1. Introduction
Cotton, a globally significant strategic resource, plays a crucial role in sustaining national economies and livelihoods [
1]. Currently, the cotton industry faces challenges, such as high labor and material costs, particularly in large-scale cotton field management and water resource allocation [
2]. Traditional methods for gathering information about cotton fields rely on manual sampling surveys, which are inefficient, prone to errors and omissions, inadequate for meeting the demands of rapid and accurate data collection [
3], and time-consuming. Annual data on cotton planting areas at the regional level primarily depend on remote sensing estimates and sampling surveys, which involve temporal delays and limit their applicability in production decision-making. Therefore, accurate and rapid monitoring of cotton-planting areas is essential to ensure robust, large-scale, and sustainable production [
4,
5].
The rapid advancement of satellite remote sensing technology has enabled innovative solutions for the precise extraction of regional crop information by providing high-frequency, multi-dimensional observational data [
6]. This technology effectively covers extensive crop cultivation areas and provides timely and dynamic crop-planting information, thus facilitating the development of precise and smart agricultural systems. When combined with machine learning algorithms, the spectral characteristics of multispectral data can improve the accuracy of the model [
7]. Maruf et al. [
8] utilized the extensive spectral imagery provided by Sentinel-2 for land cover mapping, achieving an accuracy of 74% using the maximum likelihood classification (MLC) method. They also assessed flood damage across various land categories, resulting in an overall accuracy (OA) of 89.06%. Wu et al. [
9] used gradient-boosting decision trees (GBDT) to develop a model correlating brightness temperature with associated surface parameters. Their findings indicated that the GBDT model achieved a correlation coefficient of 67.9% and demonstrated greater effectiveness during summer and in regions characterized by complex land cover. Guo et al. [
10] applied partial least squares regression (PLSR) to estimate soil organic carbon using various spectral bands and the normalized difference vegetation index (NDVI) from multiple satellite imagery datasets. Their results demonstrated that machine learning techniques outperformed traditional regression models in terms of predictive capability.
Previous studies have shown that individual satellites equipped with multispectral sensors face challenges in simultaneously achieving extensive coverage and high-precision imagery [
11]. Then, researchers have looked to fuse images from satellites equipped with various sensors to improve the accuracy of satellite-based monitoring. Zhang et al. [
12] used Landsat and Sentinel-2 images combined with logistic regression models to evaluate the effectiveness of the Enhanced Vegetation Index (EVI), NDVI, and Land Surface Water Index (LSWI) in assessing the annual start of season (SOS). Zhou et al. [
13] used images from Landsat-5, Landsat-7, Landsat-8, Sentinel-2, and Gaofen-1 satellites, demonstrating that the Plastic-Mulched Citrus Index (PMCI) performed exceptionally well in extracting plastic-mulched citrus (PMC) across various observation dates, with an OA exceeding 0.91 for both intra-annual and inter-annual PMC detection. Kang et al. [
14] integrated spectral, Synthetic Aperture Radar (SAR), topographic, and textural features using the Google Earth Engine (GEE) platform. Their findings demonstrated that incorporating SAR data along with topographic and textural features improved both user and producer accuracies for tea plantation classification, increasing from 89.3% and 85.9% to 95.8% and 91.7%, respectively. Li et al. [
15] used multi-temporal Landsat 8 Operational Land Imager (OLI) images, combined with spectral angle mapping and decision tree classification, to extract the major crop distributions in the Eastern Xinrong District of Datong City. They compared their results with those obtained using the MLC. However, research on cotton field area extraction using multi-source remote sensing data remains limited, with only a few studies successfully mapping cotton fields using high-resolution imagery. Zhang et al. [
16] extracted cotton field maps in Northern Xinjiang using Sentinel-1/2 remote sensing data, combined with a random forest classifier (RFC) and multi-scale image segmentation, achieving an OA of 0.932 and a kappa coefficient of 0.813.
