Clinical Relevance of a 16-Gene Pharmacogenetic Panel Test for Medication Management in a Cohort of 135 Patients
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Design and Ethical Approval
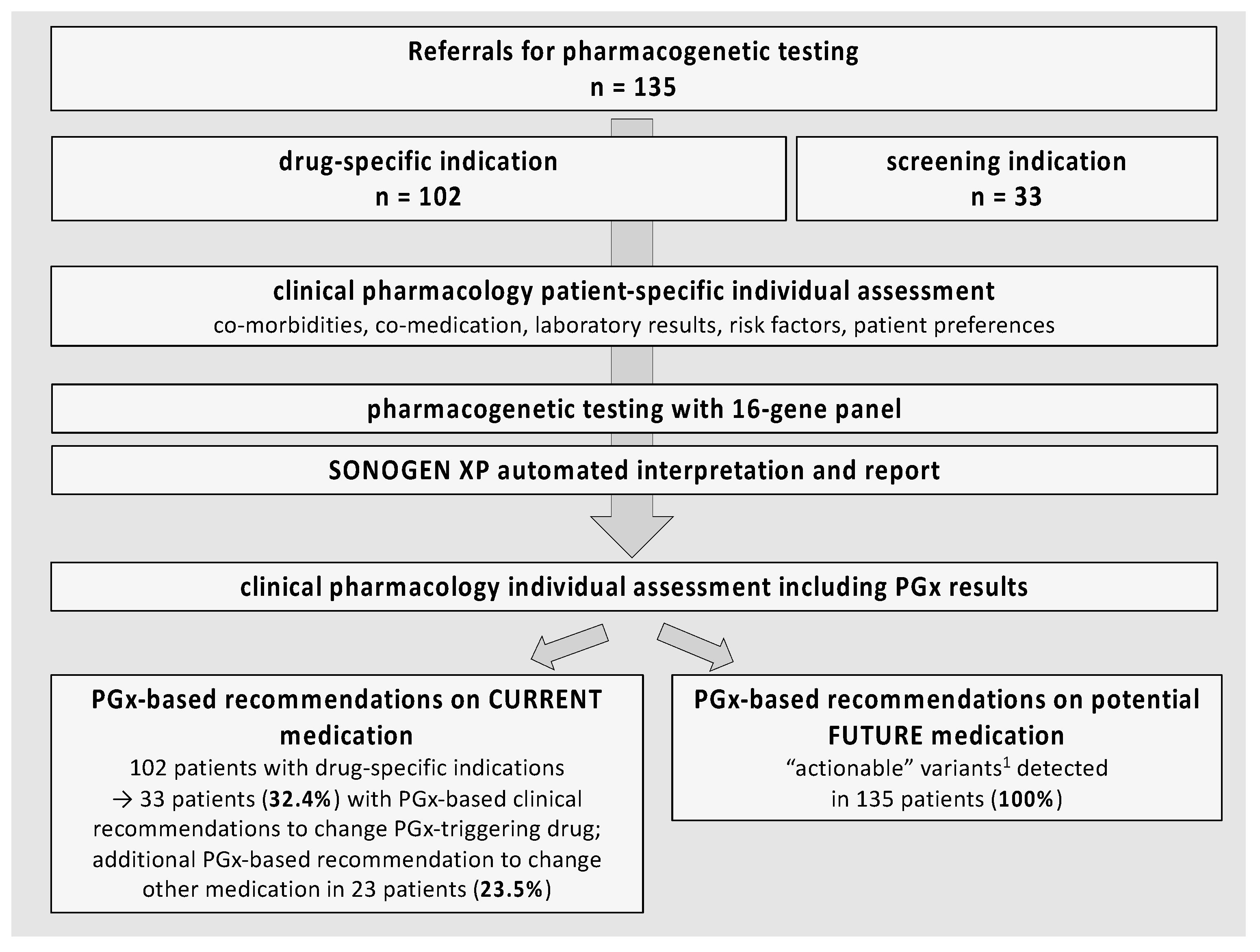
2.2. Study Population and Procedures
2.3. Genetic Analysis
2.4. PGx Expert System
2.5. Retrospective Documentation and Validation
2.6. Data Analysis
3. Results
3.1. Characteristics of the Study Population
3.2. Pharmacogenetic Variants and Their Clinical Relevance for Current Medication
3.3. Pharmacogenetic Variants and Their Clinical Relevance for Potential Future Medication
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Alshabeeb, M.A.; Deneer, V.H.M.; Khan, A.; Asselbergs, F.W. Use of Pharmacogenetic Drugs by the Dutch Population. Front. Genet. 2019, 10, 567. [Google Scholar] [CrossRef] [PubMed]
- Whirl-Carrillo, M.; McDonagh, E.M.; Hebert, J.M.; Gong, L.; Sangkuhl, K.; Thorn, C.F.; Altman, R.B.; Klein, T.E. Pharmacogenomics Knowledge for Personalized Medicine. Clin. Pharmacol. Ther. 2012, 92, 414–417. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.-J.; Chen, C.-B.; Hung, S.-I.; Ji, C.; Chung, W.-H. Pharmacogenetic Testing for Prevention of Severe Cutaneous Adverse Drug Reactions. Front. Pharmacol. 2020, 11, 969. [Google Scholar] [CrossRef]
- Relling, M.V.; Schwab, M.; Whirl-Carrillo, M.; Suarez-Kurtz, G.; Pui, C.; Stein, C.M.; Moyer, A.M.; Evans, W.E.; Klein, T.E.; Antillon-Klussmann, F.G.; et al. Clinical Pharmacogenetics Implementation Consortium Guideline for Thiopurine Dosing Based on TPMT and NUDT 15 Genotypes: 2018 Update. Clin. Pharmacol. Ther. 2018, 105, 1095–1105. [Google Scholar] [CrossRef] [Green Version]
- Claassens, D.M.; Vos, G.J.; Bergmeijer, T.; Hermanides, R.S.; Hof, A.W.V.; Van Der Harst, P.; Barbato, E.; Morisco, C.; Gin, R.M.T.J.; Asselbergs, F.; et al. A Genotype-Guided Strategy for Oral P2Y12 Inhibitors in Primary PCI. N. Engl. J. Med. 2019, 381, 1621–1631. [Google Scholar] [CrossRef]
- Drogemoller, B.I.; Wright, G.E.B.; Shih, J.; Monzon, J.G.; Gelmon, K.A.; Ross, C.J.D.; Amstutz, U.; Carleton, B.C.; Group CCR. CYP2D6 as a treatment decision aid for ER-positive non-metastatic breast cancer patients: A systematic review with accompanying clinical practice guidelines. Breast Cancer Res. Treat. 2019, 173, 521–532. [Google Scholar] [CrossRef]
- Goetz, M.P.; Sangkuhl, K.; Guchelaar, H.-J.; Schwab, M.; Province, M.; Whirl-Carrillo, M.; Symmans, W.F.; McLeod, H.L.; Ratain, M.J.; Zembutsu, H.; et al. Clinical Pharmacogenetics Implementation Consortium (CPIC) Guideline for CYP2D6 and Tamoxifen Therapy. Clin. Pharmacol. Ther. 2018, 103, 770–777. [Google Scholar] [CrossRef] [Green Version]
