A Novel Multi-Biomarker Assay for Non-Invasive Quantitative Monitoring of Kidney Injury
Abstract
1. Introduction
2. Experimental Section
2.1. Patient Selection
2.2. KIT Assay Methods
2.2.1. Sample Processing
2.2.2. KIT Biomarkers
2.3. Statistical Analysis
2.3.1. KIT Score Development
2.3.2. KIT Score Validation
3. Results
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- KDIGO 2012. Clinical Practice Guideline for the Evaluation and Management of Chronic Kidney Disease; KDIGO: Brussels, Belgium, 2013. [Google Scholar]
- Levin, A.; Stevens, P.E. Summary of KDIGO 2012 CKD Guideline: Behind the scenes, need for guidance, and a framework for moving forward. Kidney Int. 2014, 85, 49–61. [Google Scholar] [CrossRef]
- Levey, A.S.; Stevens, L.A.; Schmid, C.H.; Zhang, Y.L.; Castro, A.F.; Feldman, H.I.; Kusek, J.W.; Eggers, P.; Van Lente, F.; Greene, T.; et al. A new equation to estimate glomerular filtration rate. Ann. Intern. Med. 2009, 150, 604–612. [Google Scholar] [CrossRef] [PubMed]
- Khati, N.J.; Gorodenker, J.; Hill, M.C. Ultrasound-guided biopsies of the abdomen. Ultrasound Q. 2011, 27, 255–268. [Google Scholar] [CrossRef] [PubMed]
- Fried, L.F. Kidney Function as a Predictor of Noncardiovascular Mortality. J. Am. Soc. Nephrol. 2005, 16, 3728–3735. [Google Scholar] [CrossRef]
- Qaseem, A.; Hopkins, R.H.; Sweet, D.E.; Starkey, M.; Shekelle, P. Screening, monitoring and treatment of stage 1 to 3 CKD: A clinical practice guideline from the CGC of the ACP. Ann. Intern. Med. 2013, 159, 835–847. [Google Scholar] [PubMed]
- Mishra, J.; Dent, C.; Tarabishi, R.; Mitsnefes, M.M.; Ma, Q.; Kelly, C.; Ruff, S.M.; Zahedi, K.; Shao, M.; Bean, J.; et al. Neutrophil gelatinase-associated lipocalin (NGAL) as a biomarker for acute renal injury after cardiac surgery. Lancet 2005, 365, 1231–1238. [Google Scholar] [CrossRef]
- Parikh, C.R.; Jani, A.; Melnikov, V.Y.; Faubel, S.; Edelstein, C.L. Urinary interleukin-18 is a marker of human acute tubular necrosis. Am. J. Kidney Dis. 2004, 43, 405–414. [Google Scholar] [CrossRef] [PubMed]
- Han, W.K.; Bailly, V.; Abichandani, R.; Thadhani, R.; Bonventre, J.V. Kidney Injury Molecule-1 (KIM-1): A novel biomarker for human renal proximal tubule injury. Kidney Int. 2002, 62, 237–244. [Google Scholar] [CrossRef] [PubMed]
- Matteucci, E.; Carmellini, M.; Bertoni, C.; Boldrini, E.; Mosca, F.; Giampietro, O. Urinary Excretion Rates of Multiple Renal Indicators after Kidney Transplantation: Clinical Significance for Early Graft Outcome. Ren. Fail. 1998, 20, 325–330. [Google Scholar] [CrossRef][Green Version]
- Lybarger, J.A.; Lichtveld, M.Y.; Amler, R.W. Renal Failure Biomedical Testing of the Kidney for Persons Exposed to Hazardous Substances in the Environment. Ren. Fail. 1999, 21, 263–274. [Google Scholar] [CrossRef]
- Donadio, C.; Tramonti, G.; Lucchesi, A.; Giordani, R.; Lucchetti, A.; Bianchi, C. Gamma-glutamyltransferase is a reliable marker for tubular effects of contrast media. Ren. Fail. 1998, 20, 319–324. [Google Scholar] [CrossRef] [PubMed]
- Nouwen, E.J.; De Broe, M.E. Human intestinal versus tissue-nonspecific alkaline phosphatase as complementary urinary markers for the proximal tubule. Kidney Int. Suppl. 1994, 47, S43–S51. [Google Scholar] [PubMed]
- Gatta, A.; Bazzerla, G.; Amodio, P.; Menon, F.; Angeli, P.; Schiaffino, E.; Schmid, C. Detection of the Early Steps of Cadmium Nephropathy—Comparison of Light- and Electron-Microscopical Patterns with the Urinary Enzymes Excretion. Nephron 1989, 51, 20–24. [Google Scholar] [CrossRef] [PubMed]
