A Versatile Suspended Lipid Membrane System for Probing Membrane Remodeling and Disruption
Abstract
:1. Introduction
2. Results and Discussion
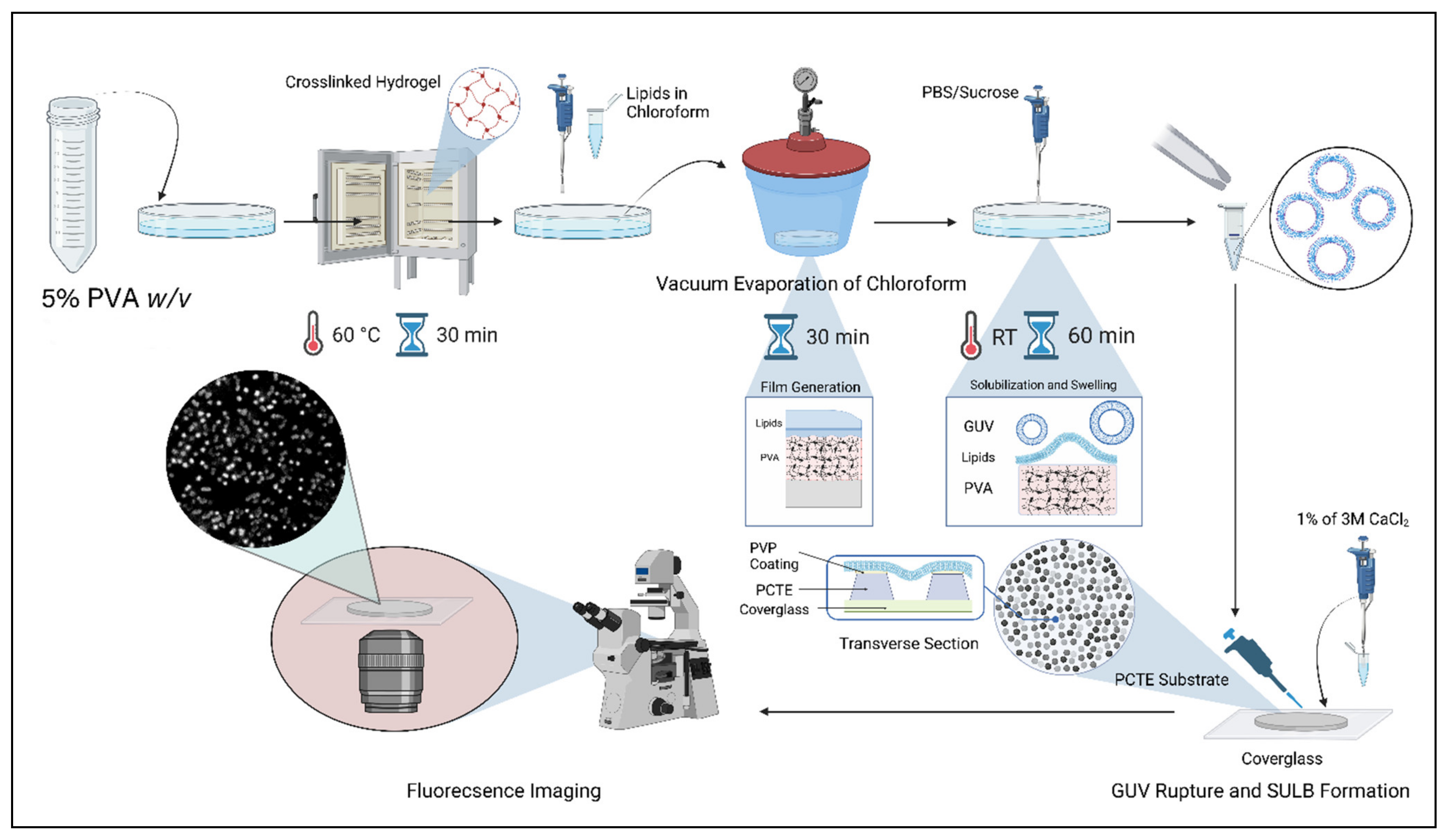
2.1. Suspended Bilayer Membranes on PCTE Substrates
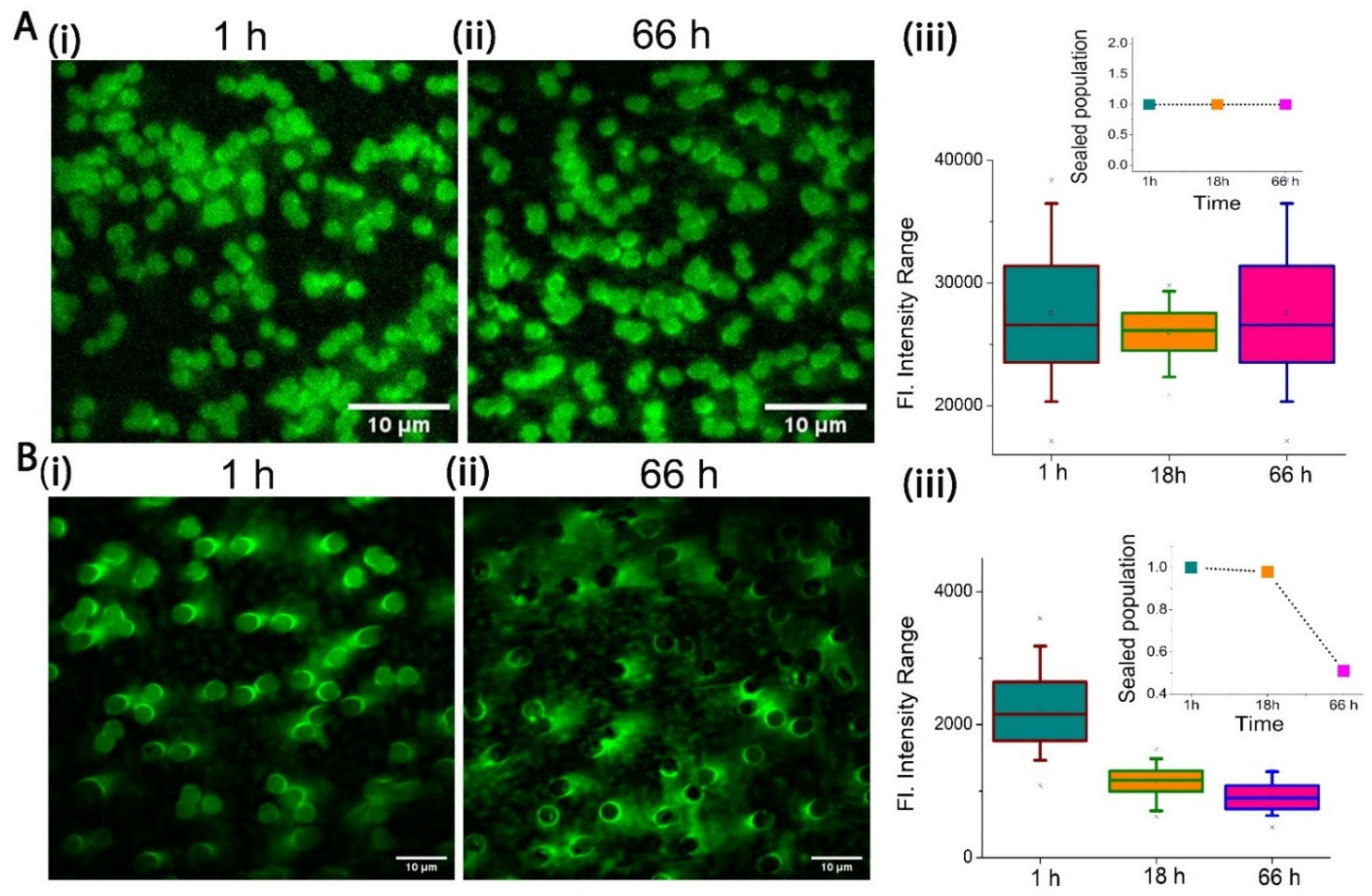
2.2. Stability of PCTE-SULB Membranes
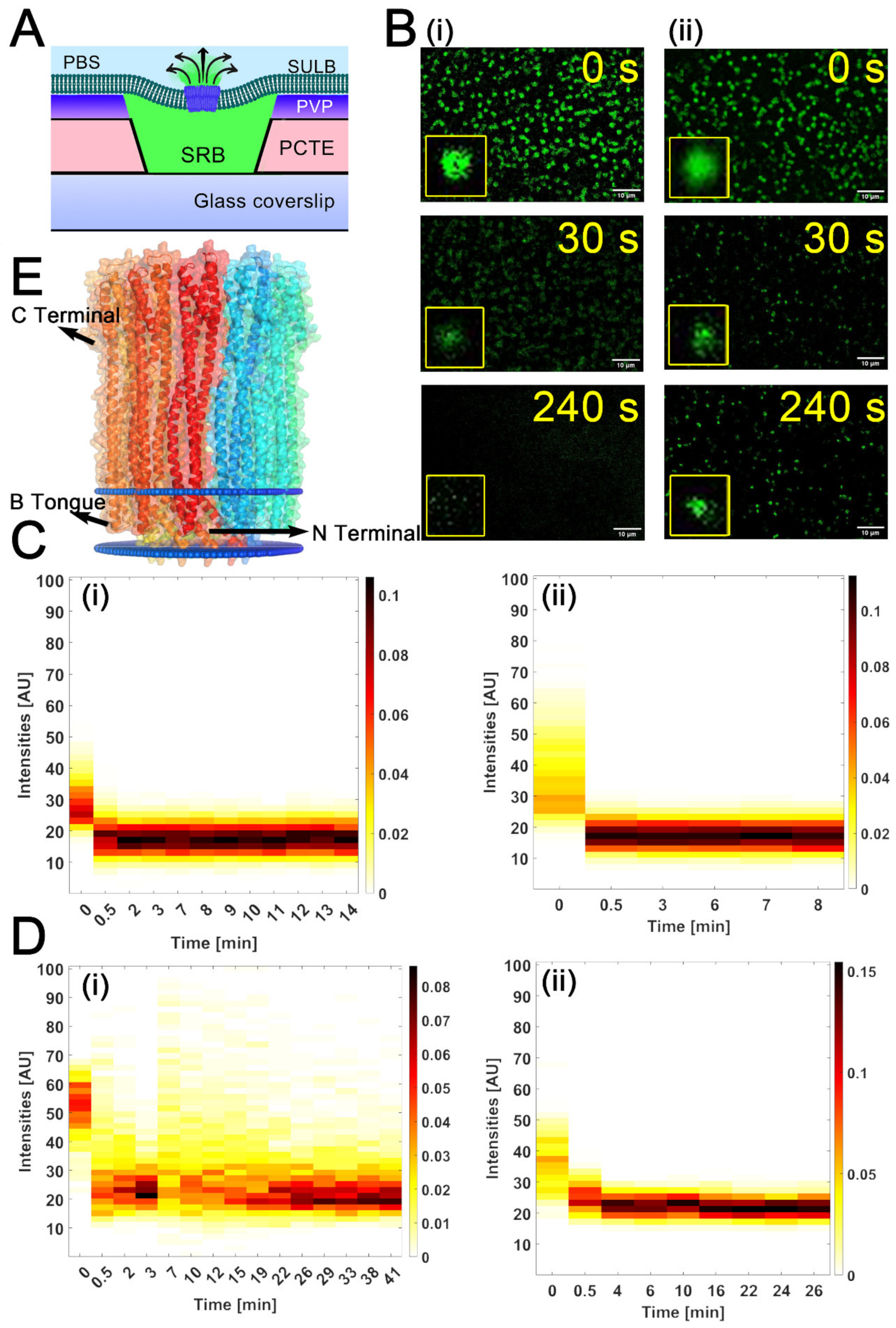
2.3. Cytolysin A Pore Formation in Ternary SULB Membranes
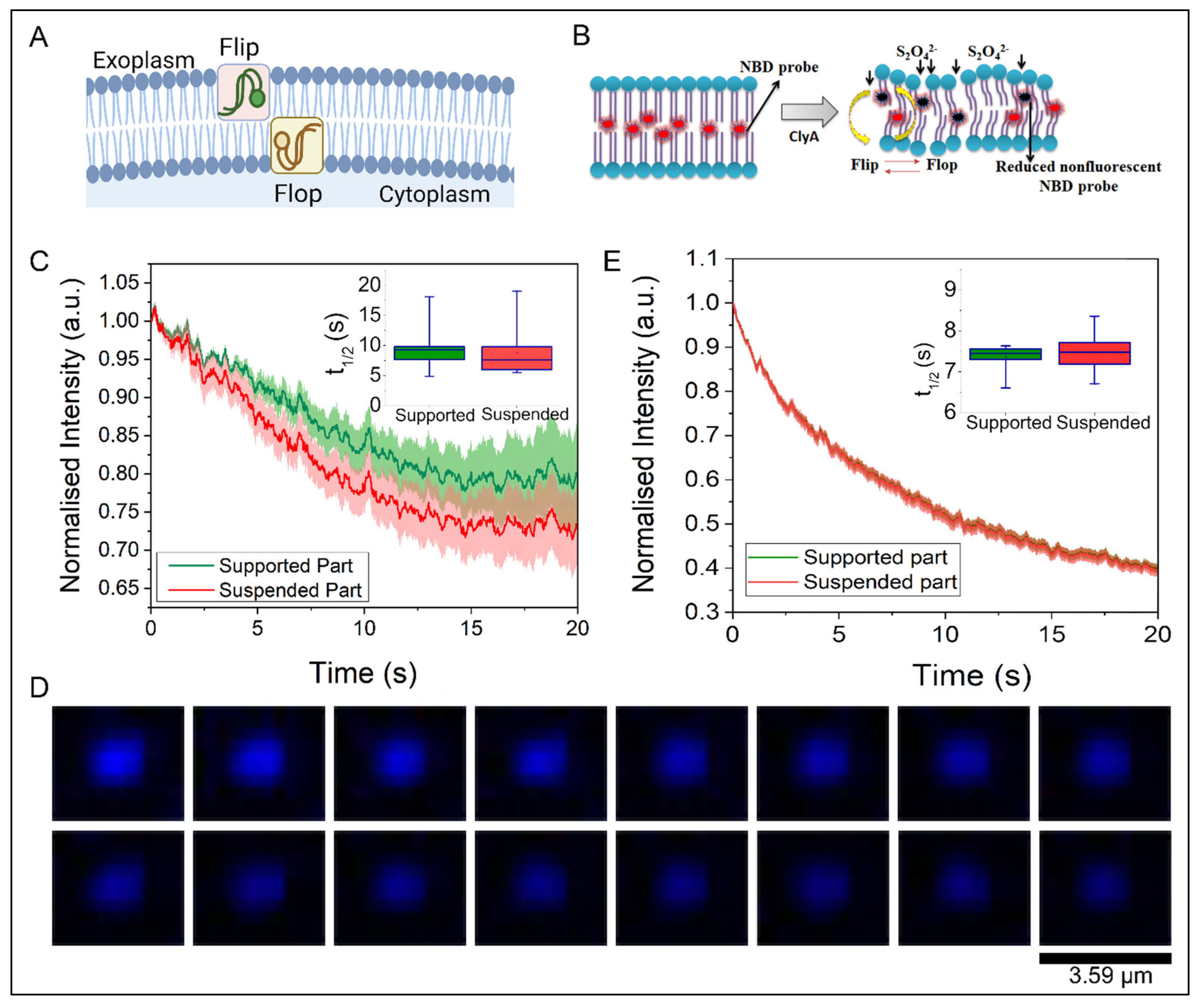
2.4. Trans-Bilayer Lipid Movement in SULB
2.5. Virus Membrane Fusion in PCTE-SULB
3. Conclusions
4. Materials and Method
4.1. Materials
4.2. GUV Formation
4.3. Atomic Force Microscopy
5. ClyA Pore Formation/Leakage Study on Suspended Lipid Bilayer
5.1. Fluorescence Microscope Setup and Analysis
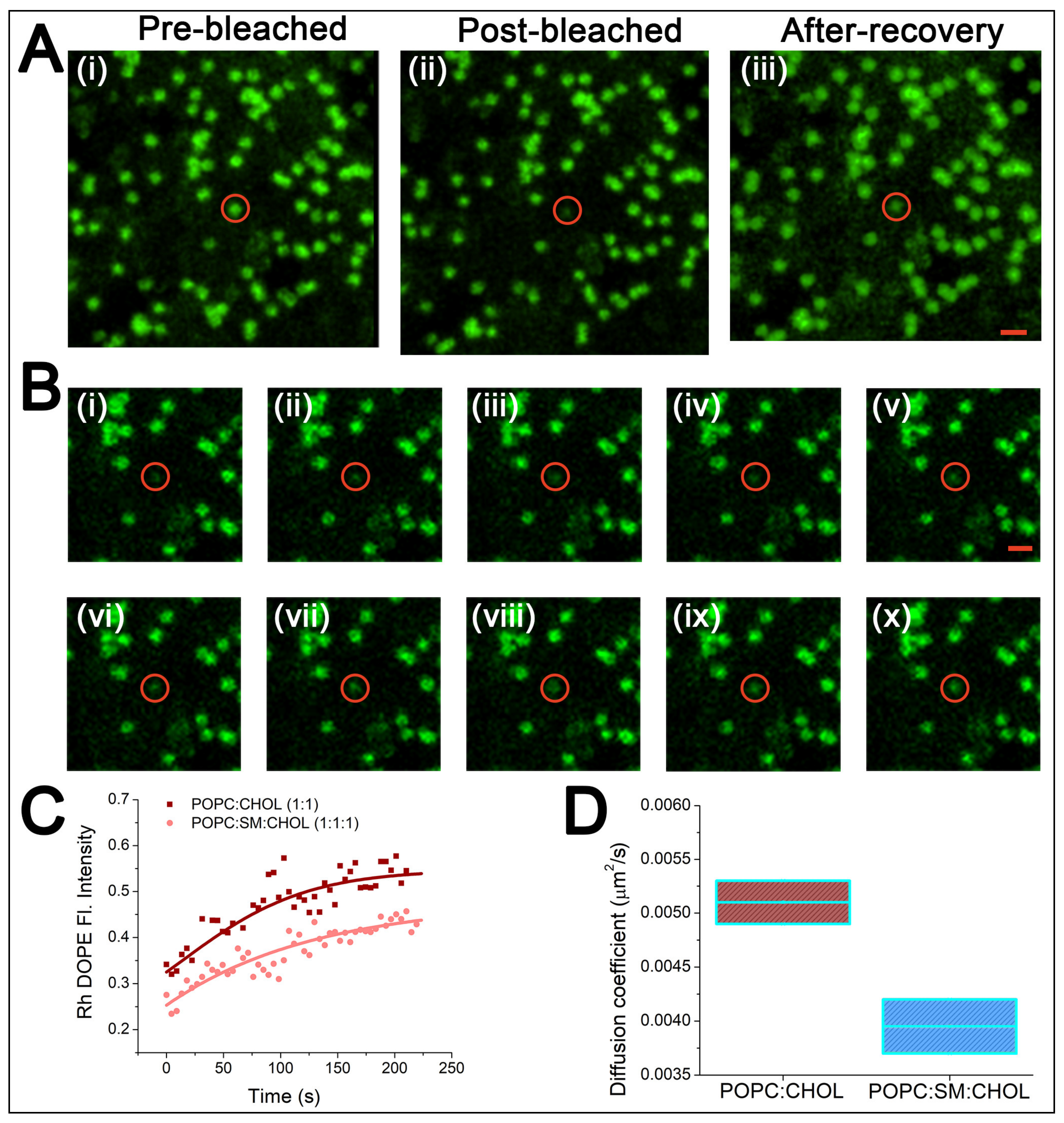
5.2. Fluorescence Recovery after Photobleaching (FRAP) of Lipid Bilayers
6. Measurement of Trans-Bilayer Movement
7. Transfection and Generation of Virus Stock
8. Virus Purification and Generation of DiD-Labeled Virus
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Siontorou, C.G.; Nikoleli, G.-P.; Nikolelis, D.P.; Karapetis, S.K. Artificial Lipid Membranes: Past, Present, and Future. Membranes 2017, 7, 38. [Google Scholar] [CrossRef]
- Ries, R.S.; Choi, H.; Blunck, R.; Bezanilla, F.; Heath, J.R. Black Lipid Membranes: Visualizing the Structure, Dynamics, and Substrate Dependence of Membranes. J. Phys. Chem. B 2004, 108, 16040–16049. [Google Scholar] [CrossRef]
- Khan, M.S.; Dosoky, N.S.; Williams, J.D. Engineering Lipid Bilayer Membranes for Protein Studies. Int. J. Mol. Sci. 2013, 14, 21561–21597. [Google Scholar] [CrossRef] [Green Version]
- Mueller, P.; Rudin, D.O.; Ti Tien, H.; Wescott, W.C. Reconstitution of Cell Membrane Structure in vitro and its Transformation into an Excitable System. Nature 1962, 194, 979–980. [Google Scholar] [CrossRef]
