Identification of Phenotypic Variation and Genetic Diversity in Rice (Oryza sativa L.) Mutants
Abstract
1. Introduction
2. Materials and Methods
2.1. Rice Materials
2.2. Rice Preparation and Field Experiments
2.3. Phenotypic Measurement
2.4. DNA Extraction
2.5. PCR Amplification
2.6. Data Analysis
3. Results
3.1. Grain Yield and Yield Components
3.2. Grain Size and Chemical Quality Traits
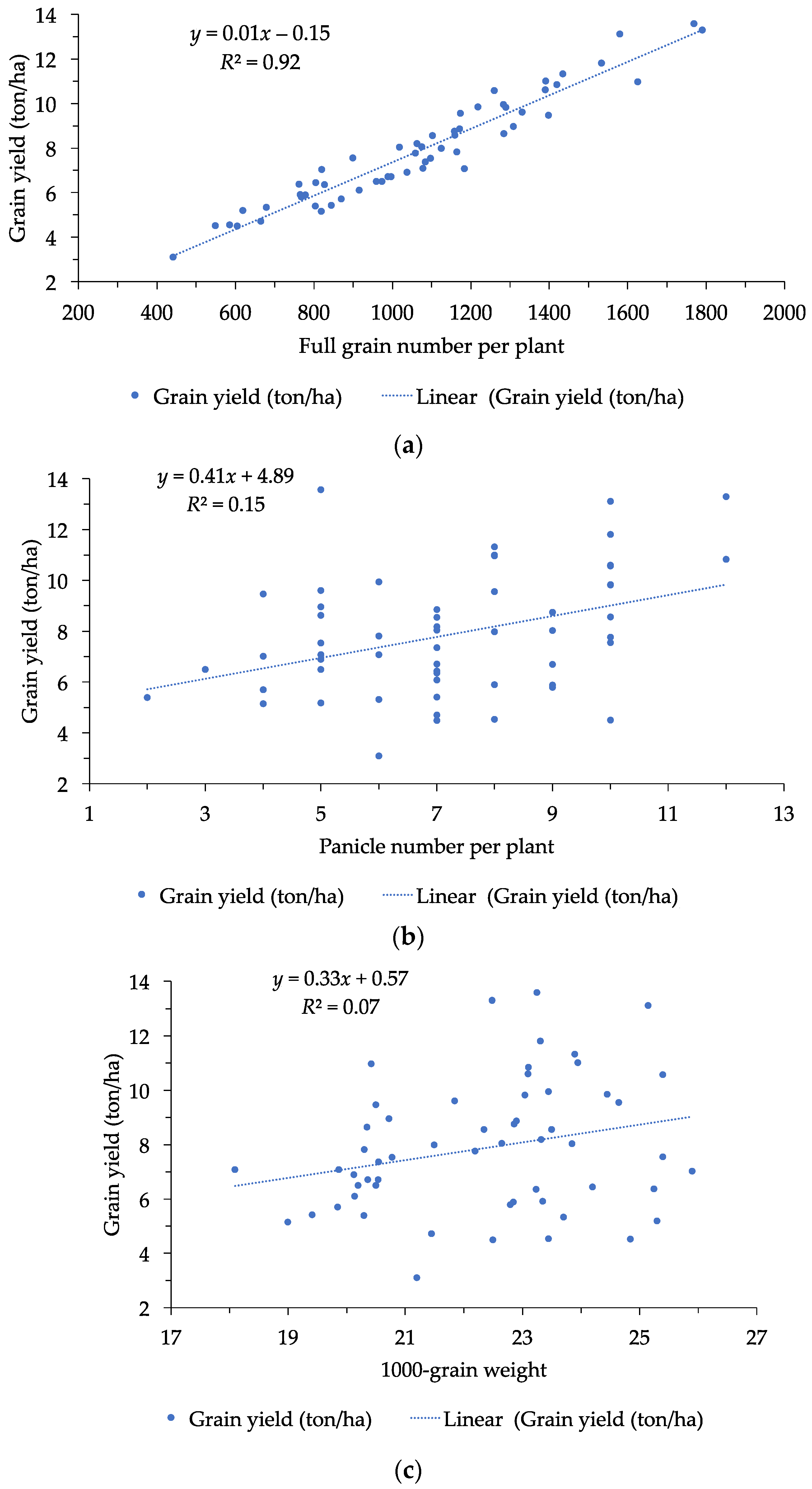
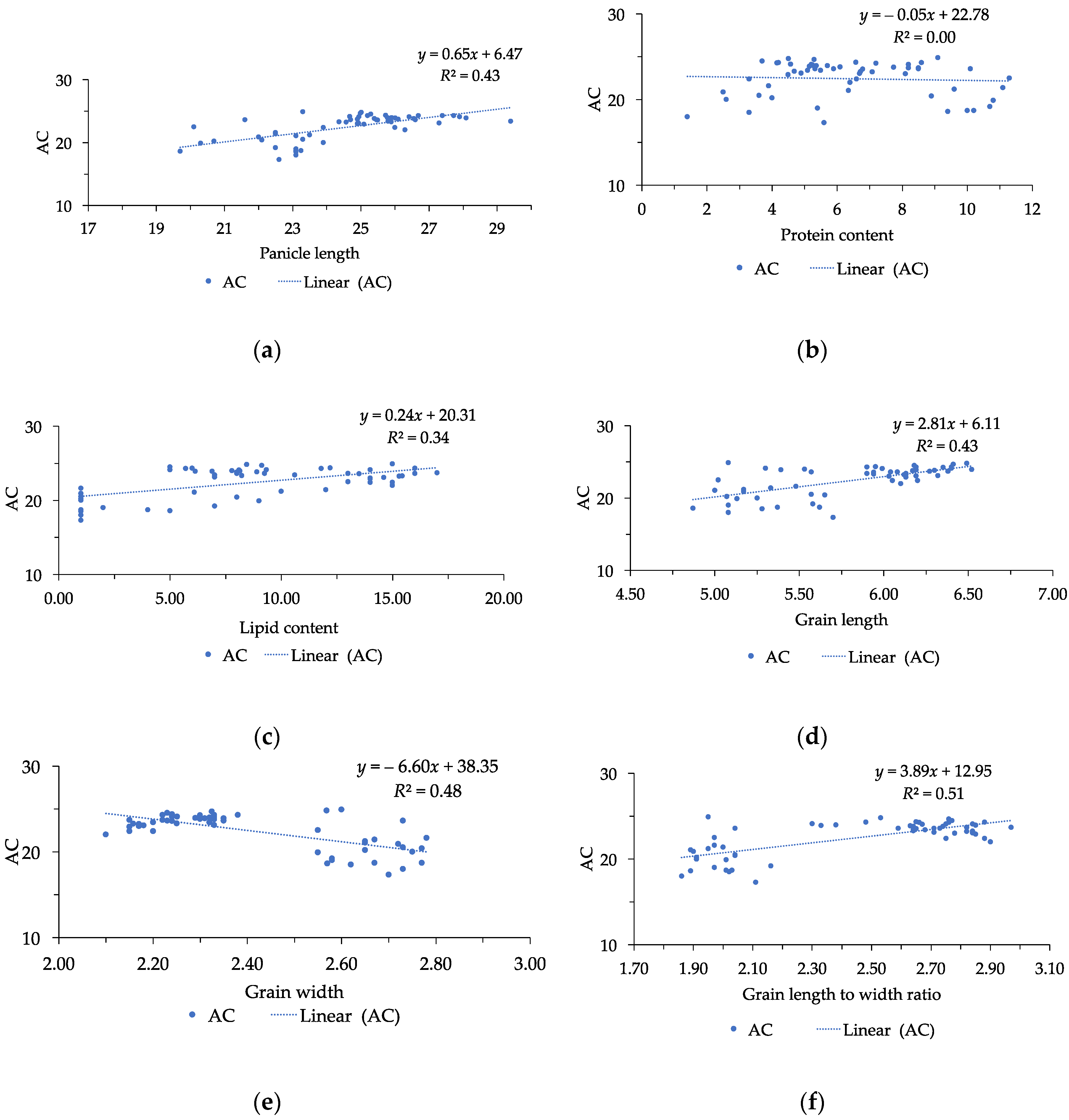
3.3. Correlation among Physico-Chemical Traits and Grain Yield
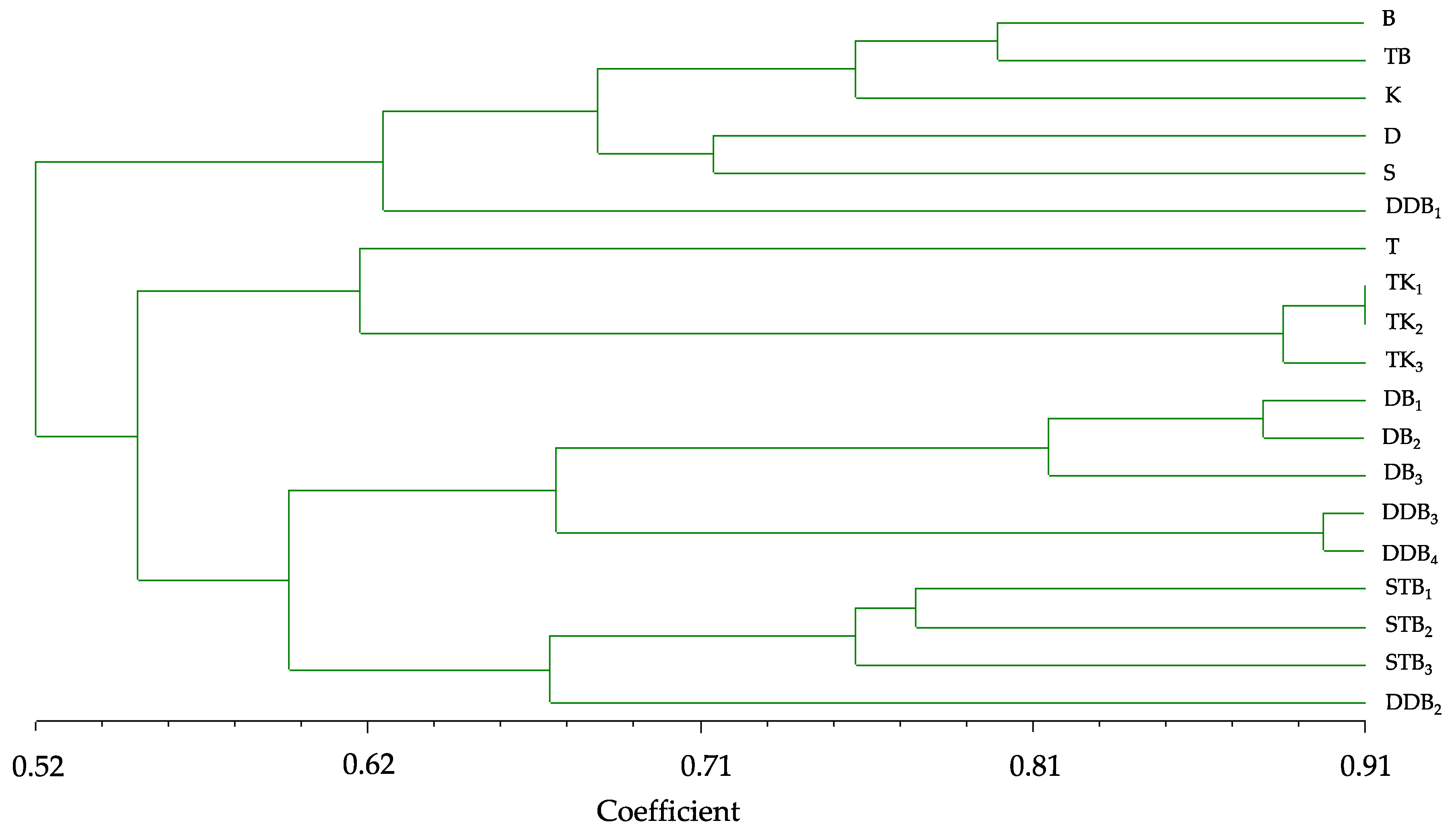
3.4. Genetic Diversity
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Bhat, F.M.; Riar, C.S. Effect of amylose, particle size & morphology on the functionality of starches of traditional rice cultivars. Int. J. Biol. Macromol. 2016, 92, 637–644. [Google Scholar] [PubMed]
- Jiang, Y.; Cai, Z.; Xie, W.; Long, T.; Yu, H.; Zhang, Q. Rice functional genomics research: Progress and implications for crop genetic improvement. Biotechnol. Adv. 2012, 30, 1059–1070. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Zhou, L.; Ponce, K.; Ye, G. The usefulness of known genes/qtls for grain quality traits in an Indica population of diverse breeding lines tested using association analysis. Rice 2015, 8. [Google Scholar] [CrossRef] [PubMed]
- Parry, M.A.J.; Madgwick, P.J.; Bayon, C.; Tearall, K.; Hernandez-Lopez, A.; Baudo, M.; Rakszege, M.; Hamada, W.; Al-Yassin, A.; Ouabbou, H.; et al. Mutation discovery for crop improvement. J. Exp. Bot. 2009, 60, 2817–2825. [Google Scholar] [CrossRef] [PubMed]
- Sharma, A.; Singh, S.K. Induced mutation—A tool for creation of genetic variability in rice (Oryza sativa L.). J. Crop Weed. 2013, 9, 132–138. [Google Scholar]
- Abacar, J.D.; Zhao-miao, L.; Xin-cheng, Z.; Cheng-qiang, D.; She, T.; Zheng-hui, L.; Shao-hua, W.; Yan-feng, D. Variation in yield and physicochemical quality traits among mutants of Japonica rice cultivar Wuyujing 3. Rice Sci. 2016, 23, 33–41. [Google Scholar] [CrossRef]
- Oladosu, Y.; Rafii, M.Y.; Abdullah, N.; Hussin, G.; Ramli, A.; Rahim, H.A.; Miah, G.; Usman, M. Principle and application of plant mutagenesis in crop improvement: A review. Biotechnol. Biotechnol. Equip. 2016, 30, 1–16. [Google Scholar] [CrossRef]
- Reig-Valiente, J.L.; Viruel, J.; Sales, E.; Marqués, L.; Terol, J.; Gut, M.; Derdak, S.; Talon, M.; Domingo, C. Genetic diversity and population structure of rice varieties cultivated in temperate regions. Rice 2016, 9. [Google Scholar] [CrossRef] [PubMed]