- Van Der Wouden, C.H.; Paasman, E.; Teichert, M.; Crone, M.R.; Guchelaar, H.-J.; Swen, J.J. Assessing the Implementation of Pharmacogenomic Panel-Testing in Primary Care in the Netherlands Utilizing a Theoretical Framework. J. Clin. Med. 2020, 9, 814. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Van Westrhenen, R.; Aitchison, K.J.; Ingelman-Sundberg, M.; Jukić, M.M. Pharmacogenomics of Antidepressant and Antipsychotic Treatment: How Far Have We Got and Where Are We Going? Front. Psychiatry 2020, 11, 94. [Google Scholar] [CrossRef] [Green Version]
- Cardoso, F.; Kyriakides, S.; Ohno, S.; Penault-Llorca, F.; Poortmans, P.; Rubio, I.; Zackrisson, S.; Senkus, E. Early breast cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 2019, 30, 1194–1220. [Google Scholar] [CrossRef] [Green Version]
- Harris, L.N.; Ismaila, N.; McShane, L.M.; Andre, F.; Collyar, D.E.; Gonzalez-Angulo, A.M.; Hammond, E.H.; Kuderer, N.M.; Liu, M.C.; Mennel, R.G.; et al. Use of Biomarkers to Guide Decisions on Adjuvant Systemic Therapy for Women With Early-Stage Invasive Breast Cancer: American Society of Clinical Oncology Clinical Practice Guideline. J. Clin. Oncol. 2016, 34, 1134–1150. [Google Scholar] [CrossRef] [Green Version]
- Park, S.K.; Thigpen, J.; Lee, I.J. Coverage of pharmacogenetic tests by private health insurance companies. J. Am. Pharm. Assoc. 2020, 60, 352–356.e3. [Google Scholar] [CrossRef]
- Somogyi, A.A.; Phillips, E. Genomic testing as a tool to optimise drug therapy. Aust. Prescr. 2017, 40, 101–104. [Google Scholar] [CrossRef]
- Haga, S.B.; Burke, W.; Ginsburg, G.S.; Mills, R.; Agans, R. Primary care physicians’ knowledge of and experience with pharmacogenetic testing. Clin. Genet. 2012, 82, 388–394. [Google Scholar] [CrossRef] [Green Version]
- Hodges, L.M.; Markova, S.M.; Chinn, L.; Gow, J.M.; Kroetz, D.L.; Klein, T.E.; Altman, R.B. Very important pharmacogene summary. Pharmacogenet. Genom. 2011, 21, 152–161. [Google Scholar] [CrossRef] [Green Version]
- Caudle, K.E.; Dunnenberger, H.M.; Freimuth, R.; Peterson, J.F.; Burlison, J.D.; Whirl-Carrillo, M.; Scott, S.A.; Rehm, H.L.; Williams, M.S.; Klein, T.E.; et al. Standardizing terms for clinical pharmacogenetic test results: Consensus terms from the Clinical Pharmacogenetics Implementation Consortium (CPIC). Genet. Med. 2016, 19, 215–223. [Google Scholar] [CrossRef]
