MicroRNAs: Emerging Novel Clinical Biomarkers for Hepatocellular Carcinomas
Abstract
:1. Introduction
2. MicroRNA (miRNA)
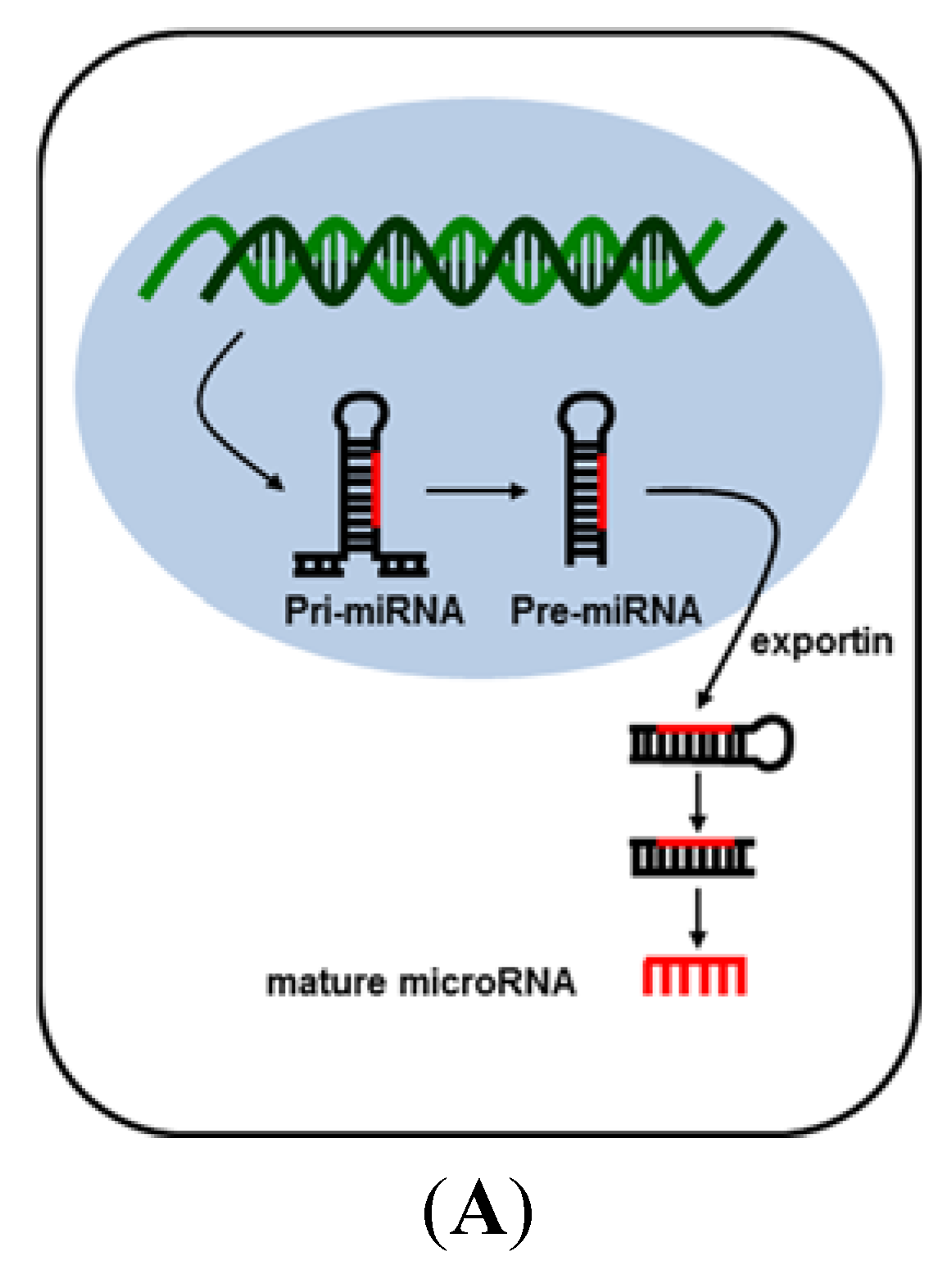
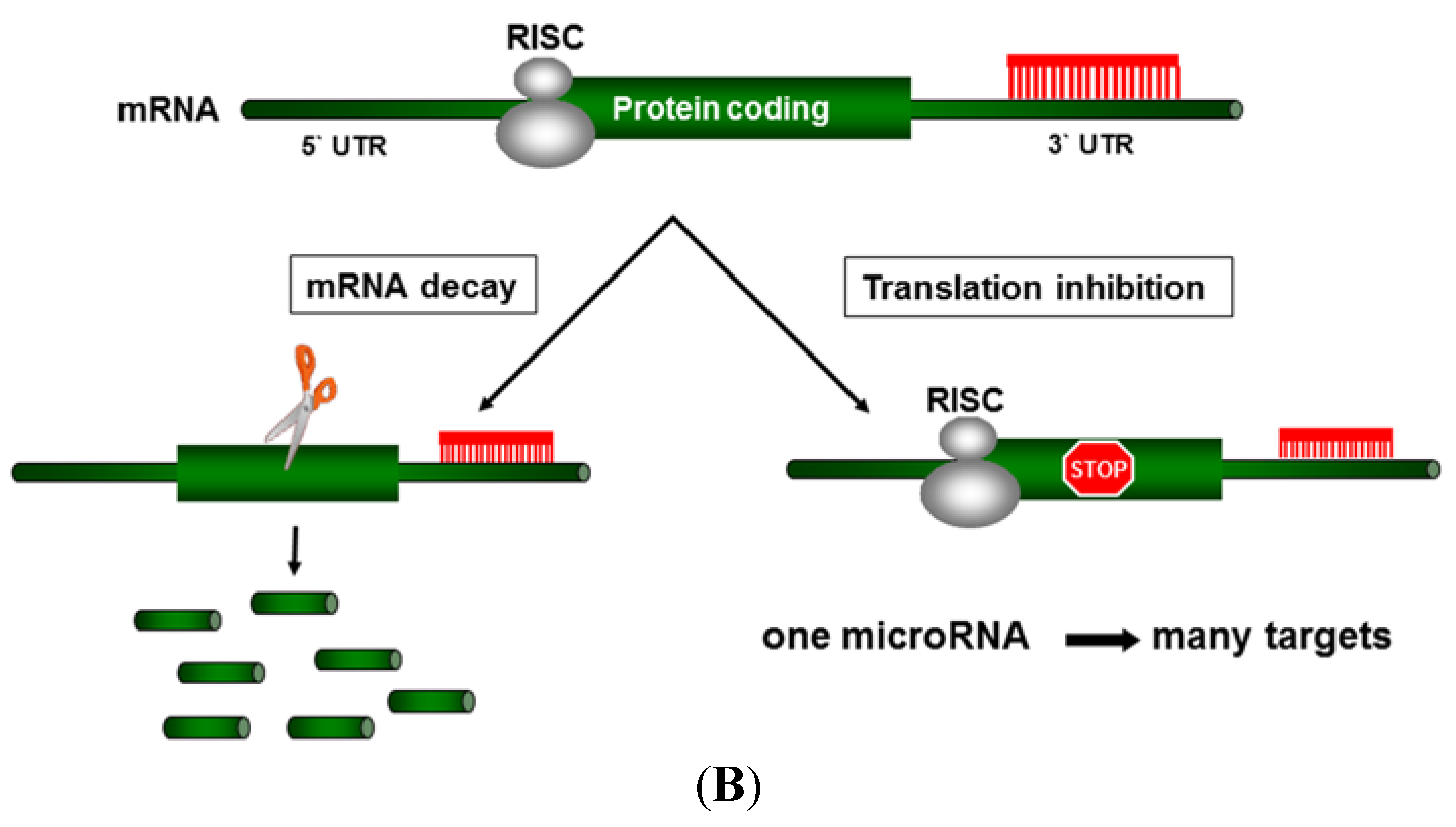
3. MicroRNAs in Liver Carcinogenesis
4. MicroRNAs as Diagnostic Markers
4.1. Primary Tissue Specimens
4.2. Serum
4.3. Plasma
5. MicroRNA Profiling for Prognosis in HCC
6. MicroRNAs as Therapeutic Targets and for Monitoring Therapeutic Response
7. Future Directions
| Diagnostic Biomarkers | ||||
|---|---|---|---|---|
| MicroRNA | Regulation | Source | Information | Ref |
| miR-106 | Up | Plasma | Differentiate HCC from healthy control and chronic liver disease | [41] |
| miR-122 | Up | Serum | Differentiate HCC from healthy control | [35] |
| miR-15b, miR-130b | Up | Serum | Differentiate HCC from healthy control | [38] |
| miR-16, miR-199a | Down | Serum | Differentiated HCC from chronic hepatitis and healthy control | [37] |
| miR-183 | Up | Tissue | Differentiate benign and malignant liver tumor | [39] |
| miR-15b, miR-130b | Up | Serum | Differentiate HCC and healthy patients and reduce after surgery | [38] |
| miR-18a | Up | Serum | Differentiate HCC and healthy patients | [93] |
| miR-122, miR192, miR-21, miR-223, miR-26a, miR-27a, miR-801 | Signature | Plasma | Differentiated HCC from cirrhosis, chronic liver patients, and healthy controls | [40] |
| miR-21 | Up | Serum, plasma | Differentiate HCC from cirrhosis and healthy controls | [59,94] |
| miR-375 | Up | Serum | Differentiated HBV- and HCV-related HCC from healthy controls | [36] |
| miR-483 | Up | Plasma | Differentiated HCC patients from healthy controls | [95] |
| miR-618/650 | Up | Urine | Differentiate HCC and control | [96] |
| miR-885 | Up | Serum | Differentiate HCC, cirrhosis, and chronic liver patients from healthy controls | [97] |
| miR-92a | Down | Plasma | Differentiated HCC from healthy control | [98] |
| miR-25, miR-375, let-7f | Up | Serum | Differentiate HCC from healthy control | [36] |
| miR-20a-5p, miR-320a, miR-324-3p and miR-375 | Up | Plasma | Differentiate HCC from non-cancerous lesions | [42] |
| miR-29a, miR-29c, miR-133a, miR-143, miR-145, miR-192, and miR-505 | Signature | Serum | Detect early stage HCC and AFP-negative HCC | [99] |
| MicroRNAs as Prognostic Markers | ||||
| miR-10b | Up | Tissue | Poor prognosis | [100] |
| miR-122 | Down | Tissue | Poor prognosis | [101] |
| miR-124 | Down | Tissue | Poor prognosis and aggressive type | [102] |
| miR-135a | Up | Tissue | Shorter overall survival and disease-free survival | [103] |
| miR-139 | Down | Tissue | Metastasis and poor prognosis | [104] |
| miR-155 | Up | Tissue | Poor prognosis, recurrence, micro-vascular invasion | [61] |
| miR-182 | Up | Tissue | Intrahepatic metastasis and poor prognosis | [62] |
| miR-199b-5p | Down | Tissue | Shorter overall survival | [105] |
| miR-203 | Up | Tissue | Better prognosis, longer survival | [106] |
| mi-21, miR-221 | Up | Tissue | Tumor stage and poor prognosis | [107] |
| miR-22 | Down | Tissue | Poor survival | [108] |
| miR-221 | Up | Serum | Poor survival | [58] |
| miR-29 | Down | Tissue | Shorter disease-free survival | [64] |
| miR-29a-5p | Up | Tissue | Recurrence in early stage HCC | [63] |
| miR-99a | Down | Tissue | Shorter survival | [109] |
| let-7g | Down | Tissue | Poor survival | [110] |
| DLK1-DIO3 miRNA cluster | Up | Tissue | Poor prognosis | [111] |
| C19MC microRNA cluster | Up | Tissue | Poor clinico-pathological features, recurrence, and shorter overall survival | [112] |
| miR-155, miR-15a, miR-432, miR-486-3p, miR-15b, miR-30b | Up | Tissue | Recurrence-free survival | [113] |
| miR-19a, miR-886, miR-126, miR-223, miR-24, and miR-147 | Signature | Tissue | Overall survival and recurrent free survival | [65] |
| 67 miRs signature | Signature | Tissue | Differentiate recurrence after liver transplantation | [114] |
| miR signatures in tumor and non-tumor tissues | Signature | Tissue | Differentiate early and late recurrence | [115] |
| miR-326, miR-3677, miR-511-1, miR-511-2, miR-9-1, and miR-9-2 | Signature | Tissue | Negatively associated with overall survival | [116] |
| Predictive Therapeutic Response Markers | ||||
| miR-122 | Down | Cells, tissue | Decreased sensitivity to Doxorubicin | [81] |
| miR-122 | Down | Cells, tissue | Decreased sensitivity to Adriamycin, Vincristin | [80] |
| miR-122 | Down | Cells, tissue | Suppressed sensitivity to sorafenib | [76] |
| miR-146a | Up | Cells | Suppresses sensitivity to interferon-α | [71] |
| miR-193a-3p | Down | Cells, tissue | Resistance to 5-FU | [84] |
