13-gene DNA Methylation Analysis from Oral Brushing: A Promising Non Invasive Tool in the Follow-up of Oral Cancer Patients
Abstract
1. Introduction
2. Experimental Section
2.1. Ethics Statement
2.2. Population Study
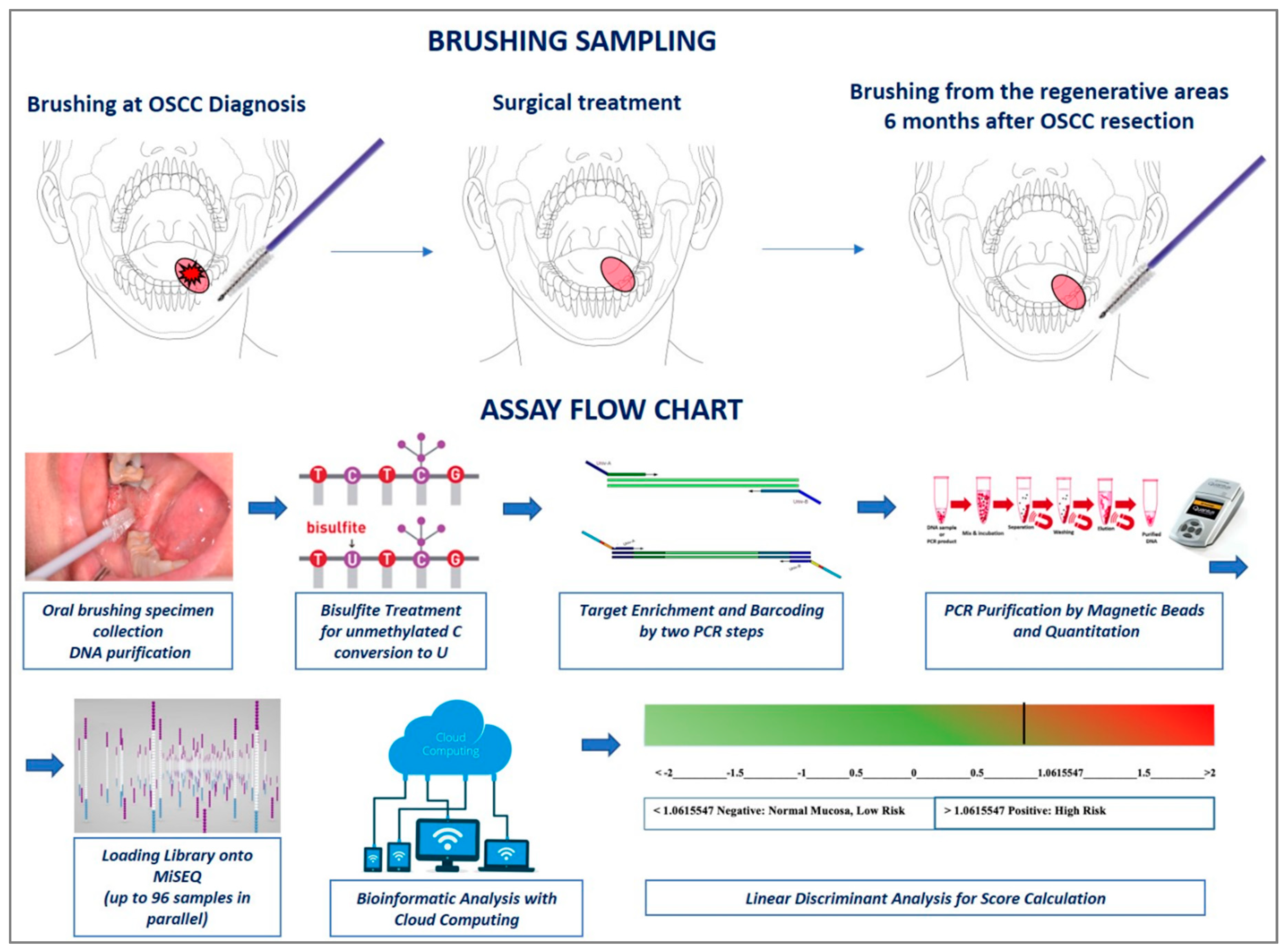