- Diatchenko, L.; Slade, G.D.; Nackley, A.G.; Bhalang, K.; Sigurdsson, A.; Belfer, I.; Goldman, D.; Xu, K.; Shabalina, S.; Shagin, D.; et al. Genetic basis for individual variations in pain perception and the development of a chronic pain condition. Hum. Mol. Genet. 2004, 14, 135–143. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Karnes, J.H.; Rettie, A.E.; Somogyi, A.A.; Huddart, R.; Fohner, A.E.; Formea, C.M.; Lee, M.T.M.; Llerena, A.; Whirl-Carrillo, M.; Klein, T.E.; et al. Clinical Pharmacogenetics Implementation Consortium (CPIC) Guideline for CYP2C9 and HLA-B Genotypes and Phenytoin Dosing: 2020 Update. Clin. Pharmacol. Ther. 2020, 109, 302–309. [Google Scholar] [CrossRef]
- Theken, K.; Lee, C.; Gong, L.; Caudle, K.E.; Formea, C.M.; Gaedigk, A.; Klein, T.E.; Agundez, J.; Grosser, T. Clinical Pharmacogenetics Implementation Consortium Guideline (CPIC) for CYP2C9 and Nonsteroidal Anti-Inflammatory Drugs. Clin. Pharmacol. Ther. 2020, 108, 191–200. [Google Scholar] [CrossRef]
- Hicks, J.K.; Sangkuhl, K.; Swen, J.; Ellingrod, V.L.; Müller, D.J.; Shimoda, K.; Bishop, J.R.; Kharasch, E.D.; Skaar, T.C.; Gaedigk, A.; et al. Clinical pharmacogenetics implementation consortium guideline (CPIC) for CYP2D6 and CYP2C19 genotypes and dosing of tricyclic antidepressants: 2016 update. Clin. Pharmacol. Ther. 2017, 102, 37–44. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lima, J.J.; Thomas, C.D.; Barbarino, J.; Desta, Z.; Van Driest, S.L.; El Rouby, N.; Johnson, J.A.; Cavallari, L.H.; Shakhnovich, V.; Thacker, D.L.; et al. Clinical Pharmacogenetics Implementation Consortium (CPIC) Guideline for CYP2C19 and Proton Pump Inhibitor Dosing. Clin. Pharmacol. Ther. 2020, 109, 1417–1423. [Google Scholar] [CrossRef]
- Caudle, K.E.; Sangkuhl, K.; Whirl-Carrillo, M.; Swen, J.; Haidar, C.E.; Klein, T.E.; Gammal, R.; Relling, M.V.; Scott, S.A.; Hertz, D.L.; et al. Standardizing CYP 2D6 Genotype to Phenotype Translation: Consensus Recommendations from the Clinical Pharmacogenetics Implementation Consortium and Dutch Pharmacogenetics Working Group. Clin. Transl. Sci. 2019, 13, 116–124. [Google Scholar] [CrossRef] [Green Version]
- Birdwell, K.; Decker, B.; Barbarino, J.M.; Peterson, J.F.; Stein, C.M.; Sadee, W.; Wang, D.; Vinks, A.; He, Y.; Swen, J.; et al. Clinical Pharmacogenetics Implementation Consortium (CPIC) Guidelines forCYP3A5Genotype and Tacrolimus Dosing. Clin. Pharmacol. Ther. 2015, 98, 19–24. [Google Scholar] [CrossRef] [Green Version]
- Amstutz, U.; Henricks, L.; Offer, S.M.; Barbarino, J.; Schellens, J.H.; Swen, J.; Klein, T.E.; McLeod, H.L.; Caudle, K.E.; Diasio, R.B.; et al. Clinical Pharmacogenetics Implementation Consortium (CPIC) Guideline for Dihydropyrimidine Dehydrogenase Genotype and Fluoropyrimidine Dosing: 2017 Update. Clin. Pharmacol. Ther. 2017, 103, 210–216. [Google Scholar] [CrossRef]
