Review on Convolutional Neural Network (CNN) Applied to Plant Leaf Disease Classification
Abstract
:1. Introduction
2. Deep Learning
- Step 1. Preparing the Data and Preprocessing
- Step 2. Building, Training, and Evaluating the Model
- Step 3. Inference and Deployment
2.1. Data Preparation and Preprocessing
2.2. Building Model Architecture, Training, and Evaluating the Model
2.3. Inference and Deployment
3. Problems and Solutions
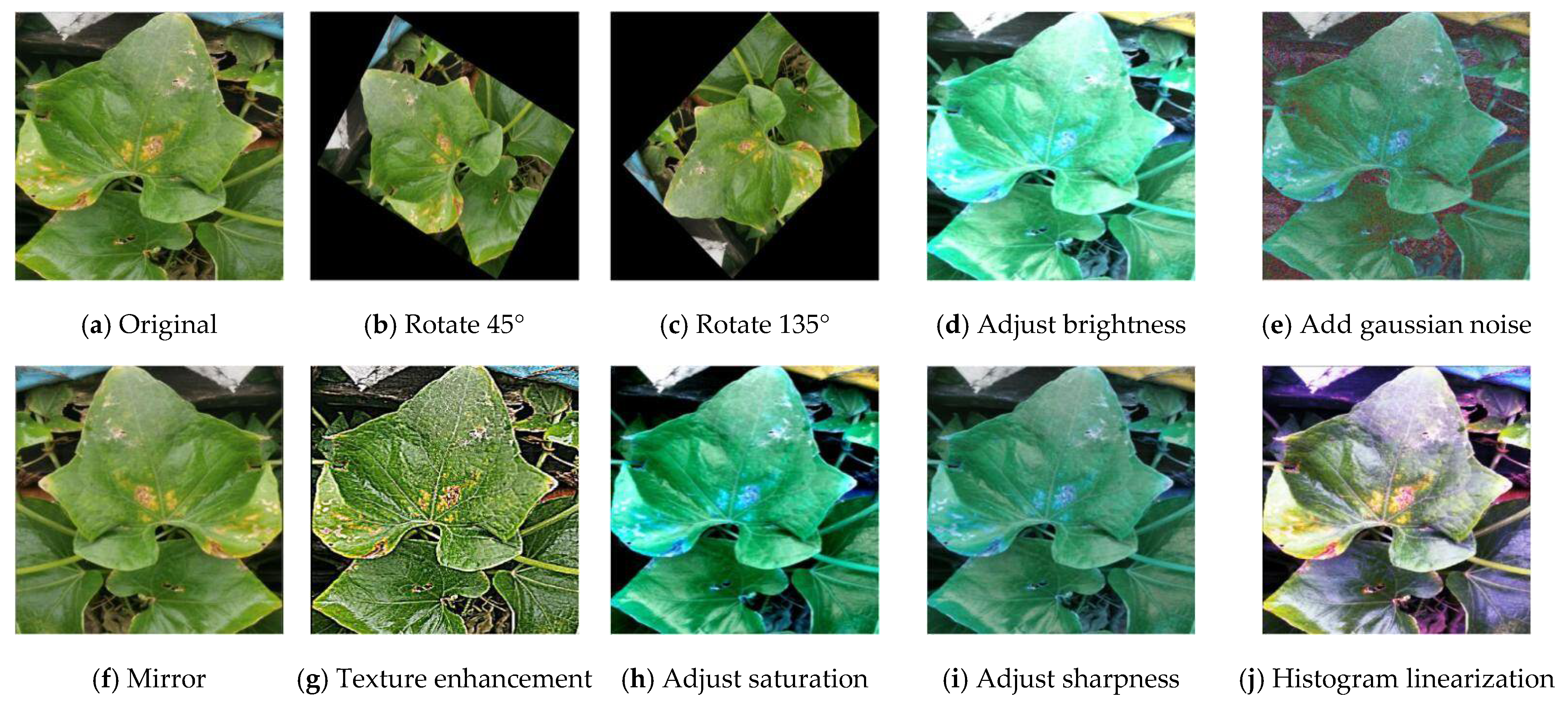
3.1. Insufficient Datasets
| Expanded Dataset | Methods | Best Accuracy | Reference |
|---|---|---|---|
| From 1053 to 13,689 images | Direction disturbance and light disturbance and PCA (Principal components analysis) jittering | 97.62% | Bin et al. (2017) [83] |
| From 10,820 to 32,460 images | Noise addition, color jittering, and radial blur | 96.17% (improved 3.15%) | Lin et al. (2018) [58] |
| From 54,309 to 87,848 images | Cropping, resizing | 99.53% | Ferentinos (2018) [11] |
| From 1567 to 46,409 images | Segmentation, resizing | 94.00% (improved 12%) | Arnal Barbedo (2019) [86] |
| From 5000 to 43,398 images | Resizing, crop, rotation, noise... | 85.98% | Fuente et al. (2017) [72] |
| From 4483 to 33,469 images | Affine transformation, perspective transformation, and rotation | 96.30% | Srdjan et al. (2016) [87] |
3.2. Nonideal Robustness
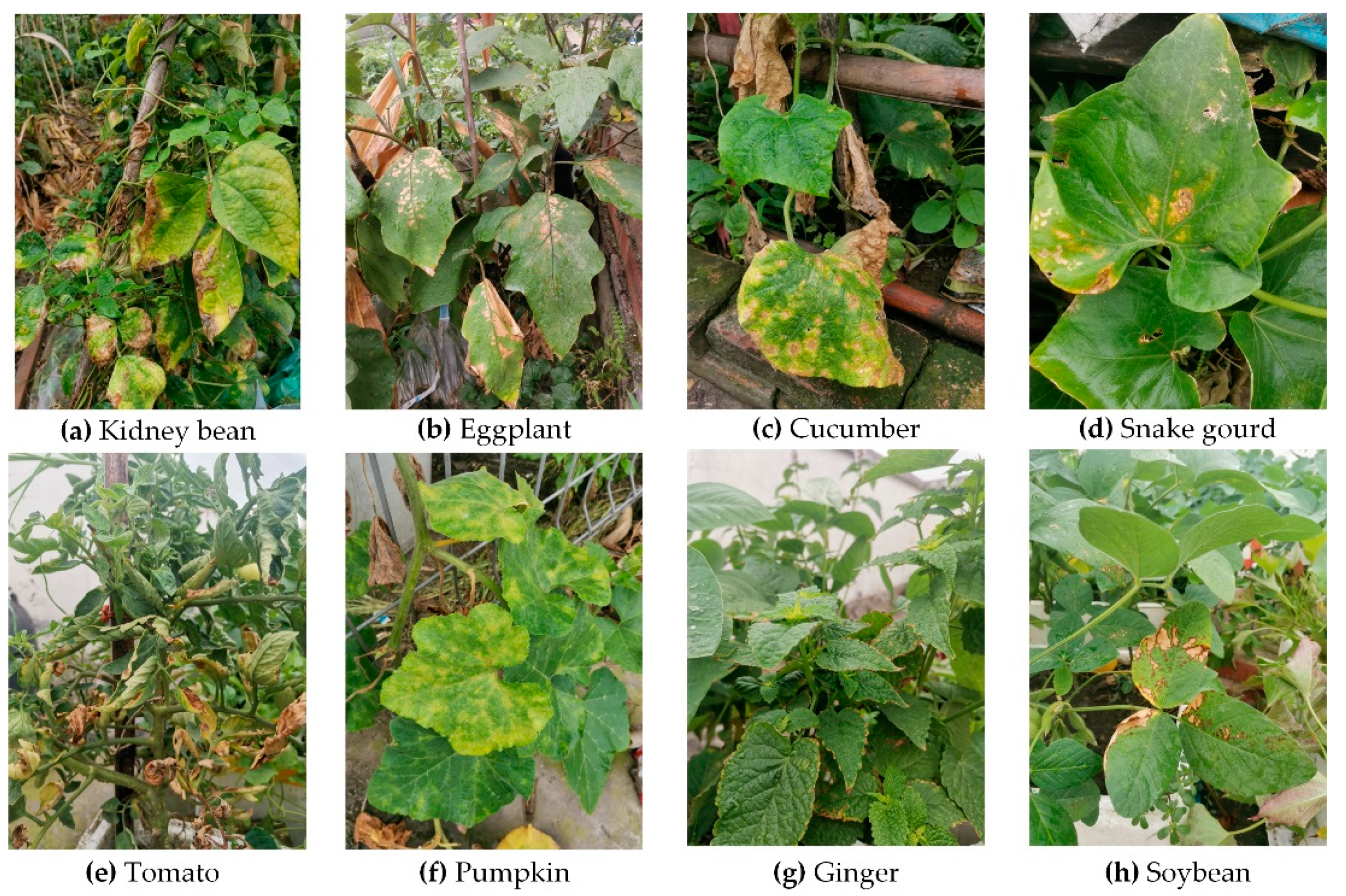
3.3. Symptom Variations
3.4. Image Background
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Mukti, I.Z.; Biswas, D. Transfer Learning Based Plant Diseases Detection Using ResNet50. In Proceedings of the 2019 4th International Conference on Electrical Information and Communication Technology (EICT), Khulna, Bangladesh, 20–22 December 2019. [Google Scholar]
- Arunnehru, J.; Vidhyasagar, B.S.; Anwar Basha, H. Plant Leaf Diseases Recognition Using Convolutional Neural Network and Transfer Learning. In International Conference on Communication, Computing and Electronics Systems; Bindhu, V., Chen, J., Tavares, J.M.R.S., Eds.; Springer: Singapore, 2020; pp. 221–229. [Google Scholar]
- Hughes, D.P.; Salathe, M. An open access repository of images on plant health to enable the development of mobile disease diagnostics. arXiv 2015, arXiv:1511.08060. [Google Scholar]
- Kianat, J.; Khan, M.A.; Sharif, M.; Akram, T.; Rehman, A.; Saba, T. A joint framework of feature reduction and robust feature selection for cucumber leaf diseases recognition. Optik 2021, 240, 166566. [Google Scholar] [CrossRef]
- Zhang, S.; Zhang, S.; Zhang, C.; Wang, X.; Shi, Y. Cucumber leaf disease identification with global pooling dilated convolutional neural network. Comput. Electron. Agric. 2019, 162, 422–430. [Google Scholar] [CrossRef]
- Agarwal, M.; Gupta, S.; Biswas, K.K. A new conv2d model with modified relu activation function for identification of disease type and severity in cucumber plant. Sustain. Comput. Inform. Syst. 2021, 30, 100473. [Google Scholar]
- Chen, J.; Zhang, D.; Zeb, A.; Nanehkaran, Y.A. Identification of rice plant diseases using lightweight attention networks. Expert Syst. Appl. 2021, 169, 114514. [Google Scholar] [CrossRef]
- Shrivastava, V.K.; Pradhan, M.K.; Minz, S.; Thakur, M.P. Rice plant disease classification using transfer learning of deep convolution neural network. Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2019, XLII-3/W6, 631–635. [Google Scholar] [CrossRef] [Green Version]
- Sun, H.; Zhai, L.; Teng, F.; Li, Z.; Zhang, Z. qRgls1. 06, a major QTL conferring resistance to gray leaf spot disease in maize. Crop. J. 2021, 9, 342–350. [Google Scholar] [CrossRef]
- Yu, C.; Ai-hong, Z.; Ai-Jun, R.; Hong-qin, M. Types of Maize virus diseases and progress in virus identification techniques in China. J. Northeast Agric. Univ. 2014, 21, 75–83. [Google Scholar] [CrossRef]
