1. Introduction
Neonatal calf diarrhea (NCD) is a global health concern for dairy calves within their first month of life. Clinical signs of NCD include watery feces, dehydration, and behavioral signs such as depression and decreased suckling behavior [
1]. NCD can lead to significant morbidity and mortality in preweaned calves which is precipitated by a cascade of complications including dehydration, acidosis, and electrolyte imbalance [
2,
3]. Estimates suggest that NCD accounts for 21.3% of all morbidity in preweaned calves and is the leading cause of calf mortality, with 53 to 57% of calf deaths attributed to diarrhea [
4,
5]. The intensity of NCD symptoms can differ considerably, with cases classified from mild to severe. Mild cases often resolve with supportive care such as electrolyte supplementation and increased monitoring. Moderate cases may require additional interventions to reverse electrolyte imbalance and dehydration, including multiple oral fluid therapy treatments. Severe cases, which constitute approximately one-third of all NCD incidents, often require more intensive treatments, such as intravenous fluid therapy and, in some cases, antibiotics [
6]. However, if left untreated or in severe instances, NCD can lead to rapid physiological decline due to escalating dehydration, metabolic disturbances, and immune suppression, ultimately increasing the risk of mortality. Therefore, early detection and the mitigation of NCD through enhanced husbandry protocols should be regarded as a primary objective within any calf rearing program.
A clinical health scoring (CHS) schema and blood gas analysis are conventional methods for identifying NCD and assessing disease severity [
1,
7,
8]. A CHS schema involves a systematic evaluation of clinical signs to ascertain the overall health status of a calf, such as fecal consistency, hydration status, behavior changes, and appetite [
9,
10]. Despite detailed scoring definitions, CHS relies heavily on subjective interpretation. This subjectivity introduces variability in the assessment process, leading to inconsistencies in NCD diagnosis and evaluation of disease severity. Moreover, the accuracy of CHS results may be influenced by the observer’s experience and expertise, further complicating its reliability as a diagnostic tool. Blood gas analysis is a diagnostic method used to assess various blood parameters, such as pH, oxygen and carbon dioxide levels, bicarbonate concentration, and oxygen saturation [
3,
11]. This analysis provides valuable information about acid–base balance and electrolyte status, which are essential for understanding the metabolic disturbances associated with NCD and guiding appropriate treatment strategies. Blood gas analysis, while considered the gold standard for evaluating metabolic disturbances in diarrheic calves, has practical limitations. This method requires blood sampling procedures, which may not be well tolerated by neonatal calves and can lead to additional stress. Additionally, blood gas analysis requires specialized equipment and trained personnel, making it resource-intensive, expensive, and less feasible for routine use in on-farm settings.
Emerging technologies in dairy farming, such as Automatic Calf Feeders (ACFs), offer promising opportunities for NCD detection and management strategies in calves during the preweaning phase [
12,
13,
14]. Calves at risk of NCD can be identified by analyzing ACF-collected feeding behavior data, including milk intake, drinking speed, and the frequency of rewarded and non-rewarded access [
15,
16]. However, ACF settings may introduce variability and complexity in the assessment of NCD risk among calves through differences in milk feeding strategies, daily allocation, housing density, and dietary intake [
16]. While these methods can provide early health warnings for calves, clinical examination of the individual calf is still required.
Suckling behavior can be used as an indicator of vitality in newborn calves and has been associated with evaluating the severity of metabolic acidosis and dehydration in cases of NCD [
7,
17,
18]. The suckling reflex is a complex neuromuscular response mediated by the brainstem and requires coordinated input from the central nervous system, cranial nerves, and orofacial muscles. As such, it is highly sensitive to systemic disturbances such as acidosis, dehydration, and energy imbalance, which are commonly observed during NCD episodes. Impairments in suckling performance may thus reflect underlying physiological stress, reduced neuromuscular function, or diminished responsiveness—making them a valuable proxy for overall health status.
Suckle strength has been employed as a qualitative measure to gauge the severity of the condition and guide treatment strategies [
3]. However, the current assessment of suckle strength involves a subjective evaluation of the calf’s response to finger placement in its mouth, since specialized sensors are not yet available. Although pressure manometers have been used to quantify suckle strength, concerns have been raised regarding their accuracy and reliability in assessing suckling behavior in calves [
19].
Recent progress in imaging modalities and machine learning algorithms has significantly expanded the potential for disease surveillance and diagnosis in veterinary contexts [
20,
21,
22]. This study endeavors to explore whether suckle pressure patterns can serve as a quantitative, objective biomarker for the early detection and prediction of NCD. We hypothesize that deviations from normal suckling behavior—quantified through pressure distributions and patterns—may precede the onset of clinical signs of NCD and thus offer predictive value. This hypothesis is supported by the physiological interplay between feeding behavior, metabolic stress, and enteric health. During the preweaning period, suckling is tightly regulated by neurological and metabolic functions, and early gastrointestinal disturbances may lead to discomfort, reduced appetite, or impaired reflexes, which can manifest as measurable changes in suckle intensity and rhythm. By capturing these changes, we seek to characterize both healthy and abnormal suckling behavior and evaluate their relationship to NCD onset.
In this observational cohort study, our objective is to assess the effectiveness of suckle pressure measurement in conjunction with machine learning techniques for NCD identification. Key image-derived features—such as pixel density, color saturation, entropy, and histogram-based characteristics—are extracted from pressure impression images to quantify variations in pressure distribution and intensity. These features are then utilized by machine learning classifiers to distinguish between healthy calves and those afflicted with NCD. This work will serve as a foundation for informing the design of animal health sensors, which are capable of real-time monitoring of subtle changes in suckle pressure and pattern.
2. Materials and Methods
2.1. Animals and Experimental Design
This study was conducted at a single commercial dairy farm in central New York state from 26 September 2023 to 5 November 2023. All animal-related procedures were conducted in accordance with the guidelines approved by the Cornell University Institutional Animal Care and Use Committee (protocol number 2023-0146).