- Nachimuthu, V.V.; Muthurajan, R.; Duraialaguraja, S.; Sivakami, R.; Pandian, B.A.; Ponniah, G.; Gunasekaran, K.; Swaminathan, M.K.S.K.; Sabariappan, R. Analysis of population structure and genetic diversity in rice germplasm using SSR markers: An initiative towards association mapping of agronomic traits in Oryza sativa. Rice 2015, 8. [Google Scholar] [CrossRef] [PubMed]
- Haritha, G.; Sudhakar, T.; Chandra, D.; Ram, T.; Divya, B.; Sarla, N. Informative ISSR markers help identify genetically distinct accessions of Oryza rufipogon in yield improvement. Rice Sci. 2016, 23, 225–241. [Google Scholar] [CrossRef]
- Ram, S.G.; Thiruvengadam, V.; Vinod, K.K. Genetic diversity among cultivars, landraces and wild relatives of rice as revealed by microsatellite markers. J. Appl. Genet. 2007, 48, 337–345. [Google Scholar] [CrossRef] [PubMed]
- Edzesi, W.M.; Dang, X.; Liang, L.; Liu, E.; Zaid, I.U.; Hong, D. Genetic diversity and elite allele mining for grain traits in rice (Oryza sativa L.) by association mapping. Front. Plant. Sci. 2016, 7, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Collard, B.C.Y.; Mackill, D.J. Marker-assisted selection: an approach for precision plant breeding in the twenty-first century. Philos. Trans. R. Soc. B 2008, 363, 557–572. [Google Scholar] [CrossRef] [PubMed]
- Ni, J.; Colowit, P.M.; Mackill, D.J. Evaluation of genetic diversity in rice subspecies using microsatellite markers. Crop Sci. 2002, 42, 601–607. [Google Scholar] [CrossRef]
- Akagi, H.; Yokozeki, Y.; Inagaki, A.; Fujimura, T. Highly polymorphic microsatellites of rice consist of AT repeats, and a classification of closely related cultivars with these microsatellite loci. Theor. Appl. Genet. 1997, 94, 61–67. [Google Scholar] [CrossRef] [PubMed]
- Sega, G.A. A review of the genetic effects of ethyl methanesulfonate. Mutat. Res. 1984, 134, 113–142. [Google Scholar] [CrossRef]
- Vogel, E.W.; Natarajan, A.T. DNA damage and repair in somatic and germ cells in vivo. Mutat. Res. 1995, 330, 183–208. [Google Scholar] [CrossRef]
- Greene, E.A.; Codomo, C.A.; Taylor, N.E.; Henikoff, J.G.; Till, B.J.; Reynolds, S.H.; Enns, L.C.; Burtner, C.; Johnson, J.E.; Odden, A.R.; et al. Spectrum of chemically induced mutations from a large-scale reversegenetic screen in Arabidopsis. Genetics 2003, 164, 731–740. [Google Scholar] [PubMed]
- Till, B.J.; Cooper, J.; Tai, T.H.; Colowit, P.; Greene, E.A.; Henikoff, S.; Comai, L. Discovery of chemical induced mutations in rice by TILLING. BMC Plant Biol. 2007, 7, 19. [Google Scholar] [CrossRef] [PubMed]