| miR-193b | Up | Cells, Tissue | Sensitivity to cisplatin | [117] |
| miR-199a-3p | Down | Cells, tissue | Increased sensitivity to Doxorubicin | [82] |
| miR-1247a | Down | Cells | Resistance to sorafenib | [118] |
| miR-21 | Up | Cells, tissue | Resistance to interferon-α/5FU in HCC cells | [74] |
| miR-34a | Down | Cells, tissue | Resistance to sorafenib | [94] |
| 13 microRNA signature | Signature | Cells, tissue | Multidrug resistance | [90] |
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Jemal, A.; Bray, F.; Center, M.M.; Ferlay, J.; Ward, E.; Forman, D. Global Cancer Statistics: 2011. CA Cancer J. Clin. 2011, 61, 69–90. [Google Scholar] [CrossRef] [PubMed]
- Altekruse, S.F.; McGlynn, K.A.; Reichman, M.E. Hepatocellular carcinoma incidence, mortality, and survival trends in the United States from 1975 to 2005. J. Clin. Oncol. 2009, 27, 1485–1491. [Google Scholar] [CrossRef] [PubMed]
- You, J.S.; Jones, P.A. Cancer Genetics and Epigenetics: Two Sides of the Same Coin? Cancer Cell 2012, 22, 9–20. [Google Scholar] [CrossRef] [PubMed]
- Ashworth, A.; Lord, C.J.; Reis-Filho, J.S. Genetic interactions in cancer progression and treatment. Cell 2011, 145, 30–38. [Google Scholar] [CrossRef] [PubMed]
- Ambros, V. microRNAs: Tiny regulators with great potential. Cell 2001, 107, 823–826. [Google Scholar] [CrossRef]
- Pogribny, I.P.; Rusyn, I. Role of epigenetic aberrations in the development and progression of human hepatocellular carcinoma. Cancer Lett. 2014, 342, 223–230. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.W.; Heegaard, N.H.H.; Orum, H. MicroRNAs in liver disease. Gastroenterology 2012, 142, 1431–1443. [Google Scholar] [CrossRef] [PubMed]
- Borel, F.; Konstantinova, P.; Jansen, P.L.M. Diagnostic and therapeutic potential of miRNA signatures in patients with hepatocellular carcinoma. J. Hepatol. 2012, 56, 1371–1383. [Google Scholar] [CrossRef] [PubMed]
- Giordano, S.; Columbano, A. MicroRNAs: New tools for diagnosis, prognosis, and therapy in hepatocellular carcinoma? Hepatology 2013, 57, 840–847. [Google Scholar] [CrossRef] [PubMed]
- De Gramont, A.; Watson, S.; Ellis, L.M.; Rodón, J.; Tabernero, J.; de Gramont, A.; Hamilton, S.R. Pragmatic issues in biomarker evaluation for targeted therapies in cancer. Nat. Rev. Clin. Oncol. 2014, 12, 197–212. [Google Scholar] [CrossRef] [PubMed]
- He, L.; Hannon, G.J. MicroRNAs: Small RNAs with a big role in gene regulation. Nat. Rev. Genet. 2004, 5, 522–531. [Google Scholar] [CrossRef] [PubMed]
- Cai, X.; Hagedorn, C.H.; Cullen, B.R. Human microRNAs are processed from capped, polyadenylated transcripts that can also function as mRNAs. 2004, 10, 1957–1966. [Google Scholar]
- Orban, T.I.; Izaurralde, E. Decay of mRNAs targeted by RISC requires XRN1, the Ski complex, and the exosome. RNA 2005, 11, 459–469. [Google Scholar] [CrossRef] [PubMed]
- Kim, V.N. MicroRNA biogenesis: Coordinated cropping and dicing. Nat. Rev. Mol. Cell Biol. 2005, 6, 376–385. [Google Scholar] [CrossRef] [PubMed]
- Rehwinkel, J.; Behm-Ansmant, I.; Gatfield, D.; Izaurralde, E. A crucial role for GW182 and the DCP1:DCP2 decapping complex in miRNA-mediated gene silencing. RNA 2005, 11, 1640–1647. [Google Scholar] [CrossRef] [PubMed]
- Lewis, B.P.; Burge, C.B.; Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005, 120, 15–20. [Google Scholar] [CrossRef] [PubMed]
- Calin, G.A.; Dumitru, C.D.; Shimizu, M.; Bichi, R.; Zupo, S.; Noch, E.; Aldler, H.; Rattan, S.; Keating, M.; Rai, K.; et al. Frequent deletions and down-regulation of micro- RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc. Natl. Acad. Sci. USA 2002, 99, 13–18. [Google Scholar] [CrossRef] [PubMed]
- Wong, C.M.; Wong, C.C.; Lee, J.M.; Fan, D.N.; Au, S.L.; Ng, I.O. Sequential alterations of microrna expression in hepatocellular carcinoma development and venous metastasis. Hepatology 2012, 55, 1453–1461. [Google Scholar] [CrossRef] [PubMed]
- Croce, C.M. Causes and consequences of microRNA dysregulation in cancer. Nat. Rev. Genet. 2009, 10, 704–714. [Google Scholar] [CrossRef] [PubMed]
- Anwar, S.L.; Albat, C.; Krech, T.; Hasemeier, B.; Schipper, E.; Schweitzer, N.; Vogel, A.; Kreipe, H.; Lehmann, U. Concordant hypermethylation of intergenic microRNA genes in human hepatocellular carcinoma as new diagnostic and prognostic marker. Int. J. Cancer 2013, 133, 660–670. [Google Scholar] [CrossRef] [PubMed]
- Anwar, S.L.; Lehmann, U. DNA methylation, microRNAs, and their crosstalk as potential biomarkers in hepatocellular carcinoma. World J. Gastroenterol. 2014, 20, 7894–7913. [Google Scholar] [CrossRef] [PubMed]
