Potential Rapid Diagnostics, Vaccine and Therapeutics for 2019 Novel Coronavirus (2019-nCoV): A Systematic Review
Abstract
1. Introduction
2. Material and Methods
2.1. Study Identification and Selection
2.2. Supplementary Strategies
3. Results
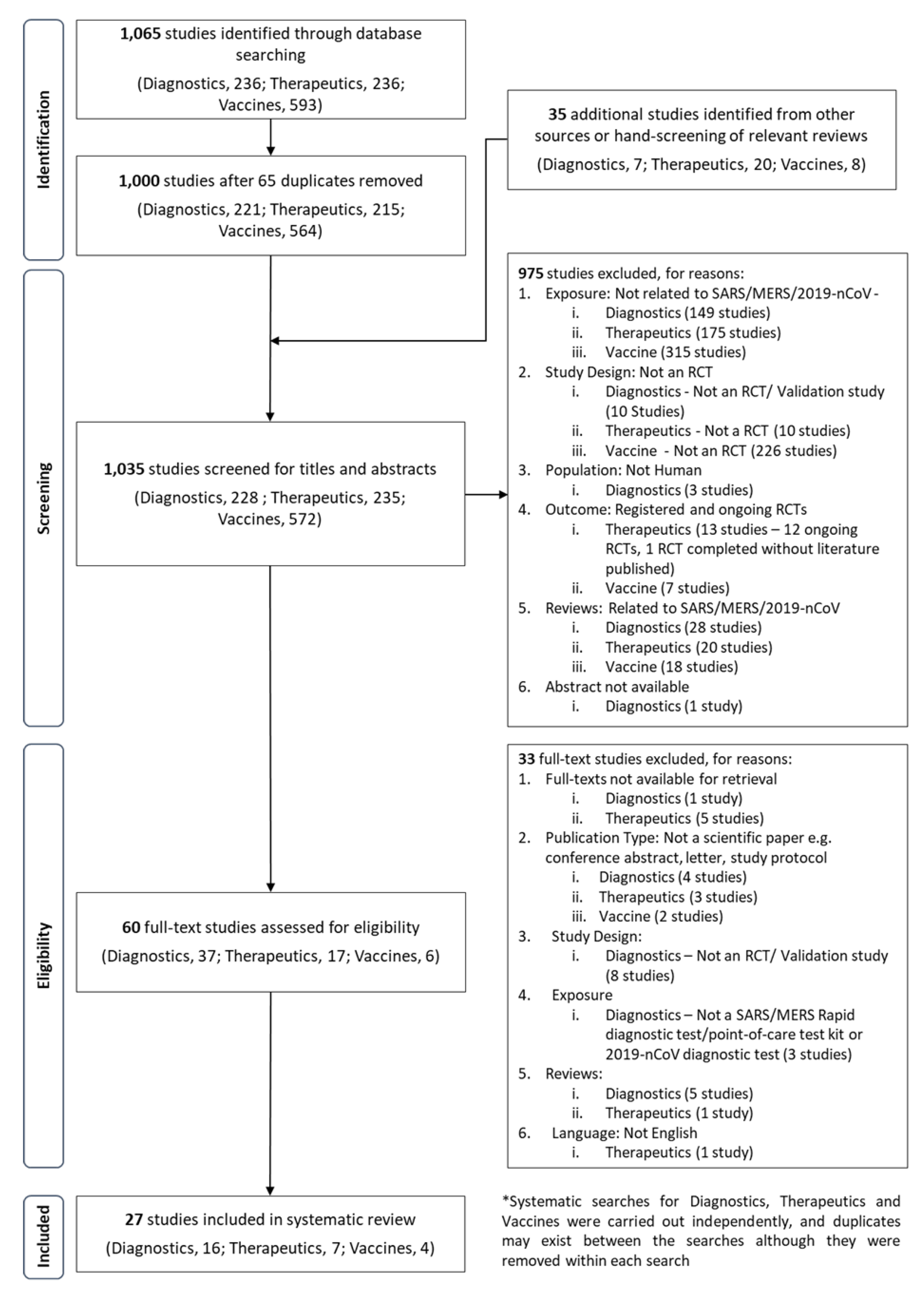
3.1. Search Results
3.2. Diagnostics
3.3. Potential Vaccines
3.4. Potential Therapeutics
4. Discussion
4.1. Rapid Diagnostics
4.1.1. Detection of Genetic Material
4.1.2. Serological Testing
4.2. Vaccines
4.3. Therapeutics
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- World Health Organization. Novel coronavirus (2019-ncov) situation report 15. Available online: https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situation-reports/ (accessed on 5 February 2020).
- Chen, N.; Zhou, M.; Dong, X.; Qu, J.; Gong, F.; Han, Y.; Qiu, Y.; Wang, J.; Liu, Y.; Wei, Y.; et al. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in wuhan, china: A descriptive study. Lancet (Lond. Engl.) 2020. [Google Scholar] [CrossRef]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in wuhan, china. Lancet (Lond. Engl.) 2020, 395, 497–506. [Google Scholar] [CrossRef]
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A novel coronavirus from patients with pneumonia in china, 2019. N. Engl. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef]
- Zaki, A.M.; van Boheemen, S.; Bestebroer, T.M.; Osterhaus, A.D.; Fouchier, R.A. Isolation of a novel coronavirus from a man with pneumonia in saudi arabia. N. Engl. J. Med. 2012, 367, 1814–1820. [Google Scholar] [CrossRef]
- Zhong, N.S.; Zheng, B.J.; Li, Y.M.; Poon, L.L.M.; Xie, Z.H.; Chan, K.H.; Li, P.H.; Tan, S.Y.; Chang, Q.; Xie, J.P.; et al. Epidemiology and cause of severe acute respiratory syndrome (sars) in guangdong, people’s republic of china, in February, 2003. Lancet (Lond. Engl.) 2003, 362, 1353–1358. [Google Scholar] [CrossRef]
- Chen, J. Pathogenicity and transmissibility of 2019-ncov-a quick overview and comparison with other emerging viruses. Microbes Infect. 2020. [Google Scholar] [CrossRef]
- Wu, J.T.; Leung, K.; Leung, G.M. Nowcasting and forecasting the potential domestic and international spread of the 2019-ncov outbreak originating in wuhan, china: A modelling study. Lancet (Lond. Engl.) 2020. [Google Scholar] [CrossRef]
- World Health Organization. Novel coronavirus (2019-ncov) technical guidance: Laboratory testing for 2019-ncov in humans. Available online: https://www.who.int/emergencies/diseases/novel-coronavirus-2019/technical-guidance/laboratory-guidance (accessed on 28 January 2020).
- Charité Virology. Diagnostic detection of wuhan coronavirus 2019 (17 January 2019). Available online: https://www.who.int/docs/default-source/coronaviruse/protocol-v2-1.pdf?sfvrsn=a9ef618c_2 (accessed on 28 January 2020).
- Corman, V.M.; Landt, O.; Kaiser, M.; Molenkamp, R.; Meijer, A.; Chu, D.K.; Bleicker, T.; Brünink, S.; Schneider, J.; Schmidt, M.L.; et al. Detection of 2019 novel coronavirus (2019-ncov) by real-time rt-pcr. Eurosurveillance 2020, 25, 2000045. [Google Scholar] [CrossRef]
- School of Public Health The University of Hong Kong. Detection of 2019 novel coronavirus (2019-ncov) in suspected human cases by rt-pcr. Available online: https://www.who.int/docs/default-source/coronaviruse/peiris-protocol-16-1-20.pdf?sfvrsn=af1aac73_4 (accessed on 28 January 2020).
- Chu, D.K.W.; Pan, Y.; Cheng, S.M.S.; Hui, K.P.Y.; Krishnan, P.; Liu, Y.; Ng, D.Y.M.; Wan, C.K.C.; Yang, P.; Wang, Q.; et al. Molecular diagnosis of a novel coronavirus (2019-ncov) causing an outbreak of pneumonia. Clin. Chem. 2020. [Google Scholar] [CrossRef]
- Chinese Center for Disease Control and Prevention. Specific primers and probes for detection 2019 novel coronavirus. Available online: http://ivdc.chinacdc.cn/kyjz/202001/t20200121_211337.html (accessed on 28 January 2020).
- Department of Medical Sciences Ministry of Public Health Thailand. Conventional rt-pcr for detection of ncov. Available online: https://www.who.int/docs/default-source/coronaviruse/conventional-rt-pcr-followed-by-sequencing-for-detection-of-ncov-rirl-nat-inst-health-t.pdf (accessed on 28 January 2020).
- Nao, N.; Shirato, K.; Katano, H.; Matsuyama, S.; Takeda, M. Detection of second case of 2019-ncov infection in japan (corrected version). Available online: https://www.who.int/docs/default-source/coronaviruse/method-niid-20200123-2.pdf?sfvrsn=fbf75320_7 (accessed on 28 January 2020).
- U.S. Department of Health & Human Services. Real-time rt-pcr panel for detection 2019-novel coronavirus. Available online: https://www.cdc.gov/coronavirus/2019-ncov/downloads/rt-pcr-panel-for-detection-instructions.pdf (accessed on 28 January 2020).
- U.S. Department of Health & Human Services. 2019-novel coronavirus (2019-ncov) real-time rrt-pcr panel primers and probes. Available online: https://www.cdc.gov/coronavirus/2019-ncov/downloads/rt-pcr-panel-primer-probes.pdf (accessed on 28 January 2020).
