Intestinal Microbiota and Perspectives of the Use of Meta-Analysis for Comparison of Ulcerative Colitis Studies
Abstract
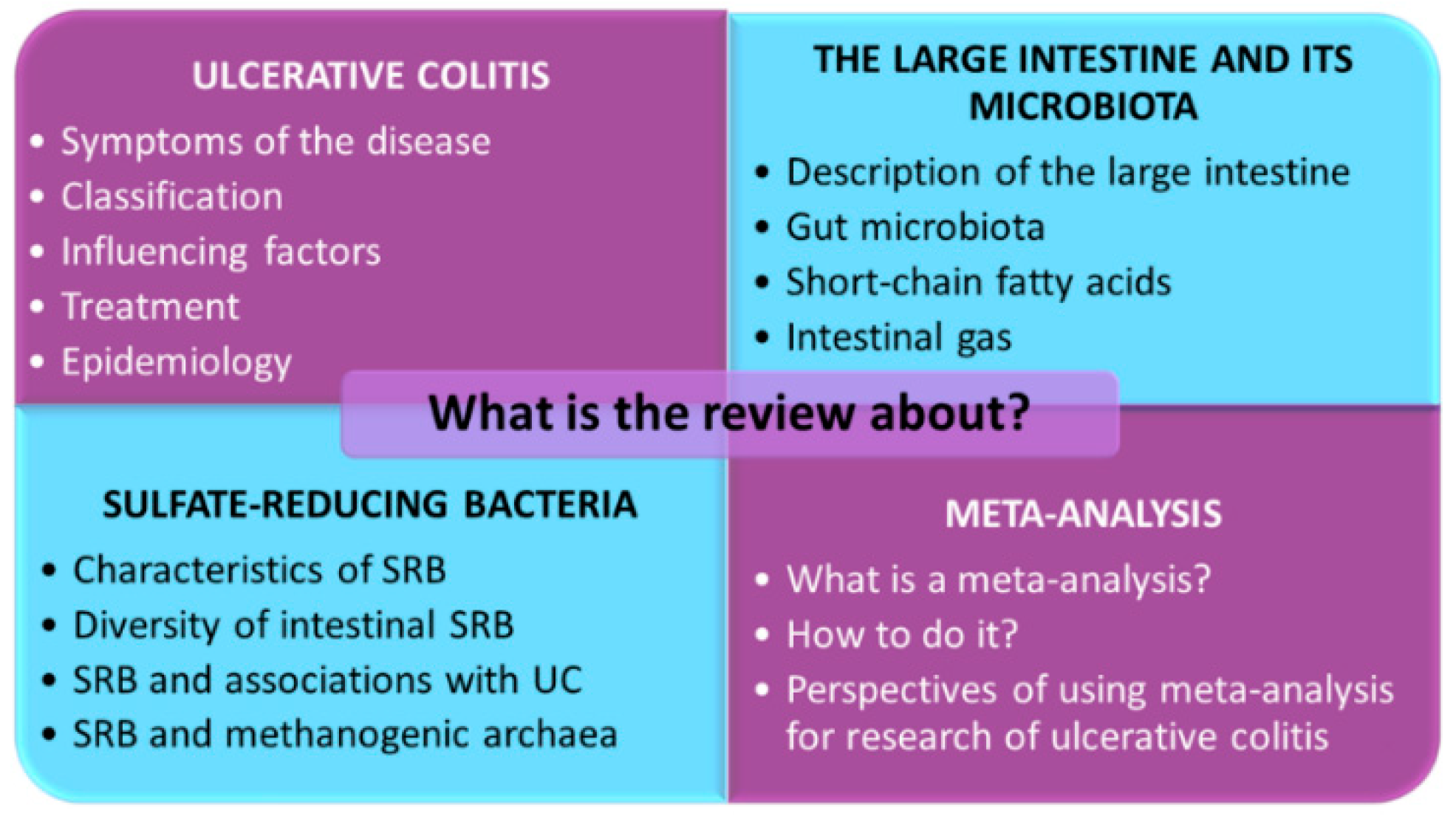
1. Introduction
2. Ulcerative Colitis
2.1. Genetic Predisposition
2.2. Environmental Factors
2.3. Microbial Effect
2.4. Immune Response
2.5. Epidemiology
3. The Large Intestine and Its Microbiota
3.1. Intestinal Microbiota
3.2. Short-Chain Fatty Acids
3.3. Intestinal Gases
4. Sulfate-Reducing Bacteria
4.1. Diversity of Intestinal Sulfate-Reducing Bacteria
4.1.1. Order Desulfovibrionales (Domain Bacteria, Phylum Proteobacteria, Class Deltaproteobacteria)
4.1.2. Family Desulfovibrionaceae, Genus Desulfovibrio
4.1.3. Family Desulfomicrobiaceae, Genus Desulfomicrobium
4.1.4. Order Desulfobacterales (Domain Bacteria, Phylum Proteobacteria, Class Deltaproteobacteria)
4.1.5. Family Desulfobacteraceae, Genus Desulfobacter
4.1.6. Family Desulfobulbaceae, Genus Desulfobulbus
4.1.7. Order Clostridiales (Domain Bacteria, Phylum Firmicutes, Class Clostridia)
4.1.8. Family Peptococcaceae, Genus Desulfotomaculum
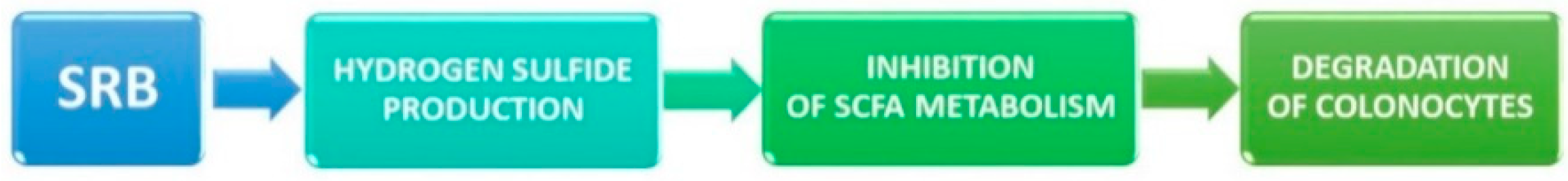
4.2. Sulfate-Reducing Bacteria and Ulcerative Colitis
4.3. Sulfate-Reducing Bacteria in Relation to Methanogenic Archaea
5. Meta-Analysis
5.1. Performing a Meta-Analysis
5.1.1. Literature Research and Data Abstraction
5.1.2. Forest Plot
6. Perspectives of Using Meta-Analysis for Ulcerative Colitis
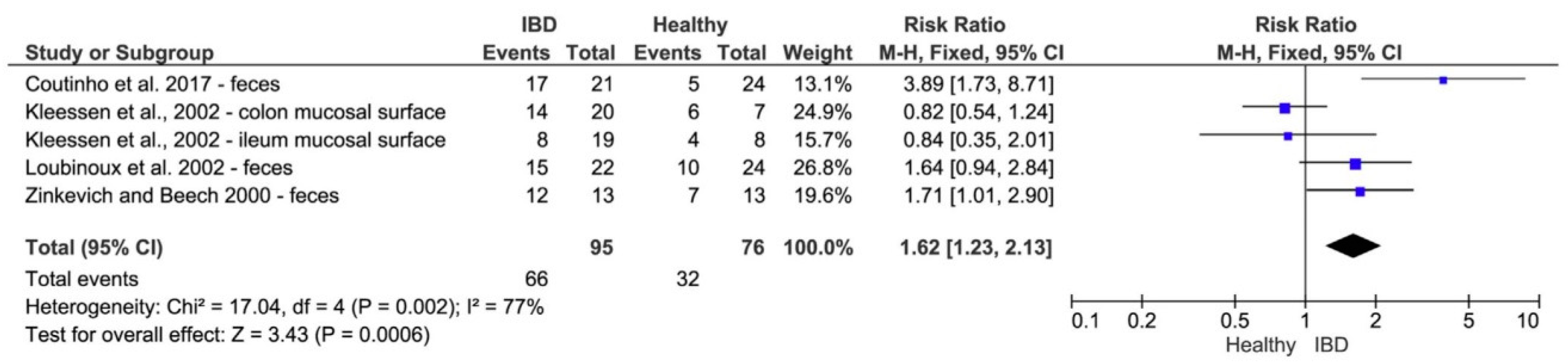
6.1. Sulfate-Reducing Bacteria and Probiotic Treatment of Ulcerative Colitis
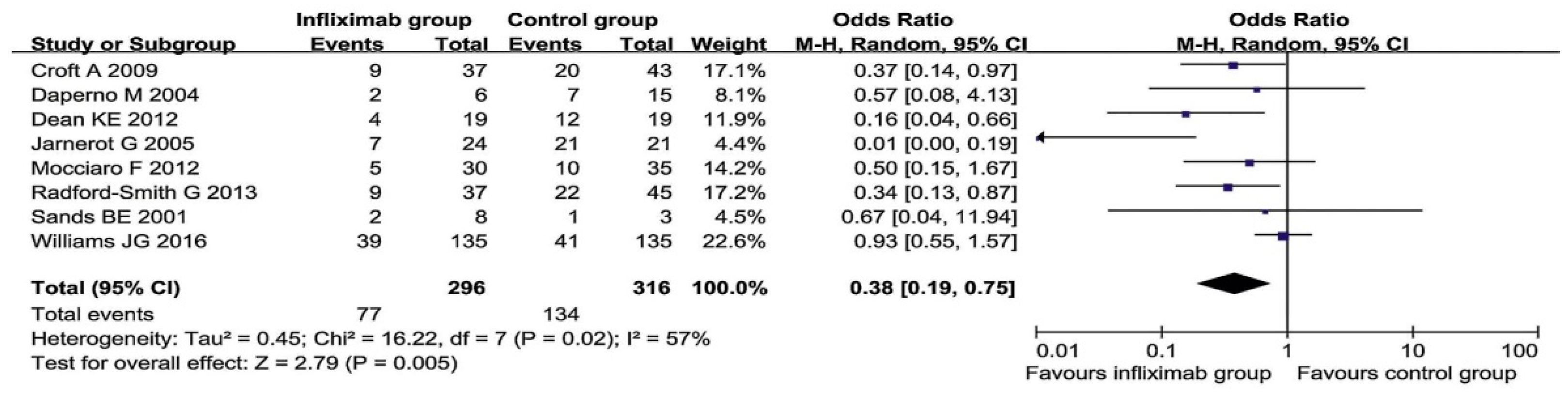
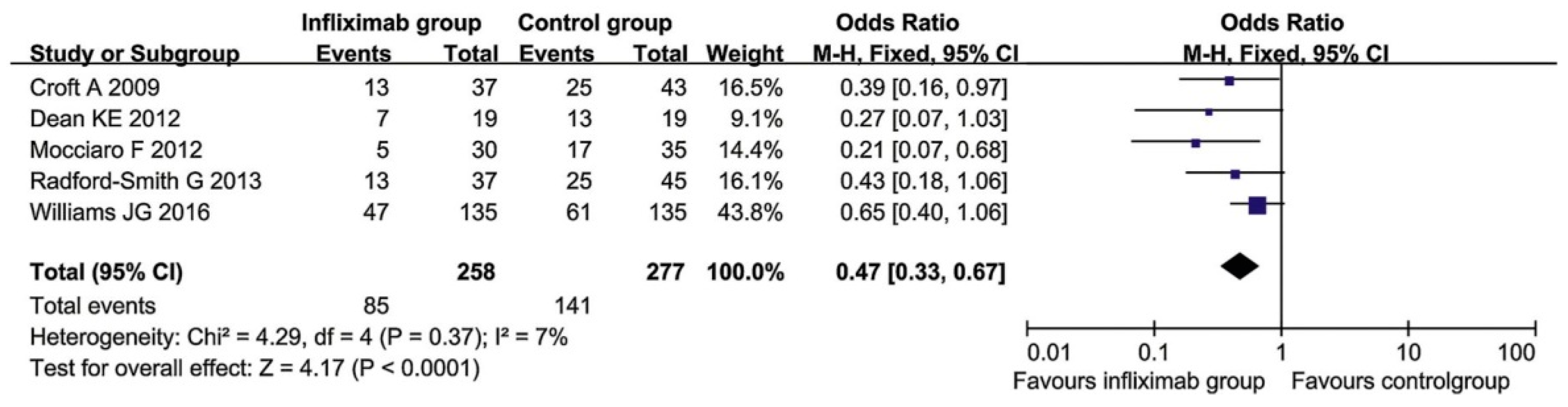
6.2. Treatment of Ulcerative Colitis with Infliximab
6.3. Clostridium difficile and Ulcerative Colitis
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Ahmed, I.; Niaz, Z. Ulcerative Colitis. In Ulcerative Colitis. Epidemiology, Pathogenesis and Complications; IntechOpen: London, UK, 2011; pp. 1–12. [Google Scholar] [CrossRef]
- Ho, G.-T.; Boyapati, R.; Satsangi, J. Ulcerative colitis. Medicine 2015, 43, 276–281. [Google Scholar] [CrossRef]
- Kushkevych, I.; Dordević, D.; Vítězová, M. Toxicity of hydrogen sulfide toward sulfate-reducing bacteria Desulfovibrio piger Vib-7. Arch. Microbiol. 2019, 201, 389–397. [Google Scholar] [CrossRef] [PubMed]
- Kushkevych, I.; Kotrsová, V.; Dordević, D.; Buňková, L.; Vítězová, M.; Amedei, A. Hydrogen Sulfide Effects on the Survival of Lactobacilli with Emphasis on the Development of Inflammatory Bowel Diseases. Biomolecules 2019, 9, 752. [Google Scholar] [CrossRef] [PubMed]
- Vilela, E.G. Evaluation of inflammatory activity in Crohn’s disease and ulcerative colitis. WJG 2012, 18, 872. [Google Scholar] [CrossRef] [PubMed]
- Gibson, G.R.; Macfarlane, S.; Macfarlane, G.T. Metabolic interactions involving sulphate-reducing and methanogenic bacteria in the human large intestine. FEMS Microbiol. Ecol. 1993, 12, 117–125. [Google Scholar] [CrossRef]
- Cummings, J.H.; Macfarlane, G.T. Colonic microflora: Nutrition and health. Nutrition 1997, 13, 476–478. [Google Scholar] [CrossRef]
- Cummings, J.H.; Macfarlane, G.T.; Macfarlane, S. Intestinal bacteria and ulcerative colitis. Curr. Issues Intestig. Microbiol. 2003, 4, 9–20. [Google Scholar]
- Malik, T.A. Inflammatory Bowel Disease. Surg. Clin. N. Am. 2015, 95, 1105–1122. [Google Scholar] [CrossRef]