- Ferentinos, K. Deep learning models for plant disease detection and diagnosis. Comput. Electron. Agric. 2018, 145, 311–318. [Google Scholar] [CrossRef]
- Abbas, A.; Jain, S.; Gour, M.; Vankudothu, S. Tomato plant disease detection using transfer learning with C-GAN synthetic images. Comput. Electron. Agric. 2021, 187, 106279. [Google Scholar] [CrossRef]
- Sankaran, S.; Mishra, A.; Ehsani, R.; Davis, C. A review of advanced techniques for detecting plant diseases. Comput. Electron. Agric. 2010, 72, 1–13. [Google Scholar] [CrossRef]
- Barbedo Arnal, J.G. An Automatic Method to Detect and Measure Leaf Disease Symptoms Using Digital Image Processing. Plant Dis. 2014, 98, 1709–1716. [Google Scholar] [CrossRef] [Green Version]
- Feng, Q.; Dongxia, L.; Bingda, S.; Liu, R.; Zhanhong, M.; Haiguang, W. Identification of Alfalfa Leaf Diseases Using Image Recognition Technology. PLoS ONE 2016, 11, e168274. [Google Scholar]
- Omrani, E.; Khoshnevisan, B.; Shamshirband, S.; Saboohi, H.; Anuar, N.B.; Nasir, M.H.N.M. Potential of radial basis function-based support vector regression for apple disease detection. Measurement 2014, 55, 512–519. [Google Scholar] [CrossRef]
- Barbedo Arnal, J.G. A new automatic method for disease symptom segmentation in digital photographs of plant leaves. Eur. J. Plant Pathol. 2017, 147, 349–364. [Google Scholar] [CrossRef]
- Springer. SVM-Based Detection of Tomato Leaves Diseases; Springer: Berlin/Heidelberg, Germany, 2015. [Google Scholar]
- Rumpf, T.; Mahlein, A.K.; Steiner, U.; Oerke, E.C.; Dehne, H.W.; Plümer, L. Early detection and classification of plant diseases with Support Vector Machines based on hyperspectral reflectance. Comput. Electron. Agric. 2010, 74, 91–99. [Google Scholar] [CrossRef]
- Hossain, E.; Hossain, M.F.; Rahaman, M.A. A Color and Texture Based Approach for the Detection and Classification of Plant Leaf Disease Using KNN Classifier. In Proceedings of the 2019 International Conference on Electrical, Computer and Communication Engineering (ECCE), Cox’s Bazar, Bangladesh, 7–9 February 2019. [Google Scholar]
- Mohana, R.M.; Reddy CK, K.; Anisha, P.R.; Murthy, B.R. Random forest algorithms for the classification of tree-based ensemble. Mater. Today Proc. 2021. [Google Scholar] [CrossRef]
- Türkoğlu, M.; Hanbay, D. Plant disease and pest detection using deep learning-based features. Turk. J. Electr. Eng. Comput. Sci. 2019, 27, 1636–1651. [Google Scholar] [CrossRef]
- Arivazhagan, S.; Shebiah, R.N.; Ananthi, S.; Varthini, S.V. Detection of unhealthy region of plant leaves and classification of plant leaf diseases using texture features. Agric. Eng. Int. CIGR J. 2013, 15, 211–217. [Google Scholar]
- Jiang, F.; Lu, Y.; Chen, Y.; Cai, D.; Li, G. Image recognition of four rice leaf diseases based on deep learning and support vector machine. Comput. Electron. Agric. 2020, 179, 105824. [Google Scholar] [CrossRef]
- Gao, J.; French, A.P.; Pound, M.P.; He, Y.; Pridmore, T.P.; Pieters, J.G. Deep convolutional neural networks for image-based Convolvulus sepium detection in sugar beet fields. Plant Methods 2020, 16, 29. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Athanikar, G.; Badar, P. Potato leaf diseases detection and classification system. Int. J. Comput. Sci. Mob. Comput. 2016, 5, 76–88. [Google Scholar]
- Yu, J.; Sharpe, S.M.; Schumann, A.W.; Boyd, N.S. Deep learning for image-based weed detection in turfgrass. Eur. J. Agron. 2019, 104, 78–84. [Google Scholar] [CrossRef]
- Bansal, P.; Kumar, R.; Kumar, S. Disease Detection in Apple Leaves Using Deep Convolutional Neural Network. Agriculture 2021, 11, 617. [Google Scholar] [CrossRef]