- Satoh, H.; Shibahara, K.; Tokunaga, T.; Nishi, A.; Tasaki, M.; Hwang, S.K.; Okita, T.W.; Kaneko, N.; Fujita, N.; Yoshida, M.; et al. Mutation of the plastidial α-glucan phosphorylase gene in rice affects the synthesis and structure of starch in the endosperm. Plant Cell 2008, 20, 1833–1849. [Google Scholar] [CrossRef] [PubMed]
- Shah, S.M.; Arif, M.; Aslam, K.; Shabir, G.; Thomson, M.J. Genetic diversity analysis of Pakistan rice (Oryza sativa) germplasm using multiplexed single nucleotide polymorphism markers. Genet. Resour. Crop Evol. 2016, 63, 1113–1126. [Google Scholar] [CrossRef]
- Xuan, T.D.; Khanh, T.D.; Anh, T.T.T. Possibility of breeding super rice cultivars using gene linkage. In Proceedings of the 9th Asian Crop Science Association Conference (9th ACSAC), Jeju, Korea, 5–7 June 2017. [Google Scholar]
- Yi, M.; New, K.T.; Vanavichit, A.; Chai-arree, W.; Toojinda, T. Marker assisted backcross breeding to improve cooking quality traits in Myanmar rice cultivar Manawthukha. Field Crops Res. 2009, 113, 178–186. [Google Scholar] [CrossRef]
- Jin, L.; Lu, Y.; Xiao, P.; Sun, M.; Corke, H.; Bao, J. Genetic diversity and population structure of a diverse set of rice germplasm for association mapping. Theor. Appl. Genet. 2010, 121, 475–487. [Google Scholar] [CrossRef] [PubMed]
- Murray, M.G.; Thompson, W.F. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 1980, 8, 4321–4325. [Google Scholar] [CrossRef] [PubMed]
- Tello-Ruiz, M.K.; Stein, J.; Wei, S.; Preece, J.; Olson, A.; Naithani, S.; Amarasinghe, V.; Dharmawardhana, P.; Mulvaney, J.; Kumari, S.; et al. Gramene 2016: Comparative plant genomics and pathway resources. Nucleic Acids Res. 2016, 44, D1133–D1140. [Google Scholar] [CrossRef] [PubMed]
- Kumbhar, S.D.; Kulwal, P.L.; Patil, J.V.; Sarawate, C.D.; Gaikwad, A.P.; Jadhav, A.S. Genetic diversity and population structure in landraces and improved rice varieties from India. Rice Sci. 2015, 22, 99–107. [Google Scholar] [CrossRef]
- Nagy, S.; Poczai, P.; Cernák, I.; Gorji, A.M.; Hegedus, G.; Taller, J. PICcalc: An online program to calculate polymorphic information content for molecular genetic studies. Biochem. Genet. 2012, 50, 670–672. [Google Scholar] [CrossRef] [PubMed]
- Peakall, R.; Smouse, P.E. GENALEX 6: Genetic analysis in Excel. Population genetic software for teaching and research. Mol. Ecol. Notes 2006, 6, 288–295. [Google Scholar] [CrossRef]
