Antibody-Oligonucleotide Conjugates: A Twist to Antibody-Drug Conjugates
Abstract
1. Introduction
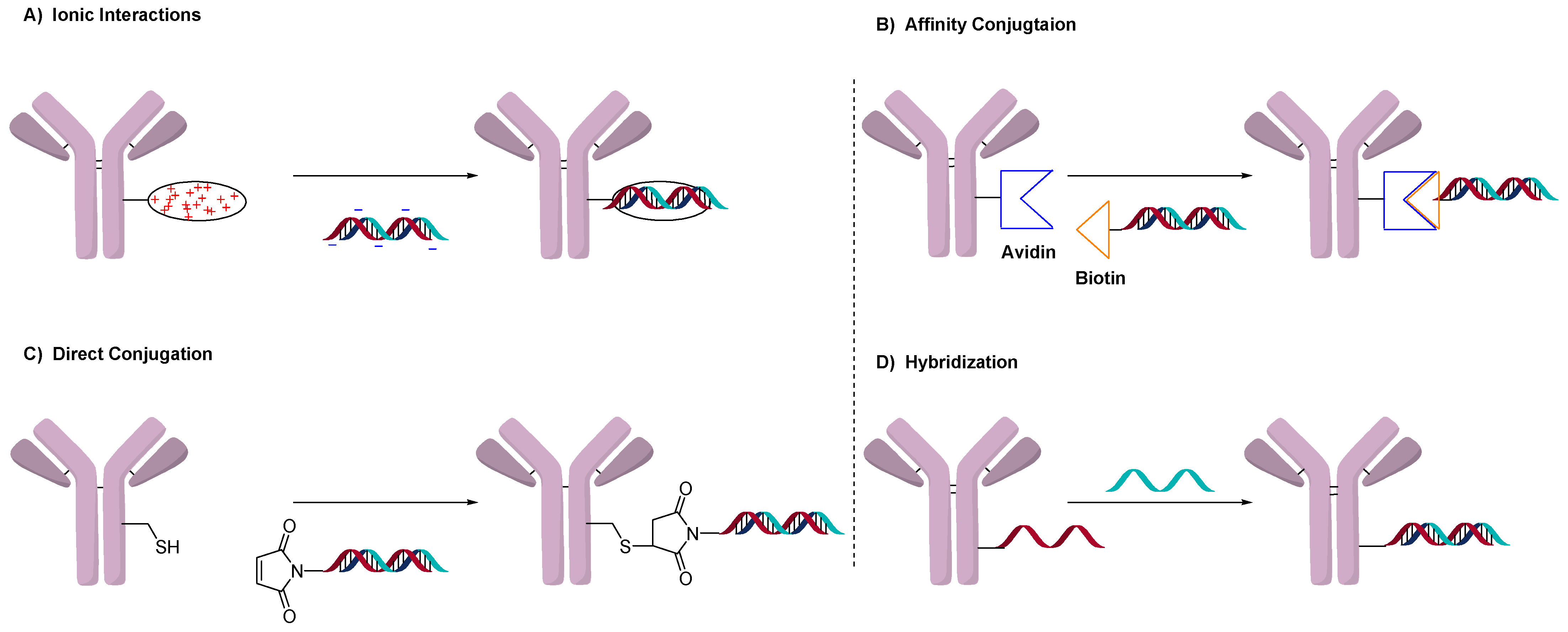
2. Conjugation of Oligonucleotides to Antibodies
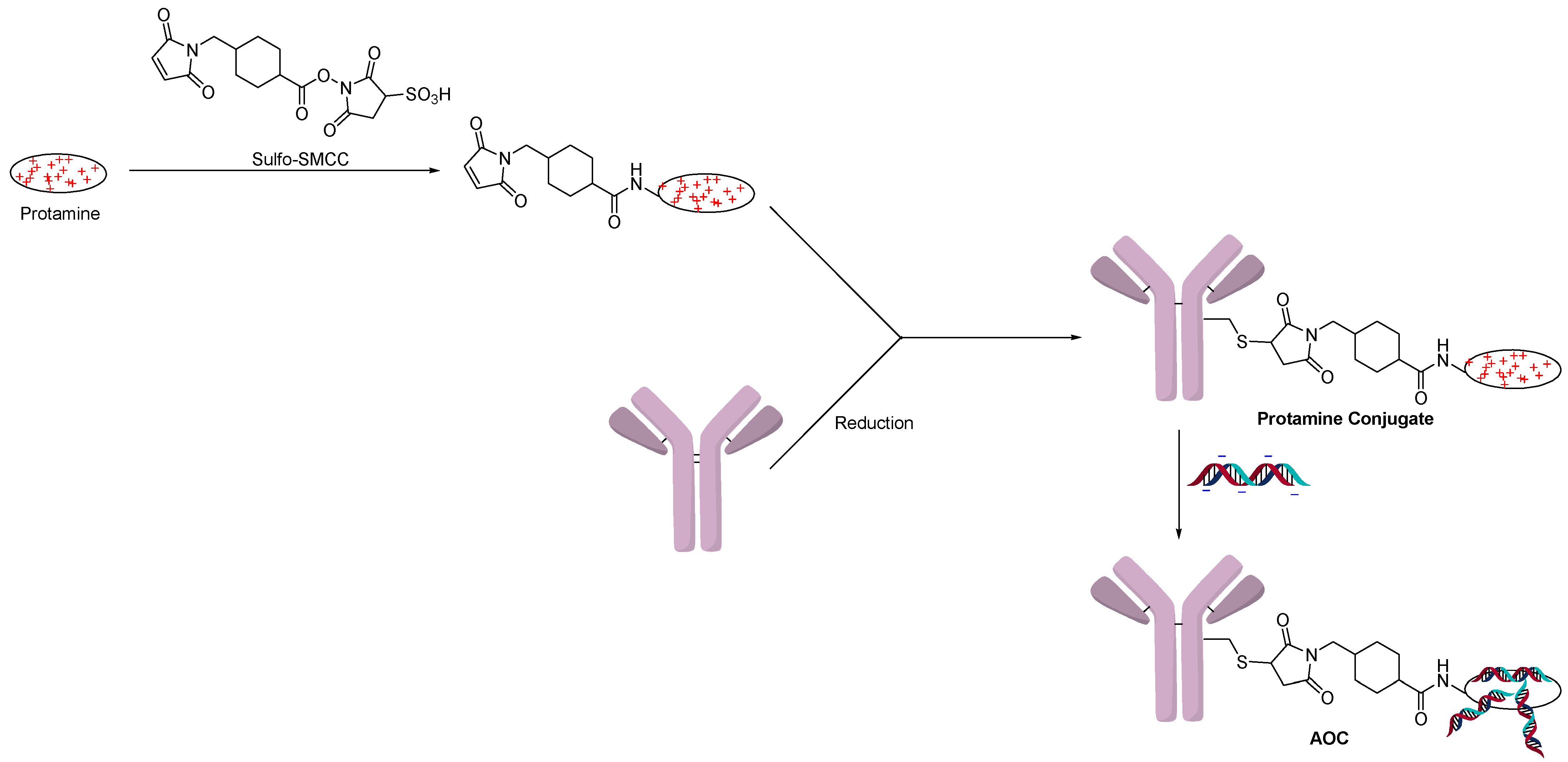
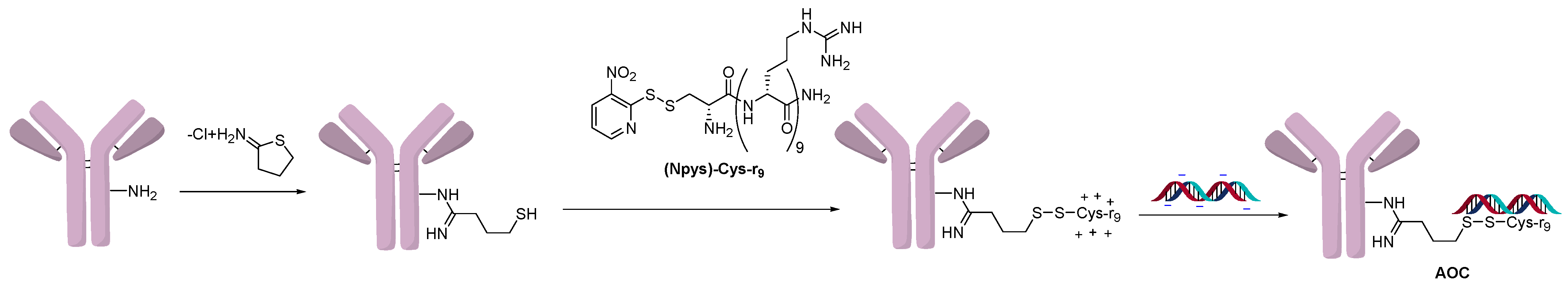
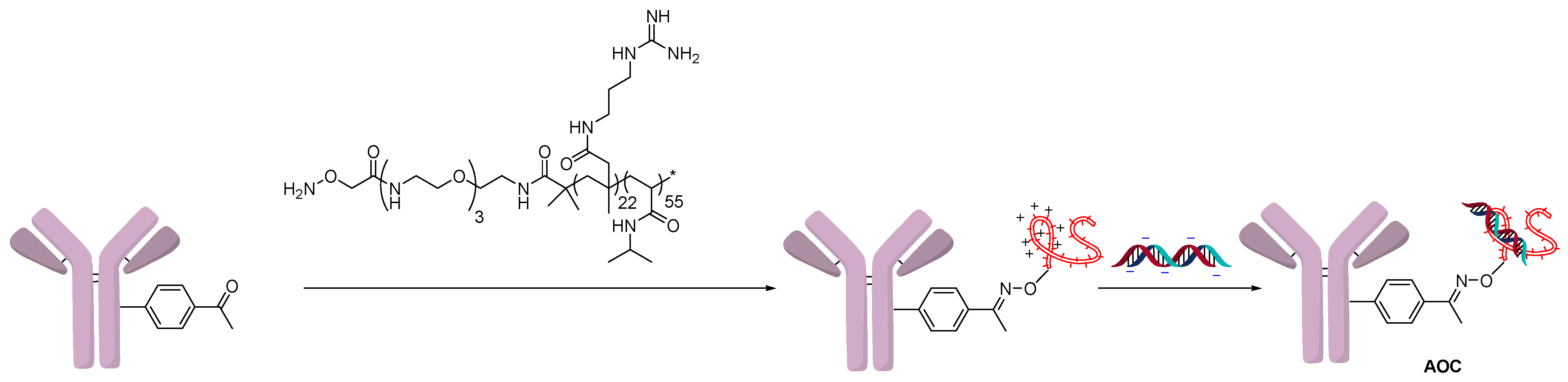
2.1. Conjugation Using Ionic Interactions
2.2. Avidin-Based Conjugation
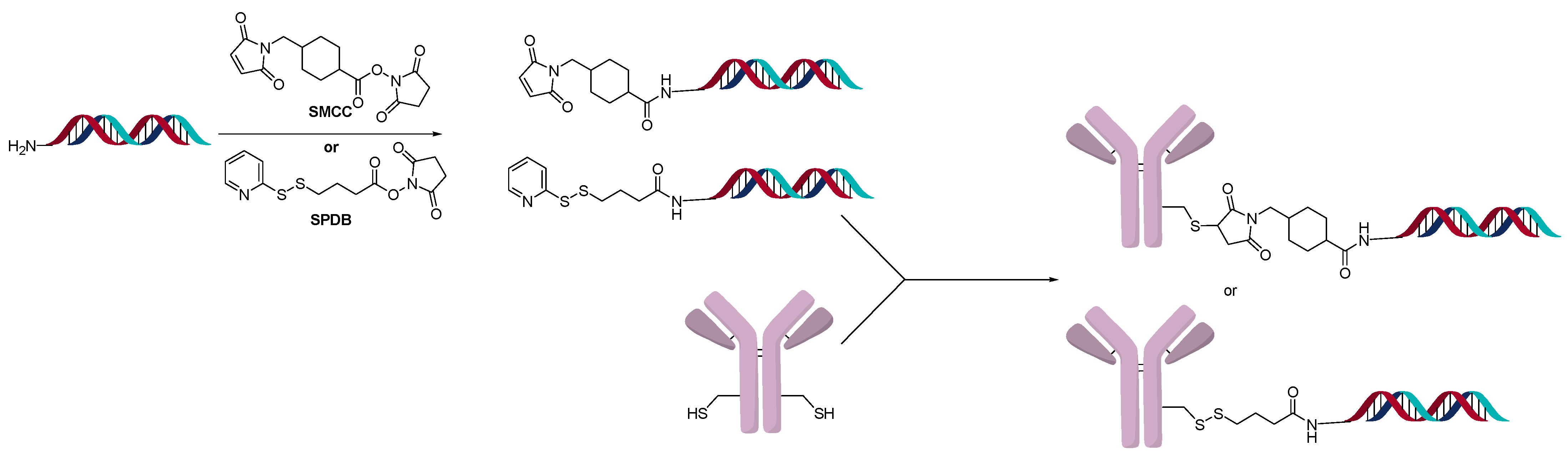
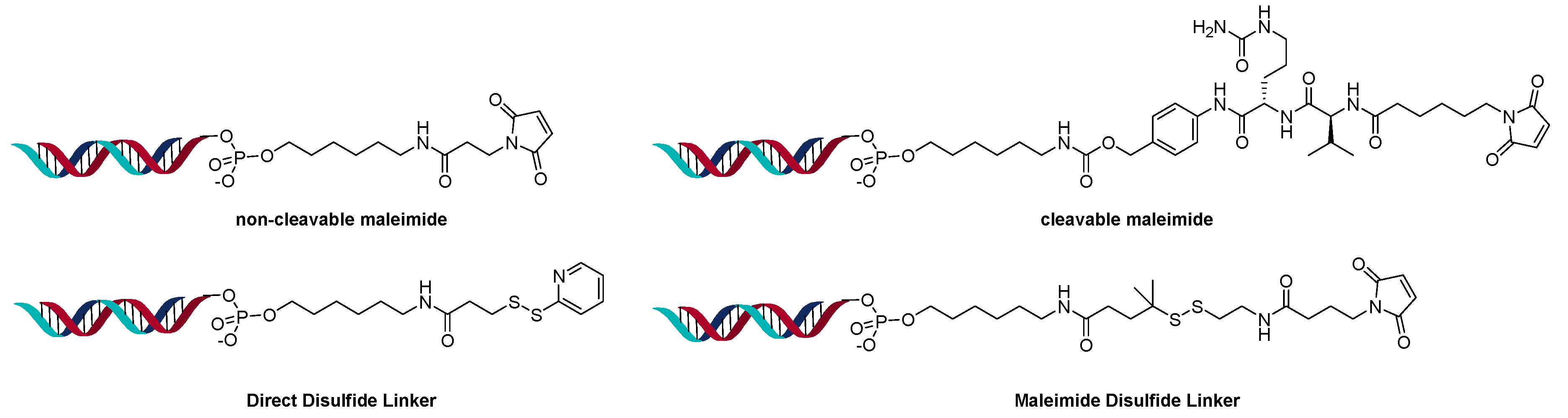
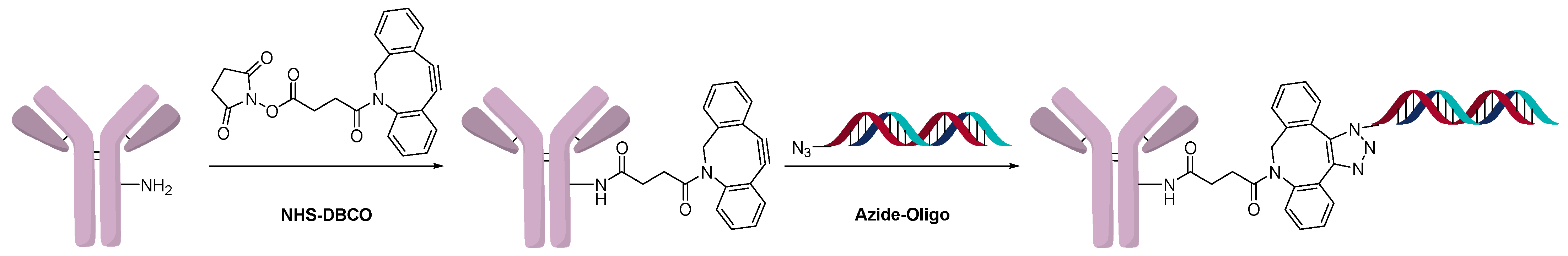
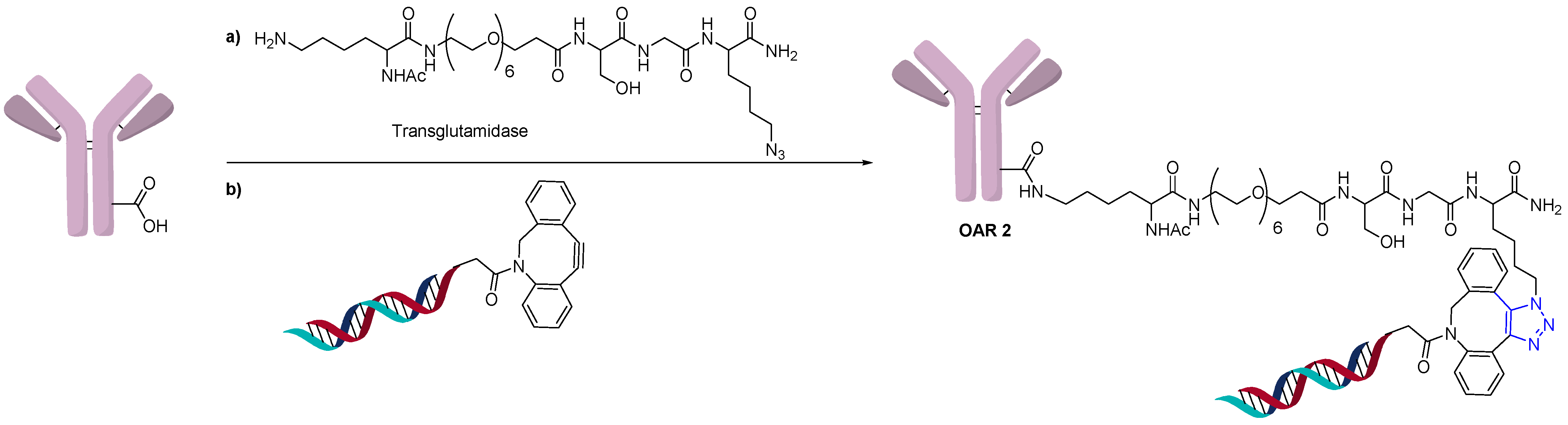
2.3. Direct Conjugation
2.4. DNA Origami or Hybridization Conjugation
2.5. Summary
3. In Vitro Data
3.1. Evaluating Gene Inhibition
3.2. Internalization and Trafficking
3.3. Cytotoxicity
3.4. Stability
3.5. Summary
4. In Vivo Data
4.1. In Vivo Imaging
4.2. Efficacy
4.3. Pharmacokinetics (PK)
4.4. Summary
5. Conclusions and Outlook
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Pysz, I.; Jackson, P.J.M.; Thurston, D.E. CHAPTER 1. Introduction to Antibody–Drug Conjugates (ADCs). In Cytotoxic Payloads for Antibody–Drug Conjugates; Thurston, D.E., Jackson, P.J.M., Eds.; Royal Society of Chemistry: Cambridge, UK, 2019; pp. 1–30. ISBN 978-1-78801-077-1. [Google Scholar]
- Beck, A.; Goetsch, L.; Dumontet, C.; Corvaïa, N. Strategies and Challenges for the next Generation of Antibody-Drug Conjugates. Nat. Rev. Drug Discov. 2017, 16, 315–337. [Google Scholar] [CrossRef]
- Khongorzul, P.; Ling, C.J.; Khan, F.U.; Ihsan, A.U.; Zhang, J. Antibody–Drug Conjugates: A Comprehensive Review. Mol. Cancer Res. 2020, 18, 3–19. [Google Scholar] [CrossRef] [PubMed]