- Gibson, G.R.; Cummings, J.H.; Macfarlane, G.T. Growth and activities of sulphate-reducing bacteria in gut contents of healthy subjects and patients with ulcerative colitis. FEMS Microbiol. Lett. 1991, 86, 103–112. [Google Scholar] [CrossRef]
- Kushkevych, I.V. Kinetic Properties of Pyruvate Ferredoxin Oxidoreductase of Intestinal Sulfate-Reducing Bacteria Desulfovibrio piger Vib-7 and Desulfomicrobium sp. Rod-9. Pol. J. Microbiol. 2015, 64, 107–114. [Google Scholar] [CrossRef]
- Kushkevych, I.V. Activity and kinetic properties of phosphotransacetylase from intestinal sulfate-reducing bacteria. Acta Biochim. Pol. 2015, 62, 103–108. [Google Scholar] [CrossRef] [PubMed]
- Kushkevych, I. Isolation and Purification of Sulfate-Reducing Bacteria. In Microorganisms; Blumenberg, M., Shaaban, M., Elgaml, A., Eds.; IntechOpen: London, UK, 2020; ISBN 978-1-83880-187-8. [Google Scholar]
- Kushkevych, I.; Kollar, P.; Suchy, P.; Parak, T.; Pauk, K.; Imramovsky, A. Activity of selected salicylamides against intestinal sulfate-reducing bacteria. Neuro Endocrinol. Lett. 2015, 36 (Suppl. 1), 106–113. [Google Scholar] [PubMed]
- Kushkevych, I.; Fafula, R.; Parák, T.; Bartoš, M. Activity of Na+/K+-activated Mg2+-dependent ATP-hydrolase in the cell-free extracts of the sulfate-reducing bacteria Desulfovibrio piger Vib-7 and Desulfomicrobium sp. Rod-9. Acta Vet. Brno 2015, 84, 3–12. [Google Scholar] [CrossRef]
- Kushkevych, I.; Kováč, J.; Vítězová, M.; Vítěz, T.; Bartoš, M. The diversity of sulfate-reducing bacteria in the seven bioreactors. Arch. Microbiol. 2018, 200, 945–950. [Google Scholar] [CrossRef] [PubMed]
- Pitcher, M.C.; Cummings, J.H. Hydrogen sulphide: A bacterial toxin in ulcerative colitis? Gut 1996, 39, 1–4. [Google Scholar] [CrossRef]
- Yuan, Y.; Hunt, R.H. Systematic Reviews: The Good, the Bad and the Ugly. Am. J. Gastroenterol. 2009, 104, 1086–1092. [Google Scholar] [CrossRef]
- Ergal, İ.; Fuchs, W.; Hasibar, B.; Thallinger, B.; Bochmann, G.; Rittmann, S.K.-M.R. The physiology and biotechnology of dark fermentative biohydrogen production. Biotechnol. Adv. 2018, 36, 2165–2186. [Google Scholar] [CrossRef]
- Rittmann, S.K.-M.R.; Seifert, A.H.; Bernacchi, S. Kinetics, multivariate statistical modelling, and physiology of CO2-based biological methane production. Appl. Energy 2018, 216, 751–760. [Google Scholar] [CrossRef]
- Dordević, D.; Jančíková, S.; Vítězová, M.; Kushkevych, I. Hydrogen sulfide toxicity in the gut environment: Meta-analysis of sulfate-reducing and lactic acid bacteria in inflammatory processes. J. Adv. Res. 2020, 27, 55–69. [Google Scholar] [CrossRef]
- Kushkevych, I.; Dordević, D.; Kollár, P. Analysis of physiological parameters of Desulfovibrio strains from individuals with colitis. Open Life Sci. 2019, 13, 481–488. [Google Scholar] [CrossRef]
- Kováč, J.; Vítězová, M.; Kushkevych, I. Metabolic activity of sulfate-reducing bacteria from rodents with colitis. Open Med. 2018, 13, 344–349. [Google Scholar] [CrossRef] [PubMed]
- Kushkevych, I.; Vítězová, M.; Kos, J.; Kollár, P.; Jampílek, J. Effect of selected 8-hydroxyquinoline-2-carboxanilides on viability and sulfate metabolism of Desulfovibrio piger. J. Appl. Biomed. 2018, 16, 241–246. [Google Scholar] [CrossRef]
- Kushkevych, I.; Dordević, D.; Vítězová, M. Analysis of pH dose-dependent growth of sulfate-reducing bacteria. Open Med. 2019, 14, 66–74. [Google Scholar] [CrossRef]
