Skin Microbiome Shifts in Various Dermatological Conditions
Abstract
1. Introduction
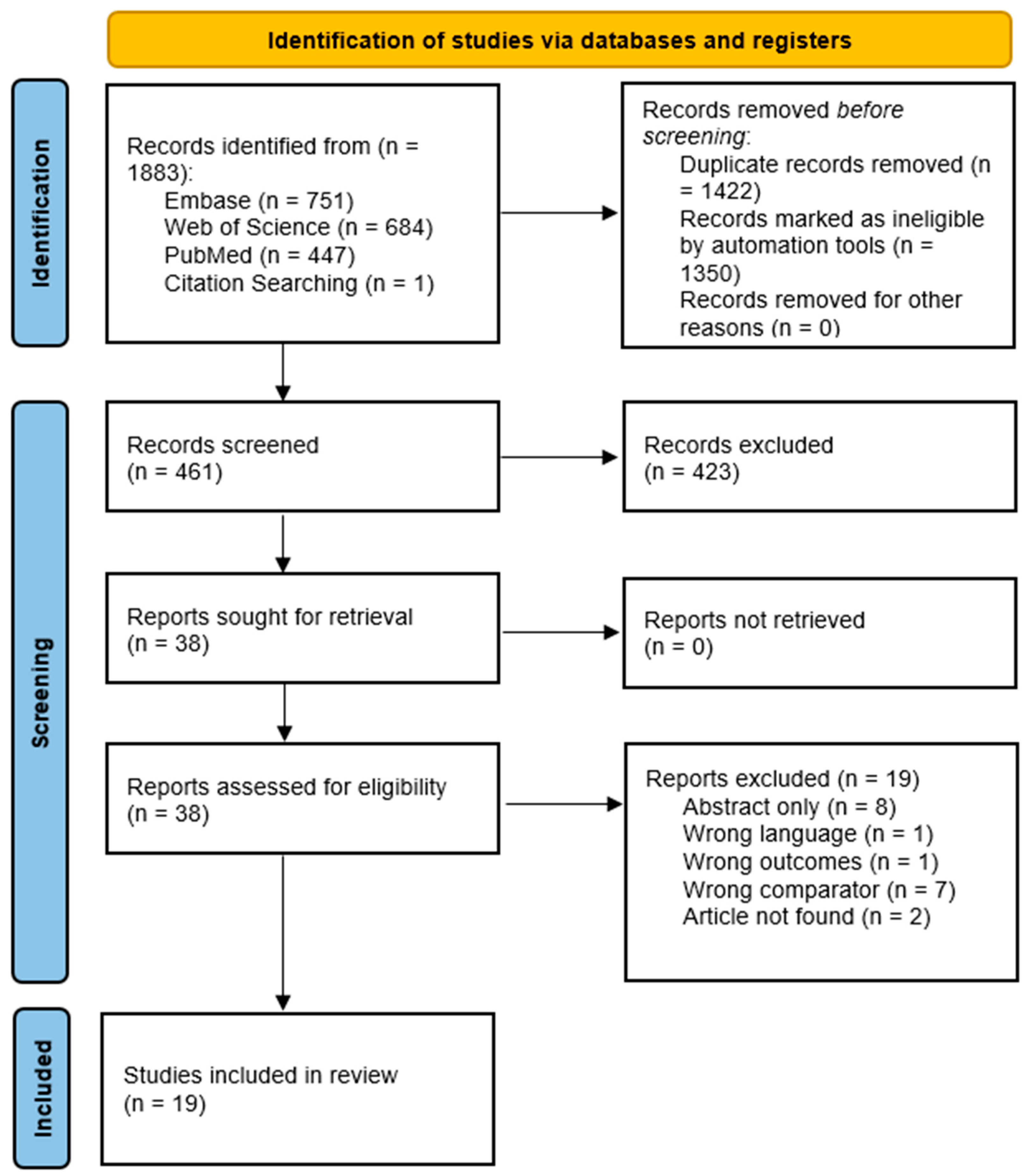
2. Materials and Methods
3. Results
3.1. Dermatologic Conditions
3.1.1. Acne Vulgaris
3.1.2. Atopic Dermatitis
3.1.3. Androgenetic Alopecia
3.1.4. Diaper Dermatitis
3.1.5. Hand Eczema
3.1.6. Lamellar Ichthyosis
3.1.7. Psoriasis
3.1.8. Rosacea
3.1.9. Seborrheic Dermatitis
4. Discussion
5. Future Studies
6. Limitations
7. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| rRNA | Ribosomal RNA |
| WGS | Whole-genome sequencing |
| PRISMA-ScR | Preferred Reporting Items for Scoping reviews and Meta-analysis Extension for Scoping Reviews |
| MRSA | Methicillin resistant staphylococcus aureus |
| LI | Lamellar Ichthyosis |
| RCT | Randomized-controlled trial |
| AD | Atopic dermatitis |
| AGA | Androgenetic alopecia |
| LefSe | Linear discriminant analysis effect size |
| SD | Seborrheic dermatitis |
| SCORAD | Shannon diversity index and Severity Scoring of Atopic Dermatitis |
| hBD-2 | Human beta defensin 2 |
| DD | Diaper dermatitis |
References
- Linehan, J.L.; Harrison, O.J.; Han, S.-J.; Byrd, A.L.; Vujkovic-Cvijin, I.; Villarino, A.V.; Sen, S.K.; Shaik, J.; Smelkinson, M.; Tamoutounour, S.; et al. Non-classical Immunity Controls Microbiota Impact on Skin Immunity and Tissue Repair. Cell 2018, 172, 784–796.e18. [Google Scholar] [CrossRef]
- Grice, E.A.; Kong, H.H.; Conlan, S.; Deming, C.B.; Davis, J.; Young, A.C.; NISC Comparative Sequencing Program; Bouffard, G.G.; Blakesley, R.W.; Murray, P.R.; et al. Topographical and temporal diversity of the human skin microbiome. Science 2009, 324, 1190–1192. [Google Scholar] [CrossRef]
- Skowron, K.; Bauza-Kaszewska, J.; Kraszewska, Z.; Wiktorczyk-Kapischke, N.; Grudlewska-Buda, K.; Kwiecińska-Piróg, J.; Wałecka-Zacharska, E.; Radtke, L.; Gospodarek-Komkowska, E. Human Skin Microbiome: Impact of Intrinsic and Extrinsic Factors on Skin Microbiota. Microorganisms 2021, 9, 543. [Google Scholar] [CrossRef]
- Bates, M. The Role of the Skin Microbiome in Health and Disease. IEEE Pulse 2022, 13, 8–13. [Google Scholar] [CrossRef]
- Gallo, R.L.; Nakatsuji, T. Microbial symbiosis with the innate immune defense system of the skin. J. Investig. Dermatol. 2011, 131, 1974–1980. [Google Scholar] [CrossRef] [PubMed]
- Sanschagrin, S.; Yergeau, E. Next-generation sequencing of 16S ribosomal RNA gene amplicons. J. Vis. Exp. 2014, 90, 51709. [Google Scholar]
- Poretsky, R.; Rodriguez-R, L.M.; Luo, C.; Tsementzi, D.; Konstantinidis, K.T.; Rodriguez-Valera, F. Strengths and limitations of 16S rRNA gene amplicon sequencing in revealing temporal microbial community dynamics. PLoS ONE 2014, 9, e93827. [Google Scholar] [CrossRef]
- Ng, P.C.; Kirkness, E.F. Whole genome sequencing. Methods Mol. Biol. 2010, 628, 215–226. [Google Scholar]
- Ranjan, R.; Rani, A.; Metwally, A.; McGee, H.S.; Perkins, D.L. Analysis of the microbiome: Advantages of whole genome shotgun versus 16S amplicon sequencing. Biochem. Biophys. Res. Commun. 2016, 469, 967–977. [Google Scholar] [CrossRef]
- Ruchti, F.; LeibundGut-Landmann, S. New insights into immunity to skin fungi shape our understanding of health and disease. Parasite Immunol. 2023, 45, e12948. [Google Scholar] [CrossRef] [PubMed]
- Tricco, A.C.; Lillie, E.; Zarin, W.; O’Brien, K.K.; Colquhoun, H.; Levac, D.; Moher, D.; Peters, M.D.J.; Horsley, T.; Weeks, L.; et al. PRISMA Extension for Scoping Reviews (PRISMA-ScR): Checklist and Explanation. Ann. Intern. Med. 2018, 169, 467–473. [Google Scholar] [CrossRef]
- Dreno, B.; Martin, R.; Moyal, D.; Henley, J.B.; Khammari, A.; Seité, S. Skin microbiome and acne vulgaris: Staphylococcus, a new actor in acne. Exp. Dermatol. 2017, 26, 798–803. [Google Scholar] [CrossRef] [PubMed]
- Coughlin, C.C.; Swink, S.M.; Horwinski, J.; Sfyroera, G.; Bugayev, J.; Grice, E.A.; Yan, A.C. The preadolescent acne microbiome: A prospective, randomized, pilot study investigating characterization and effects of acne therapy. Pediatr. Dermatol. 2017, 34, 661–664. [Google Scholar] [CrossRef] [PubMed]
