Assessing the Effectiveness of 3D-Printed Ceramic Structures for Coral Restoration: Growth, Survivorship, and Biodiversity Using Visual Surveys and eDNA
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Sites
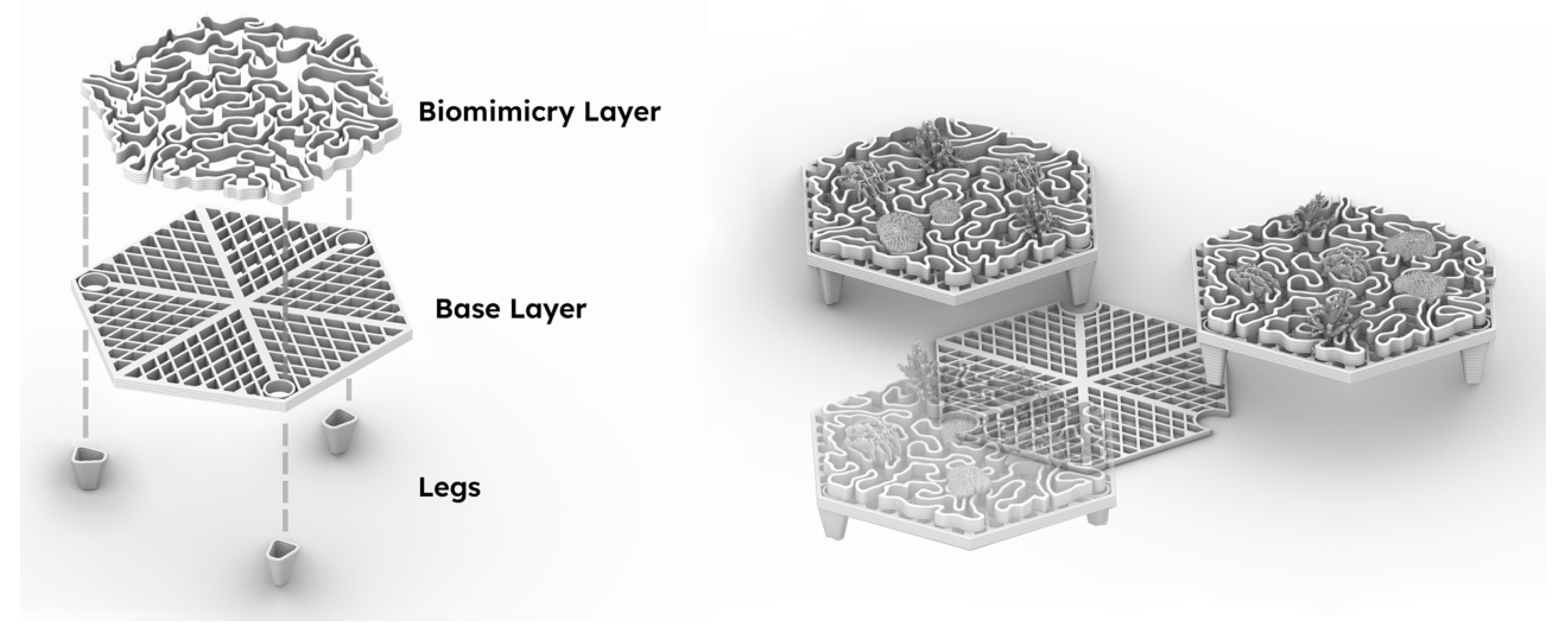
2.2. Reef Tile Design
2.3. Restoration Site Design
2.4. Monitoring Coral Survivorship and Growth
2.5. Visual Fish and Invertebrate Surveys
2.6. Environmental DNA Metabarcoding
2.6.1. DNA Sampling, Extraction, Amplification, and Sequencing
2.6.2. Bioinformatics
Sequence Analysis and Quality Control
Community Diversity Analysis
3. Results
3.1. Coral Performance
3.1.1. Coral Transplant Survival
3.1.2. Extension Rates and Breakage
3.2. Fish and Invertebrate Surveys
3.3. Environmental DNA
3.3.1. DNA Sequencing Output by Phyla
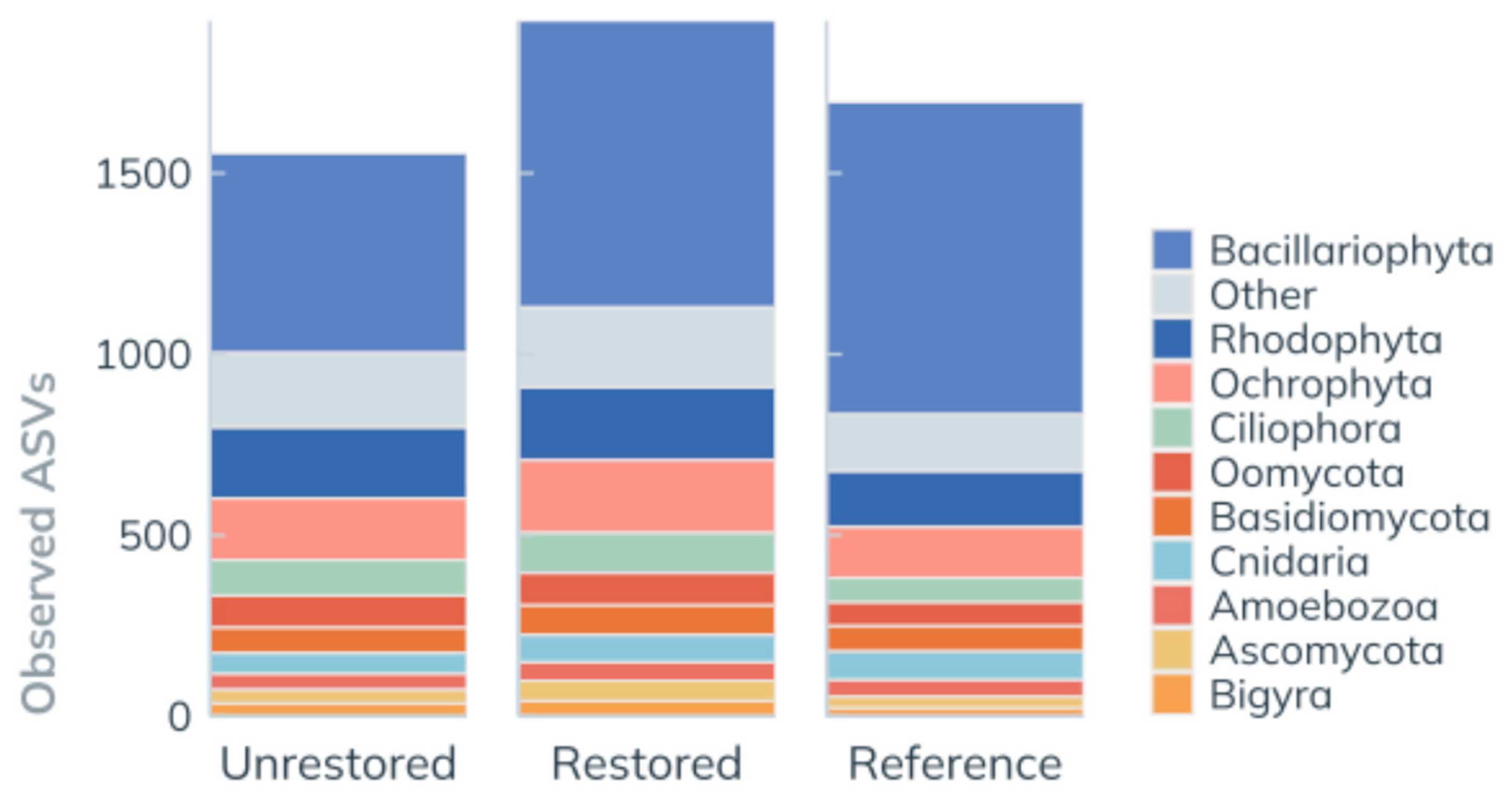
3.3.2. ASV Richness by Site
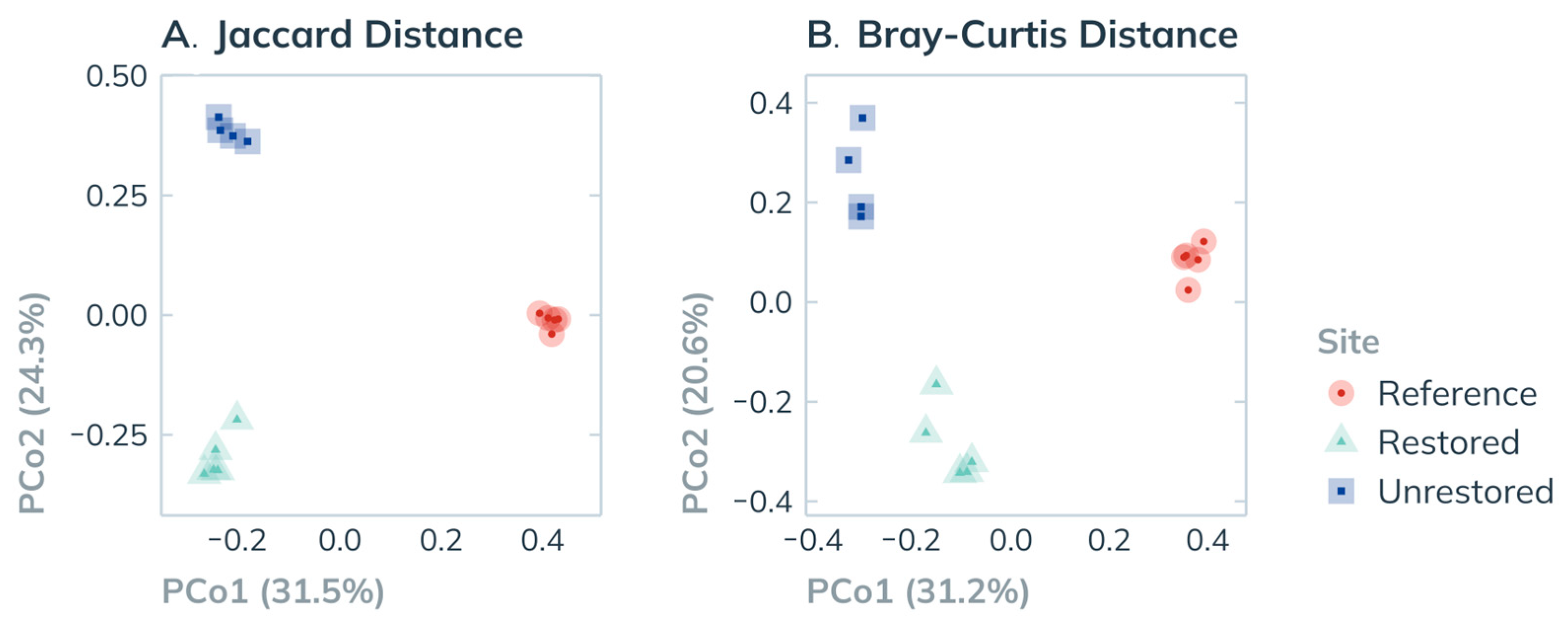
3.3.3. Community Similarity
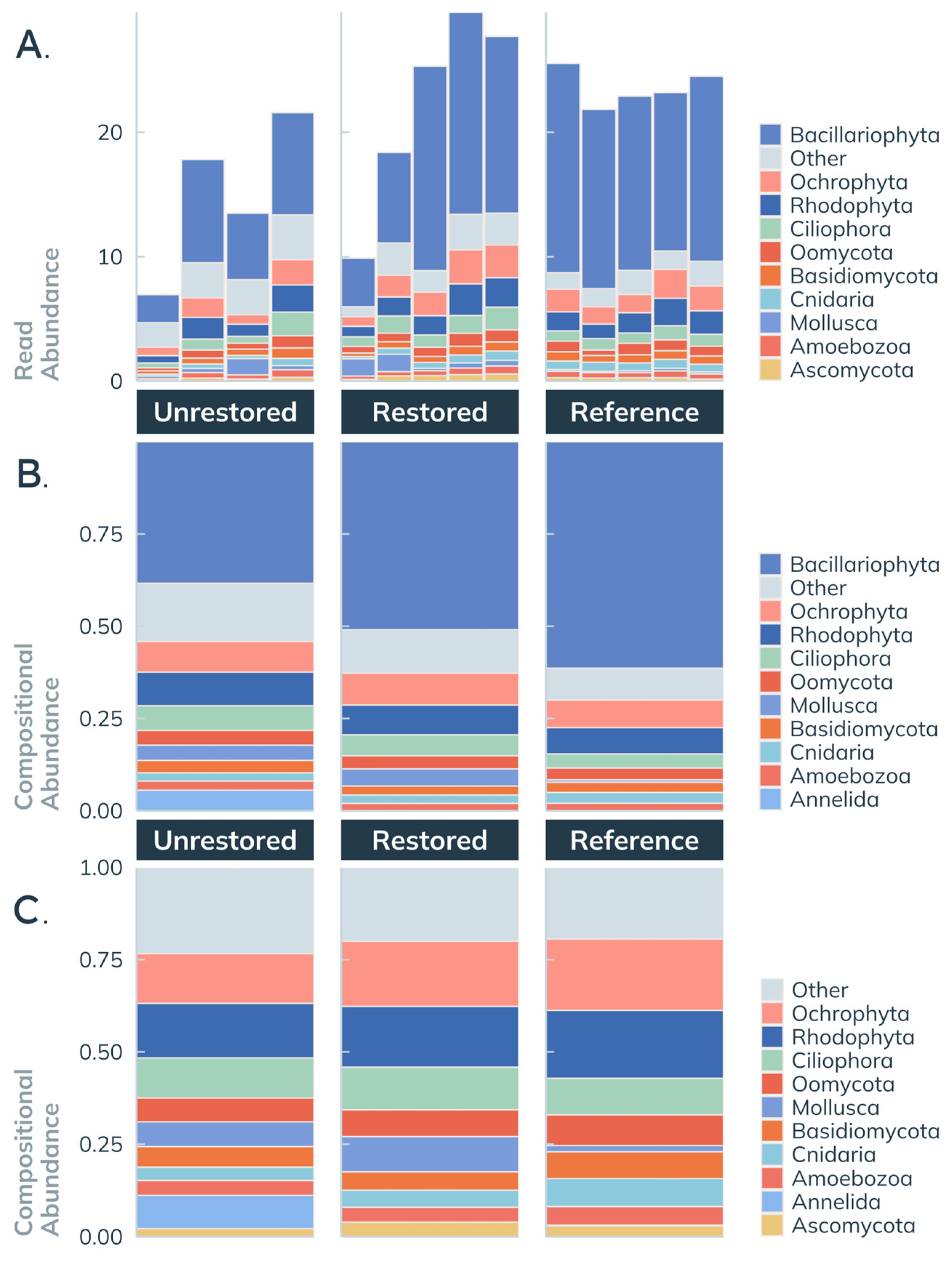
3.3.4. Taxonomic Composition by Site: Diversity and Abundance
4. Discussion
4.1. Ecological Outcomes
4.1.1. Coral Performance
4.1.2. Biodiversity
Fish and Macroinvertebrate Abundance
eDNA Cryptobiome
4.2. Measuring Coral Restoration Outcomes
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| HHWMP | Hoi Ha Wan Marine Park |
| eDNA | Environmental DNA |
Appendix A
| Model | K | logLik | AIC | ΔAIC | AIC Weight |
|---|---|---|---|---|---|
| Random: unit_no AR1: survey_no | 7 | 18,080.95 | −36,147.91 | 168.75 | <0.001 |
| Random: unit_no/tile_no AR1: survey_no | 8 | 18,102.02 | −36,188.05 | 128.61 | 0.003 |
| Random: unit_no/tile_no/coral_no AR1: survey_no | 9 | 18,167.33 | −36,316.66 | 0.00 | 0.997 |
| Acropora | Pavona | Platygyra | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Survey | H | PM | D | DTCH | H | PM | D | DTCH | H | PM | D | DTCH |
| 1 | 126 | 0 | 0 | 0 | 126 | 0 | 0 | 0 | 126 | 0 | 0 | 0 |
| 2 | 126 | 0 | 0 | 0 | 125 | 0 | 0 | 1 | 126 | 0 | 0 | 0 |
| 3 | 126 | 0 | 0 | 0 | 125 | 0 | 0 | 1 | 126 | 0 | 0 | 0 |
| 4 | 126 | 0 | 0 | 0 | 125 | 0 | 0 | 1 | 126 | 0 | 0 | 0 |
| 5 | 125 | 0 | 0 | 1 | 121 | 0 | 1 | 4 | 125 | 0 | 0 | 1 |
| 6 | 125 | 0 | 0 | 1 | 120 | 0 | 2 | 4 | 124 | 0 | 0 | 2 |
| 7 | 125 | 0 | 0 | 1 | 113 | 0 | 2 | 11 | 124 | 0 | 0 | 2 |
| 8 | 125 | 0 | 0 | 1 | 107 | 0 | 4 | 15 | 123 | 0 | 1 | 2 |
| 9 | 125 | 0 | 0 | 1 | 105 | 0 | 4 | 17 | 123 | 0 | 1 | 2 |
| 10 | 124 | 0 | 0 | 2 | 105 | 0 | 4 | 17 | 122 | 1 | 1 | 2 |
| 11 | 122 | 0 | 0 | 4 | 101 | 0 | 4 | 21 | 116 | 4 | 3 | 3 |
| 12 | 120 | 0 | 0 | 6 | 99 | 0 | 4 | 23 | 109 | 11 | 3 | 3 |
| 13 | 119 | 1 | 0 | 6 | 99 | 0 | 4 | 23 | 116 | 3 | 4 | 3 |
| 14 | 118 | 1 | 0 | 7 | 96 | 0 | 4 | 26 | 112 | 6 | 4 | 4 |
Appendix B

References
- Gardner, T.A.; Côté, I.M.; Gill, J.A.; Grant, A.; Watkinson, A.R. Long-Term Region-Wide Declines in Caribbean Corals. Science 2003, 301, 958–960. [Google Scholar] [CrossRef]
- Bruno, J.F.; Valdivia, A. Coral Reef Degradation Is Not Correlated with Local Human Population Density. Sci. Rep. 2016, 6, 29778. [Google Scholar] [CrossRef]
- Hochberg, E.J.; Gierach, M.M. Missing the Reef for the Corals: Unexpected Trends Between Coral Reef Condition and the Environment at the Ecosystem Scale. Front. Mar. Sci. 2021, 8, 727038. [Google Scholar] [CrossRef]
- Whaley, Z.; Cramer, K.; McClenachan, L.; Tewfik, A.; Alvarez-Filip, L.; McField, M.; Carilli, J.; Vardi, T. Long-Term Change in Caribbean Reef Water Quality and Ecosystem Health. Bull. Fla. Mus. Nat. Hist. 2023, 60, 126. [Google Scholar] [CrossRef]
