizing dissolved organic matter fluorescence with parallel factor analysis: A tutorial. Limnol. Oceanogr. 2008, 6, 572–579. [Google Scholar] [CrossRef]
- Andersen, C.M.; Bro, R. Practical aspects of PARAFAC modeling of fluorescence excitation-emission data. J. Chemom. 2003, 17, 200–215. [Google Scholar] [CrossRef]
- Bahram, M.; Bro, R.; Stedmon, C.; Afkhami, A. Handling of Rayleigh and Raman scatter for PARAFAC modeling of fluorescence data using interpolation. J. Chemom. 2006, 20, 99–105. [Google Scholar] [CrossRef]
- Lakowicz, J.R. Principles of Fluorescence Spectroscopy, 3rd ed.; Springer-Verlag: Berlin/Heidelberg, Germany, 2006. [Google Scholar] [CrossRef]
- Gonnelli, M.; Galletti, Y.; Marchetti, E.; Mercadante, L.; Brogi, S.R.; Ribotti, A.; Sorgente, R.; Vestri, S.; Santinelli, C. Dissolved organic matter dynamics in surface waters affected by oil spill pollution: Results from the Serious Game exercise. Deep-Sea Res. PT II 2016, 133, 88–99. [Google Scholar] [CrossRef]
- Tedetti, M.; Marie, L.; Röttgers, R.; Rodier, M.; Van Wambeke, F.; Helias, S.; Caffin, M.; Cornet-Barthaux, V.; Dupouy, C. Evolution of dissolved and particulate chromophoric materials during the VAHINE mesocosm experiment in the New Caledonian coral lagoon (south-west Pacific). Biogeosciences 2016, 13, 3283–3303. [Google Scholar] [CrossRef]
- Zappalà, G.; Bonamano, S.; Madonia, A.; Caruso, G.; Marcelli, M. Microbial risk assessment in a coastal marine environment through the use of mathematical models. WIT Trans. Ecol. Environ. 2012, 164, 3–14. [Google Scholar] [CrossRef]
- Bonamano, S.; Madonia, A.; Borsellino, C.; Stefanì, C.; Caruso, G.; De Pasquale, F.; Piermattei, V.; Zappalà, G.; Marcelli, M. Modeling the dispersion of viable and total Escherichia coli cells in the artificial semi-enclosed bathing area of Santa Marinella (Latium, Italy). Mar. Pollut. Bull. 2015, 95, 141–154. [Google Scholar] [CrossRef]
- Caruso, G.; Azzaro, M.; Caroppo, C.; Decembrini, F.; Monticelli, L.S.; Leonardi, M.; Maimone, G.; Zaccone, R.; La Ferla, R. Microbial community and its potential as descriptor of environmental status. ICES J. Mar. Sci. 2016, 73, 2174–2177. [Google Scholar] [CrossRef]
- Zaccone, R.; Caruso, G. Microbial enzymes in the Mediterranean Sea: Relationship with climate changes. AIMS Microbiol. 2019, 5, 251–271. [Google Scholar] [CrossRef] [PubMed]
- Santos, L.; Santos, E.B.H.; Dias, J.M.; Cunha, A.; Almeida, A. Photochemical and microbial alterations of DOM spectroscopic properties in the estuarine system Ria de Aveiro. Photochem. Photobiol. Sci. 2014, 13, 1146–1159. [Google Scholar] [CrossRef] [PubMed]
- Nowak, A.; Slizewska, K. B-Glucuronidase and -glucosidase activity and human fecal water genotoxicity in the presence of probiotic lactobacilli and the heterocyclic aromatic amine IQ in vitro. Env. Toxicol. Pharmacol. 2014, 37, 66–73. [Google Scholar] [CrossRef] [PubMed]
