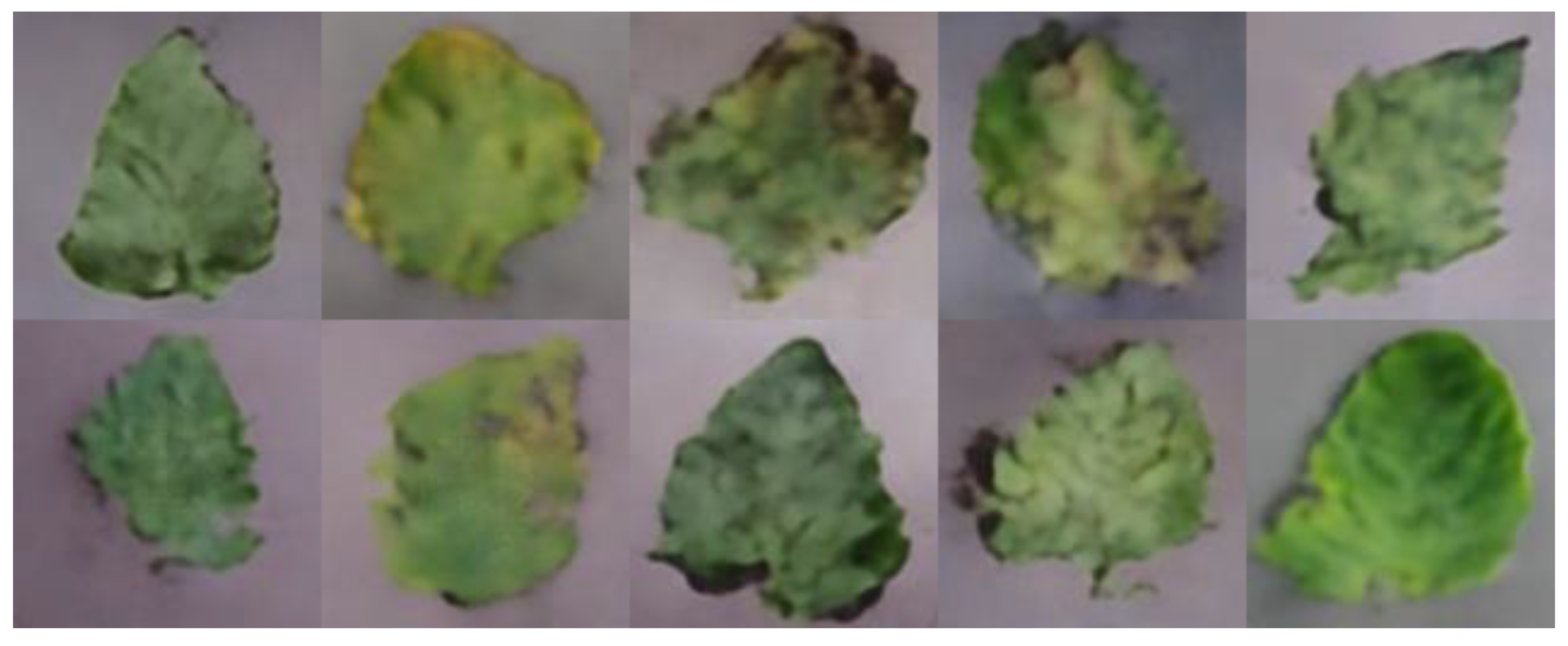
Leaf disease identification is crucial to control the spread of diseases and advance healthy development of the tomato industry. Well-timed and accurate identification of diseases is the key to early treatment, and an important prerequisite for reducing crop loss and pesticide use. Unlike traditional machine learning classification methods that manually select features, deep neural networks provide an end-to-end pipeline to automatically extract robust features, which significantly improve the availability of leaf identification. In recent years, neural network technology has been widely applied in the field of plant leaf disease identification [
1,
2,
3,
4,
5,
6,
7,
8,
9], which indicates that deep learning-based approaches have become popular. However, because the deep convolutional neural network (DCNN) has a lot of adjustable parameters, a large amount of labeled data is needed to train the model to improve its generalization ability of the model. Sufficient training images are an important requirement for models based on convolutional neural networks (CNNs) to improve generalization capability. There are little data about agriculture, especially in the field of leaf disease identification. Collecting large numbers of disease data is a waste of manpower and time, and labeling training data requires specialized domain knowledge, which makes the quantity and variety of labeled samples relatively small. Moreover, manual labeling is a very subjective task, and it is difficult to ensure the accuracy of the labeled data. Therefore, the lack of training samples is the main impediment for further improvement of leaf disease identification accuracy. How to train the deep learning model with a small amount of existing labeled data to improve the identification accuracy is a problem worth studying. In general, researchers usually solve this challenge by using traditional data augmentation methods [
10]. In computer vision, it makes perfect sense to employ data augmentation, which can change the characteristics of a sample based on prior knowledge so that the newly generated sample also conforms to, or nearly conforms to, the true distribution of the data, while maintaining the sample label. Due to the particularity of image data, additional training data can be obtained from the original image through simple geometric transformation. Common data enhancement methods include rotation, scaling, translation, cropping, noise addition, and so on. However, little additional information can be obtained from these methods.