- Callewaert, C.; Nakatsuji, T.; Knight, R.; Kosciolek, T.; Vrbanac, A.; Kotol, P.; Ardeleanu, M.; Hultsch, T.; Guttman-Yassky, E.; Bissonnette, R.; et al. IL-4Ralpha Blockade by Dupilumab Decreases Staphylococcus aureus Colonization and Increases Microbial Diversity in Atopic Dermatitis. J. Investig. Dermatol. 2020, 140, 191–202.e7. [Google Scholar] [CrossRef]
- Khadka, V.D.; Key, F.M.; Romo-González, C.; Martínez-Gayosso, A.; Campos-Cabrera, B.L.; Gerónimo-Gallegos, A.; Lynn, T.C.; Durán-McKinster, C.; Coria-Jiménez, R.; Lieberman, T.D.; et al. The Skin Microbiome of Patients With Atopic Dermatitis Normalizes Gradually During Treatment. Front. Cell Infect. Microbiol. 2021, 11, 720674. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.J.; Kim, S.E.; Shin, K.O.; Park, K.; Lee, S.E. Dupilumab Therapy Improves Stratum Corneum Hydration and Skin Dysbiosis in Patients With Atopic Dermatitis. Allergy Asthma Immunol. Res. 2021, 13, 762–775. [Google Scholar] [CrossRef]
- Chandra, J.; Retuerto, M.; Seite, S.; Martin, R.; Kus, M.; Ghannoum, M.A.; Baron, E.; Mukherjee, P.K. Effect of an Emollient on the Mycobiome of Atopic Dermatitis Patients. J. Drugs Dermatol. 2018, 17, 1039–1048. [Google Scholar]
- Gonzalez, M.E.; Schaffer, J.V.; Orlow, S.J.; Gao, Z.; Li, H.; Alekseyenko, A.V.; Blaser, M.J. Cutaneous microbiome effects of fluticasone propionate cream and adjunctive bleach baths in childhood atopic dermatitis. J. Am. Acad. Dermatol. 2016, 75, 481–493.e8. [Google Scholar] [CrossRef]
- Kwon, S.; Choi, J.Y.; Shin, J.-W.; Huh, C.-H.; Park, K.-C.; Du, M.-H.; Yoon, S.; Na, J.-I. Changes in Lesional and Non-lesional Skin Microbiome During Treatment of Atopic Dermatitis. Acta Derm. Venereol. 2019, 99, 284–290. [Google Scholar] [CrossRef] [PubMed]
- Krzysiek, J.; Żurawska-Olszewska, J.; Szczerba, I.; Lesiak, A.; Pastuszak-Lewandoska, D.; Grzegorczyk, J.; Ciążyńska, M.; Narbutt, J. Shift in skin microbiota of children with atopic dermatitis after topical gentian violet application. Postepy Dermatol. Alergol. 2023, 40, 308–314. [Google Scholar] [CrossRef]
- Zeng, J.; Dou, J.; Gao, L.; Xiang, Y.; Huang, J.; Ding, S.; Chen, J.; Zeng, Q.; Luo, Z.; Tan, W.; et al. Topical ozone therapy restores microbiome diversity in atopic dermatitis. Int. Immunopharmacol. 2020, 80, 106191. [Google Scholar] [CrossRef] [PubMed]
- Filaire, E.; Dreux-Zhiga, A.; Boutot, C.; Cabannes, M.; Ranouille, E.; Berthon, J.Y. Characteristics of healthy and androgenetic alopecia scalp microbiome: Effect of Lindera strychnifolia roots extract as a natural solution for its modulation. Int. J. Cosmet. Sci. 2020, 42, 615–621. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Wang, Q.; Ma, L.; Chen, Y.; Gao, Y.; Zhang, G.; Cui, S.; Liang, H.; Song, L.; He, C. Shifts in the skin microbiome associated with diaper dermatitis and emollient treatment amongst infants and toddlers in China. Exp. Dermatol. 2019, 28, 1289–1297. [Google Scholar] [CrossRef]
- Kuwatsuka, S.; Kuwatsuka, Y.; Tomimura, S.; Takenaka, M.; Terasaka, Y.; Izumikawa, K.; Morinaga, Y.; Yanagihara, K.; Murota, H. Impact of daily wearing of fabric gloves on the management of hand eczema: A pilot study in health-care workers. J. Dermatol. 2021, 48, 645–650. [Google Scholar] [CrossRef]
- Nørreslet, L.B.; Lilje, B.; Ingham, A.C.; Edslev, S.M.; Clausen, M.-L.; Plum, F.; Andersen, P.S.; Agner, T. Skin Microbiome in Patients with Hand Eczema and Healthy Controls: A Three-week Prospective Study. Acta Derm. Venereol. 2022, 102, adv00633. [Google Scholar] [CrossRef]
- Singh, M.; Pawar, M. A Case-Control Study of Skin Microbiome in Patients with Lamellar Ichthyosis. Serbian J. Dermatol. Venereol. 2019, 11, 111–118. [Google Scholar] [CrossRef]
- Martin, R.; Henley, J.B.; Sarrazin, P.; Seité, S. Skin Microbiome in Patients With Psoriasis Before and After Balneotherapy at the Thermal Care Center of La Roche-Posay. J. Drugs Dermatol. 2015, 14, 1400–1405. [Google Scholar]
- Xiong, J.; Chen, S.; Wang, P.; Chen, A.; Zheng, Q.; Cai, T. Characterisation of the bacterial microbiome in patients with rosacea and healthy controls. Eur. J. Dermatol. 2023, 33, 612–617. [Google Scholar] [CrossRef]