- Locatelli, F.; Del Vecchio, L.; Pozzoni, P. The importance of early detection of chronic kidney disease. Nephrol. Dial. Transplant. 2002, 17 (Suppl. 11), 2–7. [Google Scholar] [CrossRef]
- Remuzzi, G.; Ruggenenti, P.; Perico, N. Chronic renal diseases: Renoprotective benefits of renin-angiotensin system inhibition. Ann. Intern. Med. 2002, 136, 604–615. [Google Scholar] [CrossRef]
- Gansevoort, R.T. Too much nephrology? The CKD epidemic is real and concerning. A PRO view. Nephrol. Dial. Transplant. 2018, 34, 577–580. [Google Scholar] [CrossRef]
- Coresh, J.; Byrd-Holt, D.; Astor, B.C.; Briggs, J.P.; Eggers, P.W.; Lacher, D.A.; Hostetter, T.H. Chronic kidney disease awareness, prevalence, and trends among U.S. adults, 1999 to 2000. J. Am. Soc. Nephrol. 2005, 16, 180–188. [Google Scholar] [CrossRef] [PubMed]
- Coresh, J.; Wei, G.L.; McQuillan, G.; Brancati, F.L.; Levey, A.S.; Jones, C.; Klag, M.J. Prevalence of high blood pressure and elevated serum creatinine level in the United States: Findings from the third National Health and Nutrition Examination Survey (1988–1994). Arch. Intern. Med. 2001, 161, 1207–1216. [Google Scholar] [CrossRef] [PubMed]
- McClellan, W.M.; Knight, D.F.; Karp, H.; Brown, W.W. Early detection and treatment of renal disease in hospitalized diabetic and hypertensive patients: Important differences between practice and published guidelines. Am. J. Kidney Dis. 1997, 29, 368–375. [Google Scholar] [CrossRef]
- Obrador, G.T.; Ruthazer, R.; Arora, P.; Kausz, A.T.; Pereira, B.J. Prevalence of and factors associated with suboptimal care before initiation of dialysis in the United States. J. Am. Soc. Nephrol. 1999, 10, 1793–1800. [Google Scholar] [PubMed]
- Dugan, J.; Shubrook, J. International Classification of Diseases, 10th Revision, Coding for Diabetes. Clin. Diabetes 2017, 35, 232–238. [Google Scholar] [CrossRef] [PubMed]
- Levey, A.S.; Coresh, J.; Greene, T.; Stevens, L.A.; Zhang, Y.; Hendriksen, S.; Kusek, J.W.; Van Lente, F. Using Standardized Serum Creatinine Values in the Modification of Diet in Renal Disease Study Equation for Estimating Glomerular Filtration Rate. Ann. Intern. Med. 2006, 145, 247. [Google Scholar] [CrossRef] [PubMed]
- Stroun, M.; Lyautey, J.; Lederrey, C.; Olson-Sand, A.; Anker, P. About the possible origin and mechanism of circulating DNA apoptosis and active DNA release. Clin. Chim. Acta 2001, 313, 139–142. [Google Scholar] [CrossRef]
- Sarwal, M.M.; Sigdel, T.K.; Yang, J.Y.C. A Novel Immunoprobe-Based Method to Assess Organ Injury Status through a Biofluid-Based Cell-Free DNA (CFDNA) Assay. WO2018035340A1, 2 February 2018. [Google Scholar]
- Sigdel, T.K.; Vitalone, M.J.; Tran, T.Q.; Dai, H.; Hsieh, S.-C.; Salvatierra, O.; Sarwal, M.M. A rapid noninvasive assay for the detection of renal transplant injury. Transplantation 2013, 96, 97–101. [Google Scholar] [CrossRef] [PubMed]
- Sigdel, T.; Archila, F.; Constantin, T.; Prins, S.; Liberto, J.; Damm, I.; Towfighi, P.; Navarro, S.; Kirkizlar, E.; Demko, Z.; et al. Optimizing Detection of Kidney Transplant Injury by Assessment of Donor-Derived Cell-Free DNA via Massively Multiplex PCR. J. Clin. Med. 2018, 8, 19. [Google Scholar] [CrossRef]