2.3. DNA Methylation Analysis
2.4. Statistical Analysis
3. Results
3.1. Gene DNA Methylation Analysis
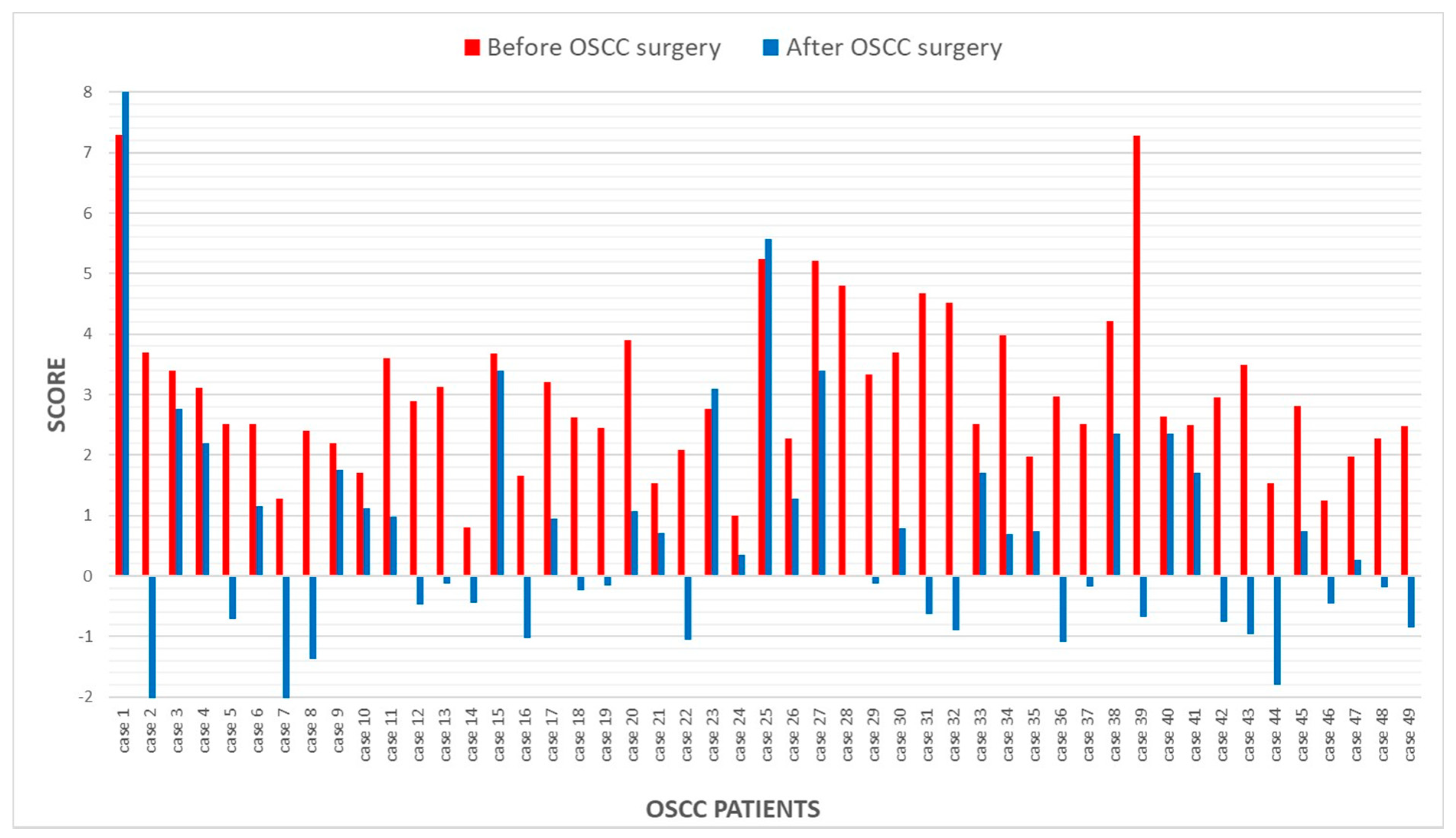
3.1.1. Before Surgery
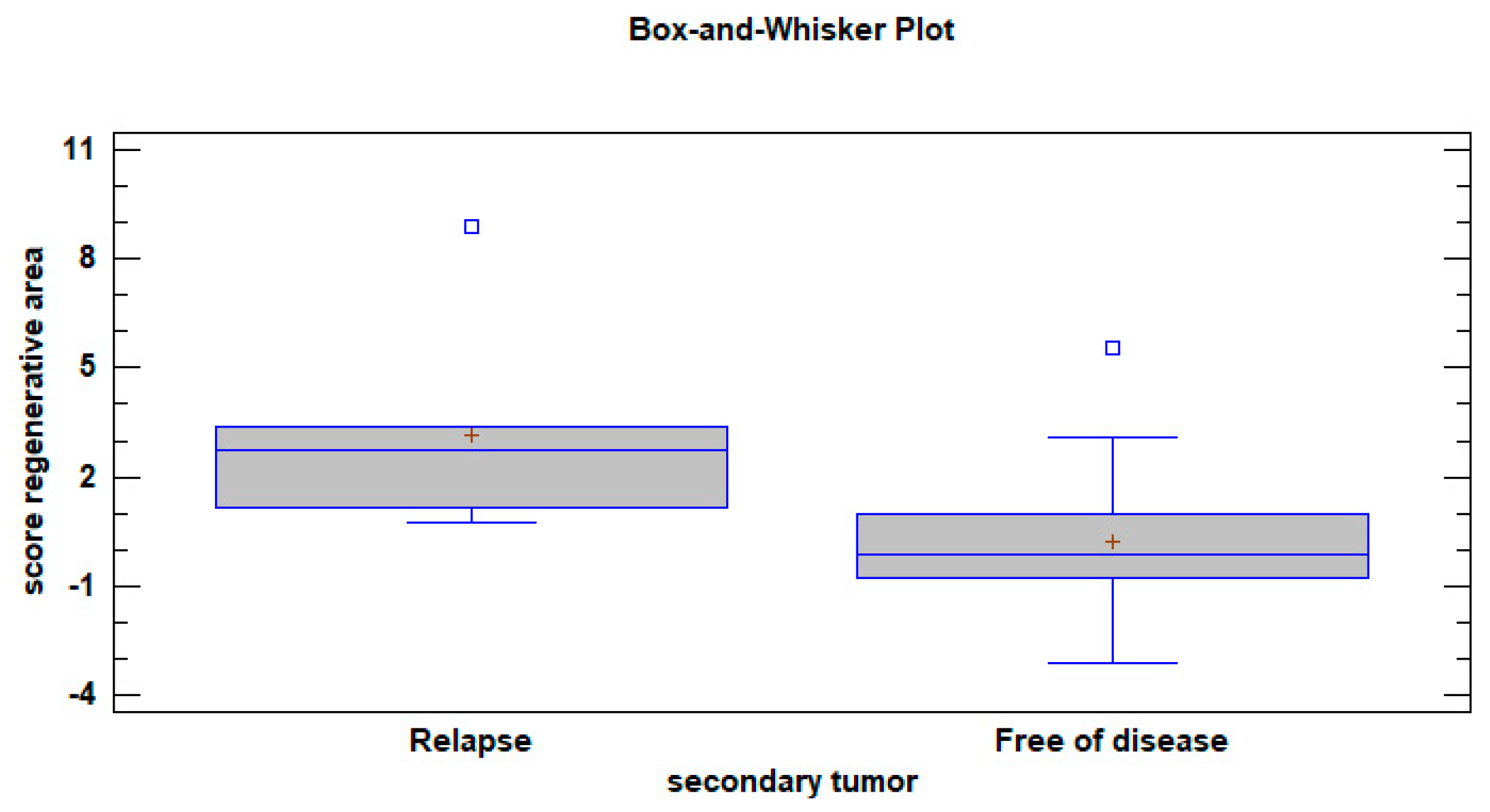
3.1.2. After Surgery
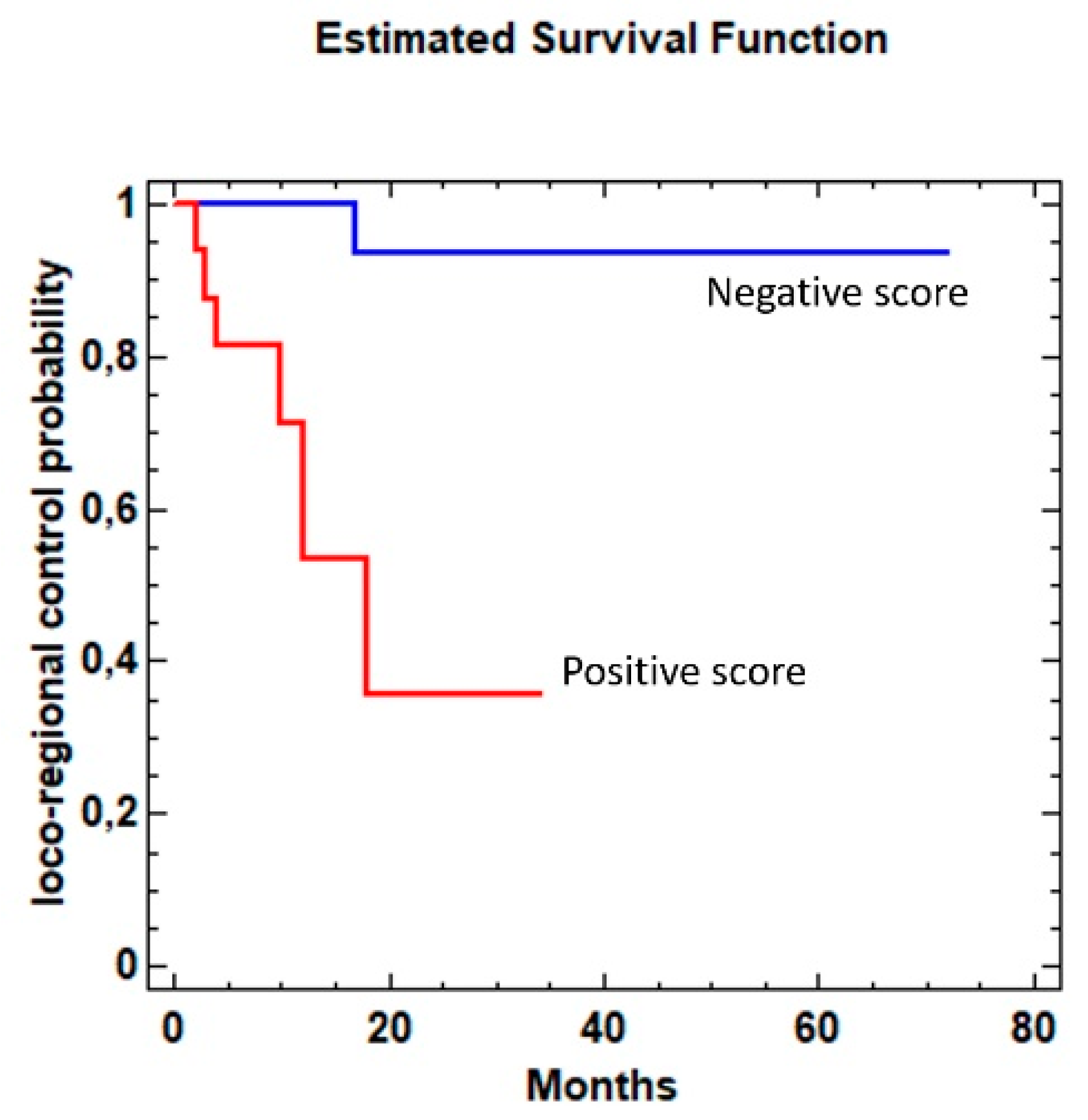
3.2. Loco-Regional Disease-Free Survival
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Cancer of the Oral Cavity and Pharynx—Cancer Stat Facts. Available online: https://seer.cancer.gov/statfacts/html/oralcav.html (accessed on 20 May 2019).