In this study, a convenience sample of 51 female Holstein calves was enrolled at birth. All female Holstein calves born between 26 September 2023 and 16 October 2023 were eligible for inclusion. We focused on the first 21 days of life, as farm records and previous observations indicate that this is the period of highest NCD incidence, likely due to immature immune function and increased pathogen exposure during the early neonatal phase. The calves were health scored daily, beginning at day 1 until day 21 of life using the University of Wisconsin Calf Health Scorer [
23]. Additionally, at 1, 3, 5, 7, 10, 14, and 21 days of life suckle pressure was collected. If calves were diagnosed with NCD upon daily health scoring, then suckle pressure was collected daily until fecal consistency returned to normal.
Calves were initially housed in a maternity area bedded with deep straw and received 4 L of colostrum via an esophageal tube within 1 h of birth. They were relocated to the calf barn three times daily, and housed in groups of five, each within separate wire enclosures (4 m × 2.5 m) lined with deep straw bedding until they were between 5 and 8 days old. During this period, they were bottle-fed 2.8 L of milk replacer (21% crude protein, 19% fat) twice daily. At 5 to 8 days of age, calves were moved to larger group pens (9.8 m × 8.9 m) that housed up to 20 calves of similar age, with ad libitum access to electrolytes (provided in a cooler), and pelleted calf starter. They were trained to an automated milk feeder (Urban Alma Pro, Wüsting, Germany), which provided up to 11 L of whole milk per day.
Calves were monitored daily for NCD during the study period by trained study personnel. To diagnose NCD, a fecal score ranging from 0 to 3 was used where a fecal score of 0 denoted normal feces and a score of 3 denoted loose watery stool that sifted through the bedding. Calves with a fecal score of 3 were diagnosed with NCD. Those diagnosed with NCD received oral meloxicam (45 mg/100 lbs) and 2 quarts of oral electrolytes via bottle three times per day until resolution. In cases of drinking refusal, electrolyte solutions were delivered using an esophageal tube feeder. Calves unable to stand due to severe NCD were given 2 L of intravenous fluids (each 1 L of sterile water mixed with 17 g of bicarbonate and 25 mL of 50% dextrose).
Figure 1 provides a visual summary of the experimental framework employed in this study.
2.2. Data Collection
Suckle pressure data were collected during the same 2 h time window from 0700 to 0900. This period was selected because it coincided with the morning feeding routine on the farm, when calves were most likely to exhibit natural and consistent suckling activity. FujiFilm Prescale impression film (Sensor Products Inc., Madison, NJ, USA), measuring 8.5 cm × 6 cm, was wrapped around the nipple of an empty calf feeding bottle to collect the data. To prevent disease transmission and safeguard the film from moisture, a clean, new disposable glove and rubber band were secured over the film for each sample. The nipple and bottle assembly were then offered to the calf, who was permitted to suckle for a duration of 15 s after latching. After sampling, the film was detached and stored in a labeled envelope until digital transfer for analysis.
2.3. Dataset Preparation
Images of the suckle pressure films were obtained using a Canon Rebel T5 Digital Single-Lens Reflex (DSLR) camera (Canon Inc., Tokyo, Japan). The images were saved in JPEG format, with resolutions ranging up to 2300 × 1800 pixels due to slight variations caused by manual adjustments during image capture. A total of 349 images were collected, of which 54 were from calves with NCD and 295 from healthy calves. NCD was observed in 29/51 calves during the trial. The fecal consistency scoring was performed by 1 of 3 trained observers using the University of Wisconsin Calf Health Scorer standards (score 0: normal, score 1: pasty, score 2: loose, score 3: watery). Calves with a fecal consistency score of 3 were labeled as having NCD for the purposes of this study. A fecal consistency score of 0 to 2 was used to identify calves without clinical signs of NCD (classified as NONCD).
2.4. Feature Extraction
Suckle pressure impressions become visible on the film as red coloration, with the intensity of the color corresponding to the magnitude of the pressure applied during suckling. To capture and characterize variations in suckle pressure within the images, color, texture, and intensity distribution features were extracted.
Density refers to the concentration and distribution of suckle pressure within the image. It indicates how tight or dispersed suckle pressure measurements are observed. Density is calculated by converting the images to grayscale, applying binary thresholding, and determining the ratio of black pixels to the total number of pixels (Equation (1)).
Saturation represents the intensity and vividness of a hue. Measurement of image saturation involves converting a color image from the red, green, and blue (RGB) color space to the Hue, Saturation, and Value (HSV) color space. Saturation values are then extracted from each pixel within the image (Equation (2)), followed by computing their average (Equation (3)). R, G, and B are the red, green, and blue pixel values (normalized to the range [0, 1]). N indicates the number of pixels within, and
corresponds to the saturation intensity of the i-th pixel.
Entropy functions as an indicator of the variability and intricacy present in the distribution of suckle pressure, mirroring the textural complexity observed within the images. The image is first converted to grayscale to simplify intensity analysis and focus on the luminance component. A histogram of pixel intensities is then generated, representing the frequency of each intensity level. The histogram values are normalized to obtain the probability distribution of pixel intensities, and entropy is calculated based on Shannon’s entropy equation (Equation (4)), which captures the uncertainty in the distribution.
is the normalized probability of the i-th intensity value, and L is the total number of intensity levels.
Histograms provide a visual depiction of the distribution of pixel intensity values within a grayscale image, showcasing the occurrence frequency of individual intensity levels. In this study, intensity levels ranged from 0 (black) to 255 (white), and the histogram recorded the frequency of each level. The mean and standard deviation of histogram values serve as metrics for assessing the overall intensity distribution and variability within the image. Specifically, the mean represents the average brightness, while the standard deviation indicates the degree of variation in intensity. These statistics help capture texture and illumination differences that may be related to changes in suckling behavior.