- Buurman, R.; Gürlevik, E.; Schäffer, V.; Eilers, M.; Sandbothe, M.; Kreipe, H.; Wilkens, L.; Schlegelberger, B.; Kühnel, F.; Skawran, B. Histone deacetylases activate hepatocyte growth factor signaling by repressing microRNA-449 in hepatocellular carcinoma cells. Gastroenterology 2012, 143, 811–820. [Google Scholar] [CrossRef] [PubMed]
- Iorio, M.V.; Croce, C.M. MicroRNA dysregulation in cancer: Diagnostics, monitoring and therapeutics. A comprehensive review. EMBO Mol. Med. 2012, 4, 143–159. [Google Scholar] [CrossRef] [PubMed]
- Wei, R.; Huang, G.L.; Zhang, M.Y.; Li, B.K.; Zhang, H.Z.; Shi, M.; Chen, X.Q.; Huang, L.; Zhou, Q.M.; Jia, W.H.; et al. Clinical significance and prognostic value of microRNA expression signature in hepatocellular carcinoma. Clin. Cancer Res. 2013, 19, 4780–4791. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.; He, X. The role of microRNAs in liver cancer progression. Br. J. Cancer 2011, 104, 235–240. [Google Scholar] [CrossRef] [PubMed]
- Law, P.T.-Y.; Wong, N. Emerging roles of microRNA in the intracellular signaling networks of hepatocellular carcinoma. J. Gastroenterol. Hepatol. 2011, 26, 437–449. [Google Scholar] [CrossRef] [PubMed]
- Ladeiro, Y.; Couchy, G.; Balabaud, C.; Bioulac-Sage, P.; Pelletier, L.; Rebouissou, S.; Zucman-Rossi, J. MicroRNA profiling in hepatocellular tumors is associated with clinical features and oncogene/tumor suppressor gene mutations. Hepatology 2008, 47, 1955–1963. [Google Scholar] [CrossRef] [PubMed]
- Salvi, A.; Abeni, E.; Portolani, N.; Barlati, S.; de Petro, G. Human hepatocellular carcinoma cell-specific miRNAs reveal the differential expression of miR-24 and miR-27a in cirrhotic/non-cirrhotic HCC. Int. J. Oncol. 2013, 42, 391–402. [Google Scholar] [PubMed]
- Fu, Y.; Wei, X.; Tang, C.; Li, J.; Liu, R.; Shen, A.; Wu, Z. Circulating microRNA-101 as a potential biomarker for hepatitis B virus-related hepatocellular carcinoma. Oncol. Lett. 2013, 6, 1811–1815. [Google Scholar] [PubMed]
- Gao, P.; Wong, C.C.; Tung, E.K.; Lee, J.M.; Wong, C.M.; Ng, I.O. Deregulation of microRNA expression occurs early and accumulates in early stages of HBV-associated multistep hepatocarcinogenesis. J. Hepatol. 2011, 54, 1177–1184. [Google Scholar] [CrossRef] [PubMed]
- Pineau, P.; Volinia, S.; McJunkin, K.; Marchio, A.; Battiston, C.; Terris, B.; Mazzaferro, V.; Lowe, S.W.; Croce, C.M.; Dejean, A. MiR-221 overexpression contributes to liver tumorigenesis. Proc. Natl. Acad. Sci. USA 2010, 107, 264–269. [Google Scholar] [CrossRef] [PubMed]
- Zhu, K.; Dai, Z.; Zhou, J. Biomarkers for hepatocellular carcinoma: Progression in early diagnosis, prognosis, and personalized therapy. Biomark. Res. 2013, 1, 10. [Google Scholar] [CrossRef] [PubMed]
- Law, P.T.Y.; Qin, H.; Ching, A.K.; Lai, K.P.; Co, N.N.; He, M.; Lung, R.W.; Chan, A.W.; Chan, T.F.; Wong, N. Deep sequencing of small RNA transcriptome reveals novel non-coding RNAs in hepatocellular carcinoma. J. Hepatol. 2013, 58, 1165–1173. [Google Scholar] [CrossRef] [PubMed]
- Hou, J.; Lin, L.; Zhou, W.; Wang, Z.; Ding, G.; Dong, Q.; Qin, L.; Wu, X.; Zheng, Y.; Yang, Y.; et al. Identification of miRNomes in Human Liver and Hepatocellular Carcinoma Reveals miR-199a/b-3p as Therapeutic Target for Hepatocellular Carcinoma. Cancer Cell 2011, 19, 232–243. [Google Scholar] [CrossRef] [PubMed]
- Qi, P.; Cheng, S.Q.; Wang, H.; Li, N.; Chen, Y.F.; Gao, C.F. Serum MicroRNAs as Biomarkers for Hepatocellular Carcinoma in Chinese Patients with Chronic Hepatitis B Virus Infection. PLoS ONE 2011, 6, e28486. [Google Scholar] [CrossRef] [PubMed]
- Li, L.M.; Hu, Z.B.; Zhou, Z.X.; Chen, X.; Liu, F.Y.; Zhang, J.F.; Shen, H.B.; Zhang, C.Y.; Zen, K. Serum microRNA profiles serve as novel biomarkers for HBV infection and diagnosis of HBV-positive hepatocarcinoma. Cancer Res. 2010, 70, 9798–9807. [Google Scholar] [CrossRef] [PubMed]
- Jiang, L.; Li, X.; Cheng, Q.; Zhang, B.H. Plasma microRNA might as a potential biomarker for hepatocellular carcinoma andchronic liver disease screening. Tumor Biol. 2015. [Google Scholar] [CrossRef] [PubMed]
- Liu, A.M.; Yao, T.J.; Wang, W.; Wong, K.F.; Lee, N.P.; Fan, S.T.; Poon, R.T.; Gao, C.; Luk, J.M. Circulating miR-15b and miR-130b in serum as potential markers for detecting hepatocellular carcinoma: A retrospective cohort study. BMJ Open 2012, 2, e000825. [Google Scholar] [CrossRef] [PubMed]
- Lin, X.J.; Chong, Y.; Guo, Z.W.; Xie, C.; Yang, X.J.; Zhang, Q.; Li, S.P.; Xiong, Y.; Yuan, Y.; Min, J.; et al. A serum microRNA classifier for early detection of hepatocellular carcinoma: A multicentre, retrospective, longitudinal biomarker identification study with a nested case-control study. Lancet Oncol. 2015. [Google Scholar] [CrossRef]