With the rapid advancement of deep learning, its applications in semantic segmentation and object detection in remote sensing imagery have become increasingly widespread [
17,
18,
19,
20]. Li et al. [
21] conducted a comparative study using Gaofen-1 satellite imagery of Zhengzhou City to evaluate four different deep-learning network models for automated land cover classification in high-resolution images. Their results demonstrated that the MS-EfficientUNet method achieved optimal performance in classifying land cover in Zhengzhou with an OA of 0.7981. Cultivated land yielded the highest intersection over union (IoU) and F1-Score values of 0.7801 and 0.8764, respectively. Zhong et al. [
22] developed two types of deep learning models, long short-term memory (LSTM) and 1D convolutional neural networks (NNs), for summer crop classification. Their study revealed that the Conv1D-based model achieved the highest accuracy of 85.54% and an F1-score of 0.73. Seydi et al. [
23] proposed a novel framework that integrates deep convolutional neural networks (CNNs) with dual attention modules (DAM) using Sentinel-2 time-series datasets to generate accurate and timely crop-type maps. This approach achieved exceptional performance, with an OA of 98.54% and a kappa coefficient of 0.981.
In summary, although numerous studies have utilized multi-source satellite image fusion for land-cover area estimation, studies specifically focusing on cotton field extraction remain limited. Although multi-source remote sensing data fusion has demonstrated improved classification accuracy for certain crops, the effectiveness of modeling varies significantly depending on the fusion methodologies and feature band selection approaches used. Furthermore, the development of training datasets is constrained by regional geographical characteristics, climatic conditions, and spatiotemporal