Composition and Clinical Significance of Exosomes in Tuberculosis: A Systematic Literature Review
Abstract
1. Introduction
2. Methods
3. Review
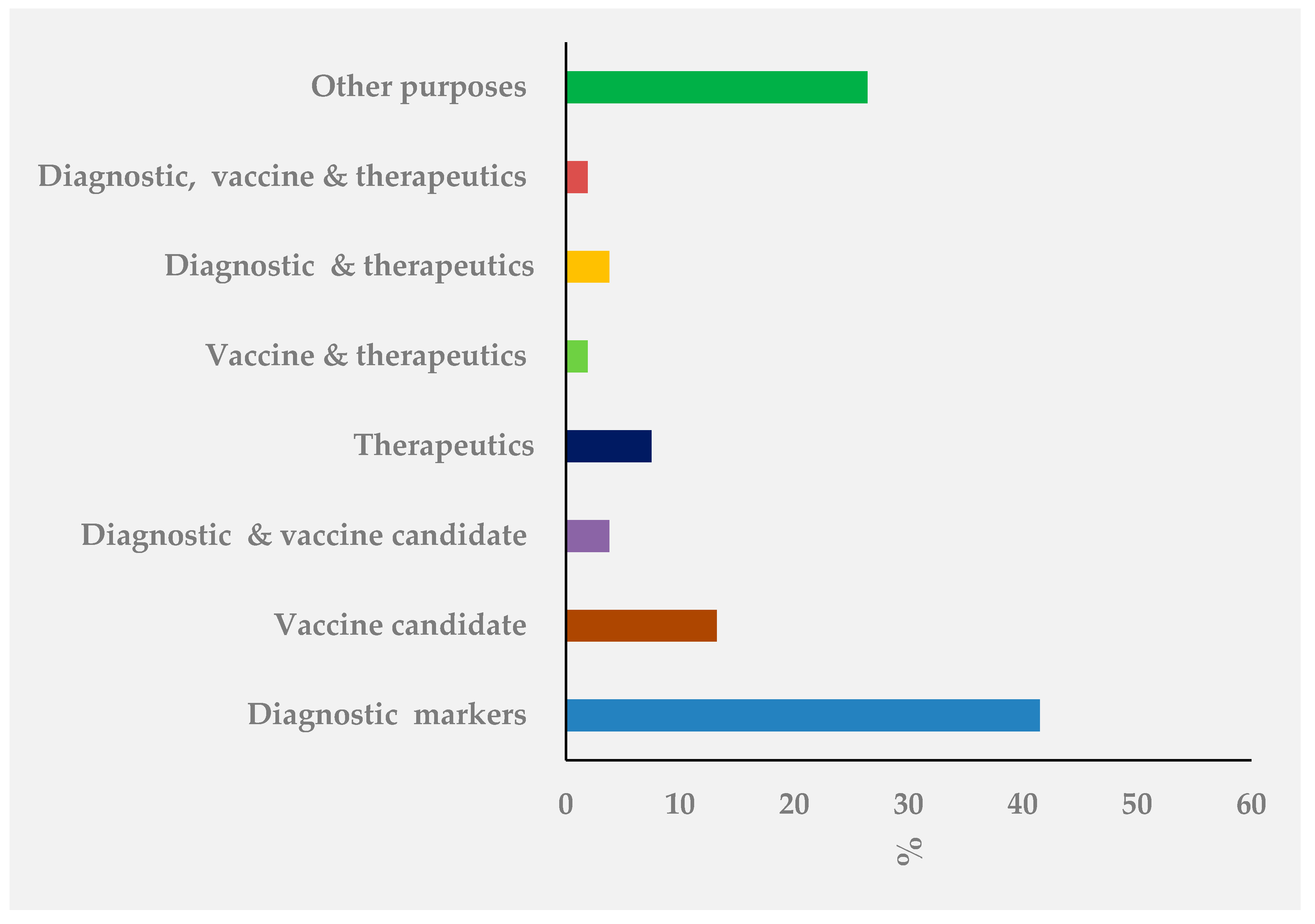
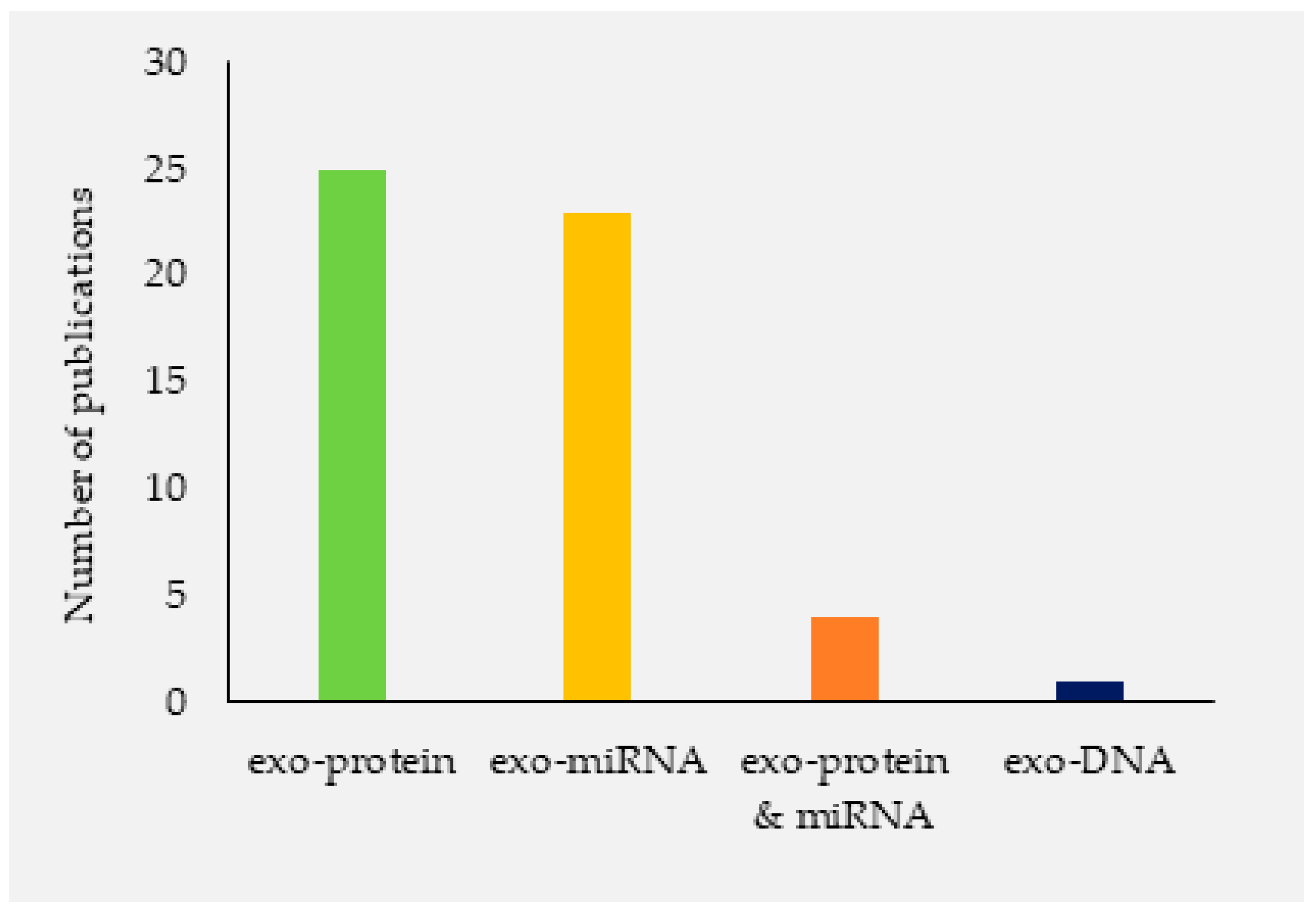
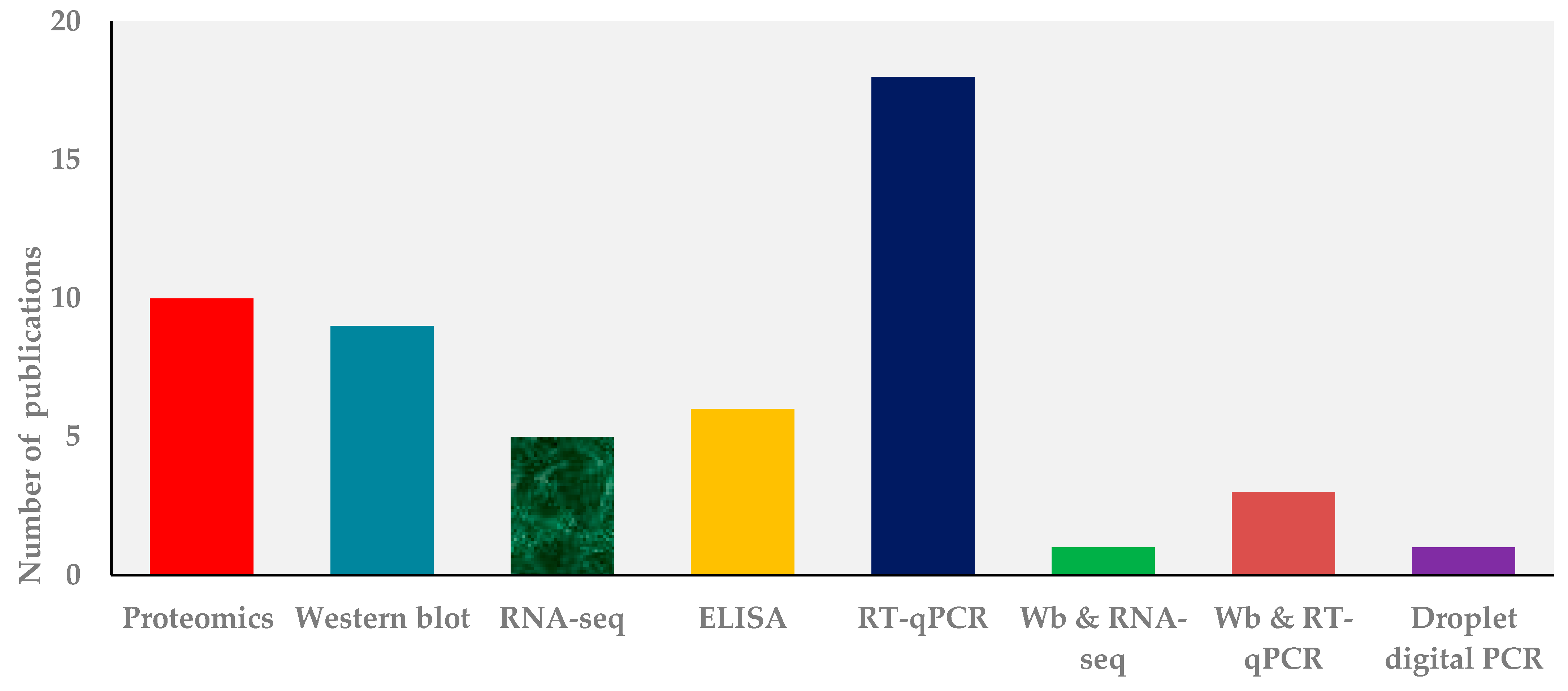
3.1. Finding
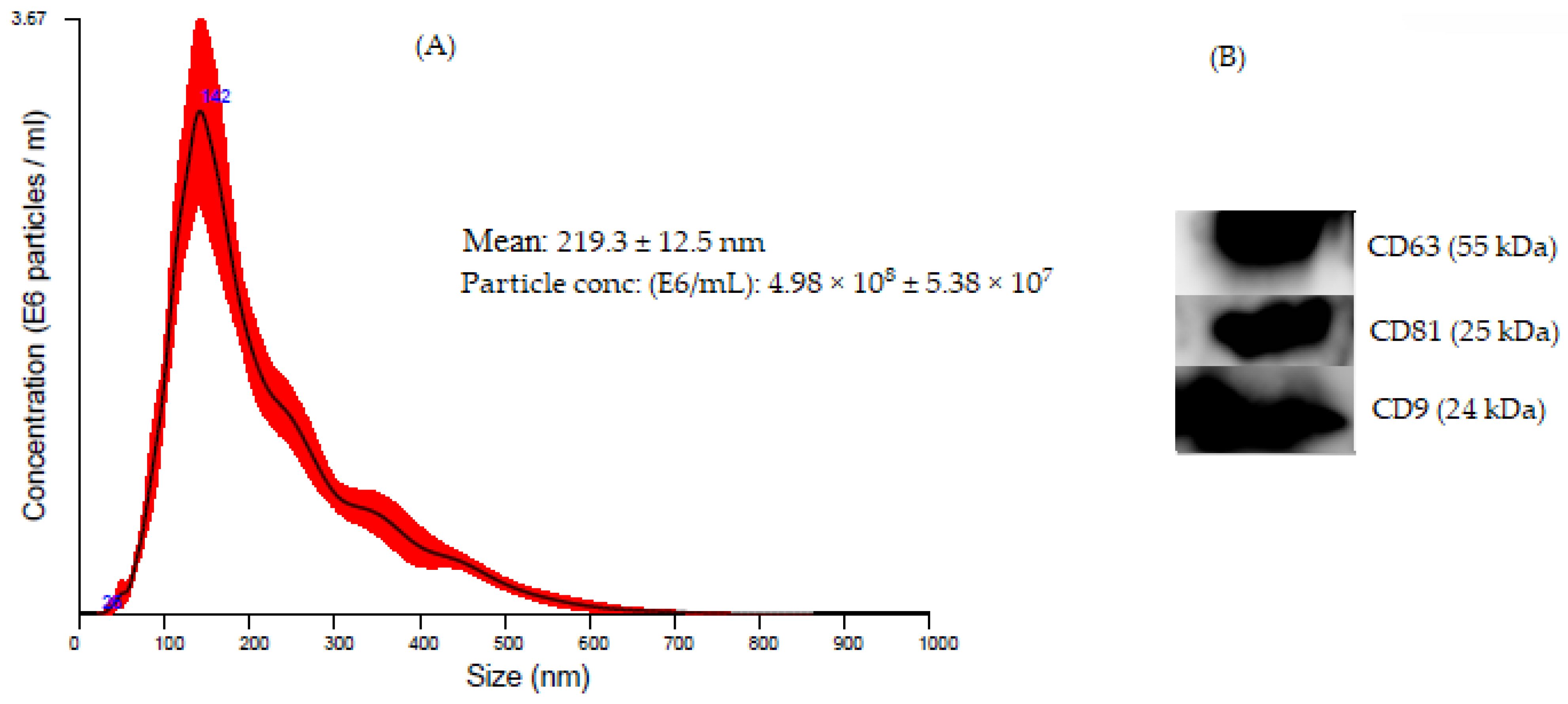
3.2. Mtb Exosomes Isolation and Characterization Methods
3.3. Composition of Host Exosomal Surface Markers by Mtb Infected Cells
3.4. Mtb Molecules Extracted from Exosomes
3.5. Exosome and Other Lung Diseases
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- World Health Organization. Global Tuberculosis Report 2018; World Health Organization: Geneva, Switzerland, 2018; ISBN 978-92-4-156564-6. [Google Scholar]
- World Health Organization. WHO Guidelines on Tuberculosis Infection Prevention and Control. 2019 Update; World Health Organization: Geneva, Switzerland, 2019; ISBN 978-92-4-155051-2. [Google Scholar]
- Flynn, J.L.; Chan, J. Tuberculosis: Latency and reactivation. Infect. Immun. 2001, 69, 4195–4201. [Google Scholar] [CrossRef]
- Wang, J.; Yao, Y.; Chen, X.; Wu, J.; Gu, T.; Tang, X. Host derived exosomes-pathogens interactions: Potential functions of exosomes in pathogen infection. Biomed. Pharmacother. 2018, 108, 1451–1459. [Google Scholar] [CrossRef]
- Borges, F.T.; Reis, L.A.; Schor, N. Extracellular vesicles: Structure, function, and potential clinical uses in renal diseases. Braz. J. Med. Biol. Res. 2013, 46, 824–830. [Google Scholar] [CrossRef] [PubMed]
- Rashed, M.H.; Bayraktar, E.; Helal, G.K.; Abd-Ellah, M.F.; Amero, P.; Chavez-Reyes, A.; Rodriguez-Aguayo, C. Exosomes: From Garbage Bins to Promising Therapeutic Targets. Int. J. Mol. Sci. 2017, 18. [Google Scholar] [CrossRef] [PubMed]
- Bebelman, M.P.; Smit, M.J.; Pegtel, D.M.; Baglio, S.R. Biogenesis and function of extracellular vesicles in cancer. Pharmacol. Ther. 2018, 188, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Raposo, G.; Stoorvogel, W. Extracellular vesicles: Exosomes, microvesicles, and friends. J. Cell Biol. 2013, 200, 373–383. [Google Scholar] [CrossRef]
- Zhang, W.; Jiang, X.; Bao, J.; Wang, Y.; Liu, H.; Tang, L. Exosomes in Pathogen Infections: A Bridge to Deliver Molecules and Link Functions. Front. Immunol. 2018, 9, 90. [Google Scholar] [CrossRef]
- Palviainen, M.; Saraswat, M.; Varga, Z.; Kitka, D.; Neuvonen, M.; Puhka, M.; Joenväärä, S.; Renkonen, R.; Nieuwland, R.; Takatalo, M.; et al. Extracellular vesicles from human plasma and serum are carriers of extravesicular cargo-Implications for biomarker discovery. PLoS ONE 2020, 15, e0236439. [Google Scholar] [CrossRef] [PubMed]
- Wiklander, O.P.B.; Brennan, M.Á.; Lötvall, J.; Breakefield, X.O.; El Andaloussi, S. Advances in therapeutic applications of extracellular vesicles. Sci. Transl. Med. 2019, 11. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Lee, H.; Park, K.; Shin, S. Rapid and Efficient Isolation of Exosomes by Clustering and Scattering. J. Clin. Med. 2020, 9. [Google Scholar] [CrossRef] [PubMed]
- Lobb, R.J.; Becker, M.; Wen, S.W.; Wong, C.S.F.; Wiegmans, A.P.; Leimgruber, A.; Möller, A. Optimized exosome isolation protocol for cell culture supernatant and human plasma. J. Extracell. Vesicles 2015, 4, 27031. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.; Qu, X.; Dong, Z.; Zeng, Q.; Ma, X.; Jia, Y.; Li, R.; Jiang, X.; Williams, C.; Wang, T.; et al. Comparison of serum exosome isolation methods on co-precipitated free microRNAs. PeerJ 2020, 8, e9434. [Google Scholar] [CrossRef]
- Wijenayake, S.; Eisha, S.; Tawhidi, Z.; Pitino, M.A.; Steele, M.A.; Fleming, A.S.; McGowan, P.O. Comparison of methods for milk pre-processing, exosome isolation, and RNA extraction in bovine and human milk. bioRxiv 2020. [Google Scholar] [CrossRef]
- Cho, S.; Yang, H.C.; Rhee, W.J. Development and comparative analysis of human urine exosome isolation strategies. Process Biochemistry 2020, 88, 197–203. [Google Scholar] [CrossRef]
- Zhou, W.; Tahir, F.; Wang, J.C.-Y.; Woodson, M.; Sherman, M.B.; Karim, S.; Neelakanta, G.; Sultana, H. Discovery of Exosomes From Tick Saliva and Salivary Glands Reveals Therapeutic Roles for CXCL12 and IL-8 in Wound Healing at the Tick-Human Skin Interface. Front. Cell Dev. Biol. 2020, 8, 554. [Google Scholar] [CrossRef] [PubMed]
- Crenshaw, B.J.; Sims, B.; Matthews, Q.L. Biological Function of Exosomes as Diagnostic Markers and Therapeutic Delivery Vehicles in Carcinogenesis and Infectious Diseases. In Nanomedicines; Akhyar Farrukh, M., Ed.; IntechOpen: Cham, Switerland, 2019; ISBN 978-1-78985-283-7. [Google Scholar] [CrossRef]
- Helwa, I.; Cai, J.; Drewry, M.D.; Zimmerman, A.; Dinkins, M.B.; Khaled, M.L.; Seremwe, M.; Dismuke, W.M.; Bieberich, E.; Stamer, W.D.; et al. A Comparative Study of Serum Exosome Isolation Using Differential Ultracentrifugation and Three Commercial Reagents. PLoS ONE 2017, 12, e0170628. [Google Scholar] [CrossRef] [PubMed]
- Admyre, C.; Johansson, S.M.; Qazi, K.R.; Filén, J.-J.; Lahesmaa, R.; Norman, M.; Neve, E.P.A.; Scheynius, A.; Gabrielsson, S. Exosomes with immune modulatory features are present in human breast milk. J. Immunol. 2007, 179, 1969–1978. [Google Scholar] [CrossRef]
- Słomka, A.; Urban, S.K.; Lukacs-Kornek, V.; Żekanowska, E.; Kornek, M. Large Extracellular Vesicles: Have We Found the Holy Grail of Inflammation? Front. Immunol. 2018, 9, 2723. [Google Scholar] [CrossRef]