- Braakhuis, B.J.M.; Tabor, M.P.; Leemans, C.R.; van der Waal, I.; Snow, G.B.; Brakenhoff, R.H. Second primary tumors and field cancerization in oral and oropharyngeal cancer: Molecular techniques provide new insights and definitions. Head Neck 2002, 24, 198–206. [Google Scholar] [CrossRef]
- Leemans, C.R.; Tiwari, R.; Nauta, J.J.; van der Waal, I.; Snow, G.B. Recurrence at the primary site in head and neck cancer and the significance of neck lymph node metastases as a prognostic factor. Cancer 1994, 73, 187–190. [Google Scholar] [CrossRef]
- Haas, I.; Hauser, U.; Ganzer, U. The dilemma of follow-up in head and neck cancer patients. Eur. Arch. Otorhinolaryngol. 2001, 258, 177–183. [Google Scholar] [CrossRef]
- Brands, M.T.; Brennan, P.A.; Verbeek, A.L.M.; Merkx, M.A.W.; Geurts, S.M.E. Follow-up after curative treatment for oral squamous cell carcinoma. A critical appraisal of the guidelines and a review of the literature. Eur. J. Surg. Oncol. 2018, 44, 559–565. [Google Scholar] [CrossRef]
- Ho, A.S.; Kraus, D.H.; Ganly, I.; Lee, N.Y.; Shah, J.P.; Morris, L.G.T. Decision making in the management of recurrent head and neck cancer. Head Neck 2014, 36, 144–151. [Google Scholar] [CrossRef]
- De Bree, R.; van der Putten, L.; Brouwer, J.; Castelijns, J.A.; Hoekstra, O.S.; Leemans, C.R. Detection of locoregional recurrent head and neck cancer after (chemo)radiotherapy using modern imaging. Oral Oncol. 2009, 45, 386–393. [Google Scholar] [CrossRef] [PubMed]
- Shaw, R.J.; Hobkirk, A.J.; Nikolaidis, G.; Woolgar, J.A.; Triantafyllou, A.; Brown, J.S.; Liloglou, T.; Risk, J.M. Molecular staging of surgical margins in oral squamous cell carcinoma using promoter methylation of p16(INK4A), cytoglobin, E-cadherin, and TMEFF2. Ann. Surg. Oncol. 2013, 20, 2796–2802. [Google Scholar] [CrossRef][Green Version]
- Schussel, J.; Zhou, X.C.; Zhang, Z.; Pattani, K.; Bermudez, F.; Jean-Charles, G.; McCaffrey, T.; Padhya, T.; Phelan, J.; Spivakovsky, S.; et al. EDNRB and DCC salivary rinse hypermethylation has a similar performance as expert clinical examination in discrimination of oral cancer/dysplasia versus benign lesions. Clin. Cancer Res. 2013, 19, 3268–3275. [Google Scholar] [CrossRef] [PubMed]
- Morandi, L.; Gissi, D.; Tarsitano, A.; Asioli, S.; Monti, V.; Del Corso, G.; Marchetti, C.; Montebugnoli, L.; Foschini, M.P. DNA methylation analysis by bisulfite next-generation sequencing for early detection of oral squamous cell carcinoma and high-grade squamous intraepithelial lesion from oral brushing. J. Craniomaxillofac. Surg. 2015, 43, 1494–1500. [Google Scholar] [CrossRef] [PubMed]
- Demokan, S.; Chang, X.; Chuang, A.; Mydlarz, W.K.; Kaur, J.; Huang, P.; Khan, Z.; Khan, T.; Ostrow, K.L.; Brait, M.; et al. KIF1A and EDNRB are differentially methylated in primary HNSCC and salivary rinses. Int. J. Cancer 2010, 127, 2351–2359. [Google Scholar] [CrossRef] [PubMed]
- Langevin, S.M.; Stone, R.A.; Bunker, C.H.; Grandis, J.R.; Sobol, R.W.; Taioli, E. MicroRNA-137 promoter methylation in oral rinses from patients with squamous cell carcinoma of the head and neck is associated with gender and body mass index. Carcinogenesis 2010, 31, 864–870. [Google Scholar] [CrossRef] [PubMed]
- Pattani, K.M.; Zhang, Z.; Demokan, S.; Glazer, C.; Loyo, M.; Goodman, S.; Sidransky, D.; Bermudez, F.; Jean-Charles, G.; McCaffrey, T.; et al. Endothelin receptor type B gene promoter hypermethylation in salivary rinses is independently associated with risk of oral cavity cancer and premalignancy. Cancer Prev. Res. 2010, 3, 1093–1103. [Google Scholar] [CrossRef] [PubMed]