- Kushkevych, I.; Castro Sangrador, J.; Dordević, D.; Rozehnalová, M.; Černý, M.; Fafula, R.; Vítězová, M.; Rittmann, S.K.-M.R. Evaluation of Physiological Parameters of Intestinal Sulfate-Reducing Bacteria Isolated from Patients Suffering from IBD and Healthy People. JCM 2020, 9, 1920. [Google Scholar] [CrossRef] [PubMed]
- Kushkevych, I.; Dordević, D.; Vítězová, M. Possible synergy effect of hydrogen sulfide and acetate produced by sulfate-reducing bacteria on inflammatory bowel disease development. J. Adv. Res. 2020, 27, 71–78. [Google Scholar] [CrossRef]
- Stange, E.F. Inflammatory bowel diseases. Preface. Dig. Dis. 2013, 31, 269. [Google Scholar] [CrossRef]
- Boyapati, R.K.; Rossi, A.G.; Satsangi, J.; Ho, G.-T. Gut mucosal DAMPs in IBD: From mechanisms to therapeutic implications. Mucosal. Immunol. 2016, 9, 567–582. [Google Scholar] [CrossRef]
- Head, K.; Jurenka, J.S. Inflammatory bowel disease. Part II: Crohn’s disease--pathophysiology and conventional and alternative treatment options. Altern. Med. Rev. 2004, 9, 360–401. [Google Scholar]
- Ek, W.E.; D’Amato, M.; Halfvarson, J. The history of genetics in inflammatory bowel disease. Ann. Gastroenterol. 2014, 27, 294–303. [Google Scholar]
- Boirivant, M.; Cossu, A. Inflammatory bowel disease: Inflammatory bowel disease. Oral Dis. 2012, 18, 1–15. [Google Scholar] [CrossRef]
- Juyal, G.; Sood, A.; Midha, V.; Thelma, B.K. Genetics of ulcerative colitis: Putting into perspective the incremental gains from Indian studies. J. Genet. 2018, 97, 1493–1507. [Google Scholar] [CrossRef] [PubMed]
- Molodecky, N.A.; Kaplan, G.G. Environmental risk factors for inflammatory bowel disease. Gastroenterol. Hepatol. (N. Y.) 2010, 6, 339–346. [Google Scholar]
- Koloski, N.-A.; Bret, L.; Radford-Smith, G. Hygiene hypothesis in inflammatory bowel disease: A critical review of the literature. World J. Gastroenterol. 2008, 14, 165–173. [Google Scholar] [CrossRef] [PubMed]
- Sonnenberg, A. Occupational distribution of inflammatory bowel disease among German employees. Gut 1990, 31, 1037–1040. [Google Scholar] [CrossRef]
- Macfarlane, M.J.; Hopkins, G.T.; Ma, S. Bacterial Growth and Metabolism on Surfaces in the Large Intestine. Microb. Ecol. Health Dis. 2000, 12, 64–72. [Google Scholar] [CrossRef]
- Macfarlane, S.; Steed, H.; Macfarlane, G.T. Intestinal bacteria and inflammatory bowel disease. Crit. Rev. Clin. Lab. Sci. 2009, 46, 25–54. [Google Scholar] [CrossRef]
- Savage, D.C. Microbial Ecology of the Gastrointestinal Tract. Annu. Rev. Microbiol. 1977, 31, 107–133. [Google Scholar] [CrossRef]
- Carbonnel, F.; Jantchou, P.; Monnet, E.; Cosnes, J. Environmental risk factors in Crohn’s disease and ulcerative colitis: An update. Gastroenterol. Clin. ET Biol. 2009, 33, S145–S157. [Google Scholar] [CrossRef]
- Campieri, M. Bacteria as the cause of ulcerative colitis. Gut 2001, 48, 132–135. [Google Scholar] [CrossRef]
- Sands, B.E. Inflammatory bowel disease: Past, present, and future. J. Gastroenterol. 2007, 42, 16–25. [Google Scholar] [CrossRef]
- Clavel, T.; Haller, D. Bacteria- and host-derived mechanisms to control intestinal epithelial cell homeostasis: Implications for chronic inflammation: Inflamm. Bowel Dis. 2007, 13, 1153–1164. [Google Scholar] [CrossRef] [PubMed]
- Ng, S.C.; Shi, H.Y.; Hamidi, N.; Underwood, F.; Tang, W.; Benchimol, E.; Panaccione, R.; Ghosh, S.; Wu, J.; Chan, F.; et al. Evolving Trends in the Epidemiology of IBD in the 21st Century: A Systematic Review of Population-Based Studies: 594. Am. J. Gastroenterol. 2017, 112, S319–S320. [Google Scholar] [CrossRef] [PubMed]