- Nct. Safety and immunogenicity of a candidate mers-cov vaccine (mers001). 2019. Available online: https://clinicaltrials.gov/ct2/show/NCT03399578?term=vaccine&cond=Mers+CoV&draw=2&rank=2 (accessed on 28 January 2020).
- Kim, M.N.; Ko, Y.J.; Seong, M.W.; Kim, J.S.; Shin, B.M.; Sung, H. Analytical and clinical validation of six commercial middle east respiratory syndrome coronavirus rna detection kits based on real-time reverse-transcription pcr. Ann. Lab. Med. 2016, 36, 450–456. [Google Scholar] [CrossRef][Green Version]
- Shirato, K.; Yano, T.; Senba, S.; Akachi, S.; Kobayashi, T.; Nishinaka, T.; Notomi, T.; Matsuyama, S. Detection of middle east respiratory syndrome coronavirus using reverse transcription loop-mediated isothermal amplification (rt-lamp). Virol. J. 2014, 11, 139. [Google Scholar] [CrossRef] [PubMed]
- Go, Y.Y.; Kim, Y.-S.; Cheon, S.; Nam, S.; Ku, K.B.; Kim, M.; Cho, N.H.; Park, H.; Alison Lee, P.-Y.; Lin, Y.-C.; et al. Evaluation and clinical validation of two field–deployable reverse transcription-insulated isothermal pcr assays for the detection of the middle east respiratory syndrome–coronavirus. J. Mol. Diagn. 2017, 19, 817–827. [Google Scholar] [CrossRef] [PubMed]
- Hashemzadeh, M.S.; Rasouli, R.; Zahraei, B.; Izadi, M.; Tat, M.; Saadat, S.H.; Najarasl, M.; Khansari Nejad, B.; Dorostkar, R. Development of dual taqman based one-step rrt-pcr assay panel for rapid and accurate diagnostic test of mers-cov: A novel human coronavirus, ahead of hajj pilgrimage. Iran Red Crescent Med. J. 2016, 18, e23874. [Google Scholar] [CrossRef][Green Version]
- Lau, S.K.P.; Che, X.-Y.; Woo, P.C.Y.; Wong, B.H.L.; Cheng, V.C.C.; Woo, G.K.S.; Hung, I.F.N.; Poon, R.W.S.; Chan, K.-H.; Peiris, J.S.M.; et al. Sars coronavirus detection methods. Emerg. Infect. Dis. 2005, 11, 1108–1111. [Google Scholar] [CrossRef]
- Lau, L.T.; Fung, Y.-W.W.; Wong, F.P.-F.; Lin, S.S.-W.; Wang, C.R.; Li, H.L.; Dillon, N.; Collins, R.A.; Tam, J.S.-L.; Chan, P.K.S.; et al. A real-time pcr for sars-coronavirus incorporating target gene pre-amplification. Biochem. Biophys. Res. Commun. 2003, 312, 1290–1296. [Google Scholar] [CrossRef]
- Jiang, S.S.; Chen, T.-C.; Yang, J.-Y.; Hsiung, C.A.; Su, I.-J.; Liu, Y.-L.; Chen, P.-C.; Juang, J.-L. Sensitive and quantitative detection of severe acute respiratory syndrome coronavirus infection by real-time nested polymerase chain reaction. Clin. Infect. Dis. 2004, 38, 293–296. [Google Scholar] [CrossRef]
- Wu, H.-S.; Chiu, S.-C.; Tseng, T.-C.; Lin, S.-F.; Lin, J.-H.; Hsu, Y.-H.; Wang, M.-C.; Lin, T.-L.; Yang, W.-Z.; Ferng, T.-L.; et al. Serologic and molecular biologic methods for sars-associated coronavirus infection, taiwan. Emerg. Infect. Dis. 2004, 10, 304–310. [Google Scholar] [CrossRef]
- He, Q.; Chong, K.H.; Chng, H.H.; Leung, B.; Ling, A.E.; Wei, T.; Chan, S.W.; Ooi, E.E.; Kwang, J. Development of a western blot assay for detection of antibodies against coronavirus causing severe acute respiratory syndrome. Clin. Diagn Lab. Immunol. 2004, 11, 417–422. [Google Scholar] [CrossRef]
- Poon, L.L.M.; Chan, K.H.; Wong, O.K.; Yam, W.C.; Yuen, K.Y.; Guan, Y.; Lo, Y.M.D.; Peiris, J.S.M. Early diagnosis of sars coronavirus infection by real time rt-pcr. J. Clin. Virol. 2003, 28, 233–238. [Google Scholar] [CrossRef]
- Guan, M.; Chan, K.H.; Peiris, J.S.; Kwan, S.W.; Lam, S.Y.; Pang, C.M.; Chu, K.W.; Chan, K.M.; Chen, H.Y.; Phuah, E.B.; et al. Evaluation and validation of an enzyme-linked immunosorbent assay and an immunochromatographic test for serological diagnosis of severe acute respiratory syndrome. Clin. Diagn Lab. Immunol. 2004, 11, 699–703. [Google Scholar] [CrossRef][Green Version]
- Hui, R.K.H.; Zeng, F.; Chan, C.M.N.; Yuen, K.Y.; Peiris, J.S.M.; Leung, F.C.C. Reverse transcriptase pcr diagnostic assay for the coronavirus associated with severe acute respiratory syndrome. J. Clin. Microbiol. 2004, 42, 1994–1999. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Mahony, J.B.; Petrich, A.; Louie, L.; Song, X.; Chong, S.; Smieja, M.; Chernesky, M.; Loeb, M.; Richardson, S. Performance and cost evaluation of one commercial and six in-house conventional and real-time reverse transcription-pcr assays for detection of severe acute respiratory syndrome coronavirus. J. Clin. Microbiol. 2004, 42, 1471–1476. [Google Scholar] [CrossRef] [PubMed]
- Liu, I.J.; Chen, P.J.; Yeh, S.H.; Chiang, Y.P.; Huang, L.M.; Chang, M.F.; Chen, S.Y.; Yang, P.C.; Chang, S.C.; Wang, W.K. Immunofluorescence assay for detection of the nucleocapsid antigen of the severe acute respiratory syndrome (sars)-associated coronavirus in cells derived from throat wash samples of patients with sars. J. Clin. Microbiol. 2005, 43, 2444–2448. [Google Scholar] [CrossRef] [PubMed]
- Lin, H.H.; Wang, S.J.; Liu, Y.C.; Lee, S.S.; Hwang, C.K.; Chen, Y.S.; Wann, S.R.; Shih, Y.L. Quantitation of severe acute respiratory syndrome coronavirus genome by real-time polymerase chain reaction assay using minor groove binder DNA probe technology. J. Microbiol. Immunol. Infect. 2004, 37, 258–265. [Google Scholar]
- Genesig. Novel coronavirus strain 2019-ncov. Available online: https://www.genesig.com/products/10037-novel-coronavirus-strain-2019-ncov. (accessed on 30 January 2020).
- Today. Brief-jiangsu bioperfectus technologies develops test kit for new china coronavirus. Available online: https://www.todayonline.com/world/brief-jiangsu-bioperfectus-technologies-develops-test-kit-new-china-coronavirus (accessed on 30 January 2020).
- Co-Diagnostics Inc. Co-diagnostics inc designs test for new coronavirus using coprimer platform. Available online: http://codiagnostics.com/co-diagnostics-designs-new-coronavirus-test-using-coprimers/ (accessed on 30 January 2020).
- Genetic Engineering & Biotechnology News. Coronavirus detection test in the works for wuhan. Available online: https://www.genengnews.com/news/coronavirus-detection-test-in-the-works-for-wuhan/ (accessed on 30 January 2020).
- altona Diagnostics. Altona diagnostics is developing a rt-pcr kit for detection of novel coronavirus (2019-ncov). Available online: https://altona-diagnostics.com/en/news/assay-for-novel-coronavirus-under-development.html (accessed on 28 January 2020).
- BGI. Bgi develops real-time fluorescent rt-pcr kit for detecting the 2019 novel coronavirus. Available online: https://www.bgi.com/global/company/news/bgi-develops-real-time-dna-based-kit-for-detecting-the-2019-novel-coronavirus/ (accessed on 30 January 2020).
- MGI. Bgi responds to novel coronavirus with real-time detection kits, deploys emergency team to wuhan. Available online: https://en.mgitech.cn/news/129/ (accessed on 30 January 2020).
- Ng, R.J. Singapore biotech firm veredus expects to have wuhan virus test by feb 1. Available online: https://www.businesstimes.com.sg/companies-markets/singapore-biotech-firm-veredus-expects-to-have-wuhan-virus-test-by-feb-1 (accessed on 30 January 2020).
- Veredus Laboratories Pte Ltd. Veredus laboratories announces the development of a lab-on-chip for the detection of 3 coronaviruses: Mers-cov, sars-cov and 2019-ncov. Available online: http://vereduslabs.com/wordpress/wp-content/uploads/2020/01/VereCoV-Press-Release-Final.pdf (accessed on 30 January 2020).