- Grottoli, A.G.; Warner, M.E.; Levas, S.J.; Aschaffenburg, M.D.; Schoepf, V.; McGinley, M.; Baumann, J.; Matsui, Y. The Cumulative Impact of Annual Coral Bleaching Can Turn Some Coral Species Winners into Losers. Glob. Change Biol. 2014, 20, 3823–3833. [Google Scholar] [CrossRef]
- Kalmus, P.; Ekanayaka, A.; Kang, E.; Baird, M.; Gierach, M. Past the Precipice? Projected Coral Habitability Under Global Heating. Earths Future 2022, 10, e2021EF002608. [Google Scholar] [CrossRef] [PubMed]
- Reaka-Kudla, M.L. The Global Biodiversity of Coral Reefs: A Comparison with Rainforests. Biodivers. II Underst. Prot. Our Biol. Resour. 1997, 2, 551. [Google Scholar]
- Knowlton, N.; Brainard, R.E.; Fisher, R.; Moews, M.; Plaisance, L.; Caley, M.J. Coral Reef Biodiversity. In Life in the World’s Oceans; Wiley-Blackwell: Hoboken, NJ, USA, 2010; pp. 65–78. ISBN 978-1-4443-2550-8. [Google Scholar]
- Battaglia, F.M. Climate Change and the Ocean: The Disruption of the Coral Reef. In Blue Planet Law; Springer: Cham, Switzerland, 2023; pp. 121–130. ISBN 978-3-031-24887-0. [Google Scholar]
- Edwards, A.; Guest, J.; Humanes, A. Rehabilitating Coral Reefs in the Anthropocene. Curr. Biol. 2024, 34, R399–R406. [Google Scholar] [CrossRef] [PubMed]
- Boström-Einarsson, L.; Babcock, R.C.; Bayraktarov, E.; Ceccarelli, D.; Cook, N.; Ferse, S.C.A.; Hancock, B.; Harrison, P.; Hein, M.; Shaver, E.; et al. Coral Restoration—A Systematic Review of Current Methods, Successes, Failures and Future Directions. PLoS ONE 2020, 15, e0226631. [Google Scholar] [CrossRef] [PubMed]
- Higgins, E.; Metaxas, A.; Scheibling, R.E. A Systematic Review of Artificial Reefs as Platforms for Coral Reef Research and Conservation. PLoS ONE 2022, 17, e0261964. [Google Scholar] [CrossRef] [PubMed]
- Sedano, F.; Navarro-Barranco, C.; Guerra-García, J.M.; Espinosa, F. Understanding the Effects of Coastal Defence Structures on Marine Biota: The Role of Substrate Composition and Roughness in Structuring Sessile, Macro- and Meiofaunal Communities. Mar. Pollut. Bull. 2020, 157, 111334. [Google Scholar] [CrossRef] [PubMed]
- Hata, T.; Madin, J.S.; Cumbo, V.R.; Denny, M.; Figueiredo, J.; Harii, S.; Thomas, C.J.; Baird, A.H. Coral Larvae Are Poor Swimmers and Require Fine-Scale Reef Structure to Settle. Sci. Rep. 2017, 7, 2249. [Google Scholar] [CrossRef] [PubMed]
- Levy, N.; Berman, O.; Yuval, M.; Loya, Y.; Treibitz, T.; Tarazi, E.; Levy, O. Emerging 3D Technologies for Future Reformation of Coral Reefs: Enhancing Biodiversity Using Biomimetic Structures Based on Designs by Nature. Sci. Total Environ. 2022, 830, 154749. [Google Scholar] [CrossRef]
- Spieler, R.E.; Gilliam, D.S.; Sherman, R.L. Artificial Substrate and Coral Reef Restoration: What Do We Need to Know to Know What We Need? Bull. Mar. Sci. 2001, 69, 1013–1030. [Google Scholar]
- Vivier, B.; Dauvin, J.-C.; Navon, M.; Rusig, A.-M.; Mussio, I.; Orvain, F.; Boutouil, M.; Claquin, P. Marine Artificial Reefs, a Meta-Analysis of Their Design, Objectives and Effectiveness. Glob. Ecol. Conserv. 2021, 27, e01538. [Google Scholar] [CrossRef]
- Done, T.J. Phase Shifts in Coral Reef Communities and Their Ecological Significance. Hydrobiologia 1992, 247, 121–132. [Google Scholar] [CrossRef]
- Wismer, S.; Hoey, A.; Bellwood, D. Cross-Shelf Benthic Community Structure on the Great Barrier Reef: Relationships between Macroalgal Cover and Herbivore Biomass. Mar. Ecol. Prog. Ser. 2009, 376, 45–54. [Google Scholar] [CrossRef]