- Koutroubas, S.D.; Mazzini, F.; Pons, B.; Ntanos, D.A. Grain quality variation and relationships with morpho-physiological traits in rice (Oryza sativa L.) genetic resources in Europe. Field Crops Res. 2004, 86, 115–130. [Google Scholar] [CrossRef]
- Huang, R.; Jiang, L.; Zheng, J.; Wang, T.; Wang, H.; Huang, Y.; Hong, Z. Genetic bases of rice grain shape: So many genes, so little known. Trends Plant Sci. 2013, 18, 218–226. [Google Scholar] [CrossRef] [PubMed]
- Tan, Y.F.; Xing, Y.Z.; Li, J.X.; Yu, S.B.; Xu, C.G.; Zhang, Q. Genetic bases of appearance quality of rice grains in Shanyou 63, an elite rice hybrid. Theor. Appl. Genet. 2000, 101, 823–829. [Google Scholar] [CrossRef]
- Xu, Z.; Chen, W.; Ma, D.; Lu, Y.; Zhou, S.; Liu, L. Correlations between rice grain shapes and main qualitative characteristics. Acta Agron. Sin. 2004, 30, 894–900. [Google Scholar]
- Lou, J.; Chen, L.; Yue, G.; Lou, Q.; Mei, H.; Xiong, L.; Luo, L. QTL mapping of grain quality traits in rice. J. Cereal Sci. 2009, 50, 145–151. [Google Scholar] [CrossRef]
- Salem, K.F.M.; Sallam, A. Analysis of population structure and genetic diversity of Egyptian and exotic rice (Oryza sativa L.) genotypes. C. R. Biol. 2016, 339, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Hashemi, F.S.G.; Rafii, M.Y.; Ismail, M.R.; Mohamed, M.T.M.; Rahim, H.A.; Latif, M.A.; Aslani, F. The genetic and molecular origin of natural variation for the fragrance trait in an elite Malaysian aromatic rice through quantitative trait loci mapping using SSR and gene-based markers. Gene 2015, 555, 101–107. [Google Scholar] [CrossRef] [PubMed]
- Lo, S.F.; Fan, M.J.; Hsing, Y.I.; Chen, L.J.; Chen, S.; Wen, I.C.; Liu, Y.L.; Chen, K.T.; Jiang, M.J.; Lin, M.K.; et al. Genetic resources offer efficient tools for rice functional genomics research. Plant Cell Environ. 2016, 39, 998–1013. [Google Scholar] [CrossRef] [PubMed]
- Anupam, A.; Imam, J.; Quatadah, S.M.; Siddaiah, A.; Das, S.P.; Variar, M.; Mandal, N.P. Genetic diversity analysis of rice germplasm in Tripura State of Northeast India using drought and blast linked markers. Rice Sci. 2017, 24, 10–20. [Google Scholar] [CrossRef]
- Hashimoto, Z.; Mori, N.; Kawamura, M.; Ishii, T.; Yoshida, S.; Ikegami, M.; Takumi, S.; Nakamura, C. Genetic diversity and phylogeny of Japanese sake-brewing rice as revealed by AFLP and nuclear and chloroplast SSR markers. Theor. Appl. Genet. 2004, 109, 1586–1596. [Google Scholar] [CrossRef] [PubMed]