- Roberts, T.C.; Langer, R.; Wood, M.J.A. Advances in Oligonucleotide Drug Delivery. Nat. Rev. Drug Discov. 2020, 19, 673–694. [Google Scholar] [CrossRef] [PubMed]
- Dovgan, I.; Koniev, O.; Kolodych, S.; Wagner, A. Antibody–Oligonucleotide Conjugates as Therapeutic, Imaging, and Detection Agents. Bioconj. Chem. 2019, 30, 2483–2501. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Moellering, R.E. A Concise, Modular Antibody–Oligonucleotide Conjugation Strategy Based on Disuccinimidyl Ester Activation Chemistry. ChemBioChem 2019, 20, cbic.201900027. [Google Scholar] [CrossRef] [PubMed]
- Song, E.; Zhu, P.; Lee, S.-K.; Chowdhury, D.; Kussman, S.; Dykxhoorn, D.M.; Feng, Y.; Palliser, D.; Weiner, D.B.; Shankar, P.; et al. Antibody Mediated in Vivo Delivery of Small Interfering RNAs via Cell-Surface Receptors. Nat. Biotechnol. 2005, 23, 709–717. [Google Scholar] [CrossRef] [PubMed]
- Chandela, A.; Ueno, Y. Systemic Delivery of Small Interfering RNA Therapeutics: Obstacles and Advances. Rev. Agric. Sci. 2019, 7, 10–28. [Google Scholar] [CrossRef]
- Hansen, B.; Linde, S.; Kølendorf, K.; Jensen, F. Absorption of Protamine-Lnsulin in Diabetic Patients—I. Preparation and Characterization of Protamine- 125 I-InsuIin. Horm. Metab. Res. 1979, 11, 85–90. [Google Scholar] [CrossRef]
- Akinc, A.; Thomas, M.; Klibanov, A.M.; Langer, R. Exploring Polyethylenimine-Mediated DNA Transfection and the Proton Sponge Hypothesis. J. Gene Med. 2005, 7, 657–663. [Google Scholar] [CrossRef]