- Danelon, C.; Perez, J.-B.; Santschi, C.; Brugger, J.; Vogel, H. Cell Membranes Suspended Across Nanoaperture Arrays. Langmuir 2006, 22, 22–25. [Google Scholar] [CrossRef]
- Kök, F.N.; Arslan Yildiz, A.; Inci, F. Biomimetic Lipid Membranes: Fundamentals, Applications, and Commercialization; Kök, F.N., Arslan Yildiz, A., Inci, F., Eds.; Springer International Publishing: Cham, Switzerland, 2019; ISBN 978-3-030-11595-1. [Google Scholar]
- De Planque, M.R.R. Lipid bilayer platforms for parallel ion channel recordings. Jpn. J. Appl. Phys. 2022, 61, SC0804. [Google Scholar] [CrossRef]
- El Kirat, K.; Morandat, S.; Dufrêne, Y.F. Nanoscale analysis of supported lipid bilayers using atomic force microscopy. Biochim. Biophys. Acta–Biomembr. 2010, 1798, 750–765. [Google Scholar] [CrossRef] [Green Version]
- Claesson, M.; Cho, N.-J.; Frank, C.W.; Andersson, M. Vesicle Adsorption on Mesoporous Silica and Titania. Langmuir 2010, 26, 16630–16633. [Google Scholar] [CrossRef]
- Ahmed, T.; van den Driesche, S.; Bafna, J.A.; Oellers, M.; Hemmler, R.; Gall, K.; Wagner, R.; Winterhalter, M.; Vellekoop, M.J. Rapid lipid bilayer membrane formation on Parylene coated apertures to perform ion channel analyses. Biomed. Microdevices 2020, 22, 32. [Google Scholar] [CrossRef]
- Coker, H.L.E.; Cheetham, M.R.; Kattnig, D.R.; Wang, Y.J.; Garcia-Manyes, S.; Wallace, M.I. Controlling Anomalous Diffusion in Lipid Membranes. Biophys. J. 2019, 116, 1085–1094. [Google Scholar] [CrossRef] [PubMed]
- Dertinger, T.; von der Hocht, I.; Benda, A.; Hof, M.; Enderlein, J. Surface Sticking and Lateral Diffusion of Lipids in Supported Bilayers. Langmuir 2006, 22, 9339–9344. [Google Scholar] [CrossRef] [Green Version]
- Castellana, E.T.; Cremer, P.S. Solid supported lipid bilayers: From biophysical studies to sensor design. Surf. Sci. Rep. 2006, 61, 429–444. [Google Scholar] [CrossRef]
- Rebaud, S.; Maniti, O.; Girard-Egrot, A.P. Tethered bilayer lipid membranes (tBLMs): Interest and applications for biological membrane investigations. Biochimie 2014, 107, 135–142. [Google Scholar] [CrossRef]
- Hertrich, S.; Stetter, F.; Rühm, A.; Hugel, T.; Nickel, B. Highly Hydrated Deformable Polyethylene Glycol-Tethered Lipid Bilayers. Langmuir 2014, 30, 9442–9447. [Google Scholar] [CrossRef]
- Andersson, J.; Köper, I. Tethered and Polymer Supported Bilayer Lipid Membranes: Structure and Function. Membranes 2016, 6, E30. [Google Scholar] [CrossRef] [Green Version]
- Höfling, F.; Franosch, T. Anomalous transport in the crowded world of biological cells. Rep. Prog. Phys. 2013, 76, 046602. [Google Scholar] [CrossRef] [Green Version]
- Campelo, F.; Arnarez, C.; Marrink, S.J.; Kozlov, M.M. Helfrich model of membrane bending: From Gibbs theory of liquid interfaces to membranes as thick anisotropic elastic layers. Adv. Colloid Interface Sci. 2014, 208, 25–33. [Google Scholar] [CrossRef]
- Sapir, A.; Avinoam, O.; Podbilewicz, B.; Chernomordik, L.V. Viral and Developmental Cell Fusion Mechanisms: Conservation and Divergence. Dev. Cell 2008, 14, 11–21. [Google Scholar] [CrossRef] [Green Version]
- Kusters, I.; van Oijen, A.M.; Driessen, A.J.M. Membrane-on-a-Chip: Microstructured Silicon/Silicon-Dioxide Chips for High-Throughput Screening of Membrane Transport and Viral Membrane Fusion. ACS Nano 2014, 8, 3380–3392. [Google Scholar] [CrossRef]
- Suzuki, H.; Takeuchi, S. Microtechnologies for membrane protein studies. Anal. Bioanal. Chem. 2008, 391, 2695–2702. [Google Scholar] [CrossRef]
- Studer, A.; Han, X.; Winkler, F.K.; Tiefenauer, L.X. Formation of individual protein channels in lipid bilayers suspended in nanopores. Colloids Surf. B Biointerfaces 2009, 73, 325–331. [Google Scholar] [CrossRef]
- Sonnleitner, A.; Schütz, G.J.; Schmidt, T. Free Brownian Motion of Individual Lipid Molecules in Biomembranes. Biophys. J. 1999, 77, 2638–2642. [Google Scholar] [CrossRef] [Green Version]
- Buchholz, K.; Tinazli, A.; Kleefen, A.; Dorfner, D.; Pedone, D.; Rant, U.; Tampé, R.; Abstreiter, G.; Tornow, M. Silicon-on-insulator based nanopore cavity arrays for lipid membrane investigation. Nanotechnology 2008, 19, 445305. [Google Scholar] [CrossRef]
- Stutzmann, M.; Garrido, J.A.; Eickhoff, M.; Brandt, M.S. Direct biofunctionalization of semiconductors: A survey. Phys. Status Solidi A 2006, 203, 3424–3437. [Google Scholar] [CrossRef]
- Ramakrishnan, S.; Gohlke, A.; Li, F.; Coleman, J.; Xu, W.; Rothman, J.E.; Pincet, F. High-Throughput Monitoring of Single Vesicle Fusion Using Freestanding Membranes and Automated Analysis. Langmuir 2018, 34, 5849–5859. [Google Scholar] [CrossRef]
- Schwenen, L.L.G.; Hubrich, R.; Milovanovic, D.; Geil, B.; Yang, J.; Kros, A.; Jahn, R.; Steinem, C. Resolving single membrane fusion events on planar pore-spanning membranes. Sci. Rep. 2015, 5, 12006. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ramakrishnan, S.; Bera, M.; Coleman, J.; Rothman, J.E.; Krishnakumar, S.S. Synergistic roles of Synaptotagmin-1 and complexin in calcium-regulated neuronal exocytosis. eLife 2020, 9, e54506. [Google Scholar] [CrossRef] [PubMed]
- Mühlenbrock, P.; Herwig, K.; Vuong, L.; Mey, I.; Steinem, C. Fusion Pore Formation Observed during SNARE-Mediated Vesicle Fusion with Pore-Spanning Membranes. Biophys. J. 2020, 119, 151–161. [Google Scholar] [CrossRef] [PubMed]
- Kuhlmann, J.W.; Junius, M.; Diederichsen, U.; Steinem, C. SNARE-Mediated Single-Vesicle Fusion Events with Supported and Freestanding Lipid Membranes. Biophys. J. 2017, 112, 2348–2356. [Google Scholar] [CrossRef] [Green Version]
- Hubrich, R.; Park, Y.; Mey, I.; Jahn, R.; Steinem, C. SNARE-Mediated Fusion of Single Chromaffin Granules with Pore-Spanning Membranes. Biophys. J. 2019, 116, 308–318. [Google Scholar] [CrossRef]
- Hennesthal, C.; Drexler, J.; Steinem, C. Membrane-Suspended Nanocompartments Based on Ordered Pores in Alumina. ChemPhysChem 2002, 3, 885–889. [Google Scholar] [CrossRef]
- Sandison, M.E.; Zagnoni, M.; Morgan, H. Air-Exposure Technique for the Formation of Artificial Lipid Bilayers in Microsystems. Langmuir 2007, 23, 8277–8284. [Google Scholar] [CrossRef]
- Steltenkamp, S.; Müller, M.M.; Deserno, M.; Hennesthal, C.; Steinem, C.; Janshoff, A. Mechanical Properties of Pore-Spanning Lipid Bilayers Probed by Atomic Force Microscopy. Biophys. J. 2006, 91, 217–226. [Google Scholar] [CrossRef] [Green Version]
- Tiron, L.G.; Vlad, M.; Baltă, Ş. Research on Hydrophilic Nature of Polyvinylpyrrolidone on Polysulfone Membrane Filtration. IOP Conf. Ser. Mater. Sci. Eng. 2018, 374, 012059. [Google Scholar] [CrossRef]
- Weinberger, A.; Tsai, F.-C.; Koenderink, G.H.; Schmidt, T.F.; Itri, R.; Meier, W.; Schmatko, T.; Schröder, A.; Marques, C. Gel-Assisted Formation of Giant Unilamellar Vesicles. Biophys. J. 2013, 105, 154–164. [Google Scholar] [CrossRef] [Green Version]
- Mey, I.; Stephan, M.; Schmitt, E.K.; Müller, M.M.; Ben Amar, M.; Steinem, C.; Janshoff, A. Local Membrane Mechanics of Pore-Spanning Bilayers. J. Am. Chem. Soc. 2009, 131, 7031–7039. [Google Scholar] [CrossRef]
- Sun, Y.; Zang, X.; Sun, Y.; Wang, L.; Gao, Z. Lipid membranes supported by planar porous substrates. Chem. Phys. Lipids 2020, 228, 104893. [Google Scholar] [CrossRef]
- De Almeida, R.F.M.; Joly, E. Crystallization around solid-like nanosized docks can explain the specificity, diversity, and stability of membrane microdomains. Front. Plant Sci. 2014, 5, 72. [Google Scholar] [CrossRef] [Green Version]
- Sumitomo, K.; Oshima, A. Liquid-Ordered/Liquid-Crystalline Phase Separation at a Lipid Bilayer Suspended over Microwells. Langmuir 2017, 33, 13277–13283. [Google Scholar] [CrossRef]
- Schütte, O.M.; Mey, I.; Enderlein, J.; Savić, F.; Geil, B.; Janshoff, A.; Steinem, C. Size and mobility of lipid domains tuned by geometrical constraints. Proc. Natl. Acad. Sci. USA 2017, 114, E6064–E6071. [Google Scholar] [CrossRef]
- Kawano, R.; Osaki, T.; Sasaki, H.; Takeuchi, S. A Polymer-Based Nanopore-Integrated Microfluidic Device for Generating Stable Bilayer Lipid Membranes. Small 2010, 6, 2100–2104. [Google Scholar] [CrossRef] [PubMed]
- Han, X.; Studer, A.; Sehr, H.; Geissbühler, I.; Di Berardino, M.; Winkler, F.K.; Tiefenauer, L.X. Nanopore Arrays for Stable and Functional Free-Standing Lipid Bilayers. Adv. Mater. 2007, 19, 4466–4470. [Google Scholar] [CrossRef]
- Sathyanarayana, P.; Maurya, S.; Behera, A.; Ravichandran, M.; Visweswariah, S.S.; Ayappa, K.G.; Roy, R. Cholesterol promotes Cytolysin A activity by stabilizing the intermediates during pore formation. Proc. Natl. Acad. Sci. USA 2018, 115, E7323–E7330. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rojko, N.; Cronin, B.; Danial, J.S.H.; Baker, M.A.B.; Anderluh, G.; Wallace, M.I. Imaging the Lipid-Phase-Dependent Pore Formation of Equinatoxin II in Droplet Interface Bilayers. Biophys. J. 2014, 106, 1630–1637. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lomize, M.A.; Pogozheva, I.D.; Joo, H.; Mosberg, H.I.; Lomize, A.L. OPM database and PPM web server: Resources for positioning of proteins in membranes. Nucleic Acids Res. 2012, 40, D370–D376. [Google Scholar] [CrossRef]
- Van Meer, G.; Voelker, D.R.; Feigenson, G.W. Membrane lipids: Where they are and how they behave. Nat. Rev. Mol. Cell Biol. 2008, 9, 112–124. [Google Scholar] [CrossRef]
- McIntyre, J.C.; Sleight, R.G. Fluorescence assay for phospholipid membrane asymmetry. Biochemistry 1991, 30, 11819–11827. [Google Scholar] [CrossRef]
- Matsuzaki, K.; Murase, O.; Fujii, N.; Miyajima, K. An antimicrobial peptide, magainin 2, induced rapid flip-flop of phospholipids coupled with pore formation and peptide translocation. Biochemistry 1996, 35, 11361–11368. [Google Scholar] [CrossRef]
- Harrison, S.C. Viral membrane fusion. Nat. Struct. Mol. Biol. 2008, 15, 690–698. [Google Scholar] [CrossRef]
- Van der Schaar, H.M.; Rust, M.J.; Chen, C.; van der Ende-Metselaar, H.; Wilschut, J.; Zhuang, X.; Smit, J.M. Dissecting the Cell Entry Pathway of Dengue Virus by Single-Particle Tracking in Living Cells. PLoS Pathog. 2008, 4, e1000244. [Google Scholar] [CrossRef]
- Floyd, D.L.; Harrison, S.C.; van Oijen, A.M. Method for Measurement of Viral Fusion Kinetics at the Single Particle Level. J. Vis. Exp. 2009, 31, e1484. [Google Scholar] [CrossRef] [Green Version]
- Zaitseva, E.; Yang, S.-T.; Melikov, K.; Pourmal, S.; Chernomordik, L.V. Dengue Virus Ensures Its Fusion in Late Endosomes Using Compartment-Specific Lipids. PLoS Pathog. 2010, 6, e1001131. [Google Scholar] [CrossRef] [Green Version]
- Zheng, A.; Yuan, F.; Kleinfelter, L.M.; Kielian, M. A toggle switch controls the low pH-triggered rearrangement and maturation of the dengue virus envelope proteins. Nat. Commun. 2014, 5, 3877. [Google Scholar] [CrossRef] [Green Version]
- Zhang, X.; Sheng, J.; Austin, S.K.; Hoornweg, T.E.; Smit, J.M.; Kuhn, R.J.; Diamond, M.S.; Rossmann, M.G. Structure of Acidic pH Dengue Virus Showing the Fusogenic Glycoprotein Trimers. J. Virol. 2014, 89, 743–750. [Google Scholar] [CrossRef] [Green Version]
- Sannigrahi, A.; Chowdhury, S.; Das, B.; Banerjee, A.; Halder, A.; Kumar, A.; Saleem, M.; Naganathan, A.N.; Karmakar, S.; Chattopadhyay, K. The metal cofactor zinc and interacting membranes modulate SOD1 conformation-aggregation landscape in an in vitro ALS model. eLife 2021, 10, e61453. [Google Scholar] [CrossRef]
- Halder, A.; Sannigrahi, A.; De, N.; Chattopadhyay, K.; Karmakar, S. Kinetoplastid Membrane Protein-11 Induces Pores in Anionic Phospholipid Membranes: Effect of Cholesterol. Langmuir 2020, 36, 3522–3530. [Google Scholar] [CrossRef]
- Sannigrahi, A.; Chall, S.; Jawed, J.J.; Kundu, A.; Majumdar, S.; Chattopadhyay, K. Nanoparticle Induced Conformational Switch Between α-Helix and β-Sheet Attenuates Immunogenic Response of MPT63. Langmuir 2018, 34, 8807–8817. [Google Scholar] [CrossRef]
- Lowe, D.G. Object recognition from local scale-invariant features. In Proceedings of the Seventh IEEE International Conference on Computer Vision, Corfu, Greece, 20–25 September 1999; Volume 2, pp. 1150–1157. [Google Scholar]
- Rizon, M.; Yazid, H.; Saad, P.; Shakaff, A.Y.M.; Saad, A.R.; Sugisaka, M.; Yaacob, S.; Mamat, M.R.; Karthigayan, M. Object Detection using Circular Hough Transform. Am. J. Appl. Sci. 2005, 2, 1606–1609. [Google Scholar] [CrossRef]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sannigrahi, A.; Rai, V.H.; Chalil, M.V.; Chakraborty, D.; Meher, S.K.; Roy, R. A Versatile Suspended Lipid Membrane System for Probing Membrane Remodeling and Disruption. Membranes 2022, 12, 1190. https://doi.org/10.3390/membranes12121190
Sannigrahi A, Rai VH, Chalil MV, Chakraborty D, Meher SK, Roy R. A Versatile Suspended Lipid Membrane System for Probing Membrane Remodeling and Disruption. Membranes. 2022; 12(12):1190. https://doi.org/10.3390/membranes12121190
Chicago/Turabian StyleSannigrahi, Achinta, Vishwesh Haricharan Rai, Muhsin Vannan Chalil, Debayani Chakraborty, Subrat Kumar Meher, and Rahul Roy. 2022. "A Versatile Suspended Lipid Membrane System for Probing Membrane Remodeling and Disruption" Membranes 12, no. 12: 1190. https://doi.org/10.3390/membranes12121190
APA StyleSannigrahi, A., Rai, V. H., Chalil, M. V., Chakraborty, D., Meher, S. K., & Roy, R. (2022). A Versatile Suspended Lipid Membrane System for Probing Membrane Remodeling and Disruption. Membranes, 12(12), 1190. https://doi.org/10.3390/membranes12121190