- Sánchez de Medina, F.; Romero-Calvo, I.; Mascaraque, C.; Martínez-Augustin, O. Intestinal Inflammation and Mucosal Barrier Function: Inflamm. Bowel Dis. 2014, 20, 2394–2404. [Google Scholar] [CrossRef] [PubMed]
- Guarner, F.; Malagelada, J.-R. Role of bacteria in experimental colitis. Best Pract. Res. Clin. Gastroenterol. 2003, 17, 793–804. [Google Scholar] [CrossRef]
- Li, X.; Bi, Y. How Many Human and Bacteria Cells Are in the Human Body? Infect. Dis. Transl. Med. 2017, 1–2. [Google Scholar] [CrossRef]
- Pushpanathan, P.; Mathew, G.; Selvarajan, S.; Seshadri, K.; Srikanth, P. Gut microbiota and its mysteries. Indian J. Med. Microbiol. 2019, 37, 268. [Google Scholar] [CrossRef]
- Ziemer, C.J. Newly Cultured Bacteria with Broad Diversity Isolated from Eight-Week Continuous Culture Enrichments of Cow Feces on Complex Polysaccharides. Appl. Environ. Microbiol. 2014, 80, 574–585. [Google Scholar] [CrossRef]
- Schroeder, B.O. Fight them or feed them: How the intestinal mucus layer manages the gut microbiota. Gastroenterol. Rep. 2019, 7, 3–12. [Google Scholar] [CrossRef]
- Johansson, M.E.V.; Gustafsson, J.K.; Holmén-Larsson, J.; Jabbar, K.S.; Xia, L.; Xu, H.; Ghishan, F.K.; Carvalho, F.A.; Gewirtz, A.T.; Sjövall, H.; et al. Bacteria penetrate the normally impenetrable inner colon mucus layer in both murine colitis models and patients with ulcerative colitis. Gut 2014, 63, 281–291. [Google Scholar] [CrossRef]
- Cummings, J.H. Short chain fatty acids in the human colon. Gut 1981, 22, 763–779. [Google Scholar] [CrossRef]
- Rowland, I.; Gibson, G.; Heinken, A.; Scott, K.; Swann, J.; Thiele, I.; Tuohy, K. Gut microbiota functions: Metabolism of nutrients and other food components. Eur. J. Nutr. 2018, 57, 1–24. [Google Scholar] [CrossRef]
- Couto, M.R.; Gonçalves, P.; Magro, F.; Martel, F. Microbiota-derived butyrate regulates intestinal inflammation: Focus on inflammatory bowel disease. Pharmacol. Res. 2020, 159, 104947. [Google Scholar] [CrossRef] [PubMed]
- Mauerhofer, L.-M.; Pappenreiter, P.; Paulik, C.; Seifert, A.H.; Bernacchi, S.; Rittmann, S.K.-M.R. Methods for quantification of growth and productivity in anaerobic microbiology and biotechnology. Folia Microbiol. 2019, 64, 321–360. [Google Scholar] [CrossRef] [PubMed]
- Roy, C.C.; Kien, C.L.; Bouthillier, L.; Levy, E. Short-Chain Fatty Acids: Ready for Prime Time? Nutr. Clin. Pract. 2006, 21, 351–366. [Google Scholar] [CrossRef] [PubMed]
- Treem, W.R.; Ahsan, N.; Shoup, M.; Hyams, J.S. Fecal short-chain fatty acids in children with inflammatory bowel disease. J. Pediatr. Gastroenterol. Nutr. 1994, 18, 159–164. [Google Scholar] [CrossRef]
- Hill, M.J. Intestinal flora and endogenous vitamin synthesis. Eur. J. Cancer Prev. 1997, 6, S43–S45. [Google Scholar] [CrossRef] [PubMed]
- Martens, H.; Barg, M.; Warren, D.; Jah, J.-H. Microbial production of vitamin B 12. Appl. Microbiol. Biotechnol. 2002, 58, 275–285. [Google Scholar] [CrossRef]
- Tremaroli, V.; Bäckhed, F. Functional interactions between the gut microbiota and host metabolism. Nature 2012, 489, 242–249. [Google Scholar] [CrossRef]
- Duncan, S.H.; Holtrop, G.; Lobley, G.E.; Calder, A.G.; Stewart, C.S.; Flint, H.J. Contribution of acetate to butyrate formation by human faecal bacteria. Br. J. Nutr. 2004, 91, 915–923. [Google Scholar] [CrossRef]