- Peer, D.; Zhu, P.; Carman, C.V.; Lieberman, J.; Shimaoka, M. Selective Gene Silencing in Activated Leukocytes by Targeting SiRNAs to the Integrin Lymphocyte Function-Associated Antigen-1. Proc. Natl. Acad. Sci. USA 2007, 104, 4095–4100. [Google Scholar] [CrossRef]
- Baumer, S.; Baumer, N.; Appel, N.; Terheyden, L.; Fremerey, J.; Schelhaas, S.; Wardelmann, E.; Buchholz, F.; Berdel, W.E.; Muller-Tidow, C. Antibody-Mediated Delivery of Anti-KRAS-SiRNA In Vivo Overcomes Therapy Resistance in Colon Cancer. Clin. Cancer Res. 2015, 21, 1383–1394. [Google Scholar] [CrossRef] [PubMed]
- Bäumer, N.; Berdel, W.E.; Bäumer, S. Immunoprotein-Mediated SiRNA Delivery. Mol. Pharm. 2017, 14, 1339–1351. [Google Scholar] [CrossRef] [PubMed]
- Shi, S.-J.; Wang, L.-J.; Han, D.-H.; Wu, J.-H.; Jiao, D.; Zhang, K.-L.; Chen, J.-W.; Li, Y.; Yang, F.; Zhang, J.-L.; et al. Therapeutic Effects of Human Monoclonal PSMA Antibody-Mediated TRIM24 SiRNA Delivery in PSMA-Positive Castration-Resistant Prostate Cancer. Theranostics 2019, 9, 1247–1263. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Kowolik, C.M.; Swiderski, P.M.; Kortylewski, M.; Yu, H.; Horne, D.A.; Jove, R.; Caballero, O.L.; Simpson, A.J.G.; Lee, F.-T.; et al. Humanized Lewis-Y Specific Antibody Based Delivery of STAT3 SiRNA. ACS Chem. Biol. 2011, 6, 962–970. [Google Scholar] [CrossRef]
- Lu, H.; Wang, D.; Kazane, S.; Javahishvili, T.; Tian, F.; Song, F.; Sellers, A.; Barnett, B.; Schultz, P.G. Site-Specific Antibody–Polymer Conjugates for SiRNA Delivery. J. Am. Chem. Soc. 2013, 135, 13885–13891. [Google Scholar] [CrossRef] [PubMed]