- Russmann, S.; Rahmany, A.; Niedrig, D.; Hatz, K.; Ludin, K.; Burden, A.; Englberger, L.; Backhaus, R.; Serra, A.; Bechir, M. Implementation and Management Outcomes of Pharmacogenetic CYP2C19 Testing for Clopidogrel Therapy in Clinical Practice. Eur. J. Clin. Pharmacol. 2021, 77, 709–716. [Google Scholar] [CrossRef] [PubMed]
- Fricke-Galindo, I.; Céspedes-Garro, C.; Rodrigues-Soares, F.; Naranjo, M.E.G.; Delgado, Á.; De Andrés, F.; López-López, M.; Peñas-Lledó, E.; Llerena, A. Interethnic variation of CYP2C19 alleles, ‘predicted’ phenotypes and ‘measured’ metabolic phenotypes across world populations. Pharm. J. 2015, 16, 113–123. [Google Scholar] [CrossRef]
- Ji, Y.; Skierka, J.M.; Blommel, J.H.; Moore, B.E.; VanCuyk, D.L.; Bruflat, J.K.; Peterson, L.M.; Veldhuizen, T.L.; Fadra, N.; Peterson, S.E.; et al. Preemptive Pharmacogenomic Testing for Precision Medicine. J. Mol. Diagn. 2016, 18, 438–445. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Naranjo, M.-E.G.; Rodrigues-Soares, F.; Peñas-Lledó, E.M.; Tarazona, E.; Fariñas, H.; Rodeiro, I.; Teran, E.; Grazina, M.; Moya, G.E.; López-López, M.; et al. Interethnic Variability in CYP2D6, CYP2C9, and CYP2C19 Genes and Predicted Drug Metabolism Phenotypes Among 6060 Ibero- and Native Americans: RIBEF-CEIBA Consortium Report on Population Pharmacogenomics. OMICS J. Integr. Biol. 2018, 22, 575–588. [Google Scholar] [CrossRef]
- Schildcrout, J.S.; Denny, J.; Bowton, E.; Gregg, W.; Pulley, J.M.; Basford, M.A.; Cowan, J.D.; Xu, H.; Ramirez, A.H.; Crawford, D.; et al. Optimizing Drug Outcomes Through Pharmacogenetics: A Case for Preemptive Genotyping. Clin. Pharmacol. Ther. 2012, 92, 235–242. [Google Scholar] [CrossRef]
- Moliterno, D.J.; Smyth, S.S.; Abdel-Latif, A. CYP2C19 Genotyping to Guide Antiplatelet Therapy After Percutaneous Coronary Interventions. JAMA 2020, 324, 747. [Google Scholar] [CrossRef] [PubMed]
- Pereira, N.L.; Farkouh, M.E.; So, D.; Lennon, R.; Geller, N.; Mathew, V.; Bell, M.; Bae, J.-H.; Jeong, M.H.; Chavez, I.; et al. Effect of Genotype-Guided Oral P2Y12 Inhibitor Selection vs Conventional Clopidogrel Therapy on Ischemic Outcomes After Percutaneous Coronary Intervention. JAMA 2020, 324, 761. [Google Scholar] [CrossRef] [PubMed]
- Jussmann, S.; Jetter, A.; Kullak-Ublick, G.A. Pharmacogenetics of drug-induced liver injury. Hepatology 2010, 52, 748–761. [Google Scholar] [CrossRef] [PubMed]
