Abstract
The ecology of the mesopelagic fish genus Trachipterus, which is rarely found in oceans, remains unclear. In this study, we found 22 eggs of T. trachypterus and T. jacksonensis around the Ulleung Basin of the East/Japan Sea during ichthyoplankton surveys from 2019 to 2021. The eggs were identified through genetic relationships with the genus Trachipterus based on partial sequences (COI and 16S) or concatenated sequences of 13 protein-coding genes and 2 rRNA genes of mitochondrial DNA. T. trachypterus eggs were discovered in all seasons, but more frequently during the winter. One T. jacksonensis egg that appeared during the autumn was the first in the northwestern Pacific Ocean. Identifying Trachipterus pelagic eggs would provide insight into their spawning ecology and biogeography.
1. Introduction
Members of the genus Trachipterus (Lampriformes, Trachipteridae), which comprise six species (T. altivelis, T. jacksonensis, T. arcticus, T. fukuzakii, T. trachypterus, and T. ishikawae) [,], have elongated and compressed bodies with large eyes and are distributed in several oceans [,]. They are rarely caught by deep fishing gear or found inshore [,]. The species of Trachipterus have been reported based on a few specimens [,,,].
The first nominal species of Trachipterus is T. trachypterus []. Trachipterus has been described using juvenile specimens, which reside in shallower waters than adults. Due to its rarity and the morphological changes at early developmental stages, T. trachypterus larvae of different sizes are often incorrectly identified []. T. trachypterus eggs were first reported as three specimens in the Gulf of Napoli []. The Mediterranean Sea, where T. trachypterus eggs, larvae, and adults have been found several times, is a known spawning ground [].
Fish eggs are a key indicator of spawning and evidence of species intrusion [,]. Because most marine teleost fish release large numbers of pelagic eggs, their egg distribution density is higher than that of spawners []. Even in rare species, the probability of finding for eggs is higher than that for adult fish during the spawning season []. In addition, eggs with a shorter pelagic duration than larvae are in or close to spawning grounds []. The spawning grounds of the Japanese eel, Anguilla japonica, were revealed through DNA barcoding-based identification of their eggs after prolonged research [].
DNA barcoding enables species-level identification of fish eggs []. Pelagic marine fish eggs are typically transparent and round []. Eggs of the same species may have morphologically different embryos depending on developmental stage []. Unlike dramatic changes in morphology, DNA remains constant throughout life history [,]. Intra- and inter-specific genetic distances, typically based on mitochondrial DNA (mtDNA) sequences (COI, 12S, 16S, etc.), are analyzed to identify fish eggs and larvae species [,,].
We conducted COI barcoding of pelagic eggs collected from the East/Japan Sea from 2019 to 2021 and surveyed the literature that applied DNA barcoding to fish eggs near the study area. The eggs in this study were finally determined to belong to T. trachypterus and T. jacksonensis based on their COI, 16S rRNA, and mitogenome sequences. Here, we report T. trachypterus spawning and the first finding of T. jacksonensis based on eggs in the East/Japan Sea.
2. Materials and Methods
2.1. Pelagic Fish Egg Collection
Pelagic fish eggs were collected from 13 stations in the East/Japan Sea (Figure 1) using a zooplankton net (mouth diameter: 80 cm; mesh size: 300 µm) during research cruises on R/V Tamgu 3 in four seasons (winter: February and March; spring: April; summer: June and August; autumn: October and November) from 2019 to 2021. The water depths at the stations are 126–2203 m. The net was towed obliquely from 10 m above the bottom to the surface at the stations with a water depth of less than 300 m (st. 103-05, 209-05, and 209-07). At the other stations, the net was lowered to a depth of 300 m and was towed as above. Samples were preserved in 95% ethanol. Temperature and salinity were measured using CTD (SBE 911plus, Sea-Bird Scientific Inc., Bellevue, WA, USA).
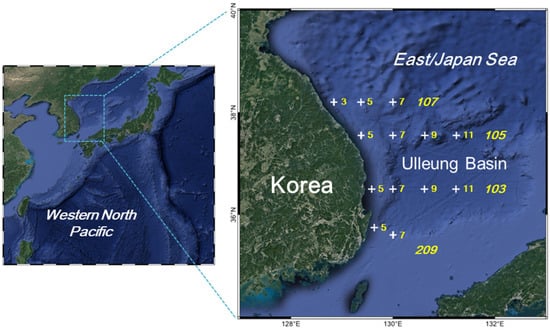
Figure 1.
Pelagic fish egg sampling stations in the East/Japan Sea, 2019–2021. Numbers with plus marks, station name; italic number, station lines. The base map is a Google Satellite map drawn using QGIS [] through the XYZ Tiles tool (https://mt1.google.com/vt/lyrs=s&x={x}&y={y}&z={z}, accessed on 10 March 2022).
Fish eggs were sorted from the samples using a stereomicroscope (M125C, Leica, Wetzlar, Germany). Among them, the 22 largest eggs were selected and photographed using a camera mounted on a stereomicroscope (SMZ18, Nikon, Tokyo, Japan).