- Rainer, B.M.; Thompson, K.G.; Antonescu, C.; Florea, L.; Mongodin, E.F.; Bui, J.; Fischer, A.H.; Pasieka, H.B.; Garza, L.A.; Kang, S.; et al. Characterization and Analysis of the Skin Microbiota in Rosacea: A Case-Control Study. Am. J. Clin. Dermatol. 2020, 21, 139–147. [Google Scholar] [CrossRef]
- Yu, R.; Lin, Q.; Zhai, Y.; Mao, Y.; Li, K.; Gao, Y.; Liu, Y.; Fu, L.; Fang, T.; Zhao, M.; et al. Recombinant human thymosin beta-4 (rhTβ4) improved scalp condition and microbiome homeostasis in seborrheic dermatitis. Microb. Biotechnol. 2021, 14, 2152–2163. [Google Scholar] [CrossRef] [PubMed]
- Oge, L.K.; Broussard, A.; Marshall, M.D. Acne Vulgaris: Diagnosis and Treatment. Am. Fam. Physician 2019, 100, 475–484. [Google Scholar] [PubMed]
- Aydemir, E.H. Acne vulgaris. Turk. Arch. Pediatr. 2014, 49, 13–16. [Google Scholar] [CrossRef]
- Baek, J.; Lee, M.G. Oxidative stress and antioxidant strategies in dermatology. Redox Rep. 2016, 21, 164–169. [Google Scholar] [CrossRef] [PubMed]
- Mayslich, C.; Grange, P.A.; Dupin, N. Cutibacterium acnes as an Opportunistic Pathogen: An Update of Its Virulence-Associated Factors. Microorganisms 2021, 9, 303. [Google Scholar] [CrossRef]
- Fishbein, A.B.; Silverberg, J.I.; Wilson, E.J.; Ong, P.Y. Update on Atopic Dermatitis: Diagnosis, Severity Assessment, and Treatment Selection. J. Allergy Clin. Immunol. Pract. 2020, 8, 91–101. [Google Scholar] [CrossRef]
- Koh, L.F.; Ong, R.Y.; Common, J.E. Skin microbiome of atopic dermatitis. Allergol. Int. 2022, 71, 31–39. [Google Scholar] [CrossRef]
- Vijaya Chandra, S.H.; Srinivas, R.; Dawson, T.L.; Common, J.E. Cutaneous Malassezia: Commensal, Pathogen, or Protector? Front. Cell Infect. Microbiol. 2020, 10, 614446. [Google Scholar] [CrossRef]
- Kobayashi, T.; Glatz, M.; Horiuchi, K.; Kawasaki, H.; Akiyama, H.; Kaplan, D.H.; Kong, H.H.; Amagai, M.; Nagao, K. Dysbiosis and Staphylococcus aureus Colonization Drives Inflammation in Atopic Dermatitis. Immunity 2015, 42, 756–766. [Google Scholar] [CrossRef]
- Gong, J.Q.; Lin, L.; Lin, T.; Hao, F.; Zeng, F.Q.; Bi, Z.G.; Yi, D.; Zhao, B. Skin colonization by Staphylococcus aureus in patients with eczema and atopic dermatitis and relevant combined topical therapy: A double-blind multicentre randomized controlled trial. Br. J. Dermatol. 2006, 155, 680–687. [Google Scholar] [CrossRef] [PubMed]
- Ho, C.H.; Sood, T.; Zito, P.M. Androgenetic Alopecia. In StatPearls; StatPearls Publishing LLC.: Treasure Island, FL, USA, 2024. [Google Scholar]
- Lolli, F.; Pallotti, F.; Rossi, A.; Fortuna, M.C.; Caro, G.; Lenzi, A.; Sansone, A.; Lombardo, F. Androgenetic alopecia: A review. Endocrine 2017, 57, 9–17. [Google Scholar] [CrossRef]
- Ho, B.S.Y.; Ho, E.X.P.; Chu, C.W.; Ramasamy, S.; Bigliardi-Qi, M.; de Sessions, P.F.; Bigliardi, P.L. Microbiome in the hair follicle of androgenetic alopecia patients. PLoS ONE 2019, 14, e0216330. [Google Scholar] [CrossRef]
- DeAngelis, Y.M.; Gemmer, C.M.; Kaczvinsky, J.R.; Kenneally, D.C.; Schwartz, J.R.; Dawson, T.L., Jr. Three etiologic facets of dandruff and seborrheic dermatitis: Malassezia fungi, sebaceous lipids, and individual sensitivity. J. Investig. Dermatol. Symp. Proc. 2005, 10, 295–297. [Google Scholar] [CrossRef]
- Saunders, C.W.; Scheynius, A.; Heitman, J. Malassezia fungi are specialized to live on skin and associated with dandruff, eczema, and other skin diseases. PLoS Pathog. 2012, 8, e1002701. [Google Scholar] [CrossRef]
- Dall’oGlio, F.; Musumeci, M.L.; Puglisi, D.F.; Micali, G. A novel treatment of diaper dermatitis in children and adults. J. Cosmet. Dermatol. 2021, 20 (Suppl. S1), 1–4. [Google Scholar] [CrossRef]
- Hertiš Petek, T.; Petek, M.; Petek, T.; Marčun Varda, N. Emerging Links between Microbiome Composition and Skin Immunology in Diaper Dermatitis: A Narrative Review. Children 2022, 9, 112. [Google Scholar] [CrossRef]
- Elston, D.M.; Ahmed, D.D.; Watsky, K.L.; Schwarzenberger, K. Hand dermatitis. J. Am. Acad. Dermatol. 2002, 47, 291–299. [Google Scholar] [CrossRef] [PubMed]