- Zhou, J.; Yu, L.; Gao, X.; Hu, J.; Wang, J.F.; Zhang, Z.; Lu, S.; Huang, X.; Wang, Z.; Qiu, S.; et al. Plasma microRNA panel to diagnose hepatitis B virus-related hepatocellular carcinoma. J. Clin. Oncol. 2011, 29, 4781–4788. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.; Ma, J.; Wu, S.; Yang, D.; Yan, Y.; Liu, K.; Wang, J.; Sun, L.; Chen, N.; Wei, H.; et al. Characterization of the liver Tissue Interstitial Fluid (TIF) proteome indicates potential for application in liver disease biomarker discovery. J. Proteome Res. 2010, 9, 1020–1031. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.; Huang, D.; Ni, S.; Peng, Z.; Sheng, W.; Du, X. Plasma microRNAs are promising novel biomarkers for early detection of colorectal cancer. Int. J. Cancer 2010, 127, 118–126. [Google Scholar] [CrossRef] [PubMed]
- Su, Z.; Zhao, J.; Rong, Z.H.; Geng, W.M.; Wu, Y.G.; Qin, C.K. Upregulation of microRNA-25 associates with prognosis in hepatocellular carcinoma. Diagn. Pathol. 2014, 9, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Liang, Z.; Gao, Y.; Shi, W.; Zhai, D.; Li, S.; Jing, L.; Guo, H.; Liu, T.; Wang, Y.; Du, Z. Expression and significance of MicroRNA-183 in hepatocellular carcinoma. Sci. World J. 2013, 2013. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Yin, Y.; Wang, F.; Wang, Y.; Zhang, L.; Tang, Y.; Sun, S. miR-17-5p Promotes migration of human hepatocellular carcinoma cells through the p38 mitogen-activated protein kinase-heat shock protein 27 pathway. Hepatology 2010, 51, 1614–1623. [Google Scholar] [CrossRef] [PubMed]
- El Tayebi, H.M.; Omar, K.; Hegy, S.; El Maghrabi, M.; El Brolosy, M.; Hosny, K.A.; Esmat, G.; Abdelaziz, A.I. Repression of miR-17-5p with elevated expression of E2F-1 and c-MYC in non-metastatic hepatocellular carcinoma and enhancement of cell growth upon reversing this expression pattern. Biochem. Biophys. Res. Commun. 2013, 434, 421–427. [Google Scholar] [CrossRef] [PubMed]
- Yoon, S.O.; Chun, S.M.; Han, E.H.; Choi, J.; Jang, S.J.; Koh, S.A.; Hwang, S.; Yu, E. Deregulated expression of microRNA-221 with the potential for prognostic biomarkers in surgically resected hepatocellular carcinoma. Hum. Pathol. 2011, 42, 1391–1400. [Google Scholar] [CrossRef] [PubMed]
- Chen, P.; Zhao, X.; Ma, L. Downregulation of microRNA-100 correlates with tumor progression and poor prognosis in hepatocellular carcinoma. Mol. Cell. Biochem. 2013, 383, 49–58. [Google Scholar] [CrossRef] [PubMed]
- Shi, C.; Xu, X. MicroRNA-22 is down-regulated in hepatitis B virus-related hepatocellular carcinoma. Biomed. Pharmacother. 2013, 67, 375–380. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.H.; Wang, Q.; Chen, J.S.; Fu, X.H.; Chen, X.L.; Chen, L.Z.; Li, W.; Bi, J.; Zhang, L.J.; Fu, Q.; et al. Bead-based microarray analysis of microRNA expression in hepatocellular carcinoma: MiR-338 is downregulated. Hepatol. Res. 2009, 39, 786–794. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Fu, H.; Tie, Y.; Hu, Z.; Kong, W.; Wu, Y.; Zheng, X. miR-34a inhibits migration and invasion by down-regulation of c-Met expression in human hepatocellular carcinoma cells. Cancer Lett. 2009, 275, 44–53. [Google Scholar] [CrossRef] [PubMed]
- Pan, L.; Huang, S.; He, R.; Rong, M.; Dang, Y.; Chen, G. Decreased expression and clinical significance of miR-148a in hepatocellular carcinoma tissues. Eur. J. Med. Res. 2014, 19, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Guo, X.; Xiong, L.; Kong, X.; Xu, Y.; Liu, C.; Zou, L.; Li, Z.; Zhao, J.; Lin, N. MicroRNA-101 suppresses SOX9-dependent tumorigenicity and promotes favorable prognosis of human hepatocellular carcinoma. FEBS Lett. 2012, 586, 4362–4370. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Zheng, W.; Hai, J. MicroRNA-148b expression is decreased in hepatocellular carcinoma and associated with prognosis. Med. Oncol. 2014, 31, 984. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Li, J.; Wang, X.; Zheng, C.; Ma, W. Downregulation of microRNA-214 and overexpression of FGFR-1 contribute to hepatocellular carcinoma metastasis. Biochem. Biophys. Res. Commun. 2013, 439, 47–53. [Google Scholar] [CrossRef] [PubMed]
- Ura, S.; Honda, M.; Yamashita, T.; Ueda, T.; Takatori, H.; Nishino, R.; Sunakozaka, H.; Sakai, Y.; Horimoto, K.; Kaneko, S. Differential microRNA expression between hepatitis B and hepatitis C leading disease progression to hepatocellular carcinoma. Hepatology 2009, 49, 1098–1112. [Google Scholar] [CrossRef] [PubMed]