- Wang, H.; Raj, B. On the origin of deep learning. arXiv 2017, arXiv:1702.07800. [Google Scholar]
- Deng, L.; Yu, D. Deep Learning: Methods and Applications. Found. Trends Signal Process. 2014, 7, 197–387. [Google Scholar] [CrossRef] [Green Version]
- Dyrmann, M.; Karstoft, H.; Midtiby, H.S. Plant species classification using deep convolutional neural network. Biosyst. Eng. 2016, 151, 72–80. [Google Scholar] [CrossRef]
- Kussul, N.; Lavreniuk, M.; Skakun, S.; Shelestov, A. Deep Learning Classification of Land Cover and Crop Types Using Remote Sensing Data. IEEE Geosci. Remote. Sens. Lett. 2017, 14, 778–782. [Google Scholar] [CrossRef]
- Wason, R. Deep learning: Evolution and expansion. Cogn. Syst. Res. 2018, 52, 701–708. [Google Scholar] [CrossRef]
- Geetharamani, G.; Pandian, A. Identification of plant leaf diseases using a nine-layer deep convolutional neural network. Comput. Electr. Eng. 2019, 76, 323–338. [Google Scholar]
- Huang, G.; Liu, Z.; Laurens, V.; Weinberger, K.Q. Densely connected convolutional networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 4700–4708. Available online: https://openaccess.thecvf.com/content_cvpr_2017/papers/Huang_Densely_Connected_Convolutional_CVPR_2017_paper.pdf (accessed on 22 July 2021).
- Too, E.C.; Yujian, L.; Njuki, S.; Yingchun, L. A comparative study of fine-tuning deep learning models for plant disease identification. Comput. Electron. Agric. 2018, 161, 272–279. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Jian, S. Identity Mappings in Deep Residual Networks. In European Conference on Computer Vision; Springer: Cham, Switzerland, 2016. [Google Scholar]
- Guan, S.; Kamona, N.; Loew, M. Segmentation of Thermal Breast Images Using Convolutional and Deconvolutional Neural Networks. In Proceedings of the 2018 IEEE Applied Imagery Pattern Recognition Workshop (AIPR), Washington, DC, USA, 9–11 October 2018. [Google Scholar]
- Fakhry, A.; Zeng, T.; Ji, S. Residual Deconvolutional Networks for Brain Electron Microscopy Image Segmentation. IEEE Trans. Med. Imaging 2017, 36, 447–456. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Wang, Y.; Li, Y.; Fu, J.; Li, J.; Lu, H. Collaborative Deconvolutional Neural Networks for Joint Depth Estimation and Semantic Segmentation. IEEE Trans. Neural Netw. Learn. Syst. 2018, 29, 5655–5666. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Wang, Z.; Tao, D.; See, S.; Wang, G. Learning Common and Specific Features for RGB-D Semantic Segmentation with Deconvolutional Networks. In European Conference on Computer Vision; Springer: Cham, Switzerland, 2016. [Google Scholar]
- Gehlot, S.; Gupta, A.; Gupta, R. SDCT-AuxNet: DCT Augmented Stain Deconvolutional CNN with Auxiliary Classifier for Cancer Diagnosis. Med. Image Anal. 2020, 61, 101661. [Google Scholar] [CrossRef]
- Duggal, R.; Gupta, A.; Gupta, R.; Mallick, P. SD-Layer: Stain Deconvolutional Layer for CNNs in Medical Microscopic Imaging; Springer: Cham, Switzerland, 2017. [Google Scholar]
- Gu, J.; Wang, Z.; Kuen, J.; Ma, L.; Wang, G. Recent Advances in Convolutional Neural Networks. Pattern Recognit. 2015, 77, 354–377. [Google Scholar] [CrossRef] [Green Version]
- Krizhevsky, A.; Sutskever, I.; Hinton, G. ImageNet Classification with Deep Convolutional Neural Networks. Adv. Neural Inf. Process. Syst. 2012, 25, 1097–1105. [Google Scholar] [CrossRef]
- Gao, Z.; Luo, Z.; Zhang, W.; Lv, Z.; Xu, Y. Deep Learning Application in Plant Stress Imaging: A Review. AgriEngineering 2020, 2, 430–446. [Google Scholar] [CrossRef]
- Bengio, Y.; Simard, P.; Frasconi, P. Learning long-term dependencies with gradient descent is difficult. IEEE Trans. Neural Netw. 1994, 5, 157–166. [Google Scholar] [CrossRef] [PubMed]