- Joosten, M.D.W.; Clabbers, J.M.K.; Jonca, N.; Mazereeuw-Hautier, J.; Gostyński, A.H. New developments in the molecular treatment of ichthyosis: Review of the literature. Orphanet J. Rare Dis. 2022, 17, 269. [Google Scholar] [CrossRef]
- Metze, D.; Traupe, H.; Süßmuth, K. Ichthyoses-A Clinical and Pathological Spectrum from Heterogeneous Cornification Disorders to Inflammation. Dermatopathology 2021, 8, 107–123. [Google Scholar] [CrossRef]
- Tham, K.C.; Lefferdink, R.; Duan, K.; Lim, S.S.; Wong, X.C.; Ibler, E.; Wu, B.; Abu-Zayed, H.; Rangel, S.M.; Del Duca, E.; et al. Distinct skin microbiome community structures in congenital ichthyosis. Br. J. Dermatol. 2022, 187, 557–570. [Google Scholar] [CrossRef]
- Malik, K.; He, H.; Huynh, T.N.; Tran, G.; Mueller, K.; Doytcheva, K.; Renert-Yuval, Y.; Czarnowicki, T.; Magidi, S.; Chou, M.; et al. Ichthyosis molecular fingerprinting shows profound T(H)17 skewing and a unique barrier genomic signature. J. Allergy Clin. Immunol. 2019, 143, 604–618. [Google Scholar] [CrossRef]
- Paller, A.S.; Renert-Yuval, Y.; Suprun, M.; Esaki, H.; Oliva, M.; Huynh, T.N.; Ungar, B.; Kunjravia, N.; Friedland, R.; Peng, X.; et al. An IL-17-dominant immune profile is shared across the major orphan forms of ichthyosis. J. Allergy Clin. Immunol. 2017, 139, 152–165. [Google Scholar] [CrossRef] [PubMed]
- Rachakonda, T.D.; Schupp, C.W.; Armstrong, A.W. Psoriasis prevalence among adults in the United States. J. Am. Acad. Dermatol. 2014, 70, 512–516. [Google Scholar] [CrossRef] [PubMed]
- Korman, N.J. Management of psoriasis as a systemic disease: What is the evidence? Br. J. Dermatol. 2020, 182, 840–848. [Google Scholar] [CrossRef]
- Olejniczak-Staruch, I.; Ciążyńska, M.; Sobolewska-Sztychny, D.; Narbutt, J.; Skibińska, M.; Lesiak, A. Alterations of the Skin and Gut Microbiome in Psoriasis and Psoriatic Arthritis. Int. J. Mol. Sci. 2021, 22, 3998. [Google Scholar] [CrossRef]
- Oge’, L.K.; Muncie, H.L.; Phillips-Savoy, A.R. Rosacea: Diagnosis and Treatment. Am. Fam. Physician 2015, 92, 187–196. [Google Scholar]
- Picardo, M.; Ottaviani, M. Skin microbiome and skin disease: The example of rosacea. J. Clin. Gastroenterol. 2014, 48 (Suppl. S1), S85–S86. [Google Scholar] [CrossRef]
- van Zuuren, E.J.; Arents, B.W.; van der Linden, M.M.; Vermeulen, S.; Fedorowicz, Z.; Tan, J. Rosacea: New Concepts in Classification and Treatment. Am. J. Clin. Dermatol. 2021, 22, 457–465. [Google Scholar] [CrossRef] [PubMed]
- Daou, H.; Daou, H.; Paradiso, M.; Hennessy, K.; Seminario-Vidal, L. Rosacea and the Microbiome: A Scoping review. Dermatol. Ther. 2021, 11, 1–12. [Google Scholar] [CrossRef]
- Nakamura, K.; O’nEill, A.M.; Williams, M.R.; Cau, L.; Nakatsuji, T.; Horswill, A.R.; Gallo, R.L. Short chain fatty acids produced by Cutibacterium acnes inhibit biofilm formation by Staphylococcus epidermidis. Sci. Rep. 2020, 10, 21237. [Google Scholar] [CrossRef]
- Tucker, D.; Masood, S. Seborrheic Dermatitis. In StatPearls; StatPearls Publishing LLC.: Treasure Island, FL, USA, 2024. [Google Scholar]
- Schwartz, J.; Messenger, A.; Tosti, A.; Todd, G.; Hordinsky, M.; Hay, R.; Wang, X.; Zachariae, C.; Kerr, K.; Henry, J.; et al. A comprehensive pathophysiology of dandruff and seborrheic dermatitis-towards a more precise definition of scalp health. Acta Derm. Venereol. 2013, 93, 131–137. [Google Scholar] [CrossRef]
- Tao, R.; Li, R.; Wang, R. Skin microbiome alterations in seborrheic dermatitis and dandruff: A scoping review. Exp. Dermatol. 2021, 30, 1546–1553. [Google Scholar] [CrossRef]
- Lin, Q.; Panchamukhi, A.; Li, P.; Shan, W.; Zhou, H.; Hou, L.; Chen, W. Malassezia and Staphylococcus dominate scalp microbiome for seborrheic dermatitis. Bioprocess. Biosyst. Eng. 2021, 44, 965–975. [Google Scholar] [CrossRef]
- Jiang, Y.; Chen, Y.; Yu, Q.; Shi, Y. Biologic and Small-Molecule Therapies for Moderate-to-Severe Psoriasis: Focus on Psoriasis Comorbidities. BioDrugs 2023, 37, 35–55. [Google Scholar] [CrossRef] [PubMed]
- Butala, S.; Castelo-Soccio, L.; Seshadri, R.; Simpson, E.L.; O’sHea, J.J.; Bieber, T.; Paller, A.S. Biologic Versus Small Molecule Therapy for Treating Moderate to Severe Atopic Dermatitis: Clinical Considerations. J. Allergy Clin. Immunol. Pract. 2023, 11, 1361–1373. [Google Scholar] [CrossRef] [PubMed]