spectral variations in observational parameters, which collectively restrict the spatial transferability of the existing cotton field mapping techniques. The Alar Reclamation Zone in Xinjiang, China, with its extensive cotton cultivation area, offers a wealth of annotated data for cotton field extraction. This presents a critical opportunity to develop high-precision, cost-effective models for cotton field mapping and acreage estimation by integrating satellite remote sensing with deep-learning technologies. The development of such models is particularly urgent to address the current agricultural needs.
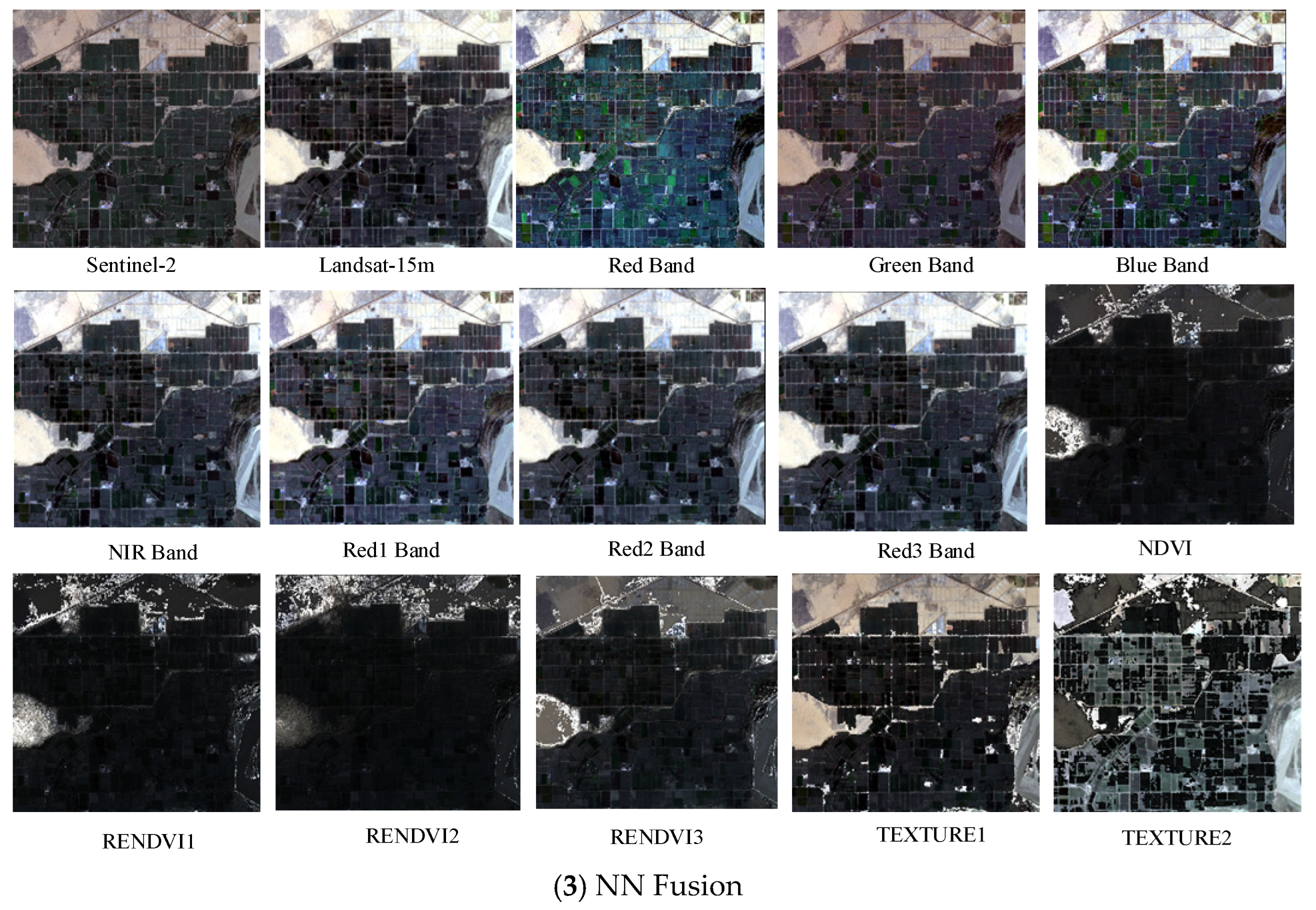
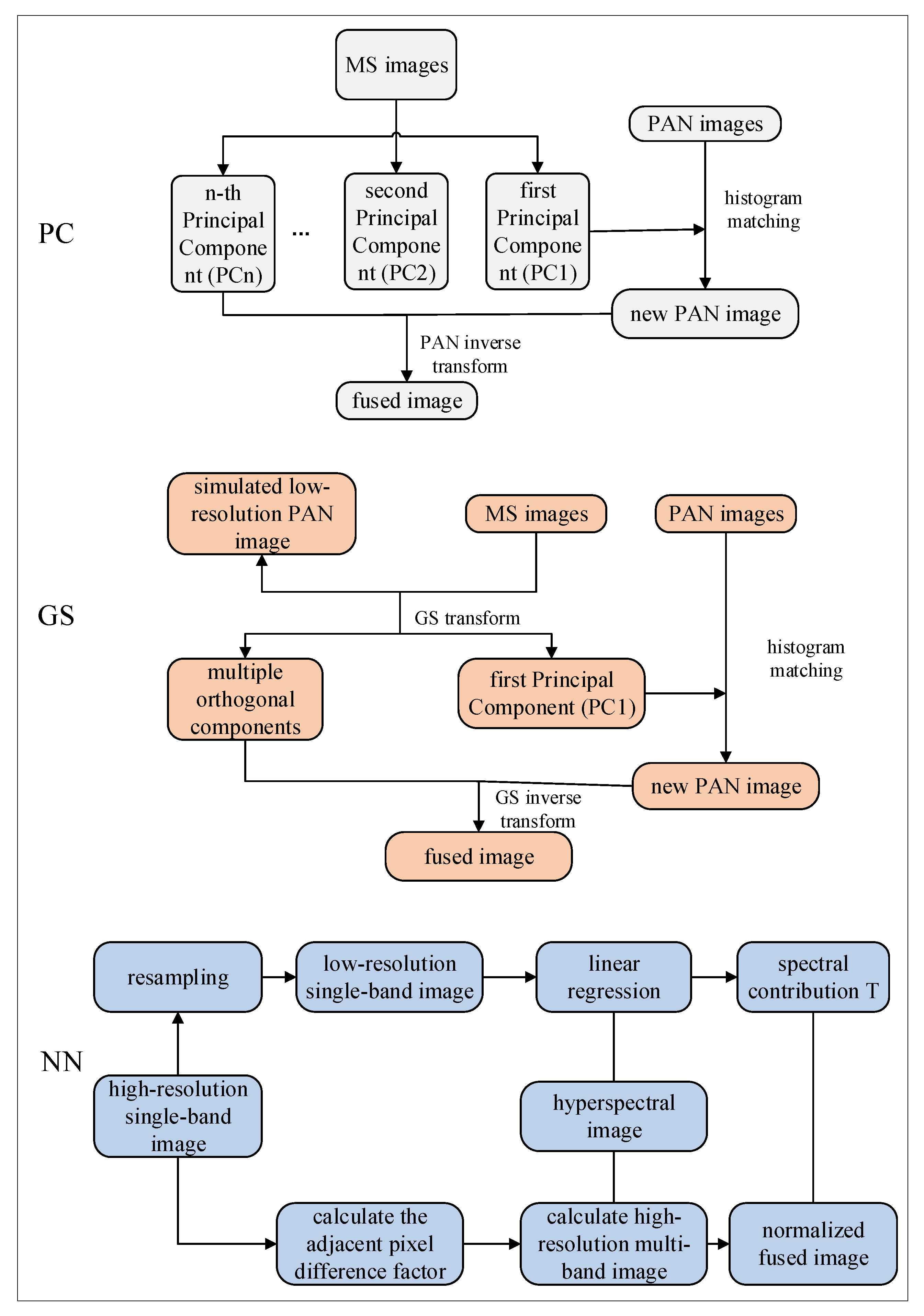
This study focused on cotton cultivation areas in Alar City, Xinjiang, using a comprehensive methodological framework to address the existing research gaps. Multi-temporal satellite imagery was collected from GF-1, Landsat 8, and Sentinel-2, implementing three distinct fusion approaches, principal component (PC), Gram–Schmidt (GS), and NN, to integrate spectral, vegetation, and texture features through pairwise image fusion. The optimal feature bands for cotton field extraction were systematically selected through quantitative evaluation using three key metrics: the mean value, standard deviation, and information entropy. A confusion matrix analysis was conducted to comparatively assess the performance of various machine-learning classifiers in cotton field extraction, using optimally fused imagery. Furthermore, the U-Net architecture was improved by incorporating both a Convolutional Block Attention Module (CBAM) and an Atrous Spatial Pyramid Pooling (ASPP) module to improve mapping accuracy. This study evaluated the estimates of cotton cultivation areas derived from various classifiers, using multi-dimensional assessment criteria. This study provides scientifically robust data support and decision-making support for multispectral remote sensing methodologies in cotton field extraction, particularly for addressing the challenges of precision agriculture in arid regions.