- Glorot, X.; Bengio, Y. Understanding the difficulty of training deep feedforward neural networks. J. Mach. Learn. Res. 2010, 9, 249–256. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. arXiv 2016, arXiv:1512.03385. [Google Scholar]
- Howard, A.G.; Zhu, M.; Chen, B.; Kalenichenko, D.; Wang, W.; Weyand, T.; Andreetto, M.; Adam, H. MobileNets: Efficient Convolutional Neural Networks for Mobile Vision Applications. arXiv 2017, arXiv:1704.04861. [Google Scholar]
- Tan, M.; Le, Q.V. EfficientNet: Rethinking Model Scaling for Convolutional Neural Networks. arXiv 2019, arXiv:1704.04861. [Google Scholar]
- Parvat, A.; Chavan, J.; Kadam, S.; Dev, S.; Pathak, V. A survey of deep-learning frameworks: 2017 International Conference on Inventive Systems and Control (ICISC). In Proceedings of the 2017 International Conference on Inventive Systems and Control (ICISC), Coimbatore, India, 19–20 January 2017. [Google Scholar]
- Mohanty, S.P.; Hughes, D.P.; Marcel, S. Using Deep Learning for Image-Based Plant Disease Detection. Front. Plant Sci. 2016, 7, 1419. [Google Scholar] [CrossRef] [Green Version]
- Brahimi, M.; Mahmoudi, S.; Boukhalfa, K.; Moussaoui, A. Deep interpretable architecture for plant diseases classification. In Proceedings of the 2019 Signal Processing: Algorithms, Architectures, Arrangements, and Applications (SPA), Poznan, Poland, 18–20 September 2019. [Google Scholar]
- Mishra, S.; Sachan, R.; Rajpal, D. Deep Convolutional Neural Network based Detection System for Real-time Corn Plant Disease Recognition. Procedia Comput. Ence 2020, 167, 2003–2010. [Google Scholar] [CrossRef]
- Darwish, A.; Ezzat, D.; Hassanien, A.E. An optimized model based on convolutional neural networks and orthogonal learning particle swarm optimization algorithm for plant diseases diagnosis. Swarm Evol. Comput. 2019, 52, 100616. [Google Scholar] [CrossRef]
- Amanda, R.; Kelsee, B.; Peter, M.C.; Babuali, A.; James, L.; Hughes, D.P. Deep Learning for Image-Based Cassava Disease Detection. Front. Plant Sci. 2017, 8, 1852. [Google Scholar]
- Lin, Z.; Mu, S.; Shi, A.; Pang, C.; Student, G.; Sun, X.; Student, G. A Novel Method of Maize Leaf Disease Image Identification Based on a Multichannel Convolutional Neural Network. Trans. ASABE 2018, 61, 1461–1474. [Google Scholar] [CrossRef]
- Yuwana, R.S.; Suryawati, E.; Zilvan, V.; Ramdan, A.; Fauziah, F. Multi-Condition Training on Deep Convolutional Neural Networks for Robust Plant Diseases Detection. In Proceedings of the 2019 International Conference on Computer, Control, Informatics and its Applications (IC3INA), Tangerang, Indonesia, 23–24 October 2019. [Google Scholar]
- Chen, J.; Chen, J.; Zhang, D.; Sun, Y.; Nanehkaran, Y.A. Using deep transfer learning for image-based plant disease identification. Comput. Electron. Agric. 2020, 173, 105393. [Google Scholar] [CrossRef]
- Fujita, E.; Kawasaki, Y.; Uga, H.; Kagiwada, S.; Iyatomi, H. Basic investigation on a robust and practical plant diagnostic system. In Proceedings of the 2016 15th IEEE International Conference on Machine Learning and Applications (ICMLA), Anaheim, CA, USA, 18–20 December 2016. [Google Scholar]
- Ghazi, M.M.; Yanikoglu, B.; Aptoula, E. Plant identification using deep neural networks via optimization of transfer learning parameters. Neurocomputing 2017, 235, 228–235. [Google Scholar] [CrossRef]
- Hidayatuloh, A.; Nursalman, M.; Nugraha, E. Identification of Tomato Plant Diseases by Leaf Image Using Squeezenet Model. In Proceedings of the 2018 International Conference on Information Technology Systems and Innovation (ICITSI), Bandung, Indonesia, 22–26 October 2018. [Google Scholar]
