Time-Dependent Response of Human Deciduous Tooth-Derived Dental Pulp Cells Treated with TheraCal LC: Functional Analysis of Gene Interactions Compared to MTA
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Isolation and Culture
2.2. Cell Viability Test
2.3. Material Preparation
2.4. RNA Isolation and Library Preparation for Next Generation Sequencing
2.5. Data Processing to Identify DEGs and GO Functional Enrichment Analysis
2.6. GSEA and PPI Network Construction
3. Results
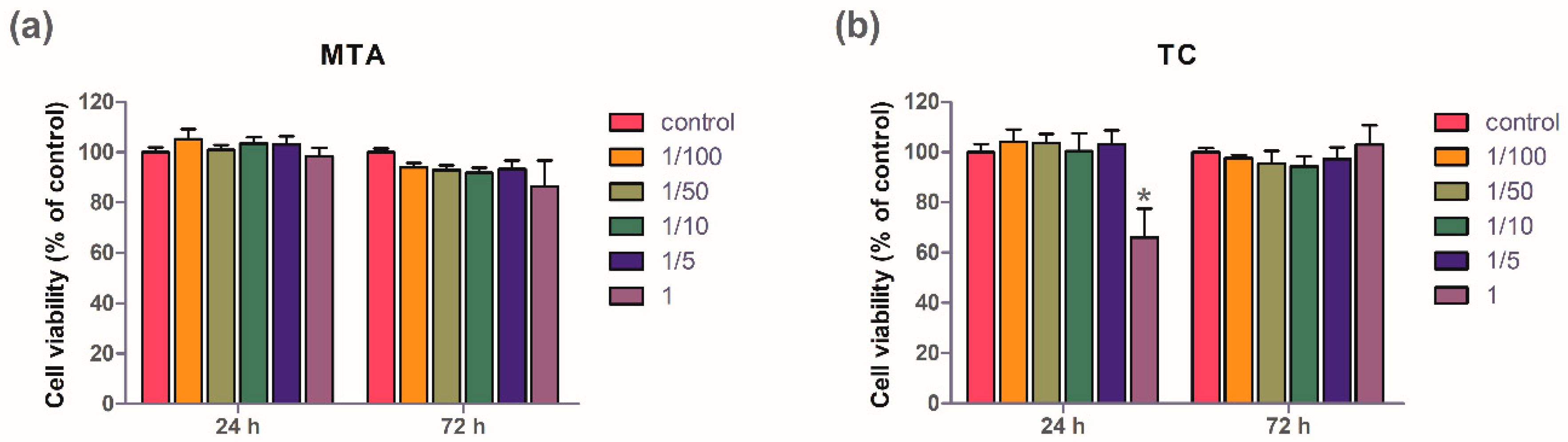
3.1. Cell Viability Test
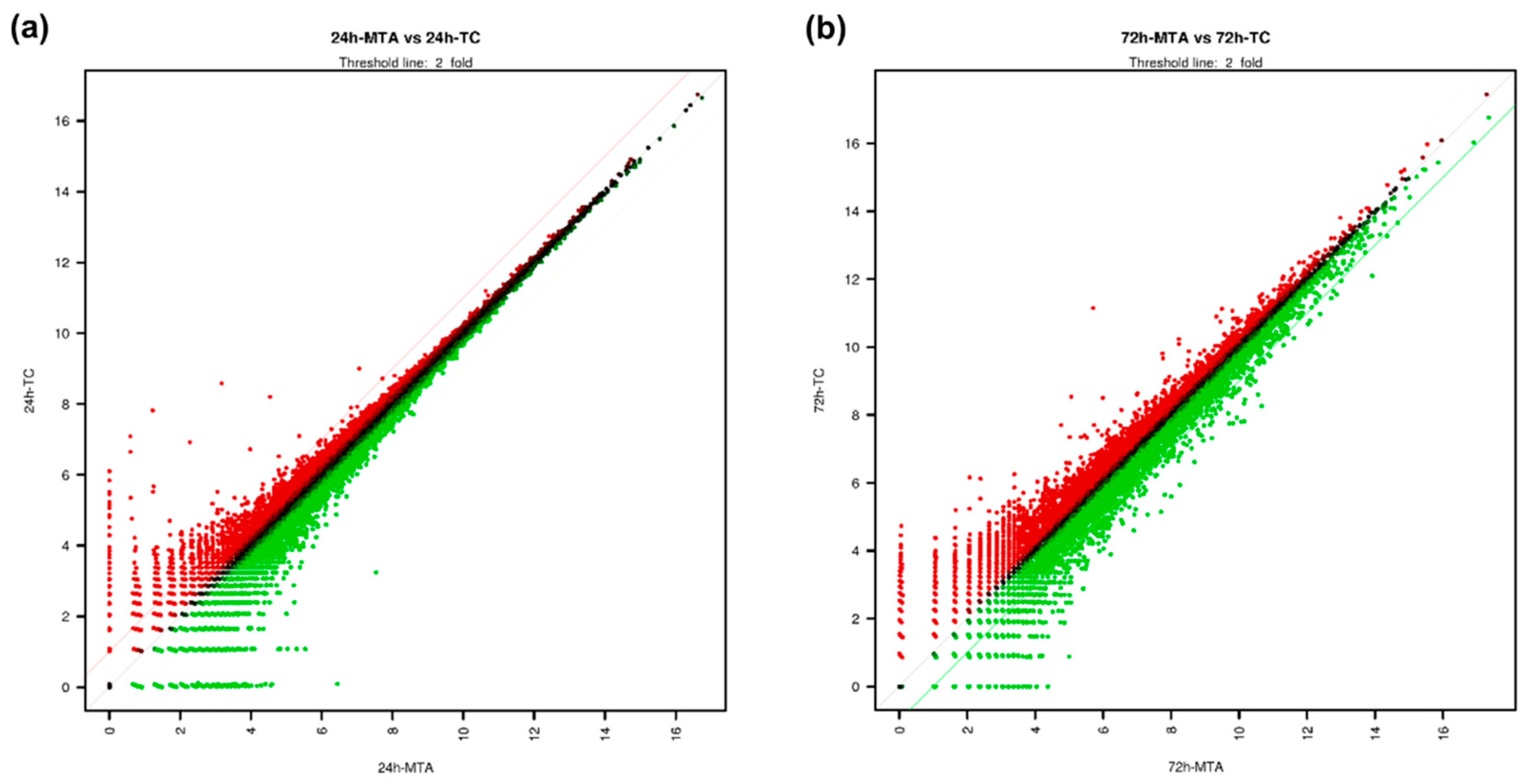
3.2. Overview of Differentially Expressed Genes (DEGs) in TC
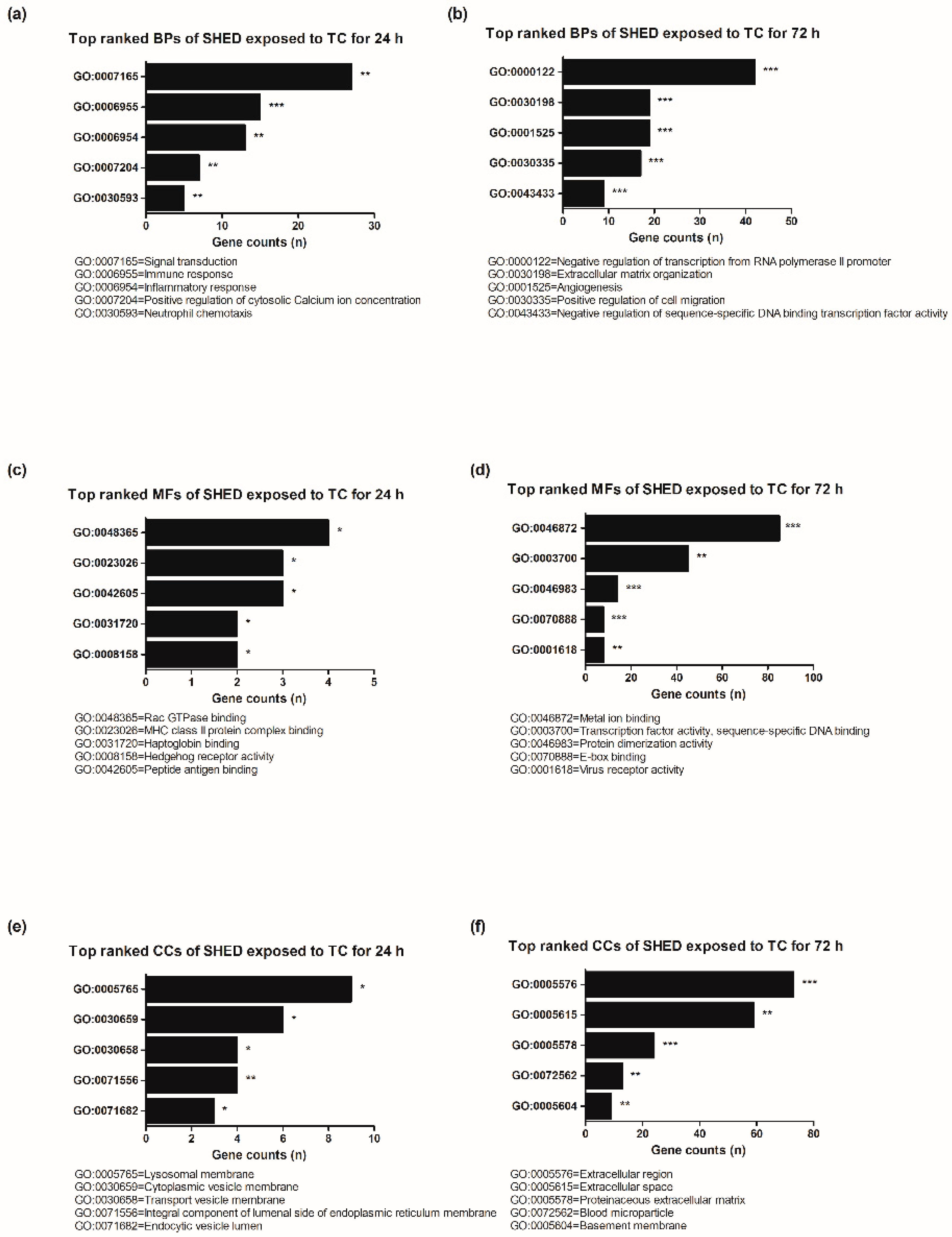
3.3. Gene Ontology (GO) Functional Enrichment Analysis
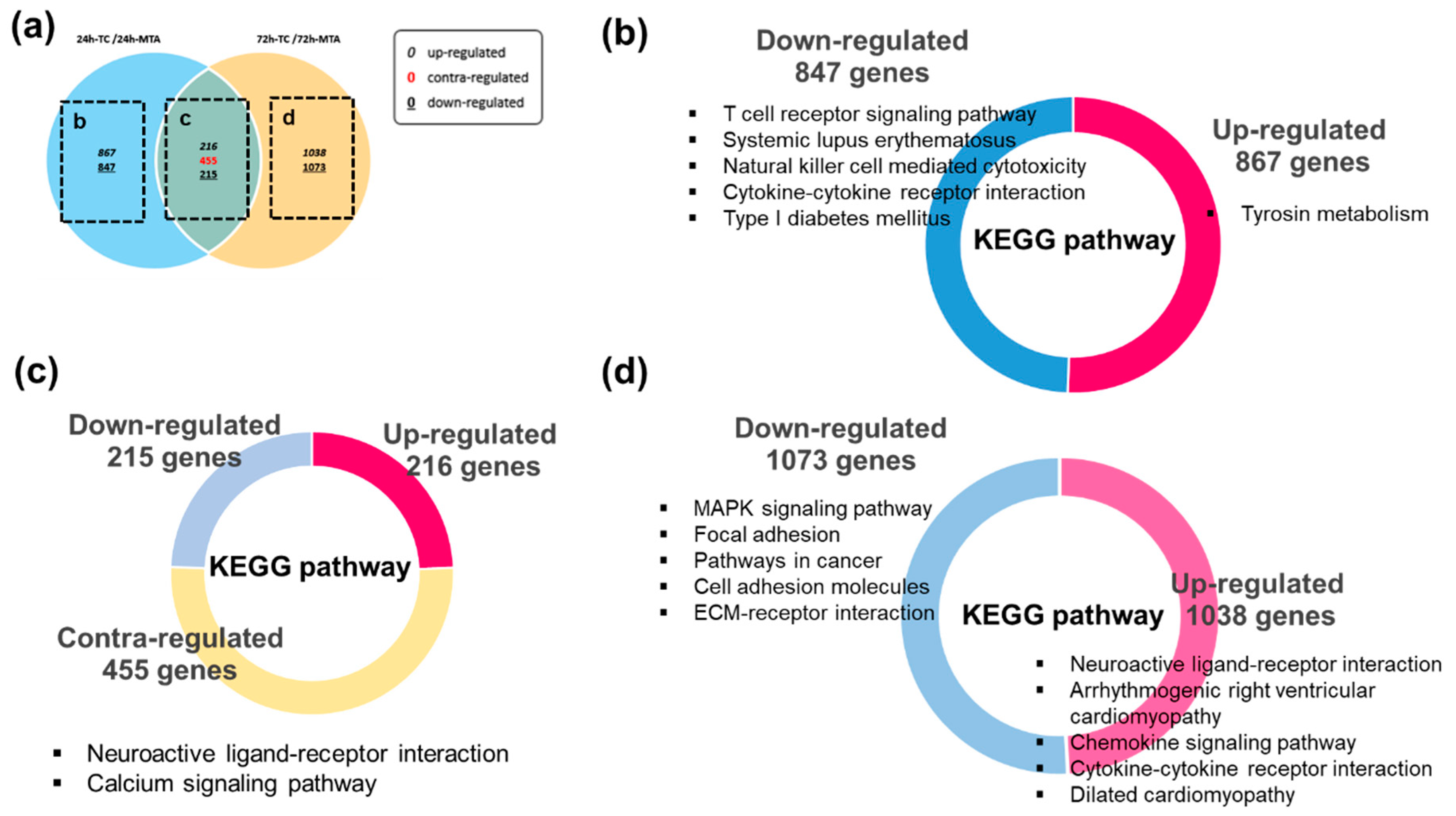
3.4. Kyoto Encyclopedia of Genes and Genomes (KEGG) Pathway Enrichment Analysis
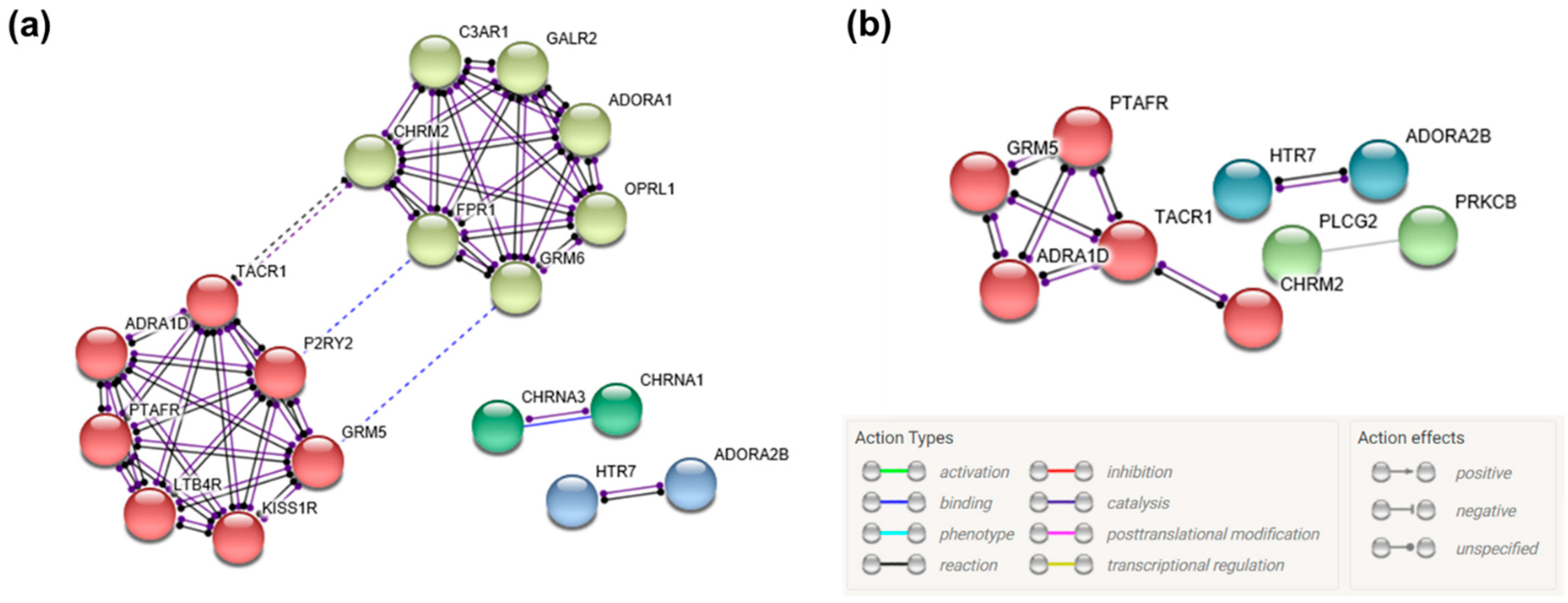
3.5. Gene Network Construction
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
Appendix A
| Gene | Description | FC2 Ratio (TC/MTA) | Normalized FC (log2) | ||||
|---|---|---|---|---|---|---|---|
| 24 h | 72 h | ||||||
| 24 h | 72 h | MTA | TC | MTA | TC | ||
| GRM5 | Glutamate metabotropic receptor 5 | 0.374 | 0.489 | 1.437 | 0.017 | 1.033 | 0.000 |
| ADORA2B | Adenosine A2b receptor | 2.889 | 0.403 | 1.332 | 2.863 | 3.217 | 1.905 |
| CHRM2 | Cholinergic receptor muscarinic 2 | 2.066 | 0.139 | 0.000 | 1.047 | 2.843 | 0.000 |
| HTR7 | 5-hydroxytryptamine receptor 7 | 0.420 | 3.231 | 4.639 | 3.386 | 2.386 | 4.078 |
| PTAFR | Platelet activating factor receptor | 3.093 | 0.244 | 0.000 | 1.629 | 2.038 | 0.000 |
| TACR1 | Tachykinin receptor 1 | 0.360 | 2.425 | 3.122 | 1.647 | 1.640 | 2.918 |
| C3AR1 | Complement C3a receptor 1 | 0.368 | 2.872 | 1.454 | 0.012 | 0.011 | 1.533 |
| CHRNA1 | Cholinergic receptor nicotinic alpha 1 subunit | 0.253 | 0.323 | 4.387 | 2.403 | 6.530 | 4.899 |
| CHRNA3 | Cholinergic receptor nicotinic alpha 3 subunit | 0.462 | 0.372 | 2.159 | 1.045 | 2.358 | 0.930 |
| CHRNB1 | Cholinergic receptor nicotinic beta 1 subunit | 2.937 | 2.494 | 2.900 | 4.454 | 3.231 | 4.550 |
| LTB4R | Leukotriene B4 receptor | 0.351 | 0.475 | 5.485 | 3.977 | 5.671 | 4.598 |
| ADORA1 | Adenosine A1 receptor | 4.749 | 2.766 | 0.795 | 3.043 | 1.046 | 2.514 |
| FPR1 | Formyl peptide receptor 1 | 0.197 | 11.638 | 4.009 | 1.666 | 1.657 | 5.198 |
| GABRA1 | Gamma-aminobutyric acid type A receptor alpha1 subunit | 2.056 | 4.751 | 0.000 | 1.040 | 0.017 | 2.265 |
| GLRA2 | Glycine receptor alpha 2 | 4.091 | 0.327 | 0.000 | 2.032 | 1.615 | 0.000 |
| GRM6 | Glutamate metabotropic receptor | 0.298 | 4.122 | 2.808 | 1.059 | 1.052 | 3.096 |