- Nagata, S.; Hamada, T.; Yamada, N.; Yokoyama, S.; Kitamoto, S.; Kanmura, Y.; Nomura, M.; Kamikawa, Y.; Yonezawa, S.; Sugihara, K. Aberrant DNA methylation of tumor-related genes in oral rinse: A noninvasive method for detection of oral squamous cell carcinoma. Cancer 2012, 118, 4298–4308. [Google Scholar] [CrossRef] [PubMed]
- Morandi, L.; Gissi, D.; Tarsitano, A.; Asioli, S.; Gabusi, A.; Marchetti, C.; Montebugnoli, L.; Foschini, M.P. CpG location and methylation level are crucial factors for the early detection of oral squamous cell carcinoma in brushing samples using bisulfite sequencing of a 13-gene panel. Clin. Epigenet. 2017, 9, 85. [Google Scholar] [CrossRef]
- Rettori, M.M.; de Carvalho, A.C.; Bomfim Longo, A.L.; de Oliveira, C.Z.; Kowalski, L.P.; Carvalho, A.L.; Vettore, A.L. Prognostic significance of TIMP3 hypermethylation in post-treatment salivary rinse from head and neck squamous cell carcinoma patients. Carcinogenesis 2013, 34, 20–27. [Google Scholar] [CrossRef]
- Wiklund, E.D.; Gao, S.; Hulf, T.; Sibbritt, T.; Nair, S.; Costea, D.E.; Villadsen, S.B.; Bakholdt, V.; Bramsen, J.B.; Sørensen, J.A.; et al. MicroRNA alterations and associated aberrant DNA methylation patterns across multiple sample types in oral squamous cell carcinoma. PLoS ONE 2011, 6, e27840. [Google Scholar] [CrossRef]
- Viet, C.T.; Schmidt, B.L. Methylation array analysis of preoperative and postoperative saliva DNA in oral cancer patients. Cancer Epidemiol. Prev. Biomark. 2008, 17, 3603–3611. [Google Scholar] [CrossRef]
- Cheng, Y.-S.L.; Rees, T.; Wright, J. A review of research on salivary biomarkers for oral cancer detection. Clin. Transl. Med. 2014, 3, 3. [Google Scholar] [CrossRef]
- Kademani, D.; Bell, R.B.; Bagheri, S.; Holmgren, E.; Dierks, E.; Potter, B.; Homer, L. Prognostic factors in intraoral squamous cell carcinoma: The influence of histologic grade. J. Oral Maxillofac. Surg. 2005, 63, 1599–1605. [Google Scholar] [CrossRef]
- Gissi, D.B.; Morandi, L.; Gabusi, A.; Tarsitano, A.; Marchetti, C.; Cura, F.; Palmieri, A.; Montebugnoli, L.; Asioli, S.; Foschini, M.P.; et al. A Noninvasive Test for MicroRNA Expression in Oral Squamous Cell Carcinoma. Int. J. Mol. Sci. 2018, 19, 1789. [Google Scholar] [CrossRef]
- Amin, M.B.; Edge, S. AJCC Cancer Staging Manual, 8th ed.; Greene, F., Byrd, D.R., Brookland, R.K., Washington, M.K., Gershenwald, J.E., Compton, C.C., Hess, K.R., Sullivan, D.C., Jessup, J.M., Brierley, J.D., et al., Eds.; Springer International Publishing: New York, NY, USA, 2017; ISBN 978-3-319-40617-6. [Google Scholar]
- Van den Brekel, M.W.; Stel, H.V.; Castelijns, J.A.; Nauta, J.J.; van der Waal, I.; Valk, J.; Meyer, C.J.; Snow, G.B. Cervical lymph node metastasis: Assessment of radiologic criteria. Radiology 1990, 177, 379–384. [Google Scholar] [CrossRef] [PubMed]
- Helliwell, T.R.; Woolgar, J.A. Datasets for Histopathology Reports on Head and Neck Carcinoma and Salivary Neoplasms, 2nd ed.; Royal College of Pathologists: London, UK, 2005. [Google Scholar]
- Morandi, L.; Righi, A.; Maletta, F.; Rucci, P.; Pagni, F.; Gallo, M.; Rossi, S.; Caporali, L.; Sapino, A.; Lloyd, R.V.; et al. Somatic mutation profiling of hobnail variant of papillary thyroid carcinoma. Endocr. Relat. Cancer 2017, 24, 107–117. [Google Scholar] [CrossRef] [PubMed]
- Afgan, E.; Baker, D.; Batut, B.; van den Beek, M.; Bouvier, D.; Cech, M.; Chilton, J.; Clements, D.; Coraor, N.; Grüning, B.A.; et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2018 update. Nucleic Acids Res. 2018, 46, W537–W544. [Google Scholar] [CrossRef] [PubMed]
- Hu, K.; Ting, A.H.; Li, J. BSPAT: A fast online tool for DNA methylation co-occurrence pattern analysis based on high-throughput bisulfite sequencing data. BMC Bioinform. 2015, 16, 220. [Google Scholar] [CrossRef] [PubMed]