- Edwards, A.; Job, S.; Wells, S. Learning Lessons from Past Reef-Rehabilitation Projects. In Reef Rehabilitation Manual; Coral Reef Targeted Research & Capacity Building for Management Program: St. Lucia, QLD, Australia, 2010; pp. 129–166. ISBN 978-1-921317-05-7. [Google Scholar]
- Hein, M.Y.; Willis, B.L.; Beeden, R.; Birtles, A. The Need for Broader Ecological and Socioeconomic Tools to Evaluate the Effectiveness of Coral Restoration Programs. Restor. Ecol. 2017, 25, 873–883. [Google Scholar] [CrossRef]
- Bourne, D.G.; Morrow, K.M.; Webster, N.S. Insights into the Coral Microbiome: Underpinning the Health and Resilience of Reef Ecosystems. Annu. Rev. Microbiol. 2016, 70, 317–340. [Google Scholar] [CrossRef]
- Leray, M.; Knowlton, N. DNA Barcoding and Metabarcoding of Standardized Samples Reveal Patterns of Marine Benthic Diversity. Proc. Natl. Acad. Sci. USA 2015, 112, 2076–2081. [Google Scholar] [CrossRef]
- Beng, K.C.; Corlett, R.T. Applications of Environmental DNA (eDNA) in Ecology and Conservation: Opportunities, Challenges and Prospects. Biodivers. Conserv. 2020, 29, 2089–2121. [Google Scholar] [CrossRef]
- Thompson, S.; Jarman, S.; Griffin, K.; Spencer, C.; Cummins, G.; Partridge, J.; Langlois, T. Novel Drop-Sampler for Simultaneous Collection of Stereo-Video, Environmental DNA and Oceanographic Data. Ecol. Evol. 2024, 14, e70705. [Google Scholar] [CrossRef] [PubMed]
- Duprey, N.N.; McIlroy, S.E.; Ng, T.P.T.; Thompson, P.D.; Kim, T.; Wong, J.C.Y.; Wong, C.W.M.; Husa, S.M.; Li, S.M.H.; Williams, G.A.; et al. Facing a Wicked Problem with Optimism: Issues and Priorities for Coral Conservation in Hong Kong. Biodivers. Conserv. 2017, 26, 2521–2545. [Google Scholar] [CrossRef]
- Xie, J.Y.; Yeung, Y.H.; Kwok, C.K.; Kei, K.; Ang, P.; Chan, L.L.; Cheang, C.C.; Chow, W.; Qiu, J.-W. Localized Bleaching and Quick Recovery in Hong Kong’s Coral Communities. Mar. Pollut. Bull. 2020, 153, 110950. [Google Scholar] [CrossRef] [PubMed]
- Duprey, N.N.; Yasuhara, M.; Baker, D.M. Reefs of Tomorrow: Eutrophication Reduces Coral Biodiversity in an Urbanized Seascape. Glob. Change Biol. 2016, 22, 3550–3565. [Google Scholar] [CrossRef]
- Fabricius, K.E. Effects of Terrestrial Runoff on the Ecology of Corals and Coral Reefs: Review and Synthesis. Mar. Pollut. Bull. 2005, 50, 125–146. [Google Scholar] [CrossRef]
- Cybulski, J.D.; Husa, S.M.; Duprey, N.N.; Mamo, B.L.; Tsang, T.P.N.; Yasuhara, M.; Xie, J.Y.; Qiu, J.-W.; Yokoyama, Y.; Baker, D.M. Coral Reef Diversity Losses in China’s Greater Bay Area Were Driven by Regional Stressors. Sci. Adv. 2020, 6, eabb1046. [Google Scholar] [CrossRef] [PubMed]
- Yeung, Y.H.; Xie, J.Y.; Kwok, C.K.; Kei, K.; Ang, P.; Chan, L.L.; Dellisanti, W.; Cheang, C.C.; Chow, W.K.; Qiu, J.-W. Hong Kong’s Subtropical Scleractinian Coral Communities: Baseline, Environmental Drivers and Management Implications. Mar. Pollut. Bull. 2021, 167, 112289. [Google Scholar] [CrossRef] [PubMed]
- Hua, F.L.; Tsang, Y.F.; Chua, H. Progress of Water Pollution Control in Hong Kong. Aquat. Ecosyst. Health Manag. 2008, 11, 225–229. [Google Scholar] [CrossRef]
- Chan, A.; Chan, K.; Choi, C.; McCorry, D.; Lee, M.; Ang, P. Field Guide to Hard Corals of Hong Kong; Agriculture, Fisheries and Conservation Department, The Hong Kong SAR Government: Hong Kong, China, 2005.
- Geeraert, N.; Archana, A.; Xu, M.N.; Kao, S.-J.; Baker, D.M.; Thibodeau, B. Investigating the Link between Pearl River-Induced Eutrophication and Hypoxia in Hong Kong Shallow Coastal Waters. Sci. Total Environ. 2021, 772, 145007. [Google Scholar] [CrossRef]