- Flores, R.; Shi, J.; Gail, M.H.; Ravel, J.; Goedert, J.J. Assessment of the human fecal microbiota: I. Measurement and reproducibility of selected enzymatic activities. Eur. J. Clin. Investig. 2012, 42, 848–854. [Google Scholar] [CrossRef]
- Singh, S.; D’Sa, E.J.; Swenson, E.M. Chromophoric dissolved organic matter (CDOM) variability in Barataria Basin using excitation–emission matrix (EEM) fluorescence and parallel factor analysis (PARAFAC). Sci. Total Environ. 2010, 408, 3211–3222. [Google Scholar] [CrossRef]
- Moore, C.; Barnard, A.; Fietzek, P.; Lewis, M.R.; Sosik, H.M.; White, S.; Zielinski, O. Optical tools for ocean monitoring and research. Ocean Sci. 2009, 5, 661–684. [Google Scholar] [CrossRef]
- Zielinski, O.; Busch, J.A.; Cembella, A.D.; Daly, K.L.; Engelbrektsson, J.; Hannides, A.K.; Schmidt, H. Detecting marine hazardous substances and organisms: Sensors for pollutants, toxins, and pathogens. Ocean Sci. 2009, 5, 329–349. [Google Scholar] [CrossRef]
- Conmy, R.N.; Del Castillo, C.E.; Downing, B.D.; Chen, R.F. Experimental design and quality assurance: In situ fluorescence instrumentation. In Aquatic Organic Matter Fluorescence; Coble, P.G., Lead, J.R., Baker, A., Reynolds, D.M., Spencer, R.G.M., Eds.; Cambridge University Press: New York, NY, USA, 2014; pp. 190–230. [Google Scholar]
- Miloslavich, P.; Bax, N.; Simmons, S.; Klein, E.; Appeltans, W.; Aburto-Oropeza, O.; Andersen Garcia, M.; Batten, S.; Benedetti-Cecchi, L.; Checkley, D.; et al. Essential Ocean Variables for global sustained observations of biodiversity and ecosystem changes. Glob. Chang. Biol. 2018, 24, 2416–2433. [Google Scholar] [CrossRef]
- Marcelli, M.; Piermattei, V.; Madonia, A.; Mainardi, U. Design and application of new low-cost instruments for marine environmental research. Sensors 2014, 14, 23348–23364. [Google Scholar] [CrossRef]
- Marcelli, M.; Piermattei, V.; Madonia, A.; Lacava, T.; Mainardi, U. T-FLaP advances: Instrumental and operative implementation. J. Oper. Oceanogr. 2016, 9, 185–192. [Google Scholar] [CrossRef]
- Caruso, G.; Denaro, R.; Genovese, M.; Giuliano, L.; Mancuso, M.; Yakimov, M. New methodological strategies for detecting bacterial indicators. Chem. Ecol. 2004, 20, 167–181. [Google Scholar] [CrossRef]
- Ammerman, J.W.; Glover, W.B. Continuous underway measurement of microbial ectoenzyme activities in aquatic ecosystems. Mar. Ecol. Progr. Ser. 2000, 201, 1–12. [Google Scholar] [CrossRef]
- Piermattei, V.; Madonia, A.; Bonamano, S.; Martellucci, R.; Bruzzone, G.; Ferretti, R.; Odetti, A.; Azzaro, M.; Zappalà, G.; Marcelli, M. Cost-Effective Technologies to Study the Arctic Ocean Environment. Sensors 2018, 18, 2257. [Google Scholar] [CrossRef] [PubMed]
| Station | Year | Temp | Salinity | a280 | a355 | E. coli MF | E. coli IF | BG | LA | AP |
|---|---|---|---|---|---|---|---|---|---|---|
| - | °C | - | m−1 | m−1 | CFU 100 mL−1 | Cells 100 mL−1 | ng L−1 h−1 | ng L−1 h−1 | ng L−1 h−1 | |
| SC0 | 2015 | 29.2 | 0.527 | 38.01 | 18.20 | 20,000 | 448,862 | 51.21 | 29.25 | 12.46 |