- Mao-bai, L.; Hui, W.; Li-ming, C. Evaluation of population structure, genetic diversity and origin of Northeast Asia weedy rice based on simple sequence repeat markers. Rice Sci. 2015, 22, 180–188. [Google Scholar] [CrossRef]
| Sample Codes | Origins | Mutants | Descriptions |
|---|---|---|---|
| B | M2-Bao thai | + | Originated from Bao Thai cultivar with good quality, cold tolerance, cultivated in Northern Midland and Mountainous region in Vietnam |
| D | DT84 | − | A traditional sticky rice with good quality in the north of Vietnam |
| DB1 | M2-DT84/Bao thai | + | F2 mutant line |
| DB2 | M2-DT84/Bao thai | + | F2 mutant line |
| DB3 | M2-DT84/Bao thai | + | F2 mutant line |
| DDB1 | M2-DT84DB | + | F2 mutant line |
| DDB2 | M2-DT84DB | + | F2 mutant line |
| DDB3 | M2-DT84DB | + | F2 mutant line |
| DDB4 | M2-DT84DB | + | F2 mutant line |
| TK1 | M2-TBR1/Khang dan | + | F2 mutant line |
| TK2 | M2-TBR1/Khang dan | + | F2 mutant line |
| TK3 | M2-TBR1/Khang dan | + | F2 mutant line |
| T | TBR1 | − | Rice leaf blight tolerance, a high yield cultivar, extensively cultivated in Vietnam |
| K | Khang dan | + | High quality cultivar, broadly cultivated in the Northern and the central of Vietnam |
| S | SKLo | − | A high yield cultivar, widely cultivated in Vietnam |
| TB | BC15TB | − | Good quality, high yield, resistance to bacterial leaf blight and brown plant hopper, intensively cultivated in Vietnam |
| STB1 | M2-BC15TB/SKLo | + | F2 mutant line |
| STB2 | M2-BC15TB/SKLo | + | F2 mutant line |
| STB3 | M2-BC15TB/SKLo | + | F2 mutant line |
| Characteristics | Mean | Range | SD | CV (%) |
|---|---|---|---|---|
| Panicle number per plant | 7.23 | 2–12 | 2.24 | 3.22 |
| Panicle length (cm) | 24.58 | 19.70–29.40 | 2.08 | 11.80 |
| Grain number per plant | 1,730 | 815–3081 | 543 | 3.18 |
| 1000-grain weight (g) | 22.35 | 18.10–25.90 | 1.90 | 1.90 |
| Full grain number per plant | 1,067 | 442–1791 | 306 | 3.48 |
| Grain yield per hecta (ton) | 7.87 | 2.64–15.31 | 2.39 | 3.29 |
| Primary branching number per panicle | 13.46 | 10–18 | 1.74 | 7.76 |
| Secondary branching number per panicle | 56.25 | 27–87 | 14.96 | 3.76 |
| Grain length (mm) | 5.81 | 4.87–6.52 | 0.48 | 12.10 |
| Grain width (mm) | 2.41 | 2.10–2.78 | 0.21 | 11.23 |
| Grain length to width ratio (mm) | 2.44 | 1.86–2.97 | 0.38 | 6.49 |
| Amylose content (%) | 22.43 | 17.30–24.90 | 2.06 | 10.91 |
| Protein content (%) | 6.41 | 1.40–11.30 | 2.41 | 2.66 |
| Lipid content (%) | 8.78 | 1–17 | 4.97 | 1.77 |
| Traits | PNP | PL | GPP | KGW | FGP | GY | PB | SB | GL | GW | LWR | PC | AC |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| PL | −0.45 ** | 1 | |||||||||||