- Juncheng, M.; Keming, D.; Feixiang, Z.; Lingxian, Z.; Zhihong, G.; Zhongfu, S. A recognition method for cucumber diseases using leaf symptom images based on deep convolutional neural network. Comput. Electron. Agric. 2018, 154, 18–24. [Google Scholar]
- Bollis, E.; Pedrini, H.; Avila, S. Weakly Supervised Learning Guided by Activation Mapping Applied to a Novel Citrus Pest Benchmark. In Proceedings of the 2020 IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops (CVPRW), Seattle, WA, USA, 14–19 June 2020. [Google Scholar]
- Ge, M.; Su, F.; Zhao, Z.; Su, D. Deep Learning Analysis on Microscopic Imaging in Materials Science. Mater. Today Nano 2020, 11, 100087. [Google Scholar] [CrossRef]
- Brahimi, M.; Boukhalfa, K.; Moussaoui, A. Deep Learning for Tomato Diseases: Classification and Symptoms Visualization. Appl. Artif. Intell. 2017, 31, 1–17. [Google Scholar] [CrossRef]
- Arsenovic, M.; Karanovic, M.; Sladojevic, S.; Anderla, A.; Stefanovic, D. Solving Current Limitations of Deep Learning Based Approaches for Plant Disease Detection. Symmetry 2019, 11, 939. [Google Scholar] [CrossRef] [Green Version]
- Peng, Y.; Wang, Y. An industrial-grade solution for agricultural image classification tasks. Comput. Electron. Agric. 2021, 187, 106253. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, J.; Zhang, W.; Zhan, Y.; Guo, S.; Zheng, Q.; Wang, X. A survey on deploying mobile deep learning applications: A systemic and technical perspective. Digit. Commun. Netw. 2021. [Google Scholar] [CrossRef]
- Gao, J.; Westergaard, J.C.; Sundmark, E.H.R.; Bagge, M.; Liljeroth, E.; Alexandersson, E. Automatic late blight lesion recognition and severity quantification based on field imagery of diverse potato genotypes by deep learning. Knowl. Based Syst. 2021, 214, 106723. [Google Scholar] [CrossRef]
- Fuentes, A.; Yoon, S.; Kim, S.C.; Park, D.S. A Robust Deep-Learning-Based Detector for Real-Time Tomato Plant Diseases and Pests Recognition. Sensors 2017, 17, 2022. [Google Scholar] [CrossRef] [Green Version]
- Barbedo, J.G.A. Factors influencing the use of deep learning for plant disease recognition. Biosyst. Eng. 2018, 172, 84–91. [Google Scholar] [CrossRef]
- Coulibaly, S.; Kamsu-Foguem, B.; Kamissoko, D.; Traore, D. Deep neural networks with transfer learning in millet crop images. Comput. Ind. 2019, 108, 115–120. [Google Scholar] [CrossRef] [Green Version]
- Abdalla, A.; Cen, H.; Wan, L.; Rashid, R.; He, Y. Fine-tuning convolutional neural network with transfer learning for semantic segmentation of ground-level oilseed rape images in a field with high weed pressure. Comput. Electron. Agric. 2019, 167, 105091. [Google Scholar] [CrossRef]
- Thenmozhi, K.; Reddy, U.S. Crop pest classification based on deep convolutional neural network and transfer learning. Comput. Electron. Agric. 2019, 164, 104906. [Google Scholar] [CrossRef]
- Suh, H.K.; IJsselmuiden, J.; Hofstee, J.W.; van Henten, E.J. Transfer learning for the classification of sugar beet and volunteer potato under field conditions. Biosyst. Eng. 2018, 174, 50–65. [Google Scholar] [CrossRef]
- Hendrycks, D.; Mu, N.; Cubuk, E.D.; Zoph, B.; Gilmer, J.; Lakshminarayanan, B. AugMix: A Simple Data Processing Method to Improve Robustness and Uncertainty. arXiv 2019, arXiv:1912.02781. [Google Scholar]
- Ho, D.; Liang, E.; Stoica, I.; Abbeel, P.; Chen, X. Population Based Augmentation: Efficient Learning of Augmentation Policy Schedules. In Proceedings of the International Conference on Machine Learning, Long Beach, CA, USA, 9–15 June 2019; pp. 2731–2741. [Google Scholar]
- Lim, S.; Kim, I.; Kim, T.; Kim, C.; Kim, S. Fast AutoAugment. Adv. Neural Inf. Process. Syst. 2019, 32, 6665–6675. [Google Scholar]