- Kang, H.; Kim, J.; Park, J. Methods to isolate extracellular vesicles for diagnosis. Micro Nano Syst. Lett. 2017, 5. [Google Scholar] [CrossRef]
- Rodrigues, M.; Fan, J.; Lyon, C.; Wan, M.; Hu, Y. Role of Extracellular Vesicles in Viral and Bacterial Infections: Pathogenesis, Diagnostics, and Therapeutics. Theranostics 2018, 8, 2709–2721. [Google Scholar] [CrossRef]
- Słomka, A.; Mocan, T.; Wang, B.; Nenu, I.; Urban, S.K.; Gonzales-Carmona, M.; Schmidt-Wolf, I.G.H.; Lukacs-Kornek, V.; Strassburg, C.P.; Spârchez, Z.; et al. EVs as Potential New Therapeutic Tool/Target in Gastrointestinal Cancer and HCC. Cancers 2020, 12, 19. [Google Scholar] [CrossRef] [PubMed]
- Zhou, B.; Xu, K.; Zheng, X.; Chen, T.; Wang, J.; Song, Y.; Shao, Y.; Zheng, S. Application of exosomes as liquid biopsy in clinical diagnosis. Signal Transduct. Target. Ther. 2020, 5, 144. [Google Scholar] [CrossRef] [PubMed]
- Maisano, D.; Mimmi, S.; Russo, R.; Fioravanti, A.; Fiume, G.; Vecchio, E.; Nisticò, N.; Quinto, I.; Iaccino, E. Uncovering the Exosomes Diversity: A Window of Opportunity for Tumor Progression Monitoring. Pharmaceuticals 2020, 13, 180. [Google Scholar] [CrossRef]
- Krishnan, S.R.; Luk, F.; Brown, R.D.; Suen, H.; Kwan, Y.; Bebawy, M. Isolation of Human CD138(+) Microparticles from the Plasma of Patients with Multiple Myeloma. Neoplasia 2016, 18, 25–32. [Google Scholar] [CrossRef] [PubMed]
- Boukouris, S.; Mathivanan, S. Exosomes in bodily fluids are a highly stable resource of disease biomarkers. Proteomics Clin. Appl. 2015, 9, 358–367. [Google Scholar] [CrossRef]
- Iaccino, E.; Mimmi, S.; Dattilo, V.; Marino, F.; Candeloro, P.; Di Loria, A.; Marimpietri, D.; Pisano, A.; Albano, F.; Vecchio, E.; et al. Monitoring multiple myeloma by idiotype-specific peptide binders of tumor-derived exosomes. Mol. Cancer 2017, 16, 159. [Google Scholar] [CrossRef]
- Manna, I.; Iaccino, E.; Dattilo, V.; Barone, S.; Vecchio, E.; Mimmi, S.; Filippelli, E.; Demonte, G.; Polidoro, S.; Granata, A.; et al. Exosome-associated miRNA profile as a prognostic tool for therapy response monitoring in multiple sclerosis patients. FASEB J. 2018, 32, 4241–4246. [Google Scholar] [CrossRef]
- Tauro, B.J.; Greening, D.W.; Mathias, R.A.; Ji, H.; Mathivanan, S.; Scott, A.M.; Simpson, R.J. Comparison of ultracentrifugation, density gradient separation, and immunoaffinity capture methods for isolating human colon cancer cell line LIM1863-derived exosomes. Methods 2012, 56, 293–304. [Google Scholar] [CrossRef]
- Doyle, L.M.; Wang, M.Z. Overview of Extracellular Vesicles, Their Origin, Composition, Purpose, and Methods for Exosome Isolation and Analysis. Cells 2019, 8, 727. [Google Scholar] [CrossRef]
- Yakimchuk, K. Exosomes: Isolation methods and specific markers. Mater Methods 2015, 5. [Google Scholar] [CrossRef]
- Mathieu, M.; Martin-Jaular, L.; Lavieu, G.; Théry, C. Specificities of secretion and uptake of exosomes and other extracellular vesicles for cell-to-cell communication. Nat. Cell Biol. 2019, 21, 9–17. [Google Scholar] [CrossRef] [PubMed]
- Dickhout, A.; Koenen, R.R. Extracellular Vesicles as Biomarkers in Cardiovascular Disease; Chances and Risks. Front. Cardiovasc. Med. 2018, 5, 113. [Google Scholar] [CrossRef] [PubMed]
- Palacios, A.; Sampedro, L.; Sevilla, I.A.; Molina, E.; Gil, D.; Azkargorta, M.; Elortza, F.; Garrido, J.M.; Anguita, J.; Prados-Rosales, R. Mycobacterium tuberculosis extracellular vesicle-associated lipoprotein LpqH as a potential biomarker to distinguish paratuberculosis infection or vaccination from tuberculosis infection. BMC Vet. Res. 2019, 15, 188. [Google Scholar] [CrossRef] [PubMed]
- Lyu, L.; Zhang, X.; Li, C.; Yang, T.; Wang, J.; Pan, L.; Jia, H.; Li, Z.; Sun, Q.; Yue, L.; et al. Small RNA Profiles of Serum Exosomes Derived From Individuals With Latent and Active Tuberculosis. Front. Microbiol. 2019, 10, 1174. [Google Scholar] [CrossRef] [PubMed]
- Singh, P.P.; LeMaire, C.; Tan, J.C.; Zeng, E.; Schorey, J.S. Exosomes released from M. tuberculosis infected cells can suppress IFN-γ mediated activation of naïve macrophages. PLoS ONE 2011, 6, e18564. [Google Scholar] [CrossRef] [PubMed]
- Prados-Rosales, R.; Baena, A.; Martinez, L.R.; Luque-Garcia, J.; Kalscheuer, R.; Veeraraghavan, U.; Camara, C.; Nosanchuk, J.D.; Besra, G.S.; Chen, B.; et al. Mycobacteria release active membrane vesicles that modulate immune responses in a TLR2-dependent manner in mice. J. Clin. Investig. 2011, 121, 1471–1483. [Google Scholar] [CrossRef] [PubMed]
- Tyagi, P.; Pal, V.K.; Agrawal, R.; Singh, S.; Srinivasan, S.; Singh, A. Mycobacterium tuberculosis Reactivates HIV-1 via Exosome-Mediated Resetting of Cellular Redox Potential and Bioenergetics. mBio 2020, 11. [Google Scholar] [CrossRef]
- Lv, L.; Li, C.; Zhang, X.; Ding, N.; Cao, T.; Jia, X.; Wang, J.; Pan, L.; Jia, H.; Li, Z.; et al. RNA Profiling Analysis of the Serum Exosomes Derived from Patients with Active and Latent Mycobacterium tuberculosis Infection. Front. Microbiol. 2017, 8, 1051. [Google Scholar] [CrossRef]
- Baranyai, T.; Herczeg, K.; Onódi, Z.; Voszka, I.; Módos, K.; Marton, N.; Nagy, G.; Mäger, I.; Wood, M.J.; El Andaloussi, S.; et al. Isolation of Exosomes from Blood Plasma: Qualitative and Quantitative Comparison of Ultracentrifugation and Size Exclusion Chromatography Methods. PLoS ONE 2015, 10, e0145686. [Google Scholar] [CrossRef]