| OPRL1 | Opioid related nociception receptor 1 | 0.304 | 2.337 | 4.371 | 2.652 | 2.383 | 3.608 |
| P2RY2 | Purinergic receptor P2Y2 | 2.652 | 2.344 | 2.556 | 3.963 | 2.382 | 3.611 |
| TRPV1 | Transient receptor potential cation channel subfamily V member 1 | 0.311 | 3.241 | 3.346 | 1.661 | 2.384 | 4.081 |
| KISS1R | KISS1 receptor | 0.365 | 0.327 | 1.463 | 0.010 | 1.612 | 0.000 |
| GALR2 | Galanin receptor 2 | 0.378 | 0.374 | 1.423 | 0.021 | 2.356 | 0.936 |
| Gene | Description | FC2 Ratio (TC/MTA) | Normalized FC (log2) | ||||
|---|---|---|---|---|---|---|---|
| 24 h | 72 h | ||||||
| 24 h | 72 h | MTA | TC | MTA | TC | ||
| GRM5 | Glutamate metabotropic receptor 5 | 0.374 | 0.489 | 1.437 | 0.017 | 1.033 | 0.000 |
| ADORA2B | Adenosine A2b receptor | 2.889 | 0.403 | 1.332 | 2.863 | 3.217 | 1.905 |
| ADRA1D | Adrenoceptor alpha | 3.113 | 0.134 | 2.325 | 3.964 | 4.370 | 1.469 |
| CHRM2 | Cholinergic receptor muscarinic 2 | 2.066 | 0.139 | 0.000 | 1.047 | 2.843 | 0.000 |
| HTR7 | 5-hydroxytryptamine receptor 7 | 0.420 | 3.231 | 4.639 | 3.386 | 2.386 | 4.078 |
| PTAFR | Platelet activating factor receptor | 3.093 | 0.244 | 0.000 | 1.629 | 2.038 | 0.000 |
| TACR1 | Tachykinin receptor 1 | 0.360 | 2.425 | 3.122 | 1.647 | 1.640 | 2.918 |
| PRKCB | Protein kinase C beta | 0.221 | 2.862 | 2.191 | 0.014 | 0.013 | 1.530 |
| PLCG2 | Phospholipase C gamma | 0.405 | 2.285 | 2.368 | 1.064 | 2.052 | 3.244 |
| CAMK2A | Calcium/calmodulin dependent protein kinase II alpha | 0.305 | 6.701 | 2.783 | 1.068 | 1.642 | 4.387 |
| CAMK4 | Calcium/calmodulin dependent protein kinase IV | 4.339 | 4.554 | 0.744 | 2.861 | 0.045 | 2.232 |
| PDE1A | Phosphodiesterase 1A | 0.455 | 0.387 | 2.786 | 1.649 | 2.856 | 1.485 |
| ATP2B2 | ATPase plasma membrane Ca2+ transporting 2 | 0.288 | 9.435 | 1.822 | 0.028 | 0.025 | 3.263 |
| TNNC2 | Troponin C2, fast skeletal type | 0.480 | 3.034 | 3.119 | 2.060 | 1.641 | 3.242 |
References
- Cohenca, N.; Paranjpe, A.; Berg, J. Vital pulp therapy. Dent. Clin. N. Am. 2013, 57, 59–73. [Google Scholar] [CrossRef]
- Tuna, D.; Ölmez, A. Clinical long-term evaluation of MTA as a direct pulp capping material in primary teeth. Int. Endod. J. 2008, 41, 273–278. [Google Scholar] [CrossRef] [PubMed]
- Huang, G.T.; Gronthos, S.; Shi, S. Mesenchymal stem cells derived from dental tissues vs. those from other sources: Their biology and role in regenerative medicine. J. Dent. Res. 2009, 88, 792–806. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.H.; Kim, S.Y.; Woo, S.M.; Jeong, H.N.; Jung, J.Y.; Kim, S.M.; Lim, H.S. Combination of mineral trioxide aggregate and propolis promotes odontoblastic differentiation of human dental pulp stem cells through ERK signaling pathway. Food Sci. Biotechnol. 2019, 28, 1801–1809. [Google Scholar] [CrossRef] [PubMed]