- Lange, C. Rethinking Artificial Reef Structures through a Robotic 3D Clay Printing Method. In Proceedings of the 25th International Conference of the Association for Computer-Aided Architectural Design Research in Asia (CAADRIA), Bangkok, Thailand, 5–6 August 2020; Association for Computer-Aided Architectural Design Research in Asia (CAADRIA): Hong Kong, 2020; Volume 2, pp. 463–472. [Google Scholar]
- Brooks, M.E.; Kristensen, K.; van Benthem, K.J.; Magnusson, A.; Berg, C.W.; Nielsen, A.; Skaug, H.J.; Mächler, M.; Bolker, B.M. glmmTMB Balances Speed and Flexibility Among Packages for Zero-Inflated Generalized Linear Mixed Modeling. R J. 2017, 9, 378–400. [Google Scholar] [CrossRef]
- Bates, D.; Mächler, M.; Bolker, B.; Walker, S. Fitting Linear Mixed-Effects Models Using Lme4. J. Stat. Softw. 2015, 67, 1–48. [Google Scholar] [CrossRef]
- Lenth, R. emmeans: Estimated Marginal Means, Aka Least-Squares Means, version 1.11.2-8; R package. 2025. Available online: https://rvlenth.github.io/emmeans/ (accessed on 19 August 2025).
- Hodgson, G. Reef Check California Instruction Manual: A Guide to Monitoring California’s Rocky Reefs, 1st ed.; Reef Check Foundation: Pacific Palisades, CA, USA, 2006; ISBN 978-0-9723051-9-8. [Google Scholar]
- Sadovy, Y.; Cornish, A.S. Reef Fishes of Hong Kong; Hong Kong University Press: Hong Kong, China, 2000; ISBN 962-209-480-5. [Google Scholar]
- Azevedo, J.M.N.; Rodrigues, J.B.; Mendizabal, M.; Arruda, L.M. Study of a Sample of Dusky Groupers, Epinephelus Marginatus (Lowe, 1834), Caught in a Tide Pool at Lajes Do Pico, Azores. Bol. Mus. Munic. Funchal. 1995, 4, 55–64. [Google Scholar]
- Moore, A.; Ndobe, S.; Ambo-Rappe, R.; Jompa, J.; Yasir, I. Dietary Preference of Key Microhabitat Diadema Setosum: A Step towards Holistic Banggai Cardinalfish Conservation. IOP Conf. Ser. Earth Environ. Sci. 2019, 235, 012054. [Google Scholar] [CrossRef]
- Vadas, R.L. Preferential Feeding: An Optimization Strategy in Sea Urchins. Ecol. Monogr. 1977, 47, 337–371. [Google Scholar] [CrossRef]
- Tsuchiya, M.; Nishihira, M.; Poung-in, S.; Choohabandit, S. Feeding Behavior of the Urchin-Eating Urchin Salmacis Sphaeroides. Galaxea J. Coral Reef Stud. 2009, 11, 149–153. [Google Scholar] [CrossRef]
- Ahmed, Q.; Ali, Q.; Bat, L.; Öztekin, A.; Ghory, F.; Shaikh, I.; Qazi, H.; Baloch, A. Gut Content Analysis in Holothuria Leucospilota and Holothuria Cinerascens(Echinodermata: Holothuroidea: Holothuriidae) From Karachi Coast. J. Mater. Env. Sci. 2023, 14, 31–40. [Google Scholar]
- Woods, C.M.C. Natural Diet of the Seahorse Hippocampus Abdominalis. N. Z. J. Mar. Freshw. Res. 2002, 36, 655–660. [Google Scholar] [CrossRef]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Reproducible, Interactive, Scalable and Extensible Microbiome Data Science Using QIIME 2. Nat. Biotechnol. 2019, 37, 852–857. [Google Scholar] [CrossRef] [PubMed]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-Resolution Sample Inference from Illumina Amplicon Data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef]
- Wang, S.; Meyer, E.; McKay, J.K.; Matz, M.V. 2b-RAD: A Simple and Flexible Method for Genome-Wide Genotyping. Nat. Methods 2012, 9, 808–810. [Google Scholar] [CrossRef]
- Porter, T.M.; Hajibabaei, M. Over 2.5 Million COI Sequences in GenBank and Growing. PLoS ONE 2018, 13, e0200177. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing, version 4.5.0; R Foundation for Statistical Computing: Vienna, Austria, 2021.