2.5. Machine Learning Methods
Machine learning has become an essential tool for disease detection in livestock, enabling early intervention by automating classification based on physiological and behavioral data. In this study, machine learning classifiers were applied to distinguish between calves diagnosed with NCD and NONCD calves using features extracted from suckle pressure images.
Suckle pressure images capture subtle variations in feeding behavior, which can serve as early indicators of NCD. Key features extracted from these images including pixel density, color saturation, entropy, and histogram-based attributes were used as inputs for machine learning classification. However, two key challenges arose in this classification task: data imbalance and nonlinear feature relationships. The dataset was inherently imbalanced, as the number of NONCD calves significantly exceeded those diagnosed with NCD, potentially biasing model training and reducing sensitivity to positive cases. Additionally, the extracted features exhibited nonlinear patterns, requiring models capable of capturing complex relationships to effectively differentiate between NONCD and NCD.
To address these challenges, five machine learning models were selected based on their effectiveness in classification tasks, ability to capture complex relationships in image-derived data, and adaptability to imbalanced datasets: K-Nearest Neighbors (KNN) [
24], Random Forest (RF) [
25], Support Vector Machine (SVM) [
26], Gradient Boosting (GB) [
27], and Easy Ensemble (EE) [
28].
KNN was selected as a baseline model for its simplicity and effectiveness in handling small datasets. KNN classifies new samples based on their similarity to training data points using distance metrics. However, KNN is susceptible to performance degradation in high-dimensional spaces due to the curse of dimensionality and struggles with class imbalance, where majority-class samples dominate the nearest neighbors.
SVM offers a robust approach for defining decision boundaries in complex feature spaces. By employing kernel functions, it maps input data into higher-dimensional representations, facilitating the classification of nonlinearly separable data. While well suited to smaller datasets, SVM can still be sensitive to noise in the feature space and may require careful hyperparameter tuning to avoid overfitting.
RF, GB, and EE were selected as ensemble learning methods to enhance classification performance and model robustness. These approaches integrate several weak learners to achieve improved predictive performance and broader generalization capability. RF constructs multiple decision trees trained on randomly sampled subsets of the data and aggregates their predictions to reduce variance and improve robustness. It is particularly effective for handling high-dimensional datasets with mixed continuous and categorical features. However, RF may struggle to detect minority-class instances in imbalanced datasets, as the majority class tends to dominate the voting process.
GB improves classification performance through sequential training, where each decision tree corrects the errors of its predecessor. It belongs to the family of boosting algorithms that build an additive model in a forward stage-wise manner, using decision trees as weak learners to minimize a specified loss function through gradient descent. This iterative learning process enhances accuracy but requires careful hyperparameter tuning to prevent overfitting. While GB often outperforms individual decision trees, it can be computationally demanding and sensitive to noisy data.
EE was specifically chosen to address class imbalance, as NCD cases were underrepresented compared to NONCD cases. EE is an ensemble method designed for imbalanced classification, which combines multiple AdaBoost classifiers trained on different balanced subsets obtained by random under-sampling of the majority class. By focusing on the minority class during training, EE improves sensitivity in detecting NCD cases, making it particularly suitable for datasets with a low prevalence of positive cases.
2.6. Evaluation Metrics
Model evaluation was conducted based on five metrics: precision, recall, accuracy, F1 score, and receiver operating characteristic (ROC) curve [
29,
30]. NCD is the class of interest and is considered the positive class, meaning calves with NCD are labeled as positive, while healthy calves are labeled as negative. Precision represents the proportion of NCD calves correctly identified among all calves predicted to have NCD (Equation (5)). Recall denotes the proportion of true NCD cases that are successfully detected among all calves with NCD (Equation (6)). Accuracy refers to the ratio of correctly labeled calves (NCD or NONCD) among the total population of calves (Equation (7)). F1 score combines precision and recall into a single measure by computing their harmonic mean, offering a balanced evaluation (Equation (8)). The ROC curve visualizes the trade-off between the true positive rate (TPR) (Equation (9)) and the false positive rate (FPR) (Equation (10)) across different thresholds [
31]. The Area Under the ROC Curve (AUC) summarizes the ROC curve, indicating the overall discriminatory power of the model. Higher AUC values suggest better model performance, with an AUC of 1 representing perfect classification and an AUC of 0.5 suggesting no discrimination beyond random class assignment. Ideally, the ROC curve should be closer to the upper left corner, indicating high TPR while keeping FPR low, thereby effectively distinguishing between positive and negative samples.
True positives (TPs) refer to calves accurately identified as having NCD, whereas true negatives (TNs) correspond to calves correctly identified as NONCD. False positives (FPs) indicate instances incorrectly classified as having NCD when they are, in fact, NONCD, whereas false negatives (FNs) signify instances incorrectly classified as NONCD when they have NCD.
2.7. Model Implementation
The experiments were performed on a desktop computer with an Intel Core i9-10980XE CPU @ 3.00 GHz and 128 GB RAM, operating on Windows 11 Enterprise. Python version 3.8 served as the main programming language for executing all experimental procedures. The libraries employed included OpenCV (version 3.8) for image processing [
32], Numpy (version 1.26.2) for numerical computations [
33], Scikit-image (version 0.22.0) for image analysis [
34], and Scipy (version 1.11.4) for scientific computing [
35]. The implementations of KNN, RF, SVM, and GB in this work were conducted utilizing the scikit-learn library (version 1.3.2) [
36], whereas EE was implemented using the imbalance-learn library (version 0.12.0) [
37].
The training process for all machine learning models utilized a grid search technique combined with 5-fold cross-validation to determine the optimal hyperparameters. Specifically, the training data were randomly split into five groups, each containing an equal or nearly equal number of instances. The model was then trained and validated five times, with each iteration using a different subset as the validation set and the remaining subsets as the training set. Grid search was performed independently for each model, with a predefined range of hyperparameter values. The optimal set of hyperparameters was selected based on average validation performance across the five folds. These selected hyperparameters were then applied to train the final model on the entire training dataset before evaluation on the holdout testing set. This systematic approach evaluated various hyperparameter combinations to ensure the best model performance.