- Budhu, A.; Jia, H.L.; Forgues, M.; Liu, C.G.; Goldstein, D.; Lam, A.; Zanetti, K.A.; Ye, Q.H.; Qin, L.X.; Croce, C.M.; et al. Identification of metastasis-related microRNAs in hepatocellular carcinoma. Hepatology 2008, 47, 897–907. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Wang, Y.; Yu, W.; Chen, J.; Luo, J. Expression of serum miR-221 in human hepatocellular carcinoma and its prognostic significance. Biochem. Biophys. Res. Commun. 2011, 406, 70–73. [Google Scholar] [CrossRef] [PubMed]
- Tomimaru, Y.; Eguchi, H.; Nagano, H.; Wada, H.; Kobayashi, S.; Marubashi, S.; Tanemura, M.; Tomokuni, A.; Takemasa, I.; Umeshita, K.; et al. Circulating microRNA-21 as a novel biomarker for hepatocellular carcinoma. J. Hepatol. 2012, 56, 167–175. [Google Scholar] [CrossRef] [PubMed]
- Gu, H.; Guo, X.; Zou, L.; Zhu, H.; Zhang, J. Upregulation of microRNA-372 associates with tumor progression and prognosis in hepatocellular carcinoma. Mol. Cell. Biochem. 2013, 375, 23–30. [Google Scholar] [CrossRef] [PubMed]
- Han, Z.-B.; Chen, H.Y.; Fan, J.W.; Wu, J.Y.; Tang, H.M.; Peng, Z.H. Up-regulation of microRNA-155 promotes cancer cell invasion and predicts poor survival of hepatocellular carcinoma following liver transplantation. J. Cancer Res. Clin. Oncol. 2012, 138, 153–161. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Li, J.; Shen, J.; Wang, C.; Yang, L.; Zhang, X. MicroRNA-182 downregulates metastasis suppressor 1 and contributes to metastasis of hepatocellular carcinoma. BMC Cancer 2012, 12, 227. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.T.; Dong, Q.Z.; Sheng, Y.Y.; Wei, J.W.; Wang, G.; Zhou, H.J.; Ren, N.; Jia, H.L.; Ye, Q.H.; Qin, L.X. MicroRNA-29a-5p Is a Novel Predictor for Early Recurrence of Hepatitis B Virus-Related Hepatocellular Carcinoma after Surgical Resection. PLoS ONE 2012, 7, e52393. [Google Scholar] [CrossRef] [PubMed]
- Xiong, Y.; Fang, J.H.; Yun, J.P.; Yang, J.; Zhang, Y.; Jia, W.H.; Zhuang, S.M. Effects of microrna-29 on apoptosis, tumorigenicity, and prognosis of hepatocellular carcinoma. Hepatology 2010, 51, 836–845. [Google Scholar] [CrossRef] [PubMed]
- Han, Z.-B.; Zhong, L.; Teng, M.J.; Fan, J.W.; Tang, H.M.; Wu, J.Y.; Chen, H.Y.; Wang, Z.W.; Qiu, G.Q.; Peng, Z.H. Identification of recurrence-related microRNAs in hepatocellular carcinoma following liver transplantation. Mol. Oncol. 2012, 6, 445–457. [Google Scholar] [CrossRef] [PubMed]
- Krützfeldt, J.; Kuwajima, S.; Braich, R.; Rajeev, K.G.; Pena, J.; Tuschl, T.; Manoharan, M.; Stoffel, M. Specificity, duplex degradation and subcellular localization of antagomirs. Nucleic Acids Res. 2007, 35, 2885–2892. [Google Scholar] [CrossRef] [PubMed]
- Callegari, E.; Elamin, B.K.; Giannone, F.; Milazzo, M.; Altavilla, G.; Fornari, F.; Giacomelli, L.; D'Abundo, L.; Ferracin, M.; Bassi, C.; et al. Liver tumorigenicity promoted by microRNA-221 in a mouse transgenic model. Hepatology 2012, 56, 1025–1033. [Google Scholar] [CrossRef] [PubMed]
- Kota, J.; Chivukula, R.R.; O'Donnell, K.A.; Wentzel, E.A.; Montgomery, C.L.; Hwang, H.W.; Chang, T.C.; Vivekanandan, P.; Torbenson, M.; Clark, K.R.; et al. Therapeutic microRNA Delivery Suppresses Tumorigenesis in a Murine Liver Cancer Model. Cell 2009, 137, 1005–1017. [Google Scholar] [CrossRef] [PubMed]
- Janssen, H.L.; Reesink, H.W.; Lawitz, E.J.; Zeuzem, S.; Rodriguez-Torres, M.; Patel, K.; van der Meer, A.J.; Patick, A.K.; Chen, A.; Zhou, Y.; et al. Treatment of HCV infection by targeting microRNA. N. Engl. J. Med. 2013, 368, 1685–1694. [Google Scholar] [CrossRef] [PubMed]
- Ling, H.; Fabbri, M.; Calin, G.A. MicroRNAs and other non-coding RNAs as targets for anticancer drug development. Nat. Rev. Drug Discov. 2013, 12, 847–865. [Google Scholar] [CrossRef] [PubMed]
- Tomokuni, A.; Eguchi, H.; Tomimaru, Y.; Wada, H.; Kawamoto, K.; Kobayashi, S.; Marubashi, S.; Tanemura, M.; Nagano, H.; Mori, M.; et al. MiR-146a suppresses the sensitivity to interferon-α in hepatocellular carcinoma cells. Biochem. Biophys. Res. Commun. 2011, 414, 675–680. [Google Scholar] [CrossRef] [PubMed]
- Ji, J.; Shi, J.; Budhu, A.; Forgues, M.; Roessler, S.; Ambs, S.; Chen, Y.; Meltzer, P.S.; Croce, C.M.; Qin, L.X.; et al. MicroRNA expression, survival, and response to interferon in liver cancer. N. Engl. J. Med. 2009, 361, 1437–1447. [Google Scholar] [CrossRef] [PubMed]
- Ji, J.; Yu, L.; Yu, Z.; Forgues, M.; Uenishi, T.; Kubo, S.; Wakasa, K.; Zhou, J.; Fan, J.; Tang, Z.Y.; et al. Development of a miR-26 companion diagnostic test for adjuvant interferon-alpha therapy in hepatocellular carcinoma. Int. J. Biol. Sci. 2013, 9, 303–312. [Google Scholar] [CrossRef] [PubMed]