- Hwang, J.; Thompson, A.; Jaros, J.; Blackcloud, P.; Hsiao, J.; Shi, V.Y. Updated understanding of Staphylococcus aureus in atopic dermatitis: From virulence factors to commensals and clonal complexes. Exp. Dermatol. 2021, 30, 1532–1545. [Google Scholar] [CrossRef] [PubMed]
- Paharik, A.E.; Parlet, C.P.; Chung, N.; Todd, D.A.; Rodriguez, E.I.; Van Dyke, M.J.; Cech, N.B.; Horswill, A.R. Coagulase-Negative Staphylococcal Strain Prevents Staphylococcus aureus Colonization and Skin Infection by Blocking Quorum Sensing. Cell Host Microbe 2017, 22, 746–756.e5. [Google Scholar] [CrossRef]
- Miller, M.B.; Bassler, B.L. Quorum sensing in bacteria. Annu. Rev. Microbiol. 2001, 55, 165–199. [Google Scholar] [CrossRef]
- Jung, G.W.; Tse, J.E.; Guiha, I.; Rao, J. Prospective, randomized, open-label trial comparing the safety, efficacy, and tolerability of an acne treatment regimen with and without a probiotic supplement and minocycline in subjects with mild to moderate acne. J. Cutan. Med. Surg. 2013, 17, 114–122. [Google Scholar] [CrossRef]
- Cunha, B.A. Antibiotic side effects. Med. Clin. N. Am. 2001, 85, 149–185. [Google Scholar] [CrossRef]
- Friedman, N.D.; Temkin, E.; Carmeli, Y. The negative impact of antibiotic resistance. Clin. Microbiol. Infect. 2016, 22, 416–422. [Google Scholar] [CrossRef]
- Shallcross, L.J.; Howard, S.J.; Fowler, T.; Davies, S.C. Tackling the threat of antimicrobial resistance: From policy to sustainable action. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2015, 370, 20140082. [Google Scholar] [CrossRef]
- Tester, R.; Al-Ghazzewi, F. The role of pre-and probiotics in skin care. Inside Cosmeceuticals 2012, 1, 5–9. [Google Scholar]
- Williams, N.T. Probiotics. Am. J. Health Syst. Pharm. 2010, 67, 449–458. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Dunaway, S.; Champer, J.; Kim, J.; Alikhan, A. Changing our microbiome: Probiotics in dermatology. Br. J. Dermatol. 2020, 182, 39–46. [Google Scholar] [CrossRef]
- De Pessemier, B.; Grine, L.; Debaere, M.; Maes, A.; Paetzold, B.; Callewaert, C. Gut-Skin Axis: Current Knowledge of the Interrelationship between Microbial Dysbiosis and Skin Conditions. Microorganisms 2021, 9, 353. [Google Scholar] [CrossRef]
- Maguire, M.; Maguire, G. The role of microbiota, and probiotics and prebiotics in skin health. Arch. Dermatol. Res. 2017, 309, 411–421. [Google Scholar] [CrossRef]
- Chen, Y.-H.; Wu, C.-S.; Chao, Y.-H.; Lin, C.-C.; Tsai, H.-Y.; Li, Y.-R.; Chen, Y.-Z.; Tsai, W.-H.; Chen, Y.-K. Lactobacillus pentosus GMNL-77 inhibits skin lesions in imiquimod-induced psoriasis-like mice. J. Food Drug Anal. 2017, 25, 559–566. [Google Scholar] [CrossRef] [PubMed]
- Pagnini, C.; Saeed, R.; Bamias, G.; Arseneau, K.O.; Pizarro, T.T.; Cominelli, F. Probiotics promote gut health through stimulation of epithelial innate immunity. Proc. Natl. Acad. Sci. USA 2010, 107, 454–459. [Google Scholar] [CrossRef]
- Abrahamsson, T.R.; Jakobsson, T.; Böttcher, M.F.; Fredrikson, M.; Jenmalm, M.C.; Björkstén, B.; Oldaeus, G. Probiotics in prevention of IgE-associated eczema: A double-blind, randomized, placebo-controlled trial. J. Allergy Clin. Immunol. 2007, 119, 1174–1180. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.Y.; Kwon, J.H.; Ahn, S.H.; Lee, S.I.; Han, Y.S.; Choi, Y.O.; Lee, S.Y.; Ahn, K.M.; Ji, G.E. Effect of probiotic mix (Bifidobacterium bifidum, Bifidobacterium lactis, Lactobacillus acidophilus) in the primary prevention of eczema: A double-blind, randomized, placebo-controlled trial. Pediatr. Allergy Immunol. 2010, 21, e386–e393. [Google Scholar] [CrossRef]
- Jiang, Z.; Chen, Z.; Xu, Y.; Li, H.; Li, Y.; Peng, L.; Shan, H.; Liu, X.; Wu, H.; Wu, L.; et al. Low-Frequency Ultrasound Sensitive Piezo1 Channels Regulate Keloid-Related Characteristics of Fibroblasts. Adv. Sci. 2024, 11, 2305489. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Jiang, S.; Xiang, F.; Li, C.; Li, S.; Gao, T.; He, K.; Chen, J.; Zhang, J.; Zhang, J. White patchy skin lesion classification using feature enhancement and interaction transformer module. Biomed. Signal Process. Control. 2025, 107, 107819. [Google Scholar] [CrossRef]