- Krainer, J.; Weinhäusel, A.; Hanak, K.; Pulverer, W.; Özen, S.; Vierlinger, K.; Pabinger, S. EPIC-TABSAT: Analysis tool for targeted bisulfite sequencing experiments and array-based methylation studies. Nucleic Acids Res. 2019, 47, W166–W170. [Google Scholar] [CrossRef]
- Gruntman, E.; Qi, Y.; Slotkin, R.K.; Roeder, T.; Martienssen, R.A.; Sachidanandam, R. Kismeth: Analyzer of plant methylation states through bisulfite sequencing. BMC Bioinform. 2008, 9, 371. [Google Scholar] [CrossRef]
- Slaughter, D.P.; Southwick, H.W.; Smejkal, W. Field cancerization in oral stratified squamous epithelium; clinical implications of multicentric origin. Cancer 1953, 6, 963–968. [Google Scholar] [CrossRef]
- Gissi, D.B.; Tarsitano, A.; Leonardi, E.; Gabusi, A.; Neri, F.; Marchetti, C.; Montebugnoli, L.; Foschini, M.P.; Morandi, L. Clonal analysis as a prognostic factor in multiple oral squamous cell carcinoma. Oral Oncol. 2017, 67, 131–137. [Google Scholar] [CrossRef]
- Braakhuis, B.J.M.; Bloemena, E.; Leemans, C.R.; Brakenhoff, R.H. Molecular analysis of surgical margins in head and neck cancer: More than a marginal issue. Oral Oncol. 2010, 46, 485–491. [Google Scholar] [CrossRef]
- Barrera, J.E.; Ai, H.; Pan, Z.; Meyers, A.D.; Varella-Garcia, M. Malignancy detection by molecular cytogenetics in clinically normal mucosa adjacent to head and neck tumors. Arch. Otolaryngol. Head Neck Surg. 1998, 124, 847–851. [Google Scholar] [CrossRef]
- Preuss, S.F.; Brieger, J.; Essig, E.K.; Stenzel, M.J.; Mann, W.J. Quantitative DNA measurement in oropharyngeal squamous cell carcinoma and surrounding mucosa. ORL J. Otorhinolaryngol. Relat. Spec. 2004, 66, 320–324. [Google Scholar] [CrossRef] [PubMed]
- Szukała, K.; Brieger, J.; Bruch, K.; Biczysko, W.; Wierzbicka, M.; Szyfter, W.; Szyfter, K. Loss of heterozygosity on chromosome arm 13q in larynx cancer patients: Analysis of tumor, margin and clinically unchanged mucosa. Med. Sci. Monit. 2004, 10, CR233–CR240. [Google Scholar] [PubMed]
- Gissi, D.B.; Gabusi, A.; Tarsitano, A.; Badiali, G.; Marchetti, C.; Morandi, L.; Foschini, M.P.; Montebugnoli, L. Ki67 Overexpression in mucosa distant from oral carcinoma: A poor prognostic factor in patients with long-term follow-up. J. Craniomaxillofac. Surg. 2016, 44, 1430–1435. [Google Scholar] [CrossRef] [PubMed]
- Montebugnoli, L.; Badiali, G.; Marchetti, C.; Cervellati, F.; Farnedi, A.; Foschini, M.P. Prognostic value of Ki67 from clinically and histologically “normal” distant mucosa in patients surgically treated for oral squamous cell carcinoma: A prospective study. Int. J. Oral Maxillofac. Surg. 2009, 38, 1165–1172. [Google Scholar] [CrossRef]
- Montebugnoli, L.; Gissi, D.B.; Badiali, G.; Marchetti, C.; Cervellati, F.; Farnedi, A.; Foschini, M.P. Ki-67 from clinically and histologically “normal” distant mucosa as prognostic marker in early-stage (T1-T2N0) oral squamous cell carcinoma: A prospective study. J. Oral Maxillofac. Surg. 2011, 69, 2579–2584. [Google Scholar] [CrossRef]
- Eljabo, N.; Nikolic, N.; Carkic, J.; Jelovac, D.; Lazarevic, M.; Tanic, N.; Milasin, J. Genetic and epigenetic alterations in the tumour, tumour margins, and normal buccal mucosa of patients with oral cancer. Int. J. Oral Maxillofac. Surg. 2018, 47, 976–982. [Google Scholar] [CrossRef]
- Guerrero-Preston, R.; Michailidi, C.; Marchionni, L.; Pickering, C.R.; Frederick, M.J.; Myers, J.N.; Yegnasubramanian, S.; Hadar, T.; Noordhuis, M.G.; Zizkova, V.; et al. Key tumor suppressor genes inactivated by “greater promoter” methylation and somatic mutations in head and neck cancer. Epigenetics 2014, 9, 1031–1046. [Google Scholar] [CrossRef]