- McMurdie, P.J.; Holmes, S. Phyloseq: An R Package for Reproducible Interactive Analysis and Graphics of Microbiome Census Data. PLoS ONE 2013, 8, e61217. [Google Scholar] [CrossRef] [PubMed]
- Kandlikar, G.S.; Gold, Z.J.; Cowen, M.C.; Meyer, R.S.; Freise, A.C.; Kraft, N.J.; Moberg-Parker, J.; Sprague, J.; Kushner, D.J.; Curd, E.E. Ranacapa: An R Package and Shiny Web App to Explore Environmental DNA Data with Exploratory Statistics and Interactive Visualizations. F1000Research 2018, 7, 1734. [Google Scholar] [CrossRef] [PubMed]
- McIlroy, S.E.; Guibert, I.; Archana, A.; Chung, W.Y.H.; Duffy, J.E.; Gotama, R.; Hui, J.; Knowlton, N.; Leray, M.; Meyer, C.; et al. Life Goes on: Spatial Heterogeneity Promotes Biodiversity in an Urbanized Coastal Marine Ecosystem. Glob. Change Biol. 2023, 30, e17248. [Google Scholar] [CrossRef]
- Wickham, H. Getting Started with Ggplot2. In ggplot2: Elegant Graphics for Data Analysis; Springer: Berlin/Heidelberg, Germany, 2016; pp. 11–31. [Google Scholar]
- Oksanen, J.; Blanchet, F.G.; Kindt, R.; Legendre, P.; Minchin, P.R.; O’hara, R.; Simpson, G.L.; Solymos, P.; Stevens, M.H.H.; Wagner, H. Package ‘Vegan’. Community Ecology Package, version 2.6-6.1. Available online: http://CRAN.R-project.org/package=vegan (accessed on 19 August 2025).
- Kassambara, A. Rstatix: Pipe-Friendly Framework for Basic Statistical Tests, version 0.7.2; CRAN Contrib. Packages, 2023. Available online: https://rpkgs.datanovia.com/rstatix/ (accessed on 19 August 2025). [CrossRef]
- Ebbert, D. Chisq.Posthoc.Test: A Post Hoc Analysis for Pearson’s Chi-Squared Test for Count Data, version 0.1.2; 2019. Available online: https://cran.r-project.org/web/packages/chisq.posthoc.test/chisq.posthoc.test.pdf (accessed on 19 August 2025).
- Beasley, T.M.; Schumacker, R.E. Multiple Regression Approach to Analyzing Contingency Tables: Post Hoc and Planned Comparison Procedures. J. Exp. Educ. 1995, 64, 79–93. [Google Scholar] [CrossRef]
- Godhe, A.; Rynearson, T. The Role of Intraspecific Variation in the Ecological and Evolutionary Success of Diatoms in Changing Environments. Philos. Trans. R. Soc. B 2017, 372, 20160399. [Google Scholar] [CrossRef]
- Julius, M.L.; Theriot, E.C. Theriot, E.C. The Diatoms: A Primer. In The Diatoms: Applications for the Environmental and Earth Sciences; Smol, J.P., Stoermer, E.F., Eds.; Cambridge University Press: Cambridge, UK, 2010; pp. 8–22. ISBN 978-0-521-50996-1. [Google Scholar]
- Martinez Arbizu, P. pairwiseAdonis: Pairwise Multilevel Comparison Using Adonis, version 0.4; 2020. Available online: https://github.com/pmartinezarbizu/pairwiseAdonis (accessed on 19 August 2025).
- Goergen, E.A.; Schopmeyer, S.; Moulding, A.L.; Moura, A.; Kramer, P.; Viehman, T.S. Coral Reef Restoration Monitoring Guide: Methods to Evaluate Restoration Success from Local to Ecosystem Scales; NOS and NCCOS: Silver Spring, MD, USA, 2020. [CrossRef]
- Gladfelter, E.H.; Monahan, R.K.; Gladfelter, W.B. Growth Rates of Five Reef-Building Corals in the Northeastern Caribbean. Bull. Mar. Sci. 1978, 28, 728–734. [Google Scholar]
- Zawada, K.J.; Dornelas, M.; Madin, J.S. Quantifying Coral Morphology. Coral Reefs 2019, 38, 1281–1292. [Google Scholar] [CrossRef]
- Rossi, S.; Schubert, N.; Brown, D.; Soares, M.d.O.; Grosso, V.; Rangel-Huerta, E.; Maldonado, E. Linking Host Morphology and Symbiont Performance in Octocorals. Sci. Rep. 2018, 8, 12823. [Google Scholar] [CrossRef]
- Lough, J.; Barnes, D. Environmental Controls on Growth of the Massive Coral Porites. J. Exp. Mar. Biol. Ecol. 2000, 245, 225–243. [Google Scholar] [CrossRef]
- Browne, N. Spatial and Temporal Variations in Coral Growth on an Inshore Turbid Reef Subjected to Multiple Disturbances. Mar. Environ. Res. 2012, 77, 71–83. [Google Scholar] [CrossRef] [PubMed]
- Calle-Triviño, J.; Muñiz-Castillo, A.I.; Cortés-Useche, C.; Morikawa, M.; Sellares-Blasco, R.; Arias-González, J.E. Approach to the Functional Importance of Acropora cervicornis in Outplanting Sites in the Dominican Republic. Front. Mar. Sci. 2021, 8, 668325. [Google Scholar] [CrossRef]
- Young, C.N.; Schopmeyer, S.; Lirman, D. A Review of Reef Restoration and Coral Propagation Using the Threatened Genus Acropora in the Caribbean and Western Atlantic. Bull. Mar. Sci. 2012, 88, 1075–1098. [Google Scholar] [CrossRef]
- Highsmith, R.C. Reproduction by Fragmentation in Corals. Mar. Ecol. Prog. Ser. Oldendorf 1982, 7, 207–226. [Google Scholar] [CrossRef]
- Lirman, D. Fragmentation in the Branching Coral Acropora palmata (Lamarck): Growth, Survivorship, and Reproduction of Colonies and Fragments. J. Exp. Mar. Biol. Ecol. 2000, 251, 41–57. [Google Scholar] [CrossRef] [PubMed]
- Clark, S.; Edwards, A. Coral Transplantation as an Aid to Reef Rehabilitation: Evaluation of a Case Study in the Maldive Islands. Coral Reefs 1995, 14, 201–213. [Google Scholar] [CrossRef]
- Yap, H.T.; Alino, P.M.; Gomez, E.D. Trends in Growth and Mortality of Three Coral Species(Anthozoa: Scleractinia), Including Effects of Transplantation. Mar. Ecol. Prog. Ser. Oldendorf 1992, 83, 91–101. [Google Scholar] [CrossRef]
- Ware, M.; Garfield, E.N.; Nedimyer, K.; Levy, J.; Kaufman, L.; Precht, W.; Winters, R.S.; Miller, S.L. Survivorship and Growth in Staghorn Coral (Acropora cervicornis) Outplanting Projects in the Florida Keys National Marine Sanctuary. PLoS ONE 2020, 15, e0231817. [Google Scholar] [CrossRef]
- Cabaitan, P.C.; Yap, H.T.; Gomez, E.D. Performance of Single versus Mixed Coral Species for Transplantation to Restore Degraded Reefs. Restor. Ecol. 2015, 23, 349–356. [Google Scholar] [CrossRef]
- Conti-Jerpe, I.E.; Thompson, P.D.; Wong, C.W.M.; Oliveira, N.L.; Duprey, N.N.; Moynihan, M.A.; Baker, D.M. Trophic Strategy and Bleaching Resistance in Reef-Building Corals. Sci. Adv. 2020, 6, eaaz5443. [Google Scholar] [CrossRef]
- Cybulski, J.D. Hong Kong’s Coral Assemblages through Time: A Paleoecological and Geochemical Look at Human-Driven Change. Ph.D. Thesis, The University of Hong Kong, Hong Kong SAR, China, 2021. [Google Scholar]
- Liao, H.; Hu, S.; Yang, H.; Wang, L.; Tanaka, S.; Takigawa, I.; Li, W.; Fan, H.; Gong, J.P. Data-Driven de Novo Design of Super-Adhesive Hydrogels. Nature 2025, 644, 89–95. [Google Scholar] [CrossRef] [PubMed]