2.2. Genomic DNA Extraction, PCR, and Sequencing and Sequence Analysis
Genomic DNA was extracted from 22 eggs according to the protocol of the DNeasy Blood & Tissue Kit (Qiagen, Hilden, Germany). The COI gene of mtDNA was amplified using the primers VF2_t1 (5′-CAACCAACCACAAAGACATTGGCAC-3′), FishF2_t1 (5′-TCGACTAATCATAAAGATATCGGCAC-3′), FishR2_t1 (5′-ACTTCAGGGTGACCGAAGAATCAGAA-3′), and FR1d_t1 (5′-ACCTCAGGGTGTCCGAARAAYCARAA-3′) [,]. The 20 µL PCR mixture was composed of 10 µL of 2X DNA free-Taq Master Mix including PCR buffer, dNTPs mixture, and Taq DNA polymerase (CellSafe, Gyeonggi, Korea), 0.2 µL of each of the four primers, 2 µL of genomic DNA, and 7.2 µL of distilled water. The PCR program consisted of initial denaturation at 94 °C for 3 min; 35 cycles of denaturation at 94 °C for 30 s, annealing at 52 °C for 40 s, extension at 72 °C for 1 min; and a final extension at 72 °C for 7 min. The PCR products were sequenced on a 3730xl DNA Analyzer (Applied Biosystems, Foster City, CA, USA).
Other mtDNA regions of the 7 eggs among the 22 eggs were also amplified for comparison with those of related taxa. The 16S rRNA gene of one egg was amplified using the 16Sar (5-CGCCTGTTTATCAAAAACAT-3) and 16Sbr (5-CCGGTCTGAACTCAGATCACGT-3) primers []. PCR and sequencing methods were used for COI analysis, except that the annealing temperature was 56 °C. Six eggs were selected to analyze mitogenome sequences in the consideration of sample conditions; three eggs were collected in 2019 and 2020 each. To obtain sufficient DNA to analyze the complete mitogenome of the six eggs, the whole genome was amplified following the REPLI-g Mini Kit (Qiagen) protocol. The amplified products were sequenced using a NovaSeq 6000 (Illumina, San Diego, CA, USA). A total of 385,684,576–474,309,190 raw reads (length: 150 bp) were obtained from the six eggs. The reads were mapped to a reference sequence (GenBank accession number: NC_003166, []) using Geneious R11 [] mapper. The resulting consensus sequences were annotated using MitoFish [] and Geneious R11. The three mitogenome sequences were constructed from three eggs (2002E3–E5), except for the d-loop region (mean coverage, 112 ± 119×–7866 ± 11,917×; 16,159–16,163 bp), but COI (1467–1551 bp) and 16S (1595–1683 bp) sequences were obtained from the other three eggs (1902E1, 1902E2, and 1904E1).
The COI, 16S, and mitogenome sequences from the 22 eggs were searched using BLAST [] and BOLD systems [] to identify related taxa. The sequences of the eggs, related taxa, and outgroups were aligned using Clustal Omega []. The sequences were used to analyze the maximum likelihood (ML) tree based on the best-fit substitution model [,,] and Kimura 2-parameter distances in MEGA X (ver. 11.0.10) []. The egg sequences were submitted to NCBI GenBank under accession numbers OM527130–OM527151, OM527153, OM574770–OM574772, and ON231742–ON231747 (Table 1). We also investigated the literature using DNA barcoding for the species identification of eggs around the study area, and sequences from the eggs of Shin et al. [] were used in this study.

Table 1.
GenBank accession numbers of eggs of this study and literature.
3. Results
3.1. Genetic Identification of Eggs
Twenty-two egg specimens were identified as T. trachypterus and T. jacksonensis according to genetic relationships based on COI, 16S, and mitogenome sequences.
The COI sequences of the 22 eggs and 3 eggs from the reference [] and genus Trachipterus consisted of three distinct clades (between genetic distances: average ± standard deviation, 0.158 ± 0.057; min, 0.083; max, 0.227; Table S1), with either two or three species in the maximum likelihood (ML) tree (Figure 2). Of the three clades, COI_Clade 1 had sequences from 24 eggs, T. altivelis and T. trachypterus, and COI_Clade 2 had sequences from one egg of this study, T. arcticus, T. jacksonensis, and Trachipterus sp. Although each clade consisted of sequences from different species, the genetic distances within the clades (COI_Clade 1, 0.009 ± 0.006; COI_Clade 2, 0.010 ± 0.007; and COI_Clade 3, 0.009 ± 0.005) were less than between the clades (0.158 ± 0.057), indicating that each clade represented the species.
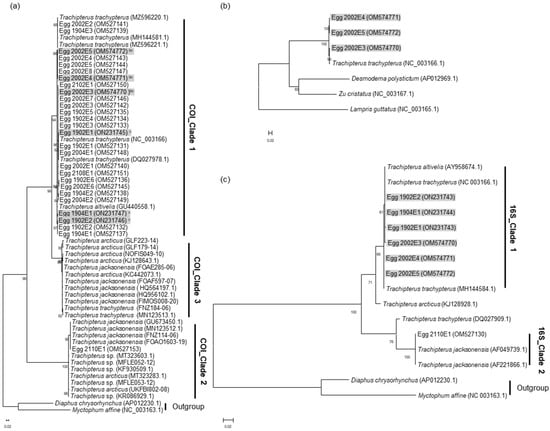
Figure 2.