The machine learning models used were configured to optimize classification performance while addressing the challenges posed by class imbalance and feature complexity. Each model’s hyperparameters were fine-tuned to enhance predictive accuracy and generalization. For KNN, the model was configured with three nearest neighbors (K = 3) for classification. The algorithm automatically selected the most efficient computation method (‘ball_tree’, ‘kd_tree’, or ‘brute’) based on the structure of the training data. The Minkowski distance metric was used, with a parameter of 2, corresponding to Euclidean distance. A distance-weighted approach was applied to give greater influence on closer neighbors, enhancing classification accuracy in feature-rich datasets.
SVM applied a linear kernel for decision boundary construction, with a regularization parameter (C) set to 1.0. The gamma parameter was automatically determined based on the dataset’s characteristics. Additionally, class weights were adjusted inversely to class frequencies to counteract data imbalance for fair representation of NCD cases.
RF was optimized with 100 trees and no predefined limit on the maximum tree depth. The selection of features at each decision node was based on the square root of the overall number of available features. Moreover, the model was configured to require a minimum of one sample per leaf node and at least two samples to split a node. Class distribution discrepancies were mitigated through the use of a balanced weighting approach.
GB was trained using 200 boosting stages with a learning rate of 0.01 to ensure gradual performance improvement while mitigating overfitting. Each tree was constrained to a maximum depth of 3 to maintain generalizability. The selection of features at each node was guided by the square root of the total number of available features. Additionally, a minimum of one sample per leaf node and at least ten samples per split were required, with the entire dataset used for fitting each base learner. Unlike RF, GB sequentially refined its predictions by focusing on misclassified instances at each iteration.
The optimal configuration for the EE used in this study comprised 20 AdaBoost learners in the ensemble and a sampling strategy set to 0.8. This sampling strategy denotes the anticipated ratio of minority-class samples to majority-class samples post-resampling.
3. Results
3.1. Descriptive Statistics
In this study, 51 calves were enrolled, with 29 diagnosed with NCD and 22 not observed to exhibit signs of NCD throughout the trial. One calf died of an intestinal torsion at 10 days of age as determined on post-mortem examination by the herd veterinarian. This calf did not have any recorded health events prior to the intestinal accident and suckle pressure images before the incident were retained for analysis. Among the calves with NCD, 15 (52%) had the condition for a single day, while 6 (21%) had NCD for two days, and another 6 (21%) for three days. Additionally, one calf (3%) had NCD for four days, and another (3%) for five days. On average, NCD lasted for 2 days (range: 1 to 5 days). The first occurrence of NCD in calves predominantly occurred within the first two weeks (28 out of 29 cases), with an average onset at 8 days of age. The earliest observation of NCD was at 1 day of age and the latest observation was at 15 days of age.
3.2. Evaluation of Machine Learning Classifiers for NCD Detection
The central goal of this research was to investigate how effectively machine learning techniques could detect NCD from suckle pressure impression images. This work compared five ML classifiers (RF, GB, KNN, SVM, and EE) based on extracted features, including density, saturation, entropy, and histograms. Suckle pressure data for both NONCD and NCD were randomly partitioned into training and testing sets in an 8:2 ratio.
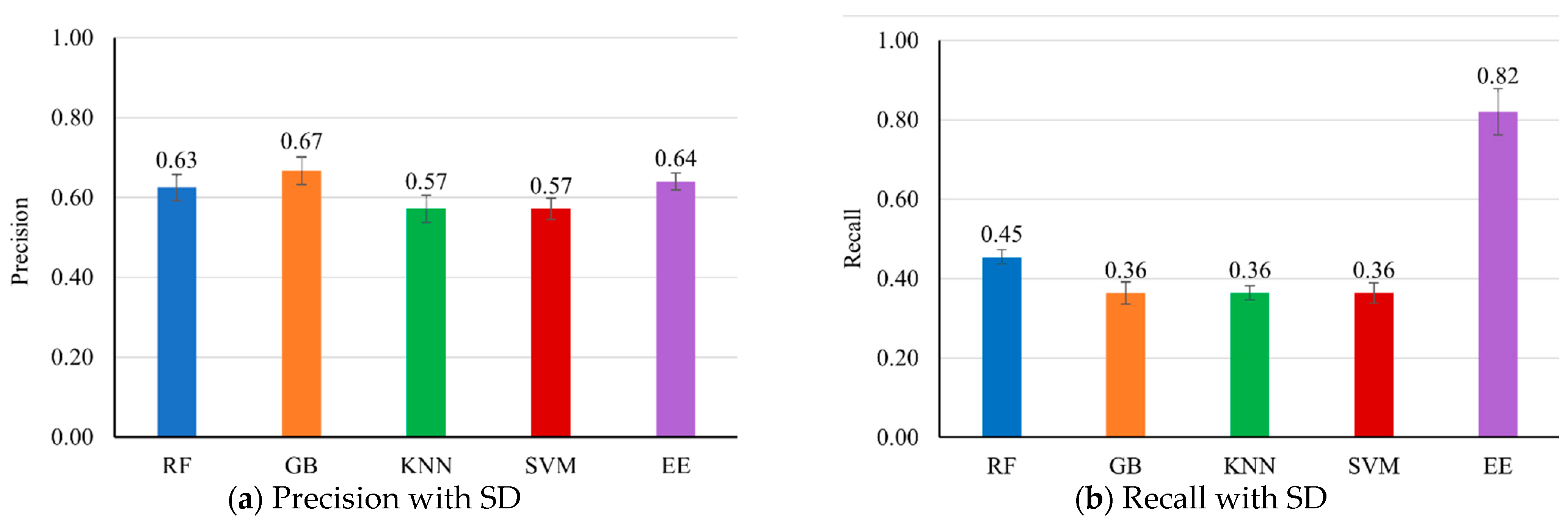
Figure 2 presents the evaluation results of multiple machine learning models on the testing dataset.