- Bohnsack, J.A.; Sutherland, D.L. Artificial Reef Research: A Review with Recommendations for Future Priorities. Bull. Mar. Sci. 1985, 37, 11–39. [Google Scholar]
- Arena, P.T.; Jordan, L.K.B.; Spieler, R.E. Fish Assemblages on Sunken Vessels and Natural Reefs in Southeast Florida, USA. Hydrobiologia 2007, 580, 157–171. [Google Scholar] [CrossRef]
- Gratwicke, B.; Speight, M.R. The Relationship between Fish Species Richness, Abundance and Habitat Complexity in a Range of Shallow Tropical Marine Habitats. J. Fish Biol. 2005, 66, 650–667. [Google Scholar] [CrossRef]
- Santos, L.N.; Araujo, F.G.; Brotto, D.S. Artificial Structures as Tools for Fish Habitat Rehabilitation in a Neotropical Reservoir. Aquat. Conserv. Mar. Freshw. Ecosyst. 2008, 18, 896. [Google Scholar] [CrossRef]
- Sherman, R.L.; Gilliam, D.S.; Spieler, R.E. Artificial Reef Design: Void Space, Complexity, and Attractants. ICES J. Mar. Sci. 2002, 59, S196–S200. [Google Scholar] [CrossRef]
- Burkepile, D.E.; Hay, M.E. Herbivore Species Richness and Feeding Complementarity Affect Community Structure and Function on a Coral Reef. Proc. Natl. Acad. Sci. USA 2008, 105, 16201–16206. [Google Scholar] [CrossRef] [PubMed]
- Hughes, T.P.; Rodrigues, M.J.; Bellwood, D.R.; Ceccarelli, D.; Hoegh-Guldberg, O.; McCook, L.; Moltschaniwskyj, N.; Pratchett, M.S.; Steneck, R.S.; Willis, B. Phase Shifts, Herbivory, and the Resilience of Coral Reefs to Climate Change. Curr. Biol. 2007, 17, 360–365. [Google Scholar] [CrossRef]
- Mumby, P.; Steneck, R. Coral Reef Management and Conservation in Light of Rapidly Evolving Ecological Paradigms. Trends Ecol. Evol. 2008, 23, 555–563. [Google Scholar] [CrossRef] [PubMed]
- Glynn, P.W.; D’Croz, L. Experimental Evidence for High Temperature Stress as the Cause of El Niño-Coincident Coral Mortality. Coral Reefs 1990, 8, 181–191. [Google Scholar] [CrossRef]
- Carreiro-Silva, M.; McClanahan, T.R. Macrobioerosion of Dead Branching Porites, 4 and 6 Years after Coral Mass Mortality. Mar. Ecol. Prog. Ser. 2012, 458, 103–122. [Google Scholar] [CrossRef]
- Yachi, S.; Loreau, M. Biodiversity and Ecosystem Productivity in a Fluctuating Environment: The Insurance Hypothesis. Proc. Natl. Acad. Sci. USA 1999, 96, 1463–1468. [Google Scholar] [CrossRef] [PubMed]
- McGrady-Steed, J.; Harris, P.M.; Morin, P.J. Biodiversity Regulates Ecosystem Predictability. Nature 1997, 390, 162–165. [Google Scholar] [CrossRef]
- Naeem, S.; Li, S. Biodiversity Enhances Ecosystem Reliability. Nature 1997, 390, 507–509. [Google Scholar] [CrossRef]
- Gann, G.D.; McDonald, T.; Walder, B.; Aronson, J.; Nelson, C.R.; Jonson, J.; Hallett, J.G.; Eisenberg, C.; Guariguata, M.R.; Liu, J. International Principles and Standards for the Practice of Ecological Restoration. Restor. Ecol. 2019, 27, S1–S46. [Google Scholar] [CrossRef]
- McDonald, T.; Gann, G.; Jonson, J.; Dixon, K. International Standards for the Practice of Ecological Restoration–Including Principles and Key Concepts; Society for Ecological Restoration: Washington, DC, USA, 2016. [Google Scholar]
- Dang, H.; Lovell, C.R. Microbial Surface Colonization and Biofilm Development in Marine Environments. Microbiol. Mol. Biol. Rev. 2015, 80, 91–138. [Google Scholar] [CrossRef] [PubMed]
- Virta, L.; Gammal, J.; Järnström, M.; Bernard, G.; Soininen, J.; Norkko, J.; Norkko, A. The Diversity of Benthic Diatoms Affects Ecosystem Productivity in Heterogeneous Coastal Environments. Ecology 2019, 100, e02765. [Google Scholar] [CrossRef]
- Perkol-Finkel, S.; Benayahu, Y. Recruitment of Benthic Organisms onto a Planned Artificial Reef: Shifts in Community Structure One Decade Post-Deployment. Mar. Environ. Res. 2005, 59, 79–99. [Google Scholar] [CrossRef]
- Spagnolo, A.; Cuicchi, C.; Punzo, E.; Santelli, A.; Scarcella, G.; Fabi, G. Patterns of Colonization and Succession of Benthic Assemblages in Two Artificial Substrates. J. Sea Res. 2014, 88, 78–86. [Google Scholar] [CrossRef]
- Pickett, S.T.A.; Cadenasso, M.L.; Rosi-Marshall, E.J.; Belt, K.T.; Groffman, P.M.; Grove, J.M.; Irwin, E.G.; Kaushal, S.S.; LaDeau, S.L.; Nilon, C.H.; et al. Dynamic Heterogeneity: A Framework to Promote Ecological Integration and Hypothesis Generation in Urban Systems. Urban Ecosyst. 2017, 20, 1–14. [Google Scholar] [CrossRef]
- Todd, P.A.; Heery, E.C.; Loke, L.H.L.; Thurstan, R.H.; Kotze, D.J.; Swan, C. Towards an Urban Marine Ecology: Characterizing the Drivers, Patterns and Processes of Marine Ecosystems in Coastal Cities. Oikos 2019, 128, 1215–1242. [Google Scholar] [CrossRef]
- Bae, S.; Ubagan, M.D.; Shin, S.; Kim, D.G. Comparison of Recruitment Patterns of Sessile Marine Invertebrates According to Substrate Characteristics. Int. J. Environ. Res. Public Health 2022, 19, 1083. [Google Scholar] [CrossRef]
- Juan, S.d.; Thrush, S.F.; Hewitt, J.E. Counting on β-Diversity to Safeguard the Resilience of Estuaries. PLoS ONE 2013, 8, e65575. [Google Scholar] [CrossRef]
- Oliver, T.H.; Heard, M.S.; Isaac, N.J.B.; Roy, D.B.; Procter, D.; Eigenbrod, F.; Freckleton, R.; Hector, A.; Orme, C.D.L.; Petchey, O.L.; et al. Biodiversity and Resilience of Ecosystem Functions. Trends Ecol. Evol. 2015, 30, 673–684. [Google Scholar] [CrossRef] [PubMed]
- Connell, S.D. Floating Pontoons Create Novel Habitats for Subtidal Epibiota. J. Exp. Mar. Biol. Ecol. 2000, 247, 183–194. [Google Scholar] [CrossRef] [PubMed]
- Perkol-Finkel, S.; Benayahu, Y. Differential Recruitment of Benthic Communities on Neighboring Artificial and Natural Reefs. J. Exp. Mar. Biol. Ecol. 2007, 340, 25–39. [Google Scholar] [CrossRef]
- Atkinson, J.; Brudvig, L.A.; Mallen-Cooper, M.; Nakagawa, S.; Moles, A.T.; Bonser, S.P. Terrestrial Ecosystem Restoration Increases Biodiversity and Reduces Its Variability, but Not to Reference Levels: A Global Meta-analysis. Ecol. Lett. 2022, 25, 1725–1737. [Google Scholar] [CrossRef]
- Deiner, K.; Walser, J.-C.; Mächler, E.; Altermatt, F. Choice of Capture and Extraction Methods Affect Detection of Freshwater Biodiversity from Environmental DNA. Biol. Conserv. 2015, 183, 53–63. [Google Scholar] [CrossRef]
- Nichols, R.V.; Vollmers, C.; Newsom, L.A.; Wang, Y.; Heintzman, P.D.; Leighton, M.; Green, R.E.; Shapiro, B. Minimizing Polymerase Biases in Metabarcoding. Mol. Ecol. Resour. 2018, 18, 927–939. [Google Scholar] [CrossRef]