Maximum likelihood (ML) tree constructed using mitochondrial DNA sequences of pelagic fish eggs, Trachipterus species, and outgroups. (a) COI ML tree (based on the HKY + G + I model) including COI sequences from 25 eggs. Sequences of eggs with superscripts (bc, c) were also analyzed in the trees in (b,c). (b) Thirteen protein-coding genes and two rRNA genes ML tree (based on the GTR + G + I model), including concatenated sequences from the three eggs. (c) 16S rRNA ML tree (based on the K2 + G model), including 16S rRNA sequences from seven eggs. Bootstrap values (1000 replicates) over 50% are shown on the branches. Sequences shaded in gray were obtained using shotgun sequencing.
Species of the two clades (COI_ Clades 1 and 2; Figure 2a), including the eggs, were re-analyzed based on 16S rRNA and mitogenome sequences. Among the eggs of COI_Clade 1, COI and 16S rRNA sequences for three eggs (1902E1, 1902E2, and 1904E1) and mitogenome sequences excluding the d-loop for three eggs (2002E3–E5) were obtained from mitogenome analysis (Table 1). The COI sequences of the six eggs were also located in COI_Clade 1. Concatenated sequences (13 protein-coding genes and two rRNAs) of the three eggs (2002E3–E5) formed a clade with those of T. trachypterus (NC_003166.1) (genetic distance, 0.007 ± 0.003; Table S2), and they were distinct from those of Desmodema polystictum and Zu cristatus of Trachipteridae (0.258 ± 0.012) (Figure 2b). The 16S rRNA sequences of the six eggs from the same samples of COI_Clade 1 and one egg (MH144584.1) from [] formed a clade with T. trachypterus and T. altivelis with very small genetic distances (0.002 ± 0.001; min, 0.000; max, 0.004; Table S3) (Figure 2c). Interestingly, the T. trachypterus 16S sequence (DQ027909.1, []) was distinct from 16S_Clade 1, although it was analyzed from the same specimen with the T. trachypterus COI sequence (DQ027978.1) of COI_Clade 1.
One egg (2110E1) formed COI_Clade 2 with T. arcticus, T. jacksonensis, T. altivelis, and Trachipterus sp. comprised a clade with only T. jacksonensis (genetic distance, 0.000 ± 0.000) in the 16S ML tree (16S_Clade 2; Figure 2c). The 16S sequence of T. arcticus (KJ128928.1) diverged sharply (0.107) from the clade of T. jacksonensis with the egg (2110E1; OM527130). The T. arcticus 16S rRNA sequence (KJ128928.1) was obtained from the same specimen as the T. arcticus COI sequence (KJ128643.1) in COI_Clade 3.
3.2. Pelagic Eggs of Two Trachipterus Species
The average diameter of T. trachypterus eggs was 3.2 ± 0.1 mm (min, 2.7 mm; max, 3.6 mm) (Figure 3; Table S4). The diameter of T. jacksonensis egg was 2.2 mm, smaller than that of T. trachypterus eggs. The internal morphology of eggs preserved in 95% ethanol was difficult to investigate. The common characteristics of T. trachypterus eggs that could be confirmed were a narrow perivitelline space and a lack of oil globules. The developed embryos had melanophores on the head, dorsal side, and on the yolk sac around the middle of the body.
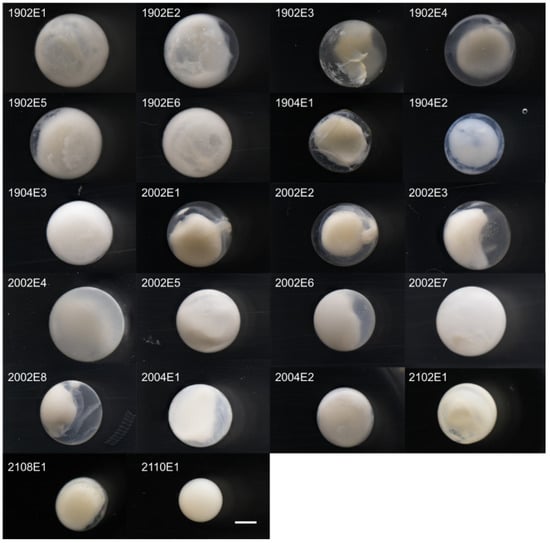
Figure 3.
Pelagic eggs of two Trachipterus species preserved in 95% ethanol. 1902E1–2108E1, T. trachypterus; 2110E1, T. jacksonensis. Scale bar, 1.0 mm.
3.3. Distribution of Trachipterus Eggs
T. trachypterus eggs appeared in all seasons around the Ulleung Basin and Dokdo, and one T. jacksonensis egg appeared once in autumn (Figure 4). The occurrence frequency of T. trachypterus eggs was highest during the winter and lowest during the summer and autumn. The surface temperature of the stations where eggs were detected ranged from 10.0 °C to 25.9 °C (Table S4). The mean surface temperature of the study area was the lowest during the winter and peaked in the summer (Table S5). Unlike the surface, the bottom temperature was constant at approximately 1 °C. Thermocline was generated by the large difference between the temperatures of the surface and bottom. The depth and strength of the thermocline varied according to season and station. The center of the thermocline was usually at a depth of 100–200 m (Figure S1). Salinity was approximately 33–34 psu, and there was no significant difference depending on season and depth, unlike the temperature.