In recent years, data expansion methods based on generative models have become a research hotspot and have been applied in various fields [
11,
12,
13,
14,
15]. For example, in [
11], the author presents an approach for learning to translate an image from a source domain X to a target domain Y in the absence of paired examples to learn a mapping G: X→Y, such that the distribution of images from G(X) is indistinguishable from the distribution Y using an adversarial loss. Usually, the two most common techniques for training generative models are the generative adversarial network (GAN) [
16] and variational auto-encoder (VAE) [
17], both of which have advantages and disadvantages. Goodfellow et al. proposed the GAN model [
16] for latent representation learning based on unsupervised learning. Through the adversarial learning of the generator and discriminator, fake data consistent with the distribution of real data can be obtained. It can overcome many difficulties, which appear in many tricky probability calculations of maximum likelihood estimation and related strategies. However, because the input z of the generator is a continuous noise signal and there are no constraints, GAN cannot use this z, which is not an interpretable representation. Radford et al. [
18] proposed DCGAN, which adds a deep convolutional network based on GAN to generate samples, and uses deep neural networks to extract hidden features and generate data. The model learns the representation from the object to the scene in the generator and discriminator. InfoGAN [
19] tried to use z to find an interpretable expression, where z is broken into incompressible noise z and interpretable implicit variable c. In order to make the correlation between x and c, it is necessary to maximize the mutual information. Based on this, the value function of the original GAN model is modified. By constraining the relationship between c and the generated data, c contains interpreted information about the data. In [
20], Arjovsky et al. proposed Wasserstein GAN (WGAN), which uses the Wasserstein distance instead of Kullback-Leibler divergence to measure the probability distribution, to solve the problem of gradient disappearance, ensure the diversity of generated samples, and balance sensitive gradient loss between the generator and discriminator. Therefore, WGAN does not need to carefully design the network architecture, and the simplest multi-layer fully connected network can do it. In [