- Tomimaru, Y.; Eguchi, H.; Nagano, H.; Wada, H.; Tomokuni, A.; Kobayashi, S.; Marubashi, S.; Takeda, Y.; Tanemura, M.; Umeshita, K.; et al. MicroRNA-21 induces resistance to the anti-tumour effect of interferon-α/5-fluorouracil in hepatocellular carcinoma cells. Br. J. Cancer 2010, 103, 1617–1626. [Google Scholar] [CrossRef] [PubMed]
- Zhou, C.; Liu, J.; Li, Y.; Liu, L.; Zhang, X.; Ma, C.Y.; Hua, S.C.; Yang, M.; Yuan, Q. MicroRNA-1274a, a modulator of sorafenib induced a disintegrin and metalloproteinase 9 (ADAM9) down-regulation in hepatocellular carcinoma. FEBS Lett. 2011, 585, 1828–1834. [Google Scholar] [CrossRef] [PubMed]
- Bai, S.; Nasser, M.W.; Wang, B.; Hsu, S.H.; Datta, J.; Kutay, H.; Yadav, A.; Nuovo, G.; Kumar, P.; Ghoshal, K. MicroRNA-122 inhibits tumorigenic properties of hepatocellular carcinoma cells and sensitizes these cells to sorafenib. J. Biol. Chem. 2009, 284, 32015–32027. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.H.; Chen, J.S.; Wang, Q.; Chen, X.L.; Wen, L.; Chen, L.Z.; Bi, J.; Zhang, L.J.; Su, Q.; Zeng, W.T. MiR-338-3p suppresses invasion of liver cancer cell by targeting smoothened. J. Pathol. 2011, 225, 463–472. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Li, Q.J.; Gong, Z.B.; Zhou, L.; You, N.; Wang, S.; Li, X.L.; Li, J.J.; An, J.Z.; Wang, D.S.; et al. MicroRNA-34a Targets Bcl-2 and Sensitizes Human Hepatocellular Carcinoma Cells to Sorafenib Treatment. Technol. Cancer Res. Treat. 2013, 13, 77–86. [Google Scholar] [PubMed]
- Yang, F.; Zhang, L.; Wang, F.; Wang, Y.; Huo, X.S.; Yin, Y.X.; Wang, Y.Q.; Zhang, L.; Sun, S.H. Modulation of the unfolded protein response is the core of microRNA-122-involved sensitivity to chemotherapy in hepatocellular carcinoma. Neoplasia 2011, 13, 590–600. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Xia, F.; Ma, L.; Shan, J.; Shen, J.; Yang, Z.; Liu, J.; Cui, Y.; Bian, X.; Bie, P.; et al. MicroRNA-122 sensitizes HCC cancer cells to adriamycin and vincristine through modulating expression of MDR and inducing cell cycle arrest. Cancer Lett. 2011, 310, 160–169. [Google Scholar] [CrossRef] [PubMed]
- Fornari, F.; Gramantieri, L.; Giovannini, C.; Veronese, A.; Ferracin, M.; Sabbioni, S.; Calin, G.A.; Grazi, G.L.; Croce, C.M.; Tavolari, S.; et al. MiR-122/cyclin G1 interaction modulates p53 activity and affects doxorubicin sensitivity of human hepatocarcinoma cells. Cancer Res. 2009, 69, 5761–5767. [Google Scholar] [CrossRef] [PubMed]
- Fornari, F.; Milazzo, M.; Chieco, P.; Negrini, M.; Calin, G.A.; Grazi, G.L.; Pollutri, D.; Croce, C.M.; Bolondi, L.; Gramantieri, L. MiR-199a-3p regulates mTOR and c-Met to influence the doxorubicin sensitivity of human hepatocarcinoma cells. Cancer Res. 2010, 70, 5184–5193. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; An, Y.; Wang, Y.; Zhang, C.; Zhang, H.; Huang, C.; Jiang, H.; Wang, X.; Li, X. MiR-101 inhibits autophagy and enhances cisplatin-induced apoptosis in hepatocellular carcinoma cells. Oncol. Rep. 2013, 29, 2019–2024. [Google Scholar] [PubMed]
- Kelong, M.; He, Y.; Zhang, H.; Fei, Q.; Niu, D.; Wang, D.; Ding, X.; Xu, H.; Chen, X.; Zhu, J. DNA methylation-regulated miR-193a-3p dictates resistance of hepatocellular carcinoma to 5-fluorouracil via repression of SRSF2 expression. J. Biol. Chem. 2012, 287, 5639–5649. [Google Scholar]
- Chen, Z.; Ma, T.; Huang, C.; Zhang, L.; Lv, X.; Xu, T.; Hu, T.; Li, J. MiR-27a modulates the MDR1/P-glycoprotein expression by inhibiting FZD7/β-catenin pathway in hepatocellular carcinoma cells. Cell. Signal. 2013, 25, 2693–2701. [Google Scholar] [CrossRef] [PubMed]
- Shi, L.; Wu, L.; Chen, Z.; Yang, J.; Chen, X.; Yu, F.; Zheng, F.; Lin, X. MiR-141 Activates Nrf2-Dependent Antioxidant Pathway via Down-Regulating the Expression of Keap1 Conferring the Resistance of Hepatocellular Carcinoma Cells to 5-Fluorouracil. Cell. Physiol. Biochem. 2015, 35, 2333–2348. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.; Zhu, M.; Tsao, S.W.; Man, K.; Zhang, Z.; Feng, Y. MiR-23a-mediated inhibition of topoisomerase 1 expression potentiates cell response to etoposide in human hepatocellular carcinoma. Mol. Cancer 2013, 12, 119. [Google Scholar] [CrossRef] [PubMed]
- Zhao, N.; Wang, R.; Zhou, L.; Zhu, Y.; Gong, J.; Zhuang, S.M. MicroRNA-26b suppresses the NF-κB signaling and enhances the chemosensitivity of hepatocellular carcinoma cells by targeting TAK1 and TAB3. Mol. Cancer 2014, 13, 35. [Google Scholar] [CrossRef] [PubMed]
- Yang, T.; Zheng, Z.M.; Li, X.N.; Li, Z.F.; Wang, Y.; Geng, Y.F.; Bai, L.; Zhang, X.B. MiR-223 modulates multidrug resistance via downregulation of ABCB1 in hepatocellular carcinoma cells. Exp. Biol. Med. (Maywood) 2013, 238, 1024–1032. [Google Scholar] [CrossRef] [PubMed]