- Xia, C.-F.; Boado, R.J.; Pardridge, W.M. Antibody-Mediated Targeting of SiRNA via the Human Insulin Receptor Using Avidin−Biotin Technology. Mol. Pharm. 2009, 6, 747–751. [Google Scholar] [CrossRef]
- Bolcato-Bellemin, A.-L.; Bonnet, M.-E.; Creusat, G.; Erbacher, P.; Behr, J.-P. Sticky Overhangs Enhance SiRNA-Mediated Gene Silencing. Proc. Natl. Acad. Sci. USA 2007, 104, 16050–16055. [Google Scholar] [CrossRef] [PubMed]
- Hauser, P.V.; Pippin, J.W.; Kaiser, C.; Krofft, R.D.; Brinkkoetter, P.T.; Hudkins, K.L.; Kerjaschki, D.; Reiser, J.; Alpers, C.E.; Shankland, S.J. Novel SiRNA Delivery System to Target Podocytes In Vivo. PLoS ONE 2010, 5, e9463. [Google Scholar] [CrossRef]
- Jain, N.; Smith, S.W.; Ghone, S.; Tomczuk, B. Current ADC Linker Chemistry. Pharm. Res. 2015, 32, 3526–3540. [Google Scholar] [CrossRef] [PubMed]
- Winkler, J. Oligonucleotide Conjugates for Therapeutic Applications. Ther. Deliv. 2013, 4, 791–809. [Google Scholar] [CrossRef]
- Junutula, J.R.; Raab, H.; Clark, S.; Bhakta, S.; Leipold, D.D.; Weir, S.; Chen, Y.; Simpson, M.; Tsai, S.P.; Dennis, M.S.; et al. Site-Specific Conjugation of a Cytotoxic Drug to an Antibody Improves the Therapeutic Index. Nat. Biotechnol. 2008, 26, 925–932. [Google Scholar] [CrossRef] [PubMed]
- Cuellar, T.L.; Barnes, D.; Nelson, C.; Tanguay, J.; Yu, S.-F.; Wen, X.; Scales, S.J.; Gesch, J.; Davis, D.; van Brabant Smith, A.; et al. Systematic Evaluation of Antibody-Mediated SiRNA Delivery Using an Industrial Platform of THIOMAB–SiRNA Conjugates. Nucleic Acids Res. 2015, 43, 1189–1203. [Google Scholar] [CrossRef]