| GPP | 0.01 | 0.43 ** | 1 | ||||||||||
| KGW | 0.48 ** | −0.25 | −0.49 ** | 1 | |||||||||
| FGP | 0.26 ** | 0.29 * | 0.44 ** | 0.09 | 1 | ||||||||
| GY | 0.39 ** | 0.22 * | 0.29 * | 0.26 | 0.96 ** | 1 | |||||||
| PB | −0.62 ** | 0.67 ** | 0.59 ** | −0.58 ** | 0.17 | 0.00 | 1 | ||||||
| SB | −0.56 ** | 0.68 ** | 0.68 ** | −0.66 ** | 0.19 | 0.0 | 0.89 ** | 1 | |||||
| GL | −0.30* | 0.74 ** | 0.48 ** | −0.28 * | 0.45 ** | 0.36 ** | 0.60 ** | 0.59 ** | 1 | ||||
| GW | 0.65 ** | −0.67 ** | −0.45 ** | 0.62 ** | −0.70 | 0.10 | −0.78 ** | −0.81 ** | −0.72 ** | 1 | |||
| LWR | −0.53 ** | 0.74 ** | 0.49 ** | −0.48 ** | 0.27 * | 0.13 | 0.75 ** | 0.75 ** | 0.92 ** | −0.93 ** | 1 | ||
| PC | −0.03 | −0.34 * | −0.17 | −0.22 | −0.15 | −0.21 | −0.09 | −0.19 | −0.13 | 0.09 | −0.12 | 1 | |
| AC | −0.31 * | 0.66 ** | 0.37 ** | −0.35 ** | 0.19 | 0.86 | 0.49 ** | 0.53 ** | 0.66 ** | −0.69 ** | 0.71 ** | −0.06 | 1 |
| LC | −0.56 ** | 0.26 | 0.20 | −0.52 ** | −0.28 | −0.18 | 0.51 ** | 0.44 ** | 0.49 ** | −0.67 ** | 0.64 ** | 0.47 ** | 0.58 ** |
| No | Marker Name | Repeat Sequence | Chromosome | Allele | Total Alleles | Ho | H | PIC |
|---|---|---|---|---|---|---|---|---|
| 1 | RM190 | (CT)11 | 6 | 2 | 20 | 0.05 | 0.48 | 0.36 |
| 2 | RM510 | (GA)15 | 6 | 2 | 18 | 0.05 | 0.44 | 0.34 |
| 3 | RM508 | (AG)17 | 6 | 3 | 18 | 0.05 | 0.58 | 0.50 |
| 4 | RM540 | (AG)16 | 6 | 2 | 19 | 1.00 | 0.33 | 0.28 |
| 5 | RM170 | (CCT)7 | 6 | 2 | 26 | 0.37 | 0.50 | 0.38 |
| 6 | RM2 | (GA)13 | 7 | 5 | 42 | 0.21 | 0.76 | 0.73 |
| 7 | RM2010 | (AT)19 | 5 | 3 | 19 | 0.05 | 0.62 | 0.54 |
| 8 | RM587 | (CTT)18 | 6 | 2 | 23 | 0.26 | 0.49 | 0.37 |
| 9 | RM219 | (CT)17 | 9 | 6 | 41 | 1.00 | 0.79 | 0.76 |
| 10 | RM107 | (GA)7 | 9 | 2 | 16 | 0.16 | 0.47 | 0.36 |
| 11 | RM225 | (CT)18 | 6 | 3 | 19 | 0.16 | 0.59 | 0.50 |
| 12 | RM111 | (GA)9 | 6 | 2 | 32 | 0.89 | 0.50 | 0.37 |
| 13 | RM229 | (TC)11(CT)5C3(CT)5 | 11 | 2 | 18 | 0.05 | 0.44 | 0.33 |
| 14 | RM434 | (TC)12 | 9 | 4 | 31 | 1.00 | 0.64 | 0.58 |
| 15 | RM584 | (CT)14 | 6 | 4 | 31 | 1.00 | 0.73 | 0.68 |
| 16 | RM5688 | (AAT)17 | 9 | 3 | 16 | 0.47 | 0.62 | 0.54 |
| 17 | RM287 | (GA)21 | 11 | 2 | 20 | 0.05 | 0.48 | 0.36 |
| 18 | RR6119 | (CGC)8 | 6 | 4 | 24 | 1.00 | 0.40 | 0.38 |
| 19 | RM402 | (ATA)7 | 6 | 3 | 21 | 0.11 | 0.61 | 0.54 |
| 20 | RM55 | (GA)17 | 3 | 3 | 14 | 0.26 | 0.50 | 0.43 |
| 21 | RM14 | (GA)18 | 1 | 2 | 18 | 0.05 | 0.20 | 0.18 |
| 22 | RM18 | (GA)4AA(GA)(AG)16 | 7 | 4 | 29 | 0.05 | 0.71 | 0.66 |