- Gentile, G.; Borro, M.; Lala, N.; Missori, S.; Simmaco, M.; Martelletti, P. Genetic polymorphisms related to efficacy and overuse of triptans in chronic migraine. J. Headache Pain 2010, 11, 431–435. [Google Scholar] [CrossRef] [Green Version]
- Mallal, S.; Phillips, E.; Carosi, G.; Molina, J.-M.; Workman, C.; Tomažič, J.; Jägel-Guedes, E.; Rugina, S.; Kozyrev, O.; Cid, J.F.; et al. HLA-B*5701 Screening for Hypersensitivity to Abacavir. N. Engl. J. Med. 2008, 358, 568–579. [Google Scholar] [CrossRef] [Green Version]
- Phillips, E.J.; Sukasem, C.; Whirl-Carrillo, M.; Müller, D.J.; Dunnenberger, H.; Chantratita, W.; Goldspiel, B.; Chen, Y.; Carleton, B.; George, A.L., Jr.; et al. Clinical Pharmacogenetics Implementation Consortium Guideline for HLA Genotype and Use of Carbamazepine and Oxcarbazepine: 2017 Update. Clin. Pharmacol. Ther. 2017, 103, 574–581. [Google Scholar] [CrossRef] [Green Version]


| All Patients with PGx Panel Testing n (%) | Patients with Specific Drug–Gene-Based Indication n (%) | Patients with PGx Screening n (%) | |
|---|---|---|---|
| n (%) | 135 (100) | 102 (75.5) | 33 (24.5) |
| Age: median (range) | 68 (25–92) | 70 (25–92) | 58 (30–83) |
| <60 | 48 (35.6) | 30 (29.4) | 18 (54.6) |
| 61–70 | 25 (18.5) | 22 (21.6) | 3 (9.1) |
| 71–80 | 41 (30.4) | 33 (32.4) | 8 (24.2) |
| >80 | 21 (15.6) | 17 (16.7) | 4 (12.1) |
| Sex | |||
| male | 81 (60) | 56 (54.9) | 25 (75.8) |
| female | 54 (40) | 46 (45.1) | 8 (24.2) |
| eGFR < 60 mL/min 1 | 19 (14.1) | 16 (15.7) | 3 (9.1) |
| Main diagnosis | |||
| Vascular disease | 68 (50.4) | 68 (66.7) | n.a. |
| Oncological disease | 17 (12.6) | 17 (16.7) | n.a. |
| Psychiatric disease | 9 (6.7) | 9 (8.8) | n.a. |
| Pain in orthopedic disease | 7 (5.2) | 7 (6.9) | n.a. |
| Gastric disease | 1 (0.7) | 1 (1.0) | n.a. |
| Indication PGx panel test | |||
| Clopidogrel | 60 (44.4) | 60 (58.8) | n.a. |
| Tamoxifen | 15 (11.1) | 15 (14.7) | n.a. |
| Polypsychopharmacotherapy | 9 (6.7) | 9 (8.8) | n.a. |
| Opioids | 7 (5.2) | 7 (6.9) | n.a. |
| Statins | 6 (4.4) | 6 (5.9) | n.a. |
| Phenprocoumon | 2 (1.5) | 2 (2.0) | n.a. |
| Chemotherapy | 2 (1.5) | 2 (2.0) | n.a. |
| Proton pump inhibitor | 1 (0.7) | 1 (1.0) | n.a. |
| Pharmacotherapy | |||
| Number of drugs, median (range) 2 | 6 (0–19) | 6 (0–19) | 3 (0–14) |
| Aspirin | 43 (31.9) | 38 (37.3) | 5 (15.2) |
| Clopidogrel | 48 (35.6) | 48 (47.1) | 0 (0) |
| Prasugrel or Ticagrelor | 1 (0.7) | 1 (1.0) | 0 (0) |
| Coumarines or NOAC | 25 (18.5) | 22 (21.6) | 3 (9.1) |
| Beta blockers | 44 (32.6) | 35 (34.3) | 9 (27.3) |
| ACE inhibitors or ARB) 3 | 60 (44.4) | 48 (47.1) | 12 (36.4) |
| Calcium channel blockers | 20 (14.8) | 16 (15.7) | 4 (12.1) |