- Lin, J.T.; Zhang, J.S.; Su, N.; Xu, J.G.; Wang, N.; Chen, J.T.; Chen, X.; Liu, Y.X.; Gao, H.; Jia, Y.P.; et al. Safety and immunogenicity from a phase i trial of inactivated severe acute respiratory syndrome coronavirus vaccine. Antivir. Ther. 2007, 12, 1107–1113. [Google Scholar]
- Martin, J.E.; Louder, M.K.; Holman, L.A.; Gordon, I.J.; Enama, M.E.; Larkin, B.D.; Andrews, C.A.; Vogel, L.; Koup, R.A.; Roederer, M.; et al. A sars DNA vaccine induces neutralizing antibody and cellular immune responses in healthy adults in a phase i clinical trial. Vaccine 2008, 26, 6338–6343. [Google Scholar] [CrossRef]
- Beigel, J.H.; Voell, J.; Kumar, P.; Raviprakash, K.; Wu, H.; Jiao, J.A.; Sullivan, E.; Luke, T.; Davey, R.T. Safety and tolerability of a novel, polyclonal human anti-mers coronavirus antibody produced from transchromosomic cattle: A phase 1 randomised, double-blind, single-dose-escalation study. Lancet Infect. Dis. 2018, 18, 410–418. [Google Scholar] [CrossRef]
- Modjarrad, K.; Roberts, C.C.; Mills, K.T.; Castellano, A.R.; Paolino, K.; Muthumani, K.; Reuschel, E.L.; Robb, M.L.; Racine, T.; Oh, M.D.; et al. Safety and immunogenicity of an anti-middle east respiratory syndrome coronavirus DNA vaccine: A phase 1, open-label, single-arm, dose-escalation trial. Lancet Infect. Dis. 2019, 19, 1013–1022. [Google Scholar] [CrossRef]
- Nct. Evaluate the safety, tolerability and immunogenicity study of gls-5300 in healthy volunteers. 2019. Available online: https://clinicaltrials.gov/ct2/show/study/NCT03721718?term=vaccine&cond=Mers+CoV&draw=2&rank=7 (accessed on 28 January 2020).
- Nct. A clinical trial to determine the safety and immunogenicity of healthy candidate mers-cov vaccine (mers002). 2019. Available online: https://clinicaltrials.gov/ct2/show/record/NCT04170829 (accessed on 28 January 2020).
- Nct. Study of safety and immunogenicity of bvrs-gamvac-combi. 2019. Available online: https://clinicaltrials.gov/show/NCT04128059 (accessed on 10 February 2020).
- Nct. Study of safety and immunogenicity of bvrs-gamvac. 2019. Available online: https://clinicaltrials.gov/show/NCT04130594 (accessed on 10 February 2020).
- Nct. Randomized, double-blind, placebo-controlled, phase ib study to assess the safety and immunogenicity of mva-mers-s_df-1. 2019. Available online: https://clinicaltrials.gov/show/NCT04119440 (accessed on 10 February 2020).
- ClinicalTrials.gov. Efficacy and safety of hydroxychloroquine for treatment of pneumonia caused by 2019-ncov (hc-ncov). Available online: https://clinicaltrials.gov/ct2/show/NCT04261517 (accessed on 10 February 2020).
- ClinicalTrials.gov. A prospective, randomized controlled clinical study of antiviral therapy in the 2019-ncov pneumonia. Available online: https://clinicaltrials.gov/ct2/show/NCT04255017 (accessed on 10 February 2020).
- ClinicalTrials.gov. A prospective, randomized controlled clinical study of interferon atomization in the 2019-ncov pneumonia. Available online: https://clinicaltrials.gov/ct2/show/NCT04254874 (accessed on 10 February 2020).
- ClinicalTrials.gov. The efficacy of lopinavir plus ritonavir and arbidol against novel coronavirus infection. Available online: https://clinicaltrials.gov/ct2/show/NCT04252885 (accessed on 10 February 2020).
- ClinicalTrials.gov. Mild/moderate 2019-ncov remdesivir rct. Available online: https://clinicaltrials.gov/ct2/show/NCT04252664 (accessed on 10 February 2020).
- ClinicalTrials.gov. Efficacy and safety of darunavir and cobicistat for treatment of pneumonia caused by 2019-ncov. Available online: https://clinicaltrials.gov/ct2/show/NCT04252274 (accessed on 10 February 2020).
- ClinicalTrials.gov. Mesenchymal stem cell treatment for pneumonia patients infected with 2019 novel coronavirus. Available online: https://clinicaltrials.gov/ct2/show/NCT04252118 (accessed on 10 February 2020).
- ClinicalTrials.gov. Treatment and prevention of traditional chinese medicines (tcms) on 2019-ncov infection. Available online: https://clinicaltrials.gov/ct2/show/NCT03301090 (accessed on 10 February 2020).
- ClinicalTrials.gov. Glucocorticoid therapy for novel coronavirus critically ill patients with severe acute respiratory failure (steroids-sari). Available online: https://clinicaltrials.gov/ct2/show/NCT04244591 (accessed on 10 February 2020).
- McKay, B.L.P. Drugmakers rush to develop vaccines against china virus the wall street journal. Available online: https://www.wsj.com/articles/drugmakers-rush-to-develop-vaccines-against-china-virus-11579813026 (accessed on 28 January 2020).
- Inovio Pharmaceuticals Inc. Inovio selected by cepi to develop vaccine against new coronavirus inovio. Available online: http://ir.inovio.com/news-and-media/news/press-release-details/2020/Inovio-Selected-by-CEPI-to-Develop-Vaccine-Against-New-Coronavirus/default.aspx (accessed on 28 January 2020).
- University of Queensland Australia. Race to develop coronavirus vaccine. Available online: https://www.uq.edu.au/news/article/2020/01/race-develop-coronavirus-vaccine (accessed on 28 January 2020).
- Hennessy, J. Australia’s been asked to make a coronavirus vaccine at ‘unprecedented speed’ business insider. Available online: https://www.sciencealert.com/australian-scientists-asked-to-make-coronavirus-vaccine-at-unprecedented-speed (accessed on 28 January 2020).
- Reinicke, C. An obscure biotech stock skyrockets 38% after saying it’s testing a coronavirus antibody (vir). Available online: https://markets.businessinsider.com/news/stocks/vir-biotechnology-stock-price-surges-coronavirus-antibody-tests-crispr-2020-1-1028847912. (accessed on 28 January 2020).
- VIR. Vir biotechnology applying multiple platforms to address public health risk from wuhan coronavirus. Available online: https://investors.vir.bio/news-releases/news-release-details/vir-biotechnology-applying-multiple-platforms-address-public (accessed on 28 January 2020).
- Xinhua. China cdc developing novel coronavirus vaccine xinhua. Available online: http://www.xinhuanet.com/english/2020-01/26/c_138734908.htm (accessed on 28 January 2020).
- Lee, J.-H.Z.W.; Zhou, L. Chinese scientists race to develop vaccine as coronavirus death toll jumps: South china morning post. Available online: https://www.scmp.com/news/china/society/article/3047676/number-coronavirus-cases-china-doubles-spread-rate-accelerates (accessed on 28 January 2020).
- Cheung, E. China coronavirus: Hong kong researchers have already developed vaccine but need time to test it, expert reveals: South china morning post. Available online: https://www.scmp.com/news/hong-kong/health-environment/article/3047956/china-coronavirus-hong-kong-researchers-have (accessed on 28 January 2020).
- China Daily. Novel coronavirus vaccine being developed: China daily. Available online: https://www.chinadaily.com.cn/a/202001/26/WS5e2d1768a310128217273481.html (accessed on 28 January 2020).
- Xinhua. China fast-tracks novel coronavirus vaccine development xinhua. Available online: http://www.xinhuanet.com/english/2020-01/28/c_138739378.htm (accessed on 28 January 2020).
- Berkeley, L., Jr. Us health officials fast-track coronavirus vaccine, hope to start clinical trial in three months: Cnbc markets. Available online: https://www.cnbc.com/2020/01/28/us-fast-tracks-coronavirus-vaccine-hopes-to-start-trial-in-three-months.html (accessed on 28 January 2020).
- Bursztynsky, J. J&j scientific officer ‘pretty confident’ they can create coronavirus vaccine as outbreak widens: Cnbc markets. Available online: https://www.cnbc.com/2020/01/27/jj-pretty-confident-it-can-create-china-coronavirus-vaccine.html (accessed on 28 January 2020).
- The Canadian Press. Saskatchewan lab joins global effort to develop coronavirus vaccine. Available online: https://www.cbc.ca/news/canada/saskatchewan/vido-intervac-working-on-coronavirus-vaccine-1.5439118 (accessed on 28 January 2020).
- GeoVax. Geovax and bravovax (wuhan, china) to collaborate on development of coronavirus vaccine. Available online: https://www.geovax.com/news/geovax-and-bravovax-wuhan-china-to-collaborate-on-development-of-coronavirus-vaccine (accessed on 28 January 2020).
- Clover Biopharmaceuticals. Clover initiates development of recombinant subunit- trimer vaccine for wuhan coronavirus (2019-ncov). Available online: http://www.cloverbiopharma.com/index.php?m=content&c=index&a=show&catid=11&id=40 (accessed on 28 January 2020).
- Precision Vaccinations. Mrna vaccines can induce immune responses for 2019- ncov. Available online: https://www.precisionvaccinations.com/curevac-mrna-platform-specifically-suitable-rapidly-provide-answer-viral-outbreak-situation-novel (accessed on 29 January 2020).
- Usdin, S. J&j developing coronavirus vaccine, at least nine other vaccines under development. Available online: https://www.biocentury.com/bc-extra/company-news/2020-01-27/jj-developing-coronavirus-vaccine-least-nine-other-vaccines-under- (accessed on 29 January 2020).