- Momen-Heravi, F.; Balaj, L.; Alian, S.; Trachtenberg, A.J.; Hochberg, F.H.; Skog, J.; Kuo, W.P. Impact of biofluid viscosity on size and sedimentation efficiency of the isolated microvesicles. Front. Physiol. 2012, 3, 162. [Google Scholar] [CrossRef]
- Théry, C.; Amigorena, S.; Raposo, G.; Clayton, A. Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Curr. Protoc. Cell Biol. 2006, 3. [Google Scholar] [CrossRef] [PubMed]
- Witwer, K.W.; Buzás, E.I.; Bemis, L.T.; Bora, A.; Lässer, C.; Lötvall, J.; Nolte-’t Hoen, E.N.; Piper, M.G.; Sivaraman, S.; Skog, J.; et al. Standardization of sample collection, isolation and analysis methods in extracellular vesicle research. J. Extracell. Vesicles 2013, 2. [Google Scholar] [CrossRef] [PubMed]
- Raposo, G.; Nijman, H.W.; Stoorvogel, W.; Liejendekker, R.; Harding, C.V.; Melief, C.J.; Geuze, H.J. B lymphocytes secrete antigen-presenting vesicles. J. Exp. Med. 1996, 183, 1161–1172. [Google Scholar] [CrossRef] [PubMed]
- Athman, J.J.; Wang, Y.; McDonald, D.J.; Boom, W.H.; Harding, C.V.; Wearsch, P.A. Bacterial Membrane Vesicles Mediate the Release of Mycobacterium tuberculosis Lipoglycans and Lipoproteins from Infected Macrophages. J. Immunol. 2015, 195, 1044–1053. [Google Scholar] [CrossRef]
- Vella, L.J.; Sharples, R.A.; Nisbet, R.M.; Cappai, R.; Hill, A.F. The role of exosomes in the processing of proteins associated with neurodegenerative diseases. Eur. Biophys. J. 2008, 37, 323–332. [Google Scholar] [CrossRef]
- García-Martínez, M.; Vázquez-Flores, L.; Álvarez-Jiménez, V.D.; Castañeda-Casimiro, J.; Ibáñez-Hernández, M.; Sánchez-Torres, L.E.; Barrios-Payán, J.; Mata-Espinosa, D.; Estrada-Parra, S.; Chacón-Salinas, R.; et al. Extracellular vesicles released by J774A.1 macrophages reduce the bacterial load in macrophages and in an experimental mouse model of tuberculosis. Int. J. Nanomed. 2019, 14, 6707–6719. [Google Scholar] [CrossRef]
- Dahiya, B.; Khan, A.; Mor, P.; Kamra, E.; Singh, N.; Gupta, K.B.; Sheoran, A.; Sreenivas, V.; Mehta, P.K. Detection of Mycobacterium tuberculosis lipoarabinomannan and CFP-10 (Rv3874) from urinary extracellular vesicles of tuberculosis patients by immuno-PCR. Pathog. Dis. 2019, 77. [Google Scholar] [CrossRef]
- Pedersen, K.W.; Kierulf, B. Direct Isolation of Exosomes from Cell Culture: Simplifying Methods for Exosome Enrichment and Analysis. Transl. Biomed. 2015, 6. [Google Scholar] [CrossRef]
- Kim, H.K.; Song, K.S.; Lee, E.S.; Lee, Y.J.; Park, Y.S.; Lee, K.R.; Lee, S.N. Optimized flow cytometric assay for the measurement of platelet microparticles in plasma: Pre-analytic and analytic considerations. Blood Coagul. Fibrinolysis 2002, 13, 393–397. [Google Scholar] [CrossRef]
- Szatanek, R.; Baj-Krzyworzeka, M.; Zimoch, J.; Lekka, M.; Siedlar, M.; Baran, J. The Methods of Choice for Extracellular Vesicles (EVs) Characterization. Int. J. Mol. Sci. 2017, 18. [Google Scholar] [CrossRef]
- Mehdiani, A.; Maier, A.; Pinto, A.; Barth, M.; Akhyari, P.; Lichtenberg, A. An innovative method for exosome quantification and size measurement. J. Vis. Exp. 2015, 50974. [Google Scholar] [CrossRef]
- Tirinato, L.; Gentile, F.; Di Mascolo, D.; Coluccio, M.L.; Das, G.; Liberale, C.; Pullano, S.A.; Perozziello, G.; Francardi, M.; Accardo, A.; et al. SERS analysis on exosomes using super-hydrophobic surfaces. Microelectron. Eng. 2012, 97, 337–340. [Google Scholar] [CrossRef]
- Sharma, S.; Gillespie, B.M.; Palanisamy, V.; Gimzewski, J.K. Quantitative nanostructural and single-molecule force spectroscopy biomolecular analysis of human-saliva-derived exosomes. Langmuir 2011, 27, 14394–14400. [Google Scholar] [CrossRef]
- Schachermeyer, S.; Ashby, J.; Zhong, W. Advances in field-flow fractionation for the analysis of biomolecules: Instrument design and hyphenation. Anal. Bioanal. Chem. 2012, 404, 1151–1158. [Google Scholar] [CrossRef] [PubMed]
- Lim, J.; Yeap, S.P.; Che, H.X.; Low, S.C. Characterization of magnetic nanoparticle by dynamic light scattering. Nanoscale Res. Lett. 2013, 8, 381. [Google Scholar] [CrossRef]
- Tang, Y.-T.; Huang, Y.-Y.; Zheng, L.; Qin, S.-H.; Xu, X.-P.; An, T.-X.; Xu, Y.; Wu, Y.-S.; Hu, X.-M.; Ping, B.-H.; et al. Comparison of isolation methods of exosomes and exosomal RNA from cell culture medium and serum. Int. J. Mol. Med. 2017, 40, 834–844. [Google Scholar] [CrossRef] [PubMed]
- Mathivanan, S.; Simpson, R.J. ExoCarta: A compendium of exosomal proteins and RNA. Proteomics 2009, 9, 4997–5000. [Google Scholar] [CrossRef] [PubMed]
- Vestad, B.; Llorente, A.; Neurauter, A.; Phuyal, S.; Kierulf, B.; Kierulf, P.; Skotland, T.; Sandvig, K.; Haug, K.B.F.; Øvstebø, R. Size and concentration analyses of extracellular vesicles by nanoparticle tracking analysis: A variation study. J. Extracell. Vesicles 2017, 6, 1344087. [Google Scholar] [CrossRef] [PubMed]
- Smith, V.L.; Cheng, Y.; Bryant, B.R.; Schorey, J.S. Exosomes function in antigen presentation during an in vivo Mycobacterium tuberculosis infection. Sci. Rep. 2017, 7, 43578. [Google Scholar] [CrossRef] [PubMed]
- Giri, P.K.; Kruh, N.A.; Dobos, K.M.; Schorey, J.S. Proteomic analysis identifies highly antigenic proteins in exosomes from M. tuberculosis-infected and culture filtrate protein-treated macrophages. Proteomics 2010, 10, 3190–3202. [Google Scholar] [CrossRef]