| Diuretics | 34 (25.2) | 28 (27.5) | 6 (18.2) |
| PPI | 45 (33.3) | 40 (39.2) | 5 (15.2) |
| Cholesterol lowering drugs | 55 (40.7) | 48 (47.1) | 7 (21.2) |
| NSAR | 12 (8.9) | 11 (10.8) | 1 (3.0) |
| Opioids | 17 (12.6) | 14 (13.7) | 3 (9.1) |
| Uric acid lowering drugs | 5 (3.7) | 3 (2.9) | 2 (6.1) |
| Benzodiazepines | 18 (13.3) | 14 (13.7) | 4 (12.1) |
| Antidepressants | 28 (20.7) | 24 (23.5) | 4 (12.1) |
| Antipsychotics | 10 (7.4) | 9 (8.8) | 1 (3.0) |
| Antiepileptics | 9 (6.7) | 8 (7.8) | 1 (3.0) |
| Antidiabetics | 22 (16.3) | 17 (16.7) | 5 (15.2) |
| Tamoxifen | 12 (8.9) | 12 (11.8) | 0 (0) |
| CYP2C19 Inhibitor 4 | 11 (8.2) | 10 (9.8) | 1 (3.0) |
| CYP2D6 Inhibitor 4 | 26 (19.3) | 20 (19.6) | 6 (18.2) |
| Gene | Drugs with Required PGx-Testing 1 | Drugs with Recommended PGx-Testing 1 | Drugs with Actionable PGx-Testing 1 | n (%) Patients with Phenotype Variants 2 |
|---|---|---|---|---|
| ABCB1 | - | - | - | 106 (78.5) 4 |
| CYP2C9 | siponimod | - | celecoxib, phenytoin, warfarin | 46 (34) |
| CYP2C19 | - | atazanavir | amitriptyline, carisoprodol, citalopram, clobazam, clomipramine, clopidogrel, desipramine, doxepin, imipramine, nortriptyline, pantoprazole, trimipramine, voriconazole | 71 (52) |
| CYP2D6 | pimozide, tetrabenazine | - | amitriptyline, aripiprazole, atomoxetine, brexpiprazole, carvedilol, cevimeline, citalopram, clomipramine, clozapine, codeine, darifenacin, desipramine, doxepin, fesoterodine, iloperodine, nortriptyline, perphenazine, propafenone, tamoxifen, thioridazine, tramadol, trimipramine, vortioxetine | 67 (50) |
| SLCO1B13 | - | - | - | 30 (22) |
| VKORC1 | - | - | warfarin | 86 (63.7) |
| COMT | - | - | - | 73 (54.1) |
| CYP1A2 | - | - | - | 65 (48.6) |
| CYP2B6 | - | - | efavirenz | 67 (49.6) |
| CYP3A4 | - | - | codeine, tamoxifen | 6 (4.4) |
| CYP3A5 | - | - | - | 17 (12.6) |
| CYP4F2 | - | - | warfarin | 66 (48,9) 4 |
| DPYD4 | - | - | capecitabine, fluorouracil | 5 (3.7) |
| OPRM1 | - | - | codeine | 34 (25.2) |
| POR | - | - | - | 72 (53.3) 4 |
| TPMT | - | azathioprine, mercaptopurine | tioguanine | 8 (5.9) |
| n | Drugs that Triggered PGx-Testing | Relevant Gene(s) | Detected Phenotype Variants 1 | Patients with SONOGEN XP Recommendation to Change Triggering Drug | Patients with Clinical Expert Recommendation to Change Triggering Drug | Patients with Additional Clinical Expert Recommendations for Current but Nontriggering Drug(s) 2 |
|---|---|---|---|---|---|---|
| 102 | All patients with specific indication | n.a. | n.a. | 36 (35.3%) | 33 (32.4%) | 23 (22.5%) |