- Bloom, R.D.; Bromberg, J.S.; Poggio, E.D.; Bunnapradist, S.; Langone, A.J.; Sood, P.; Matas, A.J.; Mehta, S.; Mannon, R.B.; Sharfuddin, A.; et al. Cell-Free DNA and Active Rejection in Kidney Allografts. J. Am. Soc. Nephrol. 2017, 28, 2221–2232. [Google Scholar] [CrossRef]
- De Vlaminck, I.; Martin, L.; Kertesz, M.; Patel, K.; Kowarsky, M.; Strehl, C.; Cohen, G.; Luikart, H.; Neff, N.F.; Okamoto, J.; et al. Noninvasive monitoring of infection and rejection after lung transplantation. Proc. Natl. Acad. Sci. USA 2015, 112, 13336–13341. [Google Scholar] [CrossRef]
- De Vlaminck, I.; Valantine, H.A.; Snyder, T.M.; Strehl, C.; Cohen, G.; Luikart, H.; Neff, N.F.; Okamoto, J.; Bernstein, D.; Weisshaar, D.; et al. Circulating cell-free DNA enables noninvasive diagnosis of heart transplant rejection. Sci. Transl. Med. 2014, 6, ra77–ra241. [Google Scholar] [CrossRef]
- Shiels, P.G.; McGuinness, D.; Eriksson, M.; Kooman, J.P.; Stenvinkel, P. The role of epigenetics in renal ageing. Nat. Rev. Nephrol. 2017, 13, 471–482. [Google Scholar] [CrossRef]
- Zhao, Y.; Ding, C.; Xue, W.; Ding, X.; Zheng, J.; Gao, Y.; Xia, X.; Li, S.; Liu, J.; Han, F.; et al. Genome-wide DNA methylation analysis in renal ischemia reperfusion injury. Gene 2017, 610, 32–43. [Google Scholar] [CrossRef]
- Lazzeri, E.; Rotondi, M.; Mazzinghi, B.; Lasagni, L.; Buonamano, A.; Rosati, A.; Pradella, F.; Fossombroni, V.; La Villa, G.; Gacci, M.; et al. High CXCL10 expression in rejected kidneys and predictive role of pretransplant serum CXCL10 for acute rejection and chronic allograft nephropathy. Transplantation 2005, 79, 1215–1220. [Google Scholar] [CrossRef]
- Schaub, S.; Nickerson, P.; Rush, D.; Mayr, M.; Hess, C.; Golian, M.; Stefura, W.; Hayglass, K. Urinary CXCL9 and CXCL10 levels correlate with the extent of subclinical tubulitis. Am. J. Transplant. 2009, 9, 1347–1353. [Google Scholar] [CrossRef]
- Yang, J.Y.C.; Verleden, S.E.; Zarinsefat, A.; Vanaudenaerde, B.M.; Vos, R.; Verleden, G.M.; Sarwal, R.D.; Sigdel, T.K.; Liberto, J.M.; Damm, I.; et al. Cell-Free DNA and CXCL10 Derived from Bronchoalveolar Lavage Predict Lung Transplant Survival. J. Clin. Med. 2019, 8, 9. [Google Scholar] [CrossRef]
- Naesens, M.; Khatri, P.; Li, L.; Sigdel, T.K.; Vitalone, M.J.; Chen, R.; Butte, A.J.; Salvatierra, O.; Sarwal, M.M. Progressive histological damage in renal allografts is associated with expression of innate and adaptive immunity genes. Kidney Int. 2011, 80, 1364–1376. [Google Scholar] [CrossRef]
- Sacreas, A.; Yang, J.Y.C.; Vanaudenaerde, B.M.; Sigdel, T.K.; Liberto, J.M.; Damm, I.; Verleden, G.M.; Vos, R.; Verleden, E.; Sarwal, M.M. The common rejection module in chronic rejection post lung transplantation. PLoS ONE 2018, 13, 1–16. [Google Scholar] [CrossRef]
- Kim, S.S.; Song, S.H.; Kim, J.H.; Jeon, Y.K.; Kim, B.H.; Kang, M.-C.; Chun, S.W.; Hong, S.H.; Chung, M.; Kim, Y.K.; et al. Urine clusterin/apolipoprotein J is linked to tubular damage and renal outcomes in patients with type 2 diabetes mellitus. Clin. Endocrinol. (Oxf.) 2017, 87, 156–164. [Google Scholar] [CrossRef]
- Dieterle, F.; Perentes, E.; Cordier, A.; Roth, D.R.; Verdes, P.; Grenet, O.; Pantano, S.; Moulin, P.; Wahl, D.; Mahl, A.; et al. Urinary clusterin, cystatin C, β2-microglobulin and total protein as markers to detect drug-induced kidney injury. Nat. Biotechnol. 2010, 28, 463–469. [Google Scholar] [CrossRef]