- Chinese Clinical Trial Registry. A randomized, open-label, blank-controlled trial for the efficacy and safety of lopinavir-ritonavir and interferon-alpha 2b in hospitalization patients with novel coronavirus infection. Available online: http://www.chictr.org.cn/showprojen.aspx?proj=48684 (accessed on 29 January 2020).
- Taylor, N.P. Gilead mulls repositioning failed ebola drug in china virus. Available online: https://www.fiercebiotech.com/biotech/gilead-mulls-repositioning-failed-ebola-drug-china-virus (accessed on 29 January 2020).
- Science Mag. Can an anti-hiv combination or other existing drugs outwit the new coronavirus? Available online: https://www.sciencemag.org/news/2020/01/can-anti-hiv-combination-or-other-existing-drugs-outwit-new-coronavirus (accessed on 29 January 2020).
- Today. Brief-ascletis pharma clarifies media reports on coronavirus treatment. Available online: https://www.todayonline.com/world/brief-ascletis-pharma-clarifies-media-reports-coronavirus-treatment?cid=todayInsideTodaypage (accessed on 29 January 2020).
- Speights, K. 5 biotech stocks to watch with the coronavirus scare. Available online: https://www.fool.com/investing/2020/01/26/what-to-expect-in-pfizers-q3-earnings-update.aspx (accessed on 29 January 2020).
- Pharmaceutical Tech. Coronavirus: Gilead, purdue university explore potential treatments. Available online: https://www.pharmaceutical-technology.com/news/coronavirus-drugs-development/ (accessed on 29 January 2020).
- Chinese Clinical Trial Registry. A prospective comparative study for xue-bi-jing injection in the treatment of pneumonia cause by novel coronavirus infection. Available online: http://www.chictr.org.cn/hvshowproject.aspx?id=21735 (accessed on 7 February 2020).
- Chinese Clinical Trial Registry. Adjunctive corticosteroid therapy for patients with severe 2019-ncov pneumonia: A randomized controlled trial. Available online: http://www.chictr.org.cn/showprojen.aspx?proj=48777 (accessed on 7 February 2020).
- Mak, E. Coronavirus outbreak pushes chinese biotech stocks higher despite market downturn. Available online: https://www.bioworld.com/articles/432890-coronavirus-outbreak-pushes-chinese-biotech-stocks-higher-despite-market-downturn (accessed on 7 February 2020).
- Zou, C. Coronavirus: Chinese researchers claim tcm herbal remedy could ‘inhibit’ 2019-ncov. Available online: https://www.bioworld.com/articles/432838-coronavirus-chinese-researchers-claim-tcm-herbal-remedy-could-inhibit-2019-ncov (accessed on 7 February 2020).
- Chinese Clinical Trial Registry. A randomized, open-label, blank-controlled, multicenter trial for shuang-huang-lian oral solution in the treatment of 2019-ncov pneumonia. Available online: http://www.chictr.org.cn/showprojen.aspx?proj=49051 (accessed on 7 February 2020).
- Chinese Clinical Trial Registry. A prospective, open-label, multiple-center study for the efficacy of chloroquine phosphate in hospitalized patients with 2019-ncov pneumonia. Available online: http://www.chictr.org.cn/showprojen.aspx?proj=49145 (accessed on 7 February 2020).
- ClinicalTrials.gov. A multi-centre, double-blinded, randomized, placebo-controlled trial on the efficacy and safety of lopinavir ritonavir plus ribavirin in the treatment of severe acute respiratory syndrome. Available online: https://clinicaltrials.gov/ct2/show/NCT00578825 (accessed on 10 February 2020).
- Hsu, C.H.; Hwang, K.C.; Chao, C.L.; Chang, S.G.; Ker, C.C.; Chien, L.C.; Ho, M.S.; Lin, J.G.; Chen, Y.M.; Chou, P. The lesson of supplementary treatment with chinese medicine on severe laboratory-confirmed sars patients. Am. J. Chin. Med. 2006, 34, 927–935. [Google Scholar] [CrossRef]
- Qiang, J.; Biao, W.; Rui-lin, Z.; Bao-guo, W.; Li-min, F.; Hai-jian, W.; Yan-nan, L.; Xiong, Y.; Hong-qiang, S.; Ying-min, M.; et al. Clinical controlled study of integrative chinese and western medicine in treating 49 cases of sars. Chin. J. Integr. Med. 2003, 9, 175–180. [Google Scholar] [CrossRef]
- Lee, N.; Allen Chan, K.C.; Hui, D.S.; Ng, E.K.; Wu, A.; Chiu, R.W.; Wong, V.W.; Chan, P.K.; Wong, K.T.; Wong, E.; et al. Effects of early corticosteroid treatment on plasma sars-associated coronavirus rna concentrations in adult patients. J. Clin. Virol. 2004, 31, 304–309. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Wang, R.; Zhang, Y.; Zhang, X.; Layon, A.J.; Li, Y.; Chen, M. Symptom combinations associated with outcome and therapeutic effects in a cohort of cases with sars. Am. J. Chin. Med. 2006, 34, 937–947. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Zhang, F.; Xu, M.; Huang, K.; Zhong, W.; Cai, W.; Yin, Z.; Huang, S.; Deng, Z.; Wei, M.; et al. Description and clinical treatment of an early outbreak of severe acute respiratory syndrome (sars) in guangzhou, pr china. J. Med. Microbiol. 2003, 52, 715–720. [Google Scholar] [CrossRef] [PubMed]
- Chu, C.M.; Cheng, V.C.; Hung, I.F.; Wong, M.M.; Chan, K.H.; Chan, K.S.; Kao, R.Y.; Poon, L.L.; Wong, C.L.; Guan, Y.; et al. Role of lopinavir/ritonavir in the treatment of sars: Initial virological and clinical findings. Thorax 2004, 59, 252–256. [Google Scholar] [CrossRef]
- ClinicalTrials.gov. Mers-cov infection treated with a combination of lopinavir ritonavir and interferon beta-1b. Available online: https://clinicaltrials.gov/ct2/show/NCT02845843 (accessed on 10 February 2020).
- ClinicalTrials.gov. A safety, tolerability, pharmacokinetics and immunogenicity trial of co-administered mers-cov antibodies regn3048 and regn3051. Available online: https://clinicaltrials.gov/ct2/show/NCT03301090 (accessed on 10 February 2020).
- Medical Technology. The importance of diagnostic tests in fighting infectious diseases. Available online: https://www.lifechanginginnovation.org/medtech-facts/importance-diagnostic-tests-fighting-infectious-diseases (accessed on 28 January 2020).
- Kelly-Cirino, C.; Mazzola, L.T.; Chua, A.; Oxenford, C.J.; Van Kerkhove, M.D. An updated roadmap for mers-cov research and product development: Focus on diagnostics. BMJ Glob. Health 2019, 4, e001105. [Google Scholar] [CrossRef]
- Public Health Laboratory Network. Phln guidance on laboratory testing for 2019-ncov 2020. Available online: https://www.health.gov.au/sites/default/files/documents/2020/01/phln-guidance-on-laboratory-testing-for-2019-ncov-phln-guidance-on-laboratory-testing-for-novel-coronavirus-2019-ncov.pdf (accessed on 28 January 2020).
- Zhang, N.; Wang, L.; Deng, X.; Liang, R.; Su, M.; He, C.; Hu, L.; Su, Y.; Ren, J.; Yu, F.; et al. Recent advances in the detection of respiratory virus infection in humans. J. Med. Virol. 2020. [Google Scholar] [CrossRef]
- World Health Organization. Laboratory testing for 2019 novel coronavirus (2019-ncov) in suspected human cases. Available online: https://www.who.int/health-topics/coronavirus/laboratory-diagnostics-for-novel-coronavirus (accessed on 28 January 2020).
- HKU Med. Detection of 2019 novel coronavirus (2019-ncov) in suspected human cases by rt-pcr lks faculty of medicine school of public health. 2020. Available online: https://www.who.int/docs/default-source/coronaviruse/peiris-protocol-16-1-20.pdf?sfvrsn=af1aac73_4. (accessed on 28 January 2020).
- CDC; RVB. Division of viral diseases. Real-time rt-pcr panel for detection 2019- novel coronavirus 2020. Available online: https://www.who.int/docs/default-source/coronaviruse/uscdcrt-pcr-panel-for-detection-instructions.pdf?sfvrsn=3aa07934_2 (accessed on 28 January 2020).
- Al Johani, S.; Hajeer, A.H. Mers-cov diagnosis: An update. J. Infect. Public Health 2016, 9, 216–219. [Google Scholar] [CrossRef]
- CEPI. Cepi to fund three programmes to develop vaccines against the novel coronavirus, ncov-2019. Available online: https://cepi.net/news_cepi/cepi-to-fund-three-programmes-to-develop-vaccines-against-the-novel-coronavirus-ncov-2019/ (accessed on 29 January 2020).
- Yong, C.Y.; Ong, H.K.; Yeap, S.K.; Ho, K.L.; Tan, W.S. Recent advances in the vaccine development against middle east respiratory syndrome-coronavirus. Front. Microbiol. 2019, 10, 1781. [Google Scholar] [CrossRef]
- Inovio’s Positive First-Inhhuman Mers Vaccine Results Published in the Lancet Infectious Diseases; Athena Information Solutions Pvt. Ltd.: Mumbai, India, 2019.