- Dammaschke, T.; Leidinger, J.; Schafer, E. Long-term evaluation of direct pulp capping--treatment outcomes over an average period of 6.1 years. Clin. Oral Investig. 2010, 14, 559–567. [Google Scholar] [CrossRef]
- Takita, T.; Hayashi, M.; Takeichi, O.; Ogiso, B.; Suzuki, N.; Otsuka, K.; Ito, K. Effect of mineral trioxide aggregate on proliferation of cultured human dental pulp cells. Int. Endod. J. 2006, 39, 415–422. [Google Scholar] [CrossRef]
- Lourenco Neto, N.; Marques, N.C.; Fernandes, A.P.; Rodini, C.O.; Sakai, V.T.; Abdo, R.C.; Machado, M.A.; Santos, C.F.; Oliveira, T.M. Immunolocalization of dentin matrix protein-1 in human primary teeth treated with different pulp capping materials. J. Biomed. Mater. Res. B Appl. Biomater. 2016, 104, 165–169. [Google Scholar] [CrossRef]
- Collado-Gonzalez, M.; Garcia-Bernal, D.; Onate-Sanchez, R.E.; Ortolani-Seltenerich, P.S.; Alvarez-Muro, T.; Lozano, A.; Forner, L.; Llena, C.; Moraleda, J.M.; Rodriguez-Lozano, F.J. Cytotoxicity and bioactivity of various pulpotomy materials on stem cells from human exfoliated primary teeth. Int. Endod. J. 2017, 50 (Suppl. 2), e19–e30. [Google Scholar] [CrossRef]
- Carrotte, P. Endodontics: Part 9. Calcium hydroxide, root resorption, endo-perio lesions. Br. Dent. J. 2004, 197, 735–743. [Google Scholar] [CrossRef]
- Roberts, H.W.; Toth, J.M.; Berzins, D.W.; Charlton, D.G. Mineral trioxide aggregate material use in endodontic treatment: A review of the literature. Dent. Mater. 2008, 24, 149–164. [Google Scholar] [CrossRef]
- Seo, M.S.; Hwang, K.G.; Lee, J.; Kim, H.; Baek, S.H. The effect of mineral trioxide aggregate on odontogenic differentiation in dental pulp stem cells. J. Endod. 2013, 39, 242–248. [Google Scholar] [CrossRef] [PubMed]
- Yoldas, S.E.; Bani, M.; Atabek, D.; Bodur, H. Comparison of the Potential Discoloration Effect of Bioaggregate, Biodentine, and White Mineral Trioxide Aggregate on Bovine Teeth: In Vitro Research. J. Endod. 2016, 42, 1815–1818. [Google Scholar] [CrossRef] [PubMed]
- Torabinejad, M.; Hong, C.U.; McDonald, F.; Pitt Ford, T.R. Physical and chemical properties of a new root-end filling material. J. Endod. 1995, 21, 349–353. [Google Scholar] [CrossRef]
- Gandolfi, M.G.; Taddei, P.; Siboni, F.; Modena, E.; Ciapetti, G.; Prati, C. Development of the foremost light-curable calcium-silicate MTA cement as root-end in oral surgery. Chemical-physical properties, bioactivity and biological behavior. Dent. Mater. 2011, 27, e134–e157. [Google Scholar] [CrossRef]