| 60 | Clopidogrel | CYP2C19 | 1 PM/19 IM/ 19 RM or UM | 20 (33.3%) | 19 (31.6%) | 16 (26.7%) |
| 15 | Tamoxifen | CYP2D6 | 3 IM/1 UM | 3 (20.0%) | 1 (6,7%) | 0 |
| 9 | Polypsycho-pharmacotherapy | CYP1A2 CYP2D6 CYP2C19 | 1A2: 6 UM CYP2D6: 7 IM/1 PM CYP2C19: 6 UM | 5 (55.6%) | 3 (33.3%) | 3 (33.3%) |
| 7 | Opioids | OPRM CYP2D6 | OPRM1 3 decreased function CYP2D6: 4 IM | 4 (57.1%) | 3 (42.9%) | 4 (57.1%) |
| 6 | Statins | SLCO1B1 | 4 decreased or poor function | 2 (33.3%) | 4 (66.7%) | 0 (%) |
| 2 | Phenprocoumon | VKORC1 CYP4F2 CYP2C9 | VKORC: 1 normal, 1 decreased function CYP2C9: 2 normal function CYP4F2: 2 normal function | 1 (50.0%) | 2 (100%) | 0 |
| 2 | Chemotherapy | DPYD | 0 | 0 | 0 | 0 |
| 1 | Proton pump inhibitor | CYP2C19 | 1 UM | 1 (100%) | 1 (100%) | 0 |
| 33 | Screening | n.a. | n.a. | n.a. | n.a. | 3 (9.1%) |
| Trigger for PGx-Testing | n Patients | n Patients with ≥1 “Required” or “Recommended” PGx Variant 1 | n SONOGEN XP Recommendations 2 per Patient Median (Range) | n Highlighted Clinical Expert Recommendations 3 per Patient Median (Range) |
|---|---|---|---|---|
| Specific PGx drug | 102 | 88 (86.3%) | 2 (2–11) | 2 (0–6) |
| Screening | 33 | 29 (87.9%) | 5 (3–9) | 3 (1–5) |
| All Patients | 135 | 117 (86.7%) | 5 (2–11) | 3 (0–6) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Niedrig, D.F.; Rahmany, A.; Heib, K.; Hatz, K.-D.; Ludin, K.; Burden, A.M.; Béchir, M.; Serra, A.; Russmann, S. Clinical Relevance of a 16-Gene Pharmacogenetic Panel Test for Medication Management in a Cohort of 135 Patients. J. Clin. Med. 2021, 10, 3200. https://doi.org/10.3390/jcm10153200
Niedrig DF, Rahmany A, Heib K, Hatz K-D, Ludin K, Burden AM, Béchir M, Serra A, Russmann S. Clinical Relevance of a 16-Gene Pharmacogenetic Panel Test for Medication Management in a Cohort of 135 Patients. Journal of Clinical Medicine. 2021; 10(15):3200. https://doi.org/10.3390/jcm10153200
Chicago/Turabian StyleNiedrig, David F., Ali Rahmany, Kai Heib, Karl-Dietrich Hatz, Katja Ludin, Andrea M. Burden, Markus Béchir, Andreas Serra, and Stefan Russmann. 2021. "Clinical Relevance of a 16-Gene Pharmacogenetic Panel Test for Medication Management in a Cohort of 135 Patients" Journal of Clinical Medicine 10, no. 15: 3200. https://doi.org/10.3390/jcm10153200
APA StyleNiedrig, D. F., Rahmany, A., Heib, K., Hatz, K.-D., Ludin, K., Burden, A. M., Béchir, M., Serra, A., & Russmann, S. (2021). Clinical Relevance of a 16-Gene Pharmacogenetic Panel Test for Medication Management in a Cohort of 135 Patients. Journal of Clinical Medicine, 10(15), 3200. https://doi.org/10.3390/jcm10153200