The EE classifier demonstrated superior performance in recall, F1 score, and accuracy. Specifically, EE achieved the highest recall (0.82 ± 0.06), indicating strong capability in correctly identifying positive cases of NCD. This high recall is crucial for disease detection tasks, where the primary concern is to minimize false negatives. The EE classifier also had the highest F1 score (0.72 ± 0.03) and accuracy (0.90 ± 0.03), suggesting a balanced performance in terms of precision and recall, and a high overall correctness in predictions.
While the EE classifier’s precision (0.64 ± 0.02) was slightly lower than that of the GB classifier (0.67 ± 0.03), this difference is relatively minor. The higher recall of the EE classifier compensates for its slightly lower precision, leading to a better overall F1 score and accuracy. This indicates that the EE classifier is more reliable in scenarios where the cost of missing a positive case (false negative) is higher than the cost of a false positive. Despite having the highest precision, the GB classifier had a lower recall (0.36 ± 0.03) and F1 score (0.47 ± 0.01), resulting in a slightly lower accuracy (0.87 ± 0.02). This suggests that GB is more conservative in predicting positive cases, leading to fewer false positives but more false negatives.
The RF classifier exhibited balanced performance, achieving a precision of 0.63 ± 0.03, recall of 0.45 ± 0.02, F1 score of 0.53 ± 0.01, and accuracy of 0.87 ± 0.02. It stands out as a strong performer, although it slightly lags behind EE. At the same time, both the KNN and SVM classifiers demonstrated similar moderate performance, with precision, recall, and F1 score at 0.57 ± 0.03, 0.36 ± 0.02, and 0.44 ± 0.02, respectively, along with an accuracy of 0.86 ± 0.03.
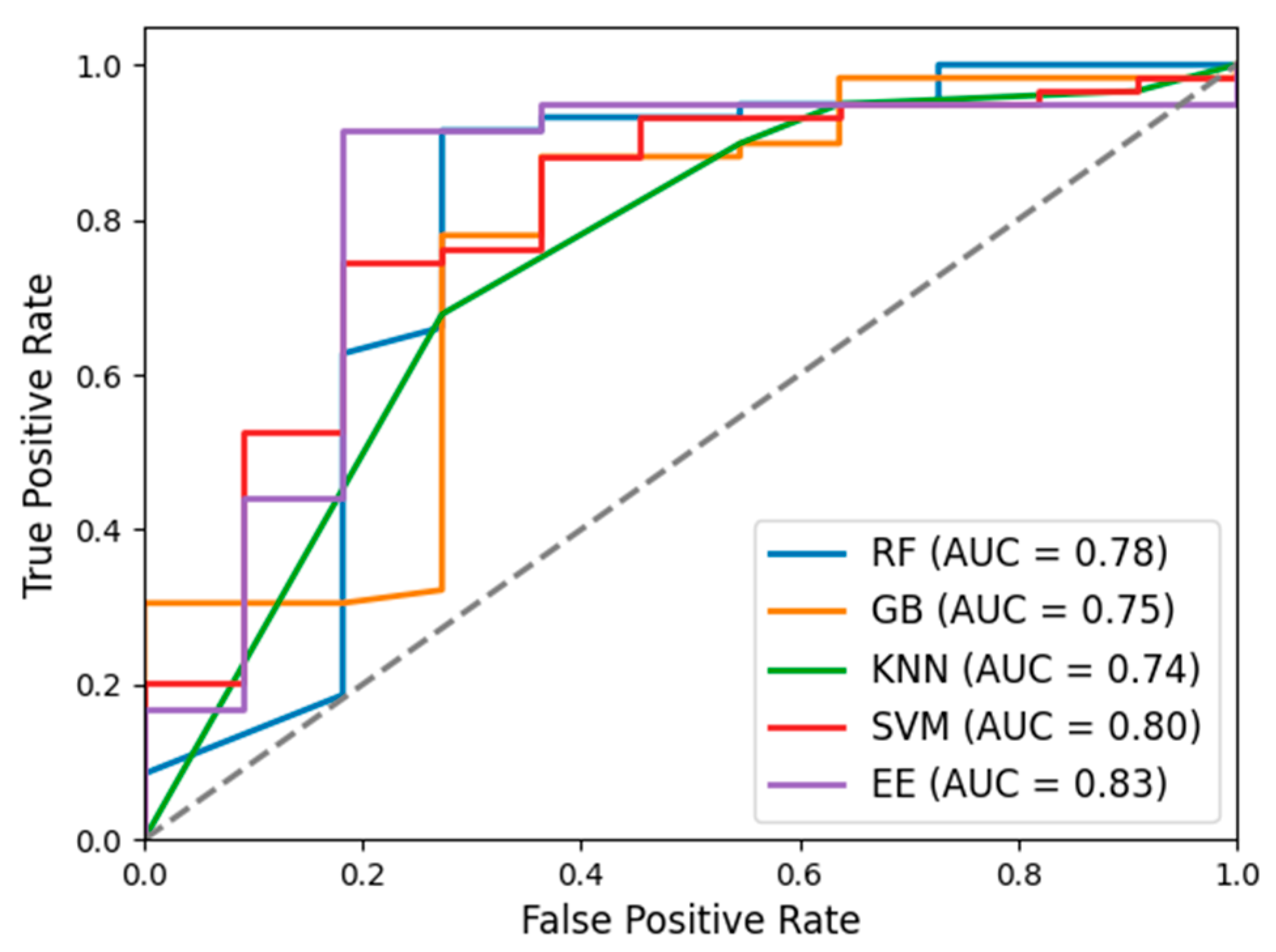
The ROC curves (
Figure 3) were generated for all five classifiers, and their respective AUC values were computed to evaluate their discriminative performance in distinguishing calves with and without NCD. The EE classifier achieved the highest AUC value (0.83 ± 0.03), indicating its robust capability in distinguishing between NONCD and NCD. This superior AUC aligns with EE’s strong performance metrics in recall, F1 score, and accuracy, reaffirming its effectiveness in detecting NCD. The SVM classifier also exhibited notable performance with an AUC value of 0.80 ± 0.09. Despite its moderate precision, recall, and F1 score, the high AUC underscores SVM’s overall capacity to differentiate between classes effectively. In contrast, the other classifiers, including RF, GB, and KNN, showed lower AUC values of 0.78 ± 0.05, 0.75 ± 0.06, and 0.74 ± 0.04, respectively. These lower AUC values indicate comparatively weaker discriminatory ability in distinguishing between NONCD and NCD cases compared to EE and SVM.