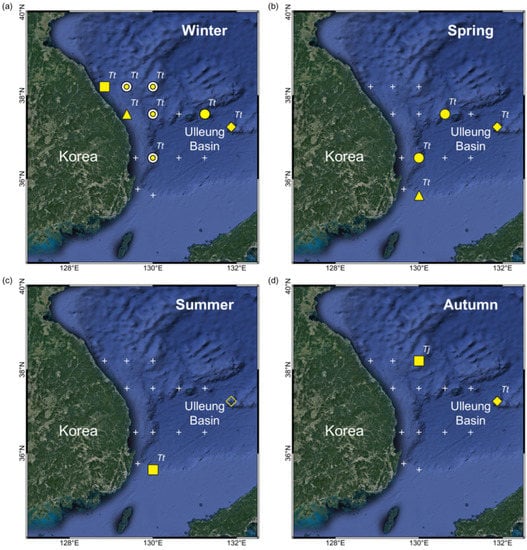
Figure 4.
Distributions of pelagic eggs of two Trachipterus species for the four seasons: (a) winter, (b) spring, (c) summer, and (d) autumn. Tt, T. trachypterus; Tj, T. jacksonensis; Sampling year of Trachipterus eggs: filled circle, 2019; fisheye, 2019 and 2020; filled triangle, 2020; filled square, 2021 of this study; filled diamond, 2017–2019 of Shin et al. []; no egg detection: cross, this study; empty diamond, Shin et al. [].
4. Discussion
DNA barcoding of fish eggs enables the investigation of the spawning ecology of various species [,]. Studies on fish eggs have been limited to a few species due to the difficulty in morphology-based egg identification []. Egg morphological characteristics were described based on larval fish hatching from eggs or eggs obtained from adult fish [,]. Recently, larval fish, which have more morphological features available for identification than eggs, have been analyzed using DNA barcoding to improve the accuracy of species identification []. However, DNA barcoding has limitations in determining species due to the lack of comparable DNA sequences, unsuitable DNA regions for barcoding, or sequences from misidentified specimens.
4.1. Genetic Identification of Eggs
The first DNA barcode for species identification of 22 eggs in this study was the COI sequence, which contains a lot of information available to identify fish. The maximum likelihood (ML) tree based on the COI sequences of the 25 eggs, including those of Shin et al. [] and Trachipterus, showed three distinct clades of Trachipterus (Figure 2a). Each clade was regarded as a species by considering the genetic distances of the intra- and inter-clades of the COI sequences of fishes [,]. However, the coexistence of sequences from two or three species within each clade was problematic: COI_Clade 1, T. altivelis, and T. trachypterus with 24 eggs; COI_Clade 2, T. arcticus, T. jacksonensis, Trachipterus sp., and one egg (2110E1); and COI_Clade 3, T. arcticus, T. jacksonensis, and T. trachypterus. This could be because COI sequences were analyzed from misidentified samples or were not appropriate for distinguishing the species.
The T. trachypterus mitogenome sequence was the key to determining the species of COI_Clade 1. The mitogenome sequences of three eggs (2002E3–E5) located in COI_Clade 1 formed a clade with that of T. trachypterus (NC_003166.1) (Figure 2b). In addition, genetic distances between the 16S rRNA sequences of the seven eggs (1902E1, 1902E2, 1904E1, 2002E3–E5, and DI_7 (MH144584.1)) and T. trachypterus (NC_003166, MH144581.1) were also very small (0.000–0.004; Table S3). These genetic relationships were a criterion for identifying 24 eggs, including those of Shin et al. [] in COI_Clade 1 as T. trachypterus.
COI_Clade 2, including one egg (2110E1; OM527153), was determined to be T. jacksonensis based on the distinct 16S_Clade 2 with sequences of T. jacksonensis and the one egg (2110E1; OM527130) in the 16S ML tree (Figure 2c). In addition, COI_Clade 3 was identified as T. arcticus based on the T. arcticus sequences (COI, KJ128643.1; 16S rRNA, KJ128928.1) showing the same position in the COI and 16S rRNA ML tree (Figure 2a,c).
Interestingly, the T. trachypterus 16S sequence (DQ027909.1, []) from the same sample with COI sequence (DQ027978.1, []) in the COI_Clade 1 was independently clustered in the 16S ML tree (Figure 2c). This indicates that the 16S rRNA sequence (DQ027909.1) was mishandled during the experiment or sequence analysis. In addition, T. altivelis (GU440558.1) of COI_Clade 1 and T. altivelis (AY958674.1) of 16S_Clade 1 would be derived from misidentification.
4.2. Trachipterus Eggs
T. trachypterus is widely distributed in the Mediterranean Sea and Atlantic Indo-Pacific Oceans [,,,,,,,,,,,,,,]. Its spawning area is the Mediterranean Sea, where eggs of Trachypterus taenia (synonym for T. trachypterus) have been found []. Lo Bianco [] collected three live eggs of T. taenia from a sampling gear lowered to a depth of 100–150 m in the Gulf of Napoli in February, May, and October 1905–1907. Spawning was assumed to occur year-round. Three eggs (diameter, 2.90–2.95 mm) were in late developmental stages. In this study, we confirmed that T. trachypterus spawned in all seasons and peaked in winter based on eggs continuously collected during the survey periods. If the T. trachypterus eggs in this study had similar ecological characteristics to the eggs from the Gulf of Napoli, the spawning depth could be estimated as the thermocline layer.