- Sugo, T.; Terada, M.; Oikawa, T.; Miyata, K.; Nishimura, S.; Kenjo, E.; Ogasawara-Shimizu, M.; Makita, Y.; Imaichi, S.; Murata, S.; et al. Development of Antibody-SiRNA Conjugate Targeted to Cardiac and Skeletal Muscles. J. Control. Release 2016, 237, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Gong, H.; Holcomb, I.; Ooi, A.; Wang, X.; Majonis, D.; Unger, M.A.; Ramakrishnan, R. Simple Method To Prepare Oligonucleotide-Conjugated Antibodies and Its Application in Multiplex Protein Detection in Single Cells. Bioconj. Chem. 2016, 27, 217–225. [Google Scholar] [CrossRef] [PubMed]
- Satake, N.; Duong, C.; Yoshida, S.; Oestergaard, M.; Chen, C.; Peralta, R.; Guo, S.; Seth, P.P.; Li, Y.; Beckett, L.; et al. Novel Targeted Therapy for Precursor B-Cell Acute Lymphoblastic Leukemia: Anti-CD22 Antibody-MXD3 Antisense Oligonucleotide Conjugate. Mol. Med. 2016, 22, 632–642. [Google Scholar] [CrossRef]
- Arnold, A.E.; Malek-Adamian, E.; Le, P.U.; Meng, A.; Martínez-Montero, S.; Petrecca, K.; Damha, M.J.; Shoichet, M.S. Antibody-Antisense Oligonucleotide Conjugate Downregulates a Key Gene in Glioblastoma Stem Cells. Mol. Ther. Nucleic Acids 2018, 11, 518–527. [Google Scholar] [CrossRef] [PubMed]
- Maerle, A.V.; Simonova, M.A.; Pivovarov, V.D.; Voronina, D.V.; Drobyazina, P.E.; Trofimov, D.Y.; Alekseev, L.P.; Zavriev, S.K.; Ryazantsev, D.Y. Development of the Covalent Antibody-DNA Conjugates Technology for Detection of IgE and IgM Antibodies by Immuno-PCR. PLoS ONE 2019, 14, e0209860. [Google Scholar] [CrossRef]
- Huggins, I.J.; Medina, C.A.; Springer, A.D.; van den Berg, A.; Jadhav, S.; Cui, X.; Dowdy, S.F. Site Selective Antibody-Oligonucleotide Conjugation via Microbial Transglutaminase. Molecules 2019, 24, 3287. [Google Scholar] [CrossRef] [PubMed]