- Cubuk, E.D.; Zoph, B.; Shlens, J.; Le, Q.V. Randaugment: Practical automated data augmentation with a reduced search space. In Proceedings of the 2020 IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops (CVPRW), Seattle, WA, USA, 14–19 June 2020. [Google Scholar]
- Yun, S.; Han, D.; Chun, S.; Oh, S.J.; Yoo, Y.; Choe, J. CutMix: Regularization Strategy to Train Strong Classifiers with Localizable Features. In Proceedings of the IEEE/CVF International Conference on Computer Vision, Seoul, Korea, 27–28 October 2019; pp. 6023–6032. Available online: https://openaccess.thecvf.com/content_ICCV_2019/papers/Yun_CutMix_Regularization_Strategy_to_Train_Strong_Classifiers_With_Localizable_Features_ICCV_2019_paper.pdf (accessed on 22 July 2021).
- Bin, L.; Yun, Z.; Dongjian, H.; Yuxiang, L. Identification of Apple Leaf Diseases Based on Deep Convolutional Neural Networks. Symmetry 2017, 10, 11. [Google Scholar]
- Douarre, C.; Crispim-Junior, C.F.; Gelibert, A.; Tougne, L.; Rousseau, D. Novel data augmentation strategies to boost supervised segmentation of plant disease. Comput. Electron. Agric. 2019, 165, 104967. [Google Scholar] [CrossRef]
- Argüeso, D.; Picon, A.; Irusta, U.; Medela, A.; Alvarez-Gila, A. Few-Shot Learning approach for plant disease classification using images taken in the field. Comput. Electron. Agric. 2020, 175, 105542. [Google Scholar] [CrossRef]
- Arnal Barbedo, J.G. Plant disease identification from individual lesions and spots using deep learning. Biosyst. Eng. 2019, 180, 96–107. [Google Scholar] [CrossRef]
- Srdjan, S.; Marko, A.; Andras, A.; Dubravko, C.; Darko, S. Deep Neural Networks Based Recognition of Plant Diseases by Leaf Image Classification. Comput. Intell. Neurosci. 2016, 2016, 3289801. [Google Scholar]
- Barbedo Arnal, J.G. A review on the main challenges in automatic plant disease identification based on visible range images. Biosyst. Eng. 2016, 144, 52–60. [Google Scholar] [CrossRef]
- Zhang, H.; Tang, Z.; Xie, Y.; Gao, X.; Chen, Q. A watershed segmentation algorithm based on an optimal marker for bubble size measurement. Measurement 2019, 138, 182–193. [Google Scholar] [CrossRef]
- Peressotti, E.; Duchene, E.; Merdinoglu, D.; Mestre, P. A semi-automatic non-destructive method to quantify grapevine downy mildew sporulation. J. Microbiol. Methods 2011, 84, 265–271. [Google Scholar] [CrossRef] [PubMed]
- Clément, A.; Verfaille, T.; Lormel, C.; Jaloux, B. A new colour vision system to quantify automatically foliar discolouration caused by insect pests feeding on leaf cells. Biosyst. Eng. 2015, 133, 128–140. [Google Scholar] [CrossRef]
- Ling, Q.; Yan, J.; Li, F.; Zhang, Y. A background modeling and foreground segmentation approach based on the feedback of moving objects in traffic surveillance systems. Neurocomputing 2014, 133, 32–45. [Google Scholar] [CrossRef]
- Du, H.; Chen, X.; Xi, J. An improved background segmentation algorithm for fringe projection profilometry based on Otsu method. Opt. Commun. 2019, 453, 124206. [Google Scholar] [CrossRef]
- Singh, B.; Toshniwal, D.; Allur, S.K. Shunt connection: An intelligent skipping of contiguous blocks for optimizing MobileNet-V2. Neural Netw. 2019, 118, 192–203. [Google Scholar] [CrossRef]
- Xu, X.; Xu, S.; Jin, L.; Song, E. Characteristic analysis of Otsu threshold and its applications. Pattern Recognit. Lett. 2011, 32, 956–961. [Google Scholar] [CrossRef]
- Gao, L.; Lin, X. Fully automatic segmentation method for medicinal plant leaf images in complex background. Comput. Electron. Agric. 2019, 164, 104924. [Google Scholar] [CrossRef]