- Lu, R.; Zhao, X.; Li, J.; Niu, P.; Yang, B.; Wu, H.; Wang, W.; Song, H.; Huang, B.; Zhu, N.; et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: Implications for virus origins and receptor binding. Lancet (Lond. Engl.) 2020, 395, 565–574. [Google Scholar] [CrossRef]
- Zhou, P.; Yang, X.L.; Wang, X.G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.R.; Zhu, Y.; Li, B.; Huang, C.L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020. [Google Scholar] [CrossRef]
- de Wit, E.; van Doremalen, N.; Falzarano, D.; Munster, V.J. Sars and mers: Recent insights into emerging coronaviruses. Nat. Rev. Microbiol. 2016, 14, 523–534. [Google Scholar] [CrossRef] [PubMed]
- Cui, J.; Li, F.; Shi, Z.L. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 2019, 17, 181–192. [Google Scholar] [CrossRef] [PubMed]
- Momattin, H.; Al-Ali, A.Y.; Al-Tawfiq, J.A. A systematic review of therapeutic agents for the treatment of the middle east respiratory syndrome coronavirus (mers-cov). Travel Med. Infect. Dis. 2019, 30, 9–18. [Google Scholar] [CrossRef] [PubMed]
- Momattin, H.; Mohammed, K.; Zumla, A.; Memish, Z.A.; Al-Tawfiq, J.A. Therapeutic options for middle east respiratory syndrome coronavirus (mers-cov) – possible lessons from a systematic review of sars-cov therapy. Int. J. Infect. Dis. 2013, 17, e792–e798. [Google Scholar] [CrossRef]
- Wong, S.S.Y.; Yuen, K.-Y. The management of coronavirus infections with particular reference to sars. J. Antimicrob. Chemother. 2008, 62, 437–441. [Google Scholar] [CrossRef]
- Mair-Jenkins, J.; Saavedra-Campos, M.; Baillie, J.K.; Cleary, P.; Khaw, F.-M.; Lim, W.S.; Makki, S.; Rooney, K.D.; Nguyen-Van-Tam, J.S.; Beck, C.R.; et al. The effectiveness of convalescent plasma and hyperimmune immunoglobulin for the treatment of severe acute respiratory infections of viral etiology: A systematic review and exploratory meta-analysis. J. Infect. Dis. 2015, 211, 80–90. [Google Scholar] [CrossRef]
- Johnson, R.F.; Bagci, U.; Keith, L.; Tang, X.; Mollura, D.J.; Zeitlin, L.; Qin, J.; Huzella, L.; Bartos, C.J.; Bohorova, N.; et al. 3b11-n, a monoclonal antibody against mers-cov, reduces lung pathology in rhesus monkeys following intratracheal inoculation of mers-cov jordan-n3/2012. Virology 2016, 490, 49–58. [Google Scholar] [CrossRef]
- Keyaerts, E.; Vijgen, L.; Maes, P.; Neyts, J.; Ranst, M.V. In vitro inhibition of severe acute respiratory syndrome coronavirus by chloroquine. Biochem. Biophys. Res. Commun. 2004, 323, 264–268. [Google Scholar] [CrossRef]
- Wu, C.-J.; Jan, J.-T.; Chen, C.-M.; Hsieh, H.-P.; Hwang, D.-R.; Liu, H.-W.; Liu, C.-Y.; Huang, H.-W.; Chen, S.-C.; Hong, C.-F.; et al. Inhibition of severe acute respiratory syndrome coronavirus replication by niclosamide. Antimicrob. Agents Chemother. 2004, 48, 2693–2696. [Google Scholar] [CrossRef]
- Cinatl, J.; Morgenstern, B.; Bauer, G.; Chandra, P.; Rabenau, H.; Doerr, H.W. Glycyrrhizin, an active component of liquorice roots, and replication of sars-associated coronavirus. Lancet 2003, 361, 2045–2046. [Google Scholar] [CrossRef]
- Chen, F.; Chan, K.H.; Jiang, Y.; Kao, R.Y.T.; Lu, H.T.; Fan, K.W.; Cheng, V.C.C.; Tsui, W.H.W.; Hung, I.F.N.; Lee, T.S.W.; et al. In vitro susceptibility of 10 clinical isolates of sars coronavirus to selected antiviral compounds. J. Clin. Virol. 2004, 31, 69–75. [Google Scholar] [CrossRef] [PubMed]
| S/N | Year | Author (Country) | Test | Population/Samples | Finding | References |
|---|---|---|---|---|---|---|
| 2019-nCoV | ||||||
| 1 | 2020 | Corman et al. (Germany) | rRT-PCR; First-line screening tool: E gene assay; Confirmatory testing: RdRp gene assay. | Respiratory samples were obtained during 2019 from patients hospitalised at Charité medical Centre. Additional samples were selected from biobanks at the Rijksinstituut voor Volksgezondheid en Milieu), Bilthoven, at Erasmus University Medical Center, Rotterdam, at Public Health England, London, and at the University of Hong Kong. | Preliminary finding with SARS-CoV strain Frankfurt-1 virions grown on Vero cells, E gene and RdRp gene assays produced the best result (5.2 and 3.8 copies per reaction at 95% detection probability, respectively) In vitro transcript RNA identical to 2019-nCoV target sequence, 3.9 copies per reaction for the E gene assay and 3.6 copies per reaction for the RdRp assay which were close to the 95% hit rate of 2.9 copies per reaction When tested for cross-reaction with other coronavirus, there was no reactivity with the assays. | [11] |
| MERS-CoV | ||||||
| 2 | 2016 | Kim et al. (Korea) | Six different commercial MERS-CoV RNA detection kits based on rRT-PCR: (i) PowerChek (Kogene Biotech, Korea); (ii) DiaPlexQ (SolGent, Korea); (iii) Anyplex (Seegene, Korea) Screening: envelope gene (upE) Confirmation: ORF1a (iv) AccuPower (Bioneer, Korea) (v) LightMix (Roche Molecular Diagnostics, Switzerland) (vi) UltraFast kits (Nanobiosys, Korea) detect upE and ORF1a simultaneously | 28 nasopharyngeal swabs that were positive for other respiratory viruses were used for specificity and 18 lower respiratory specimens for clinical sensitivity. | All six kits correctly identified 8 of 8 (100%) positive clinical specimens. However, based on the findings from the high inhibition panel, PowerChek and AccuPower were the least sensitive to the presence of PCR inhibition. | [20] |
| 3 | 2014 | Shirato et al. (Japan) | Loopamp RNA Amplification Kit (RT-LAMP; Eiken, Tokyo, Japan) | laboratory isolates MERS-CoV diluted with medium containing pharyngeal swabs obtained from healthy adults | Real-time RT-PCR was able to detect at level as low as 1.6 copies of MERS-CoV RNA while RT-LAMP was also able to detect viral RNA at levels as low as 0.7 copies, showing equivalence with the RT-PCR assay. There was no cross reactivity with other respiratory viruses, for RT-LAMP | [21] |
| 4 | 2017 | Go et al. (Korea) | RT-iiPCR Target upE and ORF1a gene | 55 sequential sputum samples collected from 12 patients infected with MERS-CoV were obtained from the Chungnam National University Hospital. | Overall agreement between RT-iiPCR assays and reference RT-qPCR assays were 98.06% (95% CI, 94.43–100%) and 99.03% (95% CI, 95.88–100%) for ORF1a and upE assays, respectively. | [22] |
| 5 | 2015 | Hashemzadeh et al. (Iran) | A onestep rRT-PCR assay, based on specific TaqMan probes | UpE and ORF1b was synthesized due to the difficulty in acquiring patient sample | The sensitivity obtained for upE was fewer than ten copies of RNA template per reaction and for ORF1b was 50 or fewer copies per reaction. | [23] |
| SARS-CoV | ||||||
| 6 | 2005 | Lau et al. (Hong Kong) | Real time qRT-PCR; Antibody-based capture ELISA | 40 SARS patients hospitalized in HK between March–May 2003 | Sensitivities for qRT-PCR (80% for fecal samples and 25% for urine samples) were higher than those of the polyclonal (50% and 5%) and monoclonal (35% and 8%) antibody-based nucleocapsid antigen capture enzyme-linked immunosorbent assays. | [24] |
| 7 | 2003 | Lau et al. (Hong Kong) | Enhanced real-time fluorescent PCR | 80 suspected or probable SARS cases between 1–3 April 2003 | The limit of detection of the enhanced real-time PCR method was 102-fold higher than the standard real-time PCR assay and 107-fold higher than conventional PCR methods In the clinical aspect, the enhanced real-time PCR method was able to detect 6 cases of SARS-CoV positive samples that were not confirmed by any other assay | [25] |
| 8 | 2004 | Jiang et al. (Taiwan) | Quantitative, real-time, nested polymerase chain reaction (PCR) | 46 patients with suspected or reported SARS April through May 2003 in Taiwan | The single round PCR yielded a minor amplification signal. Nested PCR produce signal without apparent background. The single round RT-PCR detected 15 of 46 positive cases, while the nest real-time PCR detected 17 of 46 cases. | [26] |
| 9 | 2004 | Wu et al. (Taiwan) | Neutralization test, (ELISA), (IFA), and (ICT) | 537 probable SARS patients in Taiwan | With the neutralization test as a reference method, the sensitivity, specificity, positive predictive value, and negative predictive value of the test were ELISA: 98.2%, 98.7%, 98.7%, and 98.4% IFA: 99.1%, 87.8%, 88.1% and 99.1% ICT: 33.6%, 98.2%, 95.7%, and 56.1% RT-PCR: 52.2%, 78.7%, 74.5%, and 58.1%, | [27] |
| 10 | 2003 | He et al. (Singapore) | Western blot assay with N195 protein | 274 clinical sera which were collected from patients suffering from probable or suspected SARS, dengue fever, autoimmune diseases | The specificity and sensitivity of this test were 98.3 and 90.9% and 40 of 44 clinical SARS samples were positive. | [28] |