- Wiklander, O.P.B.; Bostancioglu, R.B.; Welsh, J.A.; Zickler, A.M.; Murke, F.; Corso, G.; Felldin, U.; Hagey, D.W.; Evertsson, B.; Liang, X.-M.; et al. Systematic Methodological Evaluation of a Multiplex Bead-Based Flow Cytometry Assay for Detection of Extracellular Vesicle Surface Signatures. Front. Immunol. 2018, 9, 1326. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Wu, C.; Gu, W.; Ji, H.; Zhu, L. Serum Exosomal MicroRNAs Predict Acute Respiratory Distress Syndrome Events in Patients with Severe Community-Acquired Pneumonia. BioMed Res. Int. 2019, 2019, 3612020. [Google Scholar] [CrossRef] [PubMed]
- Alipoor, S.D.; Tabarsi, P.; Varahram, M.; Movassaghi, M.; Dizaji, M.K.; Folkerts, G.; Garssen, J.; Adcock, I.M.; Mortaz, E. Serum Exosomal miRNAs Are Associated with Active Pulmonary Tuberculosis. Dis. Markers 2019, 2019, 1907426. [Google Scholar] [CrossRef] [PubMed]
- Hu, G.; Drescher, K.M.; Chen, X.-M. Exosomal miRNAs: Biological Properties and Therapeutic Potential. Front. Genet. 2012, 3, 56. [Google Scholar] [CrossRef] [PubMed]
- Qiu, H.; Shi, S.; Wang, S.; Peng, H.; Ding, S.-J.; Wang, L. Proteomic Profiling Exosomes from Vascular Smooth Muscle Cell. Proteomics Clin. Appl. 2018, 12, e1700097. [Google Scholar] [CrossRef]
- Higginbotham, J.N.; Zhang, Q.; Jeppesen, D.K.; Scott, A.M.; Manning, H.C.; Ochieng, J.; Franklin, J.L.; Coffey, R.J. Identification and characterization of EGF receptor in individual exosomes by fluorescence-activated vesicle sorting. J. Extracell. Vesicles 2016, 5, 29254. [Google Scholar] [CrossRef]
- Tickner, J.A.; Urquhart, A.J.; Stephenson, S.-A.; Richard, D.J.; O’Byrne, K.J. Functions and therapeutic roles of exosomes in cancer. Front. Oncol. 2014, 4, 127. [Google Scholar] [CrossRef]
- Diaz, G.; Wolfe, L.M.; Kruh-Garcia, N.A.; Dobos, K.M. Changes in the Membrane-Associated Proteins of Exosomes Released from Human Macrophages after Mycobacterium tuberculosis Infection. Sci. Rep. 2016, 6, 37975. [Google Scholar] [CrossRef]
- Giri, P.K.; Schorey, J.S. Exosomes derived from M. Bovis BCG infected macrophages activate antigen-specific CD4+ and CD8+ T cells in vitro and in vivo. PLoS ONE 2008, 3, e2461. [Google Scholar] [CrossRef]
- Shekhawat, S.D.; Purohit, H.J.; Taori, G.M.; Daginawala, H.F.; Kashyap, R.S. Evaluation of heat shock proteins for discriminating between latent tuberculosis infection and active tuberculosis: A preliminary report. J. Infect. Public Health 2016, 9, 143–152. [Google Scholar] [CrossRef]
- Bhatnagar, S.; Schorey, J.S. Exosomes released from infected macrophages contain Mycobacterium avium glycopeptidolipids and are proinflammatory. J. Biol. Chem. 2007, 282, 25779–25789. [Google Scholar] [CrossRef] [PubMed]
- Singh, P.P.; Smith, V.L.; Karakousis, P.C.; Schorey, J.S. Exosomes isolated from mycobacteria-infected mice or cultured macrophages can recruit and activate immune cells in vitro and in vivo. J. Immunol. 2012, 189, 777–785. [Google Scholar] [CrossRef] [PubMed]
- Tyagi, P.; Pal, V.K.; Srinivasan, S.; Singh, A. Mycobacterium tuberculosis reactivates HIV via exosomes mediated resetting of cellular redox potential and bioenergetics. mBio 2019. [Google Scholar] [CrossRef]
- Alipoor, S.D.; Mortaz, E.; Tabarsi, P.; Farnia, P.; Mirsaeidi, M.; Garssen, J.; Movassaghi, M.; Adcock, I.M. Bovis Bacillus Calmette-Guerin (BCG) infection induces exosomal miRNA release by human macrophages. J. Transl. Med. 2017, 15, 105. [Google Scholar] [CrossRef] [PubMed]
- Arya, R.; Dabral, D.; Faruquee, H.M.; Mazumdar, H.; Patgiri, S.J.; Deka, T.; Basumatary, R.; Kupa, R.-U.; Semy, C.; Kapfo, W.; et al. Serum Small Extracellular Vesicles Proteome of Tuberculosis Patients Demonstrated Deregulated Immune Response. Proteomics Clin. Appl. 2020, 14, e1900062. [Google Scholar] [CrossRef]
- Zhang, D.; Yi, Z.; Fu, Y. Downregulation of miR-20b-5p facilitates Mycobacterium tuberculosis survival in RAW 264.7 macrophages via attenuating the cell apoptosis by Mcl-1 upregulation. J. Cell. Biochem. 2019, 120, 5889–5896. [Google Scholar] [CrossRef]
- Athman, J.J.; Sande, O.J.; Groft, S.G.; Reba, S.M.; Nagy, N.; Wearsch, P.A.; Richardson, E.T.; Rojas, R.; Boom, W.H.; Shukla, S.; et al. Mycobacterium tuberculosis Membrane Vesicles Inhibit T Cell Activation. J. Immunol. 2017, 198, 2028–2037. [Google Scholar] [CrossRef]
- Wang, J.; Yao, Y.; Xiong, J.; Wu, J.; Tang, X.; Li, G. Evaluation of the inflammatory response in macrophages stimulated with exosomes secreted by Mycobacterium avium-infected macrophages. BioMed Res. Int. 2015, 2015, 658421. [Google Scholar] [CrossRef]
- Smith, V.L.; Jackson, L.; Schorey, J.S. Ubiquitination as a Mechanism to Transport Soluble Mycobacterial and Eukaryotic Proteins to Exosomes. J. Immunol. 2015, 195, 2722–2730. [Google Scholar] [CrossRef]
- Ramachandra, L.; Qu, Y.; Wang, Y.; Lewis, C.J.; Cobb, B.A.; Takatsu, K.; Boom, W.H.; Dubyak, G.R.; Harding, C.V. Mycobacterium tuberculosis synergizes with ATP to induce release of microvesicles and exosomes containing major histocompatibility complex class II molecules capable of antigen presentation. Infect. Immun. 2010, 78, 5116–5125. [Google Scholar] [CrossRef]