- Kazane, S.A.; Sok, D.; Cho, E.H.; Uson, M.L.; Kuhn, P.; Schultz, P.G.; Smider, V.V. Site-Specific DNA-Antibody Conjugates for Specific and Sensitive Immuno-PCR. Proc. Natl. Acad. Sci. USA 2012, 109, 3731–3736. [Google Scholar] [CrossRef]
- Nanna, A.R.; Kel’in, A.V.; Theile, C.; Pierson, J.M.; Voo, Z.X.; Garg, A.; Nair, J.K.; Maier, M.A.; Fitzgerald, K.; Rader, C. Generation and Validation of Structurally Defined Antibody–SiRNA Conjugates. Nucleic Acids Res. 2020, 48, 5281–5293. [Google Scholar] [CrossRef]
- Nanna, A.R.; Li, X.; Walseng, E.; Pedzisa, L.; Goydel, R.S.; Hymel, D.; Burke, T.R.; Roush, W.R.; Rader, C. Harnessing a Catalytic Lysine Residue for the One-Step Preparation of Homogeneous Antibody-Drug Conjugates. Nat. Commun. 2017, 8. [Google Scholar] [CrossRef]
- Matsuda, Y.; Mendelsohn, B.A. An Overview of Process Development for Antibody-Drug Conjugates Produced by Chemical Conjugation Technology. Expert Opin. Biol. Ther. 2020, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, Y.; Robles, V.; Malinao, M.C.; Song, J.; Mendelsohn, B.A. Comparison of Analytical Methods for Antibody-Drug Conjugates Produced by Chemical Site-Specific Conjugation: First-Generation AJICAP. Anal. Chem. 2019, 91, 12724–12732. [Google Scholar] [CrossRef]
- Yamada, K.; Fujii, T.; Shikida, N.; Shimbo, K. Compound Comprising Substance Having Affinity for Antibody, Cleavage Site And Reactive Group, or Salt ThereoF. Japanese Patent WO2019240287A1, 19 December 2019. [Google Scholar]