| 23 | RM22 | (GA)22 | 3 | 2 | 18 | 0.16 | 0.40 | 0.32 |
| 24 | RM19 | (ATC)10 | 12 | 4 | 29 | 0.05 | 0.65 | 0.59 |
| 25 | RM222 | (GT)3(GTAT)8(GT)5 | 10 | 3 | 19 | 0.11 | 0.62 | 0.55 |
| 26 | RM239 | (AG)5TG(AG)2 | 10 | 4 | 42 | 1.00 | 0.74 | 0.69 |
| 27 | RM311 | (GT)3(GTAT)8(GT)5 | 10 | 2 | 25 | 0.95 | 0.50 | 0.37 |
| 28 | RM251 | (CT)29 | 3 | 2 | 18 | 0.05 | 0.40 | 0.32 |
| 29 | RM4 | (GA)16 | 1 | 4 | 49 | 1.00 | 0.71 | 0.67 |
| 30 | RM213 | (CT)17 | 2 | 4 | 29 | 1.00 | 0.62 | 0.58 |
| 31 | RM432 | (CATC)9 | 7 | 2 | 14 | 0.16 | 0.49 | 0.37 |
| 32 | RM38 | (GA)16 | 8 | 3 | 18 | 0.11 | 0.65 | 0.57 |
| 33 | RM52 | (AG)19 | 8 | 2 | 24 | 0.47 | 0.50 | 0.38 |
| 34 | RM218 | (TC)24ACT5(GT)11 | 3 | 3 | 20 | 1.00 | 0.62 | 0.53 |
| 35 | RM504 | (CA)9 | 3 | 5 | 59 | 0.05 | 0.78 | 0.74 |
| 36 | RM426 | (CA)10 | 3 | 7 | 50 | 1.00 | 0.81 | 0.79 |
| 37 | RM247 | (CT)16 | 12 | 2 | 20 | 0.16 | 0.32 | 0.27 |
| 38 | RM3392 | (CT)17 | 3 | 2 | 19 | 1.00 | 0.47 | 0.36 |
| 39 | RM569 | (CT)16 | 3 | 3 | 29 | 0.63 | 0.66 | 0.59 |
| 40 | RM234 | (CT)25 | 7 | 2 | 18 | 1.00 | 0.50 | 0.38 |
| 41 | RM284 | (GA)8 | 8 | 2 | 19 | 1.00 | 0.20 | 0.18 |
| 42 | RM338 | (CTT)6 | 3 | 5 | 52 | 1.00 | 0.77 | 0.74 |
| 43 | RM228 | (CA)6(GA)36 | 10 | 3 | 25 | 0.16 | 0.62 | 0.54 |
| 44 | RM431 | (AG)16 | 1 | 6 | 67 | 0.05 | 0.81 | 0.79 |
| 45 | RM333 | (TAT)19(CTT)19 | 10 | 6 | 58 | 1.00 | 0.81 | 0.78 |
| 46 | RM472 | (GA)21 | 1 | 4 | 29 | 0.05 | 0.71 | 0.66 |
| 47 | RM5626 | (AAG)11 | 3 | 3 | 35 | 0.26 | 0.66 | 0.58 |
| 48 | RM7 | (GA)19 | 3 | 2 | 17 | 0.05 | 0.41 | 0.33 |
| 49 | RM230 | (AGG)4(GA)9A(AG)13 | 8 | 2 | 24 | 0.37 | 0.50 | 0.38 |
| 50 | RM162 | (AC)20 | 6 | 2 | 21 | 0.74 | 0.47 | 0.36 |
| 51 | RM413 | (AG)11 | 5 | 2 | 19 | 1.00 | 0.47 | 0.36 |
| 52 | RM593 | (CT)15(CA)10 | 5 | 2 | 20 | 1.00 | 0.46 | 0.35 |
| 53 | RM247 | (CT)16 | 12 | 3 | 23 | 0.11 | 0.63 | 0.56 |
| 54 | RM60 | (AATT)5AATCT(AATT) | 3 | 2 | 19 | 1.00 | 0.10 | 0.09 |
| 55 | RM264 | (GA)27 | 8 | 2 | 19 | 1.00 | 0.47 | 0.36 |
| 56 | RM589 | (GT)24 | 6 | 2 | 18 | 1.00 | 0.28 | 0.24 |
| Name | B | D | DDB1 | TB | S | T | K | DB1 | DB2 | DB3 | TK1 | TK2 | TK3 | STB1 | STB2 | STB3 | DDB2 | DDB3 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| B | 1 | |||||||||||||||||
| D | 0.71 | 1 | ||||||||||||||||
| DDB1 | 0.60 | 0.67 | 1 | |||||||||||||||
| TB | 0.80 | 0.69 | 0.58 | 1 | ||||||||||||||
| S | 0.70 | 0.72 | 0.61 | 0.61 | 1 | |||||||||||||
| T | 0.54 | 0.62 | 0.56 | 0.63 | 0.69 | 1 | ||||||||||||