- Li, X.; Pedano, M.S.; Camargo, B.; Hauben, E.; De Vleeschauwer, S.; Chen, Z.; De Munck, J.; Vandamme, K.; Van Landuyt, K.; Van Meerbeek, B. Experimental tricalcium silicate cement induces reparative dentinogenesis. Dent. Mater. 2018, 34, 1410–1423. [Google Scholar] [CrossRef]
- Schneider, T.R.; Hakami-Tafreshi, R.; Tomasino-Perez, A.; Tayebi, L.; Lobner, D. Effects of dental composite resin monomers on dental pulp cells. Dent. Mater. J. 2019, 38, 579–583. [Google Scholar] [CrossRef]
- Hebling, J.; Lessa, F.C.; Nogueira, I.; Carvalho, R.M.; Costa, C.A. Cytotoxicity of resin-based light-cured liners. Am. J. Dent. 2009, 22, 137–142. [Google Scholar]
- Wang, Z.; Gerstein, M.; Snyder, M. RNA-Seq: A revolutionary tool for transcriptomics. Nat. Rev. Genet. 2009, 10, 57–63. [Google Scholar] [CrossRef]
- Fuks, A.B. Pulp therapy for the primary and young permanent dentitions. Dent. Clin. N. Am. 2000, 44, 571–596. [Google Scholar]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef]
- Quinlan, A.R.; Hall, I.M. BEDTools: A flexible suite of utilities for comparing genomic features. Bioinformatics 2010, 26, 841–842. [Google Scholar] [CrossRef] [PubMed]
- Gentleman, R.C.; Carey, V.J.; Bates, D.M.; Bolstad, B.; Dettling, M.; Dudoit, S.; Ellis, B.; Gautier, L.; Ge, Y.; Gentry, J.; et al. Bioconductor: Open software development for computational biology and bioinformatics. Genome Biol. 2004, 5, R80. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [Google Scholar] [CrossRef] [PubMed]
- Liberzon, A.; Birger, C.; Thorvaldsdottir, H.; Ghandi, M.; Mesirov, J.P.; Tamayo, P. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 2015, 1, 417–425. [Google Scholar] [CrossRef] [PubMed]
- Ranly, D.M.; Garcia-Godoy, F. Current and potential pulp therapies for primary and young permanent teeth. J. Dent. 2000, 28, 153–161. [Google Scholar] [CrossRef]
- Erfanparast, L.; Iranparvar, P.; Vafaei, A. Direct pulp capping in primary molars using a resin-modified Portland cement-based material (TheraCal) compared to MTA with 12-month follow-up: A randomised clinical trial. Eur. Arch. Paediatr. Dent. 2018, 19, 197–203. [Google Scholar] [CrossRef]
- Asl Aminabadi, N.; Satrab, S.; Najafpour, E.; Samiei, M.; Jamali, Z.; Shirazi, S. A randomized trial of direct pulp capping in primary molars using MTA compared to 3Mixtatin: A novel pulp capping biomaterial. Int. J. Paediatr. Dent. 2016, 26, 281–290. [Google Scholar] [CrossRef]
- Nilsen, B.W.; Jensen, E.; Ortengren, U.; Michelsen, V.B. Analysis of organic components in resin-modified pulp capping materials: Critical considerations. Eur. J. Oral Sci. 2017, 125, 183–194. [Google Scholar] [CrossRef]