4. Discussion
In agricultural remote sensing applications, multi-satellite data fusion technology offers comprehensive and accurate information and plays a significant role in crop monitoring and yield prediction. In this study, GF-1 and Sentinel-2 data were selected for fusion with the Landsat 8 imagery. Three fusion methods (PC, GS, and NN) were used to fuse the spectral characteristics, vegetation indices, and texture features of single-source satellite imagery. Fused imagery combining the GS-blue band of Sentinel-2 with that of Landsat 8 demonstrated superior performance in terms of both color representation and information content. In particular, spectral feature fusion significantly improved the clarity and color contrast of cotton-field images compared with the other methods. During the fusion of Sentinel-2 and Landsat 8, significant improvements were observed in mean, standard deviation, and information entropy. The fused imagery demonstrated clear advantages over single-source images for the accurate extraction of cotton fields. The results of this study demonstrated that the fusion of two satellite datasets yielded better outcomes than single-satellite data, which is consistent with the findings of Ram [
48], who reported superior vegetation mapping results in Japan using fused Sentinel-2 and Landsat 8 data compared with single-satellite approaches. Li et al. [
49] found that using advanced remote sensing algorithms and various fusion data for growing stock volume estimation, the fusion image data based on GF-2 and Sentinel-2 can effectively couple the advantages of the two and significantly improve the estimation performance of growing stock volume.