- Goodfellow, I.J.; Pouget-Abadie, J.; Mirza, M.; Xu, B.; Warde-Farley, D.; Ozair, S.; Courville, A.; Bengio, Y. Generative Adversarial Networks. Adv. Neural Inf. Process. Syst. 2014, 3, 2672–2680. [Google Scholar]
- Gunduz, H. An efficient dimensionality reduction method using filter-based feature selection and variational autoencoders on Parkinson’s disease classification. Biomed. Signal Process. Control. 2021, 66, 102452. [Google Scholar] [CrossRef]
- Li, Y.; Chao, X. Semi-supervised few-shot learning approach for plant diseases recognition. Plant Methods 2021, 17, 68. [Google Scholar] [CrossRef] [PubMed]
- Anami, B.S.; Malvade, N.N.; Palaiah, S. Deep learning approach for recognition and classification of yield affecting paddy crop stresses using field images. Artif. Intell. Agric. 2020, 4, 12–20. [Google Scholar] [CrossRef]
- Rahman, S.; Wang, L.; Sun, C.; Zhou, L. Deep Learning Based HEp-2 Image Classification: A Comprehensive Review. Med. Image Anal. 2020, 65, 101764. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Zhou, Z.; Tian, L.; Liu, Y.; Luo, X. Brown rice planthopper (Nilaparvata lugens Stal) detection based on deep learning. Precis. Agric. 2020, 21, 1385–1402. [Google Scholar] [CrossRef]
- Shre, K.C. An Approach towards Plant Electrical Signal Based External Stimuli Monitoring System. Ph.D. Thesis, University of Southampton, Southampton, UK, 2017. [Google Scholar]
- Najdenovska, E.; Dutoit, F.; Tran, D.; Plummer, C.; Raileanu, L.E. Classification of Plant Electrophysiology Signals for Detection of Spider Mites Infestation in Tomatoes. Appl. Sci. 2021, 11, 1414. [Google Scholar] [CrossRef]
- Chatterjee, S.K.; Das, S.; Maharatna, K.; Masi, E.; Santopolo, L.; Mancuso, S.; Vitaletti, A. Exploring strategies for classification of external stimuli using statistical features of the plant electrical response. J. R. Soc. Interface 2015, 12, 20141225. [Google Scholar] [CrossRef]
- Chatterjee, S.K.; Malik, O.; Gupta, S. Chemical Sensing Employing Plant Electrical Signal Response-Classification of Stimuli Using Curve Fitting Coefficients as Features. Biosensors 2018, 8, 83. [Google Scholar] [CrossRef] [PubMed] [Green Version]

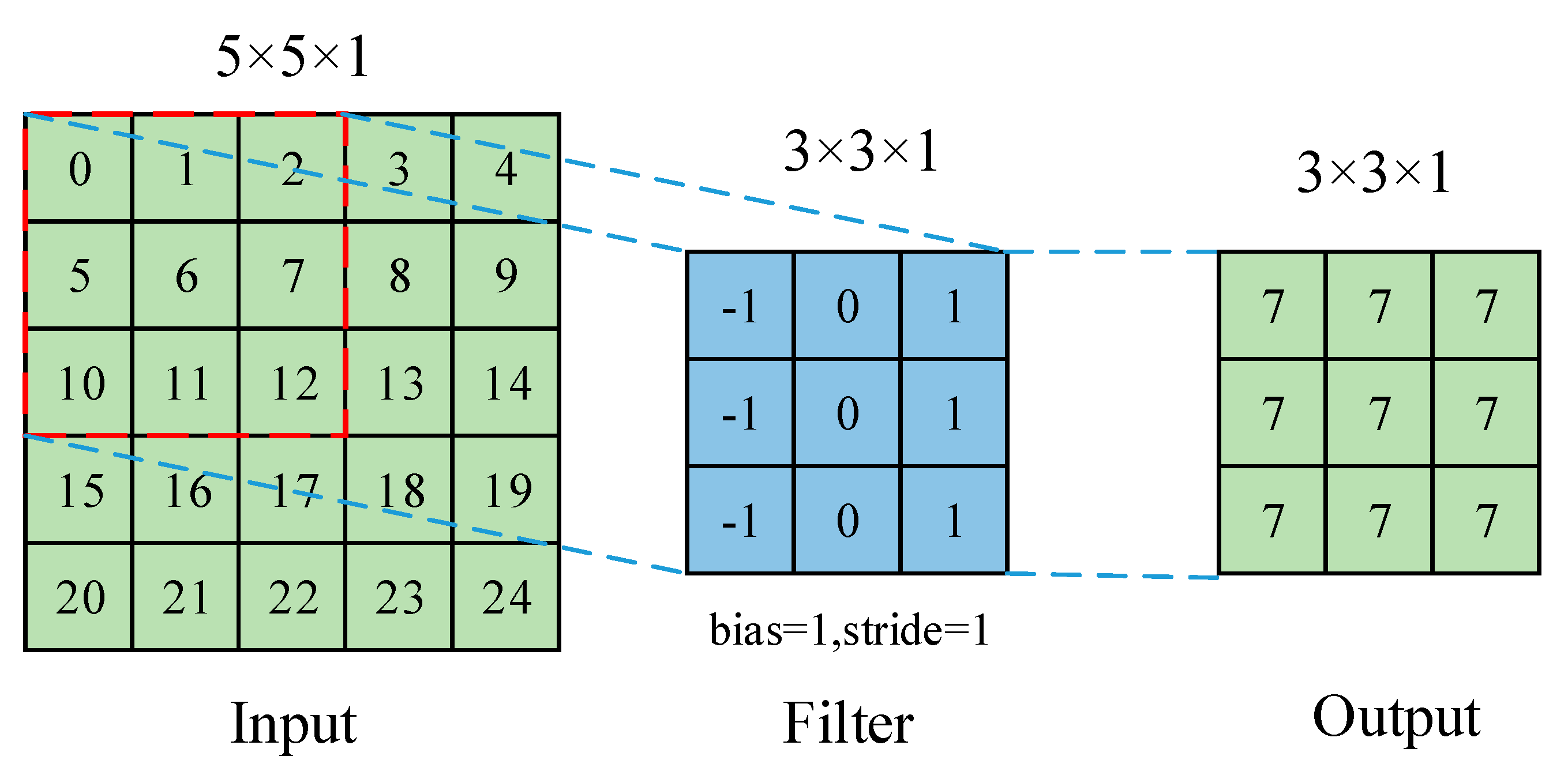
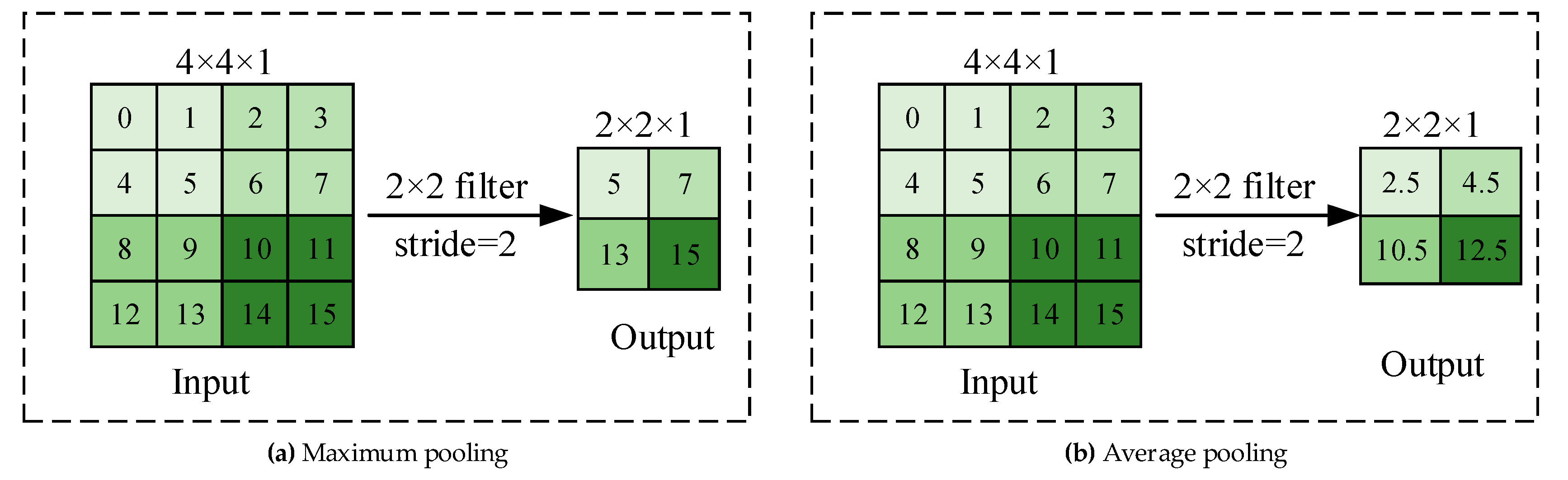
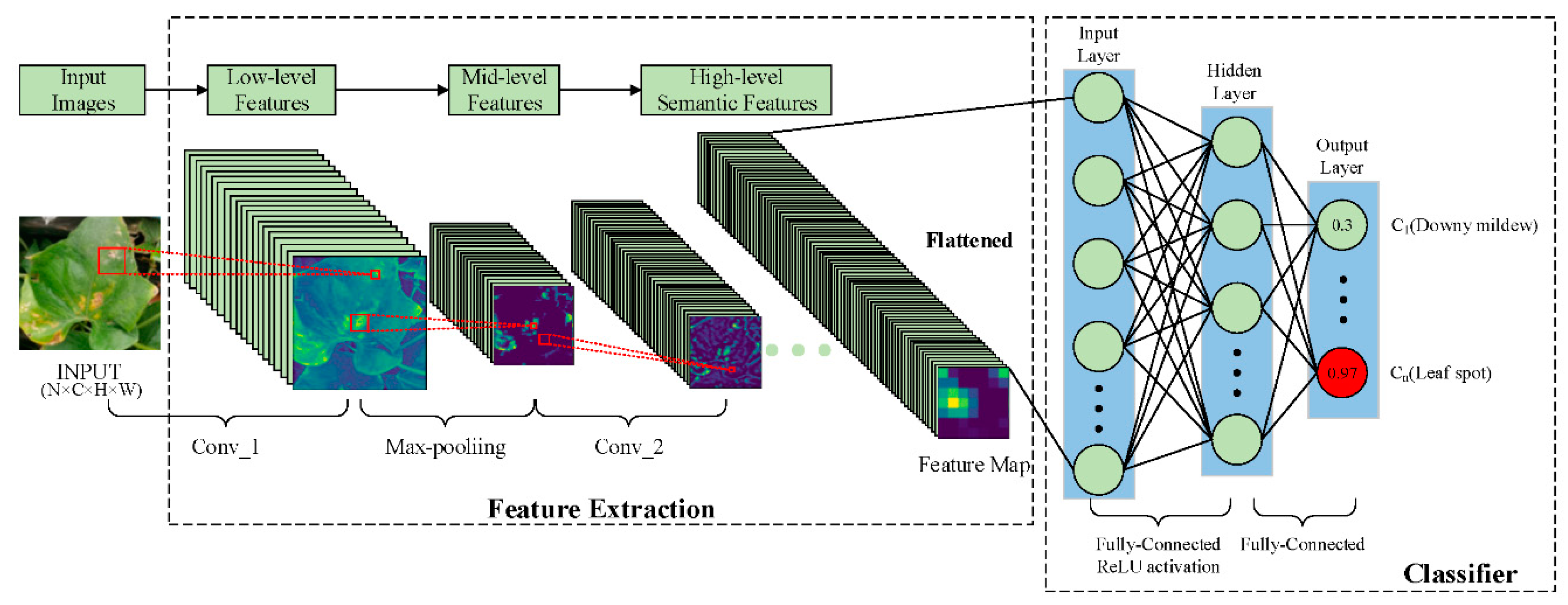
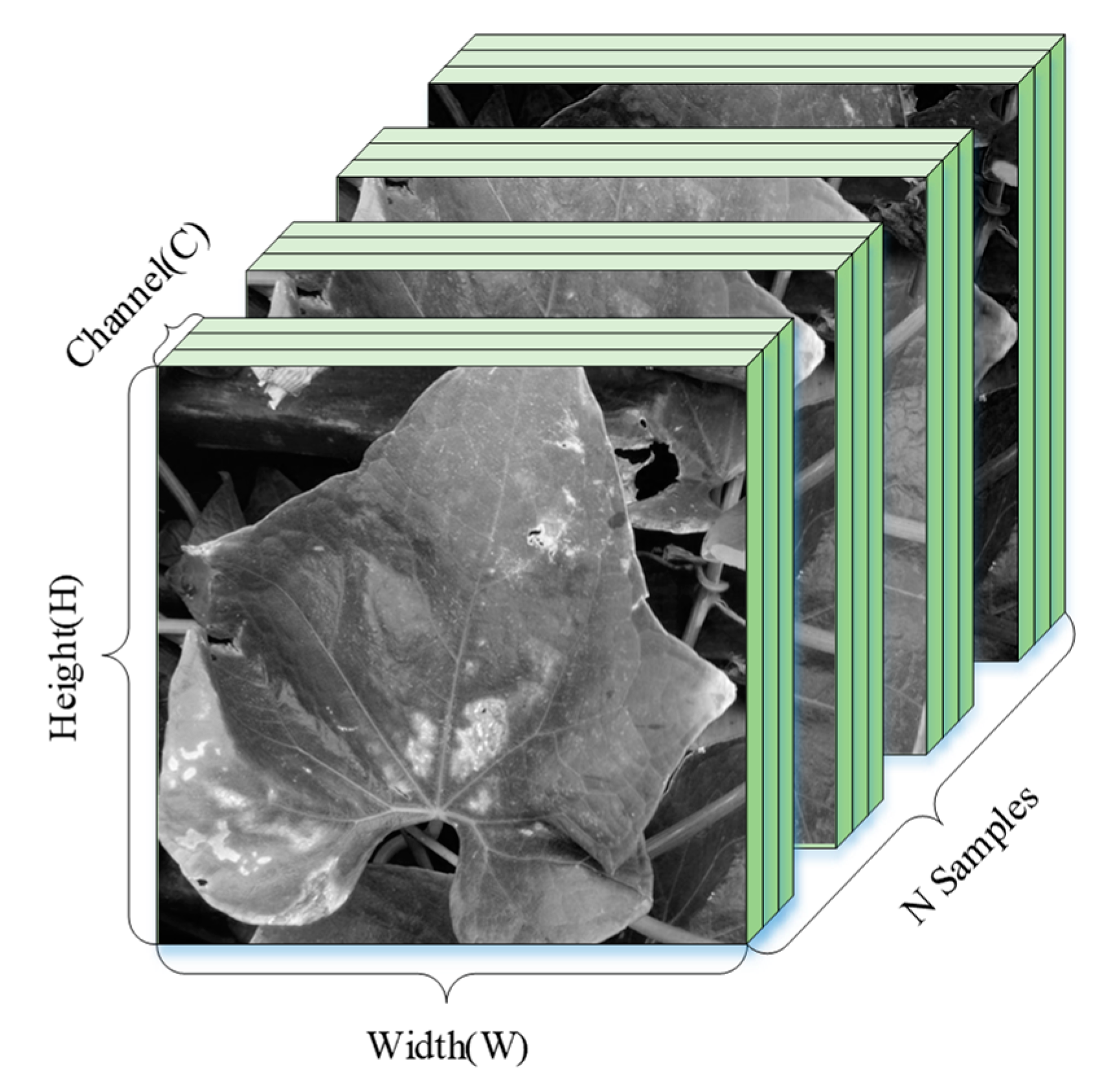

| Plant | Major Types of Disease | Reference | ||
|---|---|---|---|---|
| Fungal | Bacterial | Viral | ||
| Cucumber | Downy mildew, powdery mildew, gray mold, black spot, anthracnose | Angular spot, brown spot, target spot | Mosaic virus, yellow spot virus | Kianat et al. (2021) [4], Zhang et al. (2019) [5], Agarwal et al. (2021) [6] |
| Rice | Rice stripe blight, false smut, rice blast | Bacterial leaf blight, bacterial leaf streak | Rice leaf smut, rice black-streaked dwarf virus | Chen et al. (2021) [7], Shrivastava et al. (2019) [8] |
| Maize | Leaf spot disease, rust disease, gray leaf spot | Bacterial stalk rot, bacterial leaf streak | Rough dwarf disease, crimson leaf disease | Sun et al. (2021) [9], Yu et al. (2014) [10] |
| Tomato | Early blight, late blight, leaf mold | Bacterial wilt, soft rot, canker | Tomato yellow leaf curl virus | Ferentinos (2018) [11], Abbas et al. (2021) [12] |
| Name | Number of Images | Classes | Task | Type of View | Source |
|---|---|---|---|---|---|
| New Plant Diseases Dataset | 87,000 | 38 | Image classification | Field data | Kaggle |
| PlantVillage Dataset | 162,916 | 38 | Image classification | Uniform background | Kaggle |
| Flowers Recognition | 4242 | 4 | Image classification | Field data | Kaggle |
| Plant Seedings Dataset | 5539 | 12 | Target detection | Field data | BIFROST |
| Weed Detection in Soybean Crops | 15,336 | 4 | Target detection | Uniform background | Kaggle |