- Zakerzadeh, A.; Esnaashari, E.; Dadfar, S. In Vitro Comparison of Cytotoxicity and Genotoxicity of Three Vital Pulp Capping Materials. Iran. Endod. J. 2017, 12, 419–425. [Google Scholar]
- Lv, L.W.; Liu, Y.S.; Zhang, P.; Gu, M.; Bai, X.S.; Xiong, C.Y.; Zhou, Y.S. Transcriptomics and Functional Analysis of Graphene-Guided Osteogenic Differentiation of Mesenchymal Stem Cells. Chin. J. Dent. Res. 2018, 21, 101–111. [Google Scholar]
- Mirza, R.; DiPietro, L.A.; Koh, T.J. Selective and Specific Macrophage Ablation Is Detrimental to Wound Healing in Mice. Am. J. Pathol. 2009, 175, 2454–2462. [Google Scholar] [CrossRef] [PubMed]
- Nakatomi, M.; Morita, I.; Eto, K.; Ota, M.S. Sonic hedgehog signaling is important in tooth root development. J. Dent. Res. 2006, 85, 427–431. [Google Scholar] [CrossRef] [PubMed]
- Xia, L.; Zhang, M.; Chang, Q.; Wang, L.; Zeng, D.; Zhang, X.; Zhang, Z.; Jiang, X. Enhanced dentin-like mineralized tissue formation by AdShh-transfected human dental pulp cells and porous calcium phosphate cement. PLoS ONE 2013, 8, e62645. [Google Scholar] [CrossRef] [PubMed]
- Ma, D.; Yu, H.; Xu, S.; Wang, H.; Zhang, X.; Ning, T.; Wu, B. Stathmin inhibits proliferation and differentiation of dental pulp stem cells via sonic hedgehog/Gli. J. Cell Mol. Med. 2018, 22, 3442–3451. [Google Scholar] [CrossRef]
- Kang, J.; Bai, R.; Liu, K.; Ji, X.P.; Li, Y.; Han, J.Y. Identification of significantly different modules between permanent and deciduous teeth by network and pathway analyses. Genet. Mol. Res. 2016, 15. [Google Scholar] [CrossRef]
- Xie, F.; Dai, Q.; Liu, X.; Wang, J. Conditional Knockout of Raptor/mTORC1 Results in Dentin Malformation. Front. Physiol. 2019, 10, 250. [Google Scholar] [CrossRef]
- Xiao, Y.; Zhao, X.-P. Screening pathways and hub genes involved in osteoclastogenesis by gene expression analysis of circulating monocytes based on Gibbs sampling. Exp. Ther. Med. 2019, 17, 2529–2534. [Google Scholar] [CrossRef]
- Rufas, P.; Jeanneau, C.; Rombouts, C.; Laurent, P.; About, I. Complement C3a Mobilizes Dental Pulp Stem Cells and Specifically Guides Pulp Fibroblast Recruitment. J. Endod. 2016, 42, 1377–1384. [Google Scholar] [CrossRef]
- Yi, X.; Wang, W.; Xie, Q. Adenosine receptors enhance the ATP-induced odontoblastic differentiation of human dental pulp cells. Biochem. Biophys. Res. Commun. 2018, 497, 850–856. [Google Scholar] [CrossRef]
- Hwang, S.-Y.; Putney, J.W., Jr. Calcium signaling in osteoclasts. Biochim. Biophys. Acta 2011, 1813, 979–983. [Google Scholar] [CrossRef]
- Takayanagi, H.; Kim, S.; Koga, T.; Nishina, H.; Isshiki, M.; Yoshida, H.; Saiura, A.; Isobe, M.; Yokochi, T.; Inoue, J.; et al. Induction and activation of the transcription factor NFATc1 (NFAT2) integrate RANKL signaling in terminal differentiation of osteoclasts. Dev. Cell 2002, 3, 889–901. [Google Scholar] [CrossRef]