- Målen, H.; Berven, F.S.; Fladmark, K.E.; Wiker, H.G. Comprehensive analysis of exported proteins from Mycobacterium tuberculosis H37Rv. Proteomics 2007, 7, 1702–1718. [Google Scholar] [CrossRef] [PubMed]
- Fänge, R.; Lidman, U. Secretion of sulfuric acid in Cassidaria echinophora Lamarck (Mollusca: Mesogastropoda, marine carnivorous snail). Comp. Biochem. Physiol. A Comp. Physiol. 1976, 53, 101–103. [Google Scholar] [CrossRef]
- Kruh-Garcia, N.A.; Wolfe, L.M.; Chaisson, L.H.; Worodria, W.O.; Nahid, P.; Schorey, J.S.; Davis, J.L.; Dobos, K.M. Detection of Mycobacterium tuberculosis peptides in the exosomes of patients with active and latent M. tuberculosis infection using MRM-MS. PLoS ONE 2014, 9, e103811. [Google Scholar] [CrossRef]
- Bhatnagar, S.; Shinagawa, K.; Castellino, F.J.; Schorey, J.S. Exosomes released from macrophages infected with intracellular pathogens stimulate a proinflammatory response in vitro and in vivo. Blood 2007, 110, 3234–3244. [Google Scholar] [CrossRef] [PubMed]
- Mehaffy, C.; Kruh-Garcia, N.A.; Graham, B.; Jarlsberg, L.G.; Willyerd, C.E.; Borisov, A.; Sterling, T.R.; Nahid, P.; Dobos, K.M. Identification of Mycobacterium tuberculosis Peptides in Serum Extracellular Vesicles from Persons with Latent Tuberculosis Infection. J. Clin. Microbiol. 2020, 58. [Google Scholar] [CrossRef] [PubMed]
- Ziegenbalg, A.; Prados-Rosales, R.; Jenny-Avital, E.R.; Kim, R.S.; Casadevall, A.; Achkar, J.M. Immunogenicity of mycobacterial vesicles in humans: Identification of a new tuberculosis antibody biomarker. Tuberculosis 2013, 93, 448–455. [Google Scholar] [CrossRef]
- Rosenkrands, I.; Weldingh, K.; Jacobsen, S.; Hansen, C.V.; Florio, W.; Gianetri, I.; Andersen, P. Mapping and identification of Mycobacterium tuberculosis proteins by two-dimensional gel electrophoresis, microsequencing and immunodetection. Electrophoresis 2000, 21, 935–948. [Google Scholar] [CrossRef]
- Hu, X.; Liao, S.; Bai, H.; Wu, L.; Wang, M.; Wu, Q.; Zhou, J.; Jiao, L.; Chen, X.; Zhou, Y.; et al. Integrating exosomal microRNAs and electronic health data improved tuberculosis diagnosis. EBioMedicine 2019, 40, 564–573. [Google Scholar] [CrossRef]
- Forrellad, M.A.; Klepp, L.I.; Gioffré, A.; Sabio y García, J.; Morbidoni, H.R.; de La Paz Santangelo, M.; Cataldi, A.A.; Bigi, F. Virulence factors of the Mycobacterium tuberculosis complex. Virulence 2013, 4, 3–66. [Google Scholar] [CrossRef]
- Pandey, R.; Rodriguez, G.M. A ferritin mutant of Mycobacterium tuberculosis is highly susceptible to killing by antibiotics and is unable to establish a chronic infection in mice. Infect. Immun. 2012, 80, 3650–3659. [Google Scholar] [CrossRef]
- Fortune, S.M.; Solache, A.; Jaeger, A.; Hill, P.J.; Belisle, J.T.; Bloom, B.R.; Rubin, E.J.; Ernst, J.D. Mycobacterium tuberculosis inhibits macrophage responses to IFN-gamma through myeloid differentiation factor 88-dependent and -independent mechanisms. J. Immunol. 2004, 172, 6272–6280. [Google Scholar] [CrossRef] [PubMed]
- Bekmurzayeva, A.; Sypabekova, M.; Kanayeva, D. Tuberculosis diagnosis using immunodominant, secreted antigens of Mycobacterium tuberculosis. Tuberculosis 2013, 93, 381–388. [Google Scholar] [CrossRef] [PubMed]
- Karbalaei Zadeh Babaki, M.; Soleimanpour, S.; Rezaee, S.A. Antigen 85 complex as a powerful Mycobacterium tuberculosis immunogene: Biology, immune-pathogenicity, applications in diagnosis, and vaccine design. Microb. Pathog. 2017, 112, 20–29. [Google Scholar] [CrossRef] [PubMed]
- Singhal, N.; Sharma, P.; Kumar, M.; Joshi, B.; Bisht, D. Analysis of intracellular expressed proteins of Mycobacterium tuberculosis clinical isolates. Proteome Sci. 2012, 10, 14. [Google Scholar] [CrossRef] [PubMed]
- Castro-Garza, J.; García-Jacobo, P.; Rivera-Morales, L.G.; Quinn, F.D.; Barber, J.; Karls, R.; Haas, D.; Helms, S.; Gupta, T.; Blumberg, H.; et al. Detection of anti-HspX antibodies and HspX protein in patient sera for the identification of recent latent infection by Mycobacterium tuberculosis. PLoS ONE 2017, 12, e0181714. [Google Scholar] [CrossRef] [PubMed]
- Samanich, K.M.; Belisle, J.T.; Sonnenberg, M.G.; Keen, M.A.; Zolla-Pazner, S.; Laal, S. Delineation of human antibody responses to culture filtrate antigens of Mycobacterium tuberculosis. J. Infect. Dis. 1998, 178, 1534–1538. [Google Scholar] [CrossRef]
- Wanchu, A.; Dong, Y.; Sethi, S.; Myneedu, V.P.; Nadas, A.; Liu, Z.; Belisle, J.; Laal, S. Biomarkers for clinical and incipient tuberculosis: Performance in a TB-endemic country. PLoS ONE 2008, 3, e2071. [Google Scholar] [CrossRef]
- Cheng, Y.; Schorey, J.S. Exosomes carrying mycobacterial antigens can protect mice against Mycobacterium tuberculosis infection. Eur. J. Immunol. 2013, 43, 3279–3290. [Google Scholar] [CrossRef]
- Lee, J.; Kim, S.-H.; Choi, D.-S.; Lee, J.S.; Kim, D.-K.; Go, G.; Park, S.-M.; Kim, S.H.; Shin, J.H.; Chang, C.L.; et al. Proteomic analysis of extracellular vesicles derived from Mycobacterium tuberculosis. Proteomics 2015, 15, 3331–3337. [Google Scholar] [CrossRef]
- Singh, P.P.; Li, L.; Schorey, J.S. Exosomal RNA from Mycobacterium tuberculosis-Infected Cells Is Functional in Recipient Macrophages. Traffic 2015, 16, 555–571. [Google Scholar] [CrossRef]