- D’Amico, G.; Bazzi, C. Pathophysiology of proteinuria. Kidney Int. 2003, 63, 809–825. [Google Scholar] [CrossRef]
- Iseki, K.; Ikemiya, Y.; Iseki, C.; Takishita, S. Proteinuria and the risk of developing end-stage renal disease. Kidney Int. 2003, 63, 1468–1474. [Google Scholar] [CrossRef]
- Heymsfield, S.B.; Arteaga, C.; McManus, C.; Smith, J.; Moffitt, S. Measurement of muscle mass in humans: Validity of the 24-hour urinary creatinine method. Am. J. Clin. Nutr. 1983, 37, 478–494. [Google Scholar] [CrossRef]
- Waikar, S.S.; Sabbisetti, V.S.; Bonventre, J.V. Normalization of urinary biomarkers to creatinine during changes in glomerular filtration rate. Kidney Int. 2010, 78, 486–494. [Google Scholar] [CrossRef]
- Bauer, C.; Melamed, M.L.; Hostetter, T.H. Staging of Chronic Kidney Disease: Time for a Course Correction. J. Am. Soc. Nephrol. 2008, 19, 844–846. [Google Scholar] [CrossRef]
- Qin, S.; Rottman, J.B.; Myers, P.; Kassam, N.; Weinblatt, M.; Loetscher, M.; Koch, A.E.; Moser, B.; Mackay, C.R. The chemokine receptors CXCR3 and CCR5 mark subsets of T cells associated with certain inflammatory reactions. J. Clin. Investig. 1998, 101, 746–754. [Google Scholar] [CrossRef]
- Rabant, M.; Amrouche, L.; Lebreton, X.; Aulagnon, F.; Benon, A.; Sauvaget, V.; Bonifay, R.; Morin, L.; Scemla, A.; Delville, M.; et al. Urinary C-X-C Motif Chemokine 10 Independently Improves the Noninvasive Diagnosis of Antibody-Mediated Kidney Allograft Rejection. J. Am. Soc. Nephrol. 2015, 26, 2840–2851. [Google Scholar] [CrossRef]
- Jakiela, B.; Kosałka, J.; Plutecka, H.; Węgrzyn, A.S.; Bazan-Socha, S.; Sanak, M.; Musiał, J. Urinary cytokines and mRNA expression as biomarkers of disease activity in lupus nephritis. Lupus 2018, 27, 1259–1270. [Google Scholar] [CrossRef]
- Tyagi, P.; Killinger, K.; Tyagi, V.; Nirmal, J.; Chancellor, M.; Peters, K.M. Urinary chemokines as noninvasive predictors of ulcerative interstitial cystitis. J. Urol. 2012, 187, 2243–2248. [Google Scholar] [CrossRef]
- Marie, M.A.; Khalil, R.E.A.; Habib, H.M. Urinary CXCL10: A marker of nephritis in lupus patients. Reumatismo 2014, 73, 282–297. [Google Scholar] [CrossRef][Green Version]
- Khatri, P.; Roedder, S.; Kimura, N.; De Vusser, K.; Morgan, A.A.; Gong, Y.; Fischbein, M.P.; Robbins, R.C.; Naesens, M.; Butte, A.J.; et al. A common rejection module (CRM) for acute rejection across multiple organs identifies novel therapeutics for organ transplantation. J. Exp. Med. 2013, 210, 2205–2221. [Google Scholar] [CrossRef]
- Sigdel, T.; Yang, J.; Bestard, O.; Hsieh, S.; Roedder, S.; Damm, I.; Liberto, J.; Nandoe, S.; Sarwal, M. A Non-Invasive Urinary Common Rejection Module (uCRM) Gene Expression Score Quantifies and Differentiates Kidney Transplant Injury. Am. J. Transplant. 2017, 17, 300. [Google Scholar]
- Coresh, J.; Selvin, E.; Stevens, L.A.; Manzi, J.; Kusek, J.W.; Eggers, P.; Van Lente, F.; Levey, A.S. Prevalence of Chronic Kidney Disease in the United States. JAMA 2007, 298, 2038. [Google Scholar] [CrossRef]
- Sandilands, E.A.; Dhaun, N.; Dear, J.W.; Webb, D.J. Measurement of renal function in patients with chronic kidney disease. Br. J. Clin. Pharmacol. 2013, 76, 504–515. [Google Scholar] [CrossRef] [PubMed]
- Delanghe, J.; Speeckaert, M. Preanalytical requirements of urinalysis. Biochem. Med. 2014, 24, 89–104. [Google Scholar] [CrossRef] [PubMed]