| Pretrained Model | Dataset | Number of Class | Best Accuracy | Reference |
|---|---|---|---|---|
| ResNet50 | PlantVillage (extended) | 38 | 99.80% | Mukti and Biswas (2019) [1] |
| VGG16 | Millet crop images (own) | 7 | 95.00% | Coulibaly et al. (2019) [74] |
| VGG16 | Plant images (own) | 93.00% | Abdalla et al. (2019) [75] | |
| VGGNet | ImageNet | 9 | 91.83% | Chen et al. (2020) [60] |
| ResNet-101 | NBAIR (extended) | 40 | 95.02% | Thenmozhi et al. (2019) [76] |
| AlexNet | ImageNet (partial) | 2 | 98.00% | Suh et al. (2018) [77] |
| No. | Reference | Task | Dataset | Method | Accuracy | Pros and Cons |
|---|---|---|---|---|---|---|
| 1 | Mohanty et al. (2016) [53] | Identify 14 crop species and 26 diseases | 54,306 images from PlantVillage | AlexNet, GoogLeNet | 99.35% | Not good for practical application |
| 2 | Fuentes et al. (2017) [72] | Detect diseases and pests in tomato plants using images captured in-place by camera devices | 5000 images taken under different conditions and scenarios | VGGNet and Residual Network (ResNet) | 83% (mean) | Lacking number of samples, the precision would be lower in practical application |
| 3 | Chen et al. (2020) [60] | Identify rice and maize leaf diseases | 500 images of rice and 466 images of maize | VGGNet, Inception | 92% | Future works will focus on deploying the module on mobile devices and applying it to more real-world applications |
| 4 | Bin et al. (2017) [83] | Identify four common types of apple leaf diseases (mosaic, rust, brown spot, and Alternaria leaf spot) | 13,689 images of diseased apple leaves | A novel architecture based on AlexNet, image generation technique | 97.62% | The image generation technique proposed in this paper can enhance the robustness of the convolutional neural network model |
| 5 | Brahimi et al. (2017) [67] | Classify nine diseases of tomato leaves | 14,828 images | AlexNet, GoogLeNet | 99.18% | Lacking number of samples |
| 6 | Mishra et al. (2020) [55] | Recognize two corn leaf disease (rust, northern leaf blight) | Some of PlantVillage dataset and some real-time images | DCNN (Deep Convolutional Neural Network) | 88.46% | Only two corn diseases are identified and classified, and the dataset is not enough. |
| 7 | Darwish et al. (2019) [56] | Diagnose three maize plant diseases | 15,408 images from Kaggle | VGG16&19 | 98.2% | The diversity of dataset is not enough |
| 8 | Ferentinos (2018) [11] | Plant disease detection and diagnosis | 87,848 images (PlantVillage) | AlexNetOWTBn, VGG | 99.53% (best) | Obtaining significantly high success rate |
| 9 | Yuwana et al. (2019) [59] | Train more robust deep convolutional neural networks | 5632 images of tea | Multicondition training (MCT), AlexNet, GoogLeNet | None- | Only two segmentation methods (blur with kernel size of 5 and rotation of 40°) were used |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lu, J.; Tan, L.; Jiang, H. Review on Convolutional Neural Network (CNN) Applied to Plant Leaf Disease Classification. Agriculture 2021, 11, 707. https://doi.org/10.3390/agriculture11080707
Lu J, Tan L, Jiang H. Review on Convolutional Neural Network (CNN) Applied to Plant Leaf Disease Classification. Agriculture. 2021; 11(8):707. https://doi.org/10.3390/agriculture11080707
Chicago/Turabian StyleLu, Jinzhu, Lijuan Tan, and Huanyu Jiang. 2021. "Review on Convolutional Neural Network (CNN) Applied to Plant Leaf Disease Classification" Agriculture 11, no. 8: 707. https://doi.org/10.3390/agriculture11080707
APA StyleLu, J., Tan, L., & Jiang, H. (2021). Review on Convolutional Neural Network (CNN) Applied to Plant Leaf Disease Classification. Agriculture, 11(8), 707. https://doi.org/10.3390/agriculture11080707