3.3. Generalization of Machine Learning Classifiers for NCD Detection
To investigate the generalization capability of machine learning classifiers in practical settings, suckle pressure data from a subset of calves were used to predict the suckle pressure status of other calves. The objective was to determine the robustness and adaptability of these models beyond the dataset of calves used for initial training. The training and testing sets for both NONCD and NCD maintained an 8:2 ratio, with five different combinations of calf data used for evaluation. These combinations were generated through stratified random sampling to ensure that each dataset maintained the same proportion of NCD and NONCD. The results are summarized in
Table 1.
The EE classifier manifested significant generalization capability, as evidenced by its mean recall of 0.60 and F1 score of 0.55. The relatively low standard deviations in precision (0.17) and accuracy (0.06) further indicate the model’s stability and robustness in detecting NCD across different data subsets. However, substantial variation in recall across different data subsets was observed in the EE classifier, with a relatively large SD (0.29), suggesting sensitivity to dataset variations. Moreover, the EE classifier’s precision (0.51) and accuracy (0.84) were comparatively lower than those of other classifiers. This can be attributed to its higher recall, emphasizing the model’s tendency to identify more positive cases, potentially at the cost of increased false positives.
In comparison, the KNN classifier demonstrated the highest mean precision (0.69), but with significant variability (SD = 0.42). The high SD indicates that the performance was highly dependent on the specific data combination used. Its lower recall (0.36) and F1 score (0.47) indicate that KNN may not be as effective in reliably detecting NCD. RF and GB classifiers showed moderate mean values across all metrics, with RF having a slight edge in overall performance stability, as indicated by its higher mean values compared to GB. However, their lower recall values (0.47 and 0.42, respectively) highlight their limited effectiveness in identifying diarrheic calves. The SVM classifier presented the highest variation in performance with a mean precision of 0.69 (SD = 0.42) and a notably lower mean recall of 0.14 (SD = 0.08). The high precision but low recall suggests its capability to accurately identify a limited number of NCD cases while missing a significant portion, thereby restricting its utility for reliable NCD detection.
3.4. Prediction of NCD Occurrences from Pre-Diarrhea Data
The early detection of NCD is critical for timely intervention and improving the health outcomes of affected calves. This work evaluates the predictive performance of various machine learning classifiers in identifying NCD occurrences using pre-diarrhea data. By assessing the models’ ability to predict NCD before the onset of clinical symptoms, the purpose is to determine the earliest time point at which reliable predictions can be made.
To achieve this, the data were split into training and testing subsets based on a temporal threshold to facilitate the analysis. The training set comprised data from NCD within a specified time threshold (1, 2, 3, 4, or more days (4+)) prior to the onset of NCD, which were labeled as the NCD training set. Data from healthy calves before this time threshold were labeled as the NONCD training set. For the testing, data from both NCD and NONCD after the first onset of NCD were used.
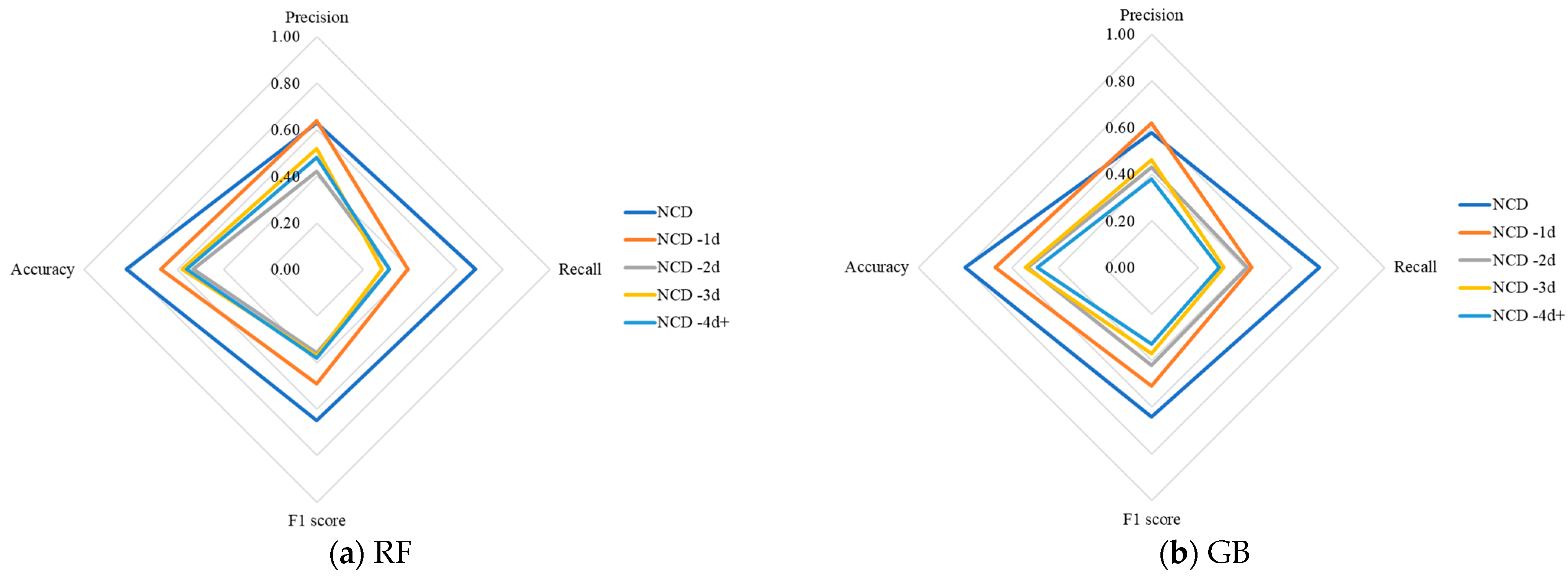
The radar charts in
Figure 4 depict the performance of five different classifiers for predicting NCD occurrences at various intervals before the onset of diarrhea. Each axis represents a different performance metric, while the lines represent different prediction intervals. The values are plotted on a scale from 0 to 1, with points closer to the outer edges of the chart indicating better performance. A larger area covered by the plotted shape signifies more robust performance across multiple metrics.