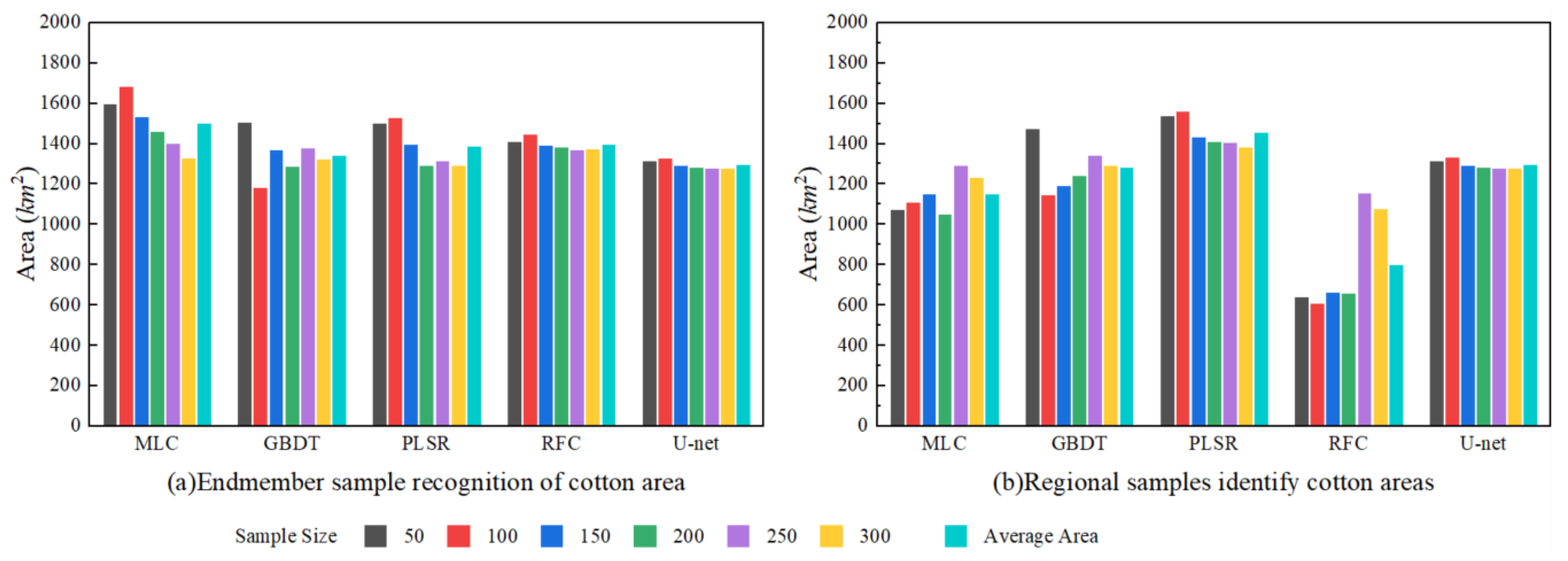
The endmember- and region-based sample extraction methods significantly improved the classification accuracy of machine learning classifiers. The four machine learning models exhibited minimal differences in kappa and OA values between the two sample types. Among all classifiers, RFC demonstrated optimal performance with endmember sample sizes of 250 and 300, achieving a kappa value of 0.963 and an OA of 97.22% (0.961 and 97.18%, respectively, for the regional samples). UA and PA metrics for the RFC model with 250 samples were 97.13% and 97.09%, respectively, successfully extracting a cotton field area of 1405.25 km
2. Although Hu [
16] achieved 89.4% OA in cotton classification within the framework of multi-crops, we improved the accuracy to 97.22% by optimizing the key methods of sample selection. The improvement of 7.82% showed that the crop-specific sample strategies (endmember/region) are better than general random sampling. When the purity of the sample is ensured, RF becomes very important. These results underscore the importance of high-quality samples in enhancing the accuracy of cotton identification.