| 11 | 2003 | Poon et al. (Hong Kong) | Real time quantitative RT-PCR modified RNA extraction method 1b region of SARS-CoVCoV | 50 NPA samples collected from days 1–3 of disease onset from SARS patients in whom SARS CoV infections was subsequently serologically confirmed | From the 50 NPA specimens collected during the first 3 days of illness, the first-generation RT-PCR assay identified 22% positive sample, modification in the RNA extraction method identified 44% positive samples and the combination of the modified RNA extraction method and real-time quantitative PCR technology, identified 80% positive sample. | [29] |
| 12 | 2004 | Guan et al. (Hong Kong) | Specific ELISA Genelabs Diagnostics Pte Ltd. utilizing two recombinant proteins (Gst-N and Gst-U274) | 227 clinical serum specimens collected from SARS patients in Hong Kong between 18 March–24 May 2003 and 385 samples from healthy donors. | For the ELISA, the overall sensitivity was 71.8% and specificity was 99.5% For the immunocromatographic test, overall rate of detection of SARS-associated specimens by the rapid test was 70.5% and its specificity was 97.7%. | [30] |
| 13 | 2003 | Hui et al. (Hong Kong) | RT-PCR | Clinical samples, NPAs (n = 131) and stool specimens (n = 5), provided by the Department of Microbiology, The University of Hong Kong. | PCR amplifying the N gene gave an average of a 26.0% (6.3 to 60.0%) stronger intensity signal than that for the 1b gene additional sensitive molecular marker for the diagnosis of the SARS coronavirus | [31] |
| 14 | 2004 | Mahony et al. (Canada) | Seven RT-PCR assays include (i) a nested assay with BNI outer and inner primers with polB gene; (ii) a two-step, non-nested assay with the BNI outer primers with polB gene; (iii) a two-step, non-nested RT-PCR assay with Cor-p-F2 and Cor-p-R1 with polB gene; (iv) a one-step RT-PCR, with BNI outer primers and polB gene with SYBR Green detection; (v) a two-step assay amplifying and nucleocapsid gene with SYBR Green detection; (vi) a one-step assay with the same nucleocapsid primers and gene with TaqMan probe and (vii) a commercial RealArt HPA CoV RT-PCR assay (Artus). | 68 specimens, including 17 respiratory tract specimens (nasopharyngeal or throat swabs), 29 urine samples, and 22 stool samples, were collected between March–April of 2003 from hospitalized patients with a probable or suspected diagnosis of SARS at Sunnybrook and Women’s College Health Sciences Centre during the Toronto outbreak of SARS. | There is no significant difference in the sensitivity and specificity for the 7 assays Assay 1: 94%; 100% Assay 2: 100%; 94% Assay 3: 94%; 100% Assay 4: 100%; 96% Assay 5: 94%; 100% Assay 6: 83%; 100% Assay 7: 94%; 100% | [32] |
| 15. | 2004 | Liu et al. (Taiwan) | Indirect IFA | Throat wash samples from 17 confirmed SARS adult patients, and 10 healthy controls admitted to the emergency department of the National Taiwan University Hospital between 16 April–1 May 2003. | SARS-CoV was detected in 11 of 17 (65%) samples from SARS patients from day 2 to day 9 of the illness but in none of the 10 samples from healthy controls | [33] |
| 16. | 2004 | Lin et al. (Taiwan) | SARS-CoV real-time PCR assay with a TaqMan minor groove binder probe developed by Applied Biosystems (Foster City, CA, USA). | 228 samples (137 sputum, 53 throat swabs or throat wash, 17 NPS, 19 stool specimen, 2 pleural fluid, 2 urine and 1 serum sample) from 151 patients with atypical pneumonia or symptoms mimicking SARS between 30 April–26 June 26 were recruited, In total, from 151 patients were tested. | • The real time PCR has a threshold sensitivity of 10 genome equivalents per reaction and it has a good reproducibility with the inter-assay coefficients of variation of 1.73 to 2.72%. • 13 specimens from 6 patients were positive with viral load range from 362 to 36,240,000 genome equivalents/mL. The real-time RT-PCR reaction was more sensitive than the nested PCR reaction, as the detection limit for the nested PCR reaction was about 103 genome equivalents in the standard cDNA control. | [34] |
| S/N | Type | Organisation | Date | Test | Sensitivity | Specificity | Availability | Turn Around | Costs | Reference |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | RT-PCR | Genesig (U.K.) | Jan 2020 | RT-PCR Kit MasterMix and q16 reaction tubes included (for genesig® q16); PCR MasterMix Kit (for other instruments) | Sensitive to < 100 copies of target Broad dynamic detection range (>6 logs) | Stated to be high but with no accompanying statistics | Sold for research use only and are not licensed for diagnostic procedures | (no info) | For use with genesig q16 tubes: £5.44/test; For use with others: £4.62/test | [35] |
| 2 | RT-PCR | Bioperfectus Technologies (China) | 14 Jan 2020 | RT-PCR test kit | Not stated | Not stated | Available as scientific research product—does not require registration | (no info) | (no info) | [36] |
| 3 | RT-PCR | Co-Diagnostics (U.S.A.) | 23 Jan 2020 | Commercial Kit RT-PCR kit | Stated to be high but with no accompanying statistics. | Claims with lower false positive1 | By Mar 2020 | (no info) | (no info) | [37,38] |
| 4 | RT-PCR | altona Diagnostics (Germany) | 23 Jan 2020 | Commercial Kit RT-PCR kit | Not stated | Not stated | (no info) | (no info) | (no info) | [39] |
| 5 | RT-PCR | BGI; Pathomics Health (distributor; China) * | 23 Jan 2020 | Fluorescent RT-PCR kit In vitro RT-PCR combining fluorescent probing 1 | Not stated | Not stated | Currently used in hospitals and local disease control centres in China. Potentially in Hong Kong, Taiwan, Brunei, Thailand, Nigeria, South Africa too. | No data stated but described as ‘can issue results in a few hours’. | (no info) | [40] |
| 6 | Combined RT-PCR and meta- genomics detection | BGI; Pathomics Health (distributor; China) * | 23 Jan 2020 | 2019-nCoV PMseq Kit A metagenomics sequencing kit based on combinatorial Probe Anchor Sythesis. Able to detect both known and novel microorganisms, Enabling monitoring of evolution during transmission 2. | Not stated | Not stated | Has been Providing technical support for the scientific clinical prevention and control of the epidemic in Wuhan. | Faster than Fluoresce nt RT- PCR kit. 128 Samples with SE50 in 5 h; 128 samples with PE100 in 22 h, | (no info) | [41] |
| 7 | Microfluidic | Veredus Laboratories (Singapore) | 24 Jan 2020 | enVision (enzyme-assisted nanocomplexes for visual identification of nucleic acids) Lab-on-Chip platform integrating PCR and microarray | Stated to be high but with no accompanying statistics. | Stated to be high but with no accompanying statistics. | (no info) | 2 h | (no info) | [42,43] |
| S/N | Company | Estimated Timeline | Technology | Stage/Funding | Reference |
|---|---|---|---|---|---|
| 1 | Moderna Therapeutics—US National Institute of Allergy and Infectious Diseases | 3 months to early stage (phase 1) clinical trial in US (earliest); much longer for full testing and regulatory approval | Messenger RNA vaccine | Preclinical Awaiting preclinical tests and phase 1 study by NIAID, Funding by CEPI. | [62] |
| 2 | Inovio Pharmaceuticals | Human testing in the next few months | INO-4800-DNA based vaccine (DNA synthesized in lab, does not require actual virus sample) | Preclinical Funding by Coalition for Epidemic Preparedness Innovations (CEPI), up to $9 million | [62,63] |
| 3 | Novavax | 3 months | Nanoparticle vaccine | Preclinical | [62] |
| 4 | University of Queensland | 6 months | Rapid Response Technology, ‘Molecular clamp’ vaccine platform (gene added to viral proteins, misleads body to generate antibodies) | Preclinical Funding by Coalition for Epidemic Preparedness Innovations (CEPI) | [64,65] |
| 5 | Vir Biotechnology | Not available | Anti-coronavirus monoclonal antibodies. Additionally, using “whole-genome CRISPR-based screening capabilities to identify the host receptor for Wuhan coronavirus” | Preclinical | [66,67] |
| 6 | Chinese Centre for Disease Control and Prevention (CDC) | At least 1 month for development, 2–3 years before availability for use | Not available Inactivated virus vaccine (postulated, not verified) | Preclinical; virus successfully isolated, currently selecting strain | [68,69,70,71] |
| 7 | Shanghai East Hospital (Tongji University)—Stermirna Therapeutics | <40 days for manufacture of vaccine samples | mRNA technology | Preclinical | [72] |
| 8 | Johnson & Johnson | 1 year to market | Adenovirus—vectored technology used for Ebola vaccine (and Zika and HIV vaccine candidates) | Preclinical | [73,74] |
| 9 | University of Hong Kong | Months for animal testing, At least 1 year for clinical trials on humans | Modified nasal spray influenza vaccine (with surface antigen of coronavirus) prevents both influenza and corona virus | Preclinical; vaccine developed | [70] |
| 10 | University of Saskatchewan (VIDO-InterVac) | Target for animal testing in 6–8 weeks, human trials in at least a year | Not available | Preclinical | [75] |
| 11 | GeoVax—BravoVax | Not available | Modified Vaccina Ankara—Virus Like Particles (MVA-VLP) vaccine platform | Preclinical | [76] |