- Chen, W.; Liu, Y.; Wang, C.; Zhu, J.; Li, G.; Liu, C.-L.; Lin, L. Cross-Modal Causal Representation Learning for Radiology Report Generation. IEEE Trans. Image Process. 2025, 34, 2970–2985. [Google Scholar] [CrossRef]
- Shu, M.; Fan, Z.; Huang, B.; Wang, C. Retrospective analysis of clinical features of pembrolizumab induced psoriasis. Investig. New Drugs 2025, 43, 582–587. [Google Scholar] [CrossRef]
- Wu, Z.; Li, X.; Huang, R.; He, B.; Wang, C. Clinical features, treatment, and prognosis of pembrolizumab-induced Stevens-Johnson syndrome / toxic epidermal necrolysis. Investig. New Drugs 2025, 43, 74–80. [Google Scholar] [CrossRef] [PubMed]
| Author | Disease | Female to Male Ratio (F to M) | Mean Age (Years) ± SD | Study Design | Comparison Groups | Sample Size, n | Method of Sequencing | Increased | Decreased | Other Changes |
|---|---|---|---|---|---|---|---|---|---|---|
| Dreno [12] | Acne | 15 to 11 | 23 ± 6.5 | DB, split-face RCT | Lesional vs. nonlesional | 26 | 16S rRNA sequencing | Increased Staphylococcus (33.87% vs. 26.85%) and Firmicutes (52.01% vs. 47.01%) in lesional skin | Decreased Proteobacteria (34.10% vs. 28.90%) in lesional skin | Similar Shannon alpha diversity index score |
| Coughlin [13] | Acne Vulgaris | 12 to 4 | NA | Prospective Pilot Study | Healthy vs. diseased | 16 | 16S rRNA sequencing | Increased Staphylococcus and Propionibacterium in diseased skin | N/A | Alpha diversity was higher in diseased skin four sites (midline forehead, dorsum of the nose, medial left cheek, and chin) |
| Callewaert [14] | AD | 24 to 29 | NA | DB RCT | Lesional vs. nonlesional | 53 | DNA extraction, 16S rRNA V4 amplicon sequencing via Quantitative PCR | Staphylococcus aureus in lesional skin | Lower Shannon alpha diversity index score in lesional skin | |
| Khadka [15] | AD | 21 to 21 | 11 | RCT | Healthy vs. diseased | 42 | 16S rRNA sequencing | Increased S. aureus | Decreased Shannon alpha diversity | Relative abundance of S. aureus positively correlated with disease severity as measured by SCORAD (rho = 0.545) S. epidermidis and S. hominis were inversely correlated with SCORAD |
| Lee [16] | AD | NA | 28.3 for healthy, 34.2 for severe AD | RCT | Healthy vs. diseased | 20 | 16S rRNA sequencing | NA | Decreased Cutibacterium and Lactobacillus in diseased skin | Increased Human beta defensin 2 (hBD-2) and lower Shannon diversity index score in lesional skin |
| Chandra [17] | AD | 32 to 17 | 10.5 | Non-RCT | Lesional vs. nonlesional | 49 | 16S rRNA sequencing, ITS1 RNA gene showed fungal composition analysis | Increased Alternaria, Coniosporium, Debaryomyce, Capnodiales | NA | Gram-positive Corynebacterium kroppenstedtiian and Staphlycoccus pettenkoferi showed significantly positive correlations with pathogenic Candida species in lesional skin; Pseudomonas spp. correlated significantly with pathogenic Aspergillus and Candida spp. |
| Gonzalez [18]) | AD | NA | NA | Single-blind RCT | Healthy vs. diseased | 35 | 16S rRNA sequencing | Increased Staphylococcus aureus and Staphylococcus species in lesional skin. Nonlesional generally had less than 25% composition of Staphylococci whereas lesional skin had 60–70%. | Decreased Corynebacterium and Propionibacterium in diseased skin | Increased baseline total bacteria density by approximately 10-fold, and decreased community richness and Shannon diversity index in diseased skin |
| Kwon [19] | AD | NA | NA | RCT | Lesional vs. nonlesional | 18 | 16S rRNA sequencing | Increased Staphylococcus aureus, Staphylococcus species in lesional skin | Decreased Shannon Diversity in lesional skin | Haemophilus parainfluenzae, Streptococcus pseudopneumoniae, P. acnes, and Corynebacterium pseudogenitalium showed significant negative correlations with S. aureus in lesional skin |