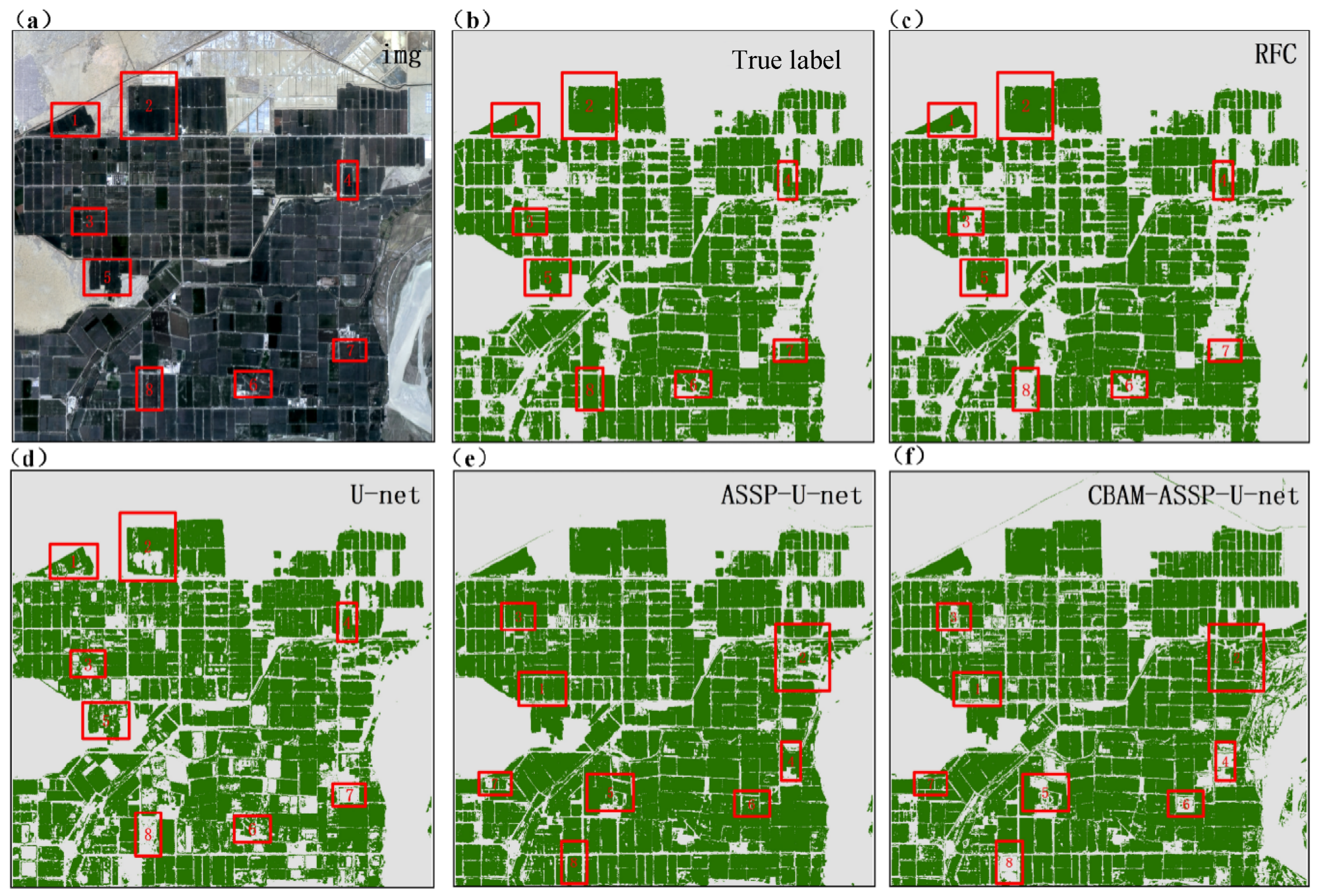
To enhance the U-Net model, this study incorporated the CBAM and ASPP modules, resulting in the CBAM-ASPP-U-Net model. When applied to cotton identification using GS-blue band images fused from Sentinel-2 and Landsat 8 data, this improved model demonstrated superior performance compared with the original U-Net, achieving higher values for kappa (0.988), OA (98.56%), UA (98.99%), and PA (100%). CBAM-ASPP-U-Net achieved 98.56% OA in cotton mapping, surpassing Seydi et al.’s [
23] time-series CNN (95.2%). Our spatial attention method performs well in the case of mixed pixels (PA = 100% versus 92.8%), which proves that GS band fusion is superior to time features in small field of view recognition. The OA gain highlights the advantages of ASPP in precision agriculture applications compared with sequential attention. The model demonstrated improved effectiveness in identifying cotton fields that were intercropped with other crops. These results indicate that the CBAM-ASPP-U-Net model can learn spatial features more effectively, thus improving its ability to recognize detailed ground objects. This study addressed the challenge of mixed-pixel extraction in small cotton fields using intercropping systems and significantly improved the accuracy of cotton field extraction. The findings of the CBAM-ASPP-U-Net model used in this study are consistent with those of Ai et al. [
50], who developed an SCA-UNet model by integrating CBAM and the Squeeze-and-Excitation Network (SE) into the U-Net algorithm for rice field levee extraction from remote sensing imagery, achieving better results than conventional U-Net and other models. Liu et al. [