| 12 | Clover Biopharmaceuticals | Not available | Highly purified recombinant 2019-nCoV S protein subunit-trimer vaccine (S-Trimer), produced using Trimer-Tag© technology | Preclinical | [77] |
| 13 | CureVac | Not available | mRNA technology | Preclinical | [78] |
| 14 | Texas Children’s Hospital Center for Vaccine Development at Baylor College of Medicine | Not available | Not available | Not available | [79] |
| 15 | Codagenix | Not available | Not available | Not available | [79] |
| S/N | Organisation; Candidate | Country of Study; Trial Type; Study Design; Study Details | Population | Outcome (Safety) | Outcome (Efficacy) | Reference | |
|---|---|---|---|---|---|---|---|
| SARS-CoV | |||||||
| 1 | Sinovac Biotech Co. Ltd.; Inactivacted SARS-CoV (ISCV) | China (Beijing); Phase I clinical trial; Randomised, double-blind and placebo controlled; 2 doses of 16 SARS-CoV units (SU) or 32 SU ISCV or placebo control vaccine, intramuscular injection of vaccines in deltoid muscle, doses were 28 days apart | 36 healthy adults between 21 and 40 years old, free of chronic diseases, immunosuppression SARS-CoV, HCV and HIV; 12 subjects were included in each intervention arm | No severe adverse reaction (grade 3) was reported. All local adverse events were mild and resolved within 47–72 h, while systemic adverse events were reported sporadically from all 3 groups and resolved within 24 h. | Seroconversion reached 100% for both vaccine groups on day 42, persisted at 100% in the group receiving 16 SU but decreased to 91.1% for the group receiving 32 SU on day 56. Geometric mean titres (GMT) of specific SARS-CoV neutralising antibody peaked 2 weeks after the second dose, but started to drop 4 weeks later (values not reported). Seroconversion and GMT of neutralising antibody levels were lower in subjects between 21–30 years old compared to those in the elder group, but without significant differences (35: seroconversion p = 0.444; GMT P = 0.528) on days 35 and 42. | [44] | |
| 2 | National Institutes of Health, National Institute of Allergy and Infectious Diseases, Vaccine Research Center; VRC-SRSDNA015-00-VP | United States (Maryland) Phase I clinical trial; Open-label study; 3 doses of vaccine (4 mg/dose), intramuscular injection into lateral deltoid muscle via the Biojector 2000® Needle-Free Injection Management System™ on study days 0, 38 and 56 | 10 subjects between 21 and 29 years old (mean age 35.5) with mean BMI of 24.6 (range 19.7 to 33.9) and were Caucasians (90%) or Asian (10%); only 9 subjects completed all 3 doses | No severe adverse reaction (grade 3), but at least 50% subjects reported at least one mild systemic symptom following vaccination. | SARS specific antibody was detected by ELISA in 8 of 10 (80%) of subjects at one or more timepoints. The neutralizing antibody response was detected in all subjects who received 3 doses of vaccine and peaked between week 8 and 12, with 6 subjects remaining positive at week 32. SARS-CoV-specific CD4+ T cell responses were detected in all vaccines between week 2 and 32, and CD8+ T cell responses in ∼20% of individuals by ICS. The peak T cell response occurred between week 8 and 12 and when present, was sustained throughout the 32 week trial. | [45] | |
| MERS-CoV | |||||||
| 4 | GeneOne Life Science and Inovio Pharmaceuticals; GLS-5300 | United States (Maryland) Phase I clinical trial; Open-label, single-arm, dose-escalation study; 3 doses of 0·67 mg, 2 mg, or 6 mg GLS-5300; intramuscular 1 mL injection followed immediately by co-localised intramuscular electroporation with CELLECTRA®-5P device at week 0, 4 and 12 with follow-up through to 48 weeks after dose 3 | 75 healthy adults between 18 and 50 years old (mean age 32.2 years), with normal screening electrocardiogram, screening laboratory findings within normal limits or be grade 0–1 findings, and have no history of clinically significant immunosuppressive or autoimmune disease, HIV, hepatitis B or C virus infection; 25 subjects were randomised into each dose group and 65 subjects completed the study, and per protocol analysis was used. | No vaccine-associated serious adverse events. 97% participants reported at least one solicited adverse event, but most solicited symptoms were reported as mild and were self-limiting (19 [76%] with 0·67 mg, 20 [80%] with 2 mg, and 17 [68%] with 6 mg); injection site reactions were the most common adverse event [92%]. | Seroconversion measured by S1-ELISA occurred in 86% and 94% participants after 2 and 3 doses, respectively, and was maintained in 79% participants up to study end at week 60. Neutralising antibodies were detected in 50% participants at one or more time points during the study, but only 3% maintained neutralisation activity to end of study. T-cell responses were detected in 71% and 76% participants after 2 and 3 doses, respectively. There were no differences in immune responses between dose groups after 6 weeks and vaccine-induced humoral and cellular responses were respectively detected in 77% and 64% participants at week 60. | [47] | |
| S/N | Company | Treatment | Stage | Reference |
|---|---|---|---|---|
| 1 | AbbVie | Lopinavir-ritonavir | Approved. Used in clinical settings. The Jin Yintan Hospital in Wuhan, China, launched a randomised, open-label, blank-controlled trial for the efficacy and safety of lopinavir-ritonavir and interferon-alpha 2b in hospitalisation of 80 patients with novel coronavirus infection. Lopinavir-ritonavir tablets (each containing 200 mg of lopinavir and 50 mg of ritonavir), twice a day, 2 tablets at a time; interferon-α2b. Assessment of effectiveness of treatment based on clinical improvement time of 28 days after randomisation. | [80] |
| 2 | Sanofi-Aventis | Teicoplanin (Targocid) | Approved. Used in clinical setting. | |
| 3 | Gilead Science | Remdesivir | Gilead is in active discussions with researchers and clinicians in the United States and China regarding the ongoing Wuhan coronavirus outbreak and the potential use of remdesivir as an investigational treatment. | [81] |
| 4 | Vir | Monoclonal antibodies | Vir is working to rapidly determine whether its previously identified anti-coronavirus monoclonal antibodies (mAbs) bind and neutralize 2019-nCoV. | [67] |
| 5 | Regeneron | Monoclonal antibodies | Regeneron Pharmaceuticals has developed monoclonal antibodies to treat MERS that are now being tested in early human studies. A company spokesperson said that researchers have begun to identify similar antibodies that might work against 2019-nCoV. With Ebola, it took Regeneron six months to develop candidate treatments and test them in animal models. | [82] |
| 6 | Ascletis | Ritonavir + ASC09 combo | Applied to include in national emergency channel on 25 January 2020. Not yet approved by regulators. | [83] |
| 7 | Biocryst Pharmaceuticals | Galidesivir | Biocryst is evaluating Galidesivir to determine if it could potentially target the coronavirus. Galidesivir is currently in a phase 1 clinical study. | [84] |
| 8 | Purdue University | Molecules that inhibit 2 coronavirus enzymes | Molecules developed by the university scientists inhibit two coronavirus enzymes and prevent its replication. The discovered drug targets are said to be more than 95% similar to enzyme targets found on the SARS virus. Researchers note that identified drugs may not be available to address the ongoing outbreak but they hope to make it accessible for future outbreaks. | [85] |
| 9 | The First Affiliated Hospital of Guangzhou Medical University | “xue bi jing” (TCM) -ChiCTR2000029381 | Approved. Recruitment of subjects has not started. | [86] |
| 10 | Chongqing Public Health Medical Center | Adjunctive steroids has a trial-ChiCTR2000029386 | Approved. Recruitment of subjects has not started. | [87] |
| 11 | Ruijin Hospital | Umefinovir (arbidol)- NCT04260594 | An antiviral treatment for influenza infection. Preliminary test in the in vitro cell showed an effective inhibition of coronavirus and a significant inhibition to the cytopathic effect. | [88] |
| 12 | Shanghai Public Health Clinical Center | Darunavir-NCT04252274 | An antiviral treatment for HIV. Study showed that it can significantly inhibit the replication of the new coronavirus. | [88] |
| 13 | Harbin Pharmaceutical Group Sanjing Pharmaceutical Holding Co., Ltd. | Oral liquid traditional Chinese medicine, Shuanghuanglian -ChiCTR2000029605 | Preliminary testing identified that it can inhibit the new coronavirus. It was previously identified to have an antiviral effect for influenza virus, SARS and MERS. | [88,89,90] |
| 14 | The Fifth Affiliated Hospital of Sun Yat-Sen University | Chloroquine phosphate -ChiCTR2000029609 | Approved. Recruitment of subjects has not started. | [88,91] |
| 15 | Shanghai Public Health Clinical Center | Hydroxychloroquine | Recruitment in process; Interventional subjects will receive hydroxychloroquine 400 mg per day for 5 days, also take conventional treatments. | [53] |