- Claus, R.; Lucas, D.M.; Ruppert, A.S.; Williams, K.E.; Weng, D.; Patterson, K.; Zucknick, M.; Oakes, C.C.; Rassenti, L.Z.; Greaves, A.W.; et al. Validation of ZAP-70 methylation and its relative significance in predicting outcome in chronic lymphocytic leukemia. Blood 2014, 124, 42–48. [Google Scholar] [CrossRef]
- Lee, D.D.; Leão, R.; Komosa, M.; Gallo, M.; Zhang, C.H.; Lipman, T.; Remke, M.; Heidari, A.; Nunes, N.M.; Apolónio, J.D.; et al. DNA hypermethylation within TERT promoter upregulates TERT expression in cancer. J. Clin. Investig. 2019, 129, 1801. [Google Scholar] [CrossRef]
- Sheffield, N.C.; Pierron, G.; Klughammer, J.; Datlinger, P.; Schönegger, A.; Schuster, M.; Hadler, J.; Surdez, D.; Guillemot, D.; Lapouble, E.; et al. DNA methylation heterogeneity defines a disease spectrum in Ewing sarcoma. Nat. Med. 2017, 23, 386–395. [Google Scholar] [CrossRef]
- Righini, C.A.; de Fraipont, F.; Timsit, J.-F.; Faure, C.; Brambilla, E.; Reyt, E.; Favrot, M.-C. Tumor-specific methylation in saliva: A promising biomarker for early detection of head and neck cancer recurrence. Clin. Cancer Res. 2007, 13, 1179–1185. [Google Scholar] [CrossRef] [PubMed]
- Langevin, S.M.; Eliot, M.; Butler, R.A.; Cheong, A.; Zhang, X.; McClean, M.D.; Koestler, D.C.; Kelsey, K.T. CpG island methylation profile in non-invasive oral rinse samples is predictive of oral and pharyngeal carcinoma. Clin. Epigenet. 2015, 7, 125. [Google Scholar] [CrossRef] [PubMed]
- Guerrero-Preston, R.; Soudry, E.; Acero, J.; Orera, M.; Moreno-López, L.; Macía-Colón, G.; Jaffe, A.; Berdasco, M.; Ili-Gangas, C.; Brebi-Mieville, P.; et al. NID2 and HOXA9 promoter hypermethylation as biomarkers for prevention and early detection in oral cavity squamous cell carcinoma tissues and saliva. Cancer Prev. Res. 2011, 4, 1061–1072. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, A.L.; Jeronimo, C.; Kim, M.M.; Henrique, R.; Zhang, Z.; Hoque, M.O.; Chang, S.; Brait, M.; Nayak, C.S.; Jiang, W.-W.; et al. Evaluation of promoter hypermethylation detection in body fluids as a screening/diagnosis tool for head and neck squamous cell carcinoma. Clin. Cancer Res. 2008, 14, 97–107. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.; Zaboli, D.; Wang, H.; Liu, Y.; Arnaoutakis, D.; Khan, T.; Khan, Z.; Koch, W.M.; Califano, J.A. Detection of TIMP3 promoter hypermethylation in salivary rinse as an independent predictor of local recurrence-free survival in head and neck cancer. Clin. Cancer Res. 2012, 18, 1082–1091. [Google Scholar] [CrossRef]
| Patient | Site of Index Tumor | Site of Secondary Tumor | TNM Stage of Index Tumor | Perineural Invasion | Vascular Invasion | Histological Grade | Status of Surgical Margin | Presence of Concomitant OPMD | DNA Methylation Score in OSCC Sample (pre-treatment) | DNA Methylation Score in Regenerative Area (post-treatment) |
|---|---|---|---|---|---|---|---|---|---|---|
| Case 1 | Palate | Palate | T1N0M0 | Yes | Yes | G2 | Clear | No | 7.29 (POSITIVE) | 8.88 (POSITIVE) |
| Case 3 | Superior gum/hard palate | Superior gum/hard palate | T2N2bM0 | No | No | G2 | Clear | No | 3.39 (POSITIVE) | 2.76 (POSITIVE) |
| Case 6 | Superior gum/hard palate | Superior gum/hard palate | T1N0M0 | No | No | G2 | Close | Lichen | 2.51 (POSITIVE) | 1.15 (POSITIVE) |
| Case 15 | Right tongue | Left tongue | T2N3M0 | Yes | No | G2 | Clear | No | 3.68 (POSITIVE) | 3.39 (POSITIVE) |
| Case 27 | Superior gum/palate | palate | T2N0M0 | Yes | Yes | G3 | Clear | No | 5.21 (POSITIVE) | 3.38 (POSITIVE) |
| Case 33 | Hard palate | Hard palate | T1N0M0 | No | No | G1 | Clear | Lichen | 2.51 (POSITIVE) | 1.69 (POSITIVE) |