| Krzysiek [20] | AD | NA | Median, 6.8 for AD group, 8.7 for healthy | Non-RCT | Healthy vs. diseased | 60 | CHROMagar plates for S. aureus and Malassezia | Increased Staphylococcus aureus, Staphylococcus species in AD skin; Increased Malassezia species in AD skin | Decreased Corynebacterium urealyticum in AD skin | The number of S. aureus on lesional skin positively correlated with severity of disease according to validated scoring systems |
| Zeng [21] | AD | 4 to 8 | 17.08 ± 6.72 | Split side RCT | Lesional vs. nonlesional | 12 | 16S rRNA sequencing | NA | NA | Lower Shannon alpha diversity index score and negative correlation between SCORAD and Shannon diversity index score in lesional skin |
| Filaire [22] | Androgenetic Alopecia | 0 to 24 | 50.5 ± 3.2 for AGA, 48.6 ± 2.1 for healthy | Non-RCT | Healthy vs. diseased | 24 | 16S rRNA sequencing, ITS1 rRNA sequencing | Increased Cutibacterium acnes (84% vs. 79%) and Stenotrophomanas geniculata (1.6% vs. 0%) in diseased skin | Decreased Staphylococcus epidermidis (10% vs. 12%) in diseased skin | Alpha diversity did not differ and ratio of Cutibacterium acnes to Staphylococcus epidermidis was significantly higher in diseased skin |
| Zheng [23] | Diaper Dermatitis | NA | NA | Non-RCT | Healthy vs. diseased | 85 | 16S rRNA sequencing | Significantly increased Shannon diversity and Chao index (richness) Significantly increased Proteobacteria, Enterococcus, Erwinia, Pseudomonas, Rhodococcus, Acinetobacter, and Ruminococcus | Significantly decreased Clostridium and Actinomyces | PCoA distribution in healthy samples were found to be more concentrated, indicating higher intra-group similarities |
| Kuwatsuka [24] | Hand Eczema | 1 to 0 | 34.3 | Non-RCT | Healthy vs. diseased | 16 | 16S rRNA sequencing | NA | NA | No difference in alpha or beta diversity between hand eczema and control groups |
| Norreslet [25] | Hand Eczema | 28 to 22 | 40.1 ± 11.7 | Non-RCT | Healthy vs. diseased | 50 | 16S rRNA sequencing | Significantly increased S. aureus in diseased skin versus healthy controls | Decreased bacterial alpha diversity in diseased skin | Disease severity was correlated with abundance of S. aureus |
| Singh [26] | Lamellar Ichthyosis | 9 to 18 | 35.56 weeks | Comparative retrospective study | Healthy vs. diseased | 27 | Qualitative bacterial culture and sensitivities from swabs | Increased methicillin resistant Staphylococcus aureus (MRSA), Fusobacterium (16.67% vs. 4.17%), Gram-negative rods consisting of Enterobacter, Proteus, and Klebsiella (52.78% vs. 51.39%), and fungal population mostly involving Candida (22.22% vs. 5.56%) in diseased skin | Decreased lipophilic diphtheroids (11.11% vs. 27.78%), Propionibacterium acnes (5.6% vs. 15.28%), and Micrococci (22.22% vs. 36.11%) in diseased skin | MRSA exclusively seen in LI patients constituting 33.33% of Staphylococcus aureus flora |
| Martin [27] | Psoriasis | 22 to 32 | 59 ± 13 | Non-RCT | Lesional vs. nonlesional | 54 | 16S rRNA sequencing | Increased Firmicute phylum compared to healthy controls | Decreased Proteobacteria phylum compared to healthy controls | No significant differences in Shannon diversity or richness between lesional and nonlesional skin |
| Xiong [28] | Rosacea | 26 to 18 | Median, 27 for rosacea, 26 for healthy | Observational case–control | Healthy vs. diseased | 44 | 16S rRNA sequencing | Increased Staphylococcus epidermidis (19.64% vs. 6.48%) in diseased skin | Decreased actinobacteria (69.07% vs. 86.09%), Cutibacterium acnes (61.79% vs. 79.69%), and firmicutes (8.05% vs. 21.19%) in diseased skin | No significant difference in diversity, statistically insignificant differences in Shannon diversity, Chao, and Simpson index |
| Rainer [29] | Rosacea | 28 to 10 | NA, range 23–65 | Observational case–control | Healthy vs. diseased | 38 | 16S rRNA sequencing | Increased relative abundance of Cutibacterium acnes in diseased skin of female patients (29.7% vs. 27.8%) | Decreased relative abundance of Cutibacterium acnes in diseased skin of male patients (23.8% vs. 57.5%) | Across all age groups, Cutibacterium acnes remained the most abundant species and Corynebacterium kroppenstedtii the second. No significant differences in ecologic diversity of microbiota |