- Gonçalves, P.; Martel, F. Butyrate and Colorectal Cancer: The Role of Butyrate Transport. CDM 2013, 14, 994–1008. [Google Scholar] [CrossRef]
- LeBlanc, J.G.; Chain, F.; Martín, R.; Bermúdez-Humarán, L.G.; Courau, S.; Langella, P. Beneficial effects on host energy metabolism of short-chain fatty acids and vitamins produced by commensal and probiotic bacteria. Microb. Cell Fact. 2017, 16, 79. [Google Scholar] [CrossRef] [PubMed]
- Oliphant, K.; Allen-Vercoe, E. Macronutrient metabolism by the human gut microbiome: Major fermentation by-products and their impact on host health. Microbiome 2019, 7, 91. [Google Scholar] [CrossRef] [PubMed]
- Clausen, M.R.; Mortensen, P.B. Kinetic studies on colonocyte metabolism of short chain fatty acids and glucose in ulcerative colitis. Gut 1995, 37, 684–689. [Google Scholar] [CrossRef] [PubMed]
- Moeinian, M. Beneficial effect of butyrate, Lactobacillus casei and L-carnitine combination in preference to each in experimental colitis. WJG 2014, 20, 10876. [Google Scholar] [CrossRef] [PubMed]
- den Besten, G.; Lange, K.; Havinga, R.; van Dijk, T.H.; Gerding, A.; van Eunen, K.; Müller, M.; Groen, A.K.; Hooiveld, G.J.; Bakker, B.M.; et al. Gut-derived short-chain fatty acids are vividly assimilated into host carbohydrates and lipids. Am. J. Physiol. Gastrointest. Liver Physiol. 2013, 305, G900–G910. [Google Scholar] [CrossRef]
- Al-Asmakh, M.; Anuar, F.; Zadjali, F.; Rafter, J.; Pettersson, S. Gut microbial communities modulating brain development and function. Gut Microbes 2012, 3, 366–373. [Google Scholar] [CrossRef]
- De Vadder, F.; Kovatcheva-Datchary, P.; Goncalves, D.; Vinera, J.; Zitoun, C.; Duchampt, A.; Bäckhed, F.; Mithieux, G. Microbiota-Generated Metabolites Promote Metabolic Benefits via Gut-Brain Neural Circuits. Cell 2014, 156, 84–96. [Google Scholar] [CrossRef]
- Abdel Azim, A.; Rittmann, S.K.-M.R.; Fino, D.; Bochmann, G. The physiological effect of heavy metals and volatile fatty acids on Methanococcus maripaludis S2. Biotechnol. Biofuels 2018, 11, 301. [Google Scholar] [CrossRef]
- Frost, G.; Sleeth, M.L.; Sahuri-Arisoylu, M.; Lizarbe, B.; Cerdan, S.; Brody, L.; Anastasovska, J.; Ghourab, S.; Hankir, M.; Zhang, S.; et al. The short-chain fatty acid acetate reduces appetite via a central homeostatic mechanism. Nat. Commun. 2014, 5, 3611. [Google Scholar] [CrossRef]
- Louis, P.; Flint, H.J. Formation of propionate and butyrate by the human colonic microbiota. Environ. Microbiol. 2017, 19, 29–41. [Google Scholar] [CrossRef]
- Levitt, M.D.; Bond, J.H. Volume, composition, and source of intestinal gas. Gastroenterology 1970, 59, 921–929. [Google Scholar] [CrossRef]
- Figliuolo, V.R.; Coutinho-Silva, R.; Coutinho, C.M.L.M. Contribution of sulfate-reducing bacteria to homeostasis disruption during intestinal inflammation. Life Sci. 2018, 215, 145–151. [Google Scholar] [CrossRef] [PubMed]
- Christl, S.U.; Murgatroyd, P.R.; Gibson, G.R.; Cummings, J.H. Production, metabolism, and excretion of hydrogen in the large intestine. Gastroenterology 1992, 102, 1269–1277. [Google Scholar] [CrossRef]
- Wolf, P.G.; Biswas, A.; Morales, S.E.; Greening, C.; Gaskins, H.R. H2 metabolism is widespread and diverse among human colonic microbes. Gut Microbes 2016, 7, 235–245. [Google Scholar] [CrossRef]
- Florin, T.H.J.; Neale, G.; Goretski, S.; Cummings, J.H. The Sulfate Content of Foods and Beverages. J. Food Compos. Anal. 1993, 6, 140–151. [Google Scholar] [CrossRef]