| Case 35 | Right Cheek | Lymph node metastasis | T3N0M0 | No | No | G3 | Clear | No | 1.97 (POSITIVE) | 0.73 (NEGATIVE) |
| Clinico-Pathological Variables | Patients | Positive Score Regenerative Mucosa | p | Relapse (LR, SPT or LNM) | ||
|---|---|---|---|---|---|---|
| N Patients | % Patients | p | ||||
| Sex | ||||||
| Male | 19 | 4 | p = 0.07 | 1 | 5.3% | p = 0.06 |
| Female | 30 | 12 | 6 | 20% | ||
| Age | ||||||
| < 65 | 18 | 4 | p = 0.234 | 1 | 5.5% | p = 0.07 |
| > 65 | 31 | 12 | 6 | 19.5% | ||
| Site | ||||||
| Tongue | 17 | 3 | p = 0.07 | 1 | 5.9% | p = 0.07 |
| Floor of mouth | 1 | 0 | 0 | 0 | ||
| Cheek | 4 | 0 | 1 | 25% | ||
| Soft palate | 0 | 0 | 0 | 0 | ||
| Superior gum/hard palate | 7 | 5 | 2 | 28.6% | ||
| Inferior gum | 19 | 8 | 3 | 15.8% | ||
| Lip | 1 | 0 | 0 | 0 | ||
| Grading | ||||||
| G1 | 20 | 7 | p = 0.37 | 1 | 5% | p = 0.06 |
| G2 | 21 | 8 | 6 | 28.6% | ||
| G3 | 8 | 1 | 0 | 0 | ||
| T stage | ||||||
| T1–2 = 0 | 28 | 9 | p = 0.93 | 6 | 21.4% | p = 0.08 |
| T3–4 = 1 | 21 | 7 | 1 | 4.7% | ||
| N positivity | ||||||
| N0 | 42 | 14 | p = 0.81 | 6 | 14.3% | p = 0.93 |
| N+ | 7 | 2 | 1 | 14.3% | ||
| Perineural invasion | ||||||
| No | 42 | 13 | p = 0.53 | 4 | 9.5% | p = 0.0004 * |
| Yes | 7 | 3 | 3 | 42.8% | ||
| Vascular invasion | ||||||
| No | 46 | 13 | p = 0.06 | 5 | 10.9% | p = 0.000006 * |
| Yes | 3 | 3 | 2 | 66.7% | ||
| Margin of resection tumor | ||||||
| Clear | 45 | 14 | p = 0.35 | 6 | 13.3% | p = 0.48 |
| Close | 3 | 2 | 1 | 33.4% | ||
| Dysplasia | 1 | 0 | 0 | 0 | ||
| Presence of OPMD | ||||||
| No | 42 | 13 | p = 0.26 | 5 | 11.9% | p = 0.38 |
| Oral Lichen Planus | 5 | 3 | 2 | 40% | ||
| Leukoplakia | 2 | 0 | 0 | 0 | ||
| Adjuvant Radiotherapy | ||||||
| No | 27 | 9 | p = 0.91 | 3 | 11.1% | p = 0.58 |
| Yes | 22 | 7 | 4 | 18.2% | ||
| Reconstructive flap | ||||||
| No | 23 | 7 | p = 0.75 | 3 | 13.1% | p = 0.73 |
| Yes | 26 | 9 | 4 | 15.4% | ||
| Score in regenerative area | ||||||
| Negative | 33 | 1 | 3.1% | p = 0.0002 * | ||
| Positive | 16 | 6 | 37.5% | |||
| Factor | Df | Hazard Ratio | p-Value |
|---|---|---|---|
| Perineural invasion | 1 | 7.84 (1.28–48.12) | 0.0279 |
| Positive score in regenerative area | 1 | 15.02 (1.7–130.2) | 0.0024 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gissi, D.B.; Tarsitano, A.; Gabusi, A.; Rossi, R.; Attardo, G.; Lenzi, J.; Marchetti, C.; Montebugnoli, L.; Foschini, M.P.; Morandi, L. 13-gene DNA Methylation Analysis from Oral Brushing: A Promising Non Invasive Tool in the Follow-up of Oral Cancer Patients. J. Clin. Med. 2019, 8, 2107. https://doi.org/10.3390/jcm8122107
Gissi DB, Tarsitano A, Gabusi A, Rossi R, Attardo G, Lenzi J, Marchetti C, Montebugnoli L, Foschini MP, Morandi L. 13-gene DNA Methylation Analysis from Oral Brushing: A Promising Non Invasive Tool in the Follow-up of Oral Cancer Patients. Journal of Clinical Medicine. 2019; 8(12):2107. https://doi.org/10.3390/jcm8122107
Chicago/Turabian StyleGissi, Davide B., Achille Tarsitano, Andrea Gabusi, Roberto Rossi, Giuseppe Attardo, Jacopo Lenzi, Claudio Marchetti, Lucio Montebugnoli, Maria P. Foschini, and Luca Morandi. 2019. "13-gene DNA Methylation Analysis from Oral Brushing: A Promising Non Invasive Tool in the Follow-up of Oral Cancer Patients" Journal of Clinical Medicine 8, no. 12: 2107. https://doi.org/10.3390/jcm8122107
APA StyleGissi, D. B., Tarsitano, A., Gabusi, A., Rossi, R., Attardo, G., Lenzi, J., Marchetti, C., Montebugnoli, L., Foschini, M. P., & Morandi, L. (2019). 13-gene DNA Methylation Analysis from Oral Brushing: A Promising Non Invasive Tool in the Follow-up of Oral Cancer Patients. Journal of Clinical Medicine, 8(12), 2107. https://doi.org/10.3390/jcm8122107