- Borel, F.; Han, R.; Visser, A.; Petry, H.; van Deventer, S.J.H.; Jansen, P.L.M.; Konstantinova, P.; Réseau Centre de Ressources Biologiques Foie (French Liver Biobanks Network), France. Adenosine triphosphate-binding cassette transporter genes up-regulation in untreated hepatocellular carcinoma is mediated by cellular microRNAs. Hepatology 2012, 55, 821–832. [Google Scholar] [CrossRef] [PubMed]
- Tiniakos, D.G.; vos, M.B.; Brunt, E.M. Nonalcoholic fatty liver disease: Pathology and pathogenesis. Annu. Rev. Pathol. 2010, 5, 145–171. [Google Scholar] [CrossRef] [PubMed]
- Kosaka, N.; Iguchi, H.; Ochiya, T. Circulating microRNA in body fluid: A new potential biomarker for cancer diagnosis and prognosis. Cancer Sci. 2010, 101, 2087–2092. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Guo, Z.; Wang, J.; Mao, Y.; Gao, Q. Serum miR-18a: A potential marker for hepatitis B virus-related hepatocellular carcinoma screening. Dig. Dis. Sci. 2012, 57, 2910–2916. [Google Scholar] [CrossRef] [PubMed]
- Bihrer, V.; Waidmann, O.; Friedrich-Rust, M.; Forestier, N.; Susser, S.; Haupenthal, J.; Welker, M.; Shi, Y.; Peveling-Oberhag, J.; Potal, A.; et al. Serum microRNA-21 as marker for necroinflammation in hepatitis C patients with and without hepatocellular carcinoma. PLoS ONE 2011, 6, e26971. [Google Scholar] [CrossRef] [PubMed]
- Shen, J.; Wang, A.; Wang, Q.; Gurvich, I.; Siegel, A.B.; Remotti, H.; Santella, R.M. Exploration of genome-wide circulating microRNA in hepatocellular carcinoma: MiR-483-5p as a potential biomarker. Cancer Epidemiol. Biomark. Prev. 2013, 22, 2364–2373. [Google Scholar] [CrossRef] [PubMed]
- Abdalla, M.A.K.; Haj-Ahmad, Y. Promising candidate urinary microrna biomarkers for the early detection of hepatocellular carcinoma among high-risk hepatitis C virus Egyptian patients. J. Cancer 2012, 3, 19–31. [Google Scholar] [CrossRef] [PubMed]
- Gui, J.; Tian, Y.; Wen, X.; Zhang, W.; Zhang, P.; Gao, J.; Run, W.; Tian, L.; Jia, X.; Gao, Y. Serum microRNA characterization identifies miR-885-5p as a potential marker for detecting liver pathologies. Clin. Sci. (Lond.) 2011, 120, 183–193. [Google Scholar] [CrossRef] [PubMed]
- Shigoka, M.; Tsuchida, A.; Matsudo, T.; Nagakawa, Y.; Saito, H.; Suzuki, Y.; Aoki, T.; Murakami, Y.; Toyoda, H.; Kumada, T.; et al. Deregulation of miR-92a expression is implicated in hepatocellular carcinoma development. Pathol. Int. 2010, 60, 351–357. [Google Scholar] [CrossRef] [PubMed]
- Wu, P.; Agnelli, L.; Walker, B.A.; Todoert, K.; Lionetti, M.; Johnson, D.C.; Kaiser, M.; Mirabella, F.; Wardell, C.; Gregory, W.M.; et al. Improved risk stratification in myeloma using a microRNA-based classifier. Br. J. Haematol. 2013, 162, 348–359. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.J.; Zhou, L.; Yang, F.; Wang, G.X.; Zheng, H.; Wang, D.S.; He, Y.; Dou, K.F. MicroRNA-10b promotes migration and invasion through CADM1 in human hepatocellular carcinoma cells. Tumor Biol. 2012, 33, 1455–1465. [Google Scholar] [CrossRef] [PubMed]
- Coulouarn, C.; Factor, V.M.; Andersen, J.B.; Durkin, M.E.; Thorgeirsson, S.S. Loss of miR-122 expression in liver cancer correlates with suppression of the hepatic phenotype and gain of metastatic properties. Oncogene 2009, 28, 3526–3536. [Google Scholar] [CrossRef] [PubMed]
- Zheng, F.; Liao, Y.J.; Cai, M.Y.; Liu, Y.H.; Liu, T.H.; Chen, S.P.; Bian, X.W.; Guan, X.Y.; Lin, M.C.; Zeng, Y.X.; et al. The putative tumour suppressor microRNA-124 modulates hepatocellular carcinoma cell aggressiveness by repressing ROCK2 and EZH2. Gut 2012, 61, 278–289. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Guo, W.; Shi, J.; Li, N.; Yu, X.; Xue, J.; Fu, X.; Chu, K.; Lu, C.; Zhao, J.; et al. MicroRNA-135a contributes to the development of portal vein tumor thrombus by promoting metastasis in hepatocellular carcinoma. J. Hepatol. 2012, 56, 389–396. [Google Scholar] [CrossRef] [PubMed]
- Wong, C.C.; Wong, C.M.; Tung, E.K.; Au, S.L.; Lee, J.M.; Poon, R.T.; Man, K.; Ng, I.O. The MicroRNA miR-139 suppresses metastasis and progression of hepatocellular carcinoma by down-regulating rho-kinase 2. Gastroenterology 2011, 140, 322–331. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Song, B.; Song, W.; Liu, J.; Sun, A.; Wu, D.; Yu, H.; Lian, J.; Chen, L.; Han, J. Underexpressed microRNA-199b-5p targets Hypoxia-Inducible Factor-1α in hepatocellular carcinoma and predicts prognosis of hepatocellular carcinoma patients. J. Gastroenterol. Hepatol. 2011, 26, 1630–1637. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.Y.; Han, Z.B.; Fan, J.W.; Xia, J.; Wu, J.Y.; Qiu, G.Q.; Tang, H.M.; Peng, Z.H. miR-203 expression predicts outcome after liver transplantation for hepatocellular carcinoma in cirrhotic liver. Med. Oncol. 2011, 29, 1859–1865. [Google Scholar] [CrossRef] [PubMed]