- Singh, Y.; Kaul, V.; Mehra, A.; Chatterjee, S.; Tousif, S.; Dwivedi, V.P.; Suar, M.; van Kaer, L.; Bishai, W.R.; Das, G. Mycobacterium tuberculosis controls microRNA-99b (miR-99b) expression in infected murine dendritic cells to modulate host immunity. J. Biol. Chem. 2013, 288, 5056–5061. [Google Scholar] [CrossRef] [PubMed]
- Kumar, R.; Halder, P.; Sahu, S.K.; Kumar, M.; Kumari, M.; Jana, K.; Ghosh, Z.; Sharma, P.; Kundu, M.; Basu, J. Identification of a novel role of ESAT-6-dependent miR-155 induction during infection of macrophages with Mycobacterium tuberculosis. Cell. Microbiol. 2012, 14, 1620–1631. [Google Scholar] [CrossRef] [PubMed]
- Sinha, A.; Yadav, A.K.; Chakraborty, S.; Kabra, S.K.; Lodha, R.; Kumar, M.; Kulshreshtha, A.; Sethi, T.; Pandey, R.; Malik, G.; et al. Exosome-enclosed microRNAs in exhaled breath hold potential for biomarker discovery in patients with pulmonary diseases. J. Allergy Clin. Immunol. 2013, 132, 219–222. [Google Scholar] [CrossRef] [PubMed]
- Gallo, A.; Tandon, M.; Alevizos, I.; Illei, G.G. The majority of microRNAs detectable in serum and saliva is concentrated in exosomes. PLoS ONE 2012, 7, e30679. [Google Scholar] [CrossRef]
- Zhang, H.; Sun, Z.; Wei, W.; Liu, Z.; Fleming, J.; Zhang, S.; Lin, N.; Wang, M.; Chen, M.; Xu, Y.; et al. Identification of serum microRNA biomarkers for tuberculosis using RNA-seq. PLoS ONE 2014, 9, e88909. [Google Scholar] [CrossRef]
- Zhang, X.; Guo, J.; Fan, S.; Li, Y.; Wei, L.; Yang, X.; Jiang, T.; Chen, Z.; Wang, C.; Liu, J.; et al. Screening and identification of six serum microRNAs as novel potential combination biomarkers for pulmonary tuberculosis diagnosis. PLoS ONE 2013, 8, e81076. [Google Scholar] [CrossRef]
- Miotto, P.; Mwangoka, G.; Valente, I.C.; Norbis, L.; Sotgiu, G.; Bosu, R.; Ambrosi, A.; Codecasa, L.R.; Goletti, D.; Matteelli, A.; et al. miRNA signatures in sera of patients with active pulmonary tuberculosis. PLoS ONE 2013, 8, e80149. [Google Scholar] [CrossRef]
- Liu, Y.; Wang, X.; Jiang, J.; Cao, Z.; Yang, B.; Cheng, X. Modulation of T cell cytokine production by miR-144* with elevated expression in patients with pulmonary tuberculosis. Mol. Immunol. 2011, 48, 1084–1090. [Google Scholar] [CrossRef]
- Qi, Y.; Cui, L.; Ge, Y.; Shi, Z.; Zhao, K.; Guo, X.; Yang, D.; Yu, H.; Cui, L.; Shan, Y.; et al. Altered serum microRNAs as biomarkers for the early diagnosis of pulmonary tuberculosis infection. BMC Infect. Dis. 2012, 12, 384. [Google Scholar] [CrossRef]
- Zhou, M.; Yu, G.; Yang, X.; Zhu, C.; Zhang, Z.; Zhan, X. Circulating microRNAs as biomarkers for the early diagnosis of childhood tuberculosis infection. Mol. Med. Rep. 2016, 13, 4620–4626. [Google Scholar] [CrossRef]
- Latorre, I.; Leidinger, P.; Backes, C.; Domínguez, J.; de Souza-Galvão, M.L.; Maldonado, J.; Prat, C.; Ruiz-Manzano, J.; Sánchez, F.; Casas, I.; et al. A novel whole-blood miRNA signature for a rapid diagnosis of pulmonary tuberculosis. Eur. Respir. J. 2015, 45, 1173–1176. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.-Y.; Chen, Y.-M.; Lin, C.-F.; Lo, C.-M.; Liu, H.-J.; Liao, T.-L. MicroRNA-889 Inhibits Autophagy To Maintain Mycobacterial Survival in Patients with Latent Tuberculosis Infection by Targeting TWEAK. mBio 2020, 11. [Google Scholar] [CrossRef] [PubMed]
- Ndzi, E.N.; Indu Viswanath, A.N.; Adzemye, N.G.; Tamgue, O.; Nsongka, M.V.; Nair, A.S.; Nkenfou, C.N. Upregulated bovine tuberculosis microRNAs Trigger oncogenic pathways: An In silico perception. Int. J. Mycobacteriol. 2019, 8, 70–74. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Xu, Y.-M.; Zou, Y.-Q.; Lin, J.; Huang, B.; Liu, J.; Li, J.; Zhang, J.; Yang, W.-M.; Min, Q.-H.; et al. Identification of differential expressed PE exosomal miRNA in lung adenocarcinoma, tuberculosis, and other benign lesions. Medicine 2017, 96, e8361. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Q.; Chen, H.; Yang, Y.; Fu, Y.; Yi, Z. miR-18a promotes Mycobacterial survival in macrophages via inhibiting autophagy by down-regulation of ATM. J. Cell. Mol. Med. 2020, 24, 2004–2012. [Google Scholar] [CrossRef]
- Mortaz, E.; Alipoor, S.D.; Folkerts, G.; Garssen, J.; Adcock, I. Exosomal miRNAs and Association with Pulmonary Tuberculosis. In Respiratory Infections; ERS International Congress 2019 Abstracts; European Respiratory Society: Lausanne, Switzerland, 2019. [Google Scholar] [CrossRef]
- Shepelkova, G.; Evstifeev, V.; Ergesheva, A.; Guskov, A.; Bagirov, M.; Yeremeev, V. Lung microRNAs as Biomarkers of Post Primary TB Variants. In Tuberculosis; ERS International Congress 2019 Abstracts; European Respiratory Society: Lausanne, Switzerland, 2019. [Google Scholar] [CrossRef]
- Pan, L.; Liu, F.; Zhang, J.; Li, J.; Jia, H.; Huang, M.; Liu, X.; Chen, W.; Ding, Z.; Wang, Y.; et al. Genome-Wide miRNA Analysis Identifies Potential Biomarkers in Distinguishing Tuberculous and Viral Meningitis. Front. Cell. Infect. Microbiol. 2019, 9, 323. [Google Scholar] [CrossRef]
- Wang, C.; Yang, S.; Sun, G.; Tang, X.; Lu, S.; Neyrolles, O.; Gao, Q. Comparative miRNA expression profiles in individuals with latent and active tuberculosis. PLoS ONE 2011, 6, e25832. [Google Scholar] [CrossRef]
- Fu, Y.; Yi, Z.; Li, J.; Li, R. Deregulated microRNAs in CD4+ T cells from individuals with latent tuberculosis versus active tuberculosis. J. Cell. Mol. Med. 2014, 18, 503–513. [Google Scholar] [CrossRef]