T. jacksonensis is distributed in the Southern Hemisphere (Australia, New Zealand, Africa, Brazil, and others) [,,,,,,,,,]. Adults have not been reported in the Northern Hemisphere. The eggs detected once in autumn in this study were the first in the Northern Hemisphere. Similarly to T. jacksonensis, Peristedion liorhynchus has not been recorded in Korean waters, but its eggs and larvae have been recorded []. This suggests that eggs may be useful for the detection of rare species.
5. Conclusions
In this study, we discovered T. trachypterus and T. jacksonensis eggs in the Ulleung Basin of the East/Japan Sea of which their sequences could be used for post verification. The Ulleung Basin is a spawning area for T. trachypterus along with the Mediterranean Sea; this is the first report of T. jacksonensis egg in the northwestern Pacific. Finding Trachipterus species eggs will elucidate their spawning ecology and geographic distribution.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/jmse10050637/s1, Figure S1: Vertical distribution of temperature at stations where Trachipterus eggs appeared in this study; Table S1: Pairwise genetic distances of the COI genes of pelagic eggs of this study, Trachipterus species, and outgroups. Outlined values belong to each clade of the maximum likelihood tree in Figure 2a; Table S2: Pairwise genetic distances of the concatenated mtDNA sequences of 13 protein-coding genes and two rRNA genes of pelagic eggs of this study, Trachipterus species, and outgroups; Table S3: Pairwise genetic distances of the 16S rRNA genes of pelagic egg of this study, Trachipterus species, and outgroups. Outlined values belong to each clade of the maximum likelihood tree in Figure 2c; Table S4: Information on Trachipterus eggs of this study and literature; Table S5: Mean and standard deviation values of temperature and salinity from 2019 to 2021 in the East/Japan Sea.
Author Contributions
Conceptualization, H.-y.C.; methodology, H.-y.C. and S.K.; validation, H.-y.C. and S.K.; formal analysis, H.-y.C. and S.K.; investigation, H.-c.C.; data curation, H.-y.C.; writing—original draft preparation, H.-y.C.; writing—review and editing, H.-y.C., H.-c.C., S.K., and S.-h.Y.; visualization, H.-y.C.; supervision, S.-h.Y.; project administration, H.-j.O.; funding acquisition, H.-j.O. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the National Institute of Fisheries Science (NIFS) grant (Study on the ecophysiology and occurrence prediction of jellyfish; R2022052).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The Sequence data that support the findings of this study are deposited in NCBI/GenBank (https://www.ncbi.nlm.nih.gov/genbank/) under accession numbers OM527130-OM527151, OM527153, OM574770-OM574772, and ON231742-ON231747.
Acknowledgments
We appreciate the captain and crews of the Tamgu3 research vessel for the survey. We are also grateful to the researchers in the NIFS for their assistance with sampling.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Integrated Taxonomic Information System (ITIS). Available online: http://www.itis.gov (accessed on 25 January 2022).
- Eschmeyer’s Catalog of Fishes. Available online: http://researcharchive.calacademy.org/research/ichthyology/catalog/fishcatmain.asp (accessed on 25 January 2022).
- Palmer, G. Trachipteridae. In Fishes of the North-Eastern Atlantic and the Mediterranean; Whitehead, P.J.P., Bauchot, M.L., Hureau, J.C., Nielsen, J., Tortonese, E., Eds.; United Nations Educational, Scientific and Cultural Organization: Paris, France, 1986; Volume 2, pp. 729–732. [Google Scholar]
- Olney, J.E.; Richards, W.J. Trachipteridae: Dealfishes and ribbonfishes. In Early Stages of Atlantic Fishes an Identification Guide for the Western Central North Atlantic; Richard, W.J., Ed.; Taylor and Francis: Boca Raton, FL, USA, 2006; pp. 1019–1026. [Google Scholar]
- Borme, D.; Voltolina, F. On the occurrence of ribbon fish Trachipterus trachypterus (Gmelin, 1789) in the Gulf of Trieste (northern Adriatic Sea). Ann. Ser. Hist. Nat. 2006, 16, 181. [Google Scholar]
- Tiralongo, F.; Lillo, A.O.; Tibullo, D.; Tondo, E.; Martire, C.L.; D’Agnese, R.; Macali, A.; Mancini, E.; Giovos, I.; Coco, S.; et al. Monitoring uncommon and non-indigenous fishes in Italian waters: One year of results for the AlienFish project. Reg. Stud. Mar. Sci. 2019, 28, 100606. [Google Scholar] [CrossRef]