- Suarez, F.; Furne, J.; Springfield, J.; Levitt, M. Insights into human colonic physiology obtained from the study of flatus composition. Am. J. Physiol. 1997, 272, G1028–G1033. [Google Scholar] [CrossRef]
- Postgate, J. The Suphate-Reducing Bacteria, 2nd ed.; Cambridge University: Cambridge, UK, 1984; Volume 1984. [Google Scholar]
- Kushkevych, I.V. Dissimilatory sulfate reduction in the intestinal sulfate-reducing bacteria. Biol. Stud. 2016, 10, 197–228. [Google Scholar] [CrossRef]
- Kushkevych, I.; Vítězová, M.; Vítěz, T.; Kováč, J.; Kaucká, P.; Jesionek, W.; Bartoš, M.; Barton, L. A new combination of substrates: Biogas production and diversity of the methanogenic microorganisms. Open Life Sci. 2018, 13, 119–128. [Google Scholar] [CrossRef]
- Kushkevych, I.; Leščanová, O.; Dordević, D.; Jančíková, S.; Hošek, J.; Vítězová, M.; Buňková, L.; Drago, L. The Sulfate-Reducing Microbial Communities and Meta-Analysis of Their Occurrence during Diseases of Small–Large Intestine Axis. JCM 2019, 8, 1656. [Google Scholar] [CrossRef]
- Černý, M.; Vítězová, M.; Vítěz, T.; Bartoš, M.; Kushkevych, I. Variation in the Distribution of Hydrogen Producers from the Clostridiales Order in Biogas Reactors Depending on Different Input Substrates. Energies 2018, 11, 3270. [Google Scholar] [CrossRef]
- Kovac, J.; Kushkevych, I. New modification of cultivation medium for isolation and growth of intestinal sulfate-reducing bacteria. In Proceedings of the MendelNet 2017, Faculty of AgriSciences, Mendel University in Brno, Brno, Czech Republic, 8–9 November 2017; Volume 2017, pp. 702–707. [Google Scholar]
- Abdulina, D.; Kováč, J.; Iutynska, G.; Kushkevych, I. ATP sulfurylase activity of sulfate-reducing bacteria from various ecotopes. 3 Biotech 2020, 10, 55. [Google Scholar] [CrossRef] [PubMed]
- Kushkevych, I.; Dordević, D.; Kollar, P.; Vítězová, M.; Drago, L. Hydrogen Sulfide as a Toxic Product in the Small–Large Intestine Axis and its Role in IBD Development. JCM 2019, 8, 1054. [Google Scholar] [CrossRef] [PubMed]
- Loubinoux, J.; Valente, F.M.A.; Pereira, I.A.C.; Costa, A.; Grimont, P.A.D.; Le Faou, A.E. Reclassification of the only species of the genus Desulfomonas, Desulfomonas pigra, as Desulfovibrio piger comb. nov. Int. J. Syst. Evol. Microbiol. 2002, 52, 1305–1308. [Google Scholar] [CrossRef] [PubMed]
- Kushkevych, I.; Dordević, D.; Vítězová, M.; Kollár, P. Cross-correlation analysis of the Desulfovibrio growth parameters of intestinal species isolated from people with colitis. Biologia 2018, 73, 1137–1143. [Google Scholar] [CrossRef]
- Kushkevych, I.; Coufalová, M.; Vítězová, M.; Rittmann, S.K.-M.R. Sulfate-Reducing Bacteria of the Oral Cavity and Their Relation with Periodontitis—Recent Advances. JCM 2020, 9, 2347. [Google Scholar] [CrossRef]
- Barton, L.L.; Fardeau, M.-L.; Fauque, G.D. Hydrogen sulfide: A toxic gas produced by dissimilatory sulfate and sulfur reduction and consumed by microbial oxidation. Met. Ions Life Sci. 2014, 14, 237–277. [Google Scholar] [CrossRef]
- Brenner, D.J.; Krieg, N.R.; Staley, J.T.; Garrity, G.M. The Proteobacteria, Part C: The Alpha-, Beta-, Delta-, and Epsilonproteobacteria. In Bergey’s Manual of Systematic Bacteriology; Springer: Boston, MA, USA, 2010; Volume 2005, p. 1388. [Google Scholar]
- Glass, G.V. Primary, Secondary, and Meta-Analysis of Research. Educ. Res. 1976, 5, 3–8. [Google Scholar] [CrossRef]