- Okahata, Y.; Kawase, M.; Niikura, K.; Ohtake, F.; Furusawa, H.; Ebara, Y. Kinetic Measurements of DNA Hybridization on an Oligonucleotide-Immobilized 27-MHz Quartz Crystal Microbalance. Anal. Chem. 1998, 70, 1288–1296. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.X.; Fang, J.Z.; Duan, W.; Wu, L.R.; Zhang, A.W.; Dalchau, N.; Yordanov, B.; Petersen, R.; Phillips, A.; Zhang, D.Y. Predicting DNA Hybridization Kinetics from Sequence. Nat. Chem. 2018, 10, 91–98. [Google Scholar] [CrossRef] [PubMed]
- Setyawati, M.I.; Kutty, R.V.; Leong, D.T. DNA Nanostructures Carrying Stoichiometrically Definable Antibodies. Small 2016, 12, 5601–5611. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.; Song, P.; Märcher, A.; Kjems, J.; Yang, C.; Gothelf, K.V. Selective Delivery of Doxorubicin to EGFR + Cancer Cells by Cetuximab–DNA Conjugates. ChemBioChem 2019, 20, 1014–1018. [Google Scholar] [CrossRef] [PubMed]
- Hsu, N.-S.; Lee, C.-C.; Kuo, W.-C.; Chang, Y.-W.; Lo, S.-Y.; Wang, A.H.J. Development of a Versatile and Modular Linker for Antibody–Drug Conjugates Based on Oligonucleotide Strand Pairing. Bioconj. Chem. 2020, 31, 1804–1811. [Google Scholar] [CrossRef]
- Dovgan, I.; Ehkirch, A.; Lehot, V.; Kuhn, I.; Koniev, O.; Kolodych, S.; Hentz, A.; Ripoll, M.; Ursuegui, S.; Nothisen, M.; et al. On the Use of DNA as a Linker in Antibody-Drug Conjugates: Synthesis, Stability and in Vitro Potency. Sci. Rep. 2020, 10, 7691. [Google Scholar] [CrossRef]