- MoriTz, T.; STüMer, D.; JaKobSen, K.; JaKobSen, J. Observations on two live specimens of Trachipterus arcticus (Lampriformes: Trachipteridae) from the Azores. Cybium 2015, 39, 78–80. [Google Scholar]
- Angulo, A.; Lopez-Sanchez, M.I. New records of lampriform fishes (Teleostei: Lampriformes) from the Pacific coast of lower Central America, with comments on the diversity, taxonomy and distribution of the Lampriformes in the eastern Pacific Ocean. Zootaxa 2017, 4236, 573–591. [Google Scholar] [CrossRef] [PubMed]
- Lipej, L.; Trkov, D.; Mavrič, B. Occurrence of ribbon fish (Trachipterus trachypterus) in Slovenian waters (northern Adriatic Sea). Ann. Ser. Hist. Nat. 2018, 28, 129–134. [Google Scholar]
- Han, K.H.; Lee, S.H.; Kim, C.C.; Yu, T.S. Description of Morphology and Osteology of the Slender Ribbonfish, Trachipterus ishikawae Jordan & Snyder, 1901. Korean J. Ichthyol. 2020, 32, 136–142. [Google Scholar]
- Martin, J.; Hilton, E.J. A taxonomic review of the family Trachipteridae (Lampridiformes), with an emphasis on taxa distributed in the Western Pacific Ocean. Zootaxa 2021, 5039, 301–351. [Google Scholar] [CrossRef]
- Lo Bianco, S. Uova e larve di Trachypterus taenia Bl. Mitt. Zool. Stn. Neapel. 1908, 19, 1–17. [Google Scholar]
- Palmer, G. The dealfishes (Trachipteridae) of the Mediterranean and north-east Atlantic. Bull. Br. Mus. Nat. Hist. Zool. 1961, 7, 335–351. [Google Scholar] [CrossRef]
- Lewis, L.A.; Richardson, D.E.; Zakharov, E.V.; Hanner, R. Integrating DNA barcoding of fish eggs into ichthyoplankton monitoring programs. Fish. Bull. 2016, 114, 153–165. [Google Scholar] [CrossRef]
- Choi, H.Y.; Chin, B.S.; Park, G.S.; Kim, S. Evidence of intrusion of a rare Species, Peristedion liorhynchus, into Korean waters based on high-throughput sequencing of the mixed fish eggs. Korean J. Icthyol. 2022, 1, 8–15. [Google Scholar] [CrossRef]
- Houde, E.D. Fish early life dynamics and recruitment variability. Am. Fish. Soc. Symp. 2018, 2, 17–29. [Google Scholar]
- Choi, H.Y.; Oh, J.; Kim, S. Genetic identification of eggs from four species of Ophichthidae and Congridae (Anguilliformes) in the northern East China Sea. PLoS ONE 2018, 13, e0195382. [Google Scholar] [CrossRef] [PubMed]
- Tsukamoto, K.; Chow, S.; Otake, T.; Kurogi, H.; Mochioka, N.; Miller, M.J.; Aoyama, J.; Kimura, S.; Watanabe, S.; Yoshinaga, T.; et al. Oceanic spawning ecology of freshwater eels in the western North Pacific. Nat. Commun. 2011, 2, 179. [Google Scholar] [CrossRef]
- Shao, K.T.; Chen, K.C.; Wu, J.H. Identification of marine fish eggs in Taiwan using light microscopy, scanning electric microscopy and mtDNA sequencing. Mar. Freshw. Res. 2002, 53, 355–365. [Google Scholar] [CrossRef]
- Ahlstrom, E.H.; Moser, H.G. Characters useful in identification of pelagic marine fish eggs. CalCOFI Rep. 1980, 21, 121–131. [Google Scholar]
- Kendall, A.W.; Ahlstrom, E.H., Jr.; Moser, H.G. Early Life History Stages of Fishes and Their Characters. In Ontogeny and Systematics of Fishes; Moser, H.G., Richards, W.J., Cohen, D.M., Fahay, M.P., Kendall, A.W., Richardson, S.L., Eds.; American Society of Ichthyologists and Herpetologists: Kansas, MO, USA, 1984; pp. 31–33. [Google Scholar]
- Avise, J.C. Molecular Markers, Natural History and Evolution; Springer: Boston, MA, USA, 1994. [Google Scholar]
- Hebert, P.D.; Cywinska, A.; Ball, S.L.; DeWaard, J.R. Biological identifications through DNA barcodes. Proc. R. Soc. B Biol. Sci. 2003, 270, 313–321. [Google Scholar] [CrossRef]
- Choi, H.Y.; Jang, Y.S.; Oh, J.N.; Kim, S. Morphology of a Larval Hammerjaw Omosudis lowii Gunther 1887 (Aulopiformes, Omosudidae) Identified by Partial Mitochondrial 12S rRNA Gene Analysis. Korean J. Ichthyol. 2020, 32, 239–244. [Google Scholar] [CrossRef]
- Hou, G.; Xu, Y.; Chen, Z.; Zhang, K.; Huang, W.; Wang, J.; Zhou, J. Identification of Eggs and Spawning Zones of Hairtail Fishes Trichiurus (Pisces: Trichiuridae) in Northern South China Sea, Using DNA Barcoding. Front. Environ. Sci. 2021, 9, 703029. [Google Scholar] [CrossRef]
- Habib, K.A.; Neogi, A.K.; Rahman, M.; Oh, J.; Lee, Y.H.; Kim, C.G. DNA barcoding of brackish and marine water fishes and shellfishes of Sundarbans, the world’s largest mangrove ecosystem. PLoS ONE 2021, 16, e0255110. [Google Scholar] [CrossRef]
- QGIS Geographic Information System. Available online: http://www.qgis.org (accessed on 25 January 2022).