- Zinger, L.; Bonin, A.; Alsos, I.G.; Bálint, M.; Bik, H.; Boyer, F.; Chariton, A.A.; Creer, S.; Coissac, E.; Deagle, B.E.; et al. DNA Metabarcoding—Need for Robust Experimental Designs to Draw Sound Ecological Conclusions. Mol. Ecol. 2019, 28, 1857–1862. [Google Scholar] [CrossRef]
- Shelton, A.O.; Gold, Z.J.; Jensen, A.J.; D′Agnese, E.; Andruszkiewicz Allan, E.; Van Cise, A.; Gallego, R.; Ramón-Laca, A.; Garber-Yonts, M.; Parsons, K.; et al. Toward Quantitative Metabarcoding. Ecology 2023, 104, e3906. [Google Scholar] [CrossRef]
- Skelton, J.; Cauvin, A.; Hunter, M.E. Environmental DNA Metabarcoding Read Numbers and Their Variability Predict Species Abundance, but Weakly in Non-Dominant Species. Environ. DNA 2023, 5, 1092–1104. [Google Scholar] [CrossRef]
- Joseph, C.; Faiq, M.E.; Li, Z.; Chen, G. Persistence and Degradation Dynamics of eDNA Affected by Environmental Factors in Aquatic Ecosystems. Hydrobiologia 2022, 849, 4119–4133. [Google Scholar] [CrossRef]
- Xie, R.; Zhao, G.; Yang, J.; Wang, Z.; Xu, Y.; Zhang, X.; Wang, Z. eDNA Metabarcoding Revealed Differential Structures of Aquatic Communities in a Dynamic Freshwater Ecosystem Shaped by Habitat Heterogeneity. Environ. Res. 2021, 201, 111602. [Google Scholar] [CrossRef] [PubMed]
- Shogren, A.J.; Tank, J.L.; Andruszkiewicz, E.; Olds, B.; Mahon, A.R.; Jerde, C.L.; Bolster, D. Controls on eDNA Movement in Streams: Transport, Retention, and Resuspension. Sci. Rep. 2017, 7, 5065. [Google Scholar] [CrossRef] [PubMed]
- Turner, C.R.; Uy, K.L.; Everhart, R.C. Fish Environmental DNA Is More Concentrated in Aquatic Sediments than Surface Water. Biol. Conserv. 2015, 183, 93–102. [Google Scholar] [CrossRef]
- Prosser, C.M.; Hedgpeth, B.M. Effects of Bioturbation on Environmental DNA Migration through Soil Media. PLoS ONE 2018, 13, e0196430. [Google Scholar] [CrossRef]
- Jeunen, G.-J.; Knapp, M.; Spencer, H.G.; Lamare, M.D.; Taylor, H.R.; Stat, M.; Bunce, M.; Gemmell, N.J. Environmental DNA (eDNA) Metabarcoding Reveals Strong Discrimination among Diverse Marine Habitats Connected by Water Movement. Mol. Ecol. Resour. 2019, 19, 426–438. [Google Scholar] [CrossRef] [PubMed]
- URycki, D.R.; Kirtane, A.A.; Aronoff, R.; Avila, C.C.; Blackman, R.C.; Carraro, L.; Evrard, O.; Good, S.P.; Hoyos, J.D.C.; López-Rodríguez, N.; et al. A New Flow Path: eDNA Connecting Hydrology and Biology. WIREs Water 2024, 11, e1749. [Google Scholar] [CrossRef]
- Bayraktarov, E.; Stewart-Sinclair, P.J.; Brisbane, S.; Boström-Einarsson, L.; Saunders, M.I.; Lovelock, C.E.; Possingham, H.P.; Mumby, P.J.; Wilson, K.A. Motivations, Success, and Cost of Coral Reef Restoration. Restor. Ecol. 2019, 27, 981–991. [Google Scholar] [CrossRef]
- Sato, M.; Inoue, N.; Nambu, R.; Furuichi, N.; Imaizumi, T.; Ushio, M. Quantitative Assessment of Multiple Fish Species around Artificial Reefs Combining Environmental DNA Metabarcoding and Acoustic Survey. Sci. Rep. 2021, 11, 19477. [Google Scholar] [CrossRef] [PubMed]
- Inoue, N.; Sato, M.; Furuichi, N.; Imaizumi, T.; Ushio, M. The Relationship between eDNA Density Distribution and Current Fields around an Artificial Reef in the Waters of Tateyama Bay, Japan. Metabarcoding Metagenomics 2022, 6, e87415. [Google Scholar] [CrossRef]
- Krolow, A.P. Assessing the Diversity of Fish Communities at or Around Artificial Reefs Along the Louisiana Coast Through the Use of Environmental DNA (eDNA). Master’s Thesis, Southeastern Louisiana University, Hammond, LA, USA, 2019. [Google Scholar]
- Krolow, A.D.; Geheber, A.D.; Piller, K.R. If You Build It, Will They Come? An Environmental DNA Assessment of Fish Assemblages on Artificial Reefs in the Northern Gulf of Mexico. Trans. Am. Fish. Soc. 2022, 151, 297–321. [Google Scholar] [CrossRef]
- Miyajima-Taga, Y.; Sato, M.; Oi, K.; Furuichi, N.; Inoue, N. Fine-Scale Spatial Distribution of a Fish Community in Artificial Reefs Investigated Using an Underwater Drone and Environmental DNA Analysis. Mar. Ecol. Prog. Ser. 2024, 740, 123–144. [Google Scholar] [CrossRef]
- Levy, N.; Simon-Blecher, N.; Ben-Ezra, S.; Yuval, M.; Doniger, T.; Leray, M.; Karako-Lampert, S.; Tarazi, E.; Levy, O. Evaluating Biodiversity for Coral Reef Reformation and Monitoring on Complex 3D Structures Using Environmental DNA (eDNA) Metabarcoding. Sci. Total Environ. 2023, 856, 159051. [Google Scholar] [CrossRef] [PubMed]
- Knoester, E.; Rienstra, J.; Schürmann, Q.; Wolma, A.; Murk, A.; Osinga, R. Community-Managed Coral Reef Restoration in Southern Kenya Initiates Reef Recovery Using Various Artificial Reef Designs. Front. Mar. Sci. 2023, 10, 1152106. [Google Scholar] [CrossRef]
- Harrington, L.; Fabricius, K.; De’ath, G.; Negri, A. Recognition and Selection of Settlement Substrata Determine Post-Settlement Survival in Corals. Ecology 2004, 85, 3428–3437. [Google Scholar] [CrossRef]
- Webster, N.S.; Soo, R.; Cobb, R.; Negri, A.P. Elevated Seawater Temperature Causes a Microbial Shift on Crustose Coralline Algae with Implications for the Recruitment of Coral Larvae. ISME J. 2011, 5, 759–770. [Google Scholar] [CrossRef]
- Bessey, C.; Jarman, S.N.; Berry, O.; Olsen, Y.S.; Bunce, M.; Simpson, T.; Power, M.; McLaughlin, J.; Edgar, G.J.; Keesing, J. Maximizing Fish Detection with eDNA Metabarcoding. Environ. DNA 2020, 2, 493–504. [Google Scholar] [CrossRef]
- Clark, D.E.; Pilditch, C.A.; Pearman, J.K.; Ellis, J.I.; Zaiko, A. Environmental DNA Metabarcoding Reveals Estuarine Benthic Community Response to Nutrient Enrichment—Evidence from an in-Situ Experiment. Environ. Pollut. 2020, 267, 115472. [Google Scholar] [CrossRef]
- Couton, M.; Lévêque, L.; Daguin-Thiébaut, C.; Comtet, T.; Viard, F. Water eDNA Metabarcoding Is Effective in Detecting Non-Native Species in Marinas, but Detection Errors Still Hinder Its Use for Passive Monitoring. Biofouling 2022, 38, 367–383. [Google Scholar] [CrossRef] [PubMed]
- Liang, D.; Xia, J.; Song, J.; Sun, H.; Xu, W. Using eDNA to Identify the Dynamic Evolution of Multi-Trophic Communities under the Eco-Hydrological Changes in River. Front. Environ. Sci. 2022, 10, 929541. [Google Scholar] [CrossRef]