- Yang, S.; Li, Y.P. RGS10-null mutation impairs osteoclast differentiation resulting from the loss of [Ca2+] i oscillation regulation. Genes Dev. 2007, 21, 1803–1816. [Google Scholar] [CrossRef] [PubMed]
- Takayanagi, H. Osteoimmunology: Shared mechanisms and crosstalk between the immune and bone systems. Nat. Rev. Immunol. 2007, 7, 292–304. [Google Scholar] [CrossRef] [PubMed]
- Ang, E.S.; Zhang, P.; Steer, J.H.; Tan, J.W.; Yip, K.; Zheng, M.H.; Joyce, D.A.; Xu, J. Calcium/calmodulin-dependent kinase activity is required for efficient induction of osteoclast differentiation and bone resorption by receptor activator of nuclear factor kappa B ligand (RANKL). J. Cell Physiol. 2007, 212, 787–795. [Google Scholar] [CrossRef]
- Alagarsamy, S.; Saugstad, J.; Warren, L.; Mansuy, I.M.; Gereau, R.W.; Conn, P.J. NMDA-induced potentiation of mGluR5 is mediated by activation of protein phosphatase 2B/calcineurin. Neuropharmacology 2005, 49, 135–145. [Google Scholar] [CrossRef]
- Mao, D.; Epple, H.; Uthgenannt, B.; Novack, D.V.; Faccio, R. PLCgamma2 regulates osteoclastogenesis via its interaction with ITAM proteins and GAB2. J. Clin. Investig. 2006, 116, 2869–2879. [Google Scholar] [CrossRef]
- Ratner, B.D. The Engineering of Biomaterials Exhibiting Recognition and Specificity. J. Mol. Recognit. 1996, 9, 617–625. [Google Scholar] [CrossRef]
- Smith, M.J.; White, K.L.; Smith, D.C.; Bowlin, G.L. In vitro evaluations of innate and acquired immune responses to electrospun polydioxanone–elastin blends. Biomaterials 2009, 30, 149–159. [Google Scholar] [CrossRef]
- Smith, M.J.; Smith, D.C.; Bowlin, G.L.; White, K.L. Modulation of murine innate and acquired immune responses following in vitro exposure to electrospun blends of collagen and polydioxanone. J. Biomed. Mater. Res. A 2010, 93, 793–806. [Google Scholar]
- Jewett, A.; Arasteh, A.; Tseng, H.C.; Behel, A.; Arasteh, H.; Yang, W.; Cacalano, N.A.; Paranjpe, A. Strategies to rescue mesenchymal stem cells (MSCs) and dental pulp stem cells (DPSCs) from NK cell mediated cytotoxicity. PLoS ONE 2010, 5, e9874. [Google Scholar] [CrossRef]
- Johnson, G.L.; Lapadat, R. Mitogen-activated protein kinase pathways mediated by ERK, JNK, and p38 protein kinases. Science 2002, 298, 1911–1912. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.H.; Sharrocks, A.D.; Whitmarsh, A.J. Transcriptional regulation by the MAP kinase signaling cascades. Gene 2003, 320, 3–21. [Google Scholar] [CrossRef]