| 16 | Tongji Hospital | Abidol Hydrochloride combined with Interferon atomization | Recruitment of subjects has not started; Interventional subjects will receive standard symptomatic support therapy (SMT) plus abidol hydrochloride (0.2 g, 3 times a day) or Abidol Hydrochloride combined with Interferon (PegIFN-α-2b) atomization (45 ug) | [55] |
| 17 | Tongji Hospital | Drug: Abidol hydrochloride, Oseltamivir, Lopinavir/ritonavir | Recruitment of subjects has not started; Interventional subjects will receive either Abidol hydrochloride 0.2 g once, 3 times a day, 2 weeks or Oseltamivir 75 mg once, twice a day, 2 weeks or Lopinavir/ritonavir 500 mg once, twice a day, 2 weeks | [54] |
| 18 | Guangzhou 8th People’s Hospital | Lopinavir Plus Ritonavir and Arbidol | Recruitment in process; Interventional subjects will receive either standard treatment plus a regimen of lopinavir (200 mg) and ritonavir (50 mg) (oral, q12h, every time 2 tablets of each, taking for 7–14 days) or Standard treatment plus a regimen of arbidol (100 mg) (oral, tid, 200 mg each time, taking for 7–14 days). | [56] |
| 19 | China-Japan Friendship Hospital | Remdesivir | Recruitment of subjects has not started | [57] |
| 20 | Shanghai Public Health Clinical Center | Darunavir & Cobicistat | Recruitment of subjects has not started; Interventional subjects will receive darunavir and cobicistat one tablet per day for 5 days plus conventional treatments. | [58] |
| 21 | Beijing 302 Hospital | Mesenchymal Stem Cell (MSC) | Recruitment in process; Interventional subjects will receive conventional treatment plus 3 times of MSCs (0.5–1.0 × 10E6 MSCs/kg body weight intravenously at Day 0, Day 3, Day 6). | [59] |
| 22 | Beijing 302 Hospital | Traditional Chinese Medicine | Recruitment in process; Interventional subjects will receive oxygen therapy, antiviral therapy (alfa interferon via aerosol inhalation, and lopinavir/ritonavir, 400 mg/100 mg, p.o, bid) for 14 days plus Traditional Chinese Medicines (TCMs) granules: one bag, p.o, bid, for 14 days. | [60] |
| 23 | Medical ICU, Peking Union Medical College Hospital | Methylprednisolone | Recruitment in process; Interventional subjects will receive standard care plus methylprednisolone therapy (40 mg q12h for 5 days) | [61] |
| S/N | Date | Country | No. of Participants | Treatment | Outcome | Reference |
|---|---|---|---|---|---|---|
| SARS-CoV | ||||||
| 1 | Recruitment: 24 March–28 April 2003 | Hong Kong | 152/152 | Historical control: 4 g oral loading dose followed by 1.2 g every 8 h, or 8 mg/kg intravenously every 8 h if the patient could not tolerate oral treatment) with a reducing regimen of corticosteroid for 21 days Treatment group: combination of lopinavir (400 mg)/ritonavir (100 mg) orally every 12 h for 14 days Both groups given ribavirin & corticosteroid according to the same protocol | Primary outcome: composite adverse outcome at 21 days, severe hypoxaemia to fraction of inspired oxygen or death The 21 day adverse outcome rate was therefore 28.8% for the historical controls and 2.4% for the treatment group, giving an effect size of 26.4% (95% confidence interval 16.8 to 36.0, p = 0.001) for lopinavir/ritonavir treatment. Apparent reduction in also viral load, steroid dose, incidence of nocosomial infections in treatment group compared to control group. | [98] |
| 2 | Recruitment: 24 April–30 June 2003 | Taiwan | 4/4 | CM A: Composition of 13 herbs CM B: Popular health care product in Taiwan | Death in patient 1 (placebo control), recovery in remaining 3 patients | [93] |
| 3 | Recruitment: 10 April–31 May 2003 | China | 115/123 | Western medicine: oxygen supplementation, hemofiltration, ribavirin, antibacterials (azithromycin, cefuroxime, metronidazole), and immunoregulation with thymosin injection. Combined treatment: Herba houttuyniae with western medicine, and when necessary, TCM treatment like heat clearing and detoxifying, qi supplementing, blood regulating prescription. | Patients with early symptoms experienced a longer hospital stay (p = 0.028), a non-statistically significant mortality rate decrease (combined treatment: 9.6% versus WM: 11.1%), and a significant improvement of arthralgia and myalgia (p < 0.05) when on combined treatment compared with a strictly WM treatment. Combined treatment also improved arterial oxyhemoglobin saturation significantly at day 22 (p < 0.05) compared to WM. | [96] |
| 4 | 20 April 2003–30 May 2003 | Hong Kong | 16/17 | 9 (Early hydrocortisone treatment) & 7 (Saline placebo) | Median time for SARS-CoV RNA to become undetectable In plasma was 12 days vs. 8 days in the hydrocortisone and placebo groups. Corticosteroid treatment early in the treatment was associated with higher subsequent plasma viral load compared to placebo (AUC; Mann–Whitney, p = 0.023), delaying viral clearance | [95] |
| 5 | 28 January–28 February 2003 | China | 190/190 | 40 (A; Ribavirin, cefoperazone/sulbactam); 30 (B; fluoroquinolone, azithromycin, recombinant interferon alpha and restricted steroid use); 60 (C; quinolone, azithromycin, some given recombinant interferon alpha, steroid use when symptoms worsen); 60 (D; levofloxacin, azithromycin, of which 45 were given recombinant interferon alpha) | Treatment D presented with the most favourable outcome. The shortest mean time to discharge, 20.7 (SD, 4.6) days, was observed in treatment group D, compared to 24.8, 24.8 and 22.4 days for treatment groups A, B and C respectively. None of the patients on treatment D needed mechanical ventilation, all recovered and were discharged from the hospital compared to 2, 2, and 7 deaths from patients on treatments A, B and C respectively | [97] |
| 6 | Not reported | China | 49/52 | 29 (Control: Ribavirin, Levofloxacin, Thyopentin, Azithromycin, methylpredisolone) 20 (Treatment: Control group treatment + TCM Recipes) | All patients recovered. Significantly shorted time from the disease onset to the symptom improvement in treatment (5.10 ± 2.83 days) compared to control group (7.62 ± 2.27 days) (p < 0.05) No significant difference in blood routine improvement, pulmonary chest shadow in chest film improvement and corticosteroid usgae between the 2 groups. However, particularly in the respect of improving clinical symptoms, elevating quality of life, promoting immune function recovery, promoting absorption of pulmonary inflammation, reducing the dosage of cortisteroid and shortening the therapeutic course, treatment with integrative chinese and western medicine treatment had obvious superiority compared with using control treatment alone. | [94] |
| MERS-CoV | ||||||
| 7 | 2 June 2016–4 January 2017 | United States | 38/43 | Each group comprises 6 cohorts (Escalating doses 1 mg/kg, 2.5 mg/kg, 5 mg/kg, 10 mg/kg, 20 mg/kg & 50 mg/kg in SAB-001 treatment group) | A total of 97 adverse events (AEs) were reported, with a mean of 2.3 AEs per participant in the SAB-301 group and a mean of 3.3 AEs per participant in the placebo group. No serious adverse event related to the study intervention was observed. Single dose pharmacokinetics demonstrated relatively linear and dose-proportional increases in maximal concentration and area-under-the-concentration-time curve. Microneutralization titres also correlated to the SAB-301 levels in serum. Single infusions of SAB-301 up to 50 mg/kg appear to be safe and well-tolerated in healthy participants. | [46] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pang, J.; Wang, M.X.; Ang, I.Y.H.; Tan, S.H.X.; Lewis, R.F.; Chen, J.I.-P.; Gutierrez, R.A.; Gwee, S.X.W.; Chua, P.E.Y.; Yang, Q.; et al. Potential Rapid Diagnostics, Vaccine and Therapeutics for 2019 Novel Coronavirus (2019-nCoV): A Systematic Review. J. Clin. Med. 2020, 9, 623. https://doi.org/10.3390/jcm9030623
Pang J, Wang MX, Ang IYH, Tan SHX, Lewis RF, Chen JI-P, Gutierrez RA, Gwee SXW, Chua PEY, Yang Q, et al. Potential Rapid Diagnostics, Vaccine and Therapeutics for 2019 Novel Coronavirus (2019-nCoV): A Systematic Review. Journal of Clinical Medicine. 2020; 9(3):623. https://doi.org/10.3390/jcm9030623
Chicago/Turabian StylePang, Junxiong, Min Xian Wang, Ian Yi Han Ang, Sharon Hui Xuan Tan, Ruth Frances Lewis, Jacinta I-Pei Chen, Ramona A Gutierrez, Sylvia Xiao Wei Gwee, Pearleen Ee Yong Chua, Qian Yang, and et al. 2020. "Potential Rapid Diagnostics, Vaccine and Therapeutics for 2019 Novel Coronavirus (2019-nCoV): A Systematic Review" Journal of Clinical Medicine 9, no. 3: 623. https://doi.org/10.3390/jcm9030623
APA StylePang, J., Wang, M. X., Ang, I. Y. H., Tan, S. H. X., Lewis, R. F., Chen, J. I.-P., Gutierrez, R. A., Gwee, S. X. W., Chua, P. E. Y., Yang, Q., Ng, X. Y., Yap, R. K. S., Tan, H. Y., Teo, Y. Y., Tan, C. C., Cook, A. R., Yap, J. C.-H., & Hsu, L. Y. (2020). Potential Rapid Diagnostics, Vaccine and Therapeutics for 2019 Novel Coronavirus (2019-nCoV): A Systematic Review. Journal of Clinical Medicine, 9(3), 623. https://doi.org/10.3390/jcm9030623