The results show that as the prediction window lengthens, there is a noticeable decline in performance metrics across all classifiers. This consistent pattern across classifiers indicates a general challenge in maintaining high NCD prediction accuracy as the lead time increases. Furthermore, with the extension of the prediction window, the overall area covered by each classifier’s performance shape in the radar charts also diminishes. This indicates a reduction in the robustness and reliability of predictions over longer intervals. Notably, the radar charts reveal that all classifiers have their highest performance metrics when using data from the initial NCD event for subsequent predictions. However, making predictions before the first occurrence of NCD is more critical as it allows for earlier interventions, potentially preventing the onset of severe symptoms. Among the different prediction intervals, “One day before NCD” (NCD-1d) consistently delivers the most favorable results across the evaluated classifiers. In particular, the EE classifier achieved the highest performance at this interval, with precision, recall, F1 score, and accuracy values of 0.67, 0.70, 0.68, and 0.74, respectively.
In comparison, the KNN classifier also demonstrated high precision (0.71) at NCD-1d but had a significantly lower recall (0.22), resulting in an F1 score of 0.33 and accuracy of 0.66. This suggests that while KNN can accurately identify some positive cases, it fails to detect a large proportion of true instances of NCD, which compromises its effectiveness in early diagnosis and intervention efforts. The RF and GB classifiers showed relatively strong performance at NCD-1d, with RF achieving a precision of 0.64, recall of 0.39, F1 score of 0.49, and accuracy of 0.67, and GB achieving a precision of 0.62, recall of 0.43, F1 score of 0.51, and accuracy of 0.67. Although their performance metrics are lower than those of EE, they still indicate a reasonable capacity for early NCD prediction. The SVM classifier exhibited the lowest performance at NCD-1d, with precision, recall, F1 score, and accuracy of 0.36, 0.08, 0.13, and 0.54, respectively. This poor performance underscores the limitations of SVM for early NCD detection, especially in practical applications where high precision and recall are required.
As the prediction interval extends from NCD -2d to NCD -4d+, all classifiers experienced a marked decline in performance metrics. For instance, EE’s performance metrics dropped to a precision of 0.38, recall of 0.47, F1 score of 0.42, and accuracy of 0.46 at NCD -2d. This downward trend continued, with EE showing even lower values at NCD -3d (precision: 0.42, recall: 0.49, F1 score: 0.45, accuracy: 0.49) and NCD -4d+ (precision: 0.36, recall: 0.41, F1 score: 0.38, accuracy: 0.44). Similar declines were observed for KNN, RF, GB, and SVM, reflecting the increasing difficulty of making accurate predictions as the interval before the first NCD event increases.
4. Discussion
The present study investigated whether variations in suckle pressure could serve as an early marker to identify and predict NCD. In an investigation of an NCD outbreak and several retrospective case–control studies, it was observed that calves with NCD exhibited differences in suckle reflex compared to healthy calves, with varying degrees of NCD corresponding to distinct patterns in suckle reflex [
11,
38]. Typically, changes in feeding behavior and suckle pressure precede the appearance of clinical symptoms of NCD. Farm staff commonly monitor calves for visible indicators of illness, such as changes in attitude, changes in feed intake, or signs of dehydration. However, these clinical signs may not be readily apparent until a calf is severely affected. By the time these clinical signs are present, intestinal damage may have already occurred, which can compromise health and require extended recovery, even though mucosal repair is possible with timely intervention [
4,
15]. Thus, identifying quantifiable, subtle behavioral changes before overt clinical symptoms appear could significantly enhance calf health management and welfare.
Supervised machine learning is extensively applied in predictive and classification tasks. Supervised learning involves training models on labeled data to make accurate predictions or classifications for new, unseen data [
39]. This method enables the identification of relationships between input features and known output labels, thereby facilitating the generalization of patterns for informed predictions. Deep learning models, while powerful in capturing complex patterns, require substantial amounts of data to generalize well and avoid overfitting [
40], which was not feasible with our limited dataset size. Traditional supervised learning models such as RF, GB, KNN, SVM, and EE were utilized in this work as they are well suited for smaller datasets and offer better interpretability. Multiple machine learning models, including RF, GB, KNN, SVM, and EE, were assessed based on features derived from the pressure patterns generated during calf suckling, aiming to identify calves experiencing or about to experience NCD. This approach expands practical tools available to dairy producers, enabling timely, objective detection of NCD without relying on subjective methods such as manual evaluation of suckle strength.
Our results highlight the potential of the proposed approach for analyzing suckle pressure images to enable early detection and prediction of NCD onset. The comparative analysis of different machine learning models demonstrated that the EE algorithm consistently outperformed others. The superior performance of EE can be attributed to its ability to effectively handle imbalanced data, where NONCD significantly outnumbers cases of NCD. Data imbalance is a common issue in agricultural settings [
41,
42], where certain conditions, such as NCD in calves, may not occur frequently but have significant impacts when they do. This imbalance often leads to biased predictions in favor of the majority class, increasing the risk of false negatives. Missing true NCD cases can have serious consequences, as delayed intervention may lead to severe dehydration, electrolyte imbalance, and increased mortality risk [
43]. EE addresses this by combining multiple weak learners, each trained on balanced subsets of the data, thus enhancing the model’s sensitivity to the minority class [
44]. This enhanced recall is particularly valuable in disease detection, where early identification of affected calves allows timely intervention to prevent complications. While false positives may lead to unnecessary preventive measures, such as electrolyte supplementation, the associated costs and risks are minimal compared to the potential harm of missing true cases [
45]. EE’s ability to improve recall while maintaining high precision offers potential for early NCD detection in dairy calves through suckle pressure analysis.