- Liberati, A.; Altman, D.G.; Tetzlaff, J.; Mulrow, C.; Gøtzsche, P.C.; Ioannidis, J.P.A.; Clarke, M.; Devereaux, P.J.; Kleijnen, J.; Moher, D. The PRISMA statement for reporting systematic reviews and meta-analyses of studies that evaluate healthcare interventions: Explanation and elaboration. BMJ 2009, 339, b2700. [Google Scholar] [CrossRef]
- Pabalan, N.; Jarjanazi, H.; Steiner, T.S. Meta-analysis in microbiology. Indian J. Med. Microbiol. 2014, 32, 229–235. [Google Scholar] [CrossRef]
- Haidich, A.B. Meta-analysis in medical research. Hippokratia 2010, 14, 29–37. [Google Scholar]
- Goodman, C.S. Introduction to Health Technology Assessment; National Library of Medicine: Rockville, MD, USA, 2014. [Google Scholar]
- Kushkevych, I.; Vítězová, M.; Fedrová, P.; Vochyanová, Z.; Paráková, L.; Hošek, J. Kinetic properties of growth of intestinal sulphate-reducing bacteria isolated from healthy mice and mice with ulcerative colitis. Acta Vet. Brno 2017, 86, 405–411. [Google Scholar] [CrossRef]
- Ishikawa, H.; Akedo, I.; Umesaki, Y.; Tanaka, R.; Imaoka, A.; Otani, T. Randomized Controlled Trial of the Effect of Bifidobacteria-Fermented Milk on Ulcerative Colitis. J. Am. Coll. Nutr. 2003, 22, 56–63. [Google Scholar] [CrossRef] [PubMed]
- Miele, E.; Pascarella, F.; Giannetti, E.; Quaglietta, L.; Baldassano, R.N.; Staiano, A. Effect of a Probiotic Preparation (VSL#3) on Induction and Maintenance of Remission in Children with Ulcerative Colitis. Am. J. Gastroenterol. 2009, 104, 437–443. [Google Scholar] [CrossRef]
- Sood, A.; Midha, V.; Makharia, G.K.; Ahuja, V.; Singal, D.; Goswami, P.; Tandon, R.K. The Probiotic Preparation, VSL#3 Induces Remission in Patients with Mild-to-Moderately Active Ulcerative Colitis. Clin. Gastroenterol. Hepatol. 2009, 7, 1202–1209.e1. [Google Scholar] [CrossRef] [PubMed]
- Rembacken, B.; Snelling, A.; Hawkey, P.; Chalmers, D.; Axon, A. Non-pathogenic Escherichia coli versus mesalazine for the treatment of ulcerative colitis: A randomised trial. Lancet 1999, 354, 635–639. [Google Scholar] [CrossRef]
- Fratila, O.C.; Craciun, C. Ultrastructural evidence of mucosal healing after infliximab in patients with ulcerative colitis. J. Gastrointest. Liver Dis. 2010, 19, 147–153. [Google Scholar]
- Ungar, B.; Mazor, Y.; Weisshof, R.; Yanai, H.; Ron, Y.; Goren, I.; Waizbard, A.; Yavzori, M.; Fudim, E.; Picard, O.; et al. Induction infliximab levels among patients with acute severe ulcerative colitis compared with patients with moderately severe ulcerative colitis. Aliment. Pharm. 2016, 43, 1293–1299. [Google Scholar] [CrossRef]
- Guo, C.; Wu, K.; Liang, X.; Liang, Y.; Li, R. Infliximab clinically treating ulcerative colitis: A systematic review and meta-analysis. Pharmacol. Res. 2019, 148, 104455. [Google Scholar] [CrossRef]
- Dinesen, L.C.; Walsh, A.J.; Protic, M.N.; Heap, G.; Cummings, F.; Warren, B.F.; George, B.; Mortensen, N.J.M.; Travis, S.P.L. The pattern and outcome of acute severe colitis. J. Crohns Colitis 2010, 4, 431–437. [Google Scholar] [CrossRef]
- Roberts, S.E.; Williams, J.G.; Yeates, D.; Goldacre, M.J. Mortality in patients with and without colectomy admitted to hospital for ulcerative colitis and Crohn’s disease: Record linkage studies. BMJ 2007, 335, 1033. [Google Scholar] [CrossRef]