- Karakatsanis, A.; Papaconstantinou, I.; Gazouli, M.; Lyberopoulou, A.; Polymeneas, G.; Voros, D. Expression of microRNAs, miR-21, miR-31, miR-122, miR-145, miR-146a, miR-200c, miR-221, miR-222, and miR-223 in patients with hepatocellular carcinoma or intrahepatic cholangiocarcinoma and its prognostic significance. Mol. Carcinog. 2013, 52, 297–303. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Yang, Y.; Yang, T.; Liu, Y.; Li, A.; Fu, S.; Wu, M.; Pan, Z.; Zhou, W. microRNA-22, downregulated in hepatocellular carcinoma and correlated with prognosis, suppresses cell proliferation and tumourigenicity. Br. J. Cancer 2010, 103, 1215–1220. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Liu, X.; Lin, L.; Hou, J.; Li, N.; Wang, C.; Wang, P.; Zhang, Q.; Zhang, P.; Zhou, W.; et al. MicroRNA-99a inhibits hepatocellular carcinoma growth and correlates with prognosis of patients with hepatocellular carcinoma. J. Biol. Chem. 2011, 286, 36677–36685. [Google Scholar] [CrossRef] [PubMed]
- Ji, J.; Zhao, L.; Budhu, A.; Forgues, M.; Jia, H.L.; Qin, L.X.; Ye, Q.H.; Yu, J.; Shi, X.; Tang, Z.Y.; et al. Let-7g targets collagen type I alpha2 and inhibits cell migration in hepatocellular carcinoma. J. Hepatol. 2010, 52, 690–697. [Google Scholar] [CrossRef] [PubMed]
- Luk, J.M.; Burchard, J.; Zhang, C.; Liu, A.M.; Wong, K.F.; Shek, F.X.; Lee, N.P.; Fan, S.T.; Poon, R.T.; Ivanovska, I.; et al. DLK1-DIO3 genomic imprinted microRNA cluster at 14q32.2 defines a stemlike subtype of hepatocellular carcinoma associated with poor survival. J. Biol. Chem. 2011, 286, 30706–30713. [Google Scholar] [CrossRef] [PubMed]
- Augello, C.; Vaira, V.; Caruso, L.; Destro, A.; Maggioni, M.; Park, Y.N.; Montorsi, M.; Santambrogio, R.; Roncalli, M.; Bosari, S. MicroRNA profiling of hepatocarcinogenesis identifies C19MC cluster as a novel prognostic biomarker in hepatocellular carcinoma. Liver Int. 2012, 32, 772–782. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.H.; Lin, K.H.; Chen, H.C.; Chang, M.L.; Hsu, C.W.; Lai, M.W.; Chen, T.C.; Lee, W.C.; Tseng, Y.H.; Yeh, C.T. Identification of postoperative prognostic microRNA predictors in hepatocellular carcinoma. PLoS ONE 2012, 7, e37188. [Google Scholar] [CrossRef] [PubMed]
- Barry, C.T.; D'Souza, M.; McCall, M.; Safadjou, S.; Ryan, C.; Kashyap, R.; Marroquin, C.; Orloff, M.; Almudevar, A.; Godfrey, T.E. Micro RNA expression profiles as adjunctive data to assess the risk of hepatocellular carcinoma recurrence after liver transplantation. Am. J. Transpl. 2012, 12, 428–437. [Google Scholar] [CrossRef] [PubMed]
- Sato, F.; Hatano, E.; Kitamura, K.; Myomoto, A.; Fujiwara, T.; Takizawa, S.; Tsuchiya, S.; Tsujimoto, G.; Uemoto, S.; Shimizu, K. MicroRNA profile predicts recurrence after resection in patients with hepatocellular carcinoma within the Milan criteria. PLoS ONE 2011, 6, e16435. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Chong, C.C.N.; Chen, G.G.; Lai, P.B.S. A Seven-microRNA Expression Signature Predicts Survival in Hepatocellular Carcinoma. PLoS ONE 2015, 10, e0128628. [Google Scholar] [CrossRef] [PubMed]
- Yin, W.; Nie, Y.; Zhang, Z.; Xie, L.; He, X. miR-193b acts as a cisplatin sensitizer via the caspase-3-dependent pathway in HCC chemotherapy. Oncol. Rep. 2015, 34, 368–374. [Google Scholar] [CrossRef] [PubMed]
- Zhou, C.; Liu, J.; Li, Y.; Liu, L.; Zhang, X.; Ma, C.Y.; Hua, S.C.; Yang, M.; Yuan, Q. MicroRNA-1274a, a modulator of sorafenib induced a disintegrin and metalloproteinase 9 (ADAM9) down-regulation in hepatocellular carcinoma. FEBS Lett. 2011, 585, 1828–1834. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Anwar, S.L.; Lehmann, U. MicroRNAs: Emerging Novel Clinical Biomarkers for Hepatocellular Carcinomas. J. Clin. Med. 2015, 4, 1631-1650. https://doi.org/10.3390/jcm4081631
Anwar SL, Lehmann U. MicroRNAs: Emerging Novel Clinical Biomarkers for Hepatocellular Carcinomas. Journal of Clinical Medicine. 2015; 4(8):1631-1650. https://doi.org/10.3390/jcm4081631
Chicago/Turabian StyleAnwar, Sumadi Lukman, and Ulrich Lehmann. 2015. "MicroRNAs: Emerging Novel Clinical Biomarkers for Hepatocellular Carcinomas" Journal of Clinical Medicine 4, no. 8: 1631-1650. https://doi.org/10.3390/jcm4081631
APA StyleAnwar, S. L., & Lehmann, U. (2015). MicroRNAs: Emerging Novel Clinical Biomarkers for Hepatocellular Carcinomas. Journal of Clinical Medicine, 4(8), 1631-1650. https://doi.org/10.3390/jcm4081631