- Luo, Z.; Kohli, M.R.; Yu, Q.; Kim, S.; Qu, T.; He, W.-X. Biodentine Induces Human Dental Pulp Stem Cell Differentiation through Mitogen-activated Protein Kinase and Calcium-/Calmodulin-dependent Protein Kinase II Pathways. J. Endod. 2014, 40, 937–942. [Google Scholar] [CrossRef] [PubMed]
- Rathinam, E.; Rajasekharan, S.; Chitturi, R.T.; Martens, L.; De Coster, P. Gene Expression Profiling and Molecular Signaling of Dental Pulp Cells in Response to Tricalcium Silicate Cements: A Systematic Review. J. Endod. 2015, 41, 1805–1817. [Google Scholar] [CrossRef]
- Chang, S.W.; Kim, J.Y.; Kim, M.J.; Kim, G.H.; Yi, J.K.; Lee, D.W.; Kum, K.Y.; Kim, E.C. Combined effects of mineral trioxide aggregate and human placental extract on rat pulp tissue and growth, differentiation and angiogenesis in human dental pulp cells. Acta Odontol. Scand. 2016, 74, 298–306. [Google Scholar] [CrossRef]
- Zhao, X.; He, W.; Song, Z.; Tong, Z.; Li, S.; Ni, L. Mineral trioxide aggregate promotes odontoblastic differentiation via mitogen-activated protein kinase pathway in human dental pulp stem cells. Mol. Biol. Rep. 2012, 39, 215–220. [Google Scholar] [CrossRef]
- Arandi, N.Z.; Rabi, T. TheraCal LC: From Biochemical and Bioactive Properties to Clinical Applications. Int. J. Dent. 2018, 2018, 3484653. [Google Scholar] [CrossRef]

| Entry ID | KEGG Pathway | Number of Genes | Involved Genes | p-Value | FDR q-Value1 |
|---|---|---|---|---|---|
| hsa04080 | Neuroactive ligand–receptor interaction | 22 | GRM5, ADORA2B, ADRA1D, CHRM2, HTR7, PTAFR, TACR1, C3AR1, CHRNA1, CHRNA3, CHRNB1, LTB4R, ADORA1, FPR1, GABRA1, GLRA2, GRM6, OPRL1, P2RY2, TRPV1, KISS1R, GALR2 | 1.19 × 10−7 | 2.22 × 10−5 |
| hsa04020 | Calcium signaling pathway | 14 | GRM5, ADORA2B, ADRA1D, CHRM2, HTR7, PTAFR, TACR1, PRKCB, PLCG2, CAMK2A, CAMK4, PDE1A, ATP2B2, TNNC2 | 2.96 × 10−5 | 2.75 × 10−3 |
| Entry ID | KEGG Pathway | Number of Genes | p-Value | FDR q-Value1 |
|---|---|---|---|---|
| Upregulated in 24 h2 | ||||
| hsa00350 | Tyrosine metabolism | 6 | 1.85 × 10−4 | 3.43 × 10−2 |
| Downregulated in 24 h | ||||
| hsa04660 | T cell receptor signaling pathway | 17 | 7.8 × 10−11 | 1.45 × 10−8 |
| hsa05322 | Systemic lupus erythematosus | 17 | 3.85 × 10−9 | 3.58 × 10−7 |
| hsa04650 | Natural killer cell mediated cytotoxicity | 16 | 2.38 × 10−8 | 1.48 × 10−6 |
| hsa04060 | Cytokine–cytokine receptor interaction | 21 | 1.6 × 10−7 | 6.71 × 10−6 |
| hsa04940 | Type I diabetes mellitus | 9 | 1.8 × 10−7 | 6.71 × 10−6 |
| Upregulated in 72 h | ||||
| hsa04080 | Neuroactive ligand–receptor interaction | 28 | 3.52 × 10−10 | 6.54 × 10−8 |
| hsa05412 | Arrhythmogenic right ventricular cardiomyopathy | 11 | 2.30 × 10−6 | 2.14 × 10−4 |
| hsa04062 | Chemokine signaling pathway | 15 | 9.26 × 10−5 | 5.74 × 10−3 |