- Thomas, C.E.; Sexton, W.; Benson, K.; Sutphen, R.; Koomen, J. Urine Collection and Processing for Protein Biomarker Discovery and Quantification. Cancer Epidemiol. Prev. Biomark. 2010, 19, 953–959. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.Y.C.; Sigdel, T.K.; Sarwal, M.M. Self-antigens and rejection: A proteomic analysis. Curr. Opin. Organ Transplant. 2016, 21, 362–367. [Google Scholar] [CrossRef]
- Sigdel, T.K.; Gao, Y.; He, J.; Wang, A.; Nicora, C.D.; Fillmore, T.L.; Shi, T.; Webb-Robertson, B.-J.; Smith, R.D.; Qian, W.-J.; et al. Mining the human urine proteome for monitoring renal transplant injury. Kidney Int. 2016, 89, 1244–1252. [Google Scholar] [CrossRef]
- Sigdel, T.K.; Lee, S.; Sarwal, M.M. Profiling the proteome in renal transplantation. PROTEOMICS Clin. Appl. 2011, 5, 269–280. [Google Scholar] [CrossRef]
- Sigdel, T.K.; Salomonis, N.; Nicora, C.D.; Ryu, S.; He, J.; Dinh, V.; Orton, D.J.; Moore, R.J.; Hsieh, S.-C.; Dai, H.; et al. The Identification of Novel Potential Injury Mechanisms and Candidate Biomarkers in Renal Allograft Rejection by Quantitative Proteomics. Mol. Cell. Proteom. 2014, 13, 621–631. [Google Scholar] [CrossRef]
- Keslar, K.S.; Lin, M.; Zmijewska, A.A.; Sigdel, T.K.; Tran, T.Q.; Ma, L.; Bhasin, M.; Rao, P.; Ding, R.; Iklé, D.N.; et al. Multicenter evaluation of a standardized protocol for noninvasive gene expression profiling. Am. J. Transplant. 2013, 13, 1891–1897. [Google Scholar] [CrossRef][Green Version]
- Sigdel, T.K.; Kaushal, A.; Gritsenko, M.; Norbeck, A.D.; Qian, W.-J.; Xiao, W.; Camp, D.G.; Smith, R.D.; Sarwal, M.M. Shotgun proteomics identifies proteins specific for acute renal transplant rejection. PROTEOMICS Clin. Appl. 2010, 4, 32–47. [Google Scholar] [CrossRef] [PubMed]
- Sigdel, T.K.; Lau, K.; Schilling, J.; Sarwal, M. Optimizing protein recovery for urinary proteomics, a tool to monitor renal transplantation. Clin. Transplant. 2008, 22, 617–623. [Google Scholar] [CrossRef] [PubMed]
- Sigdel, T.K.; Nicora, C.D.; Hsieh, S.-C.; Dai, H.; Qian, W.-J.; Camp, D.G.; Sarwal, M.M. Optimization for peptide sample preparation for urine peptidomics. Clin. Proteom. 2014, 11, 7. [Google Scholar] [CrossRef]
- Sigdel, T.K.; Ng, Y.W.; Lee, S.; Nicora, C.D.; Qian, W.-J.; Smith, R.D.; Camp, D.G.; Sarwal, M.M. Perturbations in the Urinary Exosome in Transplant Rejection. Front. Med. 2015, 1, 57. [Google Scholar] [CrossRef]
- Hashimoto, T.; Yoshida, K.; Hashimoto, N.; Nakai, A.; Kaneshiro, K.; Suzuki, K.; Kawasaki, Y.; Shibanuma, N.; Hashiramoto, A. Circulating cell free DNA: A marker to predict the therapeutic response for biological DMARDs in rheumatoid arthritis. Int. J. Rheum. Dis. 2017, 20, 722–730. [Google Scholar] [CrossRef]
- Perkins, B.A.; Caskey, C.T.; Brar, P.; Dec, E.; Karow, D.S.; Kahn, A.M.; Hou, Y.-C.C.; Shah, N.; Boeldt, D.; Coughlin, E.; et al. Precision medicine screening using whole-genome sequencing and advanced imaging to identify disease risk in adults. Proc. Natl. Acad. Sci. USA 2018, 201706096. [Google Scholar] [CrossRef]
- Schwarzenbach, H.; Hoon, D.S.B.; Pantel, K. Cell-free nucleic acids as biomarkers in cancer patients. Nat. Rev. Cancer 2011, 11, 426–437. [Google Scholar] [CrossRef]
- Jiang, P.; Lo, Y.M.D. The Long and Short of Circulating Cell-Free DNA and the Ins and Outs of Molecular Diagnostics. Trends Genet. 2016, 32, 360–371. [Google Scholar] [CrossRef]