- Wiener, J.; Kokotek, D.; Rosowski, S.; Lickert, H.; Meier, M. Preparation of Single- and Double-Oligonucleotide Antibody Conjugates and Their Application for Protein Analytics. Sci. Rep. 2020, 10, 1457. [Google Scholar] [CrossRef] [PubMed]
- Uckun, F.M.; Qazi, S.; Dibirdik, I.; Myers, D.E. Rational Design of an Immunoconjugate for Selective Knock-down of Leukemia-Specific E2A–PBX1 Fusion Gene Expression in Human Pre-B Leukemia. Integr. Biol. 2013, 5, 122–132. [Google Scholar] [CrossRef] [PubMed]
- Hamblett, K.J.; Senter, P.D.; Chace, D.F.; Sun, M.M.C.; Lenox, J.; Cerveny, C.G.; Kissler, K.M.; Bernhardt, S.X.; Kopcha, A.K.; Zabinski, R.F.; et al. Effects of Drug Loading on the Antitumor Activity of a Monoclonal Antibody Drug Conjugate. Clin. Cancer Res. 2004, 10, 7063–7070. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dugal-Tessier, J.; Thirumalairajan, S.; Jain, N. Antibody-Oligonucleotide Conjugates: A Twist to Antibody-Drug Conjugates. J. Clin. Med. 2021, 10, 838. https://doi.org/10.3390/jcm10040838
Dugal-Tessier J, Thirumalairajan S, Jain N. Antibody-Oligonucleotide Conjugates: A Twist to Antibody-Drug Conjugates. Journal of Clinical Medicine. 2021; 10(4):838. https://doi.org/10.3390/jcm10040838
Chicago/Turabian StyleDugal-Tessier, Julien, Srinath Thirumalairajan, and Nareshkumar Jain. 2021. "Antibody-Oligonucleotide Conjugates: A Twist to Antibody-Drug Conjugates" Journal of Clinical Medicine 10, no. 4: 838. https://doi.org/10.3390/jcm10040838
APA StyleDugal-Tessier, J., Thirumalairajan, S., & Jain, N. (2021). Antibody-Oligonucleotide Conjugates: A Twist to Antibody-Drug Conjugates. Journal of Clinical Medicine, 10(4), 838. https://doi.org/10.3390/jcm10040838