| Yu [30] | SD | NA | NA | Prospective Cohort | Healthy vs. diseased | 92 | 16S rRNA sequencing, LEfSe analysis | Increased amount of 5 fungal genera (Malassezia, Alternaria, Nagnishia, Hanseniaspora, Cladophialophora) and 5 bacterial genera (Staphylococcus, Blautia, Bifidobacterium Xylanimicrobium, Fusobacterium, Lysobacter) in diseased skin | Decreased enrichment in 4 fungal genera (Mycosphaerella, Cladosporium, Rhodotorula, Debaryomyces) in diseased skin | Decreased Shannon diversity, PD_whole_tree index, and relative abundance of microorganisms in diseased skin |
| Disease | Increased | Decreased | Other Changes |
|---|---|---|---|
| Acne | Increased Staphylococcus, Firmicutes, and Cutibacterium | Decreased Proteobacteria | Inconclusive Shannon alpha diversity score differences |
| Atopic Dermatitis | Increased Staphylococcus aureus, Staphylococcus species, Alternaria, Coniosporium, Debaryomyce, Capnodiales, and Malassezia species | Decreased Cutibacterium, Lactobacillus, Corynebacterium, and Propionibacterium | Decreased Shannon diversity index |
| Androgenetic Alopecia | Increased Cutibacterium acnes (84% vs. 79%) and Stenotrophomanas geniculata (1.6% vs. 0%) | Decreased Staphylococcus epidermidis (10% vs. 12%) | Alpha diversity did not differ and ratio of Cutibacterium acnes to Staphylococcus epidermidis was significantly higher in diseased skin |
| Diaper Dermatitis | Increased Proteobacteria, Enterococcus, Erwinia, Pseudomonas, Rhodococcus, Acinetobacter, and Ruminococcus | Significantly decreased Clostridium and Actinomyces | Significantly increased Shannon diversity and Chao index (richness). PCoA distribution in healthy samples were found to be more concentrated, indicating higher intra-group similarities. |
| Hand Eczema | Increased S. aureus | NA | Decreased bacterial alpha diversity in diseased skin. Disease severity was correlated with abundance of S. aureus. No difference in alpha or beta diversity between hand eczema and control groups. |
| Lamellar Ichthyosis | Increased methicillin resistant Staphylococcus aureus (MRSA), Fusobacterium (16.67% vs. 4.17%), Gram negative rods consisting of Enterobacter, Proteus, and Klebsiella (52.78% vs. 51.39%), and fungal population mostly involving Candida (22.22% vs. 5.56%) | Decreased lipophilic diphtheroids (11.11% vs. 27.78%), Propionibacterium acnes (5.6% vs. 15.28%), and Micrococci (22.22% vs. 36.11%) | MRSA exclusively seen in LI patients constituting 33.33% of Staphylococcus aureus flora |
| Psoriasis | Increased Firmicute phylum | Decreased Proteobacteria phylum | No significant differences in Shannon diversity or richness |
| Rosacea | Increased Staphylococcus epidermidis | Decreased Cutibacterium acnes | No significant difference in diversity, statistically insignificant differences in Shannon diversity, Chao, and Simpson index |
| Seborrheic Dermatitis | Increased amount of 5 fungal genera (Malassezia, Alternaria, Nagnishia, Hanseniaspora, Cladophialophora) and 5 bacterial genera (Staphylococcus, Blautia, Bifidobacterium Xylanimicrobium, Fusobacterium, Lysobacter) | Decreased enrichment in 4 fungal genera (Mycosphaerella, Cladosporium, Rhodotorula, Debaryomyces) | Decreased Shannon diversity, PD_whole_tree index, and relative abundance of microorganisms in diseased skin |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, C.H.; Min, M.; Jin, S.S.; Sivamani, R.K. Skin Microbiome Shifts in Various Dermatological Conditions. J. Clin. Med. 2025, 14, 6137. https://doi.org/10.3390/jcm14176137
Lee CH, Min M, Jin SS, Sivamani RK. Skin Microbiome Shifts in Various Dermatological Conditions. Journal of Clinical Medicine. 2025; 14(17):6137. https://doi.org/10.3390/jcm14176137
Chicago/Turabian StyleLee, Conan H., Mildred Min, Sami S. Jin, and Raja K. Sivamani. 2025. "Skin Microbiome Shifts in Various Dermatological Conditions" Journal of Clinical Medicine 14, no. 17: 6137. https://doi.org/10.3390/jcm14176137
APA StyleLee, C. H., Min, M., Jin, S. S., & Sivamani, R. K. (2025). Skin Microbiome Shifts in Various Dermatological Conditions. Journal of Clinical Medicine, 14(17), 6137. https://doi.org/10.3390/jcm14176137