- Larson, E.R.; Graham, B.M.; Achury, R.; Coon, J.J.; Daniels, M.K.; Gambrell, D.K.; Jonasen, K.L.; King, G.D.; LaRacuente, N.; Perrin-Stowe, T.I.; et al. From eDNA to Citizen Science: Emerging Tools for the Early Detection of Invasive Species. Front. Ecol. Environ. 2020, 18, 194–202. [Google Scholar] [CrossRef]
- Uthicke, S.; Robson, B.; Doyle, J.R.; Logan, M.; Pratchett, M.S.; Lamare, M. Developing an Effective Marine eDNA Monitoring: eDNA Detection at Pre-Outbreak Densities of Corallivorous Seastar (Acanthaster Cf. Solaris). Sci. Total Environ. 2022, 851, 158143. [Google Scholar] [CrossRef] [PubMed]
- Ip, Y.C.A.; Chang, J.J.M.; Tun, K.P.P.; Meier, R.; Huang, D. Multispecies Environmental DNA Metabarcoding Sheds Light on Annual Coral Spawning Events. Mol. Ecol. 2023, 32, 6474–6488. [Google Scholar] [CrossRef] [PubMed]





| Fish | Invertebrate | ||||
|---|---|---|---|---|---|
| Common Name | Scientific Name | Diet | Common Name | Scientific Name | Diet |
| Grouper | Epinephelinae | Crustaceans, fish, molluscs [41] | Long-spined sea urchin | Diadema setosum | Macroalgae, seagrasses, diatoms [42] |
| Wrasse | Labridae | Benthic invertebrates, small fish, and (in cleaner wrasse) ectoparasites [40] | Decorator urchin | Salmacis sphaeroides | Macroalgae, seagrasses, detritus, jellyfish, conspecifics [43,44] |
| Sweetlips | Plectorhinchinae | Small fish, benthic invertebrates [40] | Black sea cucumber | Holothuria leucospilota | Detritus (scavenged organics in sediment) [45] |
| Seahorse | Hippocampus | Amphipods, copepods, shrimp, plankton [40,46] | |||
| Breakage | est. | SE | df | Z-Score (Glmer) | p-Value |
| A-Pv A-Pt Pv-Pt | 2.435 3.209 0.775 | 0.308 0.429 0.504 | Inf Inf Inf | 7.0899 7.472 1.527 | <0.0001 <0.0001 0.2734 |
| Extension Rate | Z-Score (glmmTMB) | ||||
| A-Pv A-Pt Pv-Pt | 0.517 0.913 0.396 | 0.0493 0.0479 0.0493 | Inf Inf Inf | 10.482 19.071 8.030 | <0.0001 <0.0001 <0.0001 |
| Site | Reference | Restored | Unrestored | |||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Taxa | Type | Total Count | Mean Count | SD | Total Count | Mean Count | SD | Total Count | Mean Count | SD |
| Grouper | Fish | 65 | 4.6 | 2.6 | 86 | 6.1 | 4.0 | 10 | 0.7 | 1.1 |
| Other Fish | Fish | 87 | 14.5 | 2.4 | 103 | 17.2 | 7.3 | 12 | 2.0 | 0.9 |
| Sweetlips | Fish | 10 | 0.7 | 1.2 | 20 | 1.4 | 2.5 | 4 | 0.3 | 0.6 |
| Wrasse | Fish | 47 | 3.4 | 3.9 | 37 | 2.6 | 3.2 | 7 | 0.5 | 0.9 |
| Black sea cucumber | Invert | 992 | 70.9 | 32.6 | 1937 | 138.4 | 52.5 | 1263 | 90.2 | 43.3 |
| Decorator urchin | Invert | 1336 | 95.4 | 92.8 | 974 | 69.6 | 82.2 | 447 | 31.9 | 39.8 |
| Long-spined urchin | Invert | 269 | 19.2 | 44.8 | 341 | 24.4 | 31.6 | 227 | 16.2 | 19.9 |
| H | df | p-Value | W (Pairwise) | p-Value (Pairwise) | |
|---|---|---|---|---|---|
| Fish | 13.6 | 2 | 0.001 | Ref-Res: 94 Ref-Un: 166 Res-Un: 168 | Ref-Res: 0.872 Ref-Un: 0.003 Res-Un: 0.003 |
| Invertebrate | 3.1 | 2 | 0.071 | Ref-Res: 73 Ref-Un: 124 Res-Un: 147 | Ref-Res: 0.260 Ref-Un: 0.260 Res-Un: 0.077 |
| df | SS | MS | F | p-Value | |
|---|---|---|---|---|---|
| Site | 2 | 243,432 | 121,716 | 2.225 | 0.154 |
| Residuals | 11 | 601,744 | 54,704 |
| Observed ASVs | % of All ASVs | % of Reads | % of Reads | ||
|---|---|---|---|---|---|
| Hellinger Transformed | No Transformation | ||||
| Unique | Unrestored | 880 | 22% | 11% | 18% |
| Restored | 1059 | 27% | 14% | 7% | |
| Reference | 1064 | 27% | 19% | 13% | |
| Shared | Unrestored—Restored—Reference | 213 | 5% | 23% | 27% |
| Unrestored—Restored | 341 | 9% | 14% | 19% | |
| Unrestored—Reference | 121 | 3% | 5% | 6% | |
| Restored—Reference | 303 | 8% | 15% | 11% | |
| Total | 3981 | 76% |
| df | SS | R2 | F | p | ||
|---|---|---|---|---|---|---|
| Jaccard | Site | 2 | 2.377 | 0.555 | 6.859 | 0.001 |
| Residual | 11 | 1.906 | 0.445 | |||
| Total | 13 | 4.284 | 1.000 | |||
| Bray-Curtis | Site | 2 | 1.867 | 0.510 | 5.714 | 0.001 |
| Residual | 11 | 1.797 | 0.490 | |||
| Total | 13 | 3.664 | 1.000 |
| Restored–Reference | Restored–Unrestored | Reference–Unrestored | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| df | SS | R2 | F | p | df | SS | R2 | F | p | df | SS | R2 | F | p | ||
| Jaccard | Site | 1 | 1.28 | 0.50 | 7.91 | 0.01 | 1 | 1.03 | 0.45 | 5.73 | <0.01 | 1 | 1.24 | 0.50 | 6.92 | <0.01 |
| Residual | 8 | 1.30 | 0.50 | 7 | 1.26 | 0.55 | 7 | 1.26 | 0.50 | |||||||
| Total | 9 | 2.58 | 1.00 | 8 | 2.29 | 1.00 | 8 | 2.50 | 1.00 | |||||||
| Bray-Curtis | Site | 1 | 0.95 | 0.45 | 6.42 | 0.01 | 1 | 0.76 | 0.38 | 4.23 | <0.01 | 1 | 1.10 | 0.49 | 6.60 | <0.01 |
| Residual | 8 | 1.18 | 0.55 | 7 | 1.25 | 0.62 | 7 | 1.16 | 0.51 | |||||||
| Total | 9 | 2.12 | 1.00 | 8 | 2.01 | 1.00 | 8 | 2.26 | 1.00 | |||||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yu, V.; Corley, A.D.; Lau, H.; Thompson, P.D.; Wan, Z.W.; Wong, J.C.Y.; Wong, Z.K.T.; Li, L.W.H.; McIlroy, S.E.; Baker, D.M. Assessing the Effectiveness of 3D-Printed Ceramic Structures for Coral Restoration: Growth, Survivorship, and Biodiversity Using Visual Surveys and eDNA. J. Mar. Sci. Eng. 2025, 13, 1605. https://doi.org/10.3390/jmse13091605
Yu V, Corley AD, Lau H, Thompson PD, Wan ZW, Wong JCY, Wong ZKT, Li LWH, McIlroy SE, Baker DM. Assessing the Effectiveness of 3D-Printed Ceramic Structures for Coral Restoration: Growth, Survivorship, and Biodiversity Using Visual Surveys and eDNA. Journal of Marine Science and Engineering. 2025; 13(9):1605. https://doi.org/10.3390/jmse13091605
Chicago/Turabian StyleYu, Vriko, Alison D. Corley, Horace Lau, Philip D. Thompson, Zhongyue Wilson Wan, Jane C. Y. Wong, Zoe Kwan Ting Wong, Louise Wai Hung Li, Shelby E. McIlroy, and David M. Baker. 2025. "Assessing the Effectiveness of 3D-Printed Ceramic Structures for Coral Restoration: Growth, Survivorship, and Biodiversity Using Visual Surveys and eDNA" Journal of Marine Science and Engineering 13, no. 9: 1605. https://doi.org/10.3390/jmse13091605
APA StyleYu, V., Corley, A. D., Lau, H., Thompson, P. D., Wan, Z. W., Wong, J. C. Y., Wong, Z. K. T., Li, L. W. H., McIlroy, S. E., & Baker, D. M. (2025). Assessing the Effectiveness of 3D-Printed Ceramic Structures for Coral Restoration: Growth, Survivorship, and Biodiversity Using Visual Surveys and eDNA. Journal of Marine Science and Engineering, 13(9), 1605. https://doi.org/10.3390/jmse13091605