- Spinelli, S.V.; Diaz, A.; D’Attilio, L.; Marchesini, M.M.; Bogue, C.; Bay, M.L.; Bottasso, O.A. Altered microRNA expression levels in mononuclear cells of patients with pulmonary and pleural tuberculosis and their relation with components of the immune response. Mol. Immunol. 2013, 53, 265–269. [Google Scholar] [CrossRef]
- Lin, J.; Wang, Y.; Zou, Y.-Q.; Chen, X.; Huang, B.; Liu, J.; Xu, Y.-M.; Li, J.; Zhang, J.; Yang, W.-M.; et al. Differential miRNA expression in pleural effusions derived from extracellular vesicles of patients with lung cancer, pulmonary tuberculosis, or pneumonia. Tumour Biol. 2016. [Google Scholar] [CrossRef]
- Cho, S.M.; Shin, S.; Kim, Y.; Song, W.; Hong, S.G.; Jeong, S.H.; Kang, M.S.; Lee, K.A. A novel approach for tuberculosis diagnosis using exosomal DNA and droplet digital PCR. Clin. Microbiol. Infect. 2020, 26, 942.e1–942.e5. [Google Scholar] [CrossRef] [PubMed]
- El Andaloussi, S.; Mäger, I.; Breakefield, X.O.; Wood, M.J.A. Extracellular vesicles: Biology and emerging therapeutic opportunities. Nat. Rev. Drug Discov. 2013, 12, 347–357. [Google Scholar] [CrossRef] [PubMed]
- Kruh-Garcia, N.A.; Schorey, J.S.; Dobos, K.M. Exosomes: New Tuberculosis Biomarkers – Prospects from the Bench to the Clinic. In Understanding Tuberculosis—Global Experiences and Innovative Approaches to the Diagnosis; Cardona, P.-J., Ed.; InTech: Cham, Switzerland, 2012; ISBN 978-953-307-938-7. [Google Scholar] [CrossRef]
- Cheng, L.; Sharples, R.A.; Scicluna, B.J.; Hill, A.F. Exosomes provide a protective and enriched source of miRNA for biomarker profiling compared to intracellular and cell-free blood. J. Extracell. Vesicles 2014, 3. [Google Scholar] [CrossRef] [PubMed]
- Prada, I.; Meldolesi, J. Binding and Fusion of Extracellular Vesicles to the Plasma Membrane of Their Cell Targets. Int. J. Mol. Sci. 2016, 17, 1296. [Google Scholar] [CrossRef] [PubMed]
- Song, Z.; Cai, Z.; Yan, J.; Shao, Y.W.; Zhang, Y. Liquid biopsies using pleural effusion-derived exosomal DNA in advanced lung adenocarcinoma. Transl. Lung Cancer Res. 2019, 8, 392–400. [Google Scholar] [CrossRef]
- Sandfeld-Paulsen, B.; Aggerholm-Pedersen, N.; Bæk, R.; Jakobsen, K.R.; Meldgaard, P.; Folkersen, B.H.; Rasmussen, T.R.; Varming, K.; Jørgensen, M.M.; Sorensen, B.S. Exosomal proteins as prognostic biomarkers in non-small cell lung cancer. Mol. Oncol. 2016, 10, 1595–1602. [Google Scholar] [CrossRef]
- Wu, Y.; Wei, J.; Zhang, W.; Xie, M.; Wang, X.; Xu, J. Serum Exosomal miR-1290 is a Potential Biomarker for Lung Adenocarcinoma. Onco. Targets. Ther. 2020, 13, 7809–7818. [Google Scholar] [CrossRef]
- Chen, L.; Cao, P.; Huang, C.; Wu, Q.; Chen, S.; Chen, F. Serum exosomal miR-7977 as a novel biomarker for lung adenocarcinoma. J. Cell. Biochem. 2020, 121, 3382–3391. [Google Scholar] [CrossRef]
- Ueda, K.; Ishikawa, N.; Tatsuguchi, A.; Saichi, N.; Fujii, R.; Nakagawa, H. Antibody-coupled monolithic silica microtips for highthroughput molecular profiling of circulating exosomes. Sci. Rep. 2014, 4, 6232. [Google Scholar] [CrossRef]
- Jakobsen, K.R.; Paulsen, B.S.; Bæk, R.; Varming, K.; Sorensen, B.S.; Jørgensen, M.M. Exosomal proteins as potential diagnostic markers in advanced non-small cell lung carcinoma. J. Extracell. Vesicles 2015, 4, 26659. [Google Scholar] [CrossRef]
- Fujita, Y.; Kuwano, K.; Ochiya, T. Development of small RNA delivery systems for lung cancer therapy. Int. J. Mol. Sci. 2015, 16, 5254–5270. [Google Scholar] [CrossRef] [PubMed]
- Fujita, Y.; Takeshita, F.; Kuwano, K.; Ochiya, T. RNAi Therapeutic Platforms for Lung Diseases. Pharmaceuticals 2013, 6, 223–250. [Google Scholar] [CrossRef] [PubMed]
- Fujita, Y.; Kuwano, K.; Ochiya, T.; Takeshita, F. The impact of extracellular vesicle-encapsulated circulating microRNAs in lung cancer research. BioMed Res. Int. 2014, 2014, 486413. [Google Scholar] [CrossRef] [PubMed]
- Lacedonia, D.; Carpagnano, G.E.; Trotta, T.; Palladino, G.P.; Panaro, M.A.; Zoppo, L.D.; Foschino Barbaro, M.P.; Porro, C. Microparticles in sputum of COPD patients: A potential biomarker of the disease? Int. J. Chron. Obstruct. Pulmon. Dis. 2016, 11, 527–533. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, T.; Kobayashi, S.; Fujino, N.; Suzuki, T.; Ota, C.; He, M.; Yamada, M.; Suzuki, S.; Yanai, M.; Kurosawa, S.; et al. Increased circulating endothelial microparticles in COPD patients: A potential biomarker for COPD exacerbation susceptibility. Thorax 2012, 67, 1067–1074. [Google Scholar] [CrossRef]
- Guiot, J.; Struman, I.; Louis, E.; Louis, R.; Malaise, M.; Njock, M.-S. Exosomal miRNAs in Lung Diseases: From Biologic Function to Therapeutic Targets. J. Clin. Med. 2019, 8, 1345. [Google Scholar] [CrossRef]
- Alipoor, S.D.; Mortaz, E.; Garssen, J.; Movassaghi, M.; Mirsaeidi, M.; Adcock, I.M. Exosomes and Exosomal miRNA in Respiratory Diseases. Mediators Inflamm. 2016, 2016, 5628404. [Google Scholar] [CrossRef]
- Minnis, P.; Kane, R.; Anglin, R.; Walsh, S.; Worrel, J.; Khan, F.; Lumsden, R.V.; Whitty, S.; Keane, M.P. Serum Exosomes from IPF Patients Display a Fibrotic miRNA Profile that Correlates to Clinical Measures of Disease Severity. In 1.5 Diffuse Parenchymal Lung Disease; Annual Congress 2015; European Respiratory Society: Lausanne, Switzerland, 2015. [Google Scholar] [CrossRef]
- Schorey, J.S.; Bhatnagar, S. Exosome function: From tumor immunology to pathogen biology. Traffic 2008, 9, 871–881. [Google Scholar] [CrossRef]

| Year | Isolation Technique | Material | Markers | References |
|---|---|---|---|---|
| 2015 | Sucrose gradient density centrifugation | Cell culture | CD9, CD63, Alix, LAMP-1and GAPDH | [47] |
| 2010 | UC | Cell culture | LAMP-1 | [63] |
| 2016 | ExoQuick-TC | Cell culture | CD63, CD81, RAb5B, HSP70 | [71] |
| 2007 | UC, Sucrose gradient density centrifugation | Cell culture | HSP70, MHC-I, II, CD86 | [74] |
| 2019 | ExoQuick-TC Ultrafiltration | Cell culture Serum | CD63, Alix, RAb5b, LAMP-2 | [76] |
| 2017 | TEIR | Serum | CD81 | [77] |
| 2020 | ExoQuick-TC | Serum | CD63, CD9 | [78] |
| 2019 | UC | Cell culture | CD63 | [79] |
| 2017 | UC | Cell culture | CD9, CD63 | [80] |
| 2015 | ExoQuick_TC | Cell culture | LAMP-1, CD40, CD80, CD81, CD86, CD195 | [81] |
| 2015 | UC | Cell culture | TSG101, LAMP-1 | [82] |
| 2010 | UC | Cell culture | MHC-II, CD86, Rab 7, ICAM-I | [83] |
| Size (nm) | Method | Exosomal Molecules | Sources | References |
|---|---|---|---|---|
| NI | Ultrafiltration | LpqN, LprA, LprF, LprG, LpqH, MPB83, FEIII, LpqL, LppX, LppZ, PBP-1, PSTS3, phoS | Culture | [39] |
| NI | Ultrafiltration and UC | pstS1, glnA1, garA, Dank, fba, icd2, ctpD, sahH | Serum | [41] |
| 85–150 nm | Sucrose density gradient | lipoproteins (LpqH and LprG) and lipoglycans (LAM and LM) | culture | [47] |
| 30–50 nm | TEIR for urine | CFP10, lipoarabinomannan | Urine | [50] |
| 30–50 nm | Filtration and UC | KatG, GlnA, SodA, GroES, CFP10, Ag85,19kDaLpqH | culture | [63] |
| 30–50 nm | ExoQuick-TC | 60 kDa heat shock protein, mitochondrial, Moesin, 60 S acidic ribosomal protein P0, 78 kDa glucose-regulated protein, ATP-dependent RNA helicase A, Lamin-B1, Heat shock cognate 71 kDa protein | culture | [71] |
| NI | UC | Ag85 complex | Culture | [72] |
| NI | Centrifugation | HSP16 | Serum | [73] |
| 50–150 nm | UC | LAM and 19-kDa lipoprotein | culture | [74] |
| 30–50 nm | ExoQuick-TC, UC | KYAT3, SERPINA1, HP, and APOC3 | Serum | [78] |
| NI | UC | LAM, LM | Cell culture | [80] |
| 30–100 nm | ExoQuick | Ag85C, MPT51, SodA, MPT63, ESAT-6 | Culture | [81] |
| NI | Sucrose gradient | KatG, HspX, and GroES | culture | [82] |
| NI | ExoQuick | AcpM, Ag85a,b,c, Apa, BfrB, Cfp1, DnaK, Fba, GlcB, GroES, HspX, Icd2, KatG, Mpt64, SahH, PpiA | Serum | [86] |
| NI | ExoQuick | Ag85a,b,c, GlnA, Mpt32, Mpt 64, BfrB, HSPX, Cfp2, GLCB, Cfp10, Dank, GroES | Serum | [88] |
| NI | Gradient filtration and centrifugation | Ag85 A, B, C, MPT64 and LprG | Serum | [89] |
| 50–150 nm | UC | 19-kDa Lipoprotein | culture | [101] |
| 20–200 nm | Methanol precipitation | SodB, EsxN, LppX, PstS1, LpqH, KatG, GlnA1, Apa, FbpA, FadA3, Mtc28, AcpM, Fba, and Prs | Culture | [102] |
| Exosomal miRNAs/DNA | Sources | References |
|---|---|---|
| let-7e-5p, let-7d-5p, miR-450a-5p, miR-140-5p miR-1246, miR-2110, miR-370-3P, miR-28-3p miR-193b-5p | Serum | [37] |
| miR-1224, miR-1293, miR-425, miR-4467, miR-4732, miR-484, miR-5094, miR-6848,miR-6849, miR-4488 miR-96 | Culture | [77] |
| miR-20b-5p | Culture | [79] |
| miR-191, miR-20b, miR-26a, miR-106a, let-7c, miR-20a, miR-486, miR-3128, miR-1468, miR-3201, miR-8084 | Plasma | [91] |
| mir-149-3p, mir181c-5p, mir1839-3p, mir151-3p, mir241-3p, mir292-3p, mir3107-5p, mir344i, mir486-5p, mir486-3p, mir434-5p, mir714, mir877-3p, mir759, mir713 | Culture | [103] |
| let-7e, let-7f, miR-10a, miR-21, miR-26a, miR-99a, miR-140-3p, miR-150, miR-181a, miR-320, miR-339-5p, miR-425, and miR-582-5p | Serum | [111] |
| miR-210, miR-432, miR-423-5p, miR-134, miR-144, miR-335, miR-26a, miR-361-5p, miR-889 and miR-576-3p | Serum | [112] |
| miR-148a-3p, miR-451a, and miR-150-5p | Pleural effusion | [117] |
| miR-18a | Culture | [118] |
| miR-484, miR-425, and miR-96 | Serum | [119] |
| miR-155, miR-150 and miR-193 | Biopsies | [120] |
| miR-126-3p, miR-130a-3p, miR-151a-3p, miR-199a-5p, miR-642a-3p, and miR-4299 | Serum | [121] |
| miR-21, miR-223, miR-302, miR-424, miR-451 and miR-486-5p, miR-144, miR-365, miR-133a, miR-424 | Blood | [122] |
| miR-340-5p miR-451a miR-32-5p miR-27a-3p miR-29a miR-29b | Blood | [123] |
| miR-424 | Blood | [124] |
| miR-3615, miR-4516, and miR-378i | Biopsy | [125] |
| ExoDNA | Sputum | [126] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Biadglegne, F.; König, B.; Rodloff, A.C.; Dorhoi, A.; Sack, U. Composition and Clinical Significance of Exosomes in Tuberculosis: A Systematic Literature Review. J. Clin. Med. 2021, 10, 145. https://doi.org/10.3390/jcm10010145
Biadglegne F, König B, Rodloff AC, Dorhoi A, Sack U. Composition and Clinical Significance of Exosomes in Tuberculosis: A Systematic Literature Review. Journal of Clinical Medicine. 2021; 10(1):145. https://doi.org/10.3390/jcm10010145
Chicago/Turabian StyleBiadglegne, Fantahun, Brigitte König, Arne C. Rodloff, Anca Dorhoi, and Ulrich Sack. 2021. "Composition and Clinical Significance of Exosomes in Tuberculosis: A Systematic Literature Review" Journal of Clinical Medicine 10, no. 1: 145. https://doi.org/10.3390/jcm10010145
APA StyleBiadglegne, F., König, B., Rodloff, A. C., Dorhoi, A., & Sack, U. (2021). Composition and Clinical Significance of Exosomes in Tuberculosis: A Systematic Literature Review. Journal of Clinical Medicine, 10(1), 145. https://doi.org/10.3390/jcm10010145