- Ward, R.D.; Zemlak, T.S.; Innes, B.H.; Last, P.R.; Hebert, P.D. DNA barcoding Australia’s fish species. Philos. Trans. R. Soc. B Biol. Sci. 2005, 360, 1847–1857. [Google Scholar] [CrossRef] [PubMed]
- Ivanova, N.V.; Zemlak, T.S.; Hanner, R.H.; Hebert, P.D. Universal primer cocktails for fish DNA barcoding. Mol. Ecol. Notes 2007, 7, 544–548. [Google Scholar] [CrossRef]
- Palumbi, S. Nucleic acids II: The polymerase chain reaction. In Molecular Systematics; Hillis, D.M., Moritz, C., Mable, B.K., Eds.; Sinauer Associates: Sunderland, MA, USA, 1996; pp. 205–247. [Google Scholar]
- Miya, M.; Kawaguchi, A.; Nishida, M. Mitogenomic exploration of higher teleostean phylogenies: A case study for moderate-scale evolutionary genomics with 38 newly determined complete mitochondrial DNA sequences. Mol. Biol. Evol. 2001, 18, 1993–2009. [Google Scholar] [CrossRef] [PubMed]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef]
- Iwasaki, W.; Fukunaga, T.; Isagozawa, R.; Yamada, K.; Maeda, Y.; Satoh, T.P.; Sado, T.; Mabuchi, K.; Takeshima, H.; Miya, M.; et al. MitoFish and MitoAnnotator: A mitochondrial genome database of fish with an accurate and automatic annotation pipeline. Mol. Biol. Evol. 2013, 30, 2531–2540. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Ratnasingham, S.; Hebert, P.D. BOLD: The Barcode of Life Data System (http://www.barcodinglife.org). Mol. Ecol. Notes 2007, 7, 355–364. [Google Scholar] [CrossRef]
- Sievers, F.; Wilm, A.; Dineen, D.; Gibson, T.J.; Karplus, K.; Li, W.; Li, W.; Lopez, R.; McWilliam, H.; Remmert, M.; et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 2011, 7, 539. [Google Scholar] [CrossRef]
- Hasegawa, M.; Kishino, H.; Yano, T. Dating the human-ape split by a molecular clock of mitochondrial DNA. J. Mol. Evol. 1985, 22, 160–174. [Google Scholar] [CrossRef]
- Nei, M.; Kumar, S. Molecular Evolution and Phylogenetics; Oxford University Press: New York, NY, USA, 2000. [Google Scholar]
- Kimura, M. A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 1980, 16, 111–120. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547. [Google Scholar] [CrossRef]
- Shin, U.C.; Yoon, S.; Kim, J.K.; Choi, G. Species composition of Ichthyoplankton off Dokdo in the East Sea. Korean J. Fish. Aquat. Sci. 2021, 54, 498–507. [Google Scholar]
- Sparks, J.S.; Dunlap, P.V.; Smith, W.L. Evolution and diversification of a sexually dimorphic luminescent system in ponyfishes (Teleostei: Leiognathidae), including diagnoses for two new genera. Cladistics 2005, 21, 305–327. [Google Scholar] [CrossRef]
- Harada, A.E.; Lindgren, E.A.; Hermsmeier, M.C.; Rogowski, P.A.; Terrill, E.; Burton, R.S. Monitoring spawning activity in a southern California marine protected area using molecular identification of fish eggs. PLoS ONE 2015, 10, e0134647. [Google Scholar] [CrossRef]
- Lin, H.Y.; Chiu, M.Y.; Shih, Y.M.; Chen, I.S.; Lee, M.A.; Shao, K.T. Species composition and assemblages of ichthyoplankton during summer in the East China Sea. Cont. Shelf Res. 2016, 126, 64–78. [Google Scholar] [CrossRef]
- Hernandez-Vazquez, S. Distribution of eggs and larvae from sardine and anchovy off California and Baja California, 1951–1989. CalCOFI Rep. 1994, 35, 94–107. [Google Scholar]
- Leis, J.M.; Moyer, J.T. Development of eggs, larvae and pelagic juveniles of three Indo-Pacific ostraciid fishes (Tetraodontiformes): Ostracion meleagris, Lactoria fornasini and L. diaphana. Jpn. J. Icthyol. 1985, 32, 189–202. [Google Scholar]
- Kim, K.M.; Kim, S.Y.; Song, M.Y.; Song, H.Y. Morphological development of egg and larvae of Hemiculter leucisculus. Korean J. Ichthyol. 2020, 32, 222–231. [Google Scholar] [CrossRef]
- Ko, H.L.; Wang, Y.T.; Chiu, T.S.; Lee, M.A.; Leu, M.Y.; Chang, K.Z.; Chen, W.Y.; Shao, K.T. Evaluating the accuracy of morphological identification of larval fishes by applying DNA barcoding. PLoS ONE 2013, 8, e53451. [Google Scholar] [CrossRef]
- Amaoka, K.; Nakaya, K.; Yabe, M. Fishes of Usujiri and adjacent waters in southern Hokkaido, Japan. Bull. Fac. Fish. Hokkaido Univ. 1989, 40, 254–277. [Google Scholar]
- Meléndez, R.; Clément, A. Trachipterus trachypterus (Gmelin, 1789) en el sur de Chile (Pisces: Lampridiformes: Trachipteridae). Cienc. Tecnol. Mar. 1992, 15, 43–47. [Google Scholar]