17], Kingma et al. proposed a deep learning technique called VAE for learning latent expressions. VAE provides a meaningful lower bound for the log likelihood that is stable during training and during the process of encoding the data into the distribution of the hidden space. However, because the structure of VAE does not clearly learn the goal of generating real samples, it just hopes to generate data that is closest to the real samples, so the generated samples are more ambiguous. In [
21], the researchers proposed a new generative model algorithm named WAE, which minimizes the penalty form of the Wasserstein distance between the model distribution and the target distribution, and derives the regularization matrix different from that of VAE. Experiments show that WAE has many characteristics of VAE, and it generates samples of better quality as measured by FID scores at the same time. Dai et al. [
22] analyzed the reasons for the poor quality of VAE generation and concluded that although it could learn data manifold, the specific distribution in the manifold it learns is different from the real distribution. In the experiment, it shows that VAE can reconstruct training data well, but it cannot generate new samples well. Therefore, a two-stage VAE is proposed, where the first one is used to learn the position of the manifold, and the second is used to learn the specific distribution within the manifold, which improves the generation effect significantly.
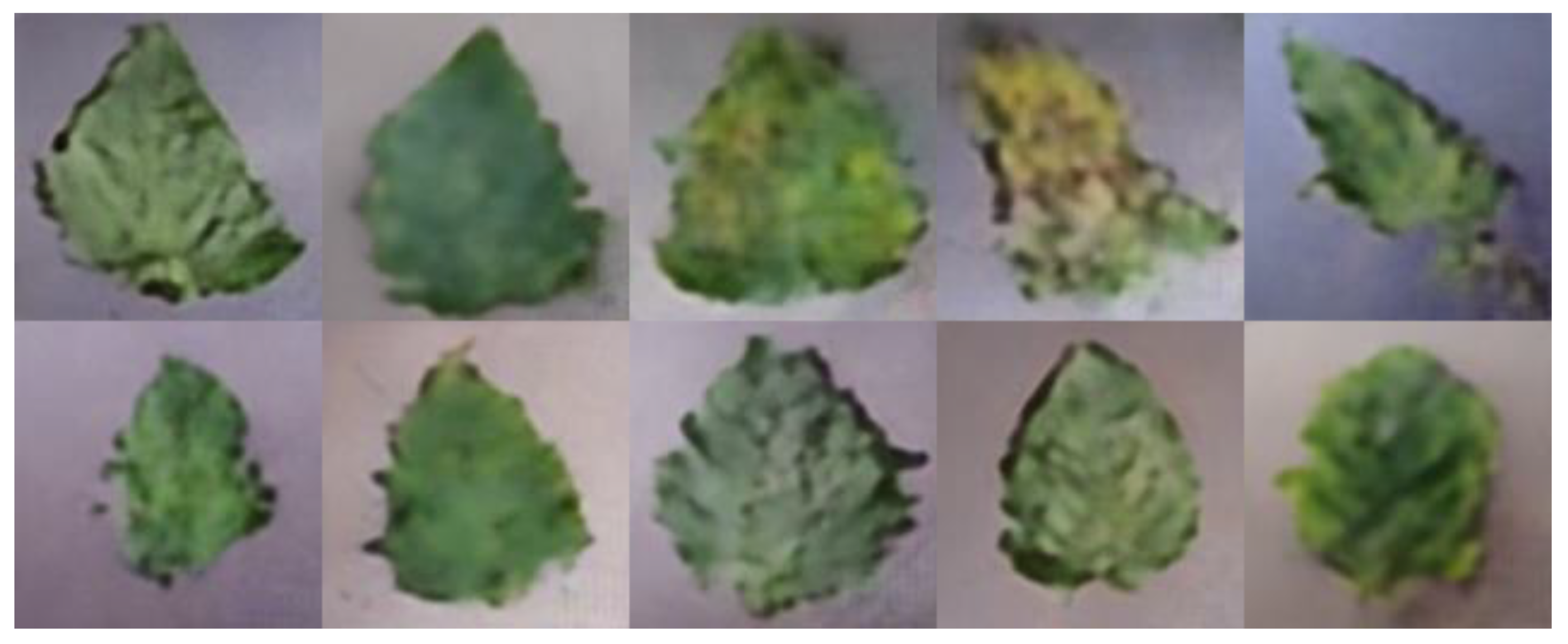
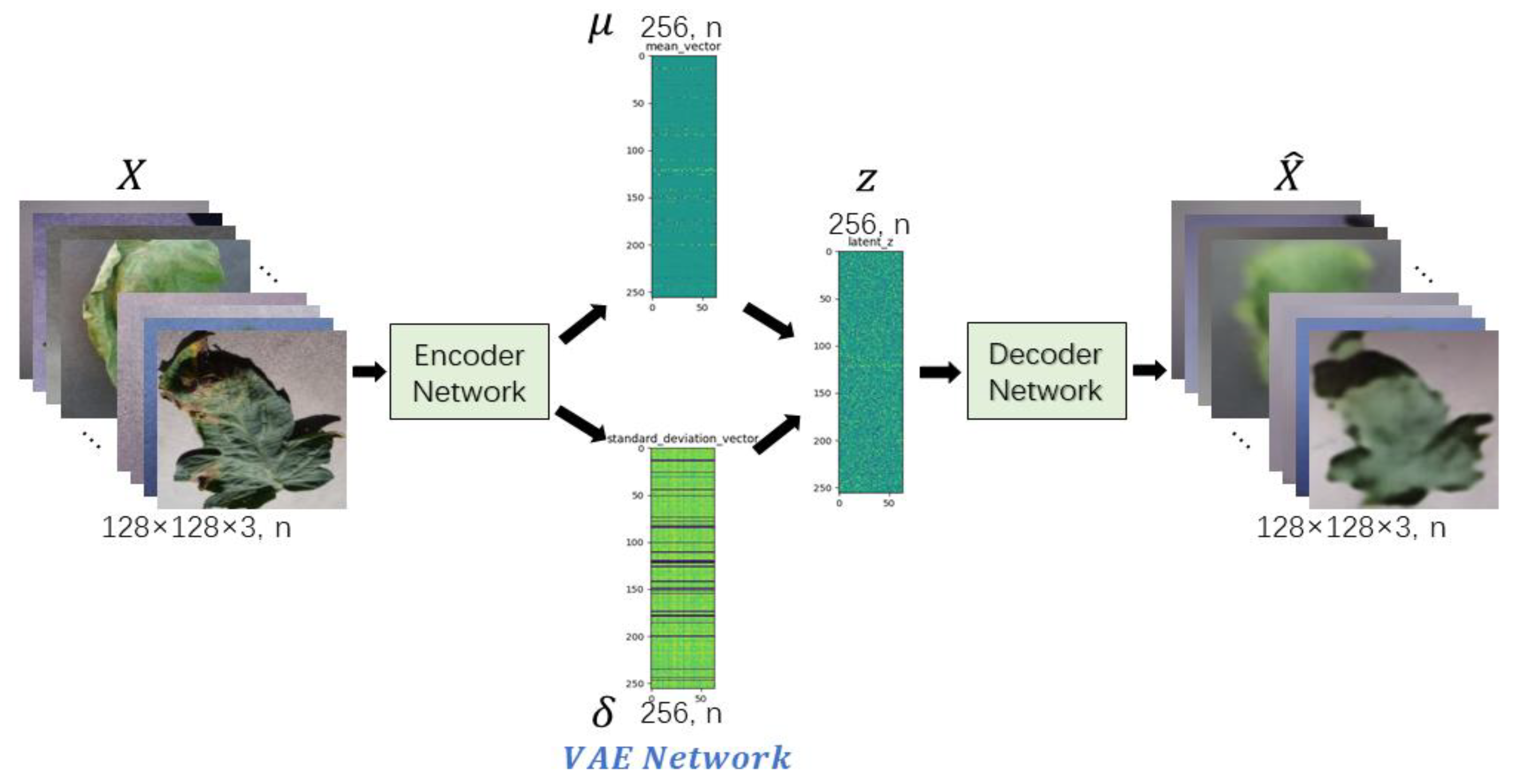
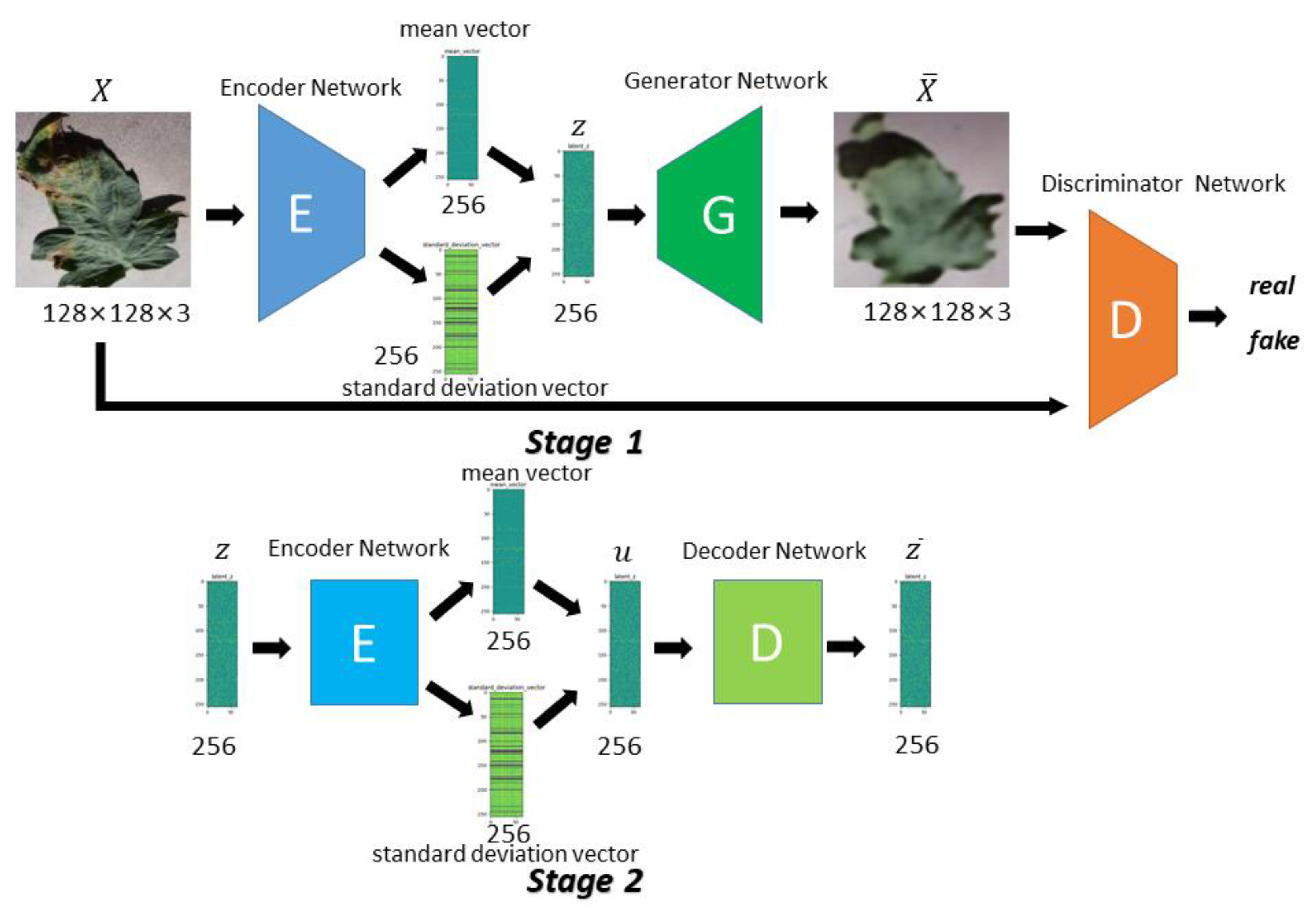
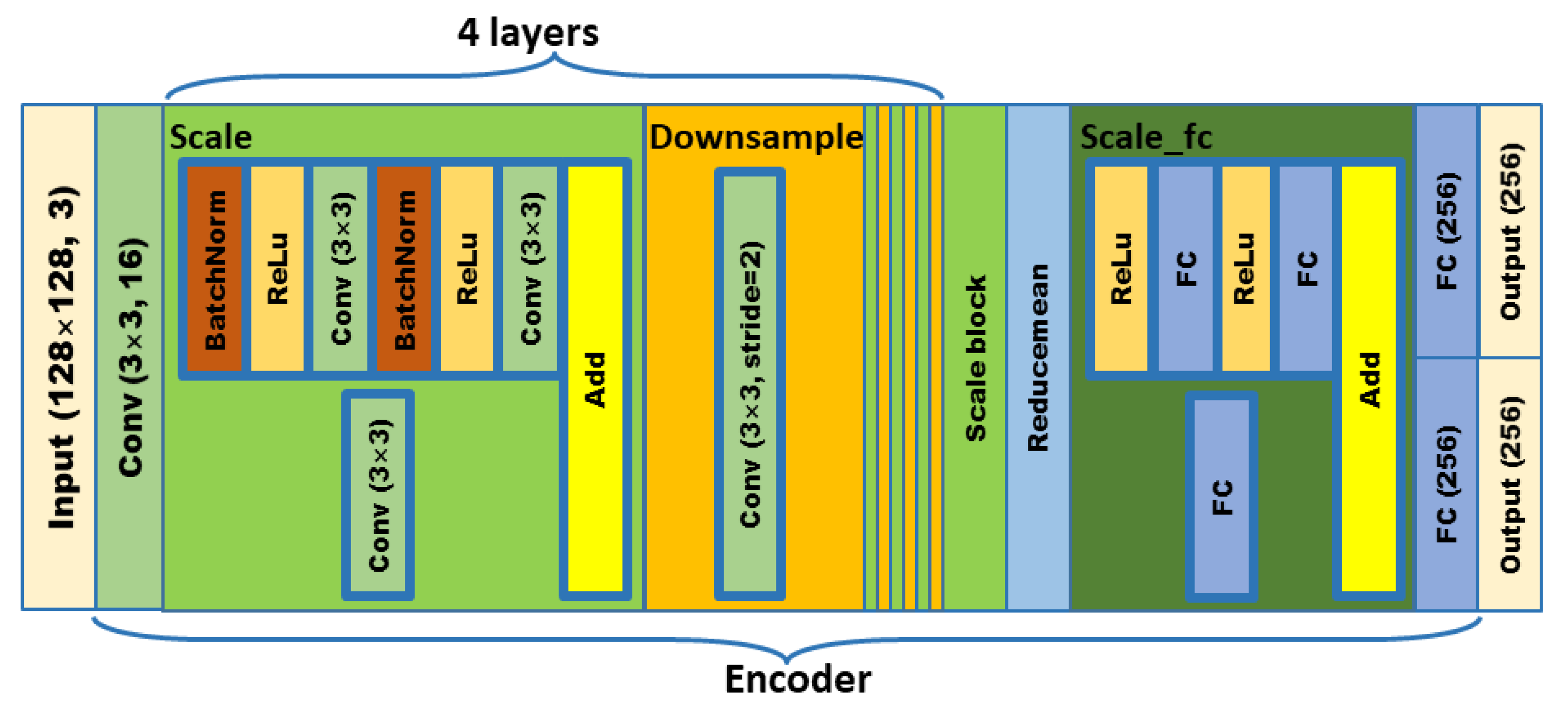