| K | 0.74 | 0.60 | 0.64 | 0.78 | 0.64 | 0.52 | 1 | |||||||||||
| DB1 | 0.48 | 0.50 | 0.49 | 0.50 | 0.51 | 0.50 | 0.48 | 1 | ||||||||||
| DB2 | 0.53 | 0.54 | 0.54 | 0.56 | 0.55 | 0.53 | 0.49 | 0.88 | 1 | |||||||||
| DB3 | 0.49 | 0.49 | 0.56 | 0.52 | 0.54 | 0.51 | 0.50 | 0.80 | 0.82 | 1 | ||||||||
| TK1 | 0.53 | 0.54 | 0.54 | 0.51 | 0.56 | 0.63 | 0.46 | 0.55 | 0.57 | 0.63 | 1 | |||||||
| TK2 | 0.50 | 0.50 | 0.41 | 0.48 | 0.49 | 0.58 | 0.41 | 0.53 | 0.54 | 0.60 | 0.91 | 1 | ||||||
| TK3 | 0.52 | 0.53 | 0.46 | 0.49 | 0.50 | 0.63 | 0.43 | 0.53 | 0.51 | 0.55 | 0.88 | 0.89 | 1 | |||||
| STB1 | 0.58 | 0.51 | 0.50 | 0.52 | 0.49 | 0.49 | 0.54 | 0.57 | 0.57 | 0.61 | 0.64 | 0.63 | 0.66 | 1 | ||||
| STB2 | 0.52 | 0.55 | 0.53 | 0.51 | 0.51 | 0.53 | 0.45 | 0.59 | 0.63 | 0.64 | 0.58 | 0.58 | 0.61 | 0.78 | 1 | |||
| STB3 | 0.56 | 0.48 | 0.47 | 0.54 | 0.51 | 0.47 | 0.49 | 0.56 | 0.57 | 0.51 | 0.59 | 0.58 | 0.59 | 0.76 | 0.75 | 1 | ||
| DDB2 | 0.54 | 0.51 | 0.51 | 0.51 | 0.50 | 0.48 | 0.49 | 0.58 | 0.57 | 0.57 | 0.54 | 0.58 | 0.60 | 0.63 | 0.70 | 0.67 | 1 | |
| DDB3 | 0.54 | 0.57 | 0.54 | 0.55 | 0.52 | 0.52 | 0.56 | 0.63 | 0.70 | 0.65 | 0.47 | 0.50 | 0.51 | 0.58 | 0.67 | 0.56 | 0.63 | 1 |
| DDB4 | 0.59 | 0.54 | 0.53 | 0.55 | 0.53 | 0.53 | 0.55 | 0.64 | 0.73 | 0.66 | 0.50 | 0.50 | 0.52 | 0.59 | 0.69 | 0.56 | 0.59 | 0.89 |
| Source | df | SS | MS | Est. Var. | % | p Value |
|---|---|---|---|---|---|---|
| Among groups | 3 | 260.66 | 86.89 | 13.04 | 34 | 0.001 |
| Within groups | 15 | 383.55 | 25.57 | 25.57 | 66 | 0.001 |
| Total | 18 | 644.21 | 38.61 | 100 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tu Anh, T.T.; Khanh, T.D.; Dat, T.D.; Xuan, T.D. Identification of Phenotypic Variation and Genetic Diversity in Rice (Oryza sativa L.) Mutants. Agriculture 2018, 8, 30. https://doi.org/10.3390/agriculture8020030
Tu Anh TT, Khanh TD, Dat TD, Xuan TD. Identification of Phenotypic Variation and Genetic Diversity in Rice (Oryza sativa L.) Mutants. Agriculture. 2018; 8(2):30. https://doi.org/10.3390/agriculture8020030
Chicago/Turabian StyleTu Anh, Truong Thi, Tran Dang Khanh, Tran Dang Dat, and Tran Dang Xuan. 2018. "Identification of Phenotypic Variation and Genetic Diversity in Rice (Oryza sativa L.) Mutants" Agriculture 8, no. 2: 30. https://doi.org/10.3390/agriculture8020030
APA StyleTu Anh, T. T., Khanh, T. D., Dat, T. D., & Xuan, T. D. (2018). Identification of Phenotypic Variation and Genetic Diversity in Rice (Oryza sativa L.) Mutants. Agriculture, 8(2), 30. https://doi.org/10.3390/agriculture8020030