51] added ASPP and CBAM dual attention mechanisms to the U-Net model to form the backbone network of the model, which enhanced the model’s ability to extract features from winter wheat information, and its results were better than FCN, U-Net, DeepLabv3, SegNet, ResUNet, and UNet, which once again proved the excellent performance of the ASPP-CBAM-U-Net model.
Table 9 shows the performance of the proposed method compared to other deep learning methods.
This study focused on selecting optimal fused imagery and identifying the most suitable cotton field classification samples and classifiers for the Aral reclamation area while acknowledging several limitations. Images from May to September were used to represent the critical periods of cotton growth. Future studies should incorporate multi-temporal imagery that covers the entire growth cycle. Although the current study distinguished cotton from other land cover types, future research could involve simultaneous analysis of multiple crop types to better address practical requirements. Although this study utilized satellite remote sensing imagery, incorporating UAV imagery into future fusion processes could improve the modeling accuracy and the economic benefits of cotton farming. Future research could explore more effective classification algorithms (e.g., deep learning models) and more representative training sample selection methods to improve the accuracy of cotton field recognition in a complex environment. In addition, the integration of field investigation and agricultural management data is helpful to verify the practical applicability of classification results and evaluate its potential to improve the economic benefits in precision cotton planting (e.g., optimizing irrigation and fertilization decisions.
5. Conclusions
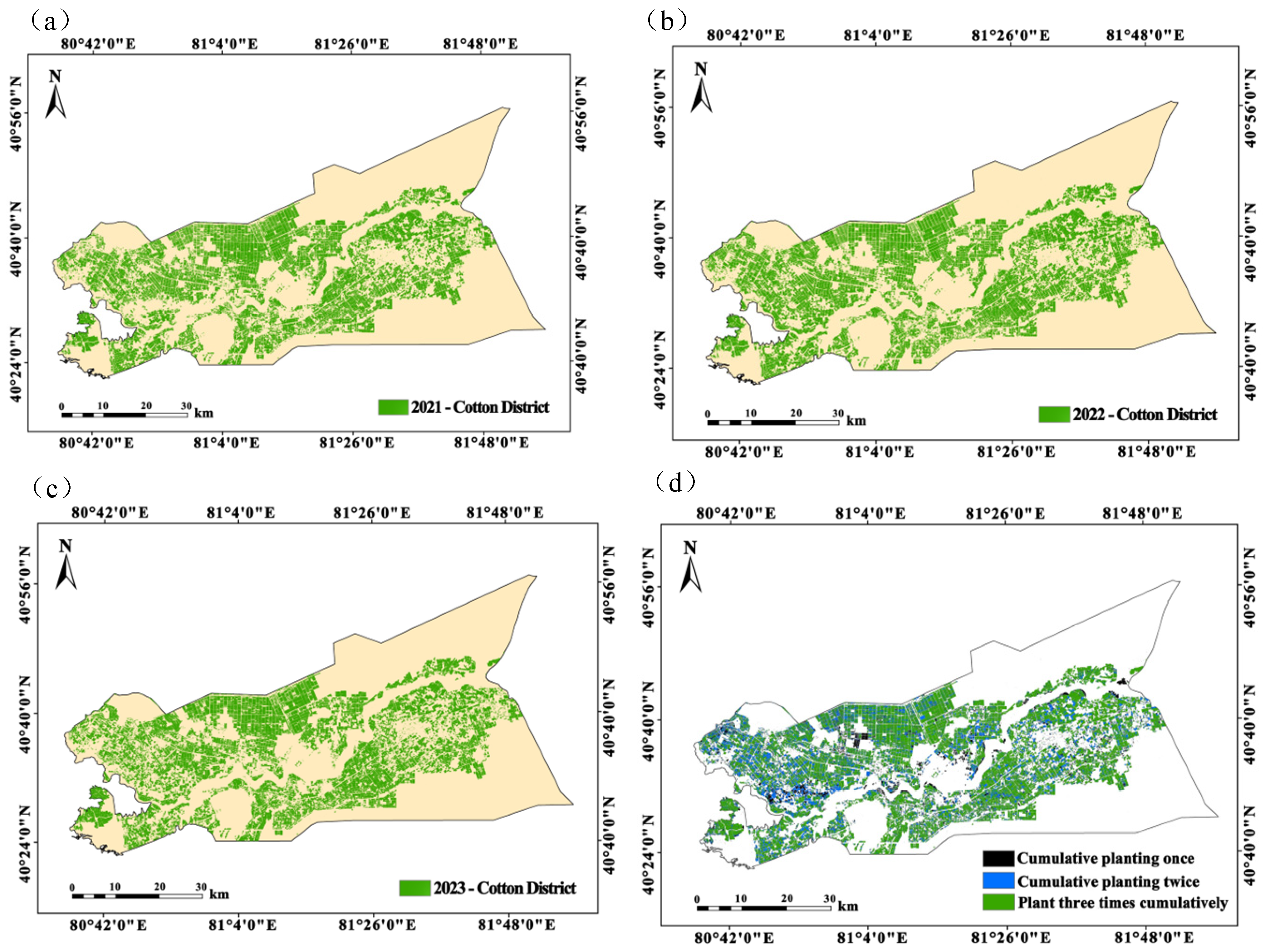