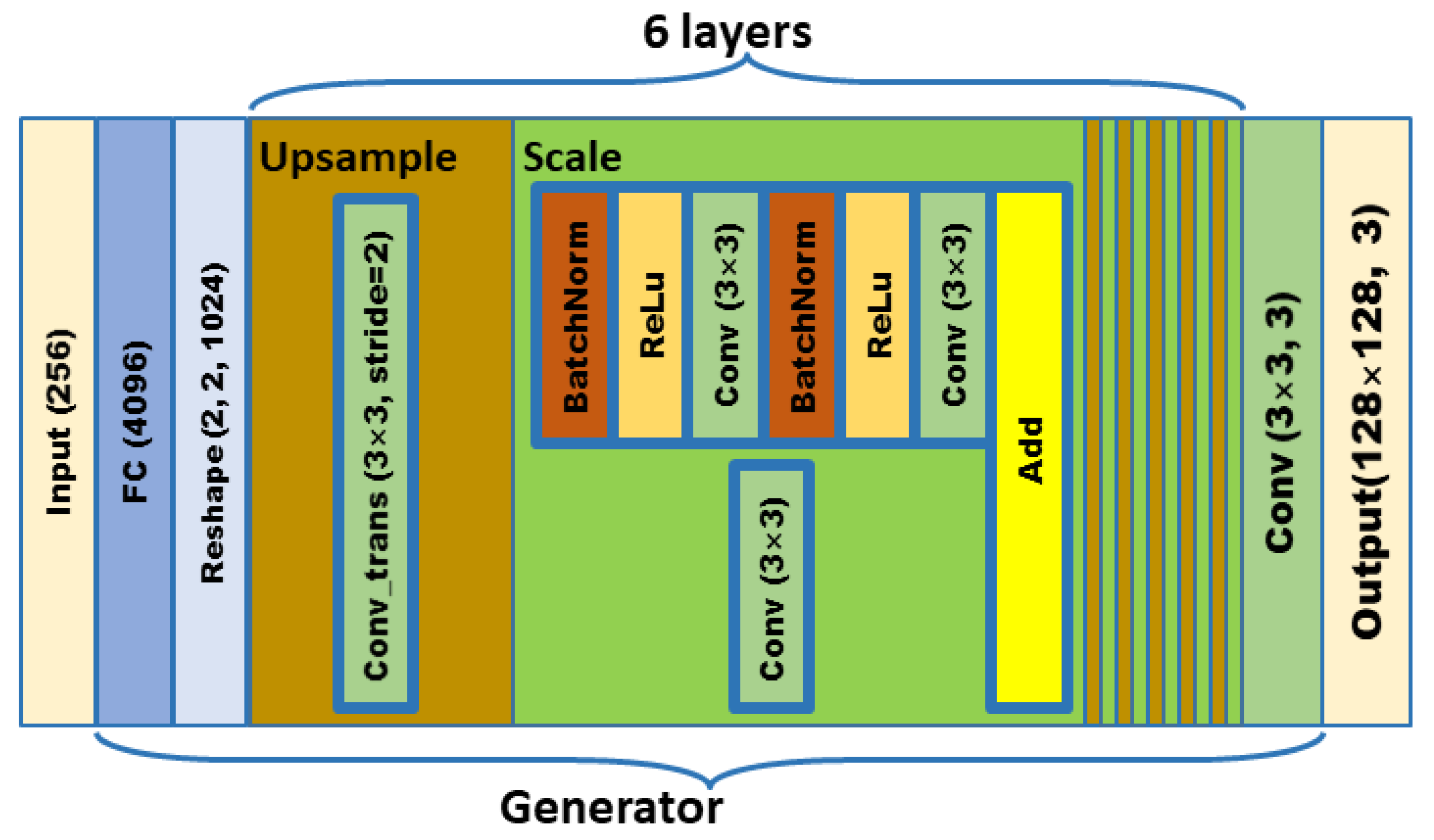
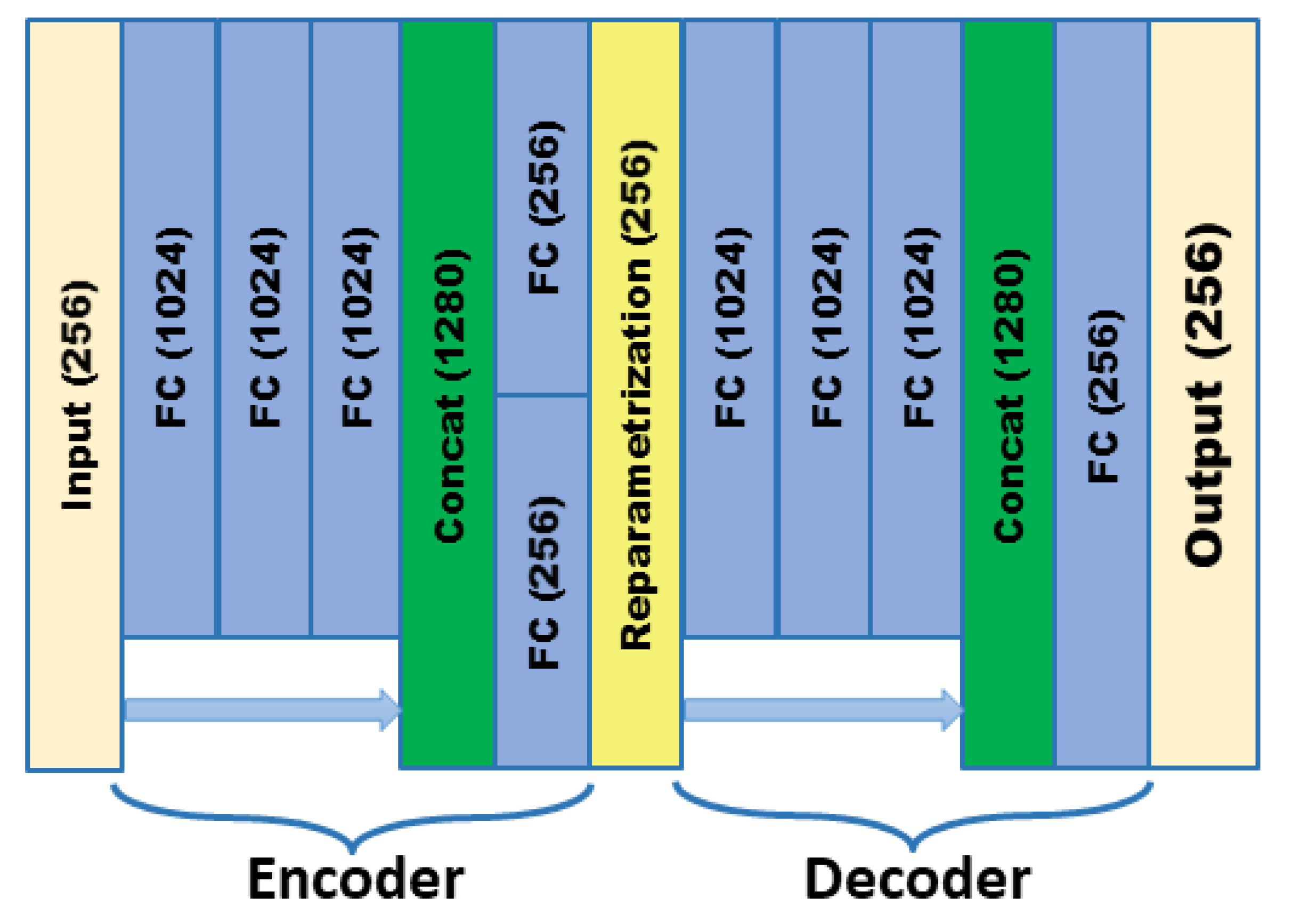
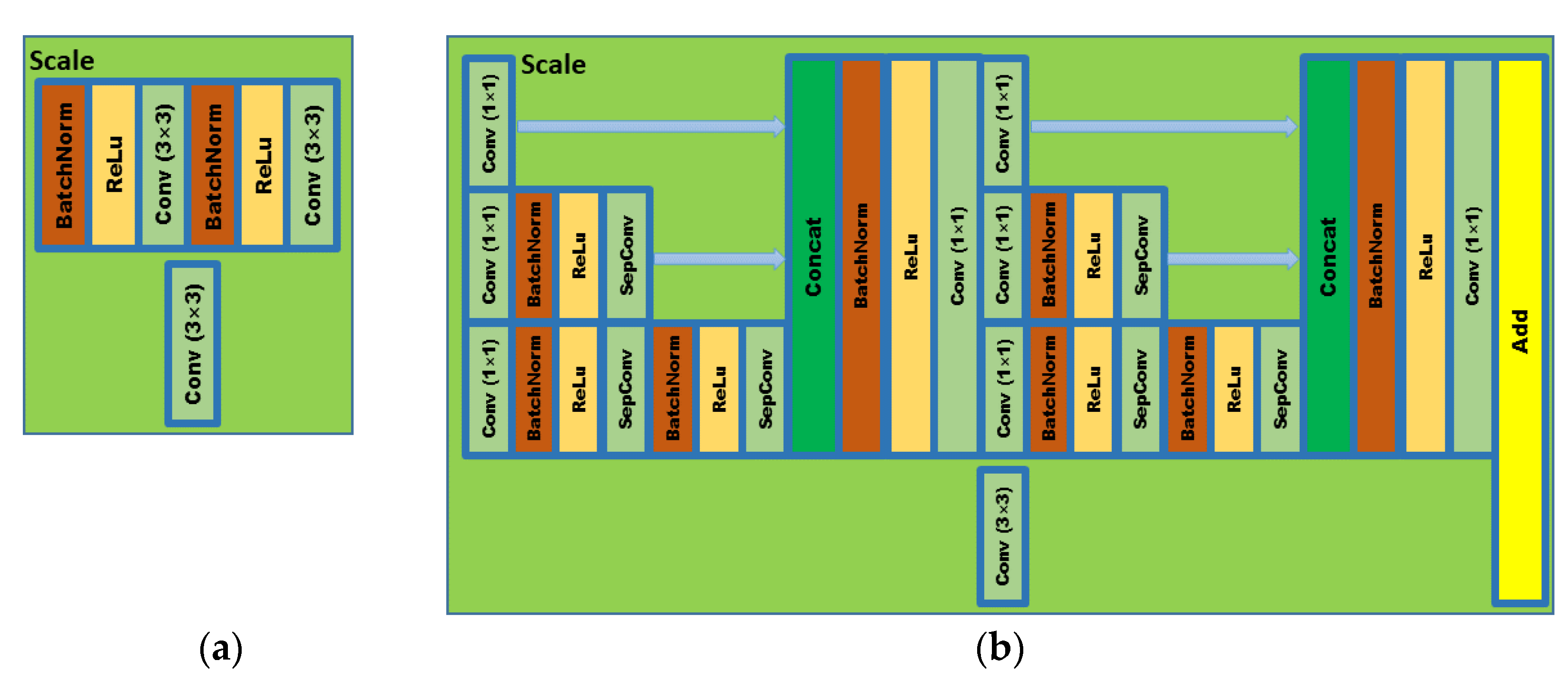
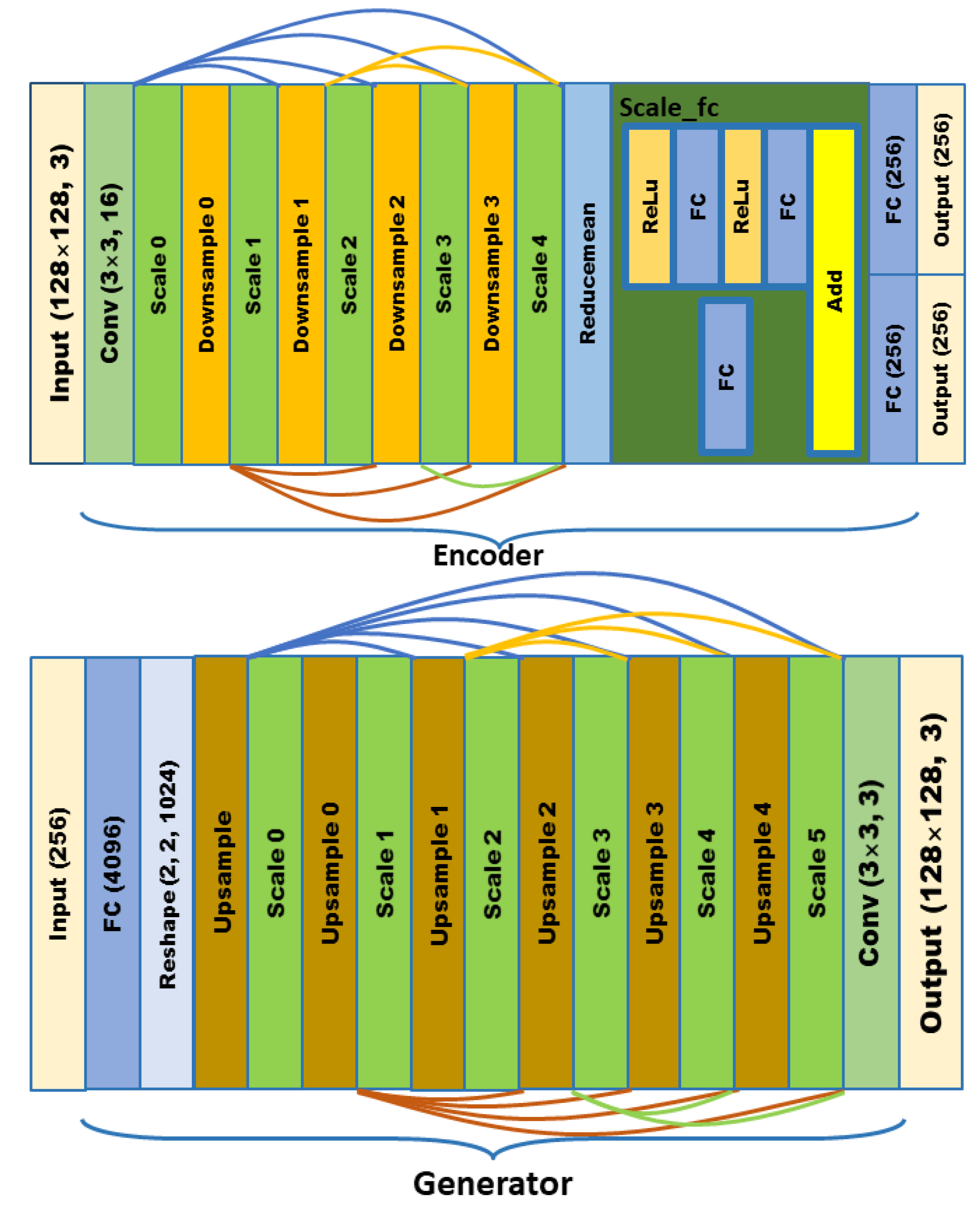
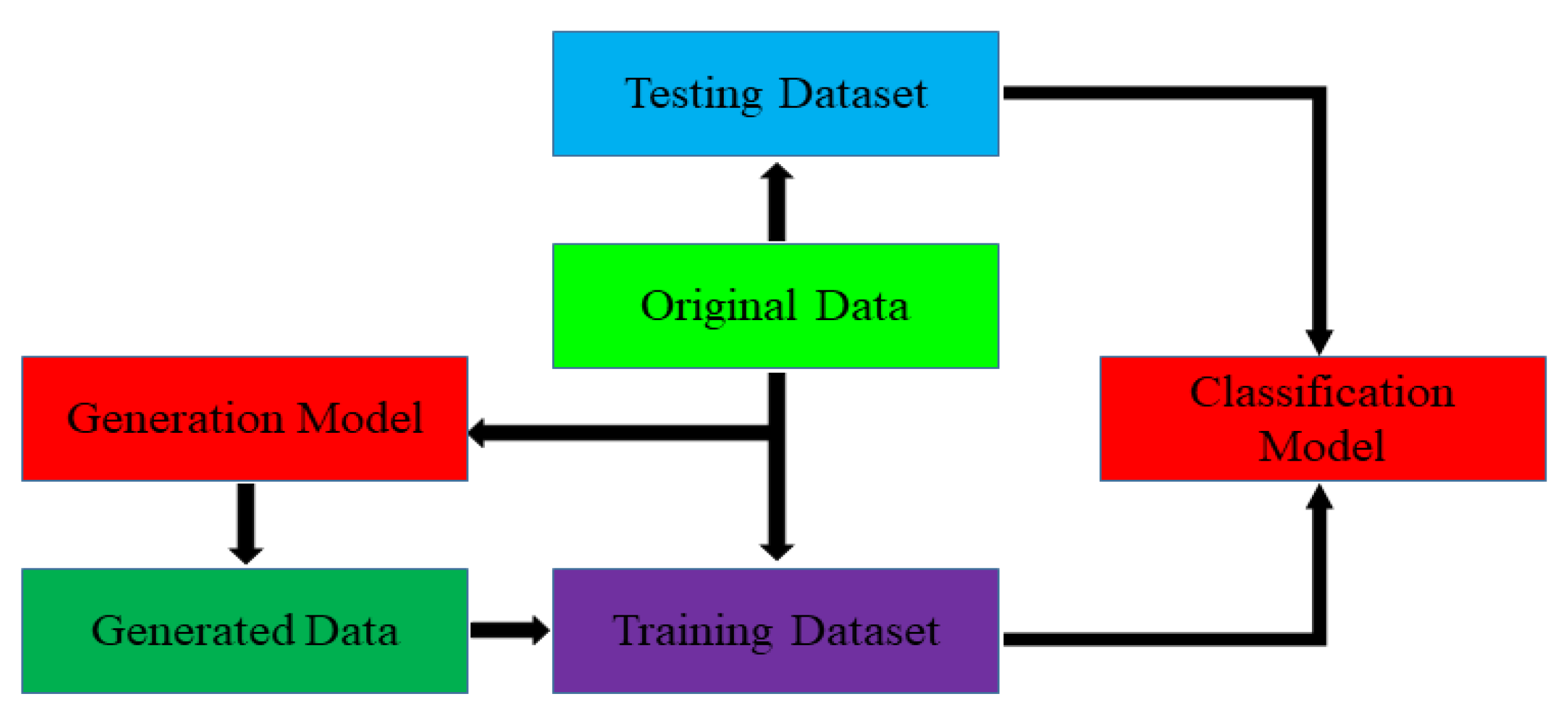
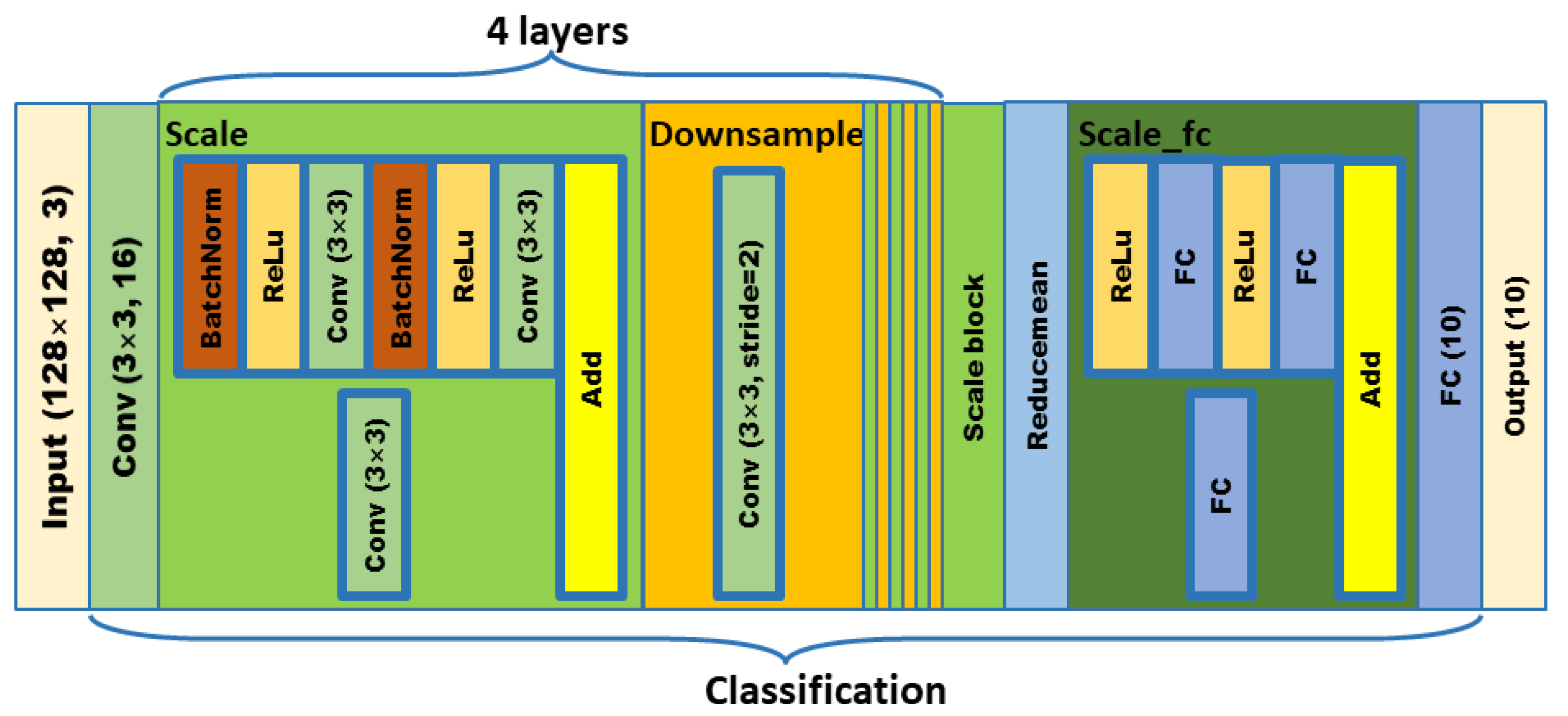
In order to meet the requirements of the training model for the large amount of image data, this paper proposes an image data generation method based on the Adversarial-VAE network model, which expands the image of tomato leaf diseases to generate images of 10 different tomato leaves, overcomes the overfitting problem caused by insufficient training data faced by the identification model. First, the Adversarial-VAE model is designed to generate images of 10 tomato leaves. Then, in view of the obvious differences in the area occupied by the leaves in the dataset and the insufficient accuracy of the feature expression of the diseased leaves using a single-size convolution kernel, the multi-scale residual learning module is used to replace the single-size convolution kernels to enhance the feature extraction ability, and the dense connection strategy is integrated into the Adversarial-VAE model to further enhance the image generative ability. The experimental results show that the tomato leaf disease images generated by Adversarial-VAE have higher quality than InfoGAN, WAE, VAE, and VAE-GAN on the FID. This method provides a solution for data enhancement of tomato leaf disease images and sufficient and high-quality tomato leaf images for different training models, improves the identification accuracy of tomato leaf disease images, and can be used in identifying similar crop leaf diseases.