| 2016 | 22.2 | 0.470 | 9.45 | 3.02 | 2509 | 13,543 | - | - | - | |
| 2017 | 26.9 | 0.550 | 59.88 | 26.37 | 21,205 | 27,460 | 20.76 | 0.88 | 32.05 | |
| median | 26.9 | 0.527 | 38.02 | 18.20 | 20,000 | 27,460 | 35.99 | 15.07 | 22.26 | |
| SC1 | 2015 | 28.67 | 31.434 | 3.74 | 1.51 | 3212.5 | 3632 | 8.94 | 4.11 | 7.31 |
| 2016 | 23 | 31.989 | 2.82 | 0.78 | 870 | 7269 | - | - | - | |
| 2017 | 24.4 | 31.855 | 19.16 | 7.44 | 17,535 | 14,355 | 14.78 | 2.7 | 19.52 | |
| median | 24.4 | 31.855 | 3.74 | 1.51 | 3213 | 7269 | 11.86 | 3.41 | 13.42 | |
| SC2- SC6 | 2015 | 28.5 | 38.433 | 2.49 | 1.02 | 25 | 4894 | 1.24 | 1.93 | 27.04 |
| 2016 | 22.7 | 38.700 | 1.97 | 0.54 | 412 | 4000 | - | - | - | |
| 2017 | 23.7 | 37.525 | 2.04 | 0.72 | 228 | 1318 | 2.6 | 0.51 | 6.42 | |
| median | 28.5 | 38.433 | 2.04 | 0.72 | 228 | 4000 | 1.92 | 1.22 | 16.73 |
| Parameter | a280 | a355 | E. coli MF | E. coli IF | BG | LA | AP |
|---|---|---|---|---|---|---|---|
| a280 | - | 1.40 × 10−55 | 1.73 × 10−18 | 2.09 × 10−2 | 6.14 × 10−5 | 0.1998 | 0.2810 |
| a355 | 0.998 ** | - | 3.77 × 10−17 | 7.65 × 10−2 | 1.88 × 10−5 | 0.1805 | 0.2740 |
| E. coli MF | 0.939 ** | 0.930 ** | - | 2.72 × 10−2 | 8.59 × 10−5 | 0.0701 | 0.4851 |
| E. coli IF | 0.619 * | 0.591 | 0.630 * | - | 7.11× 10−2 | 0.0187 | 0.9092 |
| BG | 0.764 ** | 0.780 ** | 0.785 ** | 0.651 | - | 0.0002 | 0.5895 |
| LA | 0.204 | 0.213 | 0.301 | 0.375 ** | 0.554 ** | - | 0.9547 |
| AP | 0.172 | 0.175 | 0.118 | 0.019 | 0.087 | −0.009 | - |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Madonia, A.; Caruso, G.; Piazzolla, D.; Bonamano, S.; Piermattei, V.; Zappalà, G.; Marcelli, M. Chromophoric Dissolved Organic Matter as a Tracer of Fecal Contamination for Bathing Water Quality Monitoring in the Northern Tyrrhenian Sea (Latium, Italy). J. Mar. Sci. Eng. 2020, 8, 430. https://doi.org/10.3390/jmse8060430
Madonia A, Caruso G, Piazzolla D, Bonamano S, Piermattei V, Zappalà G, Marcelli M. Chromophoric Dissolved Organic Matter as a Tracer of Fecal Contamination for Bathing Water Quality Monitoring in the Northern Tyrrhenian Sea (Latium, Italy). Journal of Marine Science and Engineering. 2020; 8(6):430. https://doi.org/10.3390/jmse8060430
Chicago/Turabian StyleMadonia, Alice, Gabriella Caruso, Daniele Piazzolla, Simone Bonamano, Viviana Piermattei, Giuseppe Zappalà, and Marco Marcelli. 2020. "Chromophoric Dissolved Organic Matter as a Tracer of Fecal Contamination for Bathing Water Quality Monitoring in the Northern Tyrrhenian Sea (Latium, Italy)" Journal of Marine Science and Engineering 8, no. 6: 430. https://doi.org/10.3390/jmse8060430
APA StyleMadonia, A., Caruso, G., Piazzolla, D., Bonamano, S., Piermattei, V., Zappalà, G., & Marcelli, M. (2020). Chromophoric Dissolved Organic Matter as a Tracer of Fecal Contamination for Bathing Water Quality Monitoring in the Northern Tyrrhenian Sea (Latium, Italy). Journal of Marine Science and Engineering, 8(6), 430. https://doi.org/10.3390/jmse8060430