- Ananthakrishnan, A.N.; McGinley, E.L.; Saeian, K.; Binion, D.G. Temporal trends in disease outcomes related to Clostridium difficile infection in patients with inflammatory bowel disease: Inflamm. Bowel Dis. 2011, 17, 976–983. [Google Scholar] [CrossRef] [PubMed]
- Ran, Z.-H.; Shen, J.; Zhu, Q.; Peng, J.-C. The impact of Clostridum difficile on surgical rate among ulcerative colitis patients: A systemic review and meta-analysis. Saudi J. Gastroenterol. 2015, 21, 208. [Google Scholar] [CrossRef] [PubMed]






| Domain/Phylum | Examples of Genera | Main Function |
|---|---|---|
| Phylum: Bakteroidetes | Bacteroidetes, Prevotella, Xylanibacter | Degradation complex glycans |
| Phylum: Firmicutes | Ruminococcus, Clostridium, Lactobacillus, Roseburia, Eubacterium, Faecalibacterium | Probiotics, butyrate producers |
| Phylum: Actinobacteria | Collinsella, Bifidobacterium | Probiotics |
| Phylum: Proteobacteria | Desulfovibrio | Sulfate-reducing bacteria |
| Phylum: Verrucomicrobia | Akkermansia | Degradation of mucin |
| Domain: Archaea | Methanobrevibacter | Methanogenesis |
| Archaea | Archaeoglobus, Caldivirga, Vulcanisaeta | |
|---|---|---|
| Bacteria | Ammonifex | Desulfomicrobium |
| Candidatus | Desulfonatronovibrio | |
| Desulfacinum | Desulfosarcina | |
| Desulfobacter | Desulfosporosinus | |
| Desulfobacterium | Desulfotomaculum | |
| Desulfobulbus | Desulfovibrio | |
| Desulfocapsa | Desulfovirga | |
| Desulfococcus | Syntrophobacter | |
| Desulfocurvus | Thermodesulfatator | |
| Desulfofustis | Thermodesulfobacterium | |
| Desulfohalobium | Thermodesulfobium | |
| Desulfoluna | Thermodesulfovibrio | |
| Scientific Discipline | Examples of Potential Use |
|---|---|
| Medicine and pharmacology | Effectiveness of treatment: infliximab and others medication, probiotics, colectomy |
| Epidemiology and statistics | Distribution in different parts of the world, age representation of patients in the population, increase or decrease in the number of patients over time |
| Genetics | Identification of risk genes |
| Immunology | Etiology of the leaking intestine, etiology of inflammatory reaction |
| Study of environmental factors | Smoking, stress, diet, influence of other health indications |
| Microbiology | Relation of microorganisms to UC: sulfate reducing bacteria, methanogenic archaea, lactic acid bacteria, Clostridium difficile |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kushkevych, I.; Martínková, K.; Vítězová, M.; Rittmann, S.K.-M.R. Intestinal Microbiota and Perspectives of the Use of Meta-Analysis for Comparison of Ulcerative Colitis Studies. J. Clin. Med. 2021, 10, 462. https://doi.org/10.3390/jcm10030462
Kushkevych I, Martínková K, Vítězová M, Rittmann SK-MR. Intestinal Microbiota and Perspectives of the Use of Meta-Analysis for Comparison of Ulcerative Colitis Studies. Journal of Clinical Medicine. 2021; 10(3):462. https://doi.org/10.3390/jcm10030462
Chicago/Turabian StyleKushkevych, Ivan, Kristýna Martínková, Monika Vítězová, and Simon K.-M. R. Rittmann. 2021. "Intestinal Microbiota and Perspectives of the Use of Meta-Analysis for Comparison of Ulcerative Colitis Studies" Journal of Clinical Medicine 10, no. 3: 462. https://doi.org/10.3390/jcm10030462
APA StyleKushkevych, I., Martínková, K., Vítězová, M., & Rittmann, S. K.-M. R. (2021). Intestinal Microbiota and Perspectives of the Use of Meta-Analysis for Comparison of Ulcerative Colitis Studies. Journal of Clinical Medicine, 10(3), 462. https://doi.org/10.3390/jcm10030462