This study investigated the identification of cotton fields in the Alar Reclamation Zone of Xinjiang and evaluated the effectiveness of multi-source remote sensing data fusion and classification algorithms for monitoring cotton cultivation. A comparative analysis of GS fusion revealed that the fusion of Sentinel-2 and Landsat 8 data in the blue band provided the most effective feature representation, significantly outperforming the fusion results of GF-1 and Landsat 8. In evaluating sampling strategies, endmember selection proved to be more effective than regional sampling, with the number of samples showing a significant positive correlation with the classification accuracy. Utilizing the GS-blue band fused imagery, the RFC achieved peak performance (kappa = 0.963, OA = 97.22%, UA = 97.13%, PA = 97.11%) for 250 endmember samples. However, it exhibited considerable error in area estimation. The CBAM-ASPP-U-Net model delivered exceptional cotton field identification using GS-blue band-fused Sentinel-2/Landsat 8 data (kappa = 0.988, OA = 98.56%, UA = 98.99%, PA = 100%), with an area estimation accuracy of 1341.25 km2 (a deviation of 25.42 km2 from the actual cultivation area). Digital mapping of continuous cotton cropping from 2021 to 2023, based on the optimal model, revealed a progressive reduction in cultivated area, confirming the effectiveness of crop rotation policies in mitigating monoculture practices. The developed “multi-source data fusion–deep feature optimization” framework successfully achieved high-precision cotton field interpretation in Alar, providing an innovative solution for crop monitoring in arid regions. These results not only validate the application value of attention mechanisms and multi-scale feature extraction in agricultural remote sensing but also establish reliable technical support for dynamic cotton acreage monitoring, yield estimation, and precision farm management in the future.