Generalization is critical for practical applications, particularly in agricultural settings where data variability is common due to differences in management practices, housing, environmental conditions, and individual animal responses. This study evaluated the ability of the proposed ML method to generalize by using suckle pressure data from one group of calves to predict the condition of different, untested calves. Ensuring that the model can reliably predict NCD across different groups of calves under various conditions is essential for its deployment in diverse farming environments. However, the prediction performance for new calves was slightly lower compared to the calves used for training. This discrepancy can be attributed to the inherent variability in agricultural settings and the relatively limited dataset size. Despite these challenges, the models still demonstrated a reasonable capability for NCD prediction, underscoring their promise for practical use in early detection and intervention.
Identifying calves at risk before clinical symptoms of NCD appear can significantly reduce morbidity and mortality rates [
4]. The results of this study indicate that the most effective NCD prediction occurred one day prior to the onset of NCD. Even with this short lead time, farm staff could implement closer monitoring and administer preventive care to at-risk calves, potentially mitigating the severity of the disease. Early intervention during these critical windows helps prevent symptom escalation, which in turn enhances calf health and reduces the economic impact associated with disease outbreaks. This proactive management approach not only supports animal welfare but also meets practical needs in the field, aligning with the objectives of sustainable livestock practices. By minimizing the need for intensive treatments and lowering the risk of widespread outbreaks, this strategy offers a practical and economically viable solution for breeders.
Previous studies have explored various methods for detecting and predicting NCD, including monitoring activity patterns and feeding and drinking behaviors [
5,
12,
15,
46,
47,
48]. A recent study employing ear tag-based accelerometers for the prediction of NCD revealed that calves with NCD exhibited decreased activity and increased lying time on the day before diagnosis [
46]. The lying time during the day prior to diagnosis showed a strong predictive capacity for NCD. These findings are consistent with our work, where the prediction performance was most accurate on the day before NCD onset. This limited prediction window likely reflects the timing of physiological changes that occur just before the clinical symptoms become apparent. However, while these methods have demonstrated some predictive capability, their clinical utility is limited, with sensitivity and specificity varying across studies due to differences in study design, environmental conditions, and feeding system variations [
5]. Additionally, the effectiveness of these techniques can vary depending on the feeding systems (ad libitum feeding or restricted feeding) in use, making them less universally applicable [
12].
Suckle pressure measurement presents several practical advantages over existing methods such as ACF and accelerometer-based monitoring. Utilizing pressure-sensitive sensors applied around the nipple to record the pressure exerted by the calf during suckling has the potential to be seamlessly integrated into standard feeding routines. Real-time feedback provided by pressure monitoring enables immediate detection of deviations from normal feeding patterns, even in facilities without automated feeders, facilitating timely interventions. Looking ahead, the design of suckle pressure measurement systems is expected to benefit from the incorporation of advanced sensor technologies. Integrating sensors with enhanced sensitivity and data transmission capabilities has the potential to enhance the precision and functionality of these systems, further improving their effectiveness in monitoring calf health.
Interpretation of the results should acknowledge certain limitations inherent in this study. A primary limitation is the relatively small sample size, which constrains the robustness and generalizability of the models. Additionally, the substantial disparity in the number of suckle pressure data collected from healthy calves versus those with NCD resulted in a significant class imbalance, posing challenges for training effective machine learning models. Another limitation lies in the relatively short duration of data collection. The study was conducted over a limited period, which may not fully capture seasonal variability in calf behavior and disease occurrence. In particular, microclimatic factors—such as elevated ambient temperatures and humidity during summer—can influence both feeding behavior and the prevalence of NCD, potentially affecting the model’s generalizability to different environmental conditions. To address these limitations, future work will focus on expanding the dataset to include a larger and more balanced representation of healthy and diarrheic calves for enhancing the model’s performance. Moreover, employing techniques to address data imbalance, such as advanced sampling strategies, could improve the accuracy and reliability of the models. Conducting longitudinal studies across different farms and environmental conditions would also help validate the model’s applicability in diverse agricultural settings, ensuring its practical utility for early NCD detection and intervention.
5. Conclusions
This preliminary study demonstrates that the integration of suckle pressure measurement with machine learning models offers a viable approach for the identification and prediction of NCD. Among the models evaluated, the EE model performed the best, especially under the constraints of limited and imbalanced data. The results indicate that the predictive performance was most effective one day prior to the clinical manifestation of NCD. From a physiological standpoint, suckling behavior reflects the calf’s overall vitality and neuromuscular coordination, which are often compromised during the early stages of disease. Decreased suckle pressure may be indicative of metabolic acidosis, dehydration, or altered feeding motivation—conditions that precede visible signs of diarrhea. Our findings suggest that subtle changes in suckle pressure can serve as a non-invasive, early signal of deteriorating health status, enabling more timely interventions before irreversible intestinal damage occurs. While these results are promising, further work is needed to enlarge the dataset, refine model parameters, and validate the findings across different production settings and environmental conditions.
The integration of suckle pressure measurement devices with automated data processing systems could be adapted for on-farm use, pending further research and validation under commercial conditions. Portable suckle pressure sensors are relatively low-maintenance and could be incorporated into routine calf feeding protocols without major disruption to farm workflows. While the actual cost would depend on device specifications, production scale, and integration with farm management software, such tools have the potential to be economically viable, particularly when balanced against savings from reduced disease incidence and improved calf performance.