- Cortes, N.; Arriaza, M.; Oyarzún, C. Nuevos registros de Trachipterus trachypterus (Gmelin, 1789) para el Pacífico suroriental, con una revisión de ejemplares congenéricos de Chile (Osteichthyes, Trachipteridae). Rev. Biol. Mar. 1995, 30, 265–273. [Google Scholar]
- Dulcic, J. First record of ribbon fish, Trachipterus trachypterus, from the eastern Adriatic. Cybium 1996, 20, 101–102. [Google Scholar]
- Farias, I.; Moura, T.; Figueiredo, I.; Vieira, A.R.; Serra-Pereira, B.; Serrano Gordo, L. Northernmost occurrence of the ribbonfish Trachipterus trachypterus (Gmelin, 1789) in the NE Atlantic: The Portuguese continental shelf. J. Appl. Ichthyol. 2010, 26, 143–144. [Google Scholar] [CrossRef]
- Guerriero, G.; Di Finizio, A.; Ciarcia, G. Biological pollution: Molecular identification of non-native species in the Central Tyrrhenian Sea. Catrina 2010, 5, 41–47. [Google Scholar]
- Shinohara, G.; Shirai, S.M.; Nazarkin, M.V.; Yabe, M. Preliminary list of the deep-sea fishes of the Sea of Japan. Bull. Natl. Mus. Nat. Sci. Ser. A 2011, 37, 35–62. [Google Scholar]
- Mytilineou, C.; Anastasopoulou, A.; Christides, G.; Bekas, P.; Smith, C.J.; Papadopoulou, K.N.; Lefkaditou, E.; Kavadas, S. New records of rare deep-water fish species in the Eastern Ionian Sea (Mediterranean Sea). J. Nat. Hist. 2013, 47, 1645–1662. [Google Scholar] [CrossRef]
- Garibaldi, F. By-catch in the mesopelagic swordfish longline fishery in the Ligurian Sea (Western Mediterranean). Collect. Vol. Sci. Pap. ICCAT 2015, 71, 1495–1498. [Google Scholar]
- Yapici, S. New and additional records of rare fish species from the Anatolian coasts of Turkey. Mugla J. Sci. Technol. 2019, 5, 13–16. [Google Scholar] [CrossRef][Green Version]
- Macali, A.; Semenov, A.; Paladini de Mendoza, F.; Dinoi, A.; Bergami, E.; Tiralongo, F. Relative Influence of Environmental Factors on Biodiversity and Behavioural Traits of a Rare Mesopelagic Fish, Trachipterus trachypterus (Gmelin, 1789), in a Continental Shelf Front of the Mediterranean Sea. J. Mar. Sci. Eng. 2020, 8, 581. [Google Scholar] [CrossRef]
- Gökoğlu, M.; Özen, M.R. First Record of Trachipterus trachypterus (Gmelin, 1789) in the Gulf of Antalya (Turkey). Acta Aquat. Turc. 2021, 17, 505–507. [Google Scholar] [CrossRef]
- Mancini, P.L.; de Amorim, A.F.; Arfelli, C.A. Observações em Trachipterus jacksonensis capturados no Brasil. In Proceedings of the III Congresso Brasileiro de Pesquisas Ambientais e Saúde (CBPAS’ 2003), Santos, Brazil, 21–23 July 2003; pp. 56–58. [Google Scholar]
- Scott, E.O.G. Observations on some Tasmanian fishes: Part XXX. Pap. Proc. R. Soc. Tasm. 1984, 118, 187–222. [Google Scholar] [CrossRef]
- Macpherson, E.; Roel, B.A. Trophic relationships in the demersal fish community off Namibia. S. Afr. J. Mar. Sci. 1987, 5, 585–596. [Google Scholar] [CrossRef]
- Roberts, C.D. Fishes of the Chatham Islands, New Zealand: A trawl survey and summary of the ichthyofauna. N. Z. J. Mar. Freshw. Res. 1991, 25, 1–19. [Google Scholar] [CrossRef]
- Mincarone, M.M.; Lima, A.T.; Soto, J.M. Sobre a ocorrência do peixe-fita Trachipterus jacksonensis (Ramsay, 1881) (Lampridiformes, Trachipteridae) na costa brasileira. Mare Magnum 2001, 1, 121–124. [Google Scholar]
- Letourneur, Y.; Chabanet, P.; DurviLLe, P.; Taquet, M.; Teissier, E.; Parmentier, M.; QUÉRO, J.C.; Pothin, K. An updated checklist of the marine fish fauna of Reunion Island, south-western Indian Ocean. Cybium 2004, 28, 199–216. [Google Scholar]
- Smith, P.J.; Steinke, D.; McMillan, P.; McVeagh, S.M.; Struthers, C.D. DNA database for commercial marine fish. N. Z. Aquat. Environ. Biodivers. 2008, 22, 1–62. [Google Scholar]
- Fricke, R.; Mulochau, T.; Durville, P.; Chabanet, P.; Tessier, E.; Letourneur, Y. Annotated checklist of the fish species (Pisces) of La Réunion, including a red list of threatened and declining species. Stuttg. Beitr. Naturkunde A 2009, 2, 1–168. [Google Scholar]
- Satapoomin, U. The fishes of southwestern Thailand, the Andaman Sea-a review of research and a provisional checklist of species. Phuket Mar. Biol. Cent. Res. 2011, 70, 29–77. [Google Scholar]
- Melo, M.R.S.D.; Caires, R.A.; Sutton, T.T. The scientific explorations for deep-sea fishes in Brazil: The known knowns, the known unknowns, and the unknown unknowns. In Brazilian Deep-Sea Biodiversity; Springer: Cham, Switzerland, 2020; pp. 153–216. [Google Scholar]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).