- Snyder, M.W.; Kircher, M.; Hill, A.J.; Daza, R.M.; Shendure, J. Cell-free DNA Comprises an in Vivo Nucleosome Footprint that Informs Its Tissues-Of-Origin. Cell 2016, 164, 57–68. [Google Scholar] [CrossRef]
- Bechtel, W.; McGoohan, S.; Zeisberg, E.M.; Müller, G.A.; Kalbacher, H.; Salant, D.J.; Müller, C.A.; Kalluri, R.; Zeisberg, M. Methylation determines fibroblast activation and fibrogenesis in the kidney. Nat. Med. 2010, 16, 544–550. [Google Scholar] [CrossRef]
- Tatapudi, R.R.; Muthukumar, T.; Dadhania, D.; Ding, R.; Li, B.; Sharma, V.K.; Lozada-Pastorio, E.; Seetharamu, N.; Hartono, C.; Serur, D.; et al. Noninvasive detection of renal allograft inflammation by measurements of mRNA for IP-10 and CXCR3 in urine. Kidney Int. 2004, 65, 2390–2397. [Google Scholar] [CrossRef]
- Yang, J.Y.C.; Sarwal, M.M. Transplant genetics and genomics. Nat. Rev. Genet. 2017, 18, 309–326. [Google Scholar] [CrossRef]
- Yang, J.Y.C.; Sarwal, R.D.; Sigdel, T.K.; Sarwal, M.M. Predicting Transplant Rejection by a Composite Urinary Injury Score. Am. J. Transplant. 2019, 17 (suppl. 3). [Google Scholar]
- Schulthess, F.T.; Paroni, F.; Sauter, N.S.; Shu, L.; Ribaux, P.; Haataja, L.; Strieter, R.M.; Oberholzer, J.; King, C.C.; Maedler, K. CXCL10 Impairs β Cell Function and Viability in Diabetes through TLR4 Signaling. Cell Metab. 2009, 9, 125–139. [Google Scholar] [CrossRef]
- Antonelli, A.; Fallahi, P.; Rotondi, M.; Ferrari, S.M.; Romagnani, P.; Ghiadoni, L.; Serio, M.; Taddei, S.; Ferrannini, E. High serum levels of CXC chemokine ligand 10 in untreated essential hypertension. J. Hum. Hypertens. 2008, 22, 579–581. [Google Scholar] [CrossRef][Green Version]
- Ghiggeri, G.M.; Bruschi, M.; Candiano, G.; Rastaldi, M.P.; Scolari, F.; Passerini, P.; Musante, L.; Pertica, N.; Caridi, G.; Ferrario, F.; et al. Depletion of clusterin in renal diseases causing nephrotic syndrome. Kidney Int. 2002, 62, 2184–2194. [Google Scholar] [CrossRef]
- Choi, J.H.; Jeong, E.; Youn, B.S.; Kim, M.S. Distinct Ultradian Rhythms in Plasma Clusterin Concentrations in Lean and Obese Korean Subjects. Endocrinol. Metab. 2018, 33, 245–251. [Google Scholar] [CrossRef]
- Judd, E.; Calhoun, D.A. Management of hypertension in CKD: Beyond the guidelines. Adv. Chronic Kidney Dis. 2015, 22, 116–122. [Google Scholar] [CrossRef]
- Intensive blood-glucose control with sulphonylureas or insulin compared with conventional treatment and risk of complications in patients with type 2 diabetes (UKPDS 33). UK Prospective Diabetes Study (UKPDS) Group. Lancet 1998, 352, 837–853. [CrossRef]
- Almaani, S.; Meara, A.; Rovin, B.H. Update on Lupus Nephritis. Clin. J. Am. Soc. Nephrol. 2017, 12, 825–835. [Google Scholar] [CrossRef]
- Sumida, K.; Molnar, M.Z.; Potukuchi, P.K.; Hassan, F.; Thomas, F.; Yamagata, K.; Kalantar-Zadeh, K.; Kovesdy, C.P. Treatment of rheumatoid arthritis with biologic agents lowers the risk of incident chronic kidney disease. Kidney Int. 2018, 93, 1207–1216. [Google Scholar] [CrossRef]
- Seidu, S.; Kunutsor, S.K.; Cos, X.; Gillani, S.; Khunti, K. For and on behalf of Primary Care Diabetes Europe SGLT2 inhibitors and renal outcomes in type 2 diabetes with or without renal impairment: A systematic review and meta-analysis. Prim. Care Diabetes 2018, 12, 265–283. [Google Scholar] [CrossRef]
- Alicic, R.Z.; Johnson, E.J.; Tuttle, K.R. SGLT2 Inhibition for the Prevention and Treatment of Diabetic Kidney Disease: A Review. Am. J. Kidney Dis. 2018, 72, 267–277. [Google Scholar] [CrossRef]
- Breyer, M.D.; Susztak, K. The next generation of therapeutics for chronic kidney disease. Nat. Rev. Drug Discov. 2016, 15, 568–588. [Google Scholar] [CrossRef] [PubMed]
- Lovasik, B.P.; Zhang, R.; Hockenberry, J.M.; Schrager, J.D.; Pastan, S.O.; Mohan, S.; Patzer, R.E. Emergency Department Use and Hospital Admissions Among Patients with End-Stage Renal Disease in the United States. JAMA Intern. Med. 2016, 176, 1563. [Google Scholar] [CrossRef] [PubMed]
- Naesens, M.; Kuypers, D.R.J.; Sarwal, M. Calcineurin inhibitor nephrotoxicity. Clin. J. Am. Soc. Nephrol. 2009, 4, 481–508. [Google Scholar] [CrossRef] [PubMed]
- Stokes, M.B.; Foster, K.; Markowitz, G.S.; Ebrahimi, F.; Hines, W.; Kaufman, D.; Moore, B.; Wolde, D.; D’Agati, V.D. Development of glomerulonephritis during anti-TNF-alpha; therapy for rheumatoid arthritis. Nephrol. Dial. Transplant. 2005, 20, 1400–1406. [Google Scholar] [CrossRef]
- Miller, R.P.; Tadagavadi, R.K.; Ramesh, G.; Reeves, W.B. Mechanisms of Cisplatin nephrotoxicity. Toxins 2010, 2, 2490–2518. [Google Scholar] [CrossRef]
- Kaloyanides, G.J.; Pastoriza-Munoz, E. Aminoglycoside nephrotoxicity. Kidney Int. 1980, 18, 571–582. [Google Scholar] [CrossRef]
- Perazella, M.A.; Shirali, A.C. Nephrotoxicity of Cancer Immunotherapies: Past, Present and Future. J. Am. Soc. Nephrol. 2018, 29, 2039–2052. [Google Scholar] [CrossRef]
- Mohammed, N.M.A.; Mahfouz, A.; Achkar, K.; Rafie, I.M.; Hajar, R. Contrast-induced Nephropathy. Heart Views 2013, 14, 106–116. [Google Scholar] [CrossRef]
- National Kidney Foundation Centers for Disease Control and Prevention. National Chronic Kidney Disease Fact Sheet; National Kidney Foundation: New York, NY, USA, 2017; pp. 1–4.
- Axelrod, D.A.; Schnitzler, M.A.; Xiao, H.; Naik, A.S.; Segev, D.L.; Dharnidharka, V.R.; Brennan, D.C.; Lentine, K.L. The Changing Financial Landscape of Renal Transplant Practice: A National Cohort Analysis. Am. J. Transplant. 2017, 17, 377–389. [Google Scholar] [CrossRef] [PubMed]
- Bowe, B.; Xie, Y.; Li, T.; Mokdad, A.H.; Xian, H.; Yan, Y.; Maddukuri, G.; Al-Aly, Z. Changes in the US Burden of Chronic Kidney Disease from 2002 to 2016. JAMA Netw. Open 2018, 1, e184412. [Google Scholar] [CrossRef]
- Yap, H.K.; Quek, C.M.; Shen, Q.; Joshi, V.; Chia, K.S. Role of urinary screening programmes in children in the prevention of chronic kidney disease. Ann. Acad. Med. Singap. 2005, 34, 3–7. [Google Scholar]
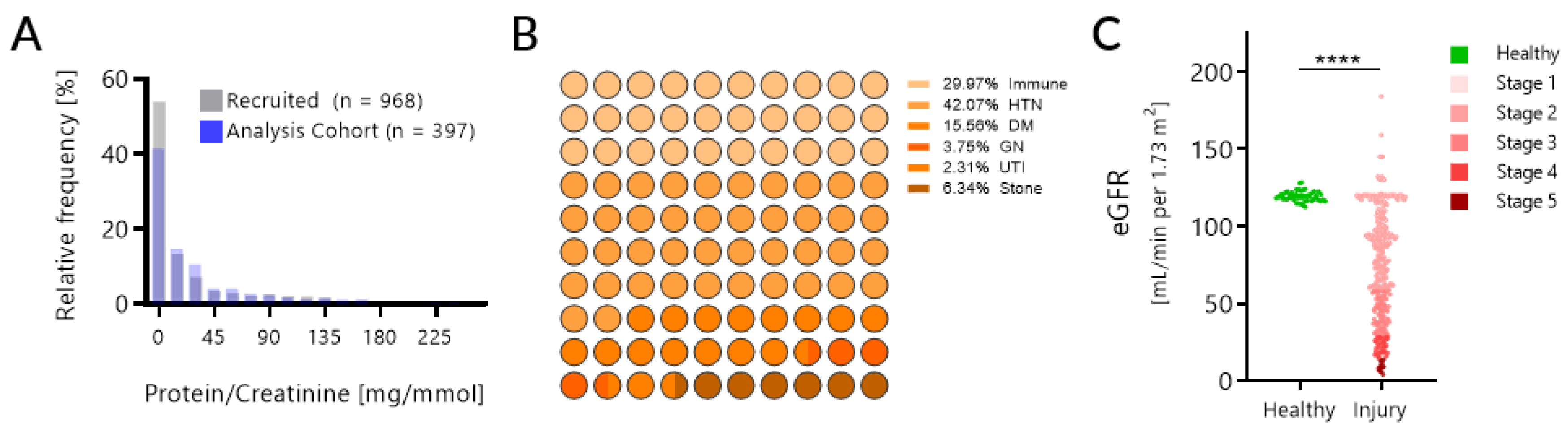
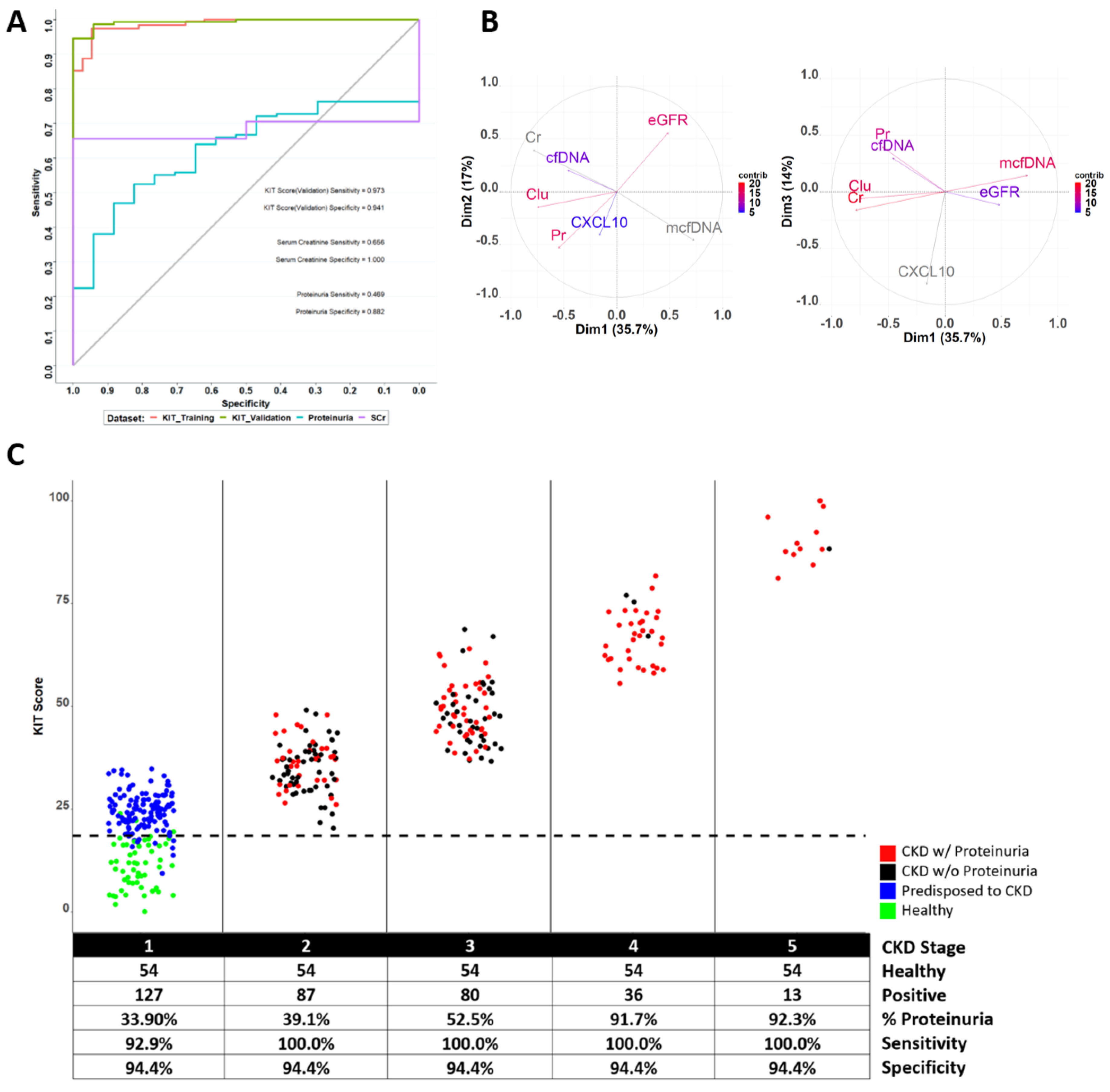
| Variable | Sampled Cohort (n = 397) Median (Range) | Training Cohort (n = 233) Median (Range) | Test Cohort (n = 164) Median (Range) |
|---|---|---|---|
| Age, years | 53 (2–98) | 53 (2–94) | 52 (4–98) |
| Gender, women, % | 49% | 47% | 53% |
| Race, % AA | 13.2% | 12.9% | 13.7% |
| Proteinuria, (mg/mmol creatinine) | 74.56 (0–8239) | 76.58 (0–8239) | 62.12 (0–3135) |
| Serum creatinine, (mg/dL) | 0.98 (0.31–9.36) | 1.06 (0.31–9.36) | 0.83 (0.32–7.13) |
| Estimated glomerular filtration rate (eGFR) (mL/min/1.73 m2) | 85 (4–184) | 73 (4–159) | 94 (6–184) |
| Cause of kidney injury, % | |||
| • Immunological | 30.0% | 34.9% | 22.5% |
| • Hypertension | 42.1% | 42.6% | 41.3% |
| • Diabetes | 15.6% | 14.8% | 16.7% |
| • Glomerulonephritis (GN) | 3.7% | 1.0% | 8.0% |
| • Urinary Tract Infection (UTI) | 2.3% | 1.9% | 2.9% |
| • Kidney Stone | 6.3% | 4.8% | 8.7% |
| Stage of chronic kidney disease (CKD), % | |||
| • Healthy | 15.6% | 15.3% | 15.9% |
| • Stage 1 | 36.6% | 28.2% | 49.3% |
| • Stage 2 | 25.1% | 26.8% | 22.5% |
| • Stage 3 | 23.1% | 26.3% | 18.1% |
| • Stage 4 | 10.4% | 12.4% | 7.2% |
| • Stage 5 | 3.7% | 5.3% | 1.4% |
| Parameter | Estimate | df | s.e. | χ2 | p-Value | OR | 2.5% | 97.5% |
|---|---|---|---|---|---|---|---|---|
| Intercept | 44.6953 | 1 | 12.8490 | 12.0999 | 0.0005 | |||
| eGFR | −8.9508 | 1 | 2.6815 | 11.1422 | 0.0008 | 1.30 × 10−4 | 1.70 × 10−7 | 9.65 × 10−3 |
| Proteinuria | 0.4971 | 1 | 0.2787 | 3.1815 | 0.0745 | 1.64 | 1.01 | 3.03 |
| Parameter | Estimate | df | s.e. | χ2 | p-Value | OR | 2.5% | 97.5% |
|---|---|---|---|---|---|---|---|---|
| Intercept | −27.1034 | 1 | 29.8369 | 0.8245 | 0.3637 | |||
| eGFR | 3.5354 | 1 | 5.5596 | 0.4045 | 0.5248 | 34.3 | 2.91 × 10−4 | 8.38 × 106 |
| Proteinuria | 1.3151 | 1 | 0.8043 | 2.6732 | 0.1020 | 3.73 | 1.01 | 27.22 |
| KIT Score | 0.7847 | 1 | 0.2834 | 7.6674 | 0.0056 | 2.19 | 1.48 | 4.75 |
| Parameter | Estimate | df | s.e. | χ2 | p-Value | OR |
|---|---|---|---|---|---|---|
| Intercept | 56.0716 | 1 | 12.9866 | 18.6451 | <0.0001 | |
| eGFR | −12.5302 | 1 | 2.7181 | 21.2521 | <0.0001 | 3.62 × 10−6 |
| Urine cfDNA | −0.2720 | 1 | 0.0973 | 7.8120 | 0.0052 | 0.76 |
| Urine m-cfDNA | −1.1260 | 1 | 0.2655 | 17.9946 | <0.0001 | 0.32 |
| Urine protein | 0.7976 | 1 | 0.1815 | 19.3702 | <0.0001 | 2.22 |
| Urine CXCL10 | 1.1304 | 1 | 0.5216 | 4.6959 | 0.0302 | 3.10 |
| Urine clusterin | −0.3506 | 1 | 0.2538 | 1.9099 | 0.1671 | 0.70 |
| Urine creatinine | 0.6448 | 1 | 0.5613 | 1.3202 | 0.2506 | 1.91 |
| CKD Stage by eGFR | Mean KIT Score/CKD Stage | % Patients w/o Proteinuria | % Patients w/o Proteinuria Who Have High-Risk for Kidney Injury (KIT Score > 18.5) |
|---|---|---|---|
| CKD Stage 1 | 24.4 | 66% (84/127) | 91% (77/84) |
| CKD Stage 2 | 35.3 | 60% (53/87) | 100% |
| CKD Stage 3 | 48.4 | 47% (38/80) | 100% |
| CKD Stage 4 | 66.9 | 8% (3/36) | 100% |
| CKD Stage 5 | 90.8 | 7% (1/13) | 100% |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Watson, D.; Yang, J.Y.C.; Sarwal, R.D.; Sigdel, T.K.; Liberto, J.M.; Damm, I.; Louie, V.; Sigdel, S.; Livingstone, D.; Soh, K.; et al. A Novel Multi-Biomarker Assay for Non-Invasive Quantitative Monitoring of Kidney Injury. J. Clin. Med. 2019, 8, 499. https://doi.org/10.3390/jcm8040499
Watson D, Yang JYC, Sarwal RD, Sigdel TK, Liberto JM, Damm I, Louie V, Sigdel S, Livingstone D, Soh K, et al. A Novel Multi-Biomarker Assay for Non-Invasive Quantitative Monitoring of Kidney Injury. Journal of Clinical Medicine. 2019; 8(4):499. https://doi.org/10.3390/jcm8040499
Chicago/Turabian StyleWatson, Drew, Joshua Y. C. Yang, Reuben D. Sarwal, Tara K. Sigdel, Juliane M. Liberto, Izabella Damm, Victoria Louie, Shristi Sigdel, Devon Livingstone, Katherine Soh, and et al. 2019. "A Novel Multi-Biomarker Assay for Non-Invasive Quantitative Monitoring of Kidney Injury" Journal of Clinical Medicine 8, no. 4: 499. https://doi.org/10.3390/jcm8040499
APA StyleWatson, D., Yang, J. Y. C., Sarwal, R. D., Sigdel, T. K., Liberto, J. M., Damm, I., Louie, V., Sigdel, S., Livingstone, D., Soh, K., Chakraborty, A., Liang, M., Lin, P.-C., & Sarwal, M. M. (2019). A Novel Multi-Biomarker Assay for Non-Invasive Quantitative Monitoring of Kidney Injury. Journal of Clinical Medicine, 8(4), 499. https://doi.org/10.3390/jcm8040499