| hsa04060 | Cytokine-cytokine receptor interaction | 17 | 4.23 × 10−4 | 1.61 × 10−2 |
| hsa05414 | Dilated cardiomyopathy | 9 | 4.34 × 10−4 | 1.61 × 10−2 |
| Downregulated in 72 h | ||||
| hsa04010 | MAPK signaling pathway | 24 | 1.52 × 10−7 | 2.82 × 10−5 |
| hsa04510 | Focal adhesion | 18 | 4.83 × 10−6 | 4.50 × 10−4 |
| hsa05200 | Pathways in cancer | 23 | 1.56 × 10−5 | 9.67 × 10−4 |
| hsa04514 | Cell adhesion molecules (CAMs) | 12 | 1.83 × 10−4 | 8.52 × 10−3 |
| hsa04512 | ECM-receptor interaction | 9 | 3.24 × 10−4 | 1.20 × 10−2 |
| Genes | Description | KEGG Pathway1 | FC2 Ratio (TC/MTA) | Normalized FC (log2) | ||||
|---|---|---|---|---|---|---|---|---|
| 24 h | 72 h | |||||||
| 24 h | 72 h | MTA | TC | MTA | TC | |||
| ADORA1 | Adenosine A1 receptor | 1 | 4.749 | 2.766 | 0.795 | 3.043 | 1.046 | 2.514 |
| ADORA2B | Adenosine A2b receptor | 1,2 | 2.889 | 0.403 | 1.332 | 2.863 | 3.217 | 1.905 |
| ATP2B2 | ATPase plasma membrane Ca2+ transporting 2 | 2 | 0.288 | 9.435 | 1.822 | 0.028 | 0.025 | 3.263 |
| CAMK4 | Calcium/calmodulin dependent protein kinase IV | 2 | 4.339 | 4.554 | 0.744 | 2.861 | 0.045 | 2.232 |
| C3AR1 | Complement C3a receptor 1 | 1 | 0.368 | 2.872 | 1.454 | 0.012 | 0.011 | 1.533 |
| GRM5 | Glutamate metabotropic receptor 5 | 1,2 | 0.374 | 0.489 | 1.437 | 0.017 | 1.033 | 0.000 |
| PLCG2 | Phospholipase C gamma 2 | 2 | 0.405 | 2.285 | 2.368 | 1.064 | 2.052 | 3.244 |
| P2RY2 | Purinergic receptor P2Y2 | 1 | 2.652 | 2.344 | 2.556 | 3.963 | 2.382 | 3.611 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nam, O.H.; Kim, J.-H.; Choi, S.C.; Kim, Y. Time-Dependent Response of Human Deciduous Tooth-Derived Dental Pulp Cells Treated with TheraCal LC: Functional Analysis of Gene Interactions Compared to MTA. J. Clin. Med. 2020, 9, 531. https://doi.org/10.3390/jcm9020531
Nam OH, Kim J-H, Choi SC, Kim Y. Time-Dependent Response of Human Deciduous Tooth-Derived Dental Pulp Cells Treated with TheraCal LC: Functional Analysis of Gene Interactions Compared to MTA. Journal of Clinical Medicine. 2020; 9(2):531. https://doi.org/10.3390/jcm9020531
Chicago/Turabian StyleNam, Ok Hyung, Jae-Hwan Kim, Sung Chul Choi, and Young Kim. 2020. "Time-Dependent Response of Human Deciduous Tooth-Derived Dental Pulp Cells Treated with TheraCal LC: Functional Analysis of Gene Interactions Compared to MTA" Journal of Clinical Medicine 9, no. 2: 531. https://doi.org/10.3390/jcm9020531
APA StyleNam, O. H., Kim, J.-H., Choi, S. C., & Kim, Y. (2020). Time-Dependent Response of Human Deciduous Tooth-Derived Dental Pulp Cells Treated with TheraCal LC: Functional Analysis of Gene Interactions Compared to MTA. Journal of Clinical Medicine, 9(2), 531. https://doi.org/10.3390/jcm